Synthesis and efficient siRNA delivery of polyamine-conjugated cationic nucleoside lipids†
Sachin Prakash
Patil
,
Jeong Wu
Yi
,
Eun-Kyoung
Bang
,
Eun Mi
Jeon
and
Byeang Hyean
Kim
*
Department of Chemistry, BK School of Molecular Science, Pohang University of Science and Technology, Pohang, 790-784, Korea. E-mail: bhkim@postech.ac.kr; Fax: +82 54 279 3399; Tel: +82 54 279 2115
First published on 7th April 2011
Abstract
To advance their use as nonviral gene delivery agents, we have synthesized cationic nucleoside lipids featuring naturally occurring polyamines conjugated at the 5′ position and oleyl groups at the 2′ and 3′ positions of uridine, through chemically stable, but biodegradable, carbamate linkers. The corresponding lipoplexes are efficient and nontoxic vectors for the delivery of siRNA into cells in vitro.
With the sequencing of the human genome, gene therapy now has enormous potential for use in the treatment of hereditary diseases and cancers.1 Synthetic small interfering RNAs (siRNAs) have emerged as important tools for the specific reduction, or silencing, of gene expression through RNA interference (RNAi), allowing particular cellular defects to be addressed.2 The use of an efficient vector for gene delivery through a cell membrane has, however, been a key step in successful gene therapy. Viral and nonviral vectors have been developed to overcome the low delivery of siRNAs through cell membranes. Although viral vectors are efficient, there is interest in developing nonviral delivery methods because of several potential risks when using viral carriers.3 Among the nonviral vectors, cationic lipids have been studied extensively ever since Felgner and co-workers reported their first results over 20 years ago.4 Cationic lipids feature three essential domains: a positive charged polar head group (often derived from an amine or polyamine), a linker (mostly possessing ether, ester, or carbamate structure), and a lipophilic tail (typically derived from cholesterol or a long-chain fatty acid).
Nucleoside-based lipids (nucleolipids) have attracted significant attention for their applications in the design of artificial molecular devices and novel therapeutic molecules.5 Nucleolipids interact with genes through hydrogen bonding, π–π stacking, and nucleobase recognition, as well as through electrostatic and hydrophobic interactions. To improve the efficiency of siRNA delivery, cationic nucleoside lipids (CNLs) have been studied extensively because they possess lower toxicity relative to the commercial vector Lipofectamine 2000.6,7 In nature, DNA is condensed through the combined actions of positively charged polyamines (putrescine, spermidine, spermine), histone, and polybasic proteins. The presence of these polyamines is essential for the growth and survival of all cells. They are widely distributed in nature and display a variety of important biological activities. Polyamines were first introduced into liposomes by Behr et al., and polyamine-mediated lipids have been synthesized as nonviral vectors for gene therapy.8
In this paper, we describe the synthesis of uridine derivatives conjugated to polyamines and oleyl groups through carbamate bonds. Carbamate-linked lipids exhibiting low toxicity have been developed previously as novel cationic lipids.9 In earlier studies, we modified nucleic acids (nucleobases, nucleosides, oligonucleotides) for potential use in biomedical applications (as drugs, drug delivery agents, and DNA/RNA probes).10 In this study, we synthesized polyamine-conjugated CNLs to enhance the favorable gene delivery properties of CNLs. Here, we used chemically stable, but biodegradable, carbamate linkages to conjugate the polyamines spermine, spermidine, and putrescine at the 5′ position and oleyl groups at the 2′ and 3′ positions of uridine. Using this approach, we obtained three CNLs featuring different cationic moieties. We found that lipoplexes containing these synthesized CNLs are efficient and nontoxic vectors for the delivery of siRNA.
To introduce hydrophobic units to the 2′ and 3′ positions of uridine selectively, we employed a DMTr unit to protect the 5′-OH group and then treated the protected sugar with oleylamine, CDI, and DMAP (Scheme 1). Because oleylamine did not directly couple to both the 2′ and 3′ positions of the uridine derivative, we obtained a singly substituted compound that we then subjected to the same conditions to obtain the doubly substituted derivative 2. After deprotection of the DMTr unit under acidic conditions, we coupled the 5′-OH group of the uridine derivative with the various polyamines (Fig. 1), followed by protection of their amino groups using Boc-anhydride, to obtain 4a–c. Deprotection of the Boc-groups (4 M HCl in dioxane) provided the desired CNLs 5a–c, which we characterized using 1H and 13C NMR spectroscopy and mass spectrometry. We then used ζ-potential measurements, dynamic light scattering (DLS), transmission electron microscopy (TEM), polyacrylamide gel electrophoresis (PAGE), enzyme-linked immunosorbent assays (ELISA), reverse transcription polymerase chain reaction (RT-PCR), and confocal microscopy to determine the physical and biological properties of the CNLs and their interactions with siRNA.
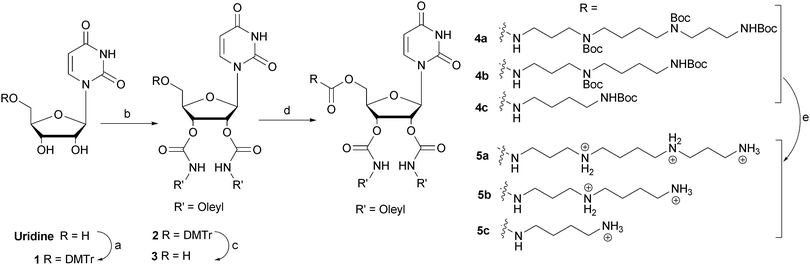 | ||
| Scheme 1 Synthesis of CNLs: (a) DMT-Cl, pyridine, rt, 4 h, 91% yield; (b) CDI, DMAP, oleylamine, DMF, rt, 12 h, 61% yield; (c) 4% TCA in CH2Cl2, 89% yield; (d) (i) CDI, DMAP, polyamine, DMF, rt, 12 h; (ii) (Boc)2O, MeOH, rt (4a, 70% yield; 4b, 68% yield; 4c, 42% yield); (e) 4 M HCl in dioxane, rt, 12 h (5a, 5b, 5c: 97% yield). | ||
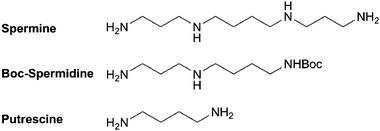 | ||
| Fig. 1 Structures of the polyamines used for the synthesis of CNLs. | ||
To investigate the applicability of these polyamine-conjugated CNLs as gene delivery agents, we first tested their ζ-potentials and sizes. Determining ζ-potentials aids predictions of the stability of the formulation as well as the ability of the positively charged nanoparticles to interact with cell membranes. The ζ-potential of a nucleoside derivative depends on several factors, including the pH, ionic charge, ion size, and concentration of ions in solution. Because the ζ-potentials of the CNL/siRNA ratios (N/P ratios) were positive (Table 1), our CNLs were stable and readily interacted with siRNA through electrostatic interactions.
| N/P ratioa | siRNA–lipoplex ζ potential (+)/mVb | ||
|---|---|---|---|
| 5a | 5b | 5c | |
| a The number of charges of the CNL divided by the number of charges of the siRNA. b Mean values from three experiments; standard deviations are given in parentheses. | |||
| 3 | 19.1 (±2.0) | 6.9 (±3.3) | 21.2 (±1.3) |
| 7.5 | 25.6 (±3.2) | 22.2 (±1.6) | 26.2 (±0.5) |
| 12 | 25.0 (±1.3) | 26.9 (±1.6) | 29.2 (±2.1) |
TEM experiments were performed with CNL/siRNA complexes using a JEM-1011 (negative staining with ammonium molybdate 1% water, Cu carbon-coated grids). TEM images of CNL/siRNA complexes formed vesicles as shown in Fig. S1†. In general, nanoparticles experience relatively higher intracellullar uptake than do their corresponding microparticles. The sizes of the CNL/siRNA complexes were in the range 20–200 nm, measured by DLS (Table 2), making them suitable for transferring siRNA into cells with high transfection efficiency.
| N/P ratioa | siRNA–lipoplex diameterb/nm | ||
|---|---|---|---|
| 5a | 5b | 5c | |
| a The number of charges of the CNL divided by the number of charges of the siRNA. b Mean values from three experiments; standard deviations are given in parentheses. | |||
| 3 | 134 (±2) | 146 (±3) | 184 (±5) |
| 7.5 | 22 (±2) | 112 (±18) | 195 (±8) |
| 12 | 36 (±3) | 153 (±5) | 191 (±5) |
We used PAGE to study the ability of CNLs to bind to siRNA. We found that each of the CNLs formed lipoplexes with siRNA at an N/P ratio of 3 (Fig. 2). Here, interactions between the CNLs and siRNA retarded the PAGE migration of the siRNA, as is typical after the formation of relatively large particles.
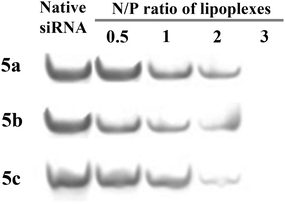 | ||
| Fig. 2 Gel retardation assays performed using the lipoplexes and siRNA. The numbers represent the N/P ratio given in Table 1. | ||
We used WST-1 assays to assess the toxicology of the CNL/siRNA complexes with respect to their N/P ratio, comparing the effect of the CNLs 5a–c on Hela cell line proliferation relative to that of the commercially available transfectant, Lipofectamine 2000. Much like other nucleoside derivatives,6,7,10 the synthesized CNLs did not exhibit significant cytotoxicity at their optimal concentration for transfection. Each of the CNLs exhibited lower cytotoxicity than that of Lipofectamine 2000, except for 5a, which provided 80% cell survival after 24 h of incubation at an N/P ratio of 12 (Fig. 3). Thus, the cell viability depends on both the structure and concentration of the CNLs employed to form the lipoplex.
![Cell viability after treatment with lipoplexes. The cells were treated with 50 nM siRNA. Control (CTL): untreated cell; negative control [Nat (−)]: natural siRNA without lipoplexes; positive control [Nat (+)]: natural siRNA treated with Lipofectamine 2000. The numbers “3,” “7.5,” and “12” indicate the N/P ratios given in Table 1.](/image/article/2011/MD/c1md00014d/c1md00014d-f3.gif) | ||
| Fig. 3 Cell viability after treatment with lipoplexes. The cells were treated with 50 nM siRNA. Control (CTL): untreated cell; negative control [Nat (−)]: natural siRNA without lipoplexes; positive control [Nat (+)]: natural siRNA treated with Lipofectamine 2000. The numbers “3,” “7.5,” and “12” indicate the N/P ratios given in Table 1. | ||
We performed an ELISA to investigate the amount of VEGF protein expressed with the CNLs 5a–c.11 In this experiment, the siRNA (50 nM) alone acted as the negative control and the siRNA with Lipofectamine 2000 as the positive control; we assessed the siRNA transfection results after 24 h. We determined the N/P ratio for the optimal transfection efficiency of each CNL by increasing the N/P ratio from 3 to 12 until the degree of transfection reached greater than 95%. Fig. 4A reveals that siRNA knockdown occurred to greater than 95% at a N/P ratio of 7.5 or 12 for all three of the CNLs. In addition, the CNL 5c provided greater than 85% gene down-regulation at an N/P ratio of 3. VEGF Elisa assay is also performed with 10 nM and 1 nM siRNA concentration (Fig. S2†). Thus, the degree of siRNA transfection was dependent on siRNA concentration, N/P ratio, and the structure of the CNLs. In order to study the specific gene silencing by CNL/siRNA complexes, we tested gene silencing by scrambled VEGF siRNA (Sc-siRNA) and only CNLs. Sc-siRNA, CNL/Sc-siRNA (N/P = 12), and only CNLs did not silenced mRNA in VEGF Elisa assay (Fig. S3†).
![VEGF expression using (A) ELISA and (B) RT-PCR assays. siRNA concentration: 50 nM (n = 3). Control (CTL): untreated cell; negative control [Nat (−)]: natural siRNA without lipoplexes; positive control [Nat (+)]: natural siRNA treated with Lipofectamine 2000. The numbers “3,” “7.5,” and “12” indicate the N/P ratios given in Table 1.](/image/article/2011/MD/c1md00014d/c1md00014d-f4.gif) | ||
| Fig. 4 VEGF expression using (A) ELISA and (B) RT-PCR assays. siRNA concentration: 50 nM (n = 3). Control (CTL): untreated cell; negative control [Nat (−)]: natural siRNA without lipoplexes; positive control [Nat (+)]: natural siRNA treated with Lipofectamine 2000. The numbers “3,” “7.5,” and “12” indicate the N/P ratios given in Table 1. | ||
We confirmed the transfection results for the siRNA through RT-PCR experiments,12 which we executed using agarose gel electrophoresis and staining with ethidium bromide (Fig. 4B). In these studies, we mixed siRNA (50 nM) with 5a, 5b, or 5c (at a N/P ratio of 3, 7.5, or 12). Based on the RNAi mechanism, the delivered VEGF siRNA induces cleavage of the VEGF mRNA, producing VEGF protein; therefore, indirect monitoring (through RT-PCR) of the quantity of mRNA is one method of estimating the siRNA transfection efficiency. The presence of siRNA alone did not affect the quantity of mRNA. In contrast, treatment of HeLa cells with siRNA and CNLs significantly decreased the amount of mRNA produced.
Finally, we examined the cellular uptake of fluorescein-tagged VEGF siRNA (Fl-siRNA), using confocal microscopy to examine their localization in HeLa cells, with Hoechst 33258 (blue) dye for nuclear staining. Here, Fl-siRNA (50 nM) was formulated with the CNLs 5a, 5b, and 5c (at an N/P ratio of 3). In the absence of the CNLs, no green fluorescence appeared in the cells, indicating the lack of siRNA uptake in cells (Fig. 5a and b). With each of the CNLs (5a–c), evidence for the presence of Fl-siRNA appeared in both the cytoplasm and nuclei of the cells, as indicated by green fluorescence (Fig. 5c–h).
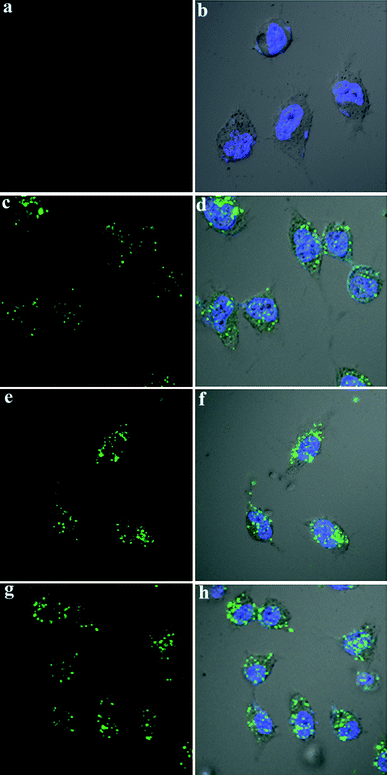 | ||
| Fig. 5 Confocal microscopy images of HeLa cells incubated with (a and b) Fl-siRNA; (c and d) Fl-siRNA and CNL 5a; (e and f) Fl-siRNA and CNL 5b; (g and h) Fl-siRNA and CNL 5c. Concentration of Fl-siRNA: 50 nM. | ||
In conclusion, we have synthesized CNLs conjugated with polyamines via chemically stable, yet biodegradable, carbamate linkers. A gel retardation assay confirmed that the CNLs formed lipoplexes with VEGF siRNA at a minimal N/P ratio of 3. These lipoplexes had diameters in the range 20–200 nm and net positive charge on their surfaces. None of these CNLs significantly affected the cell viability, as determined through a WST-1 assay. Transfection was monitored using ELISA, RT-PCR, and confocal microscopy. Each of the CNLs delivered the siRNA into cells in vitro, down-regulating target mRNA and protein; they all functioned as efficient siRNA carriers at a relatively high N/P ratio of 12, with performance almost equal to that of the commercial transfection agent Lipofectamine 2000. In addition, the CNL 5c exhibited VEGF gene down-regulation of greater than 85% relative to non-treated cells at a relatively low N/P ratio of 3. Thus, the siRNA concentration, the N/P ratio of the CNL/siRNA liposome and the structure of the polyamine were the major factors affecting transfection efficiency. It appears that the structures of these polyamine-based CNLs could be further optimized for gene delivery applications.
Acknowledgements
This study was supported by a grant (no. 2010-0002130) from the National Research Foundation (NRF) and the EPB Center (2010-0001788), funded by the Korean government (MEST). We thank Professor Sungjee Kim for assistance with the DLS and zeta potential measurements.References and notes
- (a) J. C. Venter, M. D. Adams and E. W. Myers, et al., Science, 2001, 291, 1304 CrossRef CAS; (b) P. L. Felgner and G. Rhodes, Nature, 1991, 349, 351 CrossRef CAS.
- (a) S. M. Hammond, A. A. Caudy and G. J. Hannon, Nat. Rev. Genet., 2001, 2, 110 CrossRef CAS.
- (a) C. E. Thomas, A. Ehrhardt and M. A. Kay, Nat. Rev. Genet., 2003, 4, 347 CrossRef; (b) P. P. Karmali and A. Chaudhuri, Med. Res. Rev., 2007, 27, 696 CrossRef CAS; (c) M. Z. Mintzer and E. E. Simanek, Chem. Rev., 2009, 109, 259 CrossRef CAS; (d) S.-D. Li and L. Huang, J. Controlled Release, 2007, 123, 18.
- (a) P. L. Felgner, T. R. Gadek, M. Holm, R. Roman, H. W. Chan, M. Wenz, J. P. Northrop, G. M. Ringold and M. Danielsen, Proc. Natl. Acad. Sci. U. S. A., 1987, 84, 7413; (b) A. D. Miller, Angew. Chem., Int. Ed., 1998, 37, 1768 CrossRef; (c) A. Bhattacharrya and A. Bajaj, Chem. Commun., 2009, 4632 RSC; (d) R. Srinivas, S. Samanta and A. Chaudhuri, Chem. Soc. Rev., 2009, 38, 3326 RSC.
- (a) H. Yanagawa, Y. Ogawa, H. Furuta and K. Tsuno, J. Am. Chem. Soc., 1989, 111, 4567 CrossRef CAS; (b) G. M. Whitesides, J. P. Mathias and C. T. Seto, Science, 1991, 254, 1312 CrossRef CAS.
- (a) L. Moreau, P. Barthélémy, M. E. Maataoui and M. W. Grinstaff, J. Am. Chem. Soc., 2004, 126, 7533 CrossRef CAS; (b) P. Chabaud, M. Camplo, D. Payet, G. Serin, L. Moreau, P. Barthélémy and M. W. Grinstaff, Bioconjugate Chem., 2006, 17, 466 CrossRef CAS; (c) A. Gissot, M. Camplo, M. W. Grinstaff and P. Barthélémy, Org. Biomol. Chem., 2008, 6, 1324 RSC; (d) C. Ceballos, C. H. Prata, S. Giorgio, D. Payet, P. Barthélémy, M. W. Grinstaff and M. Camplo, Bioconjugate Chem., 2009, 20, 193 CrossRef CAS; (e) P. Barthélémy, C. R. Chim., 2009, 12, 171 CrossRef CAS; (f) C. Ceballos, S. Khiati, C. A. H. Prata, X.-X. Zhang, S. Giorgio, P. Marsal, M. W. Grinstaff and M. Camplo, Bioconjugate Chem., 2010, 20, 1062 CrossRef.
- (a) S. K. Choi, T. K. Vu, J. M. Jung, S. J. Kim, H. R. Jung, T. Chang and B. H. Kim, ChemBioChem, 2005, 6, 432 CrossRef CAS; (b) S. M. Park, J. E. Woo, E. M. Jeon and B. H. Kim, Chem. Commun., 2010, 46, 1523 RSC; (c) H. W. Yang, J. W. Yi, E. K. Bang, E. M. Jeon and B. H. Kim, Org. Biomol. Chem., 2011, 9, 291 RSC.
- (a) J. P. Behr, B. Demeneix, J.-P. Loeffler and J. Perez-Mutul, Proc. Natl. Acad. Sci. U. S. A., 1989, 86, 6982 CAS; (b) O. Boussif, F. Lezoualc'h, M. A. Zanta, M. D. Mergny, D. Scherman, B. Demeneix and J. P. Behr, Proc. Natl. Acad. Sci. U. S. A., 1995, 92, 7297 CAS.
- (a) D. L. Liu, J. J. Hu, W. H. Qiao, Z. S. Li, S. B. Zhang and L. B. Cheng, Bioorg. Med. Chem., 2008, 16, 995 CrossRef CAS; (b) H. Lv, S. Zhang, B. Wang, S. Cui and J. Yan, J. Controlled Release, 2006, 114, 100 CrossRef CAS.
- (a) N. Venkatesan and B. H. Kim, Chem. Rev., 2006, 106, 3712 CrossRef; (b) E. K. Bang and B. H. Kim, Tetrahedron Lett., 2009, 50, 2545 CrossRef CAS; (c) S. M. Park, Y. Shen and B. H. Kim, Org. Biomol. Chem., 2007, 5, 610 RSC; (d) S. J. Kim, E. K. Bang, H. J. Kwon, J. S. Shim and B. H. Kim, ChemBioChem, 2004, 5, 1517 CrossRef CAS; (e) S. M. Park, Y. S. Lee and B. H. Kim, Chem. Commun., 2003, 2912 RSC; (f) S. M. Park, S. J. Nam, H. S. Jeong, W. J. Kim and B. H. Kim, Chem.–Asian J., 2011, 6, 487 CrossRef CAS.
- (a) E. Engvall and P. Perlman, Immunochemistry, 1971, 8, 871 CrossRef CAS; (b) B. K. Van Weemen and A. H. Schuurs, FEBS Lett., 1971, 15, 232 CrossRef.
- B. S. K. Miller, Promega Notes, 2001, 79, 15 Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: Experimental details and characterization data. See DOI: 10.1039/c1md00014d |
| This journal is © The Royal Society of Chemistry 2011 |
