Polyphenolic extracts from cowpea (Vigna unguiculata) protect colonic myofibroblasts (CCD18Co cells) from lipopolysaccharide (LPS)-induced inflammation – modulation of microRNA 126
Leonnard O.
Ojwang
ab,
Nivedita
Banerjee
bc,
Giuliana D.
Noratto
bc,
Gabriela
Angel-Morales
bc,
Twambo
Hachibamba
d,
Joseph M.
Awika
*ab and
Susanne U.
Mertens-Talcott
*bce
aCereal Quality Laboratory, Soil & Crop Science Department, College Station, Texas 77843, USA. E-mail: jawika@ag.tamu.edu
bDepartment of Nutrition and Food Science, College Station, Texas 77843, USA. E-mail: smtalcott@tamu.edu
cInstitute for Obesity Research and Program Evaluation, Texas A&M AgriLife Research, College Station, Texas 77843, USA
dDepartment of Food Science and Technology, University of Zambia, Lusaka, Zambia
eDepartment of Veterinary Physiology and Pharmacology, Texas A&M University, College Station, Texas 77843, USA
First published on 29th September 2014
Abstract
Cowpea (Vigna unguiculata) is a drought tolerant crop with several agronomic advantages over other legumes. This study evaluated varieties from four major cowpea phenotypes (black, red, light brown and white) containing different phenolic profiles for their anti-inflammatory property on non-malignant colonic myofibroblasts (CCD18Co) cells challenged with an endotoxin (lipopolysaccharide, LPS). Intracellular reactive oxygen species (ROS) assay on the LPS-stimulated cells revealed antioxidative potential of black and red cowpea varieties. Real-time qRT-PCR analysis in LPS-stimulated cells revealed down-regulation of proinflammatory cytokines (IL-8, TNF-α, VCAM-1), transcription factor NF-κB and modulation of microRNA-126 (specific post-transcriptional regulator of VCAM-1) by cowpea polyphenolics. The ability of cowpea polyphenols to modulate miR-126 signaling and its target gene VCAM-1 were studied in LPS-stimulated endothelial cells transfected with a specific inhibitor of miR-126, and treated with 10 mg GAE/L black cowpea extract where the extract in part reversed the effect of the miR-126 inhibitor. This suggests that cowpea may exert their anti-inflammatory activities at least in part through induction of miR-126 that then down-regulate VCAM-1 mRNA and protein expressions. Overall, Cowpea therefore is promising as an anti-inflammatory dietary component.
Introduction
Chronic inflammation is a major risk factor in cardiovascular disease (CVD) and cancers. Chronic inflammation is characterized by the presence of macrophages, lymphocytes and proliferation of vascular tissues. CVDs and cancers are the leading causes of mortality and morbidity in both industrialized and developing countries, especially in low- and middle-income countries.1 It seems that dietary factors significantly influence the incidence of chronic inflammation and associated diseases. Epidemiological evidence shows that consumption of pulses (more than 4 times per week compared with less than once a week) reduced the risk of coronary heart disease (CHD) and cardiovascular diseases (CVD) by 22 and 11%, respectively.2 Strong epidemiological evidence also suggests that regular consumption of pulses can reduce cancer risk. A comparative study of lifestyles in Jiangsu Province, China, found that people living in a low-risk area for gastric cancer consumed kidney beans more frequently than those in a high-risk area.3 Other studies4 have also shown that consumption of grain pulses such as dried beans, split peas, or lentils reduced risk of colorectal adenoma.Among the grain legumes, cowpea (Vigna unguiculata) is one of the most important food legumes of semi-arid tropics covering Asia, Africa, southern Europe, and Central and South America. It provides significant amounts of essential nutrients and energy to these populations.5 Cowpea is a heat and drought tolerant legume. Cowpea's unique traits and low agronomic input requirement are favorable to the poor and low income farmers who are increasingly vulnerable to malnutrition. Malnutrition in developing countries, especially during childhood, has been linked to various chronic diseases in adulthood,6 placing additional burdens on already overtaxed national health budgets and the poor.7 Cowpea may also be a “climate-change crop” that can help alleviate malnutrition among the poor due to its good agronomic properties, versatility and nutritional profile than cereals and tubers, thus should be exploited for its potential in prevention of chronic diseases.
We recently reported that cowpea contains relatively high amounts of flavonols (especially glycoside of quercetin and anthocyanins; as well as procyanidins) and that the seed coat color is a major determinant of flavonoid composition.8,9 Significant anti-inflammatory effects of these flavonoid classes have been demonstrated by some authors.10–12 Quercetin has been shown to potentially control intestinal inflammation in celiac disease by preventing the activation of transcription factor NF-κB and mitogen activated protein kinases (MAPK) pathways;10 consequently counteracting the expression of cytokines and inducible nitric oxide synthase (iNOS).13 Anthocyanins are also reported to exert anti-inflammatory property by inhibiting TNF-α-induced endothelial leukocyte adhesion molecule-1 (ELAM-1) and intercellular adhesion molecule-1 (ICAM-1) expression in cultured HUVEC.11 Cell adhesion molecules play a key role in monocyte recruitment that also plays a role in tumor development. Procyanidins (i.e. condensed tannins), which primarily occur in the form of catechin and epicatechin monomers, their polymers and glycosides,9 have also been shown as effective in inhibiting the AngII-induced MAPK pathways, leading to reduced adhesion molecule expression in HUVEC.12
MicroRNAs (miRs) are small non-coding RNA that post-transcriptionally regulate target genes. They can modulate inflammatory mechanisms associated with transcription factor NF-κB activation by inducing mRNA degradation or blocking translation, as well as regulating vascular inflammation.14 Endogenous miR-126 is known to regulate leukocyte adherence to endothelial cells associated with tumor development.14 Thus, miR-126 is a useful target for investigating cowpea polyphenolics in the regulation of its target gene VCAM-1.
The objective of this study was to investigate the anti-inflammatory properties of cowpea polyphenolics in LPS-stimulated nonmalignant colon (CCD18Co) cells.
Materials and methods
Chemicals and reagents
TaqMan® MicroRNA Assay kit for miR-126 was obtained from Applied Biosystems (Carlsbad, CA). The pairs of forward and reverse primers (VCAM-1, NF-κB, TNF-α and IL-8) were obtained from Integrated DNA Technologies, Inc. (San Diego, CA). Bradford reagent was obtained from Bio-Rad (Hercules, CA) and VCAM-1 (Human sVCAM-1 Immunoassay Kit) was obtained from Invitrogen Corp., Camarillo, CA. The Folin-Ciocalteu reagent, gallic acid, dichlorofluorescein diacetate (DCF-DA), and lipopolysaccharide (LPS) were purchased from Fisher Scientific (Pittsburgh, PA). Dimethyl sulfoxide (DMSO) was obtained from Sigma (St. Louis, MO). All other chemicals were analytical grade from VWR International (Bristol, CT).Plant materials and extraction
Dry cowpea seeds from four cowpea varieties were used in this study, namely, IT95K-1105-5 (black), IT97K-1042-3 (red), 09FCV-CC-27M (light brown) and Early Acre (white), and were collected at maturity in late July 2011 at university experimental station in College Station, Texas (Table 1). These varieties, representing four major cowpea phenotypes, were chosen based on their distinct differences in phenolic profiles (Table 2).8,9 Broken and damaged seeds, as well as foreign materials were removed prior to use.| Variety | Seed coat property | Seed weightb (g/100 seeds) | Amount of cowpea flour (mg)c producing 2 mg L−1 extract concentration | |
|---|---|---|---|---|
| Raw | Boiled | |||
| a Samples were grown in Texas A&M University Test Plots in College Station, TX, in 201. b Seed weight expressed as Mean ± SD of triplicate weights of 100 seeds. c Cowpea flour sample weight expressed as Mean ± SD of triplicate measurements (mg). | ||||
| IT95K-1105-5 | Black, smooth | 23.4 ± 0.37 | 1.08 ± 0.02 | 2.58 ± 0.03 |
| IT97K-1042-3 | Red, smooth | 13.1 ± 0.20 | 2.59 ± 0.04 | 2.75 ± 0.02 |
| 09FCV-CC-27M | Light brown, smooth | 14.9 ± 0.40 | 1.20 ± 0.02 | 1.62 ± 0.03 |
| EARLY ACRE | White, rough, brown-eye | 11.6 ± 0.16 | 4.46 ± 0.04 | 5.28 ± 0.05 |
| Phenotype | Black | Red | Light brown | White |
|---|---|---|---|---|
| Variety | IT95K-1105-5 | IT97K-1042-3 | 09FCV-CC27M | Early Acre |
| a Adapted from Ojwang et al.8 and Ojwang.9 b Data based on UPLC peak areas and MS/MS-MS analysis of all compounds detected in dried ground cowpea flour and expressed as Means ± SDs of triplicates on a dry weight basis. c ND – not detected; d Y – present. e λ max at 354 nm. Peaks that were not structurally identified are not included. | ||||
| Anthocyanins | ||||
| Delphinidin glycosides | 850 ± 10.5 | NDc | ND | ND |
| Cyanidin glycosides | 709 ± 13.4 | ND | ND | ND |
| Petunidin glycosides | 300 ± 4.7 | ND | ND | ND |
| Peonidin glycosides | 41.1 ± 2.1 | ND | ND | ND |
| Malvidin glycosides | 202 ± 4.8 | ND | ND | ND |
| Total anthocyanins | 2094 ± 36 | 0 | 0 | 0 |
| Flavonols | ||||
| Quercetin monoglycosides | 99.9 ± 0.7 | 165 ± 4.8 | 93.3 ± 7.5 | 5.0 ± 0.7 |
| Quercetin diglycosides | 168 ± 5.7 | 405 ± 8.1 | 333 ± 27.9 | 191 ± 19.4 |
| Quercetin triglycosides | 78.3 ± 1.8 | 127 ± 7.0 | 23.7 ± 2.9 | 43.7 ± 4.7 |
| Quercetin acyl glycosides | 121 ± 3.9 | 120 ± 4.1 | 13.6 ± 0.7 | 14.5 ± 1.6 |
| Myricetin monoglycosides | 23.2 ± 1.0 | 128 ± 5.6 | ND | ND |
| Myricetin diglycosides | 23.8 ± 1.4 | 75.4 ± 2.4 | ND | ND |
| Kaempferol diglycosides | 13.6 ± 2.4 | 38.7 ± 1.4 | ND | 15.8 ± 2.3 |
| Total flavonols | 415 ± 14 | 1060 ± 30 | 461 ± 33 | 270 ± 29 |
| Flavan-3-ols | ||||
| Catechin/epicatechin | Yd | Y | Y | ND |
| Catechin-3-O-glucoside | Y | Y | Y | ND |
| Catechin-kaempferol dimere | ND | Y | ND | ND |
| Procyanidin dimer B-type | Y | Y | Y | ND |
| Procyanidin trimer T2 | ND | Y | Y | ND |
| Procyanidin trimer C1 | ND | ND | Y | ND |
| Total flavan-3-ols | ||||
| Free phenolic acids | ||||
| Protocatechuic acid | ND | Y | ND | ND |
| trans-Feruloylaldaric acid | ND | Y | ND | ND |
| trans-Methylferuloylaldaric acid | ND | ND | Y | Y |
Sample treatment
The cowpea samples were separately soaked in water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4.5 w/v) in triplicates for 12 hours, and boiled for 15 minutes. After boiling, the seeds (including the soup) were chilled and frozen to −80 °C, freeze-dried and ground to pass through 60-mesh screen. The powders were then extracted with aqueous 70% acetone, roto-evaporated and the extracts freeze-dried. The freeze-dried extracts were stored at −20 °C until used.
4.5 w/v) in triplicates for 12 hours, and boiled for 15 minutes. After boiling, the seeds (including the soup) were chilled and frozen to −80 °C, freeze-dried and ground to pass through 60-mesh screen. The powders were then extracted with aqueous 70% acetone, roto-evaporated and the extracts freeze-dried. The freeze-dried extracts were stored at −20 °C until used.
Total soluble phenolics
Total soluble phenolic (TSP) levels (measure of total metal ion reducing capacity) of the freeze-dried boiled cowpea extracts was determined by a slightly modified Folin–Ciocalteu assay using gallic acid (GA) as the standard.15Cell culture
Non-malignant colon CCD18Co cell line was purchased from the American Type Culture Collection (Manassas, VA, USA) and cultured according to the manufacturer's recommendations. The cells were maintained at 37 °C in a humidified 5% CO2 atmosphere.Cell proliferation assay
Cells were seeded (1.5 × 104 onto a 24-well plate) and incubated for 24 h (at 37 °C/humidified 5% CO2 atmosphere) to allow cells to stabilize and attach onto the bottom of the wells. The freeze-dried cowpea extracts were re-dissolved in DMSO, and then diluted with media to known concentrations of total soluble polyphenolics ranging from 0–80 mg GAE/L.15,16 The cells were then exposed to the various concentrations of each cowpea phenolic extract for 48 h and then quantified with an electronic cell counter (Z2™ Series, Beckman Coulter, Inc). Overall, the maximum concentration of cowpea extracts that supported CCD18Co cell growth after 48 h of incubation was 20 mg GAE/L (>85% cells surviving). Therefore, extract concentrations within a dose range of 0–20 mg GAE/L were used in the subsequent assays to assess the anti-inflammatory properties of cowpea.Generation of reactive oxygen species (ROS) assay
Intracellular ROS was assessed using 2′,7′-dichlorofluorescin diacetate (DCF-DA) (Molecular Probes, Eugene, OR) as a probe as described by Meng et al.17 but with slight modifications. Cells were seeded in a black 96-well plate (3000 cells per well) for 24 h to allow for cell attachment, followed by incubation with cowpea phenolic extracts (2–20 mg GAE/L) for 24 h. The cells were then stimulated with 2 µg mL−1 LPS (in 100 µL media) to generate ROS for 2 h, followed by washing out the spent media using PBS buffer. The cells were then stained in situ with 100 µL of 10 μM DCFH, incubated at 37 °C and the fluorescence signal was monitored after 15 min at 520 nm emission and 480 nm excitation with a FLUOstar Omega plate reader (BMG Labtech Inc, Durhan, NC). Relative fluorescence units (RFU) were analyzed using Omega Microplate Data Analyse Software and normalized to control cells not treated with LPS and cowpea extracts.LPS-induced inflammation assay
Cells were seeded in a 12-well plate (80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well) for 24 h (37 °C/5% CO2) to allow cell attachment. Cowpea extracts pre-dissolved in DMSO were then diluted to known concentrations of total polyphenolics (2, 5, 10 and 20 mg GAE/L) and normalized to <0.2% DMSO in the culture medium. The cells were then pre-treated with the extracts for 3 h, and then stimulated with 2 µg mL−1 LPS for 6 h, after which messenger RNA (mRNA) and micro-RNA (miRs) were extracted from the lysated cells and analyzed.
000 cells per well) for 24 h (37 °C/5% CO2) to allow cell attachment. Cowpea extracts pre-dissolved in DMSO were then diluted to known concentrations of total polyphenolics (2, 5, 10 and 20 mg GAE/L) and normalized to <0.2% DMSO in the culture medium. The cells were then pre-treated with the extracts for 3 h, and then stimulated with 2 µg mL−1 LPS for 6 h, after which messenger RNA (mRNA) and micro-RNA (miRs) were extracted from the lysated cells and analyzed.
RNA extraction and real-time PCR analysis of mRNAs and miRNAs
| IL-8: | F: 5′-CACCGGAAGGAACCATCTCA-3′ |
| IL-8: | R: 5′-AGAGCCACGGCCAGCTT-3′ |
| TNF-α: | F: 5′-TGTGTGGCTGCAGGAAGAAC-3′ |
| TNF-α: | R: 5′-GCAATTGAAGCACTGGAAAAGG-3′ |
| VCAM-1: | F: 5′-ACAGAAGAAGTGGCCCTCCAT-3′ |
| VCAM-1: | R: 5′-TGGCATCCGTCAGGAAGTG-3′ |
| NF-κB: | F: 5′-TGGGAATGGTGAGGTCACTCT-3′ |
| NF-κB: | R: 5′-TCCTGAACTCCAGCACTCTCTTC-3′ |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 and amplified using TapMan® 2× Universal PCR Master Mix (No AmpErase® UNG) (Applied Biosystems, Foster City, CA) on a 384-well plate following the manufacturer's recommendations. The miR-NU6B small nuclear RNA was used as an endogenous control.
15 and amplified using TapMan® 2× Universal PCR Master Mix (No AmpErase® UNG) (Applied Biosystems, Foster City, CA) on a 384-well plate following the manufacturer's recommendations. The miR-NU6B small nuclear RNA was used as an endogenous control.
Protein expression
Statistical analysis
Data are reported as means ± SD for ROS and protein quantification; and means ± SE for gene expression, of 3 replicates; and analyzed using 2005 SAS (Version 9.1, SAS Inst. Inc., Cary, N.C., U.S.A.) with one-way Analysis of Variance (ANOVA). Post Hoc test (Fisher's LSD and Tukey-Kramer HSD) after ANOVA was used to compare treatments means. Significant levels were defined using p < 0.05 and 0.01.Results and discussion
Cell protection against production of reactive oxygen species (ROS)
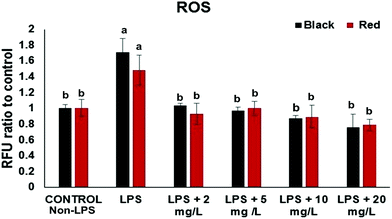
The intracellular ROS assay was performed in order to screen for cowpea varieties with greater potency in protecting cells from LPS-induced generation of ROS. At the concentration (2–20 mg GAE/L) the black and red cowpea varieties significantly inhibited the generation of ROS compared to the positive control (with LPS) (Fig. 1). Results for the light brown and white cowpea variety were not significant (data not shown). Previously, polyphenolics including the flavonols quercetin and kaempferol, inhibited TNF-α-induced generation of ROS in non-cancer human embryonic kidney HEK-293 cells.18 Another study demonstrated differential protective effect of quercetin and kaempferol against oxidative stress induced by proinflammatory stimuli in human hepatocyte-derived cell line, possibly through the modulation of antioxidant enzymes.19 Thus, results from this study suggest that cowpea polyphenolics may protect cellular components (e.g. DNA) from oxidative damage due to their ROS scavenging properties, thus, may have a role in prevention of ROS and this may be relevant to the prevention of chronic diseases.Effects of cowpea extracts on proinflammatory cytokines
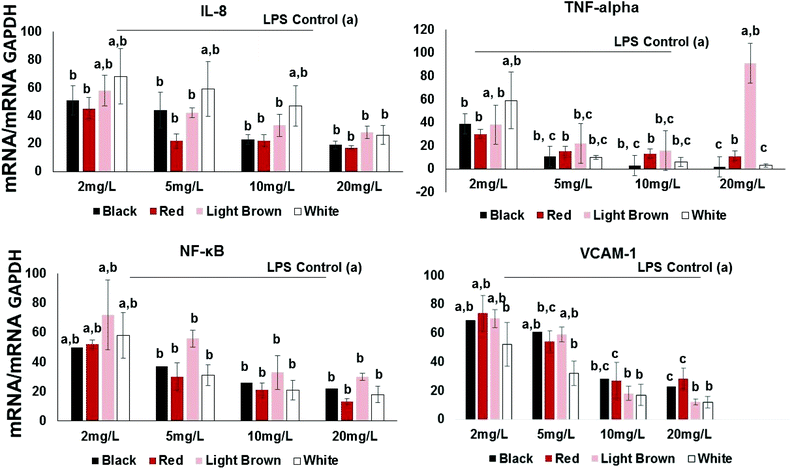
Pro-inflammatory cytokines such as interleukins (ILs) and tumor necrosis factor (TNF-α) are substances produced by resident or migrating cells (e.g. mast cells, macrophages and neutrophils).20 Thus, their inhibition by cowpea polyphenols may help prevent transcription of other pro-inflammatory molecules such as TNF-α known to activate NF-κB in immune responses. In this study, the proinflammatory cytokine/chemokine IL-8 was included in the investigation since it is known to promote phagocytosis of neutrophils, an event that causes the secretion of ROS21 and this is implicated in the etiology of several chronic disorders such as colon cancer and IBD.22 IL-8 and TNF-α, are inducible by LPS as well as transcription factors including NF-κB.Results show that all extracts concentration-dependently decreased the expression of IL-8 expression in LPS-stimulated CCD18Co cells (Fig. 2). This effect was significant across the concentration range of 2–20 mg GAE/L. At 2 mg GAE/L, the extract from black, red, brown and white cowpea varieties significantly (p < 0.05) inhibited IL-8 mRNA expression by 51, 45, 58 and 68% compared to LPS-treated cells, respectively (Fig. 2).
All varieties used in this study also showed significant inhibition of LPS-induced TNF-α gene expression (Fig. 2). At 2 mg GAE/L extracts concentration, the red variety showed the inhibition of LPS-induced TNF-α gene expression (30%), whereas both the light brown and black showed approximately 38 and 39%. The white cowpea variety had the inhibition of LPS-induced TNF-α gene expression (59%) at 2 mg GAE/L polyphenolic concentration (Fig. 2). This suggests that cowpea polyphenols inhibit proinflammatory cytokines including TNF-α and this has been shown to help prevent interactions between both circulating and resident leukocytes and the vascular endothelium by inhibiting TNF-related activation-induced cytokine (TRANCE) pathway, NF-κB activation, oxidative stress and increased expression of the CAMs.23
Effects of cowpea extracts on NF-κB gene expression
At 2 mg GAE/L extract concentration, red and black cowpea varieties induced inhibition of LPS-induced NF-κB gene expression (i.e. 50–52% inhibition compared to LPS-treated cells). The polyphenolic extracts from the light brown and white cowpea variety had no significant inhibitory effect on NF-κB mRNA (Fig. 2). In the cytoplasm, NF-κB is usually found in its inactive form bound to IκB-α, which prevents its translocation into the nucleus.24 However, proinflammatory cytokines such as IL-8 and TNF-α can activate a process where phosphorylation of IκBs, allows nuclear translocation of NF-κB.25 Activating NF-κB may induce the expression of CAMs, thus increases the transendothelial migration of leukocytes and other inflammatory cell to cell interactions.23 At 2 mg GAE/L, the extract from black, red, brown and white cowpea varieties significantly (p < 0.05) inhibited NF-κB mRNA expression by 50, 52, 72 and 58% compared to LPS-treated cells, respectively (Fig. 2). Overall, greater inhibition of NF-κB mRNA was achieved by the black and red and white cowpea varieties compared to the brown variety.Flavonols (quercetin and kaempferol) have been reported to inhibit the activation of NF-κB induced by cytokines in parenchymal liver cells, probably via protecting cells against oxidative species, inhibition of anti-inflammatory enzymes and down-regulation of the NF-κB pathway.19,26 The ability of cowpea flavonoids to down-regulate LPS-induced expression of NF-κB and TNF-α suggest their significant role in modulating downstream signaling of NF-κB dependent pathways.
Effects of cowpea extracts on VCAM-1 gene expression
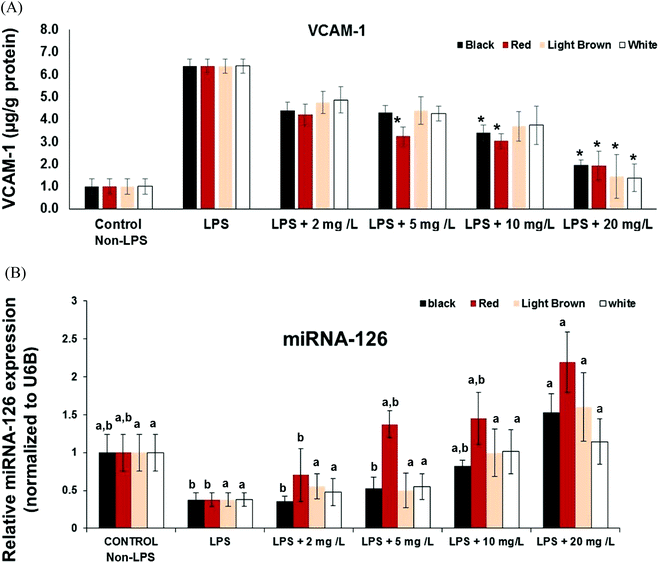
Another significant factor during inflammation is the activation of endothelial cells and subsequent leukocyte transendothelial migration and expression of cell adhesion molecules (CAMs). CAMs cause adhesive interactions between the endothelial cells and other blood constituents or extracellular matrix.27 Increased expression of CAMs (e.g. VCAM-1, ICAM-1 and Selectins) on the surface of endothelial cells is prominent when stimulated by proinflammatory molecules such as TNF-α, interleukins (e.g. IL-1), platelet-derived growth factor, and vascular endothelial growth factor (VEGF)3.28,29 Since vascular cell adhesion molecule VCAM-1 is a target gene for NF-κB, we also investigated whether cowpea polyphenolics influence the expression of VCAM-1mRNA. All cowpea extracts also had significant (p < 0.05) concentration–dependent reduction of LPS-induced VCAM-1 gene expression in the CCD18Co cells (Fig. 2), demonstrating that cowpea polyphenolics may have cardiovascular protective effects in endothelial cells. Similar results were also reported regarding açai and red Muscadine grape polyphenolics on NF-κB target genes VCAM-1, ICAM-1 and E-Selectin.30All cowpea extracts reversed LPS-induced up-regulation of VCAM-1 protein expression within the 2–20 mg GAE/L extract concentrations tested (Fig. 3A).
At the lowest polyphenolic concentration 2 mg GAE/L, the extract from black, red, brown and white cowpea varieties significantly (p < 0.05) inhibited VCAM-1 mRNA expression by 69, 74, 70 and 52% compared to LPS-treated cells, respectively (Fig. 2). Thus cowpea polyphenolics may prevent dysregulated expression of CAMs and this way prevent inflammation in non-cancer colon fibroblasts.
Effect of cowpea extracts on miR-126
MicroRNAs (miRs) are small (about 22 nt), non-coding RNA that post-transcriptionally regulate targeted mRNA and this way suppress protein synthesis by inhibiting the translation of protein from mRNA or by promoting the degradation of mRNA, thus hindering gene expression. miRNA, miR-126 is expressed in endothelial cells and also in CCD18-Co non cancer colon fibroblasts and, and VCAM-1 has been shown to be a potential target of miR-126.31 Inhibition of miR-126 reportedly increase leukocyte adherence to TNF-α-stimulated14 and LPS-stimulated cells.30 The effects of cowpea polyphenolics on regulation of cytokine signaling by miR-126 was investigated to provide further insight into its participation in modulating the anti-inflammatory activities of cowpea extracts.The expression of miR-126 in LPS-stimulated CCD18Co cells was inversely correlated to VCAM-1expression at both gene and protein levels, indicating its participation in the control of vascular inflammation (Fig. 2 and 3A). Cowpea polyphenolic extracts (2–20 mg GAE/L) concentration-dependently reversed LPS-induced induction of miR-126 (Fig. 3B). Polyphenolics from açai and red Muscadine grape seeds (5–20 mg GAE/L) have previously been reported to increase miR-126 expression in LPS-challenged HUVEC cells, and this response was inversely correlated with the expression of adhesion molecules (VCAM-1, ICAM-1, E-selectin and PECAM-1) at both gene and protein expression levels.30 This indicates miR-126 is normally involved in inhibiting VCAM-1 expression, confirming VCAM-1 regulation is mediated through miR-126.
Additionally, this study assessed the expression of VCAM-1 protein in the cell culture supernatants collected from control or miR-126-transfected CCD18Co cells using ELISA, but significant levels of VCAM-1 protein were not detected in these samples (data not shown). This does not mean that the antibody was ineffective; since VCAM-1 protein was readily detectable in LPS-treated endothelial cells (Fig. 3A); rather it showed that miRNA may also regulate stability or translation of target mRNAs as had been demonstrated by some authors.32 Previous studies14 showed that the 3′-UTR transcript for human VCAM-1 is perfectly complimentary to the nucleotides 604–625 of miR-126, which is responsible for decreasing cell adhesion and inflammation in endothelial cells. This further indicates involvement of miR-126 in modulating VCAM-1 expression during immune response.
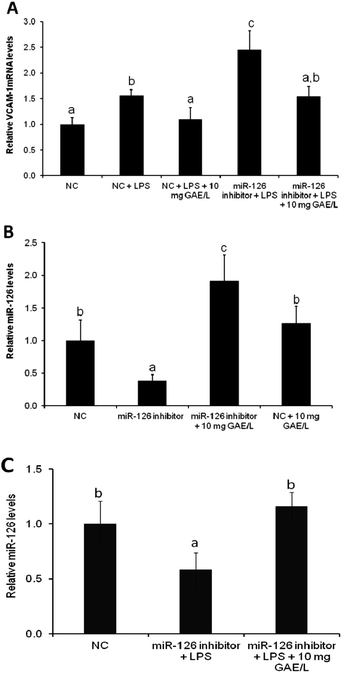
To confirm the involvement of miR-126 in cowpea-induced reduction of VCAM-1 mRNA expression of VCAM-1 was quantified in cells that were transfected with the antagomiR for miR-126 (Fig. 4) in the presence or absence of black cowpea extract at 10 mg GAE/L – a level that had been shown to significantly inhibit LPS-induced cytokine expression and ROS production. The black cowpea sample was chosen since it contained the major polyphenolic compounds identified in cowpea (Table 2). The extract decreased LPS-induced expression of VCAM-1 gene by 37% (Fig. 4A) and increased miR-126 mRNA levels by 92% (Fig. 4B) in cells transfected with the antagomiR and this suggests that the cowpea polyphenolic extracts regulated the expression of VCAM-1 mRNA, at least in part via induction of miR-126 (Fig. 4C). Since miR-126 regulate endothelial expression of VCAM-1,14 induction of miR-126 by cowpea polyphenolics following LPS stimulation demonstrates cowpea may protect endothelial cells against atherosclerotic risk factors by decreasing VCAM-1 expression as one of the underlying mechanisms.
Conclusions
This study was designed as a basis for future in vivo studies relevant to intestinal inflammation. Result from this in vitro study provide evidence of the anti-inflammatory potency of cowpea polyphenolics in modulating the level of inflammatory markers relevant to colon inflammation. Overall, cowpea polyphenolics extracts inhibited the generation of ROS and inflammation in non-cancer colon fibroblasts and the induction of miR-126 by cowpea's polyphenolic extracts at least in part seems to be involved as underlying mechanism. The anti-inflammatory effect of cowpea polyphenolic extracts on LPS-stimulated endothelial cells in vitro most likely involved the interactions of multiple polyphenolics present in cowpeas. Considering the importance of miR-126 in various physiological and pathological processes, further in vivo investigations on mechanistic and translational aspects of this mechanism would be essential in validating their relevance to human health.Acknowledgements
We thank USAID Dry Grain Pulses CRSP for the partial sponsorship of this project (grant no. 61-258), as well as Drs Jeff Ehlers and Phillip Roberts of University of California – Riverside, and Dr B. B. Singh, Texas A&M University, for generously providing the cowpea materials.References
- A. Alwan, D. R. MacLean, L. M. Riley, E. T. d'Espaignet, C. D. Mathers, G. A. Stevens and D. Bettcher, Monitoring and surveillance of chronic non-communicable diseases: progress and capacity in high-burden countries, Lancet, 2010, 376, 1861–1868 CrossRef.
- L. A. Bazzano, J. He, L. G. Ogden, C. Loria, S. Vupputuri, L. Myers and P. K. Whelton, Legume Consumption and Risk of Coronary Heart Disease in US Men and Women: NHANES I Epidemiologic Follow-up Study, Arch. Intern Med., 2001, 161, 2573–2578 CrossRef CAS.
- T. Takezaki, C. M. Gao, J. H. Ding, T. K. Liu, M. S. Li and K. Tajima, Comparative study of lifestyles of residents in high and low risk areas for gastric cancer in Jiangsu Province, China; with special reference to allium vegetables, J. Epidemiol., 1999, 9, 297–305 CrossRef CAS.
- T. Agurs-Collins, D. Smoot, J. Afful, K. Makambi and L. Adams-Campbell, Legume intake and reduced colorectal adenoma risk in African-Americans, J. Natl. Black Nurses Assoc., 2006, 17, 6–12 Search PubMed.
- A. S. Langyintuo, J. Lowenberg-DeBoer, M. Faye, D. Lambert, G. Ibro, B. Moussa, A. Kergna, S. Kushwaha, S. Musa and G. Ntoukam, Cowpea supply and demand in West and Central Africa, Field Crops Res., 2003, 82, 215–231 CrossRef.
- D. L. Pelletier, J. Frongillo, E. D. G. A. Schroeder and J. P. Habicht, The effects of malnutrition on child mortality in developing countries, Bull. W. H. O., 1995, 73, 443–448 CAS.
- WHOSIS World Health Organization Statistical Information System, World Health Statistics 2008, Geneva, Switzerland, 2008. http://www.who.int/whosis/whostat/2008/en/index.html. (Accessed on December 3 2011) Search PubMed.
- L. O. Ojwang, L. Dykes and J. M. Awika, Ultra Performance Liquid Chromatography–Tandem Quadrupole Mass Spectrometry Profiling of Anthocyanins and Flavonols in Cowpea (Vigna unguiculata) of Varying Genotypes, J. Agric. Food Chem., 2012, 60(14), 3735–3744 CrossRef CAS PubMed.
- L. O. Ojwang, Anti-inflammatory Properties of Cowpea Phenotypes with Different Phenolic Profiles, Dissertation, Texas A&M University, 2012 Search PubMed.
- D. De Stefano, M. C. Maiuri, V. Simeon, G. Grassia, A. Soscia, M. P. Cinelli and R. Carnuccio, Lycopene, quercetin and tyrosol prevent macrophage activation induced by gliadin and IFN-[gamma], Eur. J. of Pharmacol., 2007, 566, 192–199 CrossRef CAS PubMed.
- M. E. Gerritsen, W. W. Carley, G. E. Ranges, C. P. Shen, S. A. Phan, G. F. Ligon and C. A. Perry, Flavonoids inhibit cytokine-induced endothelial cell adhesion protein gene expression, Am. J. Pathol., 1995, 147, 278–292 CAS.
- Y. Naito and T. Yoshikawa, Green Tea and Heart Health, J. Cardiovas. Pharmacol., 2009, 54 Search PubMed.
- Y. D. Min, C. H. Choi, H. Bark, H. Y. Son, H. H. Park, S. Lee, J. W. Park, E. K. Park, H. I. Shin and S. H. Kim, Quercetin inhibits expression of inflammatory cytokines through attenuation of NF-κB and p38 MAPK in HMC-1 human mast cell line, Inflammation Res., 2007, 56, 210–215 CrossRef CAS PubMed.
- T. A. Harris, M. Yamakuchi, M. Ferlito, J. T. Mendell and C. J. Lowenstein, MicroRNA-126 regulates endothelial expression of vascular cell adhesion molecule 1, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 1516–1521 CrossRef CAS PubMed.
- T. Swain and W. E. Hillis, The phenolic constituents of Prunus domestica. I.—The quantitative analysis of phenolic constituents, J. Sci. Food Agric., 1959, 10, 63–68 CrossRef CAS.
- N. Banerjee, S. Talcott, S. Safe and S. U. Mertens-Talcott, Cytotoxicity of pomegranate polyphenolics in breast cancer cells in vitro and vivo: potential role of miRNA-27a and miRNA-155 in cell survival and inflammation, Breast Cancer Res. Treat., 2012, 136, 21–34 CrossRef CAS PubMed.
- Q. Meng, C. N. Velalar and R. Ruan, Effects of epigallocatechin-3-gallate on mitochondrial integrity and antioxidative enzyme activity in the aging process of human fibroblast, Free Radicals Biol. Med., 2008, 44, 1032–1041 CrossRef CAS PubMed.
- L. Soohyoung, K. Young-Jin, K. Sanghoon, L. Younghee, Y. C. Soo, P. Jinseu and K. Hyung-Joo, Inhibitory effects of flavonoids on TNF-α-induced IL-8 gene expression in HEK 293 cells, BMB Rep., 2009, 42, 265–270 CrossRef.
- I. Crespo, M. V. García-Mediavilla, M. Almar, P. González, M. J. Tuñón, S. Sánchez-Campos and J. González-Gallego, Differential effects of dietary flavonoids on reactive oxygen and nitrogen species generation and changes in antioxidant enzyme expression induced by proinflammatory cytokines in Chang Liver cells, Food Chem. Toxicol., 2008, 46, 1555–1569 CrossRef CAS PubMed.
- J. B. Calixto, M. M. Campos, M. F. Otuki and A. R. S. Santos, Anti-Inflammatory Compounds of Plant Origin. Part II. Modulation of Pro-Inflammatory Cytokines, Chemokines and Adhesion Molecules, Planta Med., 2004, 70(93), 103 Search PubMed.
- J. Kowalski, A. Samojedny, M. Paul, G. Pietsz and T. Wilczok, Effect of kaempferol on the production and gene expression of monocyte chemoattractant protein-1 in J774.2 macrophages, Pharmacol. Rep., 2005, 57, 107–112 CAS.
- M. Grimm, S. Elsbury, P. Pavli and W. Doe, Enhanced expression and production of monocyte chemoattractant protein-1 in inflammatory bowel disease mucosa, J. Leukocyte Biol., 1996, 59, 804–812 CAS.
- J.-K. Min, Y.-M. Kim, S. W. Kim, M.-C. Kwon, Y.-Y. Kong, I. K. Hwang, M. H. Won, J. Rho and Y.-G. Kwon, TNF-Related Activation-Induced Cytokine Enhances Leukocyte Adhesiveness: Induction of ICAM-1 and VCAM-1 via TNF Receptor-Associated Factor and Protein Kinase C-Dependent NF-κB Activation in Endothelial Cells, J. Immunol., 2005, 175, 531–540 CrossRef CAS.
- U. Siebenlist, Structure, regulation and function of NF-kappaB, Annu. Rev. Cell Biol., 1994, 10, 405 CrossRef CAS PubMed.
- M. Karin and Y. Ben-Neriah, Phosphorylation Meets Ubiquitination: The Control of NF-κB Activity, Annu. Rev. Immunol., 2000, 18, 621–663 CrossRef CAS PubMed.
- V. García-Mediavilla, I. Crespo, P. S. Collado, A. Esteller, S. Sánchez-Campos, M. J. Tuñón and J. González-Gallego, The anti-inflammatory flavones quercetin and kaempferol cause inhibition of inducible nitric oxide synthase, cyclooxygenase-2 and reactive C-protein, and down-regulation of the nuclear factor kappaB pathway in Chang Liver cells, Eur. J. Pharmacol., 2007, 557, 221–229 CrossRef PubMed.
- J. R. Privratsky, D. K. Newman and P. J. Newman, PECAM-1: Conflicts of interest in inflammation, Life Sci., 2010, 87, 69–82 CrossRef CAS PubMed.
- N. Morisaki, K. Takahashi, R. Shiina, M. Zenibayashi, M. Otabe, S. Yoshida and Y. Saito, Platelet-Derived Growth Factor Is a Potent Stimulator of Expression of Intercellular Adhesion Molecule-1 in Human Arterial Smooth Muscle Cells, Biochem. Biophys. Res. Commun., 1994, 200, 612–618 CrossRef CAS PubMed.
- R. J. Melder, During angiogenesis, vascular endothelial growth factor and basic fibroblast growth factor regulate natural killer cell adhesion to tumor endothelium, Nat. Med., 1996, 2, 992 CrossRef CAS PubMed.
- G. D. Noratto, G. Angel-Morales, S. T. Talcott and S. U. Mertens-Talcott, Polyphenolics from Açaí (Euterpe oleracea Mart.) and Red Muscadine Grape (Vitis rotundifolia) Protect Human Umbilical Vascular Endothelial Cells (HUVEC) from Glucose- and Lipopolysaccharide (LPS)-Induced Inflammation and Target MicroRNA-126, J. Agric. Food Chem., 2011, 59, 7999–8012 CrossRef CAS PubMed.
- N. Banerjee, H. Kim, S. Talcott and S. Mertens-Talcott, Pomegranate polyphenolics suppressed azoxymethane-induced colorectal aberrant crypt foci and inflammation: possible role of miR-126/VCAM-1 and miR-126/PI3 K/AKT/mTOR, Carcinogenesis, 2013, 34, 2814–2822 CrossRef CAS PubMed.
- J. E. Fish, M. M. Santoro, S. U. Morton, S. Yu, R.-F. Yeh, J. D. Wythe, K. N. Ivey, B. G. Bruneau, D. Y. R. Stainier and D. Srivastava, miR-126 Regulates Angiogenic Signaling and Vascular Integrity, Dev. Cell, 2008, 15, 272–284 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2015 |