2′-Substituted 2-amino-3-methylpyridine ribonucleosides in triplex-forming oligonucleotides: triplex stability is determined by chemical environment†
Chenguang
Lou
a,
Qiang
Xiao
a,
Radha R.
Tailor
a,
Nouha
Ben Gaied
a,
Nittaya
Gale
b,
Mark E.
Light
a,
Keith R.
Fox
c and
Tom
Brown
*a
aSchool of Chemistry, University of Southampton, Highfield, Southampton, SO17 1BJ, UK. E-mail: tb2@soton.ac.uk; Fax: +44 (0)2380 592991; Tel: +44 (0)2380 592974
bATDBio Ltd, School of Chemistry, University of Southampton, Highfield, Southampton, SO17 1BJ, UK
cSchool of Biological Sciences, University of Southampton, Highfield, Southampton, SO17 1BJ, UK
First published on 20th April 2011
Abstract
A new synthetic route to the phosphoramidite monomer of 2-amino-3-methyl-5-(2′-O-methyl-β-D-ribofuranosyl)pyridine (Me-MAP) and its 2′-O-methoxyethyl analogue (MOE-MAP) has been established using D-ribose and 2-amino-3-methyl-5-bromopyridine as precursors. Ultraviolet melting and DNase I footprinting studies indicate that the triplex stabilizing properties of 2′-modified MAPs are determined by the conformation of the entire oligonucleotide backbone. Me-MAP confers a higher triplex stability than 2′-deoxycytidine whereas triplex stabilization by MOE-MAP is similar to that of dC. Incorporation of Me-MAP or MOE-MAP into oligonucleotides renders them dramatically more resistant to degradation by serum nucleases than incorporation of 2-amino-3-methyl-5-(2′-deoxy-β-D-ribofuranosyl)pyridine (dMAP) or dC.
Introduction
Triplex-forming oligonucleotides (TFOs)1 have been used for the sequence-specific recognition of double helical DNA in selective gene-knockout and mutation correction experiments. Thus, optimization of the binding of TFOs to their target duplexes is of great importance.2–7 TFOs bind to DNA duplexes in the major groove adopting either a parallel (Y-R.Y) or an antiparallel (R-Y.R) orientation. In parallel triple helices the third strand binds to the purine-rich strand of the duplex by formation of C+-G.C and T-A.T triplets, each with two inter-base H-bonds. Low pH is required to protonate N3 of cytosine (pKa ≈ 4.3), to provide the second H-bond to guanine.8,9 Unfortunately, the stability of such triplexes at neutral pH is low as the second H-bond does not form. Therefore modified nucleosides have been incorporated into TFOs to improve the stability of the C+-G.C triplet and overcome the limitation caused by low triplex stability at elevated pH. Three types of chemical modifications have been introduced to achieve this aim: (1) positively charged groups on the ribose sugar moiety (e.g., 2′-aminoethoxy) that can interact with the negative charge on the phosphate backbone of the double helix;10,11 (2) analogues of cytosine that display an extended hydrogen bonding pattern;12–17 (3) analogues of cytosine that are protonated at physiological pH.18–20 Following this last strategy it has been shown that 2-amino-5-(2′-deoxy-β-D-ribofuranosyl)pyridine (dAP) and its 3-methyl derivative (dMAP)18–20 stabilize triplexes across a wide pH range. However, these analogues contain deoxyribose sugars which are not resistant to enzymatic degradation by nucleases in vivo. Studies have shown that 2′-ribose substitution with alkoxy groups considerably reduces oligonucleotide degradation21,22 and this would appear to be a simple solution to reverse the in vivo instability of TFOs containing the above C-nucleosides. However, it has been reported that TFOs containing 2-amino-3-methyl-5-(2′-O-methyl-β-D-ribofuranosyl)pyridine (Me-MAP) do not form thermodynamically stable triplexes.19 This is in contrast to TFOs that contain 2′-modified N-nucleosides, which have been shown to form very stable triple helices.9,23,24In this study we have reinvestigated the properties of triplexes formed by TFOs containing Me-MAP and MOE-MAP and we also describe a new high yielding synthetic route to the Me-MAP and MOE-MAP phosphoramidites. These monomers have been incorporated into TFOs in different base stacking and backbone environments to explore the influence of sequence context on triplex stability. The stability of these triplexes has been compared to those containing dMAP, 5-MedC and dC, and the nuclease resistance of the TFOs has been evaluated. The ability of the various TFOs to bind to complementary single-stranded DNA and form conventional antiparallel duplexes has also been investigated, to determine their selectivity for double-stranded relative to single stranded DNA. The nucleobase S has previously been used to recognize TA and CG inversions25 and we here describe the synthesis of its 2′-methoxyethyl derivative, which was incorporated in the TFO sequences to recognize both TA and CG inversions and is expected to render the TFOs resistant to enzymatic degradation. In addition, a synthetic route to the N2-bis-Fmoc-protected dMAP phosphoramidite monomer is described. The Fmoc protecting group permits dMAP to be incorporated into oligonucleotides using standard cycles and allows the oligonucleotides to be deprotected under mild conditions.
Results and discussion
Synthesis of 2′-O-modified MAP phosphoramidite monomers
In previous studies we were unable to fully deprotect oligonucleotides containing MAP when protected at N2 with phenoxyacetyl, even after extended heating in aqueous ammonia.19 Therefore in this study we opted for the more labile trifluoroacetyl protecting group.The synthesis of the TFA-protected Me-MAP and MOE-MAP phosphoramidites is shown in Scheme 1. The exocyclic amino group of 2-amino-3-methyl-5-bromopyridine was protected by reaction with 4-methoxybenzyl chloride to generate 2 in 61% yield, ready for reaction with the sugar moiety. To synthesize the required 2′-O-modified ribose moieties, the 2′-OH group of compound 326 was reacted with iodomethane or 2-bromoethyl methyl ether in the presence of NaH in DMF to give compounds 4a and 4b respectively. The synthesis of the two analogues was very similar from this point. Compound 4a or 4b was hydrolyzed to 5a/b using 80% acetic acid and a catalytic amount of conc. sulfuric acid, then oxidized to the corresponding lactone 6a/b using tetrapropylammonium perruthenate, and 4-methylmorpholine-N-oxide. The fully protected brominated methylaminopyridine 2 was coupled to sugar 6a/b using n-BuLi to give a modified C-nucleoside 7a/b as a mixture of α and β anomers (α![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) β, 1
β, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). The sugar ring was recyclized using boron trifluoride diethyletherate to afford 8a/b exclusively in the β-configuration. At this step the methoxybenzyl groups were cleaved in trifluoroacetic acid to generate free amine 9a/b. Benzyl groups were removed by hydrogenation using palladium hydroxide on activated carbon to yield compounds 10a/b. The X-ray crystal structure of 10a confirmed the β-configuration at the anomeric centre (Fig. 1B). Compounds 10a/b were selectively protected at the 5′-position by reaction with 4,4′-dimethoxytrityl chloride (DMTrCl) in pyridine to yield 11a/b, and the free exocyclic amino group (N4) was protected using trifluoroacetic anhydride to afford 12a/b. Finally, phosphitylation using 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite gave the desired monomers 13a and 13b. The overall yields from compound 3 were 10% for Me-MAP and 11% for MOE-MAP.
1). The sugar ring was recyclized using boron trifluoride diethyletherate to afford 8a/b exclusively in the β-configuration. At this step the methoxybenzyl groups were cleaved in trifluoroacetic acid to generate free amine 9a/b. Benzyl groups were removed by hydrogenation using palladium hydroxide on activated carbon to yield compounds 10a/b. The X-ray crystal structure of 10a confirmed the β-configuration at the anomeric centre (Fig. 1B). Compounds 10a/b were selectively protected at the 5′-position by reaction with 4,4′-dimethoxytrityl chloride (DMTrCl) in pyridine to yield 11a/b, and the free exocyclic amino group (N4) was protected using trifluoroacetic anhydride to afford 12a/b. Finally, phosphitylation using 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite gave the desired monomers 13a and 13b. The overall yields from compound 3 were 10% for Me-MAP and 11% for MOE-MAP.
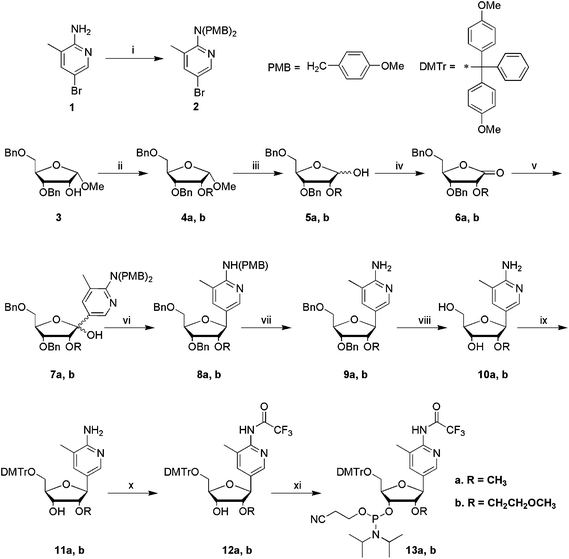 | ||
Scheme 1
Reagents and conditions: (i) 4-methoxybenzyl chloride, NaH, DMF, rt, 8 h, 61%; (ii) CH3I or BrCH2CH2OCH3, NaH, DMF, 0 °C, 8 h, 4a 73%, 4b 94%; (iii) 80% AcOH, 1% conc. H2SO4, 80 °C, 1 h, 5a 77%, 5b 94%; (iv) 4-methylmorpholine-N-oxide, tetrapropylammonium perruthenate, DCM, rt, 8 h, 6a 87%, 6b 87%; (v) 2, n-BuLi, THF, −78 °C, 4 h, 7a 69%, 7b 65%; (vi) triethylsilane, boron trifluoride diethyletherate, DCM, −78 °C, 4 h, rt, 16 h, 8a 91%, 8b n.a.; (vii) CF3COOH, rt, 5 h, 9a 89%, 9b 98% for two steps; (viii) 10aPd(OH)2 (20% on carbon), H2, anhydrous EtOH, 65 °C, 12 h, 75%; 10bPd(OH)2 (20% on carbon), HCOOH/MeOH (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1), 50 °C, overnight, 56%; (ix) DMTrCl, pyridine, rt, 3 h, 11a 98%, 11b 60%; (x) (CF3CO)2O, DIPEA, DCM, 0 °C, >2 h, 12a 90%, 12b 74%; (xi) 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite, DIPEA, DCM, rt, 3 h, 13a 70%, 13b 92%. 1), 50 °C, overnight, 56%; (ix) DMTrCl, pyridine, rt, 3 h, 11a 98%, 11b 60%; (x) (CF3CO)2O, DIPEA, DCM, 0 °C, >2 h, 12a 90%, 12b 74%; (xi) 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite, DIPEA, DCM, rt, 3 h, 13a 70%, 13b 92%. | ||
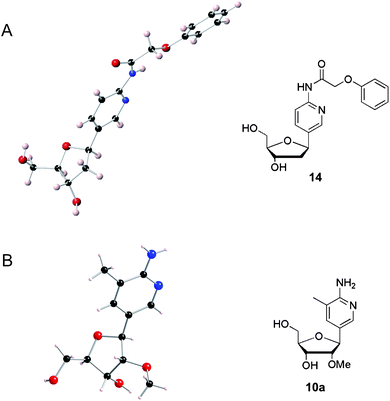 | ||
| Fig. 1 X-Ray crystal structures (left) and the corresponding chemical structures (right) of (A) N-(phenoxyacetyl) dAP and (B) Me-MAP. | ||
The X-ray crystal structures of phenoxyacetyl dAP27 (Fig. 1A) and Me-MAP (Fig. 1B) were determined and compared. The 2′-methoxy group on Me-MAP induces the S-type sugar pucker (2′-endo), which may exert a negative influence on the Hoogsteen interaction required in the Me-MAP.GC triplet, and destabilize triple helices that contain this derivative. In addition, the orientation of 3′-OH and 5′-OH groups on the sugar is affected by the 2′-methoxy modification. The 5′-OH of Me-MAP points upwards whereas that of dAP points downwards, and the 3′-OH of Me-MAP is tucked inside and below the furanose sugar to a greater extent than that of dAP. The relative disposition of the 3′-OH and 5′-OH influences the conformation of TFOs, so these observations are consistent with the difference in triplex stabilization reported for TFOs containing the ribo-C-nucleoside (Me-MAP) and the deoxyribo-C-nucleosides (dAP and dMAP).19,20
Synthesis of dMAP monomer and 2′-methoxyethyl S monomer and their incorporation into triplex-forming oligonucleotides
Standard solid-phase phosphoramidite methods were used to synthesize all oligonucleotides except those containing Me-MAP and MOE-MAP, where the acetic anhydride capping step was omitted. This was done to prevent replacement of the trifluoroacetyl protection on the 2-amino group of aminopyridine by acetyl. We have observed that this side-reaction occurs at a high level when using standard synthesis cycles. The resultant 2-acetyl MAP is very resistant to deprotection and the TFOs do not bind to their target duplexes (results not shown). Trifluoroacetyl protection at N2 of MAP has the advantage that it can be completely removed under mild conditions, e.g. concentrated aqueous ammonia at room temperature for 10 h. It is therefore fully compatible with sensitive modifications commonly used in TFOs such as the duplex crosslinker psoralen.28,29 During our studies on protection of N2 of MAP we also evaluated the Fmoc group which proved to be suitable in terms of ease of deprotection. It was removed by treatment of the synthesis resin with 20% diethylamine in acetonitrile for 20 min followed by 10 h at room temperature in concentrated aqueous ammonia. It is also compatible with standard DNA synthesis cycles including capping with acetic anhydride.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) α, 3
α, 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2). 17 β was isolated, and cleavage of the paramethoxybenzyl protecting groups was achieved using trifluoroacetic acid, to afford compound 18. This was subsequently protected at the exocyclic amino moiety using excess fluorenylmethyloxycarbonyl chloride to afford the bis-Fmoc protected compound 19. Debenzylation at the 3′ and 5′ positions of the sugar ring in compound 19 was carried out to afford compound 20, which was then reacted with DMTrCl to afford the precursor 21. Compound 21 was finally phosphitylated to give the desired monomer 22. The overall yield for the synthesis of dMAP, from the starting material 15, was 26%. The equivalent phosphoramidite monomer with a single Fmoc protecting group was also synthesized, but in contrast to compound 22 it did not give clean results in oligonucleotide synthesis.
2). 17 β was isolated, and cleavage of the paramethoxybenzyl protecting groups was achieved using trifluoroacetic acid, to afford compound 18. This was subsequently protected at the exocyclic amino moiety using excess fluorenylmethyloxycarbonyl chloride to afford the bis-Fmoc protected compound 19. Debenzylation at the 3′ and 5′ positions of the sugar ring in compound 19 was carried out to afford compound 20, which was then reacted with DMTrCl to afford the precursor 21. Compound 21 was finally phosphitylated to give the desired monomer 22. The overall yield for the synthesis of dMAP, from the starting material 15, was 26%. The equivalent phosphoramidite monomer with a single Fmoc protecting group was also synthesized, but in contrast to compound 22 it did not give clean results in oligonucleotide synthesis.
![Reagents and conditions: (i) 2-[N,N-bis(4-methoxybenzyl)]amino-3-methyl-5-bromopyridine (2), n-BuLi, THF, −35 °C, 0.5 h; −30 °C, 1 h; rt, 4.5 h, 92%; (ii) Bu3P, TMAD, THF, rt, 6 h, 88%; (iii) CF3CO2H/DCM, rt, 7 h, 99%; (iv) Fmoc-Cl (in CH3CN), pyridine, 0 °C, rt, 2 h, 96%; (v) BCl3, DCM, −78 °C, 6 h, methanol, −75 °C, 0.5 h; 4 °C, 17.5 h, 79%; (vi) DMTrCl, pyridine, rt, 3.5 h, 87%; (vii) 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite, DIPEA, DCM, rt, 2 h, 50%.](/image/article/2011/MD/c1md00068c/c1md00068c-s2.gif) | ||
| Scheme 2 Reagents and conditions: (i) 2-[N,N-bis(4-methoxybenzyl)]amino-3-methyl-5-bromopyridine (2), n-BuLi, THF, −35 °C, 0.5 h; −30 °C, 1 h; rt, 4.5 h, 92%; (ii) Bu3P, TMAD, THF, rt, 6 h, 88%; (iii) CF3CO2H/DCM, rt, 7 h, 99%; (iv) Fmoc-Cl (in CH3CN), pyridine, 0 °C, rt, 2 h, 96%; (v) BCl3, DCM, −78 °C, 6 h, methanol, −75 °C, 0.5 h; 4 °C, 17.5 h, 79%; (vi) DMTrCl, pyridine, rt, 3.5 h, 87%; (vii) 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite, DIPEA, DCM, rt, 2 h, 50%. | ||
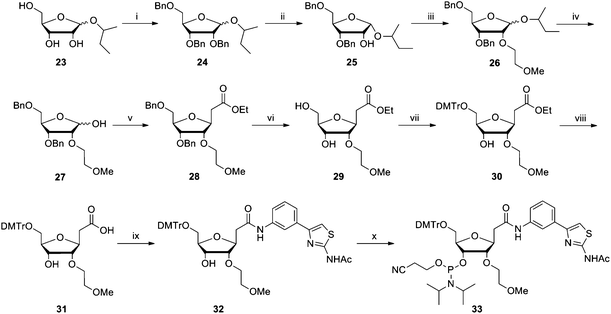 | ||
| Scheme 3 Reagents and conditions: (i) benzyl bromide, NaH, DMF, −10 °C to rt, overnight, 61%; (ii) SnCl4, DCM, rt, 1 h, 78%; (iii) Br(CH2)2OMe, NaH, DMF, −10 °C to rt, 2.5 h, 94%; (iv) CF3COOH, H2O, DCM, −10 °C to rt, 4 h, 90%; (v) PPh3CHCO2Et, THF, reflux, 3 h, then EtONa, EtOH, rt, 3 h, 44%; (vi) Pd(OH)2 (10% on carbon), H2, MeOH, rt, 20 h, 97%; (vii) DMTrCl, DMAP, pyridine, rt, 4 h, 76%; (viii) NaOH (1 M), THF, reflux, 4 h, 76%; (ix) 2-acetamido-4-(3-aminophenyl) thiazole, EDC·HCl, DMF, rt, 4 h, 36%; (x) 2-cyanoethyl-N,N-diisopropylchlorophosphoramidite, DIPEA, DCM, rt, 2 h, 58%. | ||
UV triplex melting studies
Two sets of TFOs and their corresponding hairpin DNA duplexes were synthesized. Hairpin duplexes have higher melting temperatures (Tm) than intermolecular duplexes and this puts them out of the normal range of UV melting, facilitating clear observation of triplex Tms. The TFOs for duplex 2 can be divided into two groups depending on the nature of the sugar moieties in the sequences (excluding the dC and C-nucleosides), TFOs 6–10 contain both deoxyribose and ribose sugars, whereas TFOs 11–15 contain only substituted ribose sugars. TFOs 1–5, which are designed to target duplex 1, all have mixed backbones. The dabcyl group at 5′-end on TFOs 1–5 was added to provide the added possibility of fluorescence melting if the UV meting studies proved to be inconclusive. The S34 and methoxyethoxy S-monomers (![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ) were used to recognize CG and TA inversions respectively in the target duplexes (Fig. 2). Each of these TFOs contains 2′-aminoethoxy T residues (t) as this modification enhances the stability of the T.AT triplet.33
) were used to recognize CG and TA inversions respectively in the target duplexes (Fig. 2). Each of these TFOs contains 2′-aminoethoxy T residues (t) as this modification enhances the stability of the T.AT triplet.33
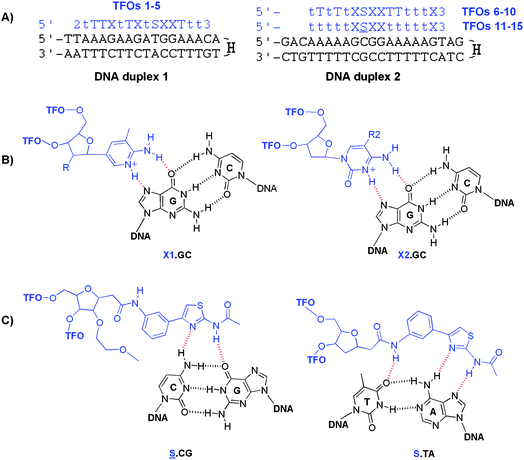 | ||
Fig. 2 Triplexes investigated in this study and triplets formed by modified bases. (A) The sequence of duplexes 1 and 2 (in black) with their TFOs (in blue). H = hexaethylene glycol linker, 2 = dabcyl, 3 = propanol, T = thymidine, t = 2′ aminoethoxy T.33 (B) Proposed structures of triplets. X1.GC triplexes where X1 = MAP analogues: MAP, R = H; Me-MAP, R = OMe; MOE-MAP, R = OEtOMe (left) and X2.GC triplexes where X2 = cytosine bases: 5-MedC, R2 = Me; dC, R2 = H (right). (C) ![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) .CG and S.TA triplexes. .CG and S.TA triplexes. | ||
Melting studies (Table 1, and Fig. 2 and 3) show that for the mixed backbone TFOs the order of stability between pH 5.5 and 7.5 is dMAP > 5-MedC > dC > Me-MAP > MOE-MAP with both duplexes 1 and 2, in agreement with the previous work19 though the rank order of dMAP and 5MedC is reversed at low pH. However, the order is different for the TFOs that contain a fully ribo-backbone (TFOs 11–15): dMAP > 5-MedC > dC > Me-MAP > MOE-MAP at low pH and dMAP > 5-MedC > Me-MAP > MOE-MAP > dC at high pH. Most notable is the observation that Me-MAP (TFO 11) formed a more stable triplex at pH 7 than the equivalent TFO containing dC (TFO 13). This observation has implications for TFO design and the best use of modified nucleotides. In all cases Me-MAP produced more stable triplexes than MOE-MAP.
| TFOs ID | Modified base (X) | pH 5.5* | pH 5.5 | pH 6.2 | pH 6.6 | pH 7.0 | pH 7.5 | pH 8.0 |
|---|---|---|---|---|---|---|---|---|
a
T
m values are an average of two melting experiments. The experiments were performed in 10 mM CH3CO2Na/CH3CO2H buffer (pH 5.5), containing 200 mM NaCl or in 10 mM NaH2PO4/Na2HPO4 buffer (pH 6.2, 6.6, 7.0, 7.5 and 8.0), containing 200 mM NaCl. The concentration of TFOs: target DNA = 3.0 µM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.0 µM. * = containing 2 mM spermine; > = higher than 75 °C; — = lower than 20 °C. Melting curves and derivatives shown below and in the ESI1. 1.0 µM. * = containing 2 mM spermine; > = higher than 75 °C; — = lower than 20 °C. Melting curves and derivatives shown below and in the ESI1.
|
||||||||
| Mixed backbone TFOs targeting duplex 1 | ||||||||
| TFO 1 | Me-MAP | 47.0 | 24.6 | — | — | — | — | — |
| TFO 2 | MOE-MAP | 41.5 | — | — | — | — | — | — |
| TFO 3 | dC | 65.4 | 50.6 | 39.6 | 31.1 | 22.3 | — | — |
| TFO 4 | 5-MedC | > | 63.1 | 52.2 | 43.5 | 35.9 | 27.0 | — |
| TFO 5 | dMAP | > | 56.6 | 50.1 | 46.5 | 41.3 | 33.2 | 26.2 |
| Mixed backbone TFOs targeting duplex 2 | ||||||||
| TFO 6 | Me-MAP | 48.9 | 29.8 | — | — | — | — | — |
| TFO 7 | MOE-MAP | 36.9 | — | — | — | — | — | — |
| TFO 8 | dC | 66.7 | 54.4 | 39.3 | 30.6 | 22.4 | — | — |
| TFO 9 | 5-MedC | > | 61.5 | 47.8 | 39.1 | 30.8 | 21.3 | — |
| TFO 10 | dMAP | > | 57.3 | 56.0 | 49.4 | 44.3 | 36.0 | 26.9 |
| Ribo-backbone TFOs targeting duplex 2 | ||||||||
| TFO 11 | Me-MAP | > | 67.3 | 61.3 | 55.5 | 49.2 | 40.3 | 30.2 |
| TFO 12 | MOE-MAP | > | 60.5 | 55.4 | 50.5 | 44.5 | 35.4 | 26.4 |
| TFO 13 | dC | > | > | 63.5 | 55.2 | 46.4 | 35.5 | 24.5 |
| TFO 14 | 5-MedC | > | > | 69.7 | 61.1 | 54.0 | 44.5 | 35.9 |
| TFO 15 | dMAP | > | > | > | 68.3 | 60.5 | 53.1 | 44.7 |
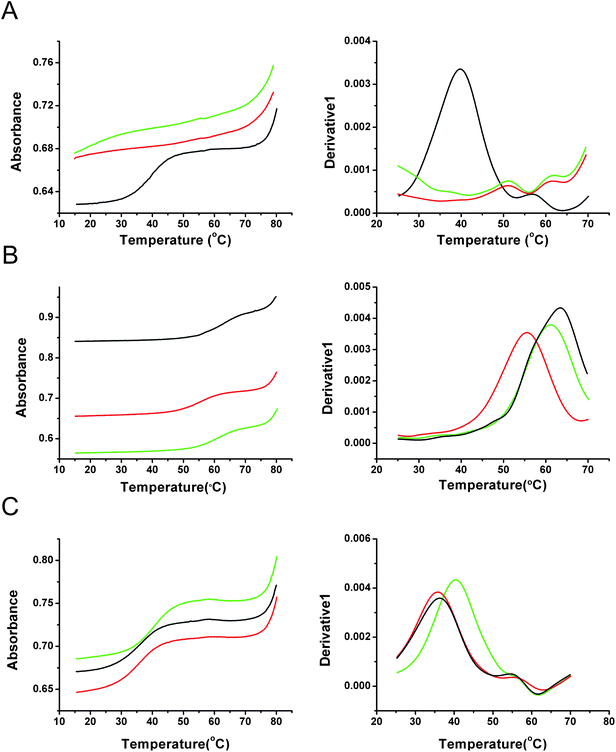 | ||
Fig. 3
UV melting curves (left) and derivatives (right) of Me-MAP and MOE-MAP TFOs with target hairpin duplex 2. (A) TFO 6 (green), TFO 7 (red) and TFO 8 (black) at pH 6.2. (B) TFO 11 (green), TFO 12 (red) and TFO 13 (black) at pH 6.2. (C) TFO 11 (green), TFO 12 (red) and TFO 13 (black) at pH 7.5. Experiments were performed in 10 mM NaH2PO4/Na2HPO4 buffer containing 200 mM NaCl pH 6.2 or 7.5. The ratio of TFO/target duplex was 3.0 µM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.0 µM. 1.0 µM. | ||
Comparing TFOs 6–10 with TFOs 11–15 shows that, as expected, replacing the four thymines with 2′-aminoethoxy T produces a large increase in the triplex stability, with an average increase in Tm of 5.3 °C per addition at pH 7.0. However this change also affected the rank order of the different modifications. In the ribo-backbone TFOs at pH 7, dMAP gave an increase in Tm of 3.5 °C/addition relative to dC and 1.6 °C/addition relative to 5-MedC. The same trends were observed in mixed-backbone TFOs targeting duplex 1 (4.8 °C/addition and 1.3 °C/addition compared with dC and 5-MedC respectively) and duplex 2 (5.5 °C/addition and 3.4 °C/addition). The mixed-backbone TFOs containing Me-MAP and MOE-MAP produced less stable triplexes than dC at all pH values.
The enhanced triplex stabilizing effect of Me-MAP in the all ribo-backbone TFOs relative to their mixed backbone counterparts can be clearly seen from these results. This indicates that the stabilizing properties of 2′-modified MAP are affected by the conformational properties of the entire TFO and the sequence context in which it is located. This suggests that when all the sugars are 2′-modified, the conformation of the TFO allows efficient interactions with the target duplex. In contrast, the presence of 2′-deoxyribose sugars in the mixed backbone TFOs disrupts this stabilizing conformation and disfavors triplex formation. This observation enables us to predict that judiciously designed 2′-modified MAP analogues might stabilize triplexes more effectively than dMAP, provided that they are surrounded by 2′-modified ribonucleotides.
Affinity of ribo-backbone TFOs for single-stranded DNA
It is known that the insertion of dMAP and Me-MAP into oligonucleotides dramatically reduces their affinity for antiparallel complementary single stranded DNA. Such oligonucleotides are unable to form stable duplexes, as MAP can only form two Watson–Crick hydrogen bonds with guanine; one between N1 of dMAP and N1 of G, and the other between NH2 of dMAP and O6 of G.19 Moreover, the first of these cannot exist in the protonated form of MAP. We therefore carried out a study to determine whether this duplex instability is also observed for the equivalent ribo-backbone TFOs. To evaluate this, we studied the binding of TFOs 11–15 to complementary antiparallel Watson–Crick single stranded DNA (where the C-nucleoside is placed opposite to G) by UV melting at pH 7.0 (Table 2).|
|
|||
|---|---|---|---|
| TFO | Nucleotide (X) | T m/°C | ΔTm (N-dC) |
| a T m values are an average of three melting temperatures. The experiments were performed in 10 mM NaH2PO4/Na2HPO4 buffer pH 7.0 containing 200 mM NaCl: single stranded DNA/TFO = 1.05 µM/1.00 µM. t = 2′-aminoethoxy T, S = S-monomer. | |||
| TFO 11 | Me-MAP | 24.9 | −18.1 |
| TFO 12 | MOE-MAP | 24.6 | −18.4 |
| TFO 13 | dC | 43.0 | — |
| TFO 14 | 5-MedC | 46.3 | +3.3 |
| TFO 15 | dMAP | 30.1 | −12.9 |
Table 2 shows that the replacement of four dC residues by dMAP leads to a large decrease in Tm (−3.2 °C per substitution). The destabilization with 2′-modified MAPs is even greater (−4.5 °C per substitution). These results show that the incorporation of dMAP, and more so its 2′-modified derivatives, renders oligonucleotides selective for double stranded DNA. Unlike TFOs containing dC or 5-MedC they will not bind significantly to single-stranded DNAin vivo.
DNase I footprinting
The results of these UV melting experiments were confirmed by DNase I footprinting with TFOs 11–13 (containing Me-MAP, MOE-MAP and dC respectively). DNase I footprinting experiments with these oligonucleotides at pH 8.0 are shown in Fig. 4. The DNA fragment contains two identical copies of the triplex target site (indicated by the bars), which are arranged in opposite orientations: the upper site shows the purine-containing strand of the target, while the lower site reveals the pyrimidine-containing strand. It can be seen that all three TFOs produce clear footprints; the protection with Me-MAP and dC persists to about 0.3 µM, while MOE-MAP binds less well to about 1 µM. The footprint at the upper site is accompanied by enhanced DNase I cleavage at the 3′-(lower) end of the target site at the triplex–duplex junction. Triplex-induced enhancements are often observed at this position on the labeled purine strand and show the same concentration dependence as the footprint itself. It should be noted that this enhanced band is one base higher with dC than with Me-MAP and MOE-MAP, suggesting that the 3′-terminal base may be fraying from the target with dC, but not with MeMAP or MOE-MAP.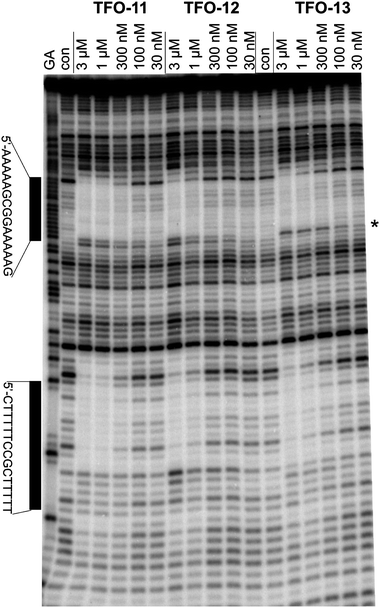 | ||
| Fig. 4 DNase I footprinting of TFOs 11–13. The experiment was performed in 10 mM Tris–HCl pH 8.0 containing 50 mM NaCl. TFO concentrations are shown at the top of each gel lane. Tracks labeled “Con” are control lanes in the absence of added TFO. The track labeled ‘GA’ is a marker specific for purines. The locations of the two triplex target sites are indicated by the bars. The position of the enhance cleavage on the 3′-(lower) end of the upper footprint is indicated by an asterisk. | ||
Resistance to nuclease degradation
The incorporation of 2′-modified N-nucleosides into TFOs renders them much more resistant to nuclease mediated degradation than the corresponding 2′-deoxyribo-N-nucleosides.21,22 In this study we determined whether the incorporation of 2′-modified MAPs (i.e. C-nucleosides) might confer similar resistance, and whether such TFOs are superior in this respect to TFOs containing dMAP.The oligonucleotides used in this degradation study (Table 3) are pyrimidine-rich and their sequences were based on previous work with C-nucleosides.18
| TFO | Sequence (5′ to 3′) |
|---|---|
| TFO 16 | TxTTxTTTTTTxTTxTTxxT |
| TFO 17 | TyTTyTTTTTTyTTyTTyyT |
| TFO 18 | TMTTMTTTTTTMTTMTTMMT |
| TFO 19 | TCTTCTTTTTTCTTCTTCCT |
TFOs 16–19 were incubated in a medium containing fetal bovine serum for various periods from 1–24 h and their stability was determined by denature PAGE analysis (Fig. 5). Interestingly, both TFO 16 and TFO 17 migrated slightly slower than TFO 18 and TFO 19. This may be because the resultant oligonucleotides became more bulky when 2′-modified MAPs were incorporated than their 2′-deoxy controls.
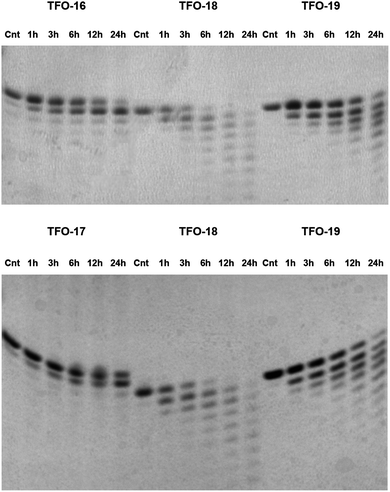 | ||
| Fig. 5 20% PAGE denaturing gels showing the time course of degradation following incubation of TFOs 16–19 in serum at pH 7.0, 37 °C. Samples were incubated for 1 h, 3 h, 6 h, 12 h and 24 h. The negative control (Cnt) was incubated at 37 °C for 24 h in the absence of serum. With TFO 18 and TFO 19 as controls, the experiments were performed for TFO 16 and TFO 17 respectively. | ||
The 2′-deoxy TFO 19 (dC) and TFO 18 (dMAP) were rapidly degraded with the appearance of shorter length bands after only 1 h incubation. The dMAP-containing TFO degraded at a greater rate than the C-containing TFO, possibly because the latter is more structured. In contrast, Me-MAP (TFO 16) and MOE-MAP (TFO 17) show much greater resistance to the nucleases present in the medium. TFO 17 was more resistant than TFO 16, probably due to the increased steric hindrance of the 2′-methoxyethoxy group relative to the 2′-methoxy group which might inhibit both enzyme binding and phosphodiester cleavage. After 6 h incubation in serum, the full-length TFO 17 was still the major component and only half of TFO 16 was degraded. For TFO 16 and TFO 17 the degradation product is seen as a band just below the intact oligonucleotide and is the 19 mer in which the 3′-nucleotide (unprotected dT residue) has been cleaved, leaving the 2′-modified MAP at the 3′-terminus. There was still a considerable amount of intact TFO 16 and TFO 17 after 24 h incubation and this shows that degradation is effectively blocked after the removal of the terminal dT residue. To summarise, as previously shown for N-nucleosides,21,22 the presence of a substituent at the 2′-oxygen of the ribose sugar confers dMAP C-nucleoside analogues with considerable resistance towards nuclease degradation. In contrast, C-nucleosides containing deoxyribose sugars are rapidly digested.
Conclusions
Me-MAP and MOE-MAP phosphoramidite monomers have been synthesized from 1-O-methyl-3,5-di-O-benzyl-α-D-ribofuranoside and 2-amino-5-bromo-3-methylpyridine. Ultraviolet melting studies and DNase I footprinting on triplexes assembled from TFOs containing these C-nucleosides indicate that the triplex-stabilizing efficacy of 2′-modified MAPs depends on the chemical composition of the entire TFO, i.e. the presence of 2′-modified ribose sugars throughout the TFO produces more stable triplexes. Me-MAP yields higher triplex stability than dC for ribo-backbone TFOs, while triplex stabilization by MOE-MAP TFOs is very similar to that of dC. The insertion of Me-MAP or MOE-MAP into oligonucleotides inhibits duplex formation with antiparallel Watson–Crick complementary strands at pH 7.0, in contrast to corresponding oligonucleotides containing dC and 5-MedC. This is important as it indicates that these C-nucleosides will have much greater selectivity for their duplex target in vivo. The incorporation of Me-MAP or MOE-MAP into TFOs renders them dramatically more resistant to degradation by serum nucleases than dMAP and dC. This is another essential consideration for in vivo applications. From these studies we predict that 2′-aminoethoxy-MAP will be a potent triplex stabilizing nucleoside, which should also make TFOs resistant to enzymatic degradation.Abbreviations
| Me-MAP | 2-amino-3-methyl-5-(2′-O-methyl-β-D-ribofuranosyl)pyridine |
| MOE-MAP | 2-amino-3-methyl-5-(2′-O-methoxyethyl-β-D-ribofuranosyl)pyridine |
| dMAP | 2-amino-3-methyl-5-(2′-deoxy-β-D-ribofuranosyl)pyridine |
| dAP | 2-amino-5-(2′-deoxy-β-D-ribofuranosyl)pyridine |
| dC | 2′-deoxycytidine |
| 5-MedC | 5-methyl-2′-deoxycytidine |
| TFO | triplex-forming oligonucleotide |
| T m | melting temperature |
| FBS | fetal bovine serum |
Acknowledgements
This work was supported by a research grant from BBSRC under the SCIBS initiative. We thank ORSAS for funding a studentship to C. Lou and ATDBio for oligonucleotide synthesis.References
- G. Felsenfeld, D. R. Davies and A. Rich, J. Am. Chem. Soc., 1957, 79, 2023–2024 CrossRef CAS.
- P. P. Chan and P. M. Glazer, J. Mol. Med., 1997, 75, 267–282 CrossRef CAS.
- L. Gorman and P. M. Glazer, Curr. Mol. Med., 2001, 1, 391–399 Search PubMed.
- M. P. Knauert and P. M. Glazer, Hum. Mol. Genet., 2001, 10, 2243–2251 CrossRef CAS.
- A. Majumdar, A. Khorlin, N. Dyatkina, F. L. M. Lin, J. Powell, J. Liu, Z. Fei, Y. Khripine, K. A. Watanabe, J. George, P. M. Glazer and M. M. Seidman, Nat. Genet., 1998, 20, 212–214 CrossRef CAS.
- S. Neidle, Anti-Cancer Drug Des., 1997, 12, 433–442 CAS.
- G. Wang, M. M. Seidman and P. M. Glazer, Science, 1996, 271, 802–805 CAS.
- D. M. Gowers and K. R. Fox, Nucleic Acids Res., 1999, 27, 1569–1577 CrossRef.
- D. A. Rusling, V. J. Broughton-Head, T. Brown and K. R. Fox, Curr. Chem. Biol., 2008, 2, 1–10 Search PubMed.
- S. Buchini and C. J. Leumann, Eur. J. Org. Chem., 2006, 3152–3168 CrossRef CAS.
- N. Puri, A. Majumdar, B. Cuenoud, F. Natt, P. Martin, A. Boyd, P. S. Miller and M. M. Seidman, J. Biol. Chem., 2001, 276, 28991–28998 CrossRef CAS.
- J. Hunziker, E. S. Priestley, H. Brunar and P. B. Dervan, J. Am. Chem. Soc., 1995, 117, 2661–2662 CrossRef CAS.
- A. Ono, P. O. P. Ts'o and L. S. Kan, J. Am. Chem. Soc., 1991, 113, 4032–4033 CrossRef CAS.
- A. Ono, P. O. P. Ts'o and L. S. Kan, J. Org. Chem., 1992, 57, 3225–3230 CrossRef CAS.
- U. von Krosigk and S. A. Benner, J. Am. Chem. Soc., 1995, 117, 5361–5362 CrossRef CAS.
- G. Xiang, W. Soussou and L. W. McLaughlin, J. Am. Chem. Soc., 1994, 116, 11155–11156 CrossRef CAS.
- J. Marfurt, S. P. Parel and C. J. Leumann, Nucleic Acids Res., 1997, 25, 1875–1882 CrossRef CAS.
- P. J. Bates, C. A. Laughton, T. C. Jenkins, D. C. Capaldi, P. D. Roselt, C. B. Reese and S. Neidle, Nucleic Acids Res., 1996, 24, 4176–4184 CrossRef CAS.
- S. Hildbrand, A. Blaser, S. P. Parel and C. J. Leumann, J. Am. Chem. Soc., 1997, 119, 5499–5511 CrossRef CAS.
- S. Hildbrand and C. Leumann, Angew. Chem., Int. Ed., 1996, 35, 1968–1970 CrossRef.
- G. P. Balow, O. L. Acevedo and P. D. Cook, U.S. patent 6093807, Isis Pharmaceuticals, Search PubMedJuly 25, 2000.
- E. Uhlmann and A. Peyman, Chem. Rev., 1990, 90, 543–584 CrossRef CAS.
- C. Escude, J. S. Sun, M. Rougee, T. Garestier and C. Helene, C. R. Acad. Sci., Ser. III, 1992, 315, 521–525 CAS.
- M. Shimizu, A. Konishi, Y. Shimada, H. Inoue and E. Ohtsuka, FEBS Lett., 1992, 302, 155–158 CrossRef CAS.
- D. Guianvarc'h, R. Benhida, J. L. Fourrey, R. Maurisse and J. S. Sun, Chem. Commun., 2001, 1814–1815 RSC.
- C. Lou, Q. Xiao, L. Brennan, M. E. Light, N. Vergara-Irigaray, E. M. Atkinson, L. M. Holden-Dye, K. R. Fox and T. Brown, Bioorg. Med. Chem., 2010, 18, 6389–6397 CrossRef CAS.
- V. E. C. Powers, PhD thesis, University of Southampton, 2005.
- P. A. Havre, E. J. Gunther, F. P. Gasparro and P. M. Glazer, Proc. Natl. Acad. Sci. U. S. A., 1993, 90, 7879–7883 CrossRef CAS.
- A. Majumdar, P. A. Muniandy, J. Liu, J. I. Liu, S. T. Liu, B. Cuenoud and M. M. Seidman, J. Biol. Chem., 2008, 283, 11244–11252 CrossRef CAS.
- W. Wierenga and H. I. Skulnick, Carbohydr. Res., 1981, 90, 41–52 CrossRef CAS.
- Y. Wang, D. A. Rusling, V. E. C. Powers, O. Lack, S. D. Osborne, K. R. Fox and T. Brown, Biochemistry, 2005, 44, 5884–5892 CrossRef CAS.
- D. Guianvarc'h, R. Benhida, J.-L. Fourrey, R. Maurisse and J.-S. Sun, Chem. Commun., 2001, 1814–1815 RSC.
- B. Cuenoud, F. Casset, D. Husken, F. Natt, R. M. Wolf, K. H. Altmann, P. Martin and H. E. Moser, Angew. Chem., Int. Ed., 1998, 37, 1288–1291 CrossRef CAS.
- D. Guianvarc'h, J.-L. Fourrey, R. Maurisse, J.-S. Sun and R. Benhida, Bioorg. Med. Chem., 2003, 11, 2751–2759 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available: Synthesis of MeMAP, MOE-MAP, 2′-methoxyethoxy-S and bis-Fmoc-protected dMAP monomers, X-ray crystallography, oligonucleotide synthesis purification and analysis, ultraviolet triplex melting studies, ultraviolet duplex melting studies, DNase I footprinting, serum stability studies, mass spectrometry data for all oligonucleotides, UV-melting curves and derivatives of TFOs 1–15 with their target duplexes. CCDC reference numbers [CCDC NUMBER(S)]. For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/c1md00068c |
| This journal is © The Royal Society of Chemistry 2011 |

