Ionic liquid effects on the activity of monooxygenase P450 BM-3
Kang Lan
Tee
a,
Danilo
Roccatano
a,
Stefan
Stolte
b,
Jürgen
Arning
b,
Bernd
Jastorff
b and
Ulrich
Schwaneberg
*c
aJacobs University, School of Engineering and Science, Campus Ring 8, 28759, Bremen, Germany
bUFT - Center for Environmental Research and Environmental Technology, Leobenerstr., 28359, Bremen, Germany
cAssociate Professor of Biochemical Engineering, Jacobs University Bremen, Campus Ring 1, 28759, Bremen, Germany. E-mail: u.schwaneberg@jacobs-university.de; Fax: +49 (421) 200 3640; Tel: +49 (421) 200 3543
First published on 20th November 2007
Abstract
P450 monooxygenases oxidize a broad range of substrates including fatty acids, alcohols, aliphatic and aromatic hydrocarbons which are often poorly soluble in water and have Km values in the millimolar range. Cosolvent and host–guest chemistry have thus been supplemented to increase catalytic efficiencies. Ionic liquids are alternative solvents and cosolvents in biocatalysis and have received an escalating interest due to unique properties such as tunable hydrophobicity and hydrophilicity. Effects of ten ionic liquids on cytochrome P450 BM-3 activity were investigated by evaluating the influence of hydrophobicity and ion pairs on P450 BM-3. Investigated ionic liquids comprised a range of four anions, four 1-alkyl-3-methylimidazolium derivatives differing in hydrophobic chain length, two altered head groups (pyridinium and pyrrolidinium) and an additional functional group in the side chain (nitrile). Results indicate that varying the chain length of the 1-alkyl-3-methylimidazolium has apparent effects on the resistance of P450 BM-3 (based on EC50), with increasing inhibition in the order of EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (232 ± 8) > BMIM
Cl (232 ± 8) > BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (148 ± 5) > HMIM
Cl (148 ± 5) > HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (32 ± 2) > OMIM
Cl (32 ± 2) > OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (9 ± 1).
Cl (9 ± 1).
Introduction
Ionic liquids have received increasing attention in recent years as alternative solvents in biocatalysis.1–4 These studies were performed on isolated/immobilized enzymes or whole cell systems, in biphasic or homogenous ionic liquid–water mixtures and pure ionic liquids.2,5–7 Stability, activity and enantioselectivity were altered through the use of ionic liquids for enzyme classes such as hydrolases (lipase,5,8 esterase6 and protease9) and oxidoreductases (formate dehydrogenase,10 horseradish peroxidase11 and cytochrome c12). Research on biocatalysis in ionic liquids is driven by solubility properties, low vapor pressure, and high thermal stability of ionic liquids which lead to lower inhalatory exposure and non-flammability. The latter are important considerations in large-scale industrial processes.Cytochrome P450s are oxidoreductases and form a ubiquitous enzyme family which plays key roles in biochemical pathways, xenobiotic metabolism and detoxification. Cytochrome P450s can hydroxylate or epoxidize substrates by inserting a single oxygen molecule from oxygen into an organic compound.13–15 These oxygenation reactions are often difficult to achieve through organic syntheses, especially when regio- or stereoselective oxygenation at ambient temperature, using oxygen as oxidant is required. P450 monooxygenases are attractive industrial biocatalysts due to their selectivity and wide spectrum of substrates accepted (alkanes, alkenes, alcohols, fatty acids, amides, polyaromatic hydrocarbons and heterocyclics).16–19 However hydrophobic substrates of P450s often have Km values in the millimolar range thus requiring the use of organic solvents or cosolvents for efficient biotransformations. P450 enzymes often cannot tolerate required amounts of organic solvents or cosolvents.20,21
P450 BM-3 is one of the most well-studied and industrially important P450 enzymes (structure,22–24catalytic efficiency18,25 and substrate specificity16,26) due to its robustness and design as a single component system (reductase and heme domain being fused on a single polypeptide27). Ionic liquids represent an alternative to solvents and cosolvents for industrial important P450 BM-3 substrates such as fatty acids, alcohols, aliphatic and aromatic hydrocarbons which are known to be totally or partially miscible with ionic liquids.28–30 The influence of ionic liquids has barely been studied for cytochrome P450s, and only one report for a multi-component system (CYP3A4) has been published.31 There have been no reports on the study of ionic liquid effects on enzymatic activity for single component P450 monooxygenases.
In this study, effects of ionic liquids and their hydrophobic side chains on P450 BM-3 activity are quantified. First hypotheses on inhibitory influences are further generated by measuring the OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl resistance of P450 BM-3 mutant F87A, in which the F87A substitution decreases tolerance of P450 BM-3 towards organic solvents.21
Cl resistance of P450 BM-3 mutant F87A, in which the F87A substitution decreases tolerance of P450 BM-3 towards organic solvents.21
Results
The pNCA (p-nitrophenoxycarboxylic acid) assay32 was used to determine the activity of P450 BM-3 under the influence of ionic liquids using p-nitrophenoxydodecanoic acid (12-pNCA) as substrate. The pH remained stable under assay conditions. Concentration range of ten ionic liquids as well as calculated EC50 values are listed in Table 1. The statistical calculations for EC50 were performed using software library drfit for the R language and environment.33 The tolerance of P450 BM-3 towards the investigated ionic liquids varies between EC50 values of 9 and 232 mM. Among the ten ionic liquids, P450 BM-3 has the highest tolerance towards EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (EC50 = 232 ± 8 mM) and the least tolerance towards OMIM
Cl (EC50 = 232 ± 8 mM) and the least tolerance towards OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (EC50 = 9 ± 1 mM).
Cl (EC50 = 9 ± 1 mM).
| Compound | Abbreviation | Concentration range/mM | EC50a/mM |
|---|---|---|---|
| a EC50 values determined based on triplicate measurements (confidence intervals α = 0.025) | |||
| 1-Ethyl-3-methylimidazolium chloride |
EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–420 | 232 ± 8 |
| 1-Ethyl-3-methylimidazolium trifluoromethanesulfonate |
EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3 CF3SO3 |
0–180 | 94 ± 6 |
| 1-Ethyl-3-methylimidazolium trifluoroacetate |
EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO CF3COO |
0–180 | 72 ± 3 |
| 1-Ethyl-3-methylimidazolium tetrafluoroborate |
EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4 BF4 |
0–100 | 42 ± 3 |
| 1-Butylpyridinium chloride |
Bpy![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–264 | 195 ± 9 |
| 1-Cyanomethyl-3-methylimidazolium chloride | 1M11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–360 | 182 ± 8 |
| 1-Butyl-1-methylpyrrolidinium chloride |
BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–300 | 175 ± 8 |
| 1-Butyl-3-methylimidazolium chloride |
BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–240 | 148 ± 5 |
| 1-Hexyl-3-methylimidazolium chloride | HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–72 | 32 ± 2 |
| 1-Octyl-3-methylimidazolium chloride | OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
0–15 | 9 ± 1 |
| Sodium chloride | NaCl | 0–600 | 202 ± 9 |
| Sodium trifluoromethanesulfonate | NaCF3SO3 | 0–300 | 166 ± 8 |
| Sodium trifluoroacetate | NaCF3COO | 0–360 | 162 ± 7 |
| Sodium tetrafluoroborate | NaBF4 | 0–360 | 119 ± 8 |
Effect of the anions and EMIM cation on P450 BM-3 activity
The EC50 values of EMIM with different anions (Cl–, CF3SO3–, CF3COO– and BF4–) show that P450 BM-3 has the lowest tolerance towards EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4. The residual activities for each anion are normalized to the activity when no ionic liquid is added and are shown in Fig. 1. It can be observed from Fig. 1 that EMIM
BF4. The residual activities for each anion are normalized to the activity when no ionic liquid is added and are shown in Fig. 1. It can be observed from Fig. 1 that EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl has a visibly lower inhibitory effect compared to the other EMIM anions. Inhibition is also reflected by the absolute decrease of EC50 from EMIM
Cl has a visibly lower inhibitory effect compared to the other EMIM anions. Inhibition is also reflected by the absolute decrease of EC50 from EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl to EMIM
Cl to EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3, which is at least 2.7-fold higher than any difference between the three other investigated EMIM salts (EMIM
CF3SO3, which is at least 2.7-fold higher than any difference between the three other investigated EMIM salts (EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3, EMIM
CF3SO3, EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO and EMIM
CF3COO and EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4). In fact Fig. 1 shows that the order of inhibition for EMIM
BF4). In fact Fig. 1 shows that the order of inhibition for EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3 and EMIM
CF3SO3 and EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO changes around 150 mM suggesting very similar inhibitions by these two ionic liquids.
CF3COO changes around 150 mM suggesting very similar inhibitions by these two ionic liquids.
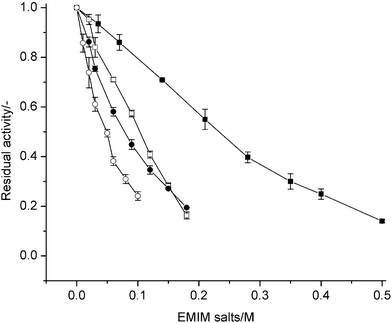 | ||
Fig. 1 Residual activities of P450 BM-3 in presence of EMIM and anions (Cl–, CF3SO3–. CF3COO–, BF4–). Residual activity is given as ratio of initial catalytic turnover number in the presence of ionic liquids to that in the absence of ionic liquids. –■–: EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; –□–: EMIM Cl; –□–: EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3; –●–: EMIM CF3SO3; –●–: EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO; –○–: EMIM CF3COO; –○–: EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4. BF4. | ||
The Na+ salts of the above four anions (Cl–, CF3SO3–, CF3COO– and BF4–) show lower inhibitory influences than the corresponding EMIM salts except NaCl (EC50 value 0.9-fold of the EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl). These results suggest that reduced activity of P450 BM-3 can mainly be attributed to an anion effect. Differences in the extend of inhibition indicate that inhibitory differences between the Na+ and EMIM salts are not solely due to an anion effect, but rather depend on EMIM and Na+ counter ion effects on corresponding anions.
Cl). These results suggest that reduced activity of P450 BM-3 can mainly be attributed to an anion effect. Differences in the extend of inhibition indicate that inhibitory differences between the Na+ and EMIM salts are not solely due to an anion effect, but rather depend on EMIM and Na+ counter ion effects on corresponding anions.
As the chloride anion has the lowest inhibition among the tested anion, it was used as the counter ion for all subsequent tests on ionic liquids with different cations.
Effect of the alkyl-chain length of EMIM cations on P450 BM-3 activity
Effects of alkyl chain length for the imidazolium cation were investigated by using 1-alkyl-3-methylimidazolium chlorides in which the alkyl-chain at one N-atom was stepwise elongated from ethyl- (C2) to octyl- (C8). Fig. 2 shows a decreasing tolerance of P450 BM-3 with stepwise increased alkyl chain length. From the EC50 values in Table 1, the tolerance of P450 BM-3 towards OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl is almost 20-fold lower than EMIM
Cl is almost 20-fold lower than EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl. Fig. 2 and EC50 values in Table 1 show a clear increase in inhibition in accordance to alkyl-chain elongations from C2 to C8.
Cl. Fig. 2 and EC50 values in Table 1 show a clear increase in inhibition in accordance to alkyl-chain elongations from C2 to C8.
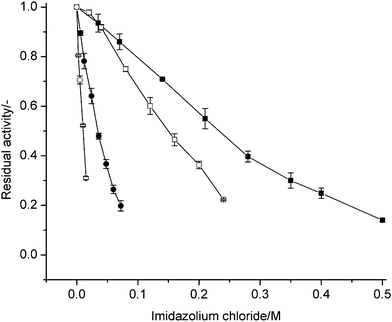 | ||
Fig. 2 Effect of hydrophobic side chain of 1-alkyl-3-methylimidazolium on residual activities of P450 BM-3. Residual activity is given as ratio of initial catalytic turnover number in the presence of ionic liquids to that in the absence of ionic liquids. –■–: EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; –□–: BMIM Cl; –□–: BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; –●–: HMIM Cl; –●–: HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; –○–: OMIM Cl; –○–: OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl. Cl. | ||
Effect of cation aromaticity, size and free electron pair in alkyl-chain of imidazolium cation
Three other cationic structures were investigated: Bpy![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, BMPyr
Cl, BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl and IM11CN
Cl and IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl. The resulted EC50 are in between EMIM
Cl. The resulted EC50 are in between EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl and BMIM
Cl and BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl. Bpy
Cl. Bpy![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl has a pyridinium head group which is larger compared to the imidazolium head group (six- compared to five-membered ring). However this does not seem to have large effects on the inhibition (EC50 ∼1.3-fold of BMIM
Cl has a pyridinium head group which is larger compared to the imidazolium head group (six- compared to five-membered ring). However this does not seem to have large effects on the inhibition (EC50 ∼1.3-fold of BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl) and EC50 remains lower than EMIM
Cl) and EC50 remains lower than EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl. BMPyr
Cl. BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl has a side chain on its cation which is the same as BMIM, but unlike the imidazolium (BMIM) head group, the pyrrolidinium (BMPyrH) head group does not have an aromatic ring system. Aromaticity however has only a minor effect on the inhibition. The BMPyr
Cl has a side chain on its cation which is the same as BMIM, but unlike the imidazolium (BMIM) head group, the pyrrolidinium (BMPyrH) head group does not have an aromatic ring system. Aromaticity however has only a minor effect on the inhibition. The BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl EC50 value is ∼1.2-fold higher than that of BMIM
Cl EC50 value is ∼1.2-fold higher than that of BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (Table 1). The side chain carrying the nitrile functional group on IM11CN
Cl (Table 1). The side chain carrying the nitrile functional group on IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl has a chain length that is in between EMIM and BMIM. Differing from the alkyl imidazolium side chains, a polar nitrile functional group was introduced. Possible electrostatic interactions have however no notable influence on P450 BM-3 inhibition (EC50 for IM11CN
Cl has a chain length that is in between EMIM and BMIM. Differing from the alkyl imidazolium side chains, a polar nitrile functional group was introduced. Possible electrostatic interactions have however no notable influence on P450 BM-3 inhibition (EC50 for IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl lies between EMIM
Cl lies between EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl and BMIM
Cl and BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl).
Cl).
k cat and Km values for P450 BM-3 in ionic liquid–water mixture
As seen in Table 1 and Fig. 2, the effect of the length of side chain has an apparent influence on the inhibition of P450 BM-3. For further insight, the kinetic parameters of the P450 BM-3 hydroxylation of 12-pNCA were determined in the presence of the imidazolium chlorides (Table 2). For each of the 1-alkyl-3-methylimidazolium chlorides (BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, HMIM
Cl, HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl and OMIM
Cl and OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl), the amount of ionic liquid equivalent to the EC50 concentration was employed. In all three ionic liquids a lower kcat and a higher Km was obtained which led to a 4.0–4.6-fold lower catalytic efficiency (kcat/Km) when compared to the P450 BM-3 activity in no ionic liquid. In detail, the kcat was reduced to 0.50–0.62-fold while the Km increased 2.0–2.6-fold. Kinetic constants (kcat, Km, Ki) were calculated by assuming mixed inhibition kinetics of P450 BM-3 (Table 2). Table 2 shows that Ki decreases 18-fold as the side chain length increases from BMIM to OMIM. Changes in Ki suggest that the increasing inhibition as side chain length increases can be attributed to a stronger association of the ionic liquid to P450 BM-3. Increasing from BMIM
Cl), the amount of ionic liquid equivalent to the EC50 concentration was employed. In all three ionic liquids a lower kcat and a higher Km was obtained which led to a 4.0–4.6-fold lower catalytic efficiency (kcat/Km) when compared to the P450 BM-3 activity in no ionic liquid. In detail, the kcat was reduced to 0.50–0.62-fold while the Km increased 2.0–2.6-fold. Kinetic constants (kcat, Km, Ki) were calculated by assuming mixed inhibition kinetics of P450 BM-3 (Table 2). Table 2 shows that Ki decreases 18-fold as the side chain length increases from BMIM to OMIM. Changes in Ki suggest that the increasing inhibition as side chain length increases can be attributed to a stronger association of the ionic liquid to P450 BM-3. Increasing from BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl to HMIM
Cl to HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl shows largest decrease in Ki (5.6-fold) and EC50 values (4.6-fold) when comparing 1-alkyl-3-methylimidazolium from C4 to C8.
Cl shows largest decrease in Ki (5.6-fold) and EC50 values (4.6-fold) when comparing 1-alkyl-3-methylimidazolium from C4 to C8.
| Mutant F87A | |||||||
|---|---|---|---|---|---|---|---|
| a k cat and Km ratio is the ratio of kinetic parameters in the presence of ionic liquids to that in the absence of ionic liquids. b K i = dissociation constant of inhibitor to enzyme. | |||||||
| No inhibitor | 0 | 404.8 ± 5.7 | 7.2 ± 0.4 | 56.2 | — | — | — |
OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl Cl |
9 | 31.4 ± 1.8 | 20.2 ± 2.8 | 1.6 | 0.08 | 2.8 | 0.3 |
Role of amino acid position 87 in P450 BM-3 ionic liquid tolerance
The amino acid residue phenylalanine 87 is important for the organic solvent resistance of P450 BM-3. Mutating this residue to alanine resulted in a significant drop in resistance of P450 BM-3 towards DMSO, THF, ethanol and acetonitrile.21 The catalytic efficiency of P450 BM-3 F87A mutant in the presence of OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl decreased 35-fold compared to when no ionic liquid was added, which is significantly larger than the 4.3-fold decrease observed in wildtype P450 BM-3 (see Table 2). The change in Km is of the same order of magnitude between the wildtype and F87A mutant while the major contributor of this drastic difference in catalytic efficiency is the 12.5-fold decrease in kcat in F87A compared to the 1.6-fold decrease seen in the wildtype. A lower Ki is also observed suggesting better association of OMIM
Cl decreased 35-fold compared to when no ionic liquid was added, which is significantly larger than the 4.3-fold decrease observed in wildtype P450 BM-3 (see Table 2). The change in Km is of the same order of magnitude between the wildtype and F87A mutant while the major contributor of this drastic difference in catalytic efficiency is the 12.5-fold decrease in kcat in F87A compared to the 1.6-fold decrease seen in the wildtype. A lower Ki is also observed suggesting better association of OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl in F87A mutant compared to the wildtype.
Cl in F87A mutant compared to the wildtype.
NADPH consumption and P450 spectra using ionic liquids as substrate
The NADPH consumption assay was used to investigate the possibility of the ionic liquids being substrates of P450 BM-3, particularly the long chain 1-alkyl-3-methylimidazolium. Except for IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, no NADPH consumption higher than the negative control was observed. For IM11CN
Cl, no NADPH consumption higher than the negative control was observed. For IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, a NADPH consumption of <12% of employed NADPH was detected after 20 min in absence of any other substrate. All employed ionic liquids except IM11CN
Cl, a NADPH consumption of <12% of employed NADPH was detected after 20 min in absence of any other substrate. All employed ionic liquids except IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl are therefore not or very poor substrates of P450 BM-3. NADPH consumption assay also shows that OMIM
Cl are therefore not or very poor substrates of P450 BM-3. NADPH consumption assay also shows that OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl is not a substrate of the F87A variant. Spectral measurements of P450 BM-3 in the presence of 1-alkyl-3-methylimidazoliums (BMIM
Cl is not a substrate of the F87A variant. Spectral measurements of P450 BM-3 in the presence of 1-alkyl-3-methylimidazoliums (BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, HMIM
Cl, HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, OMIM
Cl, OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl) did not resulted in any spectral changes (absorbance decrease around 420 nm and increase around 390 nm). These spectral changes are characteristic of substrate binding in P450 BM-3 which leads to perturbation of the sixth water ligand from the heme iron center. The iron changes thereby from a low spin to a high spin configuration that subsequently leads to an increase in the heme iron reduction potential, a pre-requisite for the electron transfer from the redox partner. All experimental results suggest a lack of inhibitor coordination to the heme iron as reported for the cosolvent DMSO.22
Cl) did not resulted in any spectral changes (absorbance decrease around 420 nm and increase around 390 nm). These spectral changes are characteristic of substrate binding in P450 BM-3 which leads to perturbation of the sixth water ligand from the heme iron center. The iron changes thereby from a low spin to a high spin configuration that subsequently leads to an increase in the heme iron reduction potential, a pre-requisite for the electron transfer from the redox partner. All experimental results suggest a lack of inhibitor coordination to the heme iron as reported for the cosolvent DMSO.22
Discussion
Principles of structure–function relationships of enzymes and biological systems in ionic liquids have been discovered and quantified.34–36 There has been so far only one study on the effects of ionic liquids on cytochrome P450 activities31 and this report does not evaluate the potential use of ionic liquids for industrial biotransformations. In this manuscript, we have performed a systematic study to quantify ionic liquid effects on P450 BM-3 activity. In detail we studied the influence of four anions and EMIM cation, the effect of alkyl-chain length of 1-alkyl-3-methylimidazolium cations, cation aromaticity and size as well as an additional free electron pair in the alkyl-chain of imidazolium cation on P450 BM-3 activity. These influences of the P450 BM-3 activity were quantified by determining the EC50 values, kcat, Km and Ki using the pNCA activity assay.32 Based on these results the EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl (EC50 = 232 ± 8 mM) is a preferred candidate for biotransformations in ionic liquids.
Cl (EC50 = 232 ± 8 mM) is a preferred candidate for biotransformations in ionic liquids.
EC50 values are at least an order of magnitude higher for P450 BM-3 wildtype when compared to studies on acetylcholinesterase.34 When compared to hydrolases P450 BM-3 resistance to ionic liquids is however low. Hydrolases even function in pure ionic liquid solutions.5,6,37 The Cl– anion has a significantly lower inhibitory effect than CF3SO3–, CF3COO– and BF4– in both cations (EMIM, Na+; see Table 1 and Fig. 1). For the cations (EMIM, Na+), no general anion trends could be observed. The EMIM and Na+ cations however have similar inhibitory effects when Cl– ion is used as counter anion (Table 1 and Fig. 1). One possible explanation of the different anion effects is the Hofmeister series which suggests that kosmotropic anions stabilize enzymes while chaotropic anions destabilize biocatalysts. According to the Hofmeister series, chaotropicity increases in the order of CF3COO– > Cl– > BF4–.38,39 The observed inhibition does show that the most chaotropic anion BF4– is most inhibitory for P450 BM-3, while CF3COO– and Cl– behave differently from the Hofmeister series. Such exceptions are also reported for bovine carbonic anhydrase, succinate dehydrogenase and NADH oxidase.38
The most significant activity effect due to changes in the cation was seen when the alkyl chain length of the 1-alkyl-3-methylimidazolium cation was increased, which has also been previously observed in acetylcholinesterase.34 Additionally, a decreased enantioselectivity of a protease from Bacillus licheniformis had been reported with increasing cation side-chain length (EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO > BMIM
CF3COO > BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO > HMIM
CF3COO > HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO).40 It had been suggested that increased destabilization with increased alkyl chain length follows the Hofmeister series where longer side-chain length increases hydrophobicity and kosmotropicity of cations.39 The Hofmeister series remains the best possible postulation for enzyme destabilization by cations but it is difficult to generalize for all the enzyme systems that have been studied so far (lipase, esterase, dehydrogenase and P450s).38 For example the kinetic resolution of 1-phenylethanol by Candida antarcticalipase A and Candida antarcticalipase B improves when the 1-alkyl-3-methyllimidazolium cation chain length increases from BMIM
CF3COO).40 It had been suggested that increased destabilization with increased alkyl chain length follows the Hofmeister series where longer side-chain length increases hydrophobicity and kosmotropicity of cations.39 The Hofmeister series remains the best possible postulation for enzyme destabilization by cations but it is difficult to generalize for all the enzyme systems that have been studied so far (lipase, esterase, dehydrogenase and P450s).38 For example the kinetic resolution of 1-phenylethanol by Candida antarcticalipase A and Candida antarcticalipase B improves when the 1-alkyl-3-methyllimidazolium cation chain length increases from BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4 to HMIM
BF4 to HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4 and OMIM
BF4 and OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4.41
BF4.41
Increasing the alkyl side chain from ethyl- to octyl- leads to more than 25-fold decrease in the EC50 value for P450 BM-3 wildtype. Table 2 summarizes the kinetic parameters (kcat, Km, Ki) of BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, HMIM
Cl, HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl and OMIM
Cl and OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl. Analysis of the P450 BM-3 structure shows a substrate binding pocket close to heme that is lined by hydrophobic residues (F42, Y51, L181, M185, L188, A328, A330 and M354).42 The increased inhibition can be attributed to the extended hydrophobicity of longer side chains which cause stronger associations with the hydrophobic patches in the P450 BM-3 substrate channel. Consequently the Km increases as substrate access is adversely affected due to competition from the 1-alkyl-3-methylimidazolium cation. Changing the cation to pyrrolidinium or addition of a nitrile functional group (Table 1 and 2) does not impact significantly on the EC50 values (Table 1 and 2).
Cl. Analysis of the P450 BM-3 structure shows a substrate binding pocket close to heme that is lined by hydrophobic residues (F42, Y51, L181, M185, L188, A328, A330 and M354).42 The increased inhibition can be attributed to the extended hydrophobicity of longer side chains which cause stronger associations with the hydrophobic patches in the P450 BM-3 substrate channel. Consequently the Km increases as substrate access is adversely affected due to competition from the 1-alkyl-3-methylimidazolium cation. Changing the cation to pyrrolidinium or addition of a nitrile functional group (Table 1 and 2) does not impact significantly on the EC50 values (Table 1 and 2).
As the full crystal structure of P450 BM-3 has not been resolved, one can only base hypotheses on the crystallized heme domain of P450 BM-3. One possible explanation of the lower Km and kcat of P450 BM-3 is the binding of 1-alkyl-3-methylimidazolium cation at the entrance of the substrate access channel. To obtain further insight, short molecular dynamics (MD) simulation of BMIM, HMIM and OMIM at the entrance of the substrate channel and inside the channel were performed. For the MD at the substrate channel entrance, the final conformations show the head group pointing towards Glu13, likely due to electrostatic interaction; the hydrophobic tail points into the substrate channel (see Fig. 3). Glu13 lies within the hydrophobic patch (P11, L14, L17, P18, P45 and A191) which is believed to be a site important for initial binding of lipophilic substrates.42 The presence of the 1-alkyl-3-methylimidazolium cation may interfere with this initial binding thereby affecting substrate binding and leading to higher Km values (Table 2). Despite the substrate channel being hydrophobic and having a positively charged R47 which coordinates the carboxylate terminal of the fatty acid substrates, it is known that positively charged substrates can enter and be converted by P450 BM-3.43 MD of the positively charged 1-alkyl-3-methylimidazoliums inside the substrate channel were performed and the positively charged head group moves away from Arg47 and orients towards Glu435 with the hydrophobic tail pointing towards the heme (see Fig. 3). OMIM, which has the longest hydrophobic tail of the investigated ionic liquids, is located ∼5 Å from the Fe iron, 9 Å from Y51 and 13 Å from R47 according to the MD simulations. The 1-alkyl-3-methylimidazoliums in the substrate channel in such a configuration can interfere with the orientation of the substrate with respect to the heme, thereby affecting the terminal hydroxylation of 12-pNCA and lead to the lower kcat values.
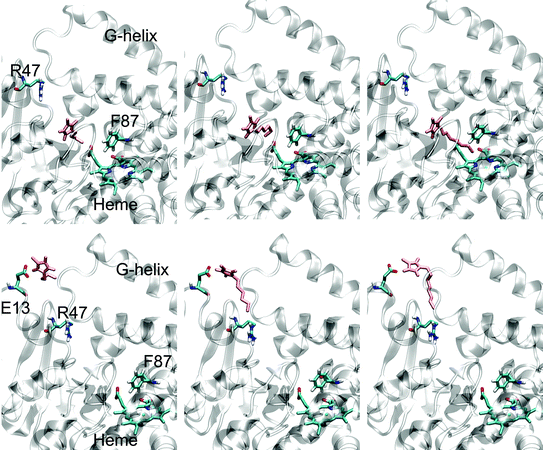 | ||
| Fig. 3 Final conformations from molecular dynamics docking simulations. Top row (substrate channel entrance; from left to right): BMIM, HMIM, OMIM; Bottom row (inside substrate binding channel; from left to right): BMIM, HMIM, OMIM. Figures on the left are labeled by highlighting important P450 BM-3 positions/regions. The 1-alkyl-3-methylimidazolium is colored in pink. | ||
The mutation F87A is known to influence P450 BM-3's tolerance towards organic solvents.21 The OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl resistance of P450 BM-3 F87A was thus determined. The change in phenylalanine 87 to alanine led to a lower resistance of P450 BM-3 to OMIM
Cl resistance of P450 BM-3 F87A was thus determined. The change in phenylalanine 87 to alanine led to a lower resistance of P450 BM-3 to OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl as reflected in the 35-fold decrease in kcat/Km compared to the 4-fold decrease observed in P450 BM-3 wildtype (Table 2). The change in kcat/Km is contributed largely by the 12.5-fold lower kcat value (Table 2). The presence of phenylalanine 87 at the end of channel can possibly interact further with the p-nitrophenyl group of the 12-pNCA substrate through π–π interaction and contribute to a catalytic “active” orientation of 12-pNCA, especially if the OMIM inside the substrate channel affects substrate orientation. Changing F87 to alanine removes this interaction and leads to a much lower kcat value. More extensive studies are currently being performed to elucidate the actual mode of inhibition by the 1-alkyl-3-methylimidazoliums.
Cl as reflected in the 35-fold decrease in kcat/Km compared to the 4-fold decrease observed in P450 BM-3 wildtype (Table 2). The change in kcat/Km is contributed largely by the 12.5-fold lower kcat value (Table 2). The presence of phenylalanine 87 at the end of channel can possibly interact further with the p-nitrophenyl group of the 12-pNCA substrate through π–π interaction and contribute to a catalytic “active” orientation of 12-pNCA, especially if the OMIM inside the substrate channel affects substrate orientation. Changing F87 to alanine removes this interaction and leads to a much lower kcat value. More extensive studies are currently being performed to elucidate the actual mode of inhibition by the 1-alkyl-3-methylimidazoliums.
Conclusion
EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl is identified as a preferred ionic liquid as cosolvent for biotransformations catalyzed by P450 BM-3. Increasing alkyl side chain from ethyl- to octyl- in the imidazolium cation significantly increases (26-fold) the inhibitory effect of ionic liquids which can likely be attributed to increase hydrophobicity of the alkyl chain. Potential binding sites lie at the hydrophobic mouth of the substrate access channel (OMIM head group pointing towards Glu13) and inside the substrate channel (OMIM head group pointing towards Glu435). P450 BM-3 shows, based on the EC50 values, a lower resistance to ionic liquids compared to previously studied organic solvents such as DMSO (50% residual activity at ∼3 M), acetonitrile (50% residual activity at ∼1 M) and ethanol (50% residual activity at ∼1 M).21 However for EMIM
Cl is identified as a preferred ionic liquid as cosolvent for biotransformations catalyzed by P450 BM-3. Increasing alkyl side chain from ethyl- to octyl- in the imidazolium cation significantly increases (26-fold) the inhibitory effect of ionic liquids which can likely be attributed to increase hydrophobicity of the alkyl chain. Potential binding sites lie at the hydrophobic mouth of the substrate access channel (OMIM head group pointing towards Glu13) and inside the substrate channel (OMIM head group pointing towards Glu435). P450 BM-3 shows, based on the EC50 values, a lower resistance to ionic liquids compared to previously studied organic solvents such as DMSO (50% residual activity at ∼3 M), acetonitrile (50% residual activity at ∼1 M) and ethanol (50% residual activity at ∼1 M).21 However for EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, IM11CN
Cl, IM11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, Bpy Cl and BMPyr
Cl, Bpy Cl and BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl, higher resistances were observed compared to THF (50% residual activity at ∼0.1 M). Changing of the cation to pyridinium and pyrrolidinium or addition of the nitrile functional group did not generate obvious changes in EC50.
Cl, higher resistances were observed compared to THF (50% residual activity at ∼0.1 M). Changing of the cation to pyridinium and pyrrolidinium or addition of the nitrile functional group did not generate obvious changes in EC50.
Directed P450 BM-3 evolution studies employing EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl are currently being performed for discovering principles on how P450 BM-3 and monooxygenases in general can be reengineered to operate efficiently in EMIM
Cl are currently being performed for discovering principles on how P450 BM-3 and monooxygenases in general can be reengineered to operate efficiently in EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl.
Cl.
Experimental
Materials
All chemicals used were of analytical reagent grade or higher quality and purchased from Sigma-Aldrich Chemie (Taufkirchen, Germany), Applichem (Darmstadt, Germany) and Carl Roth (Karlsruhe, Germany). NADPH tetrasodium salt was purchased from Codexis (Redwood city, CA, United States), p-nitrophenoxydodecanoic acid (12-pNCA) was synthesized as previously described32 and the ionic liquids used were from Merck KGaA (Darmstadt, Germany).P450 BM-3 expression and purification
P450 BM-3 wildtype and mutants were expressed in 1-L Erlenmeyer flask containing 250 mL of terrific broth (100 mg L–1ampicillin, 250 µL of trace elements as previously described21). The cells were harvested by centrifugation (4 °C, 3220 g, 15 min) and the cell pellet was resuspended in KxH3–xPO4 (25 mM, pH 7.5) and lysed by high-pressure homogenization (EmulsiFlex-C3, Avestin Inc, Ottawa, Canada; two cycles of homogenizing pressure 1500–1800 bar). Cell lysate was centrifuged (20000 g, 45 min) and the supernatant was further cleared through a 0.45 µm filter (17765 K, Sartorius, Hannover, Germany). The filtrate was loaded on a DEAE650s anion exchanger column (Tosoh Bioscience, Stuttgart, Germany) and purified as previously described.44 P450 BM-3 concentrations were measured by CO-difference spectra as reported by Omura and Sato using ε = 91 mM–1 cm–1.45Determination of pH in ionic liquid–water mixture
The pH of the reaction mixture used for activity tests was determined using a laboratory pH meter (Knick, Berlin, Germany). A reaction mixture (2.5 mL) consist of 1.5 mL buffer (0.3 M Tris pH 8.2), 0.3 mL ionic liquid (0.5 M EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.18 M EMIM
Cl; 0.18 M EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3 and EMIM
CF3SO3 and EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO; 0.096 M EMIM
CF3COO; 0.096 M EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4; 0.3 M BMPyr
BF4; 0.3 M BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.36 M 1M11CNCl; 0.24 M BMIM
Cl; 0.36 M 1M11CNCl; 0.24 M BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.072 M HMIM
Cl; 0.072 M HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.015 M OMIM
Cl; 0.015 M OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl), 75 µM 12-pNCA and 0.5% v/v DMSO. The pH was equilibrated (10 min) or until a stable reading was obtained.
Cl), 75 µM 12-pNCA and 0.5% v/v DMSO. The pH was equilibrated (10 min) or until a stable reading was obtained.
Determination of NADPH consumption and P450 spectra using ionic liquids as substrate
For NADPH consumption assay, 600 µL tris buffer (0.3 M, pH 8.2), 0.0975 nmol P450 BM-3 WT and 10 µL of ionic liquids (0.175 M EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.075 M EMIM
Cl; 0.075 M EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3SO3 and EMIM
CF3SO3 and EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CF3COO; 0.04 M EMIM
CF3COO; 0.04 M EMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BF4; 0.15 M 1M11CN
BF4; 0.15 M 1M11CN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.125 M BMPyr
Cl; 0.125 M BMPyr![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.1 M BMIM
Cl; 0.1 M BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.03 M HMIM
Cl; 0.03 M HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 0.0125 M OMIM
Cl; 0.0125 M OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl) were used. The reaction mixture was topped up by supplementing doubly-distilled H2O (900 µL), incubated (5 min) and initiated by adding NADPH (0.25 mM; final reaction volume 1 mL). NADPH consumption was followed spectrophotometrically (340 nm, 10 min; Specord200, Analytik Jena AG, Jena, Germany) in disposable cuvettes. A positive control was performed by adding 12-pNCA (75 µM) and no ionic liquids to the reaction mixture; a negative control was performed in the absence of 12-pNCA substrates. For spectra measurements, 0.2 nmol of P450 BM-3 were added to phosphate buffer (25 mM, pH 7.5) with ionic liquids (148 mM BMIM
Cl) were used. The reaction mixture was topped up by supplementing doubly-distilled H2O (900 µL), incubated (5 min) and initiated by adding NADPH (0.25 mM; final reaction volume 1 mL). NADPH consumption was followed spectrophotometrically (340 nm, 10 min; Specord200, Analytik Jena AG, Jena, Germany) in disposable cuvettes. A positive control was performed by adding 12-pNCA (75 µM) and no ionic liquids to the reaction mixture; a negative control was performed in the absence of 12-pNCA substrates. For spectra measurements, 0.2 nmol of P450 BM-3 were added to phosphate buffer (25 mM, pH 7.5) with ionic liquids (148 mM BMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 32 mM HMIM
Cl; 32 mM HMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl; 9 mM OMIM
Cl; 9 mM OMIM![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cl) and incubated for 5 min. Subsequently spectra measurements between 350 and 700 nm were recorded using a Specord200 (Analytik Jena AG).
Cl) and incubated for 5 min. Subsequently spectra measurements between 350 and 700 nm were recorded using a Specord200 (Analytik Jena AG).
Determination of P450 BM-3 activity in ionic liquid–water mixture
Residual activities of P450 BM-3 WT in the presence of ionic liquids were measured in 96-well microtiter plates (Grenier Bio-One GmbH, Frickenhausen, Germany) and ionic liquids were always prepared freshly in doubly-distilled H2O. The 200 µL reaction mixture finally consists of 0.18 M Tris buffer (pH 8.2), gradient concentrations of ionic liquids, 75 µM 12-pNCA, 0.5% DMSO and 0.0195 nmol P450 BM-3 WT. Employed concentrations of ionic liquids and the Na+ salts used for these activity measurements are listed in Table 1. Reaction mixtures were always incubated for 5 min before 12-pNCA conversion was initiated by adding NADPH (25 µL, 2.5 mM). Conversion of the 12-pNCA was monitored using the Tecan Sunrise microtiter plate reader (Tecan, Zurich, Switzerland) at 405 nm for a duration of 6–10 min with an average of 3 s interval between each reading cycle. All P450 BM-3 residual activities in ionic liquid–water mixture were determined with at least triplicate measurements.Determination of kcat and Km values for P450 BM-3 in ionic liquid–water mixture
The kinetic constants were determined using the 12-pNCA assay. The assay was performed in disposable cuvettes as previously described32 except that the tris buffer composition was changed (0.18 M, pH 8.2, final concentration of DMSO is 0.5%). Amount of P450 BM-3 per reaction was always 0.1 nmol. The activity was monitored at 410 nm using Specord 200 (Analytik Jena AG). 12-pNCA concentrations were varied from 2.5–125 µM. All measurements were made in triplicates and the kcat and Km values were determined using GraphPad Prism software (GraphPad Software, San Diego, CA, USA) with fitting to a one-site binding hyperbola equation using non-linear regression.Imidazolium cation docking in the substrate binding site of P450 BM-3
For MD simulations the atomic coordinates of P450 BM-3 wildtype were obtained using the crystal structure file (pdb code 1bu7).24 All crystallographic water and other ligand molecules were removed except the water molecule coordinating the iron in the active site of the enzyme. Polar and aromatic hydrogen atoms were added to the crystal structure. Atomic partial charges for the protein were assigned from the GROMOS96 43b1 force field.46 The initial structure of the three ionic liquids were optimized at HF level using 6-31G** basis set using Gaussian03 program (www.gaussian.com). The force field parameters for the ionic liquid were taken from published data.47 Initial docked conformations were manually generated using PBD Viewer. The initial docked conformation of 1-alkyl-3-methylimidazolium at the substrate channel entrance was placed between P45 and A191,48,49 while initial docked conformation inside the channel was selected according to the palmitoleic acid substrate (pdb code 1fag)23 close to R47 and Y51. Optimization of the docked structures was performed using standard energy minimization followed by a molecular dynamics simulation of at least 30 ps using position restraints on the protein backbone. All the simulation were performed using the program GROMACS (version 3.1) (www.gromacs.org). Final conformations were used to analyse the cation position.Notes and references
- S. Park and R. J. Kazlauskas, Curr. Opin. Biotechnol., 2003, 14, 432–437 CrossRef CAS.
- R. A. Sheldon, R. M. Lau, M. J. Sorgedrager, F. van Rantwijk and K. R. Seddon, Green Chem., 2002, 4, 147–151 RSC.
- F. van Rantwijk, R. M. Lau and R. A. Sheldon, Trends Biotechnol., 2003, 21, 131–138 CrossRef CAS.
- Z. Yang and W. B. Pan, Enzyme Microb. Technol., 2005, 37, 19–28 CrossRef CAS.
- J. L. Kaar, A. M. Jesionowski, J. A. Berberich, R. Moulton and A. J. Russell, J. Am. Chem. Soc., 2003, 125, 4125–4131 CrossRef CAS.
- M. Persson and U. T. Bornscheuer, J. Mol. Catal. B: Enzym., 2003, 22, 21–27 CrossRef CAS.
- H. Pfruender, R. Jones and D. Weuster-Botz, J. Biotechnol., 2006, 124, 182–190 CrossRef CAS.
- S. J. Nara, J. R. Harjani and M. M. Salunkhe, Tetrahedron Lett., 2002, 43, 2979–2982 CrossRef CAS.
- H. Zhao, L. Jackson, Z. Y. Song and A. Olubajo, Tetrahedron: Asymmetry, 2006, 17, 2491–2498 CrossRef CAS.
- N. Kaftzik, P. Wasserscheid and U. Kragl, Org. Process Res. Dev., 2002, 6, 553–557 Search PubMed.
- S. Sgalla, G. Fabrizi, S. Cacchi, A. Macone, A. Bonamore and A. Boffi, J. Mol. Catal. B: Enzym., 2007, 44, 144–148 CrossRef CAS.
- J. A. Laszlo and D. L. Compton, J. Mol. Catal. B: Enzym., 2002, 18, 109–120 CrossRef CAS.
- A. Chefson and K. Auclair, Mol. Biosyst., 2006, 2, 462–469 RSC.
- A. J. Fulco, Ann. Rev. Pharm. Toxicol., 1991, 31, 177–203 CrossRef CAS.
- D. F. V. Lewis and A. Wiseman, Enzyme Microb. Technol., 2005, 36, 377–384 CrossRef CAS.
- D. Appel, S. Lutz-Wahl, P. Fischer, U. Schwaneberg and R. D. Schmid, J. Biotechnol., 2001, 88, 167–171 CrossRef CAS.
- A. B. Carmichael and L. L. Wong, Eur. J. Biochem., 2001, 268, 3117–3125 CrossRef CAS.
- A. Glieder, E. T. Farinas and F. H. Arnold, Nat. Biotechnol., 2002, 20, 1135–1139 CrossRef CAS.
- S. Graham-Lorence, G. Truan, J. A. Peterson, J. R. Falck, S. Z. Wei, C. Helvig and J. H. Capdevila, J. Biol. Chem., 1997, 272, 1127–1135 CrossRef CAS.
- S. Kumar, L. Sun, B. K. Muralidhara, J. R. Halpert and J. R. Halpert, Protein Eng. Des. Sel., 2006, 19, 547–554 Search PubMed.
- T. S. Wong, F. H. Arnold and U. Schwaneberg, Biotechnol. Bioeng., 2004, 85, 351–358 CrossRef.
- J. Kuper, T. S. Wong, D. Roccatano, M. Wilmanns and U. Schwaneberg, J. Am. Chem. Soc., 2007, 129, 5786–5787 CrossRef CAS.
- H. Y. Li and T. L. Poulos, Nat. Struct. Biol., 1997, 4, 140–146 CrossRef CAS.
- I. F. Sevrioukova, H. Y. Li, H. Zhang, J. A. Peterson and T. L. Poulos, Proc. Natl. Acad. Sci. U. S. A., 1999, 96, 1863–1868 CrossRef CAS.
- K. L. Tee and U. Schwaneberg, Angew. Chem., Int. Ed., 2006, 45, 5380–5383 CrossRef CAS.
- T. S. Wong, N. Wu, D. Roccatano, M. Zacharias and U. Schwaneberg, J. Biomol. Screen., 2005, 10, 246–252 CrossRef CAS.
- L. O. Narhi and A. J. Fulco, J. Biol. Chem., 1987, 262, 6683–6690 CAS.
- U. Domanska and A. Marciniak, J. Chem. Thermodyn., 2005, 37, 577–585 CrossRef CAS.
- U. Domanska and A. Marciniak, Fluid Phase Equilib., 2006, 260, 9–18.
- S. Park, F. Viklund, K. Hult and R. J. Kazlauskas, Green Chem., 2003, 5, 715–719 RSC.
- A. Chefson and K. Auclair, ChemBioChem, 2007, 8, 1189–1197 CrossRef CAS.
- U. Schwaneberg, C. Schmidt-Dannert, J. Schmitt and R. D. Schmid, Anal. Biochem., 1999, 269, 359–366 CrossRef CAS.
- J. Ranke, Drfit: Dose–response data evaluation, 2005 Search PubMed (computer program).
- F. Stock, J. Hoffmann, J. Ranke, R. Stormann, B. Ondruschka and B. Jastorff, Green Chem., 2004, 6, 286–290 RSC.
- S. Stolte, J. Arning, U. Bottin-Weber, M. Matzke, F. Stock, K. Thiele, M. Uerdingen, U. Welz-Biermann, B. Jastorff and J. Ranke, Green Chem., 2006, 8, 621–629 RSC.
- S. Stolte, J. Arning, U. Bottin-Weber, A. Müller, W. R. Pitner, U. Welz-Biermann, B. Jastorff and J. Ranke, Green Chem., 2007, 9, 760–767 RSC.
- R. M. Lau, F. van Rantwijk, K. R. Seddon and R. A. Sheldon, Org. Lett., 2000, 2, 4189–4191 CrossRef CAS.
- H. Zhao, J. Mol. Catal. B: Enzym., 2005, 37, 16–25 CrossRef CAS.
- H. Zhao, O. Olubajo, Z. Y. Song, A. L. Sims, T. E. Person, R. A. Lawal and L. A. Holley, Bioorg. Chem., 2006, 34, 15–25 CrossRef CAS.
- H. Zhao, S. M. Campbell, L. Jackson, Z. Y. Song and O. Olubajo, Tetrahedron: Asymmetry, 2006, 17, 377–383 CrossRef CAS.
- S. H. Schofer, N. Kaftzik, P. Wasserscheid and U. Kragl, Chem. Commun., 2001, 425–426 RSC.
- K. G. Ravichandran, S. S. Boddupalli, C. A. Hasemann, J. A. Peterson and J. Deisenhofer, Science, 1993, 261, 731–736 CrossRef CAS.
- C. F. Oliver, S. Modi, W. U. Primrose, L. Y. Lian and G. C. K. Roberts, Biochem. J., 1997, 327, 537–544 CAS.
- U. Schwaneberg, A. Sprauer, C. Schmidt-Dannert and R. D. Schmid, J. Chromatogr., A, 1999, 848, 149–159 CrossRef CAS.
- T. Omura and R. Sato, J. Biol. Chem., 1964, 239, 2370–2378 CAS.
- W. F. van Gunsteren, S. R. Billeter, A. A. Eising, P. H. Hünenberger, P. Krüger, A. E. Mark, W. R. P. Scott and I. G. Tironi, in Biomolecular Simulation: The GROMOS96 Manual and User Guide, vdf Hochschulverlang, ETH Zürich, Switzerland, 1996 Search PubMed.
- N. M. Micaelo, A. M. Baptista and C. M. Soares, J. Phys. Chem. B, 2006, 110, 14444–14451 CrossRef CAS.
- M. D. Paulsen and R. L. Ornstein, Proteins: Struct., Funct., Genet., 1995, 21, 237–243 CAS.
- D. Roccatano, T. S. Wong, U. Schwaneberg and M. Zacharias, Biopolymers, 2006, 83, 467–476 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2008 |
