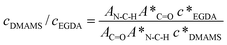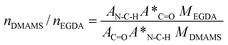Single-step fabrication of non-leaching antibacterial surfaces using vapor crosslinking†
Yumin
Ye‡
,
Qing
Song‡
and
Yu
Mao
*
Departments of Biosystems Engineering, Oklahoma State University, Stillwater, OK 74078, USA. E-mail: yu.mao@okstate.edu; Fax: +1 405 744 6059; Tel: + 1 405 744 4337
First published on 19th October 2010
Abstract
We report the fabrication of non-leaching antibacterial surfaces using a single-step vapor crosslinking method. Vapors of dimethylaminomethylstyrene (DMAMS) and ethylene glycol diacrylate (EGDA) were copolymerized to produce crosslinked polymer coatings conformally on both planar and porous substrates. The antibacterial properties of coatings with systematically controlled crosslinked degrees were studied. The crosslinked coatings kill bacteria through disruption of the bacteria membrane upon surface contact. Bacteria reduction rates as high as 99.99% were achieved against both Gram-positive Bacillus subtilis and Gram-negative Escherichia coli. The strong antibacterial activity of the highly crosslinked coatings indicates that the mobility and length of antibacterial polymer chains are not critical in determining the bactericidal activity of the P(DMAMS-co-EGDA) coatings. Leaching tests demonstrated that the P(DMAMS-co-EGDA) coatings did not leach from the surface and were stable after the durability tests.
Introduction
Bactericidal surfaces are highly desirable to prevent bacteria-associated infections in hospital and health care facilities. Silver and antibiotics have been widely used to create bactericidal surfaces.1–3 However, the bacteria killing is based on the release of the antibacterial agents into bacteria and the surrounding environment, which raises concerns since the leaching of antibiotics can contribute to the escalation of bacteria resistance. An alternative strategy is to create non-leaching antibacterial surfaces through immobilization of antibacterial agents. The non-leaching surfaces kill bacteria on contact, which was reported to reduce the probability of developing bacteria resistance.4–6Considerable research efforts have been performed to covalently immobilize polycationic compounds to create permanent antibacterial surfaces. Poly(2-(dimethylamino)ethyl methacrylate) was grafted on pretreated glass substrates using atom radical transfer polymerization.7–9 UV-induced graft copolymerization was used to immobilize poly(4-vinylpyridine) on polymeric and cellulosic materials.10 Surface derivation was reported to immobilize alkylated poly(ethylene imine).11 Among the immobilization schemes reported, most required multiple processing steps in various solvents (for both substrate surface treatment and immobilization reactions) for long periods of time (10–48 h) and at elevated temperatures.12–14 These processing conditions may affect the physical properties of substrate materials and restrict the scale-up of the immobilization process.15
Another approach to create antibacterial surfaces is to physically adsorb water-insoluble polycationic compounds on substrate materials. Antibacterial surfaces were fabricated by spraying solutions of fluorinated pyridinium block copolymers.16Polymer nanoparticles with emulsifiers consisting of a hydrophobic polystyrene block and a hydrophilic block of poly(4-vinyl-N-methylpyridinium iodide) were also developed to produce antimicrobial coatings from aqueous solutions.17 In another study, antimicrobial polymer coatings with zero solubility in water were formed using a solventless chemical vapor deposition.18
These previous studies were focused on grafted or linear antibacterial polymers. Vapor crosslinking of antibacterial polymers with well-defined structure has not been reported, and the effect of crosslinking on the antibacterial properties is not clear. In the most relevant related work, poly(ethylene imine) was thermally crosslinked in a methanol solution and subsequently casted to form coatings, which were highly effective against bacteria but showed leaching of the polymer.19 In this paper, we report immobilization of antibacterial polymers through a one-step vapor crosslinking method. Vapor molecules of a crosslinkable monomer, ethylene glycol diacrylate (EGDA), and an antimicrobial monomer, dimethylaminomethylstyrene (DMAMS), copolymerize directly on planar or porous surfaces. The reaction proceeds through a radical polymerization pathway initiated by a peroxide initiator.20 The tertiary amine groups in DMAMS units become partially protonated at neutral pH conditions, resulting in high bacterial killing efficiency.21 As a result, crosslinked coatings are formed on the surface of interest with cationic charges distributed across the polymer network (Fig. 1). Poly(dimethylaminomethylstyrene-co-ethylene glycol diacrylate) (P(DMAMS-co-EGDA)) coatings with systematic control over the crosslinking degree were synthesized, and the antibacterial activities were studied.
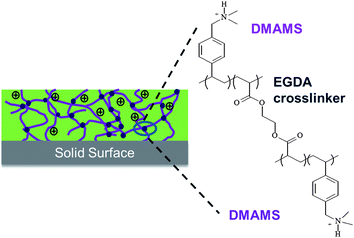 | ||
| Fig. 1 Structure of the crosslinked P(DMAMS-co-EGDA) coatings. | ||
Experimental
Materials
DMAMS (95%), EGDA (90%), and tert-butyl peroxide (TBP) (98%) were all purchased from Sigma-Aldrich. The DMAMS monomer consists of 50/50 ortho- and para-isomers and was vacuum purified prior to use to remove volatile contaminants. EGDA and TBP were directly used for deposition without further purification. Silicon wafer (MEMC Electronic Materials) and glass slides were used as the planar substrates during the vapor deposition. A nylon fabric (306 A, Testfabric) was also used as the substrate to test the applicability of the coating process on porous surfaces. Silicon wafer was used as received. Glass slides were cleaned using ultrasonication in acetone for 15 min and dried under N2 flow. Nylon fabrics with an average fiber diameter of 16–18 μm were cleaned using detergent and cut into squares of 4 × 4 cm2.Synthesis
The vapor crosslinking was carried out in a custom-built reactor (Sharon Vacuum) with a diameter of 25 cm. The reactor was equipped with a parallel array of Nichrome filament (Ni80/Cr20, Goodfellow) and water-cooled stage on which the samples were placed. The distance between the stage and filament was 2.5 cm. The monomers of DMAMS and EGDA were vaporized at 75 °C and 60 °C, respectively, and fed into the reactor through mass flow controllers (MKS, model 1153 and 1150). The initiator of TBP was vaporized at room temperature and metered into the reactor through a mass flow controller (1479A, MKS). The flow rate of TBP was kept constant at 0.4 sccm, while the flow rates of DMAMS and EGDA were varied in each deposition (Table 1). The filament was resistively heated up to 250 °C, and the temperature of the substrates was kept at 45–48 °C viawater circulation at the back side of the stage. Both temperatures were measured using thermal couples (Omega, Type K) directly attached to the filament and the stage. The pressure inside the reactor was maintained at 0.25 Torr using a throttling butterfly valve (MKS, Type 253B). The coating deposition process was monitored by measuring the increase of coating thickness on silicon wafer using an interferometry system with a 633 nm He–Ne laser (JDS Uniphase). The deposition time was approximately 45–55 min. The thickness of the coatings on silicon wafer and glass slides was kept at 800 nm ± 20 nm. Each nylon fabric sample was weighed before and after deposition to calculate the mass of the coating. The total coating mass on the two sides of each fabric sample was maintained at 50 ± 10 μg cm−2.| Sample | Flow rate (sccm) | Flow ratio | n DMAMS/nEGDAa | Crosslinking degree (%) | |
|---|---|---|---|---|---|
| DMAMS | EGDA | ||||
| a Molar ratio of DMAMS to EGDA in crosslinked coatings. | |||||
| H1 | 0.68 | — | — | — | 0 |
| H2 | — | 0.18 | — | — | 100 |
| C1 | 0.35 | 0.18 | 1.9 | 1.2 | 63 |
| C2 | 0.52 | 0.18 | 2.9 | 2.4 | 45 |
| C3 | 0.58 | 0.16 | 3.6 | 2.8 | 42 |
| C4 | 0.49 | 0.13 | 3.8 | 3.4 | 37 |
| C5 | 0.49 | 0.12 | 4.1 | 5.4 | 27 |
| C6 | 0.60 | 0.12 | 5.0 | 7.0 | 22 |
| C7 | 0.68 | 0.12 | 5.7 | 7.5 | 21 |
| C8 | 0.62 | 0.09 | 6.9 | 8.8 | 19 |
Characterization methods
A FEI Quanta 600F scanning electron microscope (SEM) was used to observe the morphology of the coated surfaces. Fourier transform infrared (FTIR) spectra of coatings on the nylon textiles and silicon surfaces were collected by a Nicolet 6700 FTIR spectrometer using a DTGS detector under attenuated total reflectance mode and transmission mode, respectively. The FTIR spectra of the coatings on nylon textiles were similar to those on silicon substrates but with more background noise. The coating thickness on silicon substrates was measured from the depth profiles of the coated surfaces with Z-shaped scratches. The depth profiles were obtained using a Veeco Multimode SPM atomic force microscope (AFM). The durability tests were performed by washing the samples in phosphate buffered saline (PBS) solutions at 200 rpm under 37 °C for 24 h and ultrasonication for 30 min.Antibacterial assessment
Live/dead viability assay (Invitrogen) is based on a two-color fluorescence method using the fluorescent dye mixture of a green SYTO9 stain and a red propidium iodide stain. The dye mixture was first incubated with Escherichia coli at 107CFU ml−1. Subsequently an aliquot of 15 μl of the bacteria solution was applied to coated and un-coated glass surfaces. The test surfaces were covered with glass coverslips and incubated in the dark for 40 min. Afterwards the bacteria were observed under an Olympus BX51 epifluorescence microscope using a green filter (excitation/emission 440–480 nm/515–540 nm) and a red filter (540–560 nm/630–660 nm).The antibacterial activities of the coatings were quantitatively evaluated according to standard ASTM E2149-01 against a Gram-negative bacterium, E. coli (ATCC 29425), and a Gram-positive bacterium, Bacillus subtilis (ATCC 6633). The bacteria were cultured for 18 h in LB liquid medium and diluted 103 times in 50 ml sterile PBS solution to 1.5–3.0 × 106 colony forming units per ml (CFU ml−1). The resultant solution was used as the working bacterial dilution. Coated and uncoated nylon fabrics were shaken at 200 rpm in the working solution for 60 min at 37 °C (E. coli) or 30 °C (B. subtilis) using an incubator shaker (Isotemp, Fisher Scientific). The supernatant was subsequently diluted to 101, 102, and 103 in series, and 100 μl of the 103 dilution was incubated in a LB agar plate for 20 h. Each sample was placed in three plates. The bacterial colonies were counted, and the percentile reduction was calculated from CFU ml−1 of uncoated textiles (U) and the CFU ml−1 of coated textiles (C): reduction rate (%) = (U − C)/U × 100 (%). Error bars were estimated as the standard deviation of the three measurements.
The zone of inhibition test was performed to examine any leaching of the antibacterial coatings. A hole with 8 mm in diameter was bored into the LB agar plate with confluent lawn of B. subtilis or E. coli (1 × 105CFU ml−1). A 100 μl aliquot of the PBS solutions after 60 min shaking with the samples were added to the holes, and the plates were incubated overnight at 30 °C (B. subtilis) or 37 °C (E. coli).
Results and discussion
Vapor crosslinking synthesis
P(DMAMS-co-EGDA) coatings (C1–C8) with systematically varied crosslinking degrees were synthesized using the vapor crosslinking method. Poly(dimethylaminomethylstyrene) (PDMAMS, H1) and poly(ethylene glycol diacrylate) (PEGDA, H2) homopolymer coatings were also synthesized using the same deposition method. The flow rates of DMAMS and EGDA vapors during each polymer synthesis were varied and listed in Table 1. By adjusting the flow ratio, the DMAMS composition in the P(DMAMS-co-EGDA) coatings was varied, which changed the crosslinking degree of the copolymers. Fig. 2 shows the FTIR spectra of P(DMAMS-co-EGDA) coatings C2, C5 and C8, compared with those of PDMAMS and PEGDA coatings. All the spectra were obtained from coatings on silicon surfaces using the transmission mode. The absorption peaks at 2768 and 2814 cm−1, assigned to the C–H stretching in the –N(CH3)2group,22 indicate that tertiary amine groups were retained during the vapor deposition process. No absorption peak of carbonyl groups at around 1730 cm−1 was found in the spectrum of the PDMAMS coating, indicating no oxidation of the DMAMS monomer during the deposition process. In addition, the absence of absorption peaks at 1630–1640 cm−1 in the spectra of copolymer coatings suggests that all the double bonds in DMAMS and EGDA have been reacted during the vapor crosslinking process. The spectra of the copolymer coating series demonstrate a gradual change in the composition: with the increase of the DMAMS/EGDA flow ratio, the intensity of the C–H stretching in the tertiary amine group (2700–2850 cm−1) increases, while the intensity of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching (1735 cm−1) in EGDA decreases.
O stretching (1735 cm−1) in EGDA decreases.
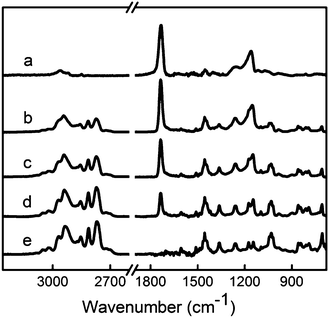 | ||
Fig. 2
FTIR spectra of polymer PEGDA (a), P(DMAMS-co-EGDA) copolymers C2 (b), C5 (c) and C8 (d), and polymer PDMAMS (e). Absorption peaks in the range of 2700–2850 cm−1 represent the C–H stretching of tertiary amine groups from DMAMS, while the absorption at 1735 cm−1 represents the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching from EGDA. O stretching from EGDA. | ||
The composition of the copolymer coatings was quantified using FTIR analysis. The FTIR spectra of PDMAMS, PEGDA, and P(DAMSM-co-EGDA) coatings were normalized to the coating thickness prior to analysis. After decoupling using the Peak Resolve tool of the Omnic software, the peak areas of the C–H stretching in the tertiary amine at 2814 cm−1 and the carbonyl absorption in EGDA at 1735 cm−1 were measured. According to the Beer–Lambert equation, the absorbance peak area of a particular mode is proportional to the concentration of the moiety and the absorption coefficient. We assume that the absorption coefficient of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching is the same in the P(DMAMS-co-EGDA) copolymers as in the homopolymer, as verified in other acrylic copolymers.23 The ratio of the EDGA mole concentration in P(DMAMS-co-EGDA) (cEGDA) to the EGDA mole concentration in PEDGA (c*EGDA) can be calculated using: cEGDA/c*EGDA = AC
O stretching is the same in the P(DMAMS-co-EGDA) copolymers as in the homopolymer, as verified in other acrylic copolymers.23 The ratio of the EDGA mole concentration in P(DMAMS-co-EGDA) (cEGDA) to the EGDA mole concentration in PEDGA (c*EGDA) can be calculated using: cEGDA/c*EGDA = AC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/A*C
O/A*C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, where AC
O, where AC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and A*C
O and A*C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O are the peak areas of the C
O are the peak areas of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption in the spectra of copolymer and homopolymer, respectively. Similarly, the ratio of the DMAMS mole concentration in P(DMAMS-co-EGDA) (cDMAMS) to the DMAMS mole concentration in PDMAMS (c*DMAMS) can be calculated using: cDMAMS/c*DMAMS = AN-C-H/A*N-C-H, where AN-C-H and A*N-C-H are the peak areas of the N–C–H absorption in the spectra of copolymer and homopolymer, respectively. Putting the two equations together, we can get:
O absorption in the spectra of copolymer and homopolymer, respectively. Similarly, the ratio of the DMAMS mole concentration in P(DMAMS-co-EGDA) (cDMAMS) to the DMAMS mole concentration in PDMAMS (c*DMAMS) can be calculated using: cDMAMS/c*DMAMS = AN-C-H/A*N-C-H, where AN-C-H and A*N-C-H are the peak areas of the N–C–H absorption in the spectra of copolymer and homopolymer, respectively. Putting the two equations together, we can get:
Since mole concentration can be calculated form the density and the molecular weight, c*EGDA/c*DMAMS is equal to MDMAMS/MEGDA under the assumption of equal density for PDMAMS and PEGDA coatings, where MDMAMS and MEGDA are the molecular weights of the DMAMS and EGDA repeating units, respectively. The molar ratio of DMAMS units to EGDA units in each copolymer coating, nDMAMS/nEGDA, is equal to cDMAMS/cEGDA and can be calculated using the equation:
The crosslinking degree (CD%) was calculated as the mole fraction of crosslinked monomer units: CD% = 2/(nDMAMS/nEGDA + 2) × 100%. There is a factor of 2 because each EDGA has two double-bond units. The calculated molar ratio of nDMAMS/nEGDA and the corresponding crosslinking degree in each copolymer coating were summarized in Table 1. With the decrease of the DMAMS/EGDA flow ratio, the molar ratio of nDMAMS/nEGDA in the copolymer coatings decreases, and the crosslinking degree increases.
Antibacterial activities
The live/dead viability assay was used to study the interaction of the P(DMAMS-co-EGDA) coatings with bacteria. The SYTO9 stain labels bacteria with both intact and damaged membranes, while the propidium iodide stain only labels bacteria with damaged membranes.24 The E. coli on the uncoated glass slides gave only green fluorescence (Fig. 3a and b), indicating that bacteria retained intact cell membranes. However, after exposure to the P(DMAMS-co-EGDA) coatings, all the E. coli bacteria showed red fluorescence and reduced green fluorescence (Fig. 3c and d), indicating that the propidium iodide stain competed for nucleic acid binding sites and displaced the SYTO9 stain inside the bacteria with damaged membranes.24 The viability assay clearly demonstrated that the bacteria membrane was disrupted after being in contact with the P(DMAMS-co-EGDA) coated surface. The bacteria solution was also examined in a confocal manner by adjusting the focal plane. The majority of bacteria were found in the suspension between the coverslip and the coated surface, while very few bacteria were observed on the focal plane around the coated surface. The SEM examination of the coated surface (not shown) confirmed that the killed bacteria did not accumulate on the surface.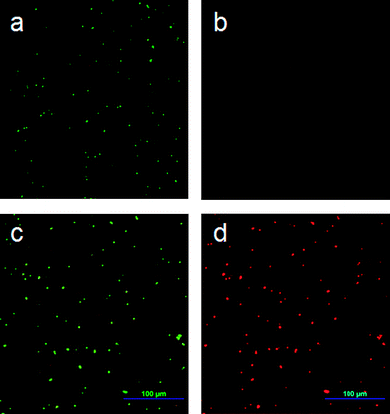 | ||
| Fig. 3 Fluorescence microscopy images of E. coli after 40 min incubation: (a, b) on uncoated glass surface using green and red filters, respectively; (c, d) on P(DMAMS-co-EGDA) (C8) coated surface using green and red filters, respectively. | ||
The antibacterial efficiencies of the P(DMAMS-co-EGDA) coatings were quantitatively measured following the standard of ASTM E2149-01. Fig. 4a shows the bacteria reduction rate of coatings with different nDMAMS/nEGDA ratios. Three features were observed: (1) the bacteria killing efficiency increases with the ratio of nDMAMS/nEGDA; (2) there is a threshold ratio of nDMAMS/nEGDA above which significant bacteria killing can be achieved; and (3) the threshold ratio is different for the killing of different bacteria. A reduction rate of more than 99% was found for the Gram-positive B. subtilis when nDMAMS/nEGDA exceeded 2.8, while a much higher ratio was needed to obtain an equivalent reduction rate of the Gram-negative E. coli. This phenomena is possibly due to the difference in the structure of the cell envelope between B. subtilis and E. coli.25 Nevertheless, copolymer coatings with a significant composition of the non-biocidal EGDA component demonstrated strong bactericidal activity. For example, the coating of C6 with a nDMAMS/nEGDA ratio of 7.0 and a crosslinking degree of 22% has 99.9% killing rate against both B. subtilis and E. coli.Fig. 4b shows the bacterial colonies formed on the agar plates from the antibacterial tests of the uncoated surface and the surface coated with C6. No colony forming unit was found in the plate of coated surface, indicating a bacterial reduction rate of close to 100%.
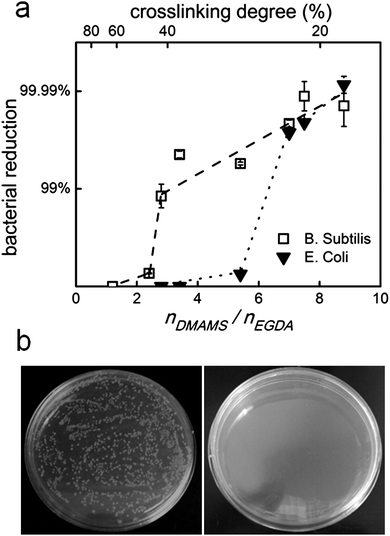 | ||
| Fig. 4 (a) Bacteria reduction rate of P(DMAMS-co-EGDA) coatings against B. subtilis and E. coli. Dashed and dotted lines only indicate the trend of data. Where not shown, error bars are smaller than the data symbols. (b) Bacterial colonies formed on the agar plates from antibacterial tests of control (left) and P(DMAMS-co-EGDA) coating C6 (right). | ||
The observation indicates that polymers with highly restricted mobility can have strong antibacterial activity, which provides several insights into the antibacterial action of the crosslinked antibacterial coatings. The least-crosslinked coating (C8) only has an average of 4.4 DMAMS repeating units between crosslinks. Such short segments will not be long enough to insert into bacteria membranes (with a thickness of 40–55 nm26,27) to induce membrane disruption. In other words, the mobility and length of polymer chains is not critical in determining the bactericidal activity of the crosslinked coatings. A possible mechanism is the release of structurally important cations within the bacteria membrane induced by the positive charges on the coating surface,8,28 which eventually results in the loss of membrane structure.
The results of antibacterial activity also suggest that the density of the DMAMS units plays a more important role in the antibacterial action of the P(DMAMS-co-EGDA) coatings than the crosslinking degree. The P(DMAMS-co-EGDA) coatings with significant bacteria killing (C3–C8) have an average of 1.4–4.4 DMAMS repeating units between crosslinks. At such high crosslink degrees, the change of bacteria killing efficiency with the coating composition is unlikely to be due to the difference in the mobility of polymer chains, but rather due to the different concentration of the effective antibacterial component. To further improve the antibacterial efficiency of the crosslinked coatings, crosslinking monomers with bactericidal moiety can be used in the coating synthesis process to increase the concentration of antibacterial components.
Non-leaching property
Thorough examinations of the possibility of leaching-induced bactericidal effect were performed. From the standard zone of inhibition test, no inhibited growth of either E. coli or B. subtilis was observed on the PDMAMS coating and the P(DMAMS-co-EGDA) coatings. Since the inhibition zone test may not be sufficiently sensitive to detect the leaching of antibacterial agents, samples were tested using a revised version of standard ASTM E2149-01 to determine if the bactericidal effect was partially due to the leaching. The coated samples were washed vigorously in PBS solutions at 200 rpm for 60 min. After removing the samples, the solutions were inoculated with E. coli and incubated for 60 min. The bactericidal efficiencies of the solutions were tested using the serial dilution method. The solutions exposed to all the crosslinked P(DMAMS-co-EGDA) coatings showed no killing efficacy towards E. coli. The test results of C6–C8 were summarized in Fig. 5. In comparison, the solution exposed to the un-crosslinked PDMAMS coating demonstrated strong bactericidal activity with more than 99.99% reduction rate. The above results clearly demonstrate that the P(DMAMS-co-EGDA) coatings did not leach into the environment to kill the bacteria. Instead, the bacteria are killed upon surface contact.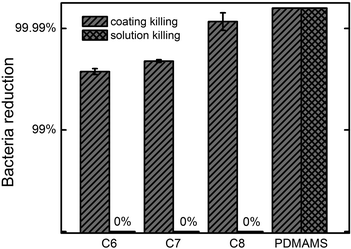 | ||
| Fig. 5 E. coli killing efficacies of P(DMAMS-co-EGDA) copolymer coatings C6, C7, C8 and homopolymer coating PDMAMS, compared with the killing efficacies of their 1 h exposure solutions. | ||
The stability of the P(DMAMS-co-EGDA) coating in aqueous solutions was studied using FTIR. The FTIR spectra of the P(DMAMS-co-EGDA) coatings after the 24 h durability tests showed that the peak position and intensity of all absorption peaks remained unchanged (Fig. 6), indicating that no leaching of polymers into the solutions occurred. By contrast, the uncrosslinked PDMAMS coating lost the majority of the FTIR absorption after the same test. The observation is in agreement with the results obtained from the solution leaching tests. The morphology of the P(DMAMS-co-EGDA) coatings after the durability test was observed using SEM (Fig. S1†). All the P(DMAMS-co-EGDA) coatings (C1–C8) were intact and smooth with no delamination or “pin-hole” formed after the 24 h durability test followed by ultrasonication for 30 min. On the contrary, most of the uncrosslinked PDMAMS coating was cleared with debris left on the fiber surface after the durability test. It is evident that the vapor crosslinking method significantly improved the durability of the P(DMAMS-co-EGDA) coatings without compromising the antibacterial functionality, enabling one-step synthesis of non-leaching antibacterial coatings directly on both planar and porous surfaces.
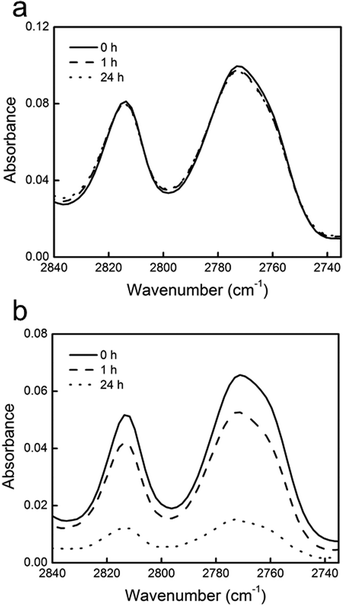 | ||
| Fig. 6 FTIR spectra of the C–H stretching absorbance in tertiary amine groups of (a) P(DMAMS-co-EGDA) coating C7 and (b) un-crosslinked PDMAMS coating before (solid line), after 1 h (dashed line) and 24 h (dotted line) washing in PBS solutions. | ||
Conclusions
Non-leaching, antibacterial P(DMAMS-co-EGDA) coatings were successfully synthesized using vapor crosslinking in one step. The deposition process formed antibacterial polymer coatings with well-defined crosslinked structure on both planar and porous surfaces. The crosslinking degree of P(DMAMS-co-EGDA) coatings was systematically varied by adjusting the flow ratio of monomer vapors. The crosslinked coatings kill bacteria through disruption of the bacteria membrane upon surface contact. Killing efficacy of more than 99.99% against both B. subtilis and E. coli was achieved. The strong antibacterial activity demonstrated by the highly crosslinked coatings indicates that the mobility and length of antibacterial polymer chains are not critical in determining the bactericidal activity of the P(DMAMS-co-EGDA) coatings. Leaching tests indicated that the crosslinked P(DMAMS-co-EGDA) coatings did not leach from the surface to kill bacteria and were stable after the durability tests.Acknowledgements
Financial support for this research was provided by the Oklahoma Center for the Advancement of Science and Technology (Award # ONAP 09-12 and AR 09.02-024). We also thank the Oklahoma State University Microscopy Laboratory for SEM experiments and Dr Astri Wayadande for the use of the fluorescence microscope.References
- K. H. Hong, J. L. Park, I. H. Sul, J. H. Youk and T. J. Kang, J. Polym. Sci., Part B: Polym. Phys., 2006, 44, 2468–2474 CrossRef CAS.
- V. Sambhy, M. M. MacBride, B. R. Peterson and A. Sen, J. Am. Chem. Soc., 2006, 128, 9798–9808 CrossRef CAS.
- W. K. Son, J. H. Youk and W. H. Park, Carbohydr. Polym., 2006, 65, 430–434 CrossRef CAS.
- K. Lewis and A. M. Klibanov, Trends Biotechnol., 2005, 23, 343–348 CrossRef CAS.
- N. M. Milovic, J. Wang, K. Lewis and A. M. Klibanov, Biotechnol. Bioeng., 2005, 90, 715–722 CrossRef CAS.
- S. Borman, Chem. Eng. News, 2002, 80, 36–38.
- S. B. Lee, R. R. Koepsel, S. W. Morley, K. Matyjaszewski, Y. J. Sun and A. J. Russell, Biomacromolecules, 2004, 5, 877–882 CrossRef CAS.
- H. Murata, R. R. Koepsel, K. Matyjaszewski and A. J. Russell, Biomaterials, 2007, 28, 4870–4879 CrossRef CAS.
- S. Lenoir, C. Pagnoulle, C. Detrembleur, M. Galleni and R. Jerome, J. Polym. Sci., Part A: Polym. Chem., 2006, 44, 1214–1224 CrossRef CAS.
- L. Cen, K. G. Neoh and E. T. Kang, Langmuir, 2003, 19, 10295–10303 CrossRef CAS.
- J. C. Tiller, C. J. Liao, K. Lewis and A. M. Klibanov, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 5981–5985 CrossRef CAS.
- S. Lenoir, C. Pagnoulle, C. Detrembleur, M. Galleni and R. Jerome, J. Polym. Sci., Part A: Polym. Chem., 2006, 44, 1214–1224 CrossRef CAS.
- E. Patrucco, S. Ouasti, C. D. Vo, P. De Leonardis, A. Pollicino, S. P. Armes, M. Scandola and N. Tirelli, Biomacromolecules, 2009, 10, 3130–3140 CrossRef CAS.
- S. J. Yuan, S. O. Pehkonen, Y. P. Ting, K. G. Neoh and E. T. Kang, Langmuir, 2010, 26, 6728–6736 CrossRef CAS.
- L. Ferreira and A. Zumbuehl, J. Mater. Chem., 2009, 19, 7796–7806 RSC.
- S. Krishnan, R. J. Ward, A. Hexemer, K. E. Sohn, K. L. Lee, E. R. Angert, D. A. Fischer, E. J. Kramer and C. K. Ober, Langmuir, 2006, 22, 11255–11266 CrossRef CAS.
- A. D. Fuchs and J. C. Tiller, Angew. Chem., Int. Ed., 2006, 45, 6759–6762 CrossRef CAS.
- T. P. Martin, S. E. Kooi, S. H. Chang, K. L. Sedransk and K. K. Gleason, Biomaterials, 2007, 28, 909–915 CrossRef CAS.
- N. Pasquier, H. Keul, E. Heine and M. Moeller, Biomacromolecules, 2007, 8, 2874–2882 CrossRef CAS.
- Y. Mao and K. K. Gleason, Langmuir, 2004, 20, 2484–2488 CrossRef CAS.
- M. A. Gelman, B. Weisblum, D. M. Lynn and S. H. Gellman, Org. Lett., 2004, 6, 557–560 CrossRef CAS.
- D. Roy, J. S. Knapp, J. T. Guthrie and S. Perrier, Biomacromolecules, 2008, 9, 91–99 CrossRef CAS.
- K. Chan and K. K. Gleason, Langmuir, 2005, 21, 8930–8939 CrossRef CAS.
- A. E. Madkour, J. A. Dabkowski, K. Nusslein and G. N. Tew, Langmuir, 2009, 25, 1060–1067 CrossRef CAS.
- L. L. Graham and T. J. Beveridge, J. Bacteriol., 1994, 176, 1413–1421 CAS.
- V. R. F. Matias and T. J. Beveridge, Mol. Microbiol., 2005, 56, 240–251 CrossRef CAS.
- S. O. Meroueh, K. Z. Bencze, D. Hesek, M. Lee, J. F. Fisher, T. L. Stemmler and S. Mobashery, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 4404–4409 CrossRef CAS.
- R. Kugler, O. Bouloussa and F. Rondelez, Microbiology, 2005, 151, 1341–1348 CrossRef CAS.
Footnotes |
| † Electronic supplementary information (ESI) available: SEM images (Fig. S1). See DOI: 10.1039/c0jm02578j |
| ‡ These two authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2011 |