Extracellular biosynthesis and characterization of silver nanoparticles using Aspergillus flavusNJP08: A mechanism perspective
Navin
Jain
a,
Arpit
Bhargava
a,
Sonali
Majumdar
a,
J. C.
Tarafdar
b and
Jitendra
Panwar
*a
aCentre for Biotechnology, Department of Biological Sciences, Birla Institute of Technology and Science, Pilani, Rajasthan 333031, India. E-mail: drjitendrapanwar@yahoo.co.in; Fax: +911596244183; Tel: +919414411654
bCentral Arid Zone Research Institute, Jodhpur, Rajasthan 342003, India
First published on 18th November 2010
Abstract
The present study demonstrates an eco-friendly and low cost protocol for synthesis of silver nanoparticles using the cell-free filtrate of Aspergillus flavusNJP08 when supplied with aqueous silver (Ag+) ions. Identification of the fungal isolate was based on nuclear ribosomal DNA internal transcribed spacer (ITS) identities. Transmission electron microscopy (TEM) and energy dispersive spectroscopy (EDS) revealed the formation of spherical metallic silver nanoparticles. The average particle size calculated using Dynamic Light Scattering measurements (DLS) was found to be 17 ± 5.9 nm. UV-Visible and Fourier transform infrared (FTIR) spectroscopy confirmed the presence of extracellular proteins. SDS-PAGE profiles of the extracellular proteins showed the presence of two intense bands of 32 and 35 kDa, responsible for the synthesis and stability of silver nanoparticles, respectively. A probable mechanism behind the biosynthesis is discussed, which leads to the possibility of using the present protocol in future “nano-factories”.
Introduction
Nanotechnology has now started leaving the confines of laboratories; and conquering new applications to change our lives. Nanoparticles possess increased structural integrity as well as unique chemical, optical, mechanical, electronic and magnetic properties compared to large particles of bulk materials.1 These unique properties are derived due to variations in specific characteristics such as size, distribution and structure of particles. Due to these incredible properties, nanoparticles have become significant in recent years and nano-products are coming to the market rapidly. The fast growing field of nanotechnology presents great potential to influence various sectors in the areas of energy, environment, agriculture, healthcare and consumer goods. Therefore, it has built great expectations not only in the academic community but also among the investors, governments, and industries. According to estimates, the worldwide nano product market is estimated to reach $1 trillion by the year 2015.2 To compete with this tremendous demand, the synthesis of nanomaterials of specific composition, shape and size is a burgeoning area of research in the field of nanotechnology.Among various metals, silver has been known since ancient times as effective antimicrobial agent for the treatment of diseases, for food preservation and to keep water safe.3 With the recent advancements in the field of nanotechnology, silver nanoparticles have been widely used as a novel therapeutic agent extending their use as antibacterial, antifungal, antiviral, anti-inflammatory and anti-cancerous agents. The broad-spectrum antimicrobial properties of silver nanoparticles encourage its use in a large number of biomedical and environmental applications as well as in growing list of cosmetics, clothing and numerous consumer products.4,5
Conventional synthesis of silver nanoparticles involves a number of chemical and physical methods including chemical reduction in aqueous or non-aqueous solution,6 microemulsion,7 template,8 sonochemical,9 and microwave-assisted10 methods. However, all these methods are energy and capital intensive, employ toxic chemicals, and often yield particles in non-polar organic solutions, thus precluding their biomedical applications. Thus the need for the development of clean, reliable, bio-compatible and benign processes to synthesize nanoparticles leads to turning of more and more researchers to exploit biological systems as possible eco-friendly “nano-factories”.
It is well known that the biological systems can provide a number of metal or metal-containing particles in the nanometre range. Thus, they can be seen as a nanophase system in their own right and as the starting point for producing other novel nanophase systems. Biological methods for nanoparticle synthesis would help circumvent many of the detrimental features by enabling synthesis at mild pH, pressure and temperature and at a substantially lower cost. A number of micro-organisms have been found to be capable of synthesizing inorganic nano-composites either intra- or extracellularly. These include magnetotactic bacteria,11,12silica deposits by diatoms,13 and gypsum and calcium layers by S-layer bacteria.14 The secrets gleaned from nature combined with academic and scientific curiosity have led to the development of biomimetic approaches for the synthesis of nanoparticles.
Whilst various studies have been commenced on identification of microorganisms as possible nanofactories,15 very little work has been conducted on the actual mechanisms of nanoparticle formation in microorganisms. The defensive mechanism of cells for silver detoxification has been suggested as a biological pathway for the synthesis of silver nanoparticles.16 Ahmad et al. had shown that NADH-dependent enzymes are responsible for the biosynthesis of nanoparticles.17 The reduction mechanism seems to be initiated by electron transfer from the NADH by NADH-dependent reductase as electron carrier.
Recent studies demonstrated that the nitrate reductase enzyme system can be responsible for the bioreduction which leads to formation of silver nanoparticles.18 Similarly, studies with Fusarium oxysporum has shown that the reduction of silver ions occurs by a nitrate-dependent reductase and a shuttle quinone extracellular process.19 However, the exact mechanism of the formation of silver nanoparticles is yet to be elucidated.
In the present study, learning from nature's own sustainable way of bioremediation of the metal ions, we have utilized a soil fungal isolate Aspergillus flavusNJP08 to develop an eco-friendly and low cost protocol for the extracellular synthesis of silver nanoparticles. The plausible mechanism behind the synthesis of silver nanoparticles and their subsequent stabilization via capping proteins is discussed.
Experimental
Materials
All chemicals were of analytical grade and procured from Sigma Aldrich (India) or Merck (India) unless otherwise stated. All culture media were purchased from HiMedia (India). Standard protein molecular weight marker (SM-0431) was obtained from MBI Fermentas, USA.Isolation and identification of fungal isolate
The fungal isolate was isolated from metal-rich regions of Udaipur, Rajasthan, India (24°21′35′′N and 73°44′15′′E). Soil samples were used as inoculums for serial dilution technique and plated on Martin Rose Bengal Agar media. The plates were incubated at 28 °C for 4 days. Individual fungal colonies were picked and further purified by sub-culturing on Potato Dextrose Agar (PDA) media.The identification of the fungal isolate was carried out by morphological and microscopic observations (such as color, texture of the mycelia, spore formation pattern, etc.) followed by nuclear ribosomal DNA internal transcribed spacer (ITS) sequencing. The genomic DNA was isolated by CTAB extraction method using standard protocols.20 Internal transcribed spacer (ITS) regions were amplified using primers ITS1 and ITS4 with 5′-TCCGTAGGTGAACCTGCGG and 5′-TCCTCCGCTTATTGATATGC sequences respectively.21 PCR amplified products were sequenced using ABI prism DNA sequencer by BigDye terminator method. The resulting sequence was entered into the BLAST algorithm of National Centre of Biological Information (NCBI) database to obtain closely related phylogenetic sequences. The phylogenetic tree was constructed using Neighbor Joining method in Mega 4.0 software.22 The substitution model was based on Jukes and Cantor.23
Extracellular biosynthesis of silver nanoparticles
The fungal isolate was maintained on PDA slants (pH 5.6) at 28 °C with regular sub-culturing on fresh media. The stock culture (4 days old) was inoculated in 100ml of MGYP medium (0.3% malt extract, 1.0% glucose, 0.3% yeast extract, 0.5% peptone; pH 7.0) in 250 ml Erlenmeyer flasks. The inoculated flasks were then incubated at 28 °C for 4 days on a rotary shaker (150 rpm). Fungal mycelium were separated from the culture medium by centrifugation (8000 rpm, 10 min, and 4 °C) and washed thrice with sterile water. Typically, 10 g of biomass (fresh weight) was resuspended in 100 ml of sterile deionized Milli-Q water and further incubated for 72 h in an Erlenmeyer flask and agitated in similar conditions as described earlier. After incubation, biomass was separated by filtration using Whatman filter paper no. 1 and the cell-free filtrate was obtained. For synthesis of silver nanoparticles, aqueous silver nitrate solution at a final concentration of 1.0 mM was added to the reaction vessels containing cell-free filtrate and incubated at 28 °C on a rotary shaker (150 rpm) without light. Controls containing cell-free filtrate (without silver nitrate) as positive and pure silver nitrate solution (without cell-free filtrate) as negative controls were also run simultaneously along with the experimental flask in three replicates.Characterization of silver nanoparticles
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100. FTIR spectrum of samples was recorded on Shimazdu IR Prestige-21 FTIR instrument with a diffuse reflectance mode (DRS-8000) attachment. All measurements were carried out in the range of 400–4000 cm−1 at a resolution of 4 cm−1.
100. FTIR spectrum of samples was recorded on Shimazdu IR Prestige-21 FTIR instrument with a diffuse reflectance mode (DRS-8000) attachment. All measurements were carried out in the range of 400–4000 cm−1 at a resolution of 4 cm−1.
High-resolution transmission electron microscope (HR-TEM) micrographs of the sample were taken using the JEOL-2100 TEM instrument having selected area electron diffraction (SAED) attachment. The instrument was operated at an accelerating voltage of 200 kV.
Protein purification and one dimensional gel electrophoresis
For the purification of proteins from aqueous cell-free filtrate, solid ammonium sulfate was added slowly to control solutions containing extracellular proteins at a final concentration of 80% (w/v). The mixture was gently stirred for overnight at 4 °C. The resulting precipitate was subsequently collected by centrifugation at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 10 min at 4 °C. The protein obtained thereafter were resuspended in copious amount of water and dialyzed using a 12-kDa cut off dialysis bag made up of cellulose acetate membrane. The dialysis bag was pre-treated as per the manufacturer's instructions. The bag was then suspended in dialysis buffer (50 mM phosphate buffer, pH 7.2) and stirred slowly. The dialysis buffer was changed 3–4 times over a 24 h period. The dialyzed protein fraction was further analyzed by one dimensional SDS-PAGE (sodium dodecyl sulfate–polyacrylamide gel electrophoresis) as per standard procedure with adequate modifications.24 All samples were denatured in sample buffer containing 60 mM Tris (pH 6.8), 25% glycerol, 2% SDS, 14.4 mM 2-mercaptoethanol, 0.1% bromophenol blue and boiled for 5 min, followed by centrifugation at 8000 rpm for 1 min at 4 °C. Stained molecular weight marker was run along with samples. Electrophoresis was performed using Bio-Rad Mini Protean gel system at a constant voltage of 100 kV for 120 min. After electrophoresis, the gel was stained with Coomassie Brilliant Blue dye and was observed in a gel imaging system (Bio-Rad, USA).
000 rpm for 10 min at 4 °C. The protein obtained thereafter were resuspended in copious amount of water and dialyzed using a 12-kDa cut off dialysis bag made up of cellulose acetate membrane. The dialysis bag was pre-treated as per the manufacturer's instructions. The bag was then suspended in dialysis buffer (50 mM phosphate buffer, pH 7.2) and stirred slowly. The dialysis buffer was changed 3–4 times over a 24 h period. The dialyzed protein fraction was further analyzed by one dimensional SDS-PAGE (sodium dodecyl sulfate–polyacrylamide gel electrophoresis) as per standard procedure with adequate modifications.24 All samples were denatured in sample buffer containing 60 mM Tris (pH 6.8), 25% glycerol, 2% SDS, 14.4 mM 2-mercaptoethanol, 0.1% bromophenol blue and boiled for 5 min, followed by centrifugation at 8000 rpm for 1 min at 4 °C. Stained molecular weight marker was run along with samples. Electrophoresis was performed using Bio-Rad Mini Protean gel system at a constant voltage of 100 kV for 120 min. After electrophoresis, the gel was stained with Coomassie Brilliant Blue dye and was observed in a gel imaging system (Bio-Rad, USA).
In order to investigate the proteins which are bound to the surface of silver nanoparticles, samples were boiled with 1% SDS solution for 10 min followed by centrifugation at 8000 rpm for 10 min for collection of supernatant. The SDS- treated and untreated samples were further analyzed by the 12% SDS-PAGE as described earlier.
Results and discussion
Identification of fungal isolate
It is pre-requisite to identify the fungal isolate up to the species level in order to fully understand its nanoparticle synthesis mechanism. Amplification and sequencing of fungal rRNA gene resulted in 564 bp long nucleotide sequence, which has been deposited in NCBI GenBank (Accession Number: HM222933). The sequence was compared using BLAST algorithm and the closely related sequences were selected followed by their analysis using molecular evolutionary computing software MEGA 4.0. The phylogenetic tree was constructed which confirms that strain NJP08 exhibited 100% similarity with Aspergillus flavus (Fig. 1).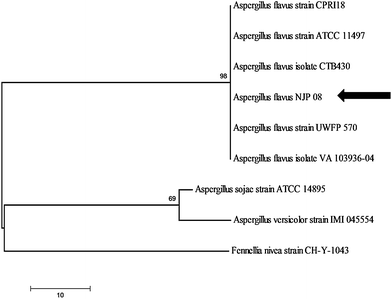 | ||
| Fig. 1 Phylogenetic tree showing genetic relationship between the isolate Aspergillus flavusNJP08 and other closely related reference microorganisms. Scale bar represents 10 nucleotide substitutions. | ||
Characterization of silver nanoparticles
Fig. 2 shows Erlenmeyer flasks containing the cell-free filtrate of Aspergillus flavusNJP08 without (A) and with silver nitrate (B) after completion of reaction at 72 h. The flask containing silver nitrate solution showed gradual change in color of reaction mixture from colorless to brown with intensity increasing during the incubation period. The negative control (pure silver nitrate solution without cell-free filtrate) did not show the characteristic change in color indicating that the synthesis is not a thermal and temporal process. | ||
| Fig. 2 Erlenmeyer flask containing cell-free filtrate of Aspergillus flavusNJP08 without (a) and with (b) silver nitrate solution (1 mM) after 72 h of reaction. | ||
UV-visible spectroscopy is one of the most widely used techniques for structural characterization of silver nanoparticles. The bio-transformed products were simultaneously characterized by UV-Vis. spectroscopy measurements performed at different time intervals to study the change in light absorption profile of the solution and increase in intensity. The absorption spectra of nanoparticles showed highly symmetric single-band absorption with peak maximum at ca. 421 nm with steadily increased in intensity as a function of time of reaction without any shift in the peak (Fig. 3). This indicates the presence of silver nanoparticles which is due to the excitation of surface plasmons; typical of silver nanoparticles.25 Inset to Fig. 3 represents the plot of absorbance at 420 nm at different time intervals of reaction. After 72 h of incubation, no further increase in intensity was recorded indicating complete reduction of precursor silver ions. It is observed that the graph follows the sigmoid kinetics which is characteristic of enzyme catalysis reactions. The kinetics of silver nanoparticles formation showed that more than 85% of the particles were formed within the 48 h of the reaction which suggests that the formation of silver nanoparticles is exponential in nature.
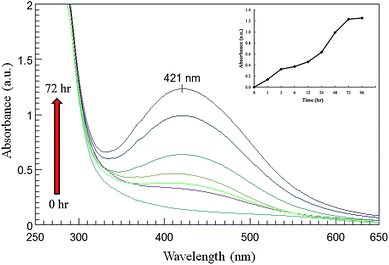 | ||
| Fig. 3 UV-Vis. spectra of cell free filtrate as a function of time. Arrow represents time of reaction (0, 3, 6, 12, 24, 48 and 72 h) with respective curves. Inset graph shows reaction saturation curve. | ||
Fig. 4 shows the UV-Vis. spectrum recorded from the reaction vessel after 72 h of reaction. An absorption peak at ca. 280 nm was observed which corresponds to aromatic amino acids of proteins (Fig. 4). It is well known that the absorbance peak at 280 nm arises due to electronic excitations in tyrosine and tryptophan residues of the protein.26,27 This observation indicates the presence of proteins secreted by fungus in the cell-free filtrate. The particles in the solution are thus stabilized by the capping agent that is likely to be proteins present in the cell-free filtrate.
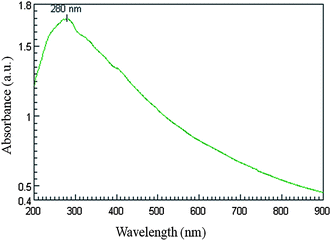 | ||
| Fig. 4 UV-Vis. spectra of cell-free filtrate showing presence of proteins. The absorption maxima at 280 nm arise due to electronic excitations in tyrosine and tryptophan residues of proteins. | ||
Stability of as-synthesized silver nanoparticles was monitored regularly for more than four months of completion of reaction. It was observed that the nanoparticle solution was extremely stable at room temperature, with no evidence of flocculation of particles as determined by UV-Vis. spectroscopy measurements. This indicates that the nanoparticles were well dispersed in the solution without aggregation. Monodispersity and chemical stability are highly desirable characteristics of the nanoparticles.28 This is an important aspect of synthesis of nanoparticles, since the lack of sufficient stability of many nanoparticles preparation has to some extent impeded the development of the real world applications of nanomaterials.29
FTIR measurements of the freeze-dried samples were carried out to identify the possible interactions between silver and bioactive molecules, which may be responsible for synthesis and stabilization (capping material) of silver nanoparticles. The amide linkages between amino acid residues in proteins give rise to well known signatures in the infrared region of the electromagnetic spectrum. FTIR spectrum reveals two bands at 1647 and 1543 cm−1 that corresponds to the bending vibrations of the amide I and amide II bands of the proteins respectively; while their corresponding stretching vibrations were seen at 3302 and 2926 cm−1 respectively (Fig. 5). The presence of the signature peaks of amino acids supports the presence of proteins in cell-free filtrate as observed in UV-Vis. spectra.
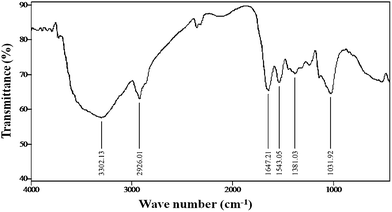 | ||
| Fig. 5 FTIR spectra of freeze-dried powder of silver nanoparticles. | ||
It is well known that protein–nanoparticle interactions can occur either through free amine groups or cysteine residues in proteins and via the electrostatic attraction of negatively charged carboxylate groups in enzymes.30 The two bands observed at 1381 and 1032 cm−1 can be assigned to the C–N stretching vibrations of the aromatic and aliphatic amines, respectively.31 These observations indicate the presence and binding of proteins with silver nanoparticles which can lead to their possible stabilization. FTIR results revealed that secondary structure of proteins have not been affected as a consequence of reaction with silver ions or binding with silver nanoparticles. It is important to understand though, that it is not just the size and shape of proteins, but the conformation of protein molecules that plays an important role.
TEM measurements were used to determine the morphology and shape of nanoparticles. Low magnification TEM micrographs (Fig. 6a) revealed that the particles are spherical in shape and uniformly distributed (monodispersed) without significant agglomeration. The particle size histogram (Fig. 7a) of silver nanoparticles shows that the particle size ranges from 10 to 35 nm and possess an average size of 17 ± 5.9 nm, although very tiny particles (ca. 5–10 nm) have also been observed that may be due to vigorous shaking. The frequency distribution observed from the histogram shows that almost 80% of the particles are in the 10- to 25-nm range. These results are in agreement with the values obtained by DLS measurements as shown in Fig. 7b.
 | ||
| Fig. 6 TEM micrographs showing uniformly distributed silver nanoparticles. (a) Low magnification image and (b) high magnification image. | ||
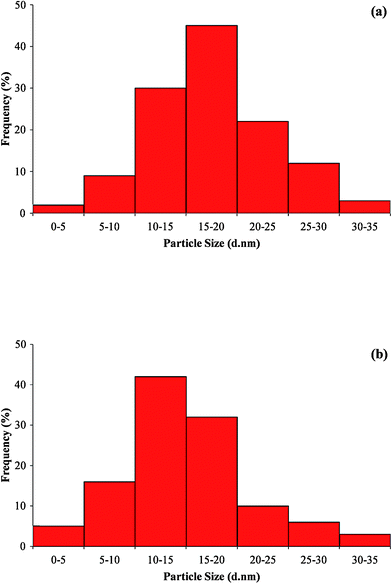 | ||
| Fig. 7 Particle size distribution histogram of silver nanoparticles from (a) transmission electron microscope (TEM) analysis and (b) particle size (DLS) analysis. | ||
The structural features of the individual silver nanoparticles can be observed more clearly in the HR-TEM images (Fig. 8). The particles are predominantly spherical with round edges and no sign of crystal defects. The inset to Fig. 8 shows the selected area electron diffraction (SAED) pattern from one of the silver nanoparticles which confirms the plane (111) of silver nanoparticles. The lattice fringes observed in micrograph also supports the crystalline nature of silver nanoparticles.
![HR-TEM
micrograph showing spherical shape of silver nanoparticle. Inset showing SAED pattern recorded from silver nanoparticle; the spot array is from [111] beam direction for a fcc crystal.](/image/article/2011/NR/c0nr00656d/c0nr00656d-f8.gif) | ||
| Fig. 8 HR-TEM micrograph showing spherical shape of silver nanoparticle. Inset showing SAED pattern recorded from silver nanoparticle; the spot array is from [111] beam direction for a fcc crystal. | ||
An EDS spectrum of drop coated film of silver nanoparticles is shown in Fig. 9. The presence of an optical absorption band at ∼3 eV reveals the presence of pure metallic silver nanoparticles.32 The spectrum shows mainly Ag (59 atom%) and only minor amounts of other elements with Nb (8.99 atom%) being the largest. Apart from this, the signals for C, N and O indicate the presence of proteins as a capping material on the surface of silver nanoparticles.
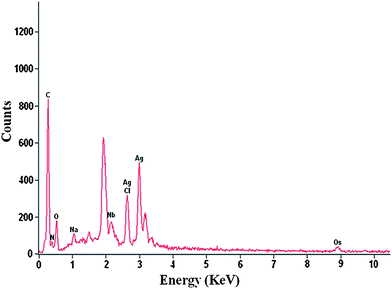 | ||
| Fig. 9 EDS spectrum showing the presence of elemental silver. | ||
Role of fungal proteins
Fungi are very well known to secrete large amount of proteins which play a major role in their life cycle. The majority of these proteins include hydrolytic enzymes such as amylases, cellulases and proteases.33 Most of the proteomic studies have focused on the proteins involved in the metabolic pathways, however, other proteins and their role still remains unknown.To identify the fungal proteins responsible for synthesis of silver nanoparticles, the cell-free filtrate was salted out overnight at 4 °C using ammonium sulfate precipitation method followed by centrifugation. The pellet fraction was subsequently dialyzed using a 12-kDa cut-off membrane. The protein profiles were compared by one dimensional SDS-PAGE followed by Coomassie Brilliant Blue staining (Fig. 10). The protein fraction clearly showed presence of two intense bands of 35 and 32 kDa (bands highlighted by arrows in Fig. 10, lane 2). These proteins can be responsible for the synthesis as well as stability of silver nanoparticles. Similarly, studies with Fusarium oxysporum showed presence of two extracellular proteins having molecular weight of 24 and 28 kDa responsible for the synthesis of zirconia nanoparticles.34
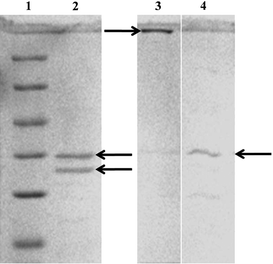 | ||
| Fig. 10 SDS-PAGE analysis of purified extracellular proteins secreted from Aspergillus flavusNJP08. Lane 1, molecular size marker (116.0 kDa β-galactosidase; 66.2 kDa, bovine serum albumin; 45 kDa, ovalbumin; 35 kDa, lactate dehydrogenase; 25 kDa, REase Bsp 98I; 18.4 kDa, β-lactoglobulin; 14.4 kDa, lysozyme). Lane 2, purified extracellular proteins. The arrows highlighted in Lane 2 indicate two intense bands of 35 and 32 kDa proteins. Protein profile bound to silver nanoparticles before (Lane 3) and after (Lane 4) boiling in 1% SDS solution. | ||
In order to identify the proteins bound to the surface of silver nanoparticles, the as-synthesized silver nanoparticles were treated with 1% SDS solution in boiling water bath for 10 min. SDS act as a denaturing agent and its treatment results in detachment of the surface bounded proteins from nanoparticles. The treated and un-treated samples were analyzed by 12% SDS-PAGE as described earlier. An intense band was observed at the transition of the stacking and resolving gel containing un-treated samples showing silver deposition (Fig. 10, lane 3). It can be predicted that the strong interactions between protein and nanoparticles prevent the migration of proteins into the resolving gel. Interestingly, the SDS treated samples showed the presence of a single intense band of ca. 35 kDa which is similar to the protein band present in lane 1 (Fig. 10, lane 4). Thus, it can be proposed that this 35 kDa protein acts as a capping material and confers stability to the silver nanoparticles. Further studies are in progress to understand the actual mechanism behind the synthesis and stability of silver nanoparticles. Efforts are underway in our laboratory to purify and characterize these two proteins.
Based on these experimental findings, a schematic presentation of the possible mechanism for synthesis of silver nanoparticles is speculated (Fig. 11). The synthesis process occurs in two steps: firstly, reduction of bulk silver ions into silver nanoparticles and, secondly, capping of the as-synthesized nanoparticles. The first step involves a 32 kDa protein which may be a reductase secreted by the fungal isolate, which may be specific for reduction of silver ions into silver nanoparticles. The second step involves 35 kDa proteins which bind with nanoparticles and confer stability. The protein–nanoparticle interactions can play a very significant role in providing stability to nanoparticles. Previous studies had also shown that the proteins bound to nanoparticle surface immediately upon contact with the nanoparticles.35 The protein adsorption has been widely studied and it has been found that protein adsorption depends on various factors such as electrostatic, hydrophobic and specific chemical interactions between the protein and the adsorbent.36FTIR results obtained during the present study also revealed that the carbonyl groups from amino acid residues and peptides of proteins have strong affinity to bind metal. However, the interaction between protein and nanoparticles is still not completely understood.
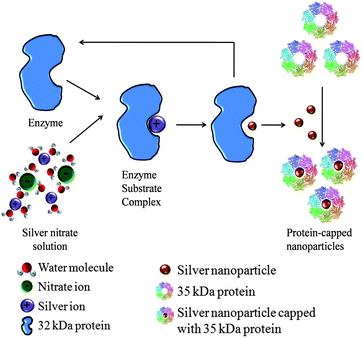 | ||
| Fig. 11 Mechanism showing the role of extracellular proteins in the synthesis of silver nanoparticles. | ||
This is the first report showing an immense study on the proteins responsible for the synthesis and stability (capping agent) of silver nanoparticles using the fungus Aspergillus flavusNJP08. The present study suggests that a protein with a molecular weight of 32 kDa may be responsible for the synthesis and one of 35 kDa is responsible for the stability of silver nanoparticles.
Conclusion
An eco-friendly and low cost protocol for biosynthesis of silver nanoparticles using cell-free filtrate of Aspergillus flavusNJP08 has been demonstrated. The formation of silver nanoparticles in extracellular secretion at room temperature can be easily handled during downstream processing. Since, the nanoparticles formed outside the cell are devoid of unnecessary cellular components, they can be directly used in various biomedical applications. The possibility of proteins as a stabilizing agent in silver nanoparticles is revealed by UV-Vis. spectroscopy and FTIR analysis. As a step towards understanding the mechanism of silver nanoparticles, the protein profile of the cell-free filtrate was analyzed. SDS-PAGE profiles of the extracellular proteins showed the presence of two intense bands of 32 and 35 kDa which are responsible for the synthesis and stability of silver nanoparticles, respectively. Efforts are underway in our laboratory to purify and characterize these two proteins to understand their mode of action and possible interactions with silver nanoparticles. Understanding the protein–nanoparticle interactions during the synthesis mechanism will lead to the possibility of using present system as future “nano-factories”. The possibility of rational use of this fungal strain isolated from metal-rich soils in synthesis of other important metal nanoparticles is an exciting and enormous future possibility.Acknowledgements
This research work was financially supported by the Indian Council of Agricultural Research, Government of India under the National Agricultural Innovation Project scheme (NAIP/C4/C-2032). Facilities provided by the Electron Microscopy & Nanoscience Laboratory, Punjab Agricultural University, Ludhiana are gratefully acknowledged. Navin Jain thanks the Council of Scientific and Industrial Research, Government of India for providing a research fellowship.Notes and references
- B. Bhushan, in Springer Handbook of Nanotechnology, Spinger-Verlag Berlin, Heidelberg, 2004 Search PubMed.
- M. C. Roco, J. Nanopart. Res., 2005, 7, 707–712 CrossRef.
- S. Silver, FEMS Microbiol. Rev., 2003, 27, 341–353 CrossRef CAS.
- M. Rai, A. Yadav and A. Gade, Biotechnol. Adv., 2009, 27, 76–83 CrossRef CAS.
- C. Marambio-Jones and E. Hoek, J. Nanopart. Res., 2010, 12, 1531–1551 CrossRef CAS.
- C. Petit, P. Lixon and M. P. Pileni, J. Phys. Chem., 1993, 97, 12974–12983 CrossRef CAS.
- J. N. Solanki and Z. V. P. Murthy, Colloids Surf., A, 2010, 359, 31–38 CrossRef CAS.
- S. Chen and D. L. Caroll, Nano Lett., 2002, 2, 1003–1007 CrossRef CAS.
- V. G. Pol, D. N. Srivastava, O. Palchik, V. Palchik, M. A. Slifkin, A. M. Weiss and A. Gedanken, Langmuir, 2002, 18, 3352–3357 CrossRef CAS.
- X. Li, L. Wang and X. Lu, J. Hazard. Mater., 2010, 177, 639–647 CrossRef CAS.
- D. R. Loveley, J. F. Stolz, G. L. Nord and E. J. P. Phillips, Nature, 1987, 330, 252–254 CAS.
- D. P. E. Dickson, J. Magn. Magn. Mater., 1999, 203, 46–49 CrossRef CAS.
- N. Kroger, R. Deutzmann and M. Sumper, Science, 1999, 286, 1129–1132 CrossRef CAS.
- D. Pum and U. B. Sleytr, Trends Biotechnol., 1999, 17, 8–12 CrossRef CAS.
- K. N. Thakkar, S. S. Mhatre, Y. Rasesh and R. Y. Parikh, Nanomed.: Nanotechnol., Biol. Med., 2010, 6, 257–262 CrossRef CAS.
- T. Klaus, R. Joerger, E. Olsson and C. G. Granqvist, Proc. Natl. Acad. Sci. U. S. A., 1999, 96, 13611–13614 CrossRef CAS.
- A. Ahmad, P. Mukherjee, S. Senapati, D. Mandal, M. I. Khan, R. Kumar and M. Sastry, Colloids Surf., B, 2003, 28, 313–318 CrossRef CAS.
- S. Anil Kumar, M. K. Abyaneh, S. W. Gosavi, S. K. Kulkarni, R. Pasricha, A. Ahmad and M. I. Khan, Biotechnol. Lett., 2007, 29, 439–445 CrossRef CAS.
- N. Duran, P. D. Marcato, O. L. Alves, G. I. De Souza and E. Esposito, J. Nanobiotechnol., 2005, 3, 8 CrossRef.
- J. Sambrook, E. F. Fritsch and T. Maniatis, Molecular Cloning: A Laboratory Manual, Cold Spring Harbour Laboratory Press, New York, 1989 Search PubMed.
- T. White, T. Bruns, S. Lee and J. Taylor, PCR Protocols: A Guide to Methods and Applications, ed. M. A. Innis, D. H. Gelfand, J. J. Shinsky, T. J. White, Academic Press, San Diego, 1990, pp. 315–322 Search PubMed.
- S. Kumar, J. Dudley, M. Nei and K. Tamura, Briefings Bioinf., 2008, 9, 299–306 Search PubMed.
- T. H. Jukes and C. R. Cantor, in Mammalian Protein Metabolism, ed. H. N. Munro, Academic Press, New York, 1969, pp. 121–123 Search PubMed.
- U. K. Laemmli, Nature, 1970, 227, 680–685 CAS.
- S. Gurunathan, K. Kalishwaralal, R. Vaidyanathan, D. Venkataraman, S. R. K. Pandian, J. Muniyandi, N. Hariharan and S. H. Eom, Colloids Surf., B, 2009, 74, 328–335 CrossRef CAS.
- J. Xie, J. Y. Lee, D. I. C. Wang and Y. P. Ting, ACS Nano, 2007, 1, 429–439 CrossRef CAS.
- N. Saifuddin, C. W. Wong and A. A. Nur Yasumira, E-J. Chem., 2009, 6, 61–70 Search PubMed.
- K. C. Bhainsa and S. F. D'Souza, Colloids Surf., B, 2006, 47, 160–164 CrossRef CAS.
- S. S. Shankar, A. Ahmad, R. Pasricha and M. Sastry, J. Mater. Chem., 2003, 13, 1822–1826 RSC.
- A. Gole, C. Dash, V. Ramakrishnan, S. R. Sainkar, A. B. Mandale, M. Rao and M. Sastry, Langmuir, 2001, 17, 1674–1679 CrossRef CAS.
- N. Vigneshwaran, A. A. Kathe, P. V. Varadarajan, R. P. Nachane and R. H. Balasubramanya, Langmuir, 2007, 23, 7113–7117 CrossRef CAS.
- P. Magudapathy, P. Gangopadhyay, B. K. Panigrahi, K. G. M. Nair and S. Dhara, Phys. B, 2001, 299, 142–146 CrossRef CAS.
- J. F. Peberdy, Trends Biotechnol., 1994, 12, 50–57 CrossRef CAS.
- V. Bansal, D. Rautaray, A. Ahmad and M. Sastry, J. Mater. Chem., 2004, 14, 3303–3305 RSC.
- J. L. Elechiguerra, J. L. Burt, J. R. Morones, A. Camacho-Bragado, X. Gao, H. H. Lara and M. J. Yacaman, J. Nanobiotechnol., 2005, 3, 6 CrossRef.
- Lynch and K. A. Dawson, Nano Today, 2008, 3, 40–47 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2011 |
