Steric control of 4-connected network topology in hydrogen bonded coordination polymers†
David R.
Turner
a,
Joshua
Strachan-Hatton
ab and
Stuart R.
Batten
*a
aSchool of Chemistry, Monash University, Clayton, Vic 3800, Australia. E-mail: stuart.batten@sci.monash.edu.au; Fax: +61(0)3 9905 4597; Tel: +61(0)3 9905 4606
bForest Hill College, 178-180 Mahoneys Road, Burwood East, Vic 3151, Australia
First published on 13th September 2007
Abstract
Two 1D coordination polymers of copper(II) acetylenedicarboxylate, [Cu(O2CCCCO2)(C5H5N)2(MeOH)] (1) and [Cu(O2CCCCO2)(Me2C5H3N)2(MeOH)]·MeOH (2), have been synthesised and structurally characterised by single crystal X-ray diffraction. In both cases the square pyramidal CuII atoms have trans coordinated pyridyl ligands and trans η1 carboxylates in the equatorial sites with methanol occupying the apical position. The linear dicarboxylate ligand results in the formation of near linear coordination polymers. The polymeric chains in 1 form 2D hydrogen bonded (4,4) sheets whereas in 2 this topology is not possible, due to the extra steric bulk of the pyridine methyl groups, and a 3D hydrogen bonded network with CdSO4 (cds) topology is formed in which the lattice methanol molecules are also incorporated into the hydrogen bonding bridges.
Introduction
The deliberate design and synthesis of coordination and hydrogen bonded networks in self-assembling systems is a highly topical research area.1 However, the reliance on self-assembly introduces a high degree of unpredictability into the synthesis.2 For example, even if, when using a net-based approach,3 a 4-connected network is both targeted and achieved, there are still a large number of networks which can form. For square planar 4-connecting nodes, possible nets include the 2D (4,4) net, and the 3D nets of (8,4), 42·84, CdSO4 (cds),4 MOF-112, NbO, quartz dual (or ‘dense’ 75·9), and USF-1.5 Thus control is not only required over the individual building blocks and their local connectivities and geometries, but over the network as a whole.Anionic ligands such as dicarboxylates have been used to bridge between metals to great effect recently in the construction of porous MOFs.6 One of the least utilised dicarboxylate ligands is acetylenedicarboxylate, –O2CCCCO2–. A search of the Cambridge Structural Database 7 finds only 26 structures in which this ligand is used as a linear bridging species between metals. The ligand is observed to bridge between either four metals (often with the metals as two dimers) using all four oxygen atoms8 or between two metals using either one oxygen atom from either extreme of the molecule 9 or with both carboxylate groups binding in an η2 fashion.10
Herein we report the synthesis and contrast the structures of two 1D CuII acetylenedicarboxylate polymers using pyridyl co-ligands. Significantly, the 1D chains are crosslinked viahydrogen bonding to form either 2D (4,4) sheet or 3D cds topology, depending on the steric bulk of pyridine co-ligands.
Results and discussion
Crystals of [Cu(O2CCCCO2)(C5H5N)2(MeOH)] (1) and [Cu(O2CCCCO2)(Me2C5H3N)2(MeOH)]·MeOH (2) were obtained from methanolic solutions of acetylenedicarboxylic acid and copper(II) chloride in the presence of pyridine and 3,5-dimethylpyridine, respectively. Reaction with 3-methylpyridine under the same conditions did not yield a product. The complexes were structurally characterised by single crystal X-ray diffraction. In both instances the product was inextricably combined with a second compound, although sufficient single crystals were isolated for IR analysis (see Experimental section). Both 1 and 2 were the only carboxylate containing products from their respective reactions with peaks attributable to the acetylene triple bond in the region 2150–2200 cm–1. The material obtained alongside 1 was identified as [CuCl2(C5H5N)2] by unit cell analysis, however, that obtained alongside 2 was amorphous and composition could not be further determined.Both complexes form one-dimensional coordination polymers although there are subtle differences in the geometry of the acetylenedicarboxylate ligands and in the relative arrangements of the pyridyl co-ligands. The polymers both contain distorted square pyramidal CuII atoms connected by the dicarboxylate ligands into 1D chains. The dicarboxylate ligands bind η1 in trans positions at the metal with the pyridyl ligands occupying the remaining two in-plane sites, Fig. 1 and 2. The apical coordination site is occupied by a methanol ligand. Bond lengths are comparable between the two complexes, Tables 1 and 2. In both cases the geometry around the CuII is significantly distorted from square pyramidal. The mean plane of four ‘in-plane’ coordinating atoms passes below the copper and oxygen atoms and above the nitrogen atoms. Surprisingly the distortion around the copper atom is less in complex 2 despite more sterically bulky dimethylpyridine co-ligands. The average nitrogen-plane distances are 0.25 and 0.19 Å and oxygen-plane distances of 0.25 and 0.19 Å for 1 and 2, respectively. The Cu-plane distances are 0.11 and 0.05 Å for the respective structures.
![A portion of the 1D coordination polymer [Cu(O2CCCCO2)(C5H5N)2(MeOH)] (1). Ellipsoids displayed at 50% probability, hydrogen atoms omitted for clarity. Symmetry equivalents used; †, x – 1, ½ – y, z – 1/2; #, 1 + x, ½ – y, ½ + z.](/image/article/2008/CE/b711891k/b711891k-f1.gif) | ||
| Fig. 1 A portion of the 1D coordination polymer [Cu(O2CCCCO2)(C5H5N)2(MeOH)] (1). Ellipsoids displayed at 50% probability, hydrogen atoms omitted for clarity. Symmetry equivalents used; †, x – 1, ½ – y, z – 1/2; #, 1 + x, ½ – y, ½ + z. | ||
![A portion of the 1D coordination polymer [Cu(O2CCCCO2)(Me2C5H3N)2(MeOH)]·MeOH (2). Ellipsoids displayed at 50% probability, hydrogen atoms omitted for clarity. Symmetry equivalents used; ‡, x – 1/2, ½ – y, ½ + z; $, ½ + x, ½ – y, z – 1/2.](/image/article/2008/CE/b711891k/b711891k-f2.gif) | ||
| Fig. 2 A portion of the 1D coordination polymer [Cu(O2CCCCO2)(Me2C5H3N)2(MeOH)]·MeOH (2). Ellipsoids displayed at 50% probability, hydrogen atoms omitted for clarity. Symmetry equivalents used; ‡, x – 1/2, ½ – y, ½ + z; $, ½ + x, ½ – y, z – 1/2. | ||
| a Symmetry equivalents used as per Fig. 1. | |||
|---|---|---|---|
| Cu(1)–O(1) | 1.967(1) | Cu(1)–N(1) | 1.998(2) |
| Cu(1)–O(3)† | 1.976(1) | Cu(1)–N(2) | 2.012(2) |
| Cu(1)–O(5) | 2.254(2) | ||
| O(1)–Cu(1)–O(3)† | 171.2(1) | O(1)–Cu(1)–N(1) | 94.1(1) |
| N(1)–Cu(1)–N(2) | 159.2(1) | N(1)–Cu(1)–O(3)† | 89.8(1) |
| O(1)–Cu(1)–O(5) | 86.5(1) | O(3)†–Cu(1)–N(2) | 90.4(1) |
| O(3)†–Cu(1)–O(5) | 85.0(1) | N(2)–Cu(1)–O(1) | 88.8 (1) |
| N(1)–Cu(1)–O(5) | 102.7(1) | N(2)–Cu(1)–O(5) | 98.0(1) |
| a Symmetry equivalents used as per Fig. 2. | |||
|---|---|---|---|
| Cu(1)–O(1) | 1.958(5) | Cu(1)–N(1) | 2.027(6) |
| Cu(1)–O(3)‡ | 1.978(5) | Cu(1)–N(2) | 2.018(4) |
| Cu(1)–O(5) | 2.258(5) | ||
| O(1)–Cu(1)–O(3)‡ | 172.2(2) | O(1)–Cu(1)–N(1) | 89.0(2) |
| N(1)–Cu(1)–N(2) | 166.4(2) | N(1)–Cu(1)–O(3)‡ | 91.0(2) |
| O(1)–Cu(1)–O(5) | 83.3(2) | O(3)‡–Cu(1)–N(2) | 91.0(2) |
| O(3)‡–Cu(1)–O(5) | 89.0(2) | N(2)–Cu(1)–O(1) | 90.9(2) |
| N(1)–Cu(1)–O(5) | 100.4(2) | N(2)–Cu(1)–O(5) | 93.1(2) |
Whilst the connectivity of the two polymers is identical the relative geometries of the ligands are different. The bridging acetylenedicarboxylate ligands adopt significantly different orientations in the two structures with the relative orientations of the terminal carboxylates with respect to each other differing. In complex 1 the angle between the two carboxylate planes is 56° whereas in 2 the angle is significantly larger at 74°. This twisting of the ligand appears to affect the relative orientation of adjacent CuII centres. The angle between adjacent N,N,O,O mean-planes is 57° in complex 1 whereas in 2 the angle is correspondingly larger, 80°. The twisting of the polymers can be predominantly attributed to the twisting of the ligand as in both 1 and 2 the angles between the planes of the binding carboxylates around the metal are smaller (19° in complex 1 and 18° in complex 2). The pyridyl ligands in the complexes are also arranged differently with respect to each other. In complex 1 the planes of the opposing pyridyl ligands are at an angle of 43°. Surprisingly the angle between the more bulky 3,5-dimethylpyridyl ligands in 2 is only 7°.
Both complexes display intermolecular hydrogen bonding involving the apical methanol ligands, Table 3. The 1D polymers in 1 pack viahydrogen bonding between the OH of the methanol ligand and the non-coordinating oxygen atom of an η1carboxylate in an adjacent chain to form (4,4) sheets, Fig. 3. The hydrogen bonding in 2 is more complex with the incorporation of the lattice methanol and the greater twist in the polymeric chain. The lattice methanol bridges between a methanol ligand and a non-binding carboxylate oxygen atom (an extended version of the hydrogen bonding in 1), Fig. 4a. The twist in the chain, however, gives rise to a 3D hydrogen bonded network rather than an expanded version of the 2D sheet in 1. The network has CdSO4 (cds) topology in which the copper atoms act as square planar 4-connecting nodes, Fig. 4b. Alternate copper atoms along the polymeric chain hydrogen bond in perpendicular directions, due to the twist in the chain, giving rise to the 3D net (highlighted in red in Fig. 4b). Packing in both complexes is further supported by π-interactions. Complex 1 contains both edge-to-face and face-to-face interactions between sheets whereas the structure of 2 contains only offset face-to-face interactions due to the presence of the methyl substituents.
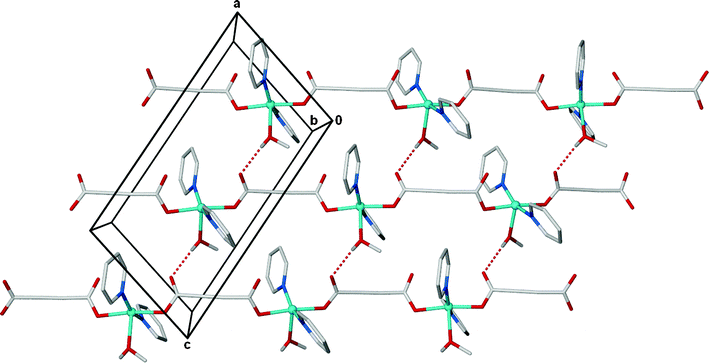 | ||
| Fig. 3 Hydrogen bonded (4,4)-sheets propagate parallel to the ac plane in the structure of 1. CH hydrogen atoms are omitted for clarity. | ||
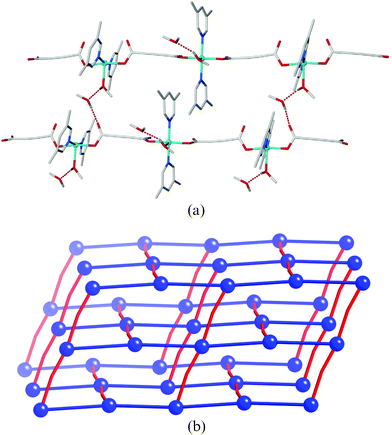 | ||
| Fig. 4 (a) The linking of two adjacent chains in 2via the hydrogen bonding bridges; all links between chains are equivalent. (b) Schematic representation of the overall cds topology. Blue spheres represent the Cu nodes, blue bonds represent the coordinated acetylenedicarboxylate bridges, and red bonds represent the hydrogen bonding bridges. | ||
The most important factor, however, in determining the network topology appears to be the steric bulk of the pyridine ligand. In 1 there are close interactions between some of the pyridine hydrogen atoms in the 3/5 positions and atoms elsewhere in the structure (H6⋯O2 = 2.637 Å, H13⋯C2 = 2.852 Å), Fig. 5. Placement of the methyl groups in these positions would create unfavourable steric clashes, and thus an alternate topology (cds) is formed. The interchain distances are also increased through the incorporation of a lattice methanol into the bridges. On the other hand, in the absence of the methyl groups, the (4,4) topology is presumably the most efficient packing arrangement.
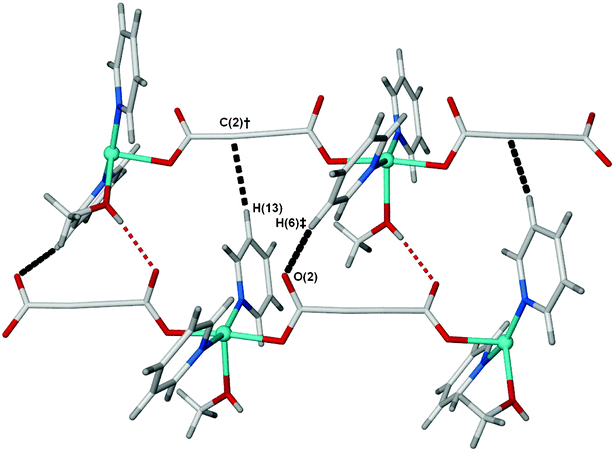 | ||
| Fig. 5 A single window in the (4,4) sheets in 1, showing the close C–H⋯X interactions (black dashed lines) for two of the four pyridine 3/5 positions. The distances are H(6)‡⋯O(2) = 2.64 Å and H(13)⋯C(2)† = 2.85 Å. Symmetry equivalents used; ‡, 1 + x, y, z; †, x, ½ – y, z – 1/2. | ||
Conclusions
The acetylenedicarboxylate ligand shows potential to act as a dianionic linear connector in coordination polymers. However, the steric bulk of the pyridine co-ligands appears to have a significant effect on the way the 1D chains are linked by hydrogen bonds to give differing 4-connected networks, both two- and three-dimensional.Experimental details
Synthesis
All reagents and solvents were purchased from standard commercial sources and used without further purification. ATR-IR were collected in the region 600–4000 cm–1 using a Bruker Equinox 55 spectrometer. Calculations of yields were possible as in both reactions two products were observed to form that could not easily be isolated from each other. A sufficient quantity of all products to perform IR was manually extracted but enough could not be separated for bulk analyses.Crystallographic details
Single crystals were mounted on fine glass fibres using viscous hydrocarbon oil. Data were collected on either a Bruker X8 Apex II CCD (1) or Nonius Kappa CCD (2) diffractometer both equipped with graphite monochromated Mo Kα radiation (λ = 0.71073 Å). Data collection temperatures were maintained at 123 K using open flow N2 cryostreams. Data were initially processed using the Apex II program suite or the DENZO package for the respective instruments with absorption corrections applied.11,12 Structures were solved using SHELXS-9713 and refined by least-squares refinement against F2 using SHELXL-9713 with the program X-Seed as a graphical interface.14 All non-hydrogen atoms were refined anisotropically. Hydrogen atoms attached to carbon were placed in idealised positions and refined using a riding model to the atom to which they are attached. Hydrogen atoms attached to methanol were located from the Fourier difference map, with those in the structure of 2 refined using DFIX restraints. Compound 2 was refined as a racemic twin. Crystallographic details and final refinement parameters are listed in Table 4.| 1 | 2 | |
|---|---|---|
| Empirical formula | C15H14Cu1N2O5 | C20H26Cu1N2O6 |
| M | 365.82 | 453.97 |
| Crystal system | Monoclinic | Monoclinic |
| Space group | P21/c | Cc |
| a/Å | 7.8631(2) | 10.5816(8) |
| b/Å | 14.2780(3) | 14.0388(10) |
| c/Å | 14.3028(3) | 14.5487(12) |
| β/° | 104.829(1) | 94.380(5) |
| V/Å3 | 1552.28(6) | 2154.9(3) |
| Z | 4 | 4 |
| µ/mm–1 | 1.433 | 1.051 |
| Unique/I > 2σI | 3438/2904 | 3318/2575 |
| R int | 0.0339 | 0.0533 |
| wR 2 (all data) | 0.0741 | 0.1130 |
| R 1 (I > 2σI) | 0.0310 | 0.0483 |
We thank the Australian Research Council for funding and for a post-doctoral fellowship (D.R.T.). We also thank CSIRO for the Student Research Scheme.
References
- J. W. Steed, D. R. Turner and K. J. Wallace, Core Concepts in Supramolecular Chemistry and Nanochemistry, 2007, John Wiley and Sons Ltd., Chichester Search PubMed.
- S. R. Batten, J. Solid State Chem., 2005, 178, 2475 CrossRef CAS.
- R. Robson, J. Chem. Soc., Dalton Trans., 2000, 3735 RSC.
- For three-letter net codes, see N. W. Ockwig, O. Delgado-Friedrichs, M. O'Keeffe and O. M. Yaghi, Acc. Chem. Res., 2005, 38, 176 Search PubMed and the associated website, http://rcsr.anu.edu.au/home.
- M.-L. Tong, X.-M. Chen and S. R. Batten, J. Am. Chem. Soc., 2003, 125, 16170 CrossRef CAS.
- J. L. C. Rowsell and O. M. Yaghi, Microporous Mesoporous Mater., 2004, 73, 3 CrossRef CAS.
- Cambridge Structural Database, v.5.28 + 1 update (Jan 2007).
- For example, (a) M. Pascu, M. Andruh, A. Muller and M. Schmidtmann, Polyhedron, 2004, 23, 673 CrossRef CAS; (b) D. Visinescu, G. I. Pascu, M. Andruh, J. Magull and H. W. Roesky, Inorg. Chim. Acta, 2002, 340, 201 CrossRef CAS.
- For example, (a) J. Kim, B. Chen, T. M. Reineke, H. Li, M. Eddaoudi, D. B. Moler, M. O'Keefe and O. M. Yaghi, J. Am. Chem. Soc., 2001, 123, 8239 CrossRef CAS; (b) I. Patenburg and U. Ruschewitz, Z. Anorg. Allg. Chem., 2002, 628, 1697 CrossRef; (c) H.-U. Wang, S. Gao, L.-H. Huo and J.-G. Zhao, Acta Crystallogr., 2006, E62, m3152 CAS.
- S. Skoulika, P. Dallas, M. G. Siskos, Y. Deligiannakis and A. Michaelides, Chem. Mater., 2003, 15, 4576 CrossRef CAS.
- Apex2 v.2.1-0, Bruker AXS, 2006.
- Z. Otwinowski and W. Minor, in Methods in Enzymology, vol. 276, edited by C. W. Carter Jr and R. M. Sweet, pp. 307–326. New York: Academic Press, 1997 Search PubMed.
- G. M. Sheldrik, SHELXS-97 and SHELXL-97, University of Göttingen, 1997.
- L. J. Barbour, J. Supramol. Chem., 2001, 1, 189 CrossRef CAS.
Footnote |
| † CCDC reference numbers 656474 and 656475. For crystallographic data in CIF or other electronic format see DOI: 10.1039/b711891k |
| This journal is © The Royal Society of Chemistry 2008 |
