Copper proteomes, phylogenetics and evolution†
Leonardo
Decaria
a,
Ivano
Bertini
ab and
Robert J. P.
Williams
*c
aMagnetic Resonance Center (CERM) – University of Florence, Via L. Sacconi 6, 50019 Sesto Fiorentino, Italy
bDepartment of Chemistry – University of Florence, Via della Lastruccia 3, 50010 Sesto Fiorentino, Italy
cDepartment of Inorganic Chemistry – University of Oxford, South Parks Road, Oxford, UK OX1 3QR. E-mail: bob.williams@chem.ox.ac.uk
First published on 1st November 2010
Abstract
This paper is a continuation of our study of the connection between the changing environment and the changing use of particular elements in organisms in the course of their combined evolution (Decaria, Bertini and Williams, Metallomics, 2010, 2, 706). Here we treat the changes in copper proteins in historically the same increasingly oxidising environmental conditions. The study is a bioinformatic analysis of the types and the numbers of copper domains of proteins from 435 DNA sequences of a wide range of organisms available in NCBI, using the method developed by Andreini, Bertini and Rosato in Accounts of Chemical Research 2009, 42, 1471. The copper domains of greatest interest are found predominantly in copper chaperones, homeostatic proteins and redox enzymes mainly used outside the cytoplasm which are in themselves somewhat diverse. The multiplicity of these proteins is strongly marked. The contrasting use of the iron and heme iron proteins in oxidations, mostly in the cytoplasm, is compared with them and with activity of zinc fingers during evolution. It is shown that evolution is a coordinated development of the chemistry of elements with use of novel and multiple copies of their proteins as their availability rises in the environment.
Introduction
It is conventional today to analyse evolution both by comparative studies of organisms following Darwin or of DNA sequences from organisms present today using mathematical methods to deduce their history. Both methods are aided by the dating of fossils. They are very effective in tracing development following the Cambrian Explosion 0.54 billion years ago. As we stated in our previous paper1 the procedures do not describe well the evolution of organisms before this time and there are few helpful fossils of dates before 0.54 billion years ago. The fossils of this earlier period are largely imprints of soft-bodied organisms difficult to classify and to relate with certainty to today's organisms and of uncertain DNA content. It is our belief that in such circumstances the most revealing evidence of evolution lies in the changing nature of the chemical environment, largely of inorganic ions, together with the deduced evidence of organism inorganic chemistry, especially the metallome, in the agreed evolutionary order of anaerobic and then aerobic prokaryotes followed by single-cell and then multicellular eukaryotes from 3.5 to 0.5 or 0.4 billion years ago (Ga).2,3 From the quantitative evidence of the amounts of trace elements, of various element ratios and of isotope distribution in sediments it has proved to be possible to give a record of the likely availability of elements in the sea as they changed with time. The principal effect is due to the gradual rise in atmospheric oxygen giving rise to more oxidising conditions in the sea. The redox potential has risen from about −0.4 (anaerobic) at 3.5 Ga in the original oceans to +0.4 volts (aerobic) today with the solubilisation of elements from sulfide minerals. In parallel with these analytical studies we and others have examined the general uses of the elements in proteins in organisms, that is their metallomes, using bioinformatic analysis of organisms extant today, e.g. modern prokaryotes, plants and animals while judging their times of evolution from general biochemical studies.4–8 This paper will give details of the presence of copper proteins in organisms looking especially at the duplication of the copper proteins which have evolved with related properties much as we did in the study of zinc.1Methods
We have investigated 435 complete proteomes, 52 from archaea, 337 from bacteria (247 aerobic and 90 anaerobic) and 46 from eukarya available in NCBI. Our chosen example here is that of the copper proteins, primarily involved in homostasic, carrier (chaperone) functions, redox reactions and electron transfer. Knowledge of the site structures allows us to recognise the copper binding domains in a protein. To obtain our starting data set, which consists of 44 proposed Cu-binding domains, some of them with one or more metal associated binding patterns (MBP) (Table S1, ESI†), we used the prediction method published by the group of one of us.7 As reported in that reference paper, we applied the HMMER program to search the NCBI refseq proteins database for matches to the hidden Markov models (HMMs) representing the selected domains. The HMMs were taken from the Pfam database without modification. We selected 10−3 as Evalue cut-off. For multi-domain proteins (e.g.ammonia monooxygenase made from 3 distinct Pfam domains) we considered as true positives only the retrieved sequences containing all the reference Pfam domains. Table 1 reports the Pfam domain composition of all the related Cu-binding proteins and their functions, and the number of ions bound.| Pfam domain | No. of bound ions | Function |
|---|---|---|
| Ald_Xan_dh_C2 | 1 | Electron_carrier |
| Monooxygenase_B | 3 | Ammonia monooxygenase |
| Biopterin_H | 1 | Aromatic-AA hydroxylase |
| CcoS | 1 | Copper_chaperone |
| CdhC | 1 | Carbon monoxide dehydrogenase |
| Cmc1 | 1 | Copper_chaperone |
| CopB | 1 | Copper_homeostasis |
| CopC | 1 | Copper_homeostasis |
| CopD | 1 | Copper_homeostasis |
| Copper-bind | 1 | Electron_carrier |
| Copper-fist | 1 | Transcription |
| COX1 | 1 | Electron_carrier |
| COX17 | 1 | Copper_chaperone |
| COX2 | 1 | COX 2 |
| CtaG_Cox11 | 1 | Copper_chaperone |
| Ctr | 1 | Copper_homeostasis |
| Cu_amine_oxid | 1 | Amine oxidase |
| Cu_bind_like | 1 | Electron_carrier |
| Cu2_monoox_C | 1 | Ascorbate dep. Monooxygenase |
| Cu2_monooxygen | 1 | |
| Cu-oxidase | 1 | Laccase-like |
| Cu-oxidase_2 | 1 | |
| Cu-oxidase_3 | 2 | |
| Cu-oxidase_4 | 4 | Laccase-like |
| CutA1 | 1 | Copper_homeostasis |
| CutC | 1 | Copper_homeostasis |
| Glyco_hydro_10 | 1 | Glycosyl hydrolase |
| Hemocyanin_M | 1 | Copper_homeostasis |
| HMA | 1 | Copper_homeostasis |
| Lysyl_oxidase | 1 | Lysyl_oxidase |
| Metallothio | 1 | Copper_homeostasis |
| Metallothio_11 | 1 | Copper_homeostasis |
| Metallothio_5 | 1 | Copper_homeostasis |
| Metallothio_Pro | 1 | Copper_homeostasis |
| Metallothionein | 1 | Copper_homeostasis |
| NlpE | 1 | Copper_homeostasis |
| NosD | 1 | Copper_chaperone |
| NosL | 1 | Copper_chaperone |
| Sod_Cu | 1 | Superoxide dismutase |
| Tyrosinase | 1 | Tyrosinase |
| Uricase | 1 | Uricase |
Results
In order to give a comparative account of the data and their analysis we have considered organisms in the following ways: the prokaryotes have been divided into the major groups of archaea and eubacteria, and both have been further divided into aerobic and anaerobic. The average content of copper proteins of each prokaryotic group has been used for comparative purposes. Amongst eukaryotes we have divided them into single-cell and multicellular examples and then considered them with respect to their complexity, using selective organisms in the order: single-cell eukaryotes S. cerevisiae, T. brucei and P. falciparine and multicell eukaryotes, C. elegans, D. melanogaster, A. thaliana and H. sapiens. Each chosen organism has been observed to be similar in DNA sequences and numbers of duplications to those in several other organisms in the group to which it belongs. The particular organism described can therefore be taken as indicative of the nature of a group.The activities of the proteins in all the organisms have been divided using their major four separate functions: homeostatic proteins, chaperones, electron transfer proteins and oxidases. The homeostatic proteins include the metallothioneins and the copper pumps. The oxidases are treated at first as a sum of all such enzymes but later we shall discuss their further functional divisions together with the superoxide dismutases. We shall not refer to either transcription factors or hydrolytic enzymes which were the major groups in our analysis of zinc proteins.
We turn now to a more detailed description of the copper oxidases which can be divided in three ways: by the number of copper sites, by the structural nature and domains in one protein, and by their organic substrates, Table 1. The numbers of copper atoms vary from 1 to 4 (and perhaps one or two more in caeruloplasmin) and the enzymes are grouped under the Enzyme Commission EC.1 label. The types of copper are also described structurally as Type I (electron transfer proteins with one copper), Type II (a single copper) with Type III (a pair of linked copper atoms) where Types II and III ions form the site of reaction of oxygen in the complicated oxidases such as lactase, EC.1.10.3, and ascorbate oxidase, EC.1.14.17. Here oxygen goes directly to water and oxidation of substrate is at a remote site. Of the other enzymes some such as galactose oxidase and amine oxidases, EC.1.4.3, have but one copper while tyrosinase, also known as catechol oxidase, EC. 1.14.18, has two linked coppers. Finally Superoxide Dismutase EC.1.15 has a copper close to a zinc site. Now these oxidases can be separately recognised in the genome by the way that copper atoms are chelated or their cofactors bind, see Table S1, ESI† and methods above. Some of the copper-dependent hydroxylases are dependent on an initial reduction of the organic substrate with release of one water molecule as they introduce one atom of oxygen only into the organic substrate much as does cytochrome P-450. We know of at least three reducing cofactors, NADH or NADPH, pteridine and ascorbate. We present the data either in terms of total numbers of copper proteins as in the Tables or as percentages of the genome as in the Figures.
There are extremely few, perhaps no, copper proteins in all the anaerobic archaea or eubacteria. There are only a few copper proteins in any of the four classes in aerobic archaea or eubacteria, one with a total genome less than 1500 and the other with a greater number. The data on EC.1 oxidases are given in Table 2. In fact there no noticeable differences in copper proteins between the bacteria of low gene and those of high gene content (not shown). Aerobic prokaryotes and all eukaryotes, animals and plants, have a copper domain in cytochrome oxidase but it is coded in the mitochondrial DNA in eukaryotes and is not included in our search. Chloroplasts in plants also have a copper electron transfer protein, plastocyanin, and it too is not included in our analysis.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (oxidoreductases) Cu–proteins for the analyzed groups of organisms. * = average value for archea, aerobic and anaerobic bacteria
1 (oxidoreductases) Cu–proteins for the analyzed groups of organisms. * = average value for archea, aerobic and anaerobic bacteria
| Proteome | No. Total | No. EC:1 | |
|---|---|---|---|
| Archea (*) | 2176 | 8 | 1 |
| Bacteria Anaerobic (*) | 2749 | 6 | 1 |
| Bacteria Aerobic (*) | 3792 | 18 | 8 |
| S. cerevisiae | 5880 | 29 | 12 |
| P. falciparum | 6265 | 7 | 1 |
| T. brucei | 9279 | 5 | 2 |
| C. elegans | 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 844 844 |
46 | 26 |
| D. melanogaster | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 513 513 |
70 | 47 |
| H. sapiens | 37![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 742 742 |
82 | 54 |
| A. thaliana | 32![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 165 165 |
245 | 144 |
Striking features in the eukaryotes are the rapid increase in the numbers of all four groups of copper proteins with complexity of the multicellular organisms and the even greater increase in plants, illustrated by Arabidopsis. The data for EC.1 oxidases are given in Table 2 but they do not show in the percentages, Fig. 1. The fungi form a group with a relatively steady number of all four kinds though in larger numbers than in the animals (not shown).
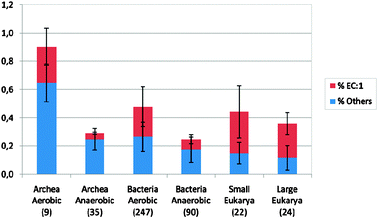 | ||
| Fig. 1 The total percentages of copper proteins including oxidases EC.1 in prokaryotes and eukaryotes. The percentages must be taken together with the total numbers so that the diversity of copper proteins, much greater in eukaryotes, Table 2, can be appreciated. The numbers in brackets refer to the toal number of organisms in each group. | ||
Discussion
The description and analysis of copper proteins and their probable evolution of them all jointly has been described in several previous publications.2–10 The main conclusions are that copper was not used by the earliest anaerobic prokaryotes, as it was not an available element before there was oxidation of sulfides. Free copper ions in organisms are known to be poisonous and hence cells have always had proteins for maintaining a very low level of total copper, especially in their membranes and cytoplasm. The control is managed through storage in homeostatic buffer proteins, such as metallothioneins in the cytoplasm and entry and exit pumps in the outer membrane. However copper became more and more valuable in cells as oxygen became more available, especially in oxidases in eukaryotic vesicles and outside cells. Here iron cannot be used since the ferrous ion is readily dissociated from proteins, oxidised and loses its function. Even the porphyrin of heme iron is susceptible to oxidation. The value of copper therefore increased externally as seen from unicellular to multicellular eukaryotes, Table 2. Its enzymes are used in the synthesis of extracellular matrices, oxidases for cross-linking phenolic units in plants and lysine oxidase in animals, absent in plants, for cross-linking of collagens. Many oxidative processes are required, especially for the production of messenger organic molecules such as adrenaline and amidated peptides in vesicles more notably in animals than in plants, Table 3. The presence of copper generally also raises risks such as chemical oxidative stress, possibly associated with Alzheimer's Disease for example, due to partial reduction oxygen to superoxide. Superoxide is removed rapidly by Cu/Zn superoxide dismutases. In this paper we have approached the problem of the evolution and these uses of copper in organisms in a different way from the above general descriptions by examining the duplication of the enzymes.| Function | H. Sapiens | A. thaliana |
|---|---|---|
| Copper_chaperone | 4 | 6 |
| Copper_homeostasis | 21 | 68 |
| Laccase | 5 | 39 |
| Monooxygenase | 0 | 0 |
| Aromatic-AA hydroxylase | 6 | 0 |
| Ascorbate dep. Monooxygenase | 12 | 0 |
| Tyrosinase | 3 | 0 |
| Ammonia monooxygenase | 0 | 0 |
| Superoxide dismutase | 3 | 8 |
| Carbon monoxide dehydrogenase | 0 | 0 |
| Amine oxidase | 4 | 10 |
| Lysyl_oxidase | 5 | 0 |
| Uricase | 0 | 1 |
| COX2 | 2 | 1 |
| Multicopper-oxidase | 2 | 2 |
| Glycosyl hydrolase | 0 | 4 |
| Electron_carrier | 3 | 23 |
| Transcription | 0 | 0 |
We shall follow the ideas which Ohno pointed out that while mutation can improve individual protein function it can hardly provide new functions without impairing the existing function of a protein.11 Duplication is therefore essential for novel functions prior to mutation.12Tables 2–4 and Fig. 1–3 show that duplication is very extensive amongst certain copper proteins as it was amongst particular but different zinc proteins. We observe first the great difference between the copper and zinc proteins described previously. There are extremely few copper transcription factors or hydrolases in marked contrast to those of zinc,1 which is in virtually no oxidases, and they are largely in different cell compartments. These features are indicative of the separate nature of the two metals. Copper is of use in oxidations as it can change valence but, as stated above, it presents a risk, especially in association with the cell nucleus. Zinc is more available and useful for hydrolytic reactions, it is nearly as powerful a Lewis acid as copper but unlike copper it cannot catalyse redox reactions. It can also act in signalling even to the nucleus in transcription factors as it is of low risk. The data show that duplication is therefore very selective to both different metal ions, proteins and enzymes and is characteristic of particular groups of organisms. For example oxidases are in greater numbers in plants but hydroxylases and transcription factors are more numerous in animals, Table 3. We therefore have to consider that the multiplied functions are for selected purpose—copper in certain oxidases, different in different organisms, and zinc in certain hydrolases and transcription factors. By far the greatest multiplications are seen in enzymes required for either the management of connective tissue and of messenger systems, copper for transmitters, zinc for hormones—both for external products. Moreover as we show in Table 4 there are large increases in the heme iron cytochrome P-450, also valuable in hormone synthesis, and the ferrous oxy-glutamate-dependent oxidases in parallel with the increases of copper oxidases. It would appear that duplication is not random though subsequent mutation may be but is perhaps preferentially in the duplicated proteins.12,13
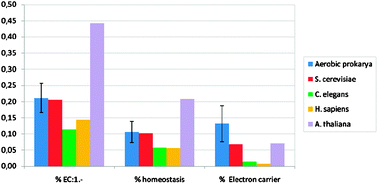 | ||
| Fig. 2 The percentage of three kinds of copper domains from five different organisms, see Table 1 for numbers. | ||
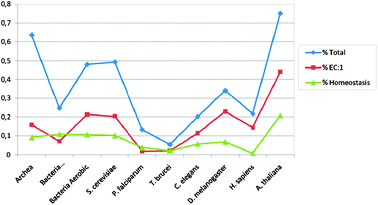 | ||
| Fig. 3 A graphical presentation of the percentage of domains of some proteins in a more extensive list of organisms. Tables 2 and 3 give numbers for these organisms and their total domain size as well as data for some proteins of very low multiplicity. | ||
| No. Cu EC:1 (oxidoreductases) | No. Fe-dependent oxygenases | No. Fe p450 | No. Heme peroxidases | |
|---|---|---|---|---|
| Note. No. Cu EC:1 is the number of copper domains and some proteins have three or four domains, see Table 1. | ||||
| Archea (*) | 1 | 0 | 0 | 0 |
| Bacteria Anaerobic (*) | 1 | 0 | 0 | 0 |
| Bacteria Aerobic (*) | 8 | 1 | 5 | 1 |
| S. cerevisiae | 12 | 1 | 3 | 1 |
| P. falciparum | 1 | 0 | 0 | 0 |
| T. brucei | 2 | 9 | 2 | 0 |
| C. elegans | 26 | 8 | 76 | 14 |
| D. melanogaster | 47 | 26 | 97 | 14 |
| H. sapiens | 54 | 9 | 70 | 16 |
| A. thaliana | 144 | 116 | 268 | 194 |
In our previous paper1 we drew attention to the parasitic organisms plasmodia and trypanosomes but they were not outstandingly different from other single-cell eukaryotes in zinc protein content. In the case of the copper enzymes we observe that the parasites have very few if any oxidases or other copper proteins except two or three for homeostasis or which act as chaperones. They then behave as single-cell eukaryotes with little oxygen chemistry. Thus they show loss of particular enzymes much as did higher eukaryotes when they became dependent on lower organisms for synthesis of many coenzymes and so require vitamins. We must ask how these developments of genes occur during evolution and at particular times such as the gain of copper enzymes with the rise in oxygen and copper and the losses of some of these enzymes with symbiosis.
Why are the oxidases of copper, cytochrome P-450 or the Fe(II)OG types all so greatly multiplied in plants relative to the numbers in animals, Table 4? These oxidases have a protective value as well as one in synthesis. It is very likely that the seed of a plant as it forms will be more exposed to adverse chemicals than the highly protected reproduction modes of animals. In particular the plants produce the oxygen used in these enzymes and accidentally produce both superoxide and hydrogen provide as well as the adventitious erroneous oxidation of the organic substrates and products of them. Oxygen and these products in cells as well as the copper and other metal ions are hazards, particularly to the enzymes themselves in plants. This gives a reason for the generally higher numbers of oxidases in plants together with those of homeostatic and chaperone proteins since the supply of copper must be kept from damaging the cytoplasm. The free copper is reduced to 10−15 in the cytoplasm of all cells while zinc is held at 10−10 M. The possible explanation of the duplication of oxygenases is that oxygen itself is the cause by two means. It can damage DNA directly but this does not explain selectivity of duplication or it can stress the production of proteins by damaging them. The most likely proteins to be damaged are those which use oxygen and when damage occurs production of them must increase. The stress affects the DNA in that increased production of a protein requires greater local exposure of its coded DNA and can lead to mismatching of DNA strands at this site during reproduction. Mismatching is a know cause of duplication. Different stresses of many kinds can affect proteins of the external matrices and also messenger systems associated with production of messengers and hormones, through damage to their copper and iron oxidases, and of messenger receptors, zinc fingers.
The most obvious simple stress due to oxygen is that of increase of both copper and zinc in the environment requiring multiplication of homeostatic and chaperone proteins. Any such possible sensitivity to stress has to be tested experimentally as a possible explanation of the particular multiplication and appearance of these useful products of oxygen generation which are also causes of stress. Is stress a major cause of evolution in the sequence
novel chemicals (from oxygen) → stress → multiplication of protective proteins with mutation of the proteins (which bind or are affected by the stress) → further multiplication followed by further mutation to give novel organisms?
Acknowledgements
We wish to thank Dr R. E. M. Rickaby and Dr L. Dupont for many valuable exchanges of views.References
- L. Decaria, I. Bertini and R. J. P. Williams, Metallomics, 2010, 2, 706–709 RSC.
- R. J. P. Williams and J. J. R. Fraùsto da Silva, The Chemistry of Evolution, Elsevier, Chichester, 2006 Search PubMed.
- R. J. P. Williams and R. E. M. Rickaby, submitted for publication.
- J. J. R. Fraùsto da Silva and R. J. P. Williams, The Biological Chemistry of the Elements, Oxford University Press, Oxford, 2nd edn, 2001 Search PubMed.
- C. L. Dupont, A. Butcher, R. E. Valas, P. E. Brown and G. Caetano-Anolles, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 10567–10572 CrossRef CAS.
- Y. Zhang and V. N. Gladychev, Chem. Rev., 2009, 109, 4828–4861 CrossRef CAS.
- C. Andreini, Lbunci, I. Bertini and A. Rosato, J. Proteome Res., 2008, 7, 209–216 CrossRef CAS.
- A. D. Anbar and A. H. Knoll, Science, 2002, 297, 1137–1142 CrossRef CAS.
- M. A. Saito, D. M. Sigman and F. M. M. Morel, Inorg. Chim. Acta, 2003, 356, 308–318 CrossRef CAS.
- D. Magnani and M. Sohoz, in Bacterial Transition Metal Homeostasis, ed. D. H. Nies and S. Silver, Springer, Heidelberg, 2007, pp. 259–285 Search PubMed.
- S. Ohno, Evolution by Gene Duplication, Springer, Heidelberg, 1970 Search PubMed.
- E. V. Kooning, Nucleic Acids Res., 2009, 37, 1011–1034.
- M. H. Serves, A. R. Kerr, T. J. McCormack and M. Riley, Biol. Direct, 2009, 4, 46–54 Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: Additional data—the 44 retrieved Cu-binding domains with the eventual Metal Binding Pattern. See DOI: 10.1039/c0mt00045k |
| This journal is © The Royal Society of Chemistry 2011 |
