Bifidobacterium animalis subsp. lactis Bbm-19 ameliorates insomnia by remodeling the gut microbiota and restoring γ-aminobutyric acid and serotonin signaling
Qiong
Wu
 abc,
Guangqi
Gao
abd,
Lai-Yu
Kwok
abc,
Jiaqi
Qiao
abc,
Ziqing
Wei
abc,
Qiuwen
He
abc and
Zhihong
Sun
*abd
abc,
Guangqi
Gao
abd,
Lai-Yu
Kwok
abc,
Jiaqi
Qiao
abc,
Ziqing
Wei
abc,
Qiuwen
He
abc and
Zhihong
Sun
*abd
aKey Laboratory of Dairy Biotechnology and Engineering, Ministry of Education, Inner Mongolia Agricultural University, Hohhot, P.R. China. E-mail: sunzhihong78@163.com; Fax: +86 471-4305357; Tel: +86 471-4308703
bKey Laboratory of Dairy Products Processing, Ministry of Agriculture and Rural Affairs, Inner Mongolia Agricultural University, Hohhot, P.R. China
cInner Mongolia Key Laboratory of Dairy Biotechnology and Engineering, Inner Mongolia Agricultural University, Hohhot, P.R. China
dCollaborative Innovative Center for Lactic Acid Bacteria and Fermented Dairy Products, Ministry of Education, Inner Mongolia Agricultural University, Hohhot, P. R. China
First published on 9th December 2025
Abstract
Insomnia is associated with dysregulation of the gut–brain axis, yet microbiome-targeted interventions remain underexplored. In this study, we investigated the effects of Bifidobacterium animalis subsp. lactis Bbm-19 (Bbm-19), a strain isolated from human breast milk, in a 4-chloro-DL-phenylalanine-induced mouse model of insomnia. Using integrated behavioral, neurochemical, immunological, and multi-omics approaches, this study demonstrates that insomnia is characterized by shortened sleep duration, prolonged sleep latency, anxiety-like behaviors, and reduced levels of serotonin and gamma-aminobutyric acid in the gut, serum, and brain. Administration of Bbm-19 significantly improved sleep parameters, reduced anxiety-like behaviors, and increased survival. Metagenomic and metabolomic analyses revealed that Bbm-19 restored gut microbiota balance, enriched beneficial taxa, including Muribaculaceae bacterium and Stercoribacter sp., and reprogrammed microbial metabolic modules, particularly those involved in amino acid metabolism (including alanine, aspartate, glutamate, arginine, proline, and tryptophan pathways). Targeted metabolomics confirmed increased levels of gamma-aminobutyric acid and serotonin in fecal and brain tissues, along with normalization of inflammatory cytokine profiles. Spearman correlation analysis linked Bbm-19-enriched taxa to improved neurotransmitter levels and sleep outcomes. Notably, Bbm-19 outperformed lorazepam in modulating gut-specific metabolic functions and synergistically enhanced its effects when co-administered. These findings demonstrate that Bbm-19 ameliorates insomnia through coordinated regulation of the gut microbiota, host metabolism, and neuroimmune signaling, highlighting its potential as a targeted psychobiotic intervention for sleep disorders.
Introduction
Sleep disorders are among the most common clinical problems,1 conventional pharmacological treatments, although widely used, are often associated with adverse effects such as dependency, daytime drowsiness, and cognitive impairment, highlighting the need for safer and more sustainable therapeutic alternatives. Emerging research has revealed that the gut microbiota plays a pivotal role in regulating central nervous system functions through the microbiota–gut–brain axis, providing a new perspective on sleep regulation.2 As core intestinal commensal bacteria, Bifidobacterium species influence host neurotransmitter balance via multiple mechanisms, including the production of microbial metabolites, immunomodulation, and neuroendocrine signaling.3–6 Among these, Bifidobacterium animalis subsp. lactis (B. animalis subsp. lactis) has garnered particular attention due to its robust intestinal colonization capacity and potential to modulate neuroactive pathways, making it a promising candidate for microbiome-based sleep interventions.4,6Clinical evidence suggests that supplementation with B. animalis subsp. lactis BB-12® increases its intestinal abundance, which has been significantly correlated with reduced crying duration and extended sleep time in infants.7 The sleep-promoting effect is believed to be mediated, at least in part, by its ability to influence key neurotransmitter systems. Several probiotic genera, including Lactobacillus, Bifidobacterium, Escherichia, and Enterococcus have been shown to synthesize critical regulators of sleep–wake cycles such as tryptophan, serotonin (5-HT) and gamma-aminobutyric acid (GABA),2,8,9 which are critical regulators of sleep–wake cycles. These microbial metabolites may serve as mediators in the bidirectional communication between the gut microbiota and the host's sleep-regulating neural circuits. Furthermore, certain probiotics, particularly Lactobacillus, Bifidobacterium, and Enterococcus, can modulate levels of inflammatory cytokines such as IL-12 and IL-4,10,11 potentially mitigating neuroinflammation that disrupts sleep centers in the brain.
Much of the existing research on probiotics and sleep has focused on multi-strain formulations. For example, a 28 day intervention with a combination of Lactobacillus brevis PBS072 and Bifidobacterium brevis BB077 improved stress response, cognitive function, and sleep quality in students facing academic stress.12 Similarly, a probiotic blend containing Lactobacillus and Bifidobacterium longum was shown to alleviate depressed mood, anger, and fatigue, while significantly enhancing Pittsburgh Sleep Quality Index scores.13 While these studies highlight the potential of multi-strain probiotics in modulating sleep and emotional well-being, they provide limited insight into the strain-specific mechanisms underlying these effects. In particular, the neurochemical pathways through which individual Bifidobacterium species, such as B. animalis subsp. lactis, influence central nervous system function remain poorly characterized. Moreover, the dynamics of gut-derived neurotransmitters, particularly GABA and 5-HT, during probiotic intervention are still controversial, with inconsistent findings across studies. Compounding these limitations, most current models lack longitudinal, multi-omics integration, making it difficult to establish causal links between microbial shifts and host physiological outcomes.
These challenges are further amplified in preclinical research, where studies on insomnia are often hindered by methodological variability and the lack of standardized animal models. To date, only a few pharmacological approaches have been widely adopted to induce insomnia-like states in rodents, with 4-chloro-DL-phenylalanine (PCPA)14–16 and caffeine17,18 being the most commonly used agents. Among these, PCPA, by selectively and irreversibly inhibiting tryptophan hydroxylase and blocking 5-HT synthesis, offers a reproducible and physiologically relevant model for studying sleep disruption linked to serotonergic dysfunction. Thus far, only a limited number of microbiota-targeted interventions have been evaluated in the PCPA-induced insomnia model. For instance, Yao et al. (2022)16 demonstrated that a 7 day treatment with Armillaria mellea fermentation broth improved sleep quality in PCPA-treated mice by modulating the 5-hydroxytryptamine system and restoring gut microbiota balance. While this study provides valuable proof-of-concept evidence for microbiome-based interventions in insomnia, it involved a complex fungal preparation rather than a defined bacterial strain, limiting mechanistic interpretation. Moreover, the short intervention period and lack of multi-omics profiling restrict insights into the longitudinal dynamics of host–microbe interactions. These limitations highlight the need for studies that employ well-defined probiotic strains, extended treatment durations, and integrative analyses to unravel the precise mechanisms by which gut microbiota influence sleep regulation.
In this study, we investigated the effects of a novel strain, B. animalis subsp. lactis Bbm-19 (Bbm-19), originally isolated from human breast milk, on sleep regulation in a PCPA-induced mouse model of insomnia. Over a 42 day intervention period, we employed integrated metabolomic and metagenomic analyses to explore the molecular basis of sleep regulation, with a focus on the tryptophan-5-HT and glutamate-GABA metabolic pathways. Our work aims to establish strain-specific mechanisms of B. animalis subsp. lactis in sleep modulation and provide foundational evidence targeted gut–brain axis interventions.
Methods
Ethics statement
All experimental procedures involving mice were approved by the Experimental Animal Ethics Committee of Inner Mongolia Agricultural University (Approval code: NND2025161) and conducted in compliance with ARRIVE guidelines 2.0.Probiotic strain
The probiotic strain Bbm-19 was isolated from human breast milk, and the packaged live probiotic powder of B. animalis subsp. lactis Bbm-19 (2 × 1011 CFU per g) was provided by the Key Laboratory of Dairy Biotechnology and Engineering, Ministry of Education, Inner Mongolia Agricultural University, China. The strain identity was verified by 16S rRNA gene sequencing and whole-genome analysis, with results confirming ≥99.9% similarity to B. animalis subsp. lactis type strains.Mice
Seventy male C57BL/6J mice (weighing 17–20 g, 6 weeks old) were obtained from SPF (Beijing) Biotechnology Co., Ltd, China (License No. SCXK (Beijing) 2019-0010). Mice were housed under controlled conditions with a 12 h light/dark cycle at a constant temperature of 22 ± 2 °C, with ad libitum access to standard chow and water.Mouse model and experimental design
The animal study was designed to evaluate the effects of Bbm-19 on PCPA-induced insomnia. All were acclimatized for one week before assignment to five experimental groups (n = 14 per group): (1) healthy control (HEL): received daily intragastric gavage of 0.1 mL saline; (2) insomnia model (INS): received intraperitoneal injections of PCPA (300 mg kg−1) and daily saline gavage; (3) probiotic treatment (Bb): received PCPA injections and daily gavage of Bbm-19 (8 × 109 CFU); (4) lorazepam treatment (LXP): received PCPA injections and lorazepam (0.5 mg kg−1) via gavage every day; (5) combination treatment (Bb + LXP): received PCPA injections and daily Bbm-19 (8 × 109 CFU) and concurrent lorazepam gavage (Fig. 1a).The study goal was to independently evaluate the therapeutic effects of probiotics following both a short-term (2 week) and a long-term (6 week) intervention period. Given that the PCPA-induced insomnia model is known to be reversible. Therefore, insomnia was induced by intraperitoneal injection of PCPA on days 6, 7, 8, 34, 35, and 36 to achieve sustained inhibition of tryptophan hydroxylase, thereby disrupting central 5-HT synthesis.
Lorazepam is a benzodiazepine derivative that enhances the function of the inhibitory neurotransmitter GABA. In addition, lorazepam is also a validated interventional agent that not only regulates sleep but has also been applied in numerous studies.19–21 Therefore, lorazepam was selected as the positive control treatment. For oral gavage, mice in the Bb and Bb + LXP groups received 0.1 mL of the freshly prepared probiotic suspension daily, delivering a final dose of 8 × 109 CFU per mouse. All interventions were administered over a 42 day period (Fig. 1a).
Sample collection and storage
Fecal, serum, colonic content, and brain tissue samples were collected aseptically. Fecal samples were collected on days 14 and 42. Prior to collection, mice were housed individually in freshly cleaned cages to allow spontaneous defecation. At the end of the trial (day 42), blood was collected via orbital sinus puncture under mild anesthesia, allowed to clot at room temperature for 2 h, and then centrifuged at 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 10 min at 4 °C to obtain serum. Colonic contents and brain tissues were aseptically harvested, immediately frozen in liquid nitrogen. All samples were stored at −80 °C until analysis. The final sample sizes included in the analysis were as follows: after the 14 day intervention, the group sizes for HEL, INS, Bb, LXP, and Bb + LXP were 12, 12, 12, 11, and 12, respectively. Following the 42 day intervention, the corresponding group sizes were 12, 12, 12, 10, and 12.
000g for 10 min at 4 °C to obtain serum. Colonic contents and brain tissues were aseptically harvested, immediately frozen in liquid nitrogen. All samples were stored at −80 °C until analysis. The final sample sizes included in the analysis were as follows: after the 14 day intervention, the group sizes for HEL, INS, Bb, LXP, and Bb + LXP were 12, 12, 12, 11, and 12, respectively. Following the 42 day intervention, the corresponding group sizes were 12, 12, 12, 10, and 12.
Pentobarbital-induced sleep test
Refer to previous research,16,22,23 this paper evaluates the sleep regulation effect of Bbm-19 through the pentobarbital compromise test, which is also a key method for improving sleep function test. Two hours after the administration of probiotics and lorazepam, sodium pentobarbital (60 mg kg−1) was intraperitoneally injected at a volume of less than 0.2 ml per 20 g body weight, a dose previously determined in preliminary experiments to induce 100% sleep onset without excessive baseline prolongation. The latency to sleep was defined as the time interval from injection to the loss of the righting reflex. Sleep duration was recorded as the time from the loss to the recovery of the righting reflex. Each mouse was monitored individually until full recovery. Prolonged sleep duration indicates test-substance synergy with pentobarbital.Open field test
An open field test was conducted on days 14 and 42 to assess locomotor activity and anxiety-like behavior. Each mouse was individually placed in the center of a square arena (40 cm × 40 cm × 40 cm) with opaque walls. After a 5 min acclimatization period, spontaneous movement was recorded for 5 min using a video tracking system. Total distance traveled and time spent in the central zone (defined as the inner 50% of the arena) were analyzed as indicators of exploratory behavior and anxiety levels, respectively. Between trials, the apparatus was thoroughly cleaned with 75% ethanol and dried with paper towels to eliminate olfactory cues.Enzyme-linked immunosorbent assays (ELISA)
Cytokine and neurotransmitter levels were quantified using ELISA. Brain tissues were homogenized in 1 mL of phosphate-buffered saline, centrifuged at 3000g for 15 min at 4 °C. Concentrations of interleukin (IL)-4, IL-6, IL-9, IL-10, and interferon-gamma (IFN-γ) in serum and brain homogenates were measured using commercial ELISA kits (Mouse IL-4, MM-0165M2; IL-6, MM-0163M2; IL-9, MM-0162M2; IL-10, MM-0176M2; IFN-γ, MM-0182M2; Jiangsu Meimian Industrial Co., Ltd, China) according to the manufacturer's instructions. For neurotransmitter analysis, 0.1 g of colonic content was suspended in 1 mL of phosphate-buffered saline, homogenized, and centrifuged (3000g, 15 min, 4 °C) to obtain supernatant. Levels of 5-HT and GABA in brain tissue, serum, and gut contents were determined using Mouse 5-HT ELISA kit (RXJ203223M) and Mouse GABA ELISA kit (RXJ99905) (Quanzhou Ruixin Biotechnology Co., Ltd, Fujian, China). Absorbance was measured per kit protocols, with concentrations calculated via standard curve construction using CurveExpert software v1.4.Shotgun metagenomic sequencing and bioinformatic analysis
Total microbial genomic DNA was extracted from fecal samples using the OMEGA Mag-Bind Soil DNA Kit (M5635-02; Omega Bio-Tek, Norcross, GA, USA) according to the manufacturer's instructions and stored at −20 °C until analysis. DNA quality was assessed using a Qubit™ 4 Fluorometer (Q33238; Invitrogen, Waltham, MA, USA) with the Qubit™ 1× Double-Stranded DNA High Sensitivity Assay (Q33231) and confirmed by agarose gel electrophoresis. Metagenomic libraries were constructed using the Illumina TruSeq Nano DNA LT Library Preparation Kit (Illumina, Inc., San Diego, CA, USA), with an average insert size of approximately 400 bp. Sequencing was performed on the Illumina NovaSeq X Plus platform to generate paired-end reads.Raw sequencing reads were subjected to stringent quality filtering using FastP (v0.23.2): reads containing more than 10% unidentified nucleotides (N content) or more than 50% bases with a Phred quality score ≤5 were discarded. Host-derived sequences were removed by aligning the reads to the mouse reference genome (GRCm39) using Minimap2 (v2.24-r1122). This yielded high-quality reads averaging 6.71 Gb per sample (total clean data: 161.08 Gb).
Bacterial interactions were modeled via Spearman correlation between SGB abundances. Significant edges (|r| > 0.8, P < 0.01) were retained to construct group-specific-occurrence networks. Network topology was analyzed to assess ecological interactions, with a focus on connection density (also referred to as network density). It was calculated as the ratio of the actual number of edges to the maximum possible number of edges in the network: network density = number of observed edges/[number of nodes × (number of nodes − 1)], providing a measure of interaction complexity and system stability.
The open reading frames (ORFs) predicted for each SGB were functionally annotated by alignment against the Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthology (KO) database to identify key metabolic modules.24 To predict bioactive gut metabolite profiles, one million reads per sample were first extracted using seqtk (v1.3-r106). These reads were then aligned via DIAMOND blastx function under stringent parameters (–query-cover 90 –id 50), and gene abundance was quantified based on the best alignment hit. Finally, metabolite abundance profiles were inferred from the gene abundance data using the MelonnPan-predict pipeline.25
Fecal non-targeted metabolomic analysis
Fecal samples (20 mg) were homogenized in 300 μL of pre-chilled methanol containing 5 ppm 2-chlorophenylalanine (internal standard) using stainless-steel beads. The mixture was vortexed for 30 s, followed by two cycles of high-throughput tissue grinding (55 Hz, 60 s) to ensure complete disruption of the sample. The homogenate was subjected to ultrasonic extraction for 10 min at 4 °C and subsequently incubated at −20 °C for 30 min to facilitate protein precipitation. After centrifugation at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 10 min at 4 °C, the supernatant was filtered through a 0.22 µm polytetrafluoroethylene membrane and transferred to a glass autosampler vial for analysis. A pooled quality control (QC) sample was prepared by mixing equal volume of filtered supernatant from each sample. Prior to sample analysis, the system was conditioned by injecting three QC samples, followed by one QC sample every 11 experimental injections to monitor instrument stability and ensure data quality.
000g for 10 min at 4 °C, the supernatant was filtered through a 0.22 µm polytetrafluoroethylene membrane and transferred to a glass autosampler vial for analysis. A pooled quality control (QC) sample was prepared by mixing equal volume of filtered supernatant from each sample. Prior to sample analysis, the system was conditioned by injecting three QC samples, followed by one QC sample every 11 experimental injections to monitor instrument stability and ensure data quality.
Chromatographic separation was performed using an ACQUITY UPLC HSS T3 column (100 Å, 1.8 µm, 2.1 mm × 100 mm; Waters Corporation, Milford, MA, USA) maintained at 40 °C, with the autosampler temperature set to 8 °C. The flow rate was 0.4 mL min−1, and a 2 µL injection volume was used. The mobile phase consisted of (A) 0.1% formic acid in water and (B) 0.1% formic acid in acetonitrile, using the following gradient (Table 1):
Metabolite detection was carried out using a Thermo Orbitrap Exploris 120 mass spectrometer (Thermo Fisher Scientific Inc., Waltham, MA, USA) in data-dependent acquisition mode, controlled by Xcalibur software (v4.7). Analysis was performed in both positive and negative electrospray ionization modes using a heated electrospray ionization source with the following parameters: spray voltage of +3.5 kV (positive) or −3.0 kV (negative); sheath gas of 40 arbitrary units; auxiliary gas of 10 arbitrary units; capillary temperature of 320 °C; auxiliary gas temperature of 300 °C; primary resolution of 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000; scan range of 70–1000 m/z; automatic gain control target set to standard; maximum injection time of 100 ms; top 4 precursor ions selected for higher-energy collisional dissociation fragmentation at 30% collision energy; dynamic exclusion duration of 4 s; secondary resolution at 15
000; scan range of 70–1000 m/z; automatic gain control target set to standard; maximum injection time of 100 ms; top 4 precursor ions selected for higher-energy collisional dissociation fragmentation at 30% collision energy; dynamic exclusion duration of 4 s; secondary resolution at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000; and secondary automatic gain control target set to standard with auto maximum injection time. All samples, including QC injections, were analyzed under identical chromatographic and mass spectrometric conditions.
000; and secondary automatic gain control target set to standard with auto maximum injection time. All samples, including QC injections, were analyzed under identical chromatographic and mass spectrometric conditions.
| Time (min) | Acetonitrile with 0.1% formic acid |
|---|---|
| 0 | 5% |
| 1 | 5% |
| 4.7 | 95% |
| 6 | 95% |
| 6.1 | 5% |
| 8.5 | 5% |
Raw data were processed using MS-DIAL software (v4.9.221218) for peak extraction, alignment, filtering, and metabolite identification. Peaks undetected in >50% of QC samples were filtered out, followed by gap filling and normalization using the software's embedded algorithms.
Targeted quantification of fecal metabolites by ultra-performance liquid chromatography-tandem mass spectrometry (UPLC-MS/MS)
Fecal samples were thawed on ice, and 50 μL aliquots were mixed with 250 μL of ice-cold 20% acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) methanol (v/v, 1
methanol (v/v, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4) for protein precipitation. The mixtures were vortexed for 3 min and centrifuged at 12
4) for protein precipitation. The mixtures were vortexed for 3 min and centrifuged at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 10 min at 4 °C. Supernatants were transferred to fresh microcentrifuge tubes, incubated at −20 °C for 30 min to further precipitate interfering substances, and centrifuged again under the same conditions. Clarified supernatants (180 μL) were analyzed using an ExionLC™ AD UPLC system coupled to a QTRAP® 6500 + mass spectrometer (AB Sciex LLC, Framingham, MA, USA) equipped with a Waters ACQUITY UPLC HSS T3 C18 column (1.8 μm, 100 × 2.1 mm). Chromatographic separation employed: mobile phase A (0.1% formic acid in water) and B (0.1% formic acid in acetonitrile) at 0.35 mL min−1 flow rate, with column temperature maintained at 40 °C and 2 μL injection volume. The gradient program was as follows: 95% A at 0 min, linearly decreased to 5% A by 8 min, held at 5% A until 9.5 min, returned to 95% A at 9.6 min, and re-equilibrated for 2.4 min until 12 min.
000g for 10 min at 4 °C. Supernatants were transferred to fresh microcentrifuge tubes, incubated at −20 °C for 30 min to further precipitate interfering substances, and centrifuged again under the same conditions. Clarified supernatants (180 μL) were analyzed using an ExionLC™ AD UPLC system coupled to a QTRAP® 6500 + mass spectrometer (AB Sciex LLC, Framingham, MA, USA) equipped with a Waters ACQUITY UPLC HSS T3 C18 column (1.8 μm, 100 × 2.1 mm). Chromatographic separation employed: mobile phase A (0.1% formic acid in water) and B (0.1% formic acid in acetonitrile) at 0.35 mL min−1 flow rate, with column temperature maintained at 40 °C and 2 μL injection volume. The gradient program was as follows: 95% A at 0 min, linearly decreased to 5% A by 8 min, held at 5% A until 9.5 min, returned to 95% A at 9.6 min, and re-equilibrated for 2.4 min until 12 min.
Mass spectra detection was operated in multiple reaction monitoring mode with dynamic switching between positive and negative electrospray ionization. Mass spectrometric parameters were set as follows: ion spray voltage of +5500 V (positive mode) and −4500 V (negative mode), source temperature of 550 °C, and curtain gas of 35 psi. Quality control samples pooled from all specimens were analyzed five times for system stability monitoring.
Raw mass spectrometry data were converted into the standardized mzXML format using the MSConvert module of the ProteoWizard software suite. Subsequent data processing, including peak detection, noise filtering, retention time correction, and cross-sample peak alignment, was performed using the RxCMS computing platform to generate a quantitative metabolite feature table. To ensure data reliability, stringent quality assessment was applied based on QC sample reproducibility, with metabolite signals retained only if their relative standard deviation (RSD) across QCs was less than 15%. For absolute quantification, calibration curves were constructed using a series of standards. The concentration of each standard was plotted against the corresponding peak area ratio of the analyte to its internal standard, enabling linear regression-based quantification of target metabolites in the samples.
Statistical analysis
Differences in other continuous variables between groups were statistically evaluated by Wilcoxon rank-sum test and one-way ANOVA with Tukey's multiple comparisons test. Statistical analysis was performed using R v4.4.2. Permutation multivariate analysis of variation (PERMANOVA) was used to perform permutation multivariate analysis of variation, significance P-value was generated based on 1000 permutations, and the effect size (R2) was calculated by analysis. Based on the Bray–Curtis distance matrix composed of intestinal microorganisms, the difference between samples was visualized using principal coordinate analysis (PCoA).Results
Bbm-19 improves sleep parameters in PCPA-induced insomniac mice
To evaluate the effects of Bbm-19 on sleep regulation in the context of the gut–microbiota–brain axis, a mouse model of insomnia was established via intraperitoneal injection of PCPA. Six-week-old male C57BL/6J mice were assigned to five experimental groups: healthy control (HEL, n = 12), insomnia model (INS, n = 12), probiotic treatment (Bb, n = 12), lorazepam treatment (LXP, n = 10), and combination treatment (Bb + LXP, n = 12), with interventions administered over a 42 day period (Fig. 1a). Sleep-related responses were assessed using a sodium pentobarbital-induced sleep test. Compared to the HEL group, the INS group exhibited significantly prolonged sleep latency (P < 0.05; Fig. 1b) and shortened sleep duration (P < 0.05; Fig. 1c), confirming the successful induction of insomnia-like phenotypes. Intervention with Bbm-19 significantly improved both parameters relative to the INS group (P < 0.05), indicating a sleep-promoting effect. Similarly, lorazepam monotherapy improved both parameters, consistent with its known sedative properties. Notably, the combination of Bbm-19 and lorazepam resulted in the most pronounced improvements, suggesting an additive or synergistic interaction between the probiotic and the drug. These results indicate that Bbm-19 supplementation ameliorates sleep disruption in PCPA-induced insomnia, with synergistic enhancement observed during combined pharmacological-probiotic therapy.Bbm-19 attenuates anxiety-like behaviors and improves survival in insomniac mice
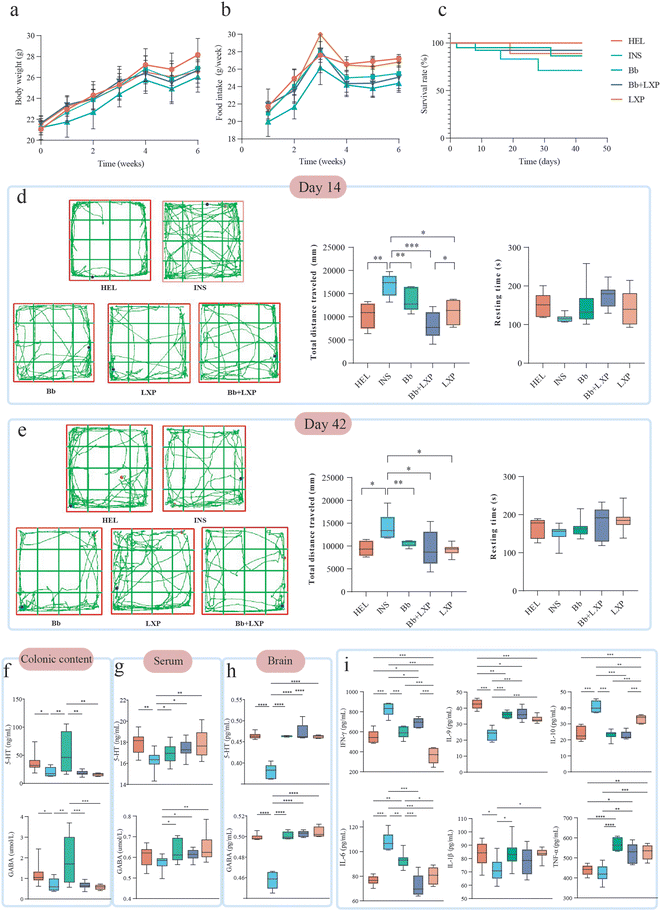
Throughout the 42 day experimental period, all mice exhibited progressive body weight gain. However, by week 5, the INS group showed a trend toward reduced weight gain compared to the HEL group, although the difference did not reach statistical significance (Fig. 2a, P > 0.05). This divergence in body weight increased over time. Food intake was slightly higher in the HEL and LXP groups compared to other groups starting at week 4, with a widening gap as the intervention progressed (Fig. 2b, P > 0.05).No mortality was observed in the HEL group. The INS group exhibited elevated mortality, which was mitigated by all active interventions, indicating a protective effect of Bbm-19 and/or LXP supplementation (Fig. 2c, P > 0.05).
Open field testing on days 14 and 42 revealed significant anxiety-like behaviors in the INS group. Compared to the HEL group, INS mice traveled a significantly greater distance (P < 0.05; Fig. 2d), indicative of hyperlocomotion, while resting time showed a similar trend, with lowest resting time in the INS group, but did not differ significantly (Fig. 2e). These behavioral changes are consistent with an anxiety-like phenotype induced by chronic insomnia. Treatment with Bbm-19, lorazepam, or their combination normalized locomotor activity, with total distance traveled no longer significantly different from the HEL group (P > 0.05). Notably, the Bb + LXP group exhibited a more pronounced reduction in locomotion than the LXP group at day 14 (P < 0.05), suggesting a stronger sedative effect with the combined drug-probiotic intervention. With the passage of time, the total distance traveled decreased and the resting time increased of mice in all groups. The total distance traveled of Bb group (P = 0.001) and LXP group (P = 0.02) decreased significantly. It shows that probiotic and drug intervention can maintain the effect.
Bbm-19 restores neurotransmitter levels in peripheral and central compartments
Neurotransmitters such as 5-HT and GABA play critical roles in regulating sleep–wake cycles. To evaluate the impact of Bbm-19 on neurotransmitter homeostasis, GABA and 5-HT levels were measured in colonic contents, serum, and brain tissue after the 42 day intervention (Fig. 2f–h).Compared to the HEL group, the INS group exhibited significantly lower 5-HT and GABA levels in both colonic contents and serum (P < 0.05; Fig. 2f and g). These peripheral deficits, combined with prolonged sleep latency in the sodium pentobarbital-induced sleep test, validate the neurochemical relevance of the insomnia model. Administration of Bbm-19 (Bb group) significantly increased GABA and 5-HT levels in colonic contents and serum compared to the INS group (P < 0.05), indicating a robust gut-targeted neuromodulatory effect. Lorazepam (LXP) and combination therapy (Bb + LXP) restored serum 5-HT and GABA to HEL-comparable levels (P > 0.05 vs. HEL group), but did not significantly alter colonic neurotransmitter concentrations (P > 0.05 vs. INS group), suggesting limited impact on gut-level dynamics.
In brain tissue, the central hub of sleep regulation, both 5-HT and GABA were significantly depleted in the INS group compared to all intervention groups (P < 0.05, Fig. 2h), reflecting central neurochemical disruption. All active treatments (Bb, LXP, Bb + LXP) comparably restored cerebral 5-HT and GABA levels (P < 0.05 vs. INS group; Fig. 2h), with no significant differences among the intervention groups (P > 0.05).
Collectively, these results demonstrate that PCPA-induced insomnia disrupts neurotransmitter homeostasis across peripheral and central compartments. While both Bbm-19 and lorazepam effectively restore brain 5-HT and GABA levels, Bbm-19 uniquely enhances neurotransmitter availability in the gut and systemic circulation, highlighting its potential as a gut-initiated modulator of the gut–brain axis.
Bbm-19 restores peripheral immune homeostasis in insomniac mice
Emerging evidence suggests that sleep disturbances can disrupt immune homeostasis. Systemic cytokine analysis after 42 days revealed significant immune dysregulation in PCPA-induced insomniac mice (INS group), characterized by distinct alterations relative to the HEL group (Fig. 2i): IL-9 and IL-1β levels decreased significantly (P < 0.05), IL-6 and IFN-γ increased significantly (P < 0.05), and a paradoxical elevation of anti-inflammatory IL-10 occurred (P < 0.05). Bbm-19 therapy (both Bb and Bb + LXP groups) effectively normalized IFN-γ, IL-1β, IL-10 and IL-6 to HEL group-comparable levels. In contrast, lorazepam treatment alone (LXP group) caused drastic decrease in IFN-γ while only partially restored the IL-10 level compared to the HEL (P < 0.05). These results establish Bbm-19 as both a restorer of insomnia-associated immune dysfunction and a modulator of drug-induced cytokine imbalances.Bbm-19 modulates gut microbiota composition and dynamics in insomniac mice
Accumulating evidence supports a critical role for the gut microbiota in regulating sleep and neurobehavioral health. To characterize the impact of PCPA-induced insomnia and Bbm-19 intervention on the murine gut microbiome, fecal samples were collected at days 14 and 42 and subjected to shotgun metagenomic sequencing. After quality filtering and removal of host-derived reads, an average of 41![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 538
538![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 654 ± 2
654 ± 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 869
869![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 953 clean reads were obtained per sample at day 14, and 41
953 clean reads were obtained per sample at day 14, and 41![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 397
397![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 482 ± 7
482 ± 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 384
384![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 312 at day 42, providing sufficient depth for species-level profiling.
312 at day 42, providing sufficient depth for species-level profiling.
Alpha diversity, assessed using the Shannon and Simpson indices, revealed a transient increase in microbial diversity in the Bb group compared to the HEL group at day 14 (P < 0.05, Fig. 3a and b), with no significant differences observed among other groups at either time point (days 14 and 42). Both indices showed a general trend of increasing over time, particularly in INS and LXP groups (Fig. 3c), suggesting time-dependent microbial succession influenced by physiological stress and pharmacological intervention. Principal coordinates analysis and similarity analysis based on Bray–Curtis distance confirmed persistent dysbiosis, demonstrating significant separation between the INS and HEL groups at both timepoints (Fig. 3d and e).
A total of 217 SGBs were reconstructed and taxonomically classified into 11 phyla, 31 families, and 85 genera (Table S1). Comparative analysis identified SGBs that were differentially abundant between the INS, HEL, and Bb groups, and the perturbations were time-dependent. At day 14, 28 SGBs differed significantly between INS and HEL, and 34 between the INS and Bb groups (Fig. 3f). Notably, members of the family Muribaculaceae, including Lepagella sp. 003979075 (SGB 142), CAG-873 sp. 009775265 (SGB 137), and Lepagella muris (SGB 91), were significantly enriched in the INS group. In contrast, other Muribaculaceae, such as Muribaculum arabinoxylanisolvens (SGB 11), Lepagella sp. 002361215 (SGB 183), and Muribaculum gordoncarteri (SGB 8), were markedly reduced. Within the Lachnospiraceae family, most SGBs were elevated in the INS group, except Suilimivivens sp. 910577275 (SGB 208) and Suilimivivens sp. 949878435 (SGB 79). Conversely, in the Bb group, nearly all SGBs of Lachnospiraceae showed increased abundance, except Petralouisia sp. 910576385 (SGB 37), with Suilimivivens sp. 910577275 (SGB 208) emerging as a shared differential taxon significantly enriched by Bbm-19 intake.
By day 42, 29 SGBs differed between the INS and HEL groups, and 19 between the INS and Bb groups (Fig. 3f and Tables S4, S5). The INS group exhibited a significant depletion of several members of the Muribaculaceae family, including Muribaculum arabinoxylanisolvens (SGB 11), Duncaniella sp. 910576785 (SGB 60), Muribaculum gordoncarteri (SGB 8), and Muribaculaceae bacterium (SGB 10). Additionally, Stercorousia sp. 910588555 (SGB 184), Bacteroides muris (SGB 44), and Parasutterella muris (SGB 86) were reduced. In contrast, taxa such as Lepagella sp. 002361215 (SGB 183), Butyribacter sp. 910586605 (SGB 104), Coproplasma sp. 910586065 and sp. 910579585 (SGBs 168, 124), and Odoribacter sp. 910578105 (SGB 191) were increased in the INS group. Importantly, Bbm-19 intervention significantly restored the abundance of Muribaculaceae bacterium (SGB 10), Odoribacter sp. 910578105 (SGB 191), Parasutterella muris (SGB 86), Coproplasma sp. 910586065 and sp. 910579585 (SGBs 168, 124), and Stercorousia sp. 910588555 (SGB 184), indicating its regulatory role in counteracting insomnia-associated dysbiosis. Notably, B. animalis subsp. lactis was significantly enriched in both the Bb and Bb + LXP groups, with abundance increasing progressively over time (P < 0.05; Fig. 3g). A comparison of the Bbm-19 strain with reconstructed SGBs revealed 99.99% genomic similarity to SGB 136, confirming successful intestinal colonization.
Longitudinal analysis revealed temporal dynamics in microbial abundance (Fig. S1a): SGBs such as Duncaniella sp. 910576785 (SGB 60), Muribaculum arabinoxylanisolvens (SGB 11), Enterococcus sp. 949082785 (SGB 68), Stercorousia sp. 910588555 (SGB184), Anaerococcus muris (SGB 201), RGIG8773 sp. 910584715 (SGB206), Parasutterella excrementihominis (SGB 214), and Pelethrinicoccus sp. 910579215 (SGB76) exhibited a time-dependent accumulation. In contrast, Lactobacillus sp. 910589675 (SGB 21), Bacteroides muris (SGB 44), Nanosyncoccus sp. 947570505 (SGB 38), Lepagella sp. 003979075 (SGB 142), and CAG-873 sp. 011959565 (SGB 17) declined over time. Notably, SGBs 184, 206, 68, and 17 showed the most pronounced temporal shifts, underscoring the dynamic nature of the gut microbiota in response to both disease and intervention.
Bbm-19 attenuates dysbiosis-associated microbial network hyperconnectivity in insomniac mice
Microbial co-occurrence network analysis revealed significant alterations in gut ecosystem dynamics associated with PCPA-induced insomnia and its treatment. Co-occurrence networks were constructed using Spearman correlation coefficients (|r| > 0.8, P < 0.01) between SGBs at days 14 and 42. The INS group exhibited significantly higher network connectivity density (0.026 at day 14; 0.035 at day 42) compared to the HEL (0.014, 0.032), Bb (0.012, 0.028), and LXP (0.014, 0.024) groups (Fig. S1b and S1c). Elevated connectivity in the INS group is consistent with a dysbiotic state characterized by increased microbial interdependencies, potentially reflecting stress-induced functional coupling or reduced ecological resilience. In contrast, the Bb and LXP groups maintained lower connectivity, suggesting a more functionally independent and stable microbial community. The combination group (Bb + LXP) displayed a unique temporal pattern: high connectivity at day 14 (0.032), followed by a decline at day 42 (0.030), indicating a shift toward network stabilization over time. Notably, microbial interaction density increased progressively in all groups except Bb + LXP, suggesting that combined intervention may mitigate the progressive dysregulation seen in untreated or monotherapy conditions.These findings indicate that chronic insomnia is associated with increased microbial co-occurrence, a hallmark of ecosystem instability. Bbm-19 administration reduces network complexity, supporting its role in promoting a more balanced and resilient gut microbiota. The distinct network trajectory of the combination group suggests that Bbm-19 may contribute to the ecological recovery of the gut microbiome during pharmacological treatment, although the underlying mechanisms require further investigation.
Bbm-19 reprograms gut–brain axis along tryptophan and glutamate metabolism
To investigate how Bbm-19 modulates microbial metabolic functions relevant to the gut–brain axis, gut metabolic modules were constructed for 217 SGBs using the MetaCyc and KEGG databases. A genome-centered metabolic network reconstruction strategy was employed to define 37 GBMs, functionally coherent sets of microbial pathways implicated in neuroactive metabolite production. Procrustes analysis revealed a significant correlation between microbial community structure and the predicted activity of GBMs (r2 > 0.51, P < 0.05; Fig. 4a), particularly at day 14, indicating strong coupling between taxonomic composition and metabolic potential during the early phase of intervention.PCPA-induced insomnia profoundly disrupted microbial metabolic pathways: at day 14, five GBMs showed differential abundance in the INS group versus the HEL group, including reduced levels of tryptophan synthesis, 17β-estradiol degradation, inositol degradation, and acetate degradation (Table S6). By day 42, dysregulation expanded to 14 GBMs (Table S7), featuring significant decreases in tryptophan degradation, GABA synthesis III, tryptophan synthesis, and glutamate synthesis II, alongside elevated inositol synthesis, nitric oxide degradation II (NO dioxygenase), and secondary bile acid biosynthesis I (Fig. 4b and S1d). These disturbances demonstrated progressive intensification over time, with pronounced impairments in tryptophan and glutamate metabolism, central hubs in the gut–brain axis.
Bbm-19 intervention counteracted these perturbations through time-dependent pathway modulation: at day 14, six differentially expressed GBMs emerged between Bb and INS groups (versus one in the LXP-INS comparison; Table S6), with probiotics significantly enhancing melatonin synthesis, GABA and glutamate degradation, and secondary bile acid biosynthesis, while reducing butyrate synthesis. By day 42, Bbm-19 substantially amplified glutamate degradation II and tryptophan degradation pathways compared to INS (Table S6 and Fig. S1d), exhibiting cumulative effects over the intervention period. In the LXP group, S-adenosylmethionine synthesis, propionate synthesis II, and corrinoid-dependent enzymes pathways were elevated, while indole-3-propionic acid synthesis and glutamate degradation II pathways were reduced (Table S7). Interestingly, in the combined treatment group (Bb + LXP), glutamate degradation II was restored (Fig. 4b and S1d), indicating that Bbm-19 counteracts the suppressing effect of lorazepam on this key catabolic pathway. To isolate the specific metabolic effects of Bbm-19, GBMs commonly differing between (i) Bb group vs. Bb + LXP group and (ii) LXP group vs. Bb + LXP group. At day 14, Bbm-19 was found to increase melatonin synthesis, tryptophan degradation, and glutamate degradation II (Fig. 4b). With prolonged intervention, these effects were sustained and intensified, particularly in tryptophan and glutamate degradation, pathways not significantly modulated by lorazepam alone.
Given that PCPA specifically targets 5-HT synthesis via inhibition of tryptophan hydroxylase, the dynamics of tryptophan degradation and synthesis pathway-encoding SGBs were analyzed. Such analysis confirmed specific enrichment of SGBs encoding tryptophan degradation enzymes by Bbm-19. While INS mice showed progressive depletion of these SGBs over 42 days, probiotic intervention significantly increased their abundance (Fig. S1e), directly linking Bbm-19 colonization to targeted modulation of tryptophan-GABA regulatory circuits central to sleep physiology.
Bbm-19 modulates gut amino acid metabolism in insomniac mice
Principal component analysis of fecal metabolites demonstrated high analytical stability, with QC samples forming tight clusters in both positive and negative ion modes (Fig. S2a and S2b). The RSD distribution of metabolite peak areas confirmed robust data quality, as 85% of metabolites exhibited RSD values below 15% (Fig. S2c and S2d). Following rigorous filtering (exclusion of metabolites with RSD >20%, >50% missing values, or low signal intensity in >80% samples), 2213 high-confidence metabolites were retained for analysis. Primary classification revealed dominant representation of lipid and lipid-like molecules, organoheterocyclic compounds, organic acids and derivatives and benzenoids (Fig. S2e). Secondary classification further resolved these into carboxylic acids and derivatives, prenol lipids, fatty acyls, benzene and substituted derivatives and steroids and steroid derivatives (Fig. S2f).Differential metabolite analysis (FDR < 0.05, |log2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) FC| > 1, VIP score >1) identified 25 downregulated and 34 upregulated metabolites in the INS group versus the HEL group at day 14 (Fig. 5a). Pathway enrichment analysis revealed significant perturbations in aldosterone synthesis and secretion, bile secretion, neurotrophin signaling, cortisol synthesis/secretion, and endocrine resistance pathways (Fig. S3), indicating early disruptions in neuroendocrine regulation and stress response pathways.
FC| > 1, VIP score >1) identified 25 downregulated and 34 upregulated metabolites in the INS group versus the HEL group at day 14 (Fig. 5a). Pathway enrichment analysis revealed significant perturbations in aldosterone synthesis and secretion, bile secretion, neurotrophin signaling, cortisol synthesis/secretion, and endocrine resistance pathways (Fig. S3), indicating early disruptions in neuroendocrine regulation and stress response pathways.
In contrast, the Bb group showed 50 downregulated and 136 upregulated metabolites compared to the HEL group at day 14 (Fig. 5a). Pathway enrichment analysis revealed that upregulated metabolites were predominantly associated with key metabolic and neuroactive pathways. Notably, amino acid metabolism was significantly enhanced, including tyrosine metabolism and the biosynthesis of branched-chain amino acids (valine, leucine, and isoleucine). Energy and carbon flux pathways such as alanine, aspartate, and glutamate metabolism, pyruvate metabolism, and glycolysis/gluconeogenesis were also upregulated, indicating increased microbial metabolic activity and host energy substrate availability (Fig. S3a).
Direct comparison of Bb versus INS groups showed seven downregulated and 24 upregulated metabolites (Fig. 5a), with pronounced upregulation of amino acid biosynthesis, lipoic acid metabolism, arginine and proline metabolism, and carbon metabolism (Fig. S3a). Downregulated pathways included neurotrophin signaling, leishmaniasis response, and adipocytokine signaling pathway (Fig. S3a), confirming an early activation of amino acid metabolic networks by Bbm-19 intake. By day 42, the number of differentially expressed metabolites decreased across groups (Fig. 5b), suggesting partial metabolic stabilization or adaptation. Compared to the HEL group, the INS group showed downregulation of phenylalanine, tyrosine, and tryptophan biosynthesis, alpha-linolenic acid metabolism, glycine, serine, and threonine metabolism, steroid biosynthesis, arginine and proline metabolism, and tryptophan metabolism (Fig. S3b), potentially reflecting a compensatory host response to accumulated metabolic stress or altered microbial metabolite load. These results suggest that prolonged PCPA-induced insomnia leads to progressive impairment of essential amino acid metabolic pathways in the gut.
Eighty-one metabolites were differentially abundant between the Bb and HEL groups on day 42 (Fig. 5b). Probiotic supplementation primarily downregulated pathways related to cofactor biosynthesis, nicotinate and nicotinamide metabolism, choline metabolism, vitamin B6 metabolism, and mineral absorption, with no significantly upregulated pathways (Fig. S3b). However, compared to the INS group, Bbm-19 intervention upregulated neurotrophin signaling, bile secretion, adipocytokine signaling, AGE-RAGE signaling, and necroptosis, while downregulating linoleic acid metabolism, and choline metabolism pathways (Fig. S3a and S3b). These shifts suggest that Bbm-19 redirects metabolic activity toward neural and immune modulation rather than nutrient biosynthesis over time.
Lorazepam also significantly altered fecal metabolism. At day 14, the LXP group showed upregulation of valine, leucine, and isoleucine biosynthesis, metabolism of nicotinate and nicotinamide, glycine, serine and threonine, propanoate, phenylalanine, caffeine, pyruvate, and tryptophan, among others (Fig. S3a), with most pathways linked to amino acid metabolism.
Compared to the LXP group, Bbm-19 supplementation further enhanced pantothenate and CoA biosynthesis, retrograde endocannabinoid signaling, carbohydrate digestion and absorption, starch and sucrose metabolism, bile acid biosynthesis, and vitamin digestion and absorption, while suppressing cortisol synthesis/secretion and Cushing syndrome pathways (Fig. S3a). As the intervention period lengthened, the Bb + LXP group exhibited increasing upregulation of amino acid metabolic pathways, including tryptophan, arginine and proline, alanine, aspartate and glutamate, and arginine biosynthesis (Fig. S3a and S3b). There was also significant upregulation of alpha-linolenic acid, and propanoate metabolism pathways, indicating synergistic metabolic reprogramming of amino acid, bile acid and neural-related pathways.
At day 42, the LXP group showed upregulation of retrograde endocannabinoid signaling, α-linolenic acid metabolism, nicotinate and nicotinamide metabolism, and multiple amino acid pathways, including glycine, serine, threonine, arginine, and proline metabolism, compared to the INS group (Fig. S3b). However, when compared to the LXP group, the Bb + LXP group showed upregulation in sphingolipid metabolism, metabolic pathways, neurotrophin signaling pathway, protein and carbohydrate digestion and absorption (Fig. S3b), suggesting that Bbm-19 fine-tunes or counterbalances the metabolic effects of lorazepam.
These results establish that Bbm-19 exerts time-dependent modulation of fecal amino acid metabolism, initially activating biosynthetic pathways (day 14) before shifting toward neural pathway regulation (day 42). Crucially, Bbm-19 demonstrates dual functionality: correcting insomnia-associated amino acid dysmetabolism while optimizing lorazepam-induced metabolic alterations through targeted pathway regulation.
Bbm-19 restores tryptophan, glutamate, organic acid, and bile acid metabolism in insomniac mice
Given the progressive metabolic and microbial changes observed over time, a targeted metabolomics analysis was conducted on fecal samples collected at day 42 to evaluate the long-term impact of Bbm-19 on key neuroactive metabolites (Fig. 5c). The results revealed significant changes in amino acid metabolism, including glutamine, tryptophan, glutamic acid, 5-HT, and GABA among the HEL, INS, and Bb groups (P < 0.05). In the INS group, fecal concentrations of tryptophan, glutamic acid, GABA and 5-HT were markedly reduced compared to the HEL group, consistent with the known disruption of serotonin synthesis by PCPA. Conversely, glutamine levels were elevated in the INS group, potentially reflecting a compensatory shift in nitrogen metabolism or impaired conversion to glutamate.Administration of Bbm-19 reversed these imbalances, partially restoring the levels of tryptophan, glutamic acid, GABA, and 5-HT levels, indicating that the probiotic enhances the availability of precursors and metabolites critical for neurotransmitter synthesis. Notably, tryptophan levels were significantly higher in the Bb + LXP group than in the Bb group (2.65 ± 0.35 μmol g−1vs. 5.49 ± 2.19 μmol g−1; P < 0.05), suggesting an additive or synergistic effect when Bbm-19 is combined with lorazepam.
In addition to amino acids, significant differences in organic acid profiles were observed. Compared to the HEL group, the Bb group showed increased fecal levels of malic acid, adipic acid, acrylic acid, gluconic acid, and lactic acid (Fig. 5c; P < 0.05). While no significant differences were detected between the Bb and INS groups for organic acids, a clear trend of accumulation was observed in both probiotic-treated groups (Bb and Bb + LXP), whereas the LXP group exhibited a general reduction in organic acid levels. This suggests that Bbm-19 promotes microbial production of organic acid, potentially contributing to gut homeostasis and metabolic signaling.
Regarding bile acids, the INS group exhibited elevated levels of cholic acid and glycoursodeoxycholic acid, with the latter showing a significant increase (P < 0.05; Fig. 5c). These changes may reflect altered bile acid metabolism under chronic insomnia. Notably, Bbm-19 intervention reduced the concentrations of these bile acids, bringing them closer to levels observed in the HEL group. This indicates that the probiotic helps normalize bile acid homeostasis, which may indirectly influence gut barrier function, inflammation, and neural signaling.
Collectively, these findings demonstrate that long-term supplementation with Bbm-19 significantly reshapes the intestinal metabolome, with a pronounced effect on the tryptophan-serotonin and glutamate-GABA pathways. The probiotic restores the availability of key neurotransmitter precursors and metabolites, outperforming lorazepam in modulating neuroactive amino acid derivatives. Additionally, Bbm-19 enhances organic acid production and normalizes bile acid profiles, further supporting its role in maintaining gut–brain axis integrity.
Integrated correlation analysis revealed the mechanism by which Bbm-19 improves sleep
Spearman correlation analysis was employed to investigate multidimensional interactions among sleep parameters, gut microbiota composition, microbial metabolites, serum cytokines, and neurotransmitter profiles in PCPA-induced insomnia mice (Fig. 6a). A significant positive correlation was observed between colonic and cerebral GABA levels (r = 0.67, P < 0.05). Similarly, cerebral 5-HT and GABA levels were positively correlated (r = 0.64, P < 0.05). Both cerebral and colonic neurotransmitter levels exhibited significant positive associations with sleep duration and specific cytokines including TNF-α, IL-1β, and IL-9 (r > 0.60, P < 0.05; Fig. 6a), while showing inverse relationships with anti-inflammatory cytokines (IL-4, IL-10). Notably, B. animals subsp. lactis (SGB 136), the administered probiotic strain, showed significant positive correlations with GABA and 5-HT levels in both the colon and brain (r > 0.55, P < 0.05). Two other SGBs, Muribaculaceae bacterium (SGB 10) and Stercorousia sp. 910588555 (SGB 184), were positively correlated with sleep duration (r > 0.53, P < 0.05). Conversely, these microbial species displayed a significant negative correlation with IFN-γ, IL-10, IL-4, and sleep latency (r < −0.64, P < 0.05).Ecological interactions among microbial taxa were also evident. SGB 10 was negatively associated with several potentially dysbiosis-associated taxa, including Lawsonibacter sp. 949840775 (SGB 161), Lachnospiraceae UBA3402 (SGB 130), Bilophila sp. 910585945 (SGB 106), Dysosmobacter sp. 947245315 (SGB 164), Butyribacter sp. 910586605 (SGB 104), and Lachnoclostridium sp. 910584265 (SGB 55) (r < −0.58, P < 0.05). Conversely, Nanosyncoccus sp. 947570505 (SGB 38) and Lepagella sp. 003979075 (SGB 142), both enriched in the INS group, exhibited a positive correlation with each other (r > 0.59, P < 0.05), suggesting a co-occurring dysbiotic cluster.
Notably, SGB 184 showed broad network of associations: negatively correlated with sleep latency and several low-abundance taxa (SGB 161, SGB 130, SGB 114, SGB164) (r < −0.59, P < 0.05), while positively correlated with colonic GABA and 5-HT, cerebral 5-HT, sleep duration, and the cytokines IL-9 and TNF-α (r > 0.54, P < 0.05).
Collectively, these correlations support a mechanistic model in which Bbm-19 intake is associated with reshaping the microbial community structure: increases in beneficial taxa, particularly B. animalis subsp. lactis (SGB 136), Muribaculaceae acteria (SGB 10), and Stercorus sp. 910588555 (SGB 184); decreases in potentially detrimental taxa such as Lepagella sp. 003979075 (SGB 142). These changes in turn modulate the levels of neuroactive metabolites (GABA, 5-HT) and systemic cytokine profiles. These peripheral changes culminate in elevated cerebral neurotransmitter levels, ultimately normalizing sleep patterns in the insomnia model (Fig. 6b).
Discussion
Bifidobacterium animalis subsp. lactis is a multifunctional probiotic with documented roles in modulating gut microbiota, supporting neurological health,3,4 and regulating immune function.5 In this study, we investigated Bbm-19, a strain isolated from healthy human breast milk and characterized by robust acid and bile salt tolerance, for its potential to ameliorate insomnia in a 42 day PCPA-induced murine model. Administered alone or in combination with lorazepam, Bbm-19 significantly improved sleep parameters, restored neurotransmitter balance, and normalized immune and microbial dysregulation. Through strain-resolved metagenomic and multi-omics analyses, we provide a comprehensive framework for understanding how this probiotic modulates the gut–brain axis to influence sleep.Mice with PCPA-induced insomnia exhibited hallmark behavioral and neurochemical deficits: reduced sleep duration, prolonged sleep latency, anxiety-like behaviors, and significant depletion of GABA and 5-HT in the gut, serum, and brain. These changes were accompanied by cytokine dysregulation, such as elevated IFN-γ, IL-6, IL-9, and IL-1β but decreased TNF-α. This pattern is indicative of a complex immune dysregulation commonly associated with sleep disruption. The elevation of the pro-inflammatory cytokines IL-6 and IFN-γ suggests a state of chronic low-grade inflammation, a well-documented feature in insomnia that contributes to hyperarousal and neuronal excitability.2,26 In addition, these findings align with clinical and preclinical evidence linking insomnia to impaired inhibitory neurotransmission and neuroimmune imbalance.16,27–29 Paradoxically, the concurrent rise in the anti-inflammatory cytokine IL-10 may represent a compensatory, yet insufficient, feedback mechanism attempting to curb the ongoing inflammation. The reduction in IL-1β, a key sleep-promoting cytokine, may directly reflect the disruption of sleep homeostasis.30,31 Similarly, the decrease in IL-9, a cytokine with diverse roles in T-cell function and neuronal survival, warrants further investigation as a potential novel player in sleep-related immune pathology. Collectively, this cytokine profile underscores that insomnia is not merely a neurological condition but one intertwined with systemic immune imbalance.
The mechanism PCPA, irreversible inhibition of tryptophan hydroxylase, directly explains the observed 5-HT deficiency. Notably, Bbm-19 intervention partially reversed these deficits, significantly improving sleep duration and latency while restoring GABA and 5-HT levels across peripheral and central compartments. Serum cytokine profiles were also normalized, indicating that Bbm-19 ameliorates both neurochemical and immune disruptions associated with chronic insomnia. These results are consistent with prior studies demonstrating the neuromodulatory potential of microbial interventions.16,32
The gut microbiota is increasingly recognized as a key regulator of brain function, often termed the “second brain” due to its extensive communication with the central nervous system.33–35 Germ-free zebrafish models exhibit impaired neuronal development in the absence of microbial colonization,36 underscoring the role of gut microbes in neurodevelopment. In Drosophila, sleep regulation via glial immune factors such as IM33 has been shown to be microbiota-dependent, with mono-colonization by Lactiplantibacillus plantarum reversing sleep deficits by increasing IM33 expression.37 This supports the concept of psychobiotics, specific bacterial strains capable of modulating neural pathways. Clinical evidence further reinforces this: an 8 week probiotic intervention in individuals with sleep and emotional symptoms significantly improved subjective sleep quality and mood, along with increased abundance of Bifidobacterium and Lactobacillus species.38 In this study, Bbm-19 administration led to a time-dependent increase in B. animalis subsp. lactis, with genomic binning confirming its identity as the administered strain (SGB 136), demonstrating successful gut colonization. This enrichment was associated with significant restructuring of the microbial community, including the expansion of beneficial taxa such as Muribaculaceae bacterium (SGB 10) and Stercoribacter sp. 910588555 (SGB 184), while reducing dysbiosis-associated species.
Microbial communities form complex ecological networks, and we report, for the first time, that PCPA-induced insomnia is associated with hyperconnectivity in gut bacterial co-occurrence networks, indicative of heightened interdependencies that may reflect ecosystem instability. This aberrant network state was attenuated by both lorazepam and Bbm-19 administration, which shifted the microbial interactome toward a more balanced and resilient configuration. Functional annotation revealed that insomnia significantly suppressed key metabolic modules linked to neuroactive metabolite production, including tryptophan synthesis and degradation, GABA synthesis III, and glutamate synthesis II. Given that enterochromaffin cells produce over 90% of the body's 5-HT,39,40 microbial tryptophan metabolism is pivotal for host serotonergic tone. Bacterial conversion of tryptophan to indole derivatives, via tryptophanase, leads to hepatic formation of indole sulfates, which activate the aryl hydrocarbon receptor in intestinal cells, promoting expression of host tryptophan hydroxylase 1 and subsequent 5-HT synthesis9,41 (Fig. 6b). Similarly, microbial GABA synthesis via glutamate decarboxylase (GadB/C) contributes to host GABA pools, potentially influencing central inhibition through vagal or systemic signaling.42 Research has found that some of the neurotransmitters (5-HT) produced by gut microbiota can enter the brain directly via the vagus nerve,43,44 where they participate in the regulation of emotions, cognition and physiological functions. Another portion, consisting of 5-HT and GABA, is absorbed into the portal venous circulation and transported to the brain via the bloodstream.45 Further research is needed to determine the specific pathways through which GABA and 5-HT influence brain function. Integrated metabolomic analyses confirmed that PCPA significantly depleted luminal and systemic levels of glutamate, tryptophan, GABA, and 5-HT. Bbm-19 intervention restored these metabolites and upregulated the corresponding microbial pathways, particularly tryptophan and glutamate metabolism, activities that worsened progressively in untreated mice, highlighting the chronic nature of the dysregulation and the therapeutic value of sustained probiotic treatment. In contrast to Bbm-19, lorazepam intervention had no significant impact on GABA levels in the colon, indicating that lorazepam does not directly act on the gut. Lorazepam enhances the affinity of the central inhibitory neurotransmitter GABA for its receptors by binding to specific sites on GABA_A receptors. This increases the frequency with which chloride channels open, leading to neuronal hyperpolarisation and producing sedative and sleep-promoting effects.
Spearman correlation analysis integrated these multi-omics findings into a unified mechanistic model (Fig. 6b). Bifidobacterium animalis subsp. lactis (SGB 136) showed strong positive correlations with GABA and 5-HT levels in the colon and brain, as well as with sleep duration, while being negatively correlated with sleep latency and pro-inflammatory cytokines (IL-9, TNF-α, IFN-γ, IL-66). These associations were mirrored by Muribaculaceae bacterium (SGB 10) and Stercoribacter sp. 910588555 (SGB 184), which also correlated improved neurochemical and behavioral outcomes. Notably, these beneficial taxa exhibited negative correlations with dysbiosis-linked species, suggesting that Bbm-19 promotes ecosystem stability through selective enrichment. Collectively, these findings support a tripartite mechanism by which Bbm-19 improves sleep: (1) direct modulation of GABAergic and serotonergic pathways, (2) restructuring of the gut microbial ecosystem to favor neuroactive metabolite-producing taxa, and (3) rebalancing of systemic immune responses to reduce neuroinflammation.
Importantly, this multi-targeted, gut-centric mechanism distinguishes Bbm-19 from conventional pharmacological agents like lorazepam, which primarily act on central GABA receptors without influencing gut microbial composition or peripheral neurotransmitter dynamics and immune responses. The synergy observed between Bbm-19 and lorazepam further suggests that probiotic co-administration may enhance the efficacy and safety profile of sedative drugs. Collectively, these results position the gut microbiome as a novel therapeutic target for insomnia and highlight the unique capacity of Bbm-19 to improve sleep through integrated microbiota–neuroimmune modulation.
This study leverages a longitudinal, multi-omics approach to dissect the strain-specific effects of Bbm-19 on the gut–brain axis, combining behavioral, neurochemical, immunological, and microbial functional data. The use of a well-defined insomnia model, combined with strain-level metagenomic resolution and targeted metabolomics, enhances the mechanistic depth and reproducibility of our findings. A key strength lies in the integration of correlation networks to link microbial dynamics with host outcomes, providing a systems-level view of probiotic action. However, some limitations should be acknowledged. PCPA, as a substance that inhibits 5-HT synthesis, is currently the classic method for preparing animal models of insomnia. However, since the PCPA model only simulates the single pathway of 5-HT deficiency leading to insomnia,46,47 it cannot encompass the complex etiology and pathophysiological processes of human insomnia. Furthermore, 5-HT levels in animals remain suppressed for at least 7 days, with near-complete recovery observed after 9 days.48 Therefore, the findings of this study cannot be generalized to all types of insomnia. While these methods are well-validated for measuring gross sleep parameters such as total sleep time and latency, they do not provide the nuanced data on sleep microstructure that EEG/EMG affords. In addition, while we demonstrate strong associations between Bbm-19, microbial shifts, and neurochemical improvements, the exact molecular mediators (such as specific microbial metabolites or host receptors) require further validation through gnotobiotic models or in vitro co-culture systems. Additionally, the role of the vagus nerve in transmitting gut-derived signals was not directly tested and remains an open question.
Future studies should validate these findings in human cohorts, particularly individuals with insomnia and gut dysbiosis. Investigating the effects of Bbm-19 in combination with cognitive behavioral therapy or low-dose pharmacotherapy could inform personalized, microbiome-targeted treatment strategies. Furthermore, exploring the epigenetic and transcriptional changes in brain regions involved in sleep regulation may uncover additional layers of gut–brain communication.
Conclusions
This study demonstrates that chronic administration of Bbm-19 effectively alleviates PCPA-induced insomnia in mice through multi-level modulation of the gut–brain axis. Bbm-19 improves sleep parameters, reduces anxiety, and enhances survival, outperforming lorazepam in restoring gut-specific metabolic functions. The probiotic reshapes the gut microbiota, enriches beneficial taxa, and reprograms microbial metabolic modules related to tryptophan and glutamate metabolism, leading to increased synthesis of GABA and 5-HT in both the gut and brain. Concurrently, Bbm-19 attenuates systemic inflammation and stabilizes microbial interaction networks. Its synergistic effects with lorazepam further highlight its therapeutic potential. Together, these findings establish Bbm-19 as a strain-specific, microbiome-targeted strategy for sleep regulation, offering a promising alternative or adjunct to conventional pharmacological treatments.Author contributions
Qiong Wu: writing – original draft. Guangqi Gao: review & editing. Lai-Yu Kwok: review & editing. Jiaqi Qiao: investigation. Ziqing Wei: investigation. Qiuwen He: project administration Zhihong Sun: conceptualization, project administration.Conflicts of interest
The authors declare no competing interests.Ethics approval and consent to participate
All experimental procedures involving animals were approved by the Experimental Animal Ethics Committee of Inner Mongolia Agricultural University (Approval code: NND2025161) and conducted in accordance with the ARRIVE guidelines 2.0 for the care and use of laboratory animals.Consent for publication
All authors have reviewed and approved the final version of the manuscript and consent to its publication.Data availability
Raw metagenomic are deposited in the Genome Sequence Archive (GSA) at the National Genomics Data Center (NGDC), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession number PRJCA043152. Additional supporting data are included in the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5fo04374c.Requests for materials should be directed to the corresponding author.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 32325040 and U25A20733), the National Key Research and Development Program of China (Grant No. 2022YFD2100702), the Inner Mongolia Agricultural University First-Class Discipline Scientific Research Special Program (YLXKZX-NND-006), and the Earmarked Fund for China Agriculture Research System (CARS36).References
- B. Karna, A. Sankari and G. Tatikonda, in StatPearls, StatPearls Publishing. Copyright © 2025, StatPearls Publishing LLC., Treasure Island (FL) ineligible companies. Disclosure: Abdulghani Sankari declares no relevant financial relationships with ineligible companies. Disclosure: Geethika Tatikonda declares no relevant financial relationships with ineligible companies, 2025 Search PubMed.
- Q. Wu, G. Gao, L.-y. Kwok, H. Lv and Z. Sun, Insomnia: the gut microbiome connection, prospects for probiotic and postbiotic therapies, and future directions, J. Adv. Res., 2025, 10:S2090-1232(25), 00503–X, DOI:10.1016/j.jare.2025.07.005.
- J. Cao, W. K. Amakye, C. Qi, X. Liu, J. Ma and J. Ren, Bifidobacterium Lactis Probio-M8 regulates gut microbiota to alleviate Alzheimer's disease in the APP/PS1 mouse model, Eur. J. Nutr., 2021, 60, 3757–3769 CrossRef CAS PubMed.
- Z. Miao, L. Chen, Y. Zhang, J. Zhang and H. Zhang, Bifidobacterium animalis subsp. lactis Probio-M8 alleviates abnormal behavior and regulates gut microbiota in a mouse model suffering from autism, mSystems, 2024, 9, e0101323 CrossRef PubMed.
- C. Lin, Y. Lin, S. Wang, J. Wang, X. Mao, Y. Zhou, H. Zhang, W. Chen and G. Wang, Bifidobacterium animalis subsp. lactis boosts neonatal immunity: unravelling systemic defences against Salmonella, Food Funct., 2024, 15, 236–254 RSC.
- S. P. Tsao, T. H. Yeh, Y. T. Lin, C. H. Pan, Y. K. Lee, C. H. Wu and H. Y. Huang, Supplementation with Bifidobacterium animalis subsp. lactis MH-022 for remission of motor impairments in a 6-OHDA-induced Parkinson's disease rat model by reducing inflammation, reshaping the gut microbiome, and fostering specific microbial taxa, Food Funct., 2024, 15, 9368–9389 RSC.
- R. Nocerino, F. De Filippis, G. Cecere, A. Marino, M. Micillo, C. Di Scala, C. de Caro, A. Calignano, C. Bruno, L. Paparo, A. M. Iannicelli, L. Cosenza, Y. Maddalena, G. Della Gatta, S. Coppola, L. Carucci, D. Ercolini and R. Berni Canani, The therapeutic efficacy of Bifidobacterium animalis subsp. lactis BB-12(®) in infant colic: A randomised, double blind, placebo-controlled trial, Aliment. Pharmacol. Ther., 2020, 51, 110–120 CrossRef CAS PubMed.
- S. M. O'Mahony, G. Clarke, Y. E. Borre, T. G. Dinan and J. F. Cryan, Serotonin, tryptophan metabolism and the brain-gut-microbiome axis, Behav. Brain Res., 2015, 277, 32–48 CrossRef PubMed.
- A. Agus, J. Planchais and H. Sokol, Gut Microbiota Regulation of Tryptophan Metabolism in Health and Disease, Cell Host Microbe, 2018, 23, 716–724 CrossRef CAS PubMed.
- T. Vanuytsel, S. van Wanrooy, H. Vanheel, C. Vanormelingen, S. Verschueren, E. Houben, S. Salim Rasoel, J. Tóth, L. Holvoet, R. Farré, L. Van Oudenhove, G. Boeckxstaens, K. Verbeke and J. Tack, Psychological stress and corticotropin-releasing hormone increase intestinal permeability in humans by a mast cell-dependent mechanism, Gut, 2014, 63, 1293–1299 CrossRef CAS PubMed.
- H. J. Zheng, J. Guo, Q. Jia, Y. S. Huang, W. J. Huang, W. Zhang, F. Zhang, W. J. Liu and Y. Wang, The effect of probiotic and synbiotic supplementation on biomarkers of inflammation and oxidative stress in diabetic patients: A systematic review and meta-analysis of randomized controlled trials, Pharmacol. Res., 2019, 142, 303–313 CrossRef CAS PubMed.
- V. Nobile, S. Giardina and F. Puoci, The Effect of a Probiotic Complex on the Gut-Brain Axis: A Translational Study, Neuropsychobiology, 2022, 81, 116–126 CrossRef PubMed.
- A. Marotta, E. Sarno, A. Del Casale, M. Pane, L. Mogna, A. Amoruso, G. E. Felis and M. Fiorio, Effects of Probiotics on Cognitive Reactivity, Mood, and Sleep Quality, Front. Psychiatry, 2019, 10, 164 CrossRef PubMed.
- H. Yu, H. Yu, L. Si, H. Meng, W. Chen, Z. Wang and A. Gula, Influence of warm acupuncture on gut microbiota and metabolites in rats with insomnia induced by PCPA, PLoS One, 2022, 17, e0267843 CrossRef CAS PubMed.
- Y. M. Liu, J. C. Li, Y. F. Gu, R. H. Qiu, J. Y. Huang, R. Xue, S. Li, Y. Zhang, K. Zhang and Y. Z. Zhang, Cannabidiol Exerts Sedative and Hypnotic Effects in Normal and Insomnia Model Mice Through Activation of 5-HT(1A) Receptor, Neurochem. Res., 2024, 49, 1150–1165 CrossRef CAS PubMed.
- L. Yao, J. Lv, C. Duan, X. An, C. Zhang, D. Li, C. Li and S. Liu, Armillaria mellea fermentation liquor ameliorates p-chlorophenylalanine-induced insomnia associated with the modulation of serotonergic system and gut microbiota in rats, J. Food Biochem., 2022, 46, e14075 CAS.
- M. Ota, Y. Maki, L. Y. Xu and T. Makino, Prolonging effects of Valeriana fauriei root extract on pentobarbital-induced sleep in caffeine-induced insomnia model mice and the pharmacokinetics of its active ingredients under conditions of glycerol fatty acid ester as emulsifiers, J. Ethnopharmacol., 2022, 298, 115625 CrossRef CAS PubMed.
- B. Huang, W. Zhao, X. Cai, Y. Zhu, Y. Lu, J. Zhao, N. Xiang, X. Wang, H. Deng, X. Tang, L. Liu, Y. Zhao and Y. Shi, Expression and Activity of the Transcription Factor CCAAT/Enhancer-Binding Protein β (C/EBPβ) Is Regulated by Specific Pulse-Modulated Radio Frequencies in Oligodendroglial Cells, Int. J. Mol. Sci., 2023, 24, 11131 CrossRef CAS PubMed.
- X. Tang, L. Yang, N. F. Fishback and L. D. Sanford, Differential effects of lorazepam on sleep and activity in C57BL/6J and BALB/cJ strain mice, J. Sleep Res., 2009, 18, 365–373 CrossRef PubMed.
- S. Porecha, T. Shah, V. Jogani, S. Naik and A. Misra, Microemulsion based intranasal delivery system for treatment of insomnia, Drug Delivery, 2009, 16, 128–134 CrossRef CAS PubMed.
- J. Röschke and J. B. Aldenhoff, A nonlinear approach to brain function: deterministic chaos and sleep EEG, Sleep, 1992, 15, 95–101 CrossRef PubMed.
- A. Lin, C.-T. Shih, C.-L. Huang, C.-C. Wu, C.-T. Lin and Y.-C. Tsai, Hypnotic Effects of Lactobacillus fermentum PS150TM on Pentobarbital-Induced Sleep in Mice, Nutrients, 2019, 11, 2409 CrossRef CAS PubMed.
- L. Yu, X. Han, S. Cen, H. Duan, S. Feng, Y. Xue, F. Tian, J. Zhao, H. Zhang, Q. Zhai and W. Chen, Beneficial effect of GABA-rich fermented milk on insomnia involving regulation of gut microbiota, Microbiol. Res., 2020, 233, 126409 CrossRef PubMed.
- Y. Darzi, G. Falony, S. Vieira-Silva and J. Raes, Towards biome-specific analysis of meta-omics data, ISME J., 2016, 10, 1025–1028 CrossRef CAS PubMed.
- H. Mallick, E. A. Franzosa, L. J. McLver, S. Banerjee, A. Sirota-Madi, A. D. Kostic, C. B. Clish, H. Vlamakis, R. J. Xavier and C. Huttenhower, Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences, Nat. Commun., 2019, 10, 3136 CrossRef PubMed.
- Z. Wang, W.-H. Chen, S.-X. Li, Z.-M. He, W.-L. Zhu, Y.-B. Ji, Z. Wang, X.-M. Zhu, K. Yuan, Y.-P. Bao, L. Shi, S.-Q. Meng, Y.-X. Xue, W. Xie, J. Shi, W. Yan, H. Wei, L. Lu and Y. Han, Gut microbiota modulates the inflammatory response and cognitive impairment induced by sleep deprivation, Mol. Psychiatry, 2021, 26, 6277–6292 CrossRef CAS PubMed.
- T. Gao, Z. Wang, Y. Dong, J. Cao, R. Lin, X. Wang, Z. Yu and Y. Chen, Role of melatonin in sleep deprivation-induced intestinal barrier dysfunction in mice, J. Pineal Res., 2019, 67, e12574 CrossRef PubMed.
- J. W. Winkelman, O. M. Buxton, J. E. Jensen, K. L. Benson, S. P. O'Connor, W. Wang and P. F. Renshaw, Reduced brain GABA in primary insomnia: preliminary data from 4T proton magnetic resonance spectroscopy (1H-MRS), Sleep, 2008, 31, 1499–1506 CrossRef PubMed.
- H. Ren, X. Kong, Y. Zhang, F. Deng, J. Li, F. Zhao, P. Li, K. Pei, J. Tan, Y. Cheng, Y. Wang, L. Zhang, Y. Wang and X. Hao, The therapeutic potential of Ziziphi Spinosae Semen and Polygalae Radix in insomnia management: Insights from gut microbiota and serum metabolomics techniques, J. Ethnopharmacol., 2024, 330, 118255 CrossRef CAS PubMed.
- T. Kubota, T. Kushikata, J. Fang and J. M. Krueger, Nuclear factor-kappaB inhibitor peptide inhibits spontaneous and interleukin-1beta-induced sleep, Am. J. Physiol.: Regul., Integr. Comp. Physiol., 2000, 279, R404–R413 CrossRef CAS PubMed.
- R. Zefferino, S. Di Gioia and M. Conese, Molecular links between endocrine, nervous and immune system during chronic stress, Brain Behav., 2021, 11, e01960 CrossRef CAS PubMed.
- Y. J. Dong, N. H. Jiang, L. H. Zhan, X. Teng, X. Fang, M. Q. Lin, Z. Y. Xie, R. Luo, L. Z. Li, B. Li, B. B. Zhang, G. Y. Lv and S. H. Chen, Soporific effect of modified Suanzaoren Decoction on mice models of insomnia by regulating Orexin-A and HPA axis homeostasis, Biomed. Pharmacother., 2021, 143, 112141 CrossRef CAS PubMed.
- K. Socała, U. Doboszewska, A. Szopa, A. Serefko, M. Włodarczyk, A. Zielińska, E. Poleszak, J. Fichna and P. Wlaź, The role of microbiota-gut-brain axis in neuropsychiatric and neurological disorders, Pharmacol. Res., 2021, 172, 105840 CrossRef PubMed.
- S. G. Sorboni, H. S. Moghaddam, R. Jafarzadeh-Esfehani and S. Soleimanpour, A Comprehensive Review on the Role of the Gut Microbiome in Human Neurological Disorders, Clin. Microbiol. Rev., 2022, 35, e0033820 CrossRef PubMed.
- J. F. Cryan, K. J. O'Riordan, K. Sandhu, V. Peterson and T. G. Dinan, The gut microbiome in neurological disorders, Lancet Neurol., 2020, 19, 179–194 CrossRef CAS PubMed.
- J. J. Bruckner, S. J. Stednitz, M. Z. Grice, D. Zaidan, M. S. Massaquoi, J. Larsch, A. Tallafuss, K. Guillemin, P. Washbourne and J. S. Eisen, The microbiota promotes social behavior by modulating microglial remodeling of forebrain neurons, PLos Biol., 2022, 20, e3001838 CrossRef CAS PubMed.
- W. Xu, J. Rustenhoven, C. A. Nelson, T. Dykstra, A. Ferreiro, Z. Papadopoulos, C. D. Burnham, G. Dantas, D. H. Fremont and J. Kipnis, A novel immune modulator IM33 mediates a glia-gut-neuronal axis that controls lifespan, Neuron, 2023, 111(20), 3244–3254 CrossRef CAS PubMed.
- H. H. Y. Chan, P. L. K. Siu, C. T. Choy, U. K. Chan, J. Zhou, C. H. Wong, Y. W. Lee, H. W. Chan, J. C. C. Tsui, S. K. F. Loo and S. K. W. Tsui, Novel Multi-Strain E3 Probiotic Formulation Improved Mental Health Symptoms and Sleep Quality in Hong Kong Chinese, Nutrients, 2023, 15(24), 5037 CrossRef CAS PubMed.
- M. S. Shajib, A. Baranov and W. I. Khan, Diverse Effects of Gut-Derived Serotonin in Intestinal Inflammation, ACS Chem. Neurosci., 2017, 8, 920–931 CrossRef CAS PubMed.
- J. M. Yabut, J. D. Crane, A. E. Green, D. J. Keating, W. I. Khan and G. R. Steinberg, Emerging Roles for Serotonin in Regulating Metabolism: New Implications for an Ancient Molecule, Endocr. Rev., 2019, 40, 1092–1107 CrossRef PubMed.
- H. M. Roager and T. R. Licht, Microbial tryptophan catabolites in health and disease, Nat. Commun., 2018, 9, 3294 CrossRef PubMed.
- I. B. A. Yogeswara, S. Maneerat and D. Haltrich, Glutamate Decarboxylase from Lactic Acid Bacteria-A Key Enzyme in GABA Synthesis, Microorganisms, 2020, 8(12), 1923 CrossRef CAS PubMed.
- E. J. Kang, M. G. Cha, G. H. Kwon, S. H. Han, S. J. Yoon, S. K. Lee, M. E. Ahn, S. M. Won, E. H. Ahn and K. T. Suk, Akkermansia muciniphila improve cognitive dysfunction by regulating BDNF and serotonin pathway in gut-liver-brain axis, Microbiome, 2024, 12, 181 CrossRef CAS PubMed.
- Y. B. Lee, Y. J. Cho and J. K. Kim, The unique role of fluoxetine in alleviating depression and anxiety by regulating gut microbiota and the expression of vagus nerve-mediated serotonin and melanocortin-4 receptors, Biomed. Pharmacother., 2025, 182, 117748 CrossRef CAS PubMed.
- C. R. Martin, V. Osadchiy, A. Kalani and E. A. Mayer, The Brain-Gut-Microbiome Axis, Cell. Mol. Gastroenterol. Hepatol., 2018, 6, 133–148 CrossRef PubMed.
- A. Dias Elpo Zomkowski, A. Oscar Rosa, J. Lin, A. R. Santos, J. B. Calixto and A. Lúcia Severo Rodrigues, Evidence for serotonin receptor subtypes involvement in agmatine antidepressant like-effect in the mouse forced swimming test, Brain Res., 2004, 1023, 253–263 CrossRef PubMed.
- N. M. Murray, G. F. Buchanan and G. B. Richerson, Insomnia Caused by Serotonin Depletion is Due to Hypothermia, Sleep, 2015, 38, 1985–1993 CrossRef PubMed.
- I. B. Deryabina, L. N. Muranova, V. V. Andrianov and K. L. Gainutdinov, Impairing of Serotonin Synthesis by P-Chlorphenylanine Prevents the Forgetting of Contextual Memory After Reminder and the Protein Synthesis Inhibition, Front. Pharmacol., 2018, 9, 607 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |