DOI:
10.1039/C0MD00246A
(Concise Article)
Med. Chem. Commun., 2011,
2, 378-384
Synthesis, in silico screening and bioevaluation of dispiro-cycloalkanones as antitubercular and mycobacterial COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase inhibitors†
Received
1st December 2010
, Accepted 21st January 2011
First published on 28th February 2011
Introduction
Despite current multidrug therapy and ongoing drug development,1–3 tuberculosis continues to be a major public health concern today. With more than 1.6 million deaths and 9.2 million new cases being reported each year, it is a leading infectious disease claiming millions of death globally.4,5Mycobacterium tuberculosis has the ability to survive for extended periods of time in a human host and thus requires prolonged drug treatment (six to nine months), resulting in low compliance. Moreover, the evolution of multidrug-resistant (MDR) and extremely drug resistant (XDR, recent mortality rate >98%) tuberculosis,6–10 and the AIDS epidemic,11–13 further makes the situation worse. In Mycobacterium tuberculosis drug resistance is not due to a common mechanism for all drugs, but different mechanisms for different classes of drugs.14,15 Almost all the conventional targets and drugs have become inadequate to control resistant TB infection and therefore the discovery of novel, sensitive and selective targets or new chemical entity is needed for development of new generation of antitubercular drugs.
DNA ligases are vital enzymes in replication and repair of DNA. They catalyze the formation of a phosphodiester linkage between adjacent termini in double stranded DNA.16 Two types of DNA ligase viz. COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent and ATP-dependent ligases are known based on their respective co-factor specificities.17 COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ ligases, commonly called LigA, occur almost exclusively in bacteria while ATP-dependent ligases are more ubiquitous and occur additionally in different viruses, archaea, eukaryotes and higher organisms.18,19 Although there is little sequence homology between the eubacterial and eukaryotic enzymes, they exhibit some structural homology in specific domains and the mechanistic steps are broadly conserved.20,21 Different steps involved in the action of DNA ligases involve large conformational changes as well as encircling and partial unwinding of the nicked DNA substrate.22–24M. tuberculosis codes for at least three different types of ATP-dependent ligases and one LigA. Gene knockout and other studies have shown LigA to be indispensable in several pathogens including E. coli, S. typhimurium, S. aureus and B. subtilis in contrast to ATP-dependent ligases which are dispensable in M. tuberculosis.25–30 To find new prototypes as specific inhibitors for COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ ligase is one of the approaches for anti-TB drug research as no drug is known to act against this enzyme so far. COMPOUND LINKS
Read more about this on ChemSpider
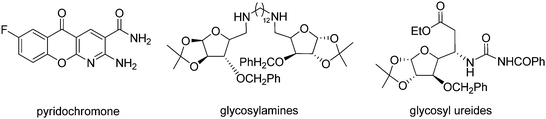
Download mol file of compoundNAD+ ligase specific inhibitors including aryl amines31 and pyridochromanones32 have been reported. We have also shown that glycosyl ureides33 and glycosyl amines34 inhibit the Mycobacterial COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ ligase and have shown bactericidal activity (Fig. 1). As a part of our continuing efforts in tuberculosis chemotherapy35–39 we have recently shown potent antitubercular activities in bis-benzylidene cycloalkanones,40 phenylcyclopropyl methanones41 and alkylaminoaryl phenyl cyclopropyl methanones42 (Fig. 2). The compounds were designed as possible inhibitors of FAS-II but the enzyme inhibition by these compounds was not very significant although few of them showed very promising antitubercular activities.
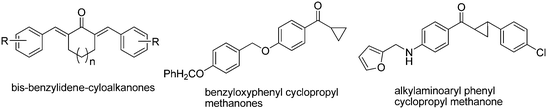 |
| | Fig. 2 Potent antitubercular compounds. | |
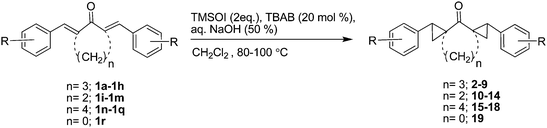
In continuation of this programme we have synthesized a series of dispiro-cycloalkanones by reacting bis benzylidene cyloalkanones with COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundtrimethylsulfoxonium iodide (TMSOI) for methylene insertion in the presence of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNaOH as base and TBAB as phase-transfer catalyst (Corey Chaykovsky reaction).43In silico screening results indicated COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ ligase as targets of these compounds and the compounds were also evaluated in vitro against M. tuberculosis H37Rv and full length COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase from M. tuberculosis.
Results and discussion
Chemistry
The starting substrates α,α′-(E,E)-bis(benzylidene)-cycloalkanones/methanones (COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound1a–1r)40 were prepared by simple condensation of two equivalents of aromatic aldehydes with one equivalent of cycloalkanones/COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundmethanone in the presence of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundKOH (5 mol%) in COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundethanol as earlier reported by us. Cyclopropanation of the double bonds in α,α′-(E,E)-bis(benzylidene)-cycloalkanones was carried out with TMSOI to give the respective disipro-cycloalkanones. In order to establish suitable reaction conditions a model reaction of α,α′-(E,E)-bis(benzylidene)-cyclohexanone (1a) with TMSOI in the presence of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundtetrabutylammonium bromide (TBAB) was carried out in different solvent and bases to give COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2,6-bis-(phenyl)-dispiro[2.1.2.3]decan-4-one (COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2) (Scheme 1) and the results are shown in Table 1.
 |
| | Scheme 1 Optimization of the cyclopropanation reaction using different solvent and base. | |
Table 1 Optimization of the cyclopropanation reaction using different solvent and base
According to the results shown in Table 1, 50% aq. COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNaOH in COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundCH2Cl2 (entry 7) is the most suitable protocol for the reaction. The success of this method is based on the solvation of the ionic species formed during the reaction which favors the reaction rate. Since solvation of ionic species is better in an aqueous COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNaOH/COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundCH2Cl2 combination than in COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundDMSO or COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundDMF as solvent, the reaction yield is accordingly enhanced in aqueous COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNaOH/COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundCH2Cl2 rather than in COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNaOH/COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundDMSO or COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundDMF combination. 44 The structure of compound COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2 was established on the basis of its spectroscopic data and microanalysis (see ESI†). The trans geometry of the cyclopropyl rings in compound COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2 was established on the basis of literature precedents where cyclopropanation of trans-propenones with TMSOI under basic conditions (Corey-Chaykovsky reaction) is always reported to result in trans products.45–47
After establishing the standard reaction conditions, we explored the scope of different substrates in the cyclopropanation. Thus, we carried out the cyclopropanation of different α,α′-(E,E)-bis(benzylidene)-cyclohexanones (1a–1h) with TMSOI to get the desired products 2–9 in moderate to good yields (Table 2, Scheme 2). To see the effect of ring size of the cycloalkanone moiety on this cyclopropanation reaction, the study was extended with α,α′-(E,E)-bis(benzylidene)-cyclopentanones (1i–1m) and α,α′-(E,E)-bis(benzylidene)-cycloheptanones (1n–1q). The reaction of different α,α′-(E,E)-bis(benzylidene)-cyclopentanones/cycloheptanones with TMSOI under similar reaction condition yielded compounds 10–18 in good yields (Scheme 2) and results are shown in Table 2. All the synthesized prototypes were well characterized by their spectroscopic data and microanalysis (see ESI†).
Table 2 Synthesized dispiro-cycloalkanones (2–19) and their in vitro antitubercular activity
| Compd. No. |
n
|

|
C logPa |
MICb (μM) M. tuberculosis H37Rv |
C logP was determined by OSIRIS Property Explorer Programme which is available at http://www.organic-chemistry.org/prog/peo/.
MIC= Minimum inhibitory concentration, the lowest concentration of the compound which inhibits the growth of mycobacterium >90%; MIC of the drugs used as control, COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundINH 4.7 and COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundethambutol 15.9 μM against M. tuberculosis H37Rv.
|
COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2
|
3 |
Phenyl |
5.20 |
>30 |
|
3
|
3 |
4-Fluorophenyl |
5.32 |
>30 |
|
4
|
3 |
4-Chlorophenyl |
6.43 |
16.8 |
|
5
|
3 |
4-Bromophenyl |
6.60 |
13.5 |
|
6
|
3 |
4-Methoxyphenyl |
4.99 |
>30 |
|
7
|
3 |
3,4-Dimethoxyphenyl |
4.78 |
29.6 |
|
8
|
3 |
3,4,5-Trimethoxyphenyl |
4.57 |
25.9 |
|
9
|
3 |
4-Benzyloxyphenyl |
7.73 |
24.3 |
|
10
|
2 |
4-Fluorophenyl |
5.00 |
>30 |
|
11
|
2 |
4-Bromophenyl |
6.28 |
28.0 |
|
12
|
2 |
4-Methoxyphenyl |
4.68 |
>30 |
|
13
|
2 |
3,4-Dimethoxyphenyl |
4.47 |
>30 |
|
14
|
2 |
4-Benzyloxyphenyl |
7.41 |
25.0 |
|
15
|
4 |
4-Chlorophenyl |
6.75 |
>30 |
|
16
|
4 |
4-Methoxyphenyl |
5.31 |
>30 |
|
17
|
4 |
3,4-Dimethoxyphenyl |
5.10 |
28.6 |
|
18
|
4 |
3,4,5-Trimethoxyphenyl |
4.89 |
25.2 |
|
19
|
0 |
4-Benzyloxyphenyl |
6.81 |
26.3 |
Biological activities
The in vitro antitubercular activity against M. tuberculosis H37Rv was determined using agar microdilution method.48 The in silico docking studies were carried out on the above synthesized dispiro-cycloalkanones using autodock tool49,50 and some of the active hits were screened for their mycobacterial COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase inhibitory activity against the full length of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent enzyme from M. tuberculosis, the major Human DNA ligase I and bacteriophage T4 DNA ligase. The compounds were assayed for their antibacterial activity via in vivo assay against S. typhimurium LT2 strain as per earlier reported protocols.51,52
(A)
In vitro antitubercular evaluation
The above synthesized dispiro-cycloalkanones 2–19 were evaluated against virulent strain M. tuberculosis H37Rv. The MIC values were determined using the agar microdilution method.48 As evident from Table 2, among all the compounds screened compounds 4 and 5 were found to possess good activity with MIC 16.8 μM and 13.5 μM against virulent strain. However, compounds 7–9, 11, 14, 17–19 displayed a moderate antitubercular activity with MIC of 25–30 μM against virulent strain, while other compounds COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2, 3, 6, 10, 12, 13, 15 and 16 possess MIC values >30 μM.
The activity results suggest that the conformation as well as the substitution pattern in the aromatic ring both govern the biological activity in the synthesized molecules. The conformational changes in the central alicyclic ring system have more impact on activity than the substitution pattern in the aromatic ring system. All these anti-TB results have good correlation with the in silico as well as in vitro enzymatic assays.
(B)
In silico screening
(C)
In vitro enzymatic assays
To identify the drug target, compounds COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2, 3, 4 and 5 which were sorted out based on the scoring function and fitness scores (minimum docking energy) as implemented in the AUTODOCK program, were assayed against the full length COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase from M. tuberculosis, the major Human DNA ligase I and bacteriophage T4 DNA ligase, for the determination of in vitro inhibitory potency. Two compounds, 3 and 5, showed selective inhibition of M. tuberculosis ligase, and were further evaluated for in vivo antibacterial activities.
In vitro inhibition of nick joining activity
DNA ligase nick joining activity was done as described earlier in the presence of varying concentrations of inhibitors. A quick screening of inhibitors was carried out with DNA ligation assay at high concentration, 100μM against both MtuLigA and T4Lig. This served as a sieve for selecting compounds with the potential to distinguish between COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+- and ATP-dependent ligases for detailed experiments. Based on the obtained results, subsequent efforts were focused on four compounds which we assayed for its in vitro inhibitory potency against the full length COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent enzyme from M. tuberculosis, the major Human DNA ligase I and bacteriophage T4 DNA ligase, respectively. In vitro inhibition data IC50 (Table 3) shows that these compounds are inhibiting MtuLigA in low micromolar range.
Out of the four compounds, compound COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compound2 bound to the Human DNA ligase (HuligI) and T4 DNA ligase (T4 lig) with low affinities; there is no selectivity of this compound for COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ and COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundATP DNA ligases. On the other hand, compound 3 distinguishes between the ATP-dependent DNA ligase and COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase of M. tuberculosis by a factor of four and between Human and M. tuberculosis enzyme by a factor of five and has high affinity for M. tuberculosis enzyme with IC50 of 8.6 μM. Compound 4 also showed greater affinity for M. tuberculosis enzyme while compound 5 showed highest the affinity among all the four compounds. For M. tuberculosis enzyme the IC50 value for this compound is 7.3 μM and it can distinguish between COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase and ATP-dependent DNA ligase by a factor of 8–10 (Fig. 4). A good correlation have been observed between the MIC of in vitro antitubercular activity and IC50 of in vitro inhibition of nick joining activity in compound 4, compound 5 is also in good agreement.
(D) Antibacterial in vivo assay
To evaluate the in vivo inhibition of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ ligases, two bacterial systems were used and the results depicted in Table 4 clearly show that the compounds inhibit more specifically the COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+- dependent DNA ligases as compared to ATP-dependent DNA ligases. Cell viability assay with the compounds again showed that the wild type S. typhimurium LT2 strain is less viable as compared to the viability of the ligase deficient variant rescued with T4Lig (Fig. 5a and 5b, Table 4). The in vivo assay results demonstrate that compounds have higher specificity for COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent ligases and strongly suggest that the observed antibacterial activities are due to in vivo inhibition of LigA.
Table 4 Antibacterial activity of compounds 3 and 5
| Compd. No. |
MIC (μM) |
|
S. typhimurium
|
S. typhimurium
|
E. coliGR501
|
E. coliGR501
|
E. coliGR501
|
|
LT2
|
TT15151
|
+pTRC99A |
+MtuNAD+ligase |
T4 DNA ligase
|
|
3
|
29.5 |
147.9 |
5.9 |
44.3 |
177.5 |
|
5
|
43.4 |
76.0 |
8.6 |
52.1 |
65.2 |
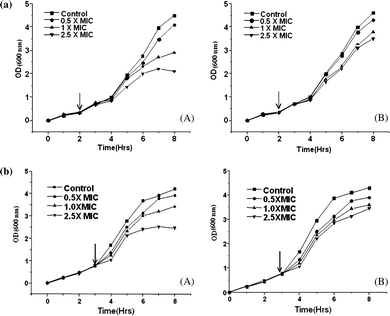 |
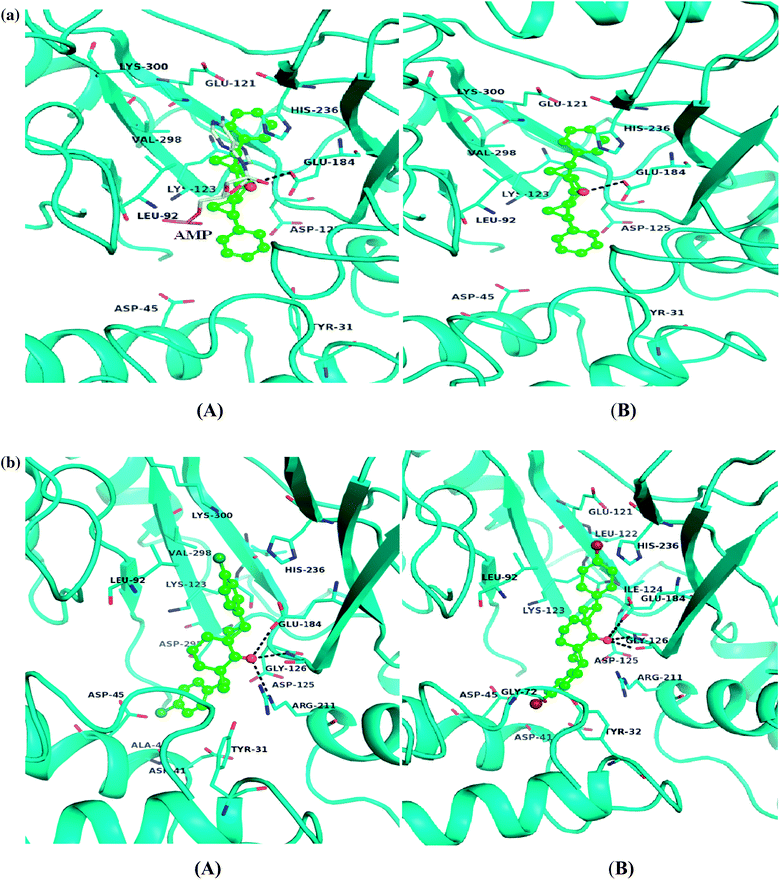
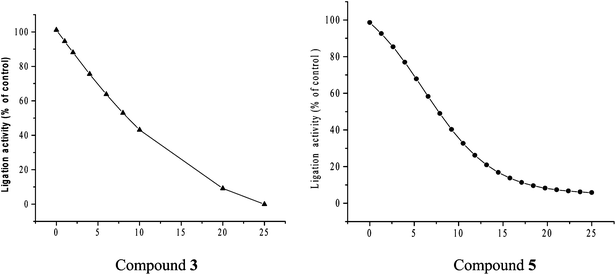
| | Fig. 5 (a). Bactericidal activity of compound 3. (A) S. typhimurium LT2 and (B) its DNA ligase minus (null) derivative TT15151 on their respective exposure to compound 3. (b) Bactericidal activity of compound 5. (A) S. typhimurium LT2 and (B) its DNA ligase minus (null) derivative TT15151 on their respective exposure to compound 5. | |
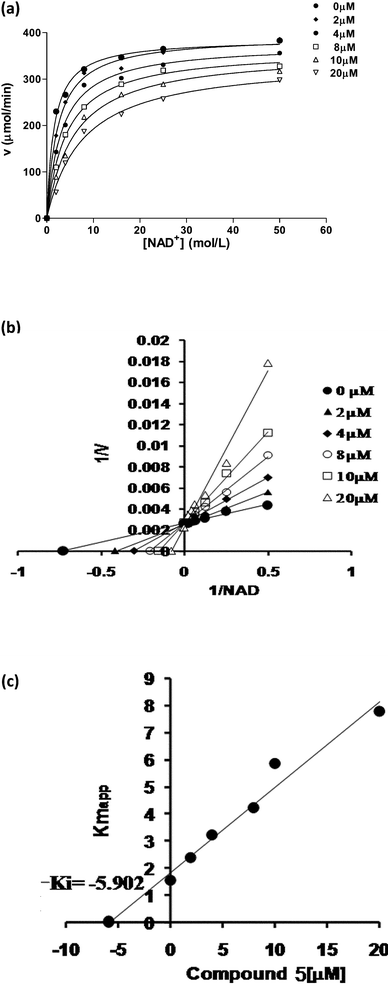
(E) Mode of inhibition
We chose compound 5 to evaluate the mode of inhibition. The standard kinetics was done with COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ in overall nick sealing reaction in vitro. When nick joining activity was measured in the presence of different concentrations of compound 5 (0–50μM) with increasing concentration of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+, the kinetics clearly indicated a competitive inhibition of COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+ by compound 5 (Fig. 6). The linear regression using the apparent Km value leads to Ki value of 5.902μM.
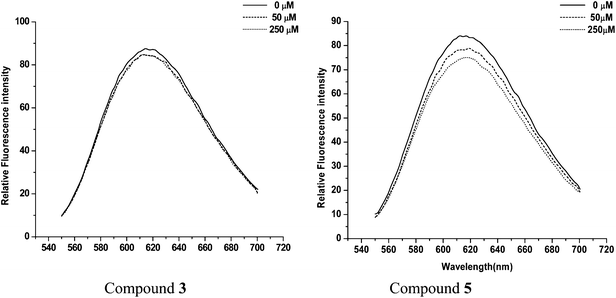
In order to check whether these two active compounds (3 and 5) are generally interacting with DNA and thereby influencing the inhibitory behavior, we carried out COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundethidium bromide displacement assays. Compounds were added to a maximum concentration of 250 μM. Even at this high concentration, representing a 50-fold excess over COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundethidium bromide (5 μM) no loss in fluorescence was observed (Fig. 7). We also carried out gel shift assays where the electrophoretic mobility of plasmid DNA was checked in the presence of increasing inhibitor concentrations. The experiments did not support any general interaction of compounds 3 and 5 with DNA.
Conclusion
In conclusion, a series of dispiro-cycloalkanones with conformationally different cycloalkyl ring systems was synthesized and evaluated against M. tuberculosis H37Rv in vitro and full length of mycobacterial COMPOUND LINKS
Read more about this on ChemSpider
Download mol file of compoundNAD+-dependent DNA ligase. A few of the compounds showed moderate to significant antitubercular and DNA ligase inhibitory activities. The possible mode of action of these compounds was also evaluated by using in silico screening, in vitro and in vivo enzymatic assays.
Acknowledgements
This is a CDRI communication no 8017. The authors thank UGC and CSIR New Delhi for the fellowship. We sincerely acknowledge the financial assistance from DRDO and DBT New Delhi. SAIF CDRI is also acknowledged for providing the spectral and microanalytical data of the compounds.
References
- M. Teresa, G. Lugo and C. A. Bewley, J. Med. Chem., 2008, 51, 2606–2612 CrossRef.
- Y. Zhang, K. P. Martens and S. Denkin, Drug Discovery Today, 2006, 11, 21–27 CrossRef CAS.
- Y. L. Janin, Bioorg. Med. Chem., 2007, 15, 2479–2513 CrossRef CAS.
- K. Duncan and C. E. Barry, Curr. Opin. Microbiol., 2004, 7, 460–465 CrossRef CAS.
-
World Health Organization. Global Tuberculosis Control: Surveillance, Planning, and Financing; WHO Report 2008; WHO Press: Geneva, Switzerland Search PubMed.
- M. Jassal and W. R. Bishai, Lancet Infect. Dis., 2009, 155, 19–30 CrossRef.
- R. P. Tripathi, N. Tewari, N. Dwivedi and V. K. Tiwari, Med. Res. Rev., 2005, 25, 93–131 CrossRef CAS.
- S. H. Gillespie, Antimicrob. Agents Chemother., 2002, 46, 267–274 CrossRef CAS.
- G. J. Ebrahim, J. Trop. Pediatr., 2007, 53, 147–149 Search PubMed.
- E. Huitric, P. Verhasselt, A. Koul, K. Andries, S. Hoffner and D. I. Andersson, Antimicrob. Agents Chemother., 2010, 54, 1022–1028 CrossRef CAS.
- D. Jones, Nat. Rev. Drug Discovery, 2005, 4, 103–103 CrossRef CAS.
- N. R. Gandhi, A. Moll and A. W. Sturm, Lancet, 2006, 4, 1575–1580 CrossRef.
- M. C. Raviglione, N. Engl. J. Med., 2008, 7, 636–638 CrossRef.
-
Y. Zhang, C. Vilcheze and W. R. Jacobs, in Tuberculosis and the tubercle bacillus, (ed.) S. T. Cole, K. D. Eisenach, D. N. McMurray and W. R. Jacobs, Jr., ASM Press, Washington, DC. 2005, pp. 115–142 Search PubMed.
- M. K. Machala, E. Rychta, A. Brzostek, H. R. Sayer, A. R. Galewicz, R. P. Bowater and J. Dziadek, Antimicrob. Agents Chemother., 2007, 51, 2888–2897 CrossRef.
- I. R. Lehman, Science, 1974, 186, 790–797 CAS.
-
M. J. Engler and C. C. Richardson, in The Enzymes, ed. P. D. Boyer, Academic Press, New York, 1982, vol. 15, pp. 3–29 Search PubMed.
- A. Wilkinson, J. Day and R. Bowater, Mol. Microbiol., 2001, 40, 1241–1248 CrossRef CAS.
- V. Sriskanda, R. W. Moyer and S. J. Shuman, J. Biol. Chem., 2001, 276, 36100–36109 CrossRef CAS.
- D. J. Timson, M. R. Singleton and D. B. Wigley, Mutat. Res., 2000, 460, 301–318 CAS.
- A. J. Doherty and S. W. Suh, Nucleic Acids Res., 2000, 28, 4051–4058 CrossRef.
- J. M. Pascal, P. J. O'Brien, A. E. Tomkinson and T. Ellenberger, Nature, 2004, 432, 473–478 CrossRef CAS.
- K. C. Gajiwala and C. Pinko, Structure, 2004, 12, 1449–1459 CrossRef CAS.
- J. Y. Lee, C. Chang, H. K. Song, J. Moon, J. K. Yang, H. K. Kim, S. T. Kwon and S. W. Suh, EMBO J., 2000, 19, 1119–1129 CrossRef CAS.
- C. Gong, A. Martins, P. Bongiorno, M. Glickman and S. Shuman, J. Biol. Chem., 2004, 279, 20594–20606 CrossRef CAS.
- A. Wilkinson, H. Sayer, D. Bullard, A. Smith, J. Day, T. Kieser and R. P. Bowater, Proteins: Struct., Funct., Bioinf., 2003, 51, 321–326 CrossRef CAS.
- J. J. Dermody, G. T. Robinson and R. Sternglanz, J. Bacteriol., 1979, 139, 701–704 CAS.
- F. S. Kaczmarek, R. P. Zaniewski, T. D. Gootz, D. E. Danley, M. N. Mansour, M. Griffor, A. V. Kamath, M. Cronan, J. Mueller, D. Sun, P. K. Martin, B. Benton, L. McDowell, D. Biek and M. B. Schmid, J. Bacteriol., 2001, 183, 3016–3024 CrossRef CAS.
- M. A. Petit and S. D. Ehrlich, Nucleic Acids Res., 2000, 28, 4642–4648 CrossRef CAS.
- C. M. Sassetti, D. H. Boyd and E. J. Rubin, Mol. Microbiol., 2003, 48, 77–84 CrossRef CAS.
- G. Ciarrocchi, D. G. MacPhee, L. W. Deady and L. Tilley, Antimicrob. Agents Chemother., 1999, 43, 2766–2772 CAS.
- H. Brotz-Oesterhelt, I. Knezevic, S. Bartel, T. Lampe, U. Warnecke-Eberz, K. Ziegelbauer, D. Habich and H. Labischiinski, J. Biol. Chem., 2003, 278, 39435–39442 CrossRef.
- S. K. Srivastava, R. P. Tripathi and R. Ramachandran, J. Biol. Chem., 2005, 280, 30273–30281 CrossRef CAS.
- S. K. Srivastava, D. Dube, N. Tewari, N. Dwivedi, R. P. Tripathi and R. Ramachandran, Nucleic Acids Res., 2005, 33, 7090–7101 CrossRef CAS.
- D. Katiyar, V. K. Tiwari, R. P. Tripathi, A. Srivastava, V. Chaturvedi, R. Srivastava and B. S. Srivastava, Bioorg. Med. Chem., 2003, 11, 4369–4375 CrossRef CAS.
- N. Tewari, V. K. Tiwari, R. P. Tripathi, V. Chaturvedi, A. Srivastava, R. Srivastava, P. K. Shukla, A. K. Chaturvedi, A. Gaikwad, S. Sinha and B. S. Srivastava, Bioorg. Med. Chem. Lett., 2004, 14, 329–332 CrossRef CAS.
- R. C. Mishra, R. Tripathi, D. Katiyar, N. Tewari, D. Singh and R. P. Tripathi, Bioorg. Med. Chem., 2003, 11, 5363–5374 CrossRef CAS.
- N. Tewari, V. K. Tiwari, R. C. Mishra, R. P. Tripathi, A. K. Srivastava, R. Ahmad, R. Srivastava and B. S. Srivastava, Bioorg. Med. Chem., 2003, 11, 2911–2922 CrossRef CAS.
- N. Tewari, V. K. Tiwari, D. Katiyar, N. Saxena, S. Sinha, A. Gaikwad, A. Srivastava, V. Chaturvedi, Y. K. Manju, R. Srivastava and B. S. Srivastava, Bioorg. Med. Chem., 2005, 13, 5668–5679 CrossRef.
- N. Singh, J. Pandey, A. Yadav, V. Chaturvedi, S. Bhatnagar, A. Gaikwad, S. Sinha, A. Kumar, P. K. Shukla and R. P. Tripathi, Eur. J. Med. Chem., 2009, 44, 1705–1709 CrossRef CAS.
- N. Dwivedi, N. Tewari, V. K. Tiwari, V. Chaturvedi, Y. K. Manju, A. Srivastava, A. Giakwad, S. Sinha and R. P. Tripathi, Bioorg. Med. Chem. Lett., 2005, 15, 4526–4530 CrossRef CAS.
- A. Ajay, V. Singh, S. Singh, S. Pandey, S. Gunjan, D. Dubey, S. K. Sinha, B. N. Singh, V. Chaturvedi, R. Tripathi, R. Ramchandran and R. P. Tripathi, Bioorg. Med. Chem., 2010, 18, 8289–8301 CrossRef CAS.
- E. J. Corey and M. Chaykovsky, J. Am. Chem. Soc., 1965, 87, 1353–1364 CrossRef CAS.
- M. H. Abraham and Y. H. Zhao, J. Org. Chem., 2004, 69, 4677–4685 CrossRef CAS.
- S. Chandrasekhar, C. Narasihmulu, V. Jagadeshwar and K. V. Reddy, Tetrahedron Lett., 2003, 44, 3629–3630 CrossRef CAS.
- R. J. Paxton and R. J. K. Taylor, Synlett, 2007, 4, 633–637.
- A. Hartikka and P. I. Arvidsson, J. Org. Chem., 2007, 72, 5874–877 CrossRef CAS.
- H. Saito, H. Tomioka, K. Sato, M. Emori, T. Yamane, K. Yamashita, K. Hosol and T. Hidaka, Antimicrob. Agents Chemother., 1991, 35, 542–547 CAS.
- ACCELRYS ver., 11. San Diego, CA, Accelrys Inc., 2000, pp. 92121–92152.
- G. M. Morris, D. S. Goodsell, R. S. Halliday, R. Huey and W. E. Hart, J. Comput. Chem., 1998, 19, 1639–1662 CrossRef CAS.
- J. B. Le-Pecq and C. Paoletti, J. Mol. Biol., 1967, 27, 87 CrossRef CAS.
- S. K. Srivastava, R. P. Tripathi and R. Ravishankar, J. Biol. Chem., 2005, 280, 30273–30281 CrossRef CAS.
Footnotes |
| † Electronic supplementary information (ESI) available: Experimental procedures, characterization data and copies of 1H NMR and 13C NMR spectra, protocols of biological assays See DOI: 10.1039/c0md00246a |
| ‡ Authors having equal contribution. |
|
| This journal is © The Royal Society of Chemistry 2011 |
Click here to see how this site uses Cookies. View our privacy policy here.