Evaluation of copper iodide and copper phosphate nanoparticles for their potential cytotoxic effect
Dipranjan
Laha†
ab,
Debalina
Bhattacharya†
a,
Arindam
Pramanik
a,
Chitta Ranjan
Santra
a,
Panchanan
Pramanik
b and
Parimal
Karmakar
*a
aDepartment of Life Science and Biotechnology, Jadavpur University, Kolkata, India. E-mail: pkarmakar_28@yahoo.co.in; Fax: +9133-24137121; Tel: +91 33-24146710
bDepartment of Chemistry, Indian Institute of Technology, Kharagpur, India. Fax: +91 3222-255303; Tel: +91 3222-255221
First published on 3rd April 2012
Abstract
To explore the potential biological activities of transition metal-based nanoparticles (NPs), we synthesized two copper-based NPs, CuI and Cu3(PO4)2. The structural features of these NPs were determined by the X-ray diffraction (XRD), dynamic light scattering (DLS) and transmission electron microscopy (TEM). The size of CuI and Cu3(PO4)2 NPs were 35 ± 4.2 nm and 67 ± 6.3 nm respectively as determined by TEM. Cell viability, generation of reactive oxygen species (ROS), cell cycle and induction of apoptosis were assessed on human breast cancer cell line MCF7 after the treatment of these NPs. Exposure of CuI and Cu3(PO4)2 NPs decreased cell viability in a dose-dependent manner. Also, CuI NPs produced more ROS compared to Cu3(PO4)2 and presence of N-acetyl cysteine (NAC) along with NPs increased the cell survival. Cell cycle analysis indicated that after exposure of these NPs at their respective LD50 doses increased Sub G1 and G2/M peak after 8 h and 24 h of treatment respectively. Apoptosis study by AnnexinV-FITC staining showed slight increased in the early and late apoptosis after 8 h of treatment and most of the cells were dead after 24 h of treatment. Thus our observations suggest that the exposure of these two NPs induced dose-dependent cytotoxicity on MCF7 cell that is associated with ROS-mediated apoptosis.
1. Introduction
Recent development of nanoscience and nanotechnology, has introduced the potential applications of inorganic nanomaterials in life science.1–7 However, there are some detrimental effects of NPs on cells.8–12 These adverse effects of NPs on cells sometimes have been utilized for killing the cancer or tumour cells. The studies in this regard have been reported with TiO2, ZrO2, CdSe, Ag2S, CuO, CuS.13,14Different metal oxide NPs are used for a large variety of applications including catalysis, sensors, electronic materials, and environmental remediation.15,16 With the growing demand of metal oxide NPs, it is important to evaluate their toxicological effects as well as their application in therapy.17 Some of the metal oxide NPs are considered as potential chemotherapeutic agents either alone or in combination with light energy or other agents.18–23
Now a day, research on nanotoxicology has focused on several transition metals (d-block element).24 In particular, d-block element have been identified as a potential candidate for anticancer application due to their wide range of oxidative states or positively charged form. This help to interact with several bio-molecules especially DNA which led to develop metal-based drugs with promising therapeutic application.
Several studies have been reported the biological role of these transition metal or transition metal-based NPs. However the individual metal has different properties. Thus while comparing different transition metal oxide-based NPs like CuO, TiO2, ZnO, CuZnFe2O4, Fe3O4, Fe2O3, it has been demonstrated that CuO are most toxic and the cells killing ability is mediated by increase in intracellular reactive oxygen species.25 At present NPs of copper compounds have promising future in different field such as heat transfer fluid, gas sensor, antimicrobial agent, anticancer agent rather than copper complexes.26–29 Recently copper oxide NPs has shown to change gene expression profile in lung epithelial cells due to the release of Cu ions from CuO NPs.30 Genotoxic potential of such NPs was attributed to oxidative stress and subsequent DNA damage. From several studies, it is concluded that CuS, CuO NPs are cytotoxic and they can be manipulated to kill cancer cells. But very limited data is available on other copper-based NPs such as CuI, Cu3(PO4)2.
In the present work, we synthesized these NPs by chemical methods and their characterizations were done by different physical methods like DLS, TEM and XRD. We hypothesized that different copper-based NPs will have different abilities to generate oxidative stress and alter cell viability based on their size and surface charge.
The aim of our study was to evaluate in vitro response of these two synthesized NPs on human cancer cell lines MCF7-a widely used breast cancer cell line, in many toxicology assay. In this work we also investigated the effect of these NPs on cell cycle progression. Our results strongly suggest that NPs of CuI and Cu3(PO4)2 have potential cytotoxic effects and may have a promising future in anticancer therapy, if they tailored properly. Moreover, the NPs of these compounds are simple to synthesize and surfaces of all these NPs can be functionalized easily for targeted delivery.
2. Experimental
2.1 Materials
Copper sulphite, potassium iodide, copper chloride, sodium di hydrogen phosphate, hydrazine, triethanol amine was received from Merck, India. Gelatine was purchased from BDH chemical. All the chemical reagents used in our experiment were analytical grade, thus no need for any further purification. The water used in this work was deionized (Millipore) water.2.2 Synthesis of copper iodide NPs
CuI NPs were prepared by co-precipitation method reported by Bokshits et al. with little modification.31 In this method 6 mM potassium iodide was added into 6 mM of copper sulphite solution containing 0.05% gelatin as a stabilizing agent. Then 30 μl hydrazine was added drop wise to the above mixture and the reaction was continued for 1 h at 60 °C. After that, the NPs were collected by centrifugation at 4000 rpm. This particle was washed several times with millipore water followed by centrifugation. Then the CuI NPs were recovered and dried at 60 °C in hot air oven for 6 h.2.3 Synthesis of copper phosphate NPs
Cu3(PO4)2 NPs was synthesized in two steps: co-precipitation and mechanical milling of co-precipitation precursors. In this method 2 M of sodium dihydrogen phosphate was mixed with 3 M of copper chloride under stirring condition in presence of triethanolamine to control the size. Then the greenish precipitation obtained which was collected by centrifugation. This precipitation was further purified by washing several times with deionized water. The powder was obtained after dried at 70 °C in hot air oven. Finally, the solid powder was ball milled in conventional planetary ball miller for 12 h to get the desired Cu3(PO4)2 NPs.2.4 Nanoparticle characterizations
The phase formation and crystallographic state of copper-based NPs were determined by XRD with an Expert Pro (Phillips) X-ray diffractometer using CoKα radiation (α = 0.178897 nm). Samples were scanned from 20° to 80° of 2θ increment of 0.04° with 2 s counting time. Copper-based NPs were dispersed in millipore water to form diluted suspension of 0.5 mg ml−1 using sonicator for 30 min. When particles were completely dispersed in water then particle was analyzed by Brookhaven 90 plus particle size analyzer (DLS). The particle size and nanostructure were studied by high-resolution transmission electron microscopy in a JEOL 3010 (HRTEM), Japan operating at 200 keV.2.5 Cell lines, culture condition and treatments
Human breast cancer cell line MCF7 were cultured in RPMI-1640 media supplemented with 10% FBS and 100 U ml−1 penicillin–streptomycin at 5% CO2 and 37 °C. At 85% confluence, cells were harvested and sub-cultured into 90 mm plates, 6-well plates or 96 well plates according to experimental requirements. Cell were seeded for 24 h prior to the treatment with CuI and Cu3(PO4)2 NPs. All the treatments were performed at 37 °C at a density allowing experimental growth.2.6 MTT assay for cell viability
Viability of human breast cancer cell line MCF7 after exposure to various concentration of copper-based NPs i.e. CuI, Cu3(PO4)2 were determine by MTT assay.32 Briefly, cells were seeded in 96 well plates at 1 × 104 cells per well and exposed to copper-based NPs at the concentrations of 0 μg ml−1, 2 μg ml−1, 5 μg ml−1, 8 μg ml−1, 10 μg ml−1, 12 μg ml−1, 15 μg ml−1, 20 μg ml−1, 25 μg ml−1, 30 μg ml−1, 35 μg ml−1, 40 μg ml−1 and 50 μg ml−1 for 24 h. After incubation, cells were washed with 1 × PBS twice and incubated with MTT solution (450 μg ml−1) for 3–4 h at 37 °C. The resulting formazan crystals were dissolved in an MTT solubilisation buffer and the absorbances were measured at 570 nm by using a microplate reader (Biorad) and the value was compared with the control cells. Each variable was assessed through MTT assay in triplicate.2.7 Reactive oxygen species (ROS) assay
To check the ability of these two NPs to induce intracellular oxidation production, a fluorescent probe 2′,7′-dichlorofluorescein diacetate (DCFHDA, Sigma) was used as a reporter of ROS generation.33 Briefly, 5 × 105 cells were treated with CuI and Cu3(PO4)2 NPs at their respective median lethal (LD50) doses for 5 h and harvested. Then DCFDA (20 mM) was added to the cell suspension and kept for 30 min at 37 °C. So that DCFDA diffuses through the cell membrane, enzymatically hydrolysed by intracellular esterases and oxidized to produce a fluorescent 2′,7′-dichlorofluorescein (DCF) in the presence of ROS. The intensity of fluorescence is measured at 529 nm by spectroflurimeter. The intensity of fluorescence is proportional to the level of intracellular reactive oxygen species.2.8 DNA damage assay by immunolabeling
After exposure of these NPs for 4 h, cells were washed twice with 1 × PBS and fixed with freshly prepared 4% paraformaldehyde (Himedia, India) for 15 min at room temperature. After the fixation, cell were washed with 1 × ice cold PBS and permeabilized with 0.2% Triton X-100. Subsequently, the cells were blocked by 5% FBS and incubated with anti-H2AX antibody diluted in wash buffer (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100) containing 0.1% BSA and 0.05% Tween 20 in PBS for overnight at 4 °C in moist chamber. After incubation, the cells were washed and labeled with appropriate secondary antibody conjugated with Alexa Fluor 568. Finally cells were observed under a fluorescence microscopy (Leica).
100) containing 0.1% BSA and 0.05% Tween 20 in PBS for overnight at 4 °C in moist chamber. After incubation, the cells were washed and labeled with appropriate secondary antibody conjugated with Alexa Fluor 568. Finally cells were observed under a fluorescence microscopy (Leica).
2.9 Cell cycle analysis
To test the effect of these NPs on the cell cycle of human breast cancer cells MCF7, 1 × 106 cells were treated with and without NPs (control) at their respective LD50 doses. After 8 h and 24 h, the cell pellets were washed with 1 × PBS and fixed with chilled 80% ethanol and kept for 2 h at 4 °C. Prior to stain with 50 μg ml−1 propidium iodide (PI, Sigma), the cells were incubated for 1 h with 100 μg ml−1 of DNAse free RNAse A (SRL, India) at 37 °C. The cell cycle was analyzed with a Becton Dickinson (FACS Calibur) flowcytometer, equipped with an air-cooled 20 mW argon laser. 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 events were counted at each data point.34
000 events were counted at each data point.34
2.10 Apoptosis measurement by AnnexinV-FITC
In this study, 1 × 106 cells ml−1 suspensions of human breast cancer cells MCF7 were induced for apoptosis by addition of LD50 doses of each copper-based NPs and untreated cells were kept as control. Both control and test samples were incubated for 8 h and 24 h in a 37 °C, 5% CO2 incubator. Following the incubation, the cells were washed twice with 1 × PBS. After that, the cells at concentration of about 1 × 106 cells ml−1 were resuspended in 1 × Annexin binding buffer. 500 μl of each cell suspension was added to a plastic 12 × 75 mm FACS tube and 2 μg ml−1 of AnnexinV-FITC (sigma) and 0.5 μg ml−1 of propidium iodide (Sigma) were added to each cell suspension. Then the tubes were incubated at room temperature for exactly 15 min at dark condition. Finally, the fluorescence of cells was immediately determined by Becton Dickinson (FACS Calibur) flow cytometer.352.11 Analysis of nuclear morphology by DAPI staining
After exposure of these two NPs for 8 h and 24 h, cells were washed three times with 1 × PBS and stained with 4′,6-diamidino-2-phenylindole (DAPI) in Vectashield (0.2 g ml−1, Vector Laboratories Inc.). The percentage of cells with rupture and decondensed nuclei were counted under a fluorescence microscope (Leica) at different time interval. At least 300 nuclei were counted per point and photographs were taken.362.12 Statistical analysis
A student's t-test was used to calculate the statistical significance of changes. In all cases, differences are significant for P < 0.05. Data analysis was performed using the Origin Pro v.8 software (Origin Lab).3. Results
The XRD pattern of each copper-based NPs were compared and interpreted with standard data of JCPDS file (JCPDS international center for diffraction data, 1991). Fig. 1A shows the XRD pattern of CuI NPs, the characteristic peaks at 2θ = 29.59°, 34.30°, 49.29°, 58.50°, 61.33° and 72.90° which are marked respectively by their indices (111), (200), (220), (311), (222), (400) agreement with JCPDS card no 83–1107. The XRD pattern of synthesized Cu3(PO4)2 NPs is shown in Fig. 1B. The characteristic peaks at 2θ = 20.95°, 25.26°, 32.21°, 36.71°, 38.1°, 51.74°, 58.52° for Cu3(PO4)2 NPs which are good agreement with JCPDS card no 36-0203.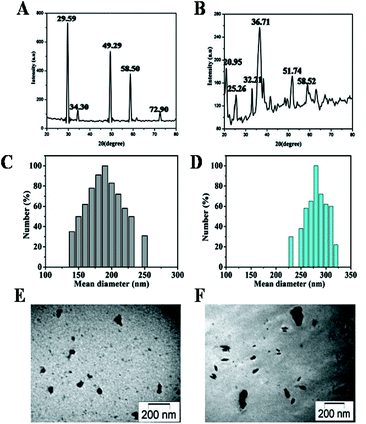 | ||
| Fig. 1 Physical characterization of CuI NPs and Cu3(PO4)2 NPs (A,B) X-ray diffraction patterns; (C,D) size distribution study; (E,F) TEM image, respectively. | ||
The hydrodynamic size of copper-based NPs was measured by DLS. Table 1 summarizes the size of two samples. Fig. 1C and 1D represent the frequency of size distribution of CuI and Cu3(PO4)2 NPs respectively. This study of these particles further confirmed the presence of stable, non aggregated NPs of copper compounds. TEM image of these copper NPs were taken. Information on mean size was calculated from measuring over 100 NPs in random field of view. Fig. 1E depicts the morphology of CuI NPs, having nearly spherical geometry with a mean size of 35 ± 4.2 nm. Fig. 1F shows the TEM image of prepared Cu3(PO4)2 NPs with diameter size 67 ± 6.3 nm. The presence of some bigger particle may be due to aggregation or overlapping of some small particles. The hydrodynamic sizes of the synthesized NPs were significantly larger than those indicated by their TEM images.
| Nanoparticles | Hydrodynamic size (nm) by DLS | Average diameter (nm) by TEM |
|---|---|---|
| CuI | 190 | 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ± ±![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4.2 4.2 |
| Cu3(PO4)2 | 280 | 67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ± ±![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6.3 6.3 |
Next, we carried out experiments to investigate the effect of synthesized NPs on human breast cancer cell lines MCF7. Cells were incubated with different concentrations (0 to 50 μg ml−1) of NPs for 24 h and then subjected to MTT assay. As seen in Fig. 2 the human breast cancer cell line MCF7 is less sensitive to Cu3(PO4)2 NPs than CuI NPs. Table 2 represents the LD50 dose of CuI NPs and Cu3(PO4)2 NPs is 2.5 μg ml−1 and 10 μg ml−1, respectively.
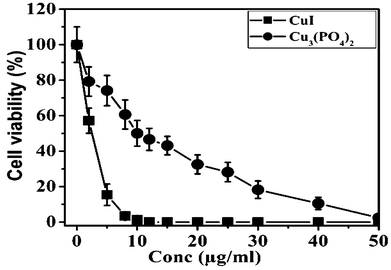 | ||
| Fig. 2 Cytotoxic effect of CuI and Cu3(PO4)2 NPs on human breast cancer cell line MCF7. Cells were incubated with increasing concentrations of CuI or Cu3(PO4)2 NPs for 24 h and their survivability was assessed by MTT. | ||
| Nanoparticles | LD50 doses of each nanoparticles (μg ml−1) in MCF7 cell |
|---|---|
| CuI | 2.5 |
| Cu3(PO4)2 | 10 |
Most of the metal or metal-based NPs are able to generate ROS and their cytotoxicity is mainly attributed to their ability to generate ROS.37,38 We next tried to determine the amount of ROS generated by these two NPs. After treating the cells with these two NPs separately at a concentration of their respective LD50 doses for 5 h, total ROS were determined. Fig. 3A presents generation of ROS increased almost 4 fold for CuI NPs, whereas for Cu3(PO4)2 NPs the ROS increased almost two fold. For ROS-mediated DNA damage, we next visualized γH2AX foci in cells treated with these two NPs. In Fig. 3B, both CuI NPs and Cu3(PO4)2 NPs treated cells showed γH2AX foci but in case of control cell no γH2AX foci were found. Also, presence of scavenger (NAC) significantly enhanced the survival of cells (Fig. 3C). The cell viability was increased up to 2.5 and 1.75 fold for CuI NPs and Cu3(PO4)2 NPs respectively as determined by MTT assay.
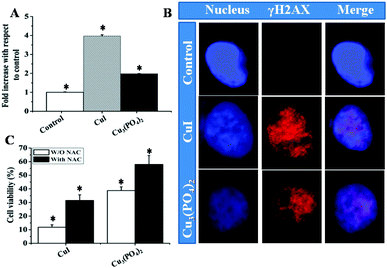 | ||
| Fig. 3 (A) Determination of reactive oxygen species (ROS) in human breast cancer cell line MCF7, treated with CuI or Cu3(PO4)2 NPs for 5 h at their respective LD50 doses; Each value represent mean ± SE of three independent experiments. * indicates p < 0.05 between control and treatment group. (B) CuI NPs and Cu3(PO4)2 NPs induce γH2AX foci in MCF7 cell. DNA counter stained with DAPI were shown in blue and γH2AX foci appeared in red. (C) Bar graph represent the MTT assay of MCF7 cells treated with CuI or Cu3(PO4)2 NPs in presence–absence of NAC for 24 h (*p < 0.05; n = 3). | ||
Cell cycle progression is impaired by DNA damage. Thus, we next analyzed cell cycle of MCF7 cells after incubating the cells with respective NPs at their LD50 doses for 8 h or 24 h. In the untreated control group, very little amount of Sub G1 peak was observed after 8 h or 24 h of incubation (Fig. 4A and 4B). However when cells were treated with respective NPs at their LD50 doses Sub G1 population of treated cells increased significantly after 8 h incubation (Fig. 4A and 4C) and in case of 24 h incubation (Fig. 4B) a slight increase in G2/M population were observed. The distribution of cells in different phases of cell cycle was shown in the Table 3.
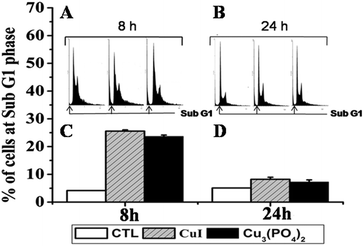 | ||
| Fig. 4 Flow cytometric cell-cycle distribution plots illustrating time course effect on the human breast cancer cell line MCF7 treated with CuI (2.5 μg ml−1) or Cu3(PO4)2 (10 μg ml−1) NPs. (A, B) Cell cycle phase distribution are given on the top of each bar; (C, D) histograms are the percentage of cell at Sub G1 peak either the untreated or CuI and Cu3(PO4)2 NPs treated cell. | ||
| G1 (%) | S (%) | G2/M (%) | ||||
|---|---|---|---|---|---|---|
| 8 h | 24 h | 8 h | 24 h | 8 h | 24 h | |
| Control | 45.03 ± 2.25 | 46.23 ± 2.84 | 16.23 ± 0.81 | 15.18 ± 0.75 | 34.18 ± 1.70 | 33.22 ± 0.85 |
| CuI | 44.14 ± 2.02 | 41.02 ± 1.89 | 7.01 ± 0.52 | 13.61 ± 0.61 | 24.00 ± 1.25 | 37.16 ± 1.25 |
| Cu3(PO4)2 | 45.74 ± 2.34 | 40.04 ± 1.75 | 4.01 ± 0.22 | 13.86 ± 0.59 | 28.06 ± 1.39 | 38.60 ± 1.54 |
We next tried to estimate the apoptosis by AnnexinV-FITC staining after incubating the cells with respective NPs at a dose, corresponding to their LD50 values, for 8 h or 24 h.
As seen in Fig. 5, in the untreated control group, very little amount of apoptosis occurred in both the time points. But about 14% and 8% cells were dead when cells treated with CuI NPs and Cu3(PO4)2NPs respectively for 8 h. After 24 h of incubation about 40–45% of the cells are dead for both the NPs and among them majority of the cells are in late apoptosis or necrosis. Finally, we studied nuclear morphology and nature of cell death by DAPI staining. About 17% and 13% cells are apoptotic in nature after CuI NPs and Cu3(PO4)2NPs treatment for 8 h respectively (Fig. 6A). Whereas about 23% and 14% cells respectively are necrotic in nature after 24 h incubation with these NPs (Fig. 6B).The necrosis after 24 h of exposure was again confirmed by ethidium bromide staining (data not shown). The time dependent effect of these two NPs suggest that NPs induced apoptotic cell death at 8 h turn into necrotic feature after long exposure time due to lack of phagocytosis in vitro.
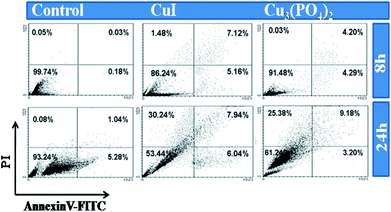 | ||
| Fig. 5 Apoptosis study by AnnexinV-FITC/PI method for human breast cancer cell line MCF7. Upper and below row represent untreated; CuI treated and Cu3(PO4)2 treated MCF7 at LD50 values for 8 h and 24 h respectively. The lower left quadrants of each panels show the viable cells, which exclude PI and are negative for FITC-AnnexinV binding. The upper right quadrants contain the non-viable dead, positive for FITC-AnnexinV binding and for PI uptake. The lower right quadrants represent the apoptotic cells, FITC-AnnexinV positive and PI negative demonstrating cytoplasmic membrane integrity. | ||
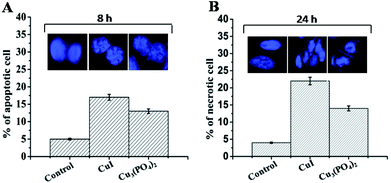 | ||
| Fig. 6 Bar graph represents the percentage of apoptotic (A) and necrotic (B) cells determine by DAPI staining. Value represents the mean ± SE. Figure in the inset represents the characteristic pattern of apoptotic and necrotic cells after DAPI staining. | ||
4. Discussion
Metallic copper and their different complexes have long been used as an algaecide, a fungicide, a nematocide and a molluscicide, as well as an antibacterial and anti-fouling agent. Numerous copper complexes have been synthesized for their potential application as chemotherapeutic drugs. Since NPs have greater biological activity than their counter complexes due to their high surface to volume ratio, we have given effort to synthesize different copper-based NPs and characterize them for their physical as well as biological activities. In the present study, we have successfully synthesized CuI and Cu3(PO4)2 NPs by co-precipitation method. CuI NPs is formed in the presence of iodide anion and hydrazine acts as reducing agent.39 Cu3(PO4)2 NPs are prepared by the reduction of grain size which is accomplished by the kinetic energy transfer from ball to powder. DLS result showed that the Cu3(PO4)2 NPs is gradually decreased with from around 1000 nm to 280 nm with increasing the ball-milling time. In the initial stage of milling, a fast decrease of grain size occurs which slows down after extended milling. Once the minimum steady state grain size is reached, further refinement ceases. But extended milling could not able to maintain the high stresses and hence reduction of grain size is limited in extended milling.Additionally their biological effects were studied by treating these two NPs on human breast cancer cell line MCF7. Interestingly, the two copper-based NPs have been shown different cytotoxic effect on MCF7 cells. Both of these NPs elicited a significant dose-dependent decrease in cell viability. Several researchers already reported that oxidative stress is the common mechanism of cell damage induced by many types of NPs. Between of these two NPs, CuI NPs produces more ROS compare to Cu3(PO4)2 NPs and consequently γH2AX foci is observed in the cells treated with these NPs. Moreover, using scavengers like NAC, the survivability of the cells increased significantly. Apart from generation of ROS, other mechanisms of action of these NPs are not elucidated yet but our study suggests that composition of NPs may play an important role in inducing cytotoxic effect. One possible explanation is that the particles are engulfed inside the cell and directly deliver to lysozyme and in the acidic pH of lysozyme these NPs may release ions. Each copper-based NPs may have different amount of charges attached with them, thus having more or less same diameter, they can penetrate inside the cells differentially. Also, the capability of releasing charges by them may differ and thus their killing ability. Moreover, different copper NPs may be metabolized differentially due to the presence of copper and after metabolize they interfere with the cellular signalling process associated with cell survival. The copper present in the NPs we have tested, may act as cofactors for the enzymes like cytochrome C, SOD and make them inactive and consequently induce their toxic effects. Also, depending on the chemical nature of the NPs, copper may directly induces DNA damage mediated apoptosis. Thus utilizing the transition metal properties of copper, a comprehensive study of copper-based NPs on different human cells needs to be done.
Generation of ROS is particularly support the sub G1 peaks in the cell cycle analysis. Since one of the major targets of ROS is cellular DNA, thus apoptosis due to DNA damage is seen after 8 h of incubation and further incubation for 24 h actually induced more dead cells. As a matter of fact, when we used LD50 doses for analyzing apoptotic cells, we have seen almost 40% of the cells are dead after 24 h, of which majority is necrosis. In our case, we have seen dead cells instead of apoptotic cells. This is perhaps due to the absence of phagocytosis in vitro, the apoptotic cells in their late phase behaves like necrotic cells. As a matter of fact, we also estimated the amount of lactate dehydrogenase (LDH) of the cells after 24 h of incubation in the presence of these NPs separately (data not shown). Amount of LDH also increased after 24 h for both the NPs. Thus, rapid generation of ROS, subsequent induction of DNA damage and apoptosis may likely the mechanisms of action of the NPs we have tested.
Thus, the cytotoxic properties of these two copper-based NPs may have a promising future in cancer therapy. Particularly, compared to other chemotherapeutic agents, their preparation is easy and cheap, raising the possibility of using them as potent chemotherapeutic agents. The results have implications for selectively tailoring the toxic effect and establishing predictive models for the design of various types of copper-based NPs with unique properties to kills specific cancer cells.
5. Conclusions
To conclude, this paper provides a special method to prepare bioactive CuI and Cu3(PO4)2 NPs and their biological effects have been explored on human breast cancer cell line MCF7. Biological assay revealed that both of them induced cell death by apoptosis followed by necrosis.Acknowledgements
The authors would like to acknowledge for financial support for this research work to the Department of Biotechnology (No. BT/PR14661/NNT/28/494/2010), Government of India and State Government fellowship scheme of Jadavpur University.Notes and references
- X. Wu, H. Liu, J. Liu, K. N. Haley, J. A. Treadway, J. P. Larson, N. Ge, F. Peale and M. P. Bruchez, Nat. Biotechnol., 2002, 21, 41–47 CrossRef
.
- M. Dahan, S. Lévi, C. Luccardini, P. Rostaing, B. Riveau and A. Triller, Science, 2003, 302, 442–445 CrossRef CAS
.
- D. S. Lidke, P. Nagy, R. Heintzmann, D. J. Arndt-Jovin, J. N. Post, H. E. Grecco, E. A. Jares-Erijman and T. M. Jovin, Nat. Biotechnol., 2004, 22, 198–203 CrossRef CAS
.
- T. J. Harris, J. J. Green, P. W. Fung, R. Langer, D. G. Anderson and S. N. Bhatia, Biomaterials, 2010, 31, 998–1006 CrossRef CAS
.
- D. Pantarotto, C. D. Partidos, J. Hoebeke, F. Brown, E. Kramer, J. P. Briand, S. Muller, M. Prato and A. Bianco, Chem. Biol., 2003, 10, 961–966 CrossRef CAS
.
- J. Nam, C. S. Thaxton and C. A. Mirkin, Science, 2003, 301, 1884–1886 CrossRef CAS
.
- R. Mahtab, J. P. Rogers and C. J. Murphy, J. Am. Chem. Soc., 1995, 117, 9099–9100 CrossRef CAS
.
- T. J. Brunner, P. Wick, P. Manser, P. Sposh, R. N. Grass, L. K. Limbech, A. Bruninink and W. J. Stark, Environ. Sci. Technol., 2006, 40, 4374–4381 CrossRef CAS
.
- C. Kirchner, T. Liedl, S. Kudera, T. Pellegrino, A. M. Javier, H. E. Gaub, S. Sto1lzle, N. Fertig and W. J. Parak, Nano Lett., 2005, 5, 331–338 CrossRef CAS
.
- Y. Jin, S. Kannan, M. Wu and J. X. Zhao, Chem. Res. Toxicol., 2007, 20, 1126–1133 CrossRef CAS
.
- K. O. Yu, C. M. Grabinski, A. M. Schrand, R. C. Murdock, W. W. B. Gu, J. J. Schlager and S. M. Hussain, J. Nanopart. Res., 2009, 11, 15–24 CrossRef CAS
.
- A. Nan, X. Bai, S. J. Son, S. B. Lee and H. Ghandehari, Nano Lett., 2008, 8, 2150–2154 CrossRef CAS
.
- Y. Guo, J. Zhang, L. Yang, H. Wang, F. Wanga and Z. Zhengb, Chem. Commun., 2010, 46, 3493–3495 RSC
.
- M. Ahamed, M. A. Siddiqui, M. J. Akhtar, I. Ahmad, A. B. Pant and H. A. Alhadlaq, Biochem. Biophys. Res. Commun., 2010, 396, 578–583 CrossRef CAS
.
- M. R. Hoffmann, S. T. Martin, W. Choi and D. W. Bahnemannt, Chem. Rev., 1995, 95, 69–96 CrossRef CAS
.
- P. V. Kamat, Chem. Rev., 1993, 93, 267–300 CrossRef CAS
.
- G. Grass, L. Rensingb and C. Rensing, Metallomics, 2011, 3, 1095–1097 RSC
.
- X. Yang, J. Liu, H. He, L. Zhou, C. Gong, X. Wang, L. Yang, J. Yuan, H. Huang, L. He, B. Zhang and Z. Zhuang, Part. Fibre Toxicol., 2010, 7, 1 CrossRef
.
- J. L. Yiyi Ye, J. Xu, L. Sun, M. Chen and M. Lan, Toxicol. in Vitro, 2010, 24, 751–758 CrossRef
.
- C. C. Huang, R. S. Aronstam, D. R. Chen and Y. W. Huang, Toxicol. in Vitro, 2010, 24, 45–55 CrossRef CAS
.
- C. Lee, C. Hong, H. Kim, J. Kang and H. M. Zheng, Photochem. Photobiol., 2010, 86, 981–989 CrossRef CAS
.
- B. Kang, M. A. Mackey and M. A. El-Sayed, J. Am. Chem. Soc., 2010, 132, 1517–1519 CrossRef CAS
.
- V. Chekhun, I. Todor, N. Lukyanova, S. Shpyleva, L. Naleskina, I. Khaetsky and G. Kulik, Exp Oncol., 2009, 3, 163–167 Search PubMed
.
- S. Rafique, M. Idrees, A. Nasim, H. Akbar and A. Athar, Biotechnol. Mol. Biol. Rev., 2010, 5, 38–45 CAS
.
- H. L. Karlsson, P. Cronholm, J. Gustafsson and L. Moller, Chem. Res. Toxicol., 2008, 21, 1726–1732 CrossRef CAS
.
- J. Zhang, J. Liu, Q. Peng, X. Wang and Y. Li, Chem. Mater., 2006, 18, 867–871 CrossRef CAS
.
- G. Rena, D. Hub, E. C. Chengb, M. A. Vargas-Reusc, P. Reipd and R. P. Allakerc, Int. J. Antimicrob. Agents, 2009, 33, 587–590 CrossRef
.
- Y. Xuan and Q. Li, Int. J. Heat Fluid Flow, 2000, 21, 58–64 CrossRef CAS
.
- M. Yanga, J. Hea, X. Hua, C. Yana, Z. Chenga, Y. Zhaoa and G. Zuo, Sens. Actuators, B, 2011 Search PubMed
.
- N. Hanagata, F. Zhuang, S. Connolly, J. Li, N. Ogawa and M. Xu, ACS Nano, 2011 Search PubMed
.
- Yu. V. Bokshits, Y. A. Fedutik and G. P. Shevchenko, Colloid J., 2004, 66, 30–33 Search PubMed
.
- M. Jo, T. H. Kim, D. Seol, J. E. Esplen, K. Dorko, T. R. Billiar and S. C. Strom, Nat. Med., 2000, 6, 564–567 CrossRef CAS
.
- B. Fahmy and S. A. Cormier, Toxicol. in Vitro, 2009, 23, 1365–1371 CrossRef CAS
.
- L. L. Vindelov, I. J. Christensen and N. I. Nissen, Cytometry, 1983, 3, 323–327 CrossRef CAS
.
- S. S. Fard, M. Jeddi-Tehrani, M. M. Akhondi, M. Hashemi and A. M. Ardekani, Avicenna J. Med. Biotechnol., 2010, 2, 53–61 Search PubMed
.
- B. Saha, A. Mukherjee, S. Samanta, P. Saha, A. K. Ghosh, C. R. Santra and P. Karmakar, Toxicol. in Vitro, 2009, 23, 1100–1109 CrossRef CAS
.
- I. Pujalté, I. Passagne, B. Brouillaud, M. Tréguer, E. Durand, C. Ohayon-Courtès and B. L'Azou, Particl. Fibre Toxicol., 2011, 3, 8–10 Search PubMed
.
- H. Yang, C. Liu, D. Yang, H. Zang and Z. Xi, J. Appl. Toxicol., 2009, 29, 69–78 CrossRef CAS
.
- S. Kapoor, R. Joshi and T. Mukherjee, Chem. Phys. Lett., 2002, 354, 443 CrossRef CAS
.
Footnote |
| † These two authors contributed equally in this work. |
| This journal is © The Royal Society of Chemistry 2012 |
