Open Access Article
Open Access ArticlePhenotypic profiling reveals polystyrene nanoplastics elicit sublethal and lethal effects on cellular morphology in rainbow trout gill epithelial cells†
Lissett G.
Diaz
 and
Rebecca
Klaper
and
Rebecca
Klaper
 *
*
School of Freshwater Sciences, University of Wisconsin-Milwaukee, Milwaukee, Wisconsin, USA. E-mail: rklaper@uwm.edu
First published on 24th January 2025
Abstract
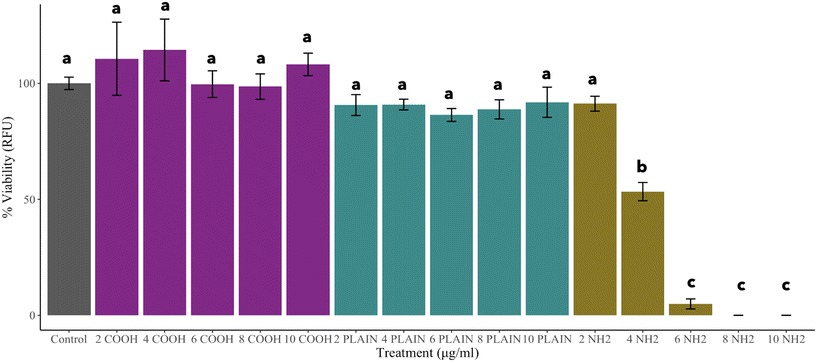
Extensive knowledge is available on the impacts of both engineered nanomaterials (ENMs) and microplastics (MPs), yet there remains a critical gap in understanding the impacts of nanoplastics, and the cellular and subcellular effects at sublethal concentrations. This study investigates the impacts of polystyrene nanoplastics (PS NPs) on Oncorhynchus mykiss (rainbow trout) gill epithelial cells, emphasizing the crucial role of surface charge in nano–bio interactions. The current study employs both traditional and non-traditional toxicological techniques presenting an image-based study to examine PS NP–cellular interactions at sublethal and lethal doses. Our findings demonstrated that relative to the uncharged and negatively charged PS NPs, the positively charged PS NPs significantly decreased cell viability at 4 μg mL−1 (EC50: 4.31 μg mL−1). However, at a sublethal concentration of 2 μg mL−1, phenotypic profiling analysis indicates that positively charged PS NPs elicit a significant change to cellular morphology and suggests key interactions with subcellular components. As the impacts measured are novel, further research into the underlying mechanisms will contribute to our understanding of nanoparticle toxicity in vertebrate species guiding both the policy and sustainable design of nanoparticles.
Environmental significanceThe presence of plastics in aquatic ecosystems has emerged as a worldwide concern due to production rates and inadequate waste management. Of particular concern are nanoscale plastics which can enter aquatic ecosystems either directly or through the degradation of macro- and microscale plastics already present. Their unique risks stem from their ability to penetrate biological barriers and acquire distinct properties that can influence their toxicity. This work explores the role that surface charge plays in mechanistic pathways underlying the toxicity of nanoplastics in rainbow trout gill epithelia by emphasizing the relevance and applicability of image-based profiling techniques for assessing environmental impacts in aquatic vertebrates. More importantly, model particles are used in this study to isolate these key mechanisms. |
1. Introduction
Synthetic polymers are man-made materials that are used extensively in many commercial products due to their diverse physical and chemical properties and have provided numerous societal benefits.1 However, their unsustainable use and inappropriate waste management has led to the pervasive accumulation of plastics in the environment.2,3 Due to the abundance of plastics and their slow degradation process, it is necessary to address their environmental impacts. During the degradation process, macroplastics (larger than 5 mm in size) break down and fragment into micro (<5 mm) and nano (<1000 nm) size particles due to both abiotic and biotic processes such as UV-radiation and bacterial degradation, respectively.4 In the environment, MPs behave differently than NPs in biological systems. NPs have a large surface area-to-volume ratio resulting in various changes to their physical properties (such as size, shape, and surface chemistry) that can influence their toxicity in aquatic organisms and can subsequently lead to detrimental ecosystem impacts.4,5Among the diverse chemical properties of NPs, research has shown that surface charge plays a major role in interactions of NPs at the nano–bio interface. These studies have been conducted in multiple cell types and plants, and on whole organisms,6–14 and have characterized the impacts of both anionic and cationic NPs. However, a multitude of these studies highlight the acute effects at lethal doses without attributing as much attention to the potential negative effects at sublethal concentrations. It is imperative to study mechanisms of toxicity in various particle types for the development of sustainable nanotechnologies that can minimize harm to ecosystems and human health under sublethal exposure conditions.
In environmentally relevant studies concerning human and ecosystem health, changes in organelle and subcellular structures have been reported to be disrupted by ENMs and PS NPs.9,15–22 The mitochondrion, an extensively studied organelle, is responsible for the synthesis of ATP (adenosine triphosphate) and amino acid, lipid metabolism, and apoptosis and can partake in cellular signaling.23 Evidence suggests that internalized NPs can contribute to mitochondrial damage, cytotoxicity, the generation of ROS (reactive oxygen species), and mitochondrial dysfunction. Changes to cytoskeletal organization in cells have also been observed in response to NP exposure.15,24–26 The cytoskeleton plays a large role in both the movement and the structure of the cell to enable the mechanical support to conduct essential cellular functions.27 Ma et al., 2017, found that mitochondrial damage can also be induced, even at sublethal concentrations suggesting morphological changes to sub-cellular compartments subsequently effecting cytoskeleton-associated signaling.15 Furthermore, in addition to cellular disorganization, NP treatment has been associated with cell shrinkage and nuclear condensation.25,26 The nucleus, a ubiquitous eukaryotic cell component, plays crucial roles in cellular metabolism, survival and death.28 Despite these insights, there remains a critical knowledge gap in understanding how PS NPs perturb mitochondrial-related response pathways at sublethal concentrations leading to morphological changes. Addressing this gap is essential for comprehensively assessing the adverse effects of PS NPs of varying surface charge on the cellular function and morphology.
To explore the role of surface charge, a variety of methods have been developed to decipher biological responses of NPs and various existing environmental contaminants. With technological advancements, phenotypic profiling coupled with high-content imaging (HCI), both high-throughput methods, have been commonly employed in drug discovery and delivery models.29–32 More importantly, while traditional biological assays such as ROS and cell viability offer valuable insights,33,34 HCI is used to identify trends or patterns in specific measured features (e.g., cell size, cell count, nuclear intensity) in cellular morphology. In contrast, phenotypic profiling is an unbiased robust tool that enables the ability to capture hundreds of features (e.g., granularity, texture) at the resolution of a single cell and can often be coupled with machine learning techniques to score similarities and differences across a relevant phenotype.32 While these methods have primarily found applications in drug discovery and delivery, the U.S. Environmental Protection Agency (EPA) advocates for the use of high-throughput screening that can identify cellular perturbations and can inform mechanisms of toxicity in environmentally relevant organisms.35
Here, we aimed to employ both traditional toxicology and automated fluorescence microscopy methods to investigate the cellular and subcellular impacts of polystyrene nanoparticles. An epithelial cell line (RTgill-W1) from rainbow trout gills, Oncorhynchus mykiss, was selected as the in vitro ecotoxicological model for this study since the gills represent the first point of contact for nanoparticle exposure in most aquatic vertebrates. We compared the effects across three model particle types of different functional groups representing positive, negative and neutral surface charges, since the nanoparticle surface charge is well-established as an important factor in determining biological impacts. The effects of PS NPs on cell viability and the morphology of cellular structures (nuclei, mitochondria, actin cytoskeleton) were investigated. Additionally, phenotypic profiling was conducted to identify significant morphological changes after particle treatment and separating their responses by concentration with respect to the control. While the nanoplastics in this study were used as models to ascertain mechanisms of nanoplastic toxicity, it should be noted that these particular nanoplastics do not represent all of the nanoplastics found in the environment due to varying sizes, shapes, compositions, and surface chemistries.
2. Materials and methods
2.1 Polystyrene nanoparticle characterization and preparation
Commercial nanoparticles used were 25 nm DiagPoly™ plain green fluorescent polystyrene particles (PS-Plain), 25 nm DiagPoly™ carboxyl green fluorescent polystyrene particles (PS-COOH), and 25 nm DiagPoly™ amine green fluorescent polystyrene particles (PS-NH2) (CD Bioparticles). All the commercial NP dispersions were custom ordered without surfactants and were stored at 4 °C. Before each experiment, the NP dispersions were sonicated for 45 minutes and immediately vortexed for 10 minutes prior to their use. All experiments were performed using the same batch of NPs.The characterization of each polystyrene nanoparticle was carried out using Dynamic Light Scattering (DLS) with a Zetasizer Nano device (Malvern PANalytical, United Kingdom) using 10 × 10 × 45 mm polystyrene cuvettes (SARSTEDT, 67.754). Size measurements and zeta potential measurements were performed in UltraPure™ distilled water (Invitrogen), phenol red- and fetal bovine serum (FBS)-free Leibovitz's L-15 growth medium (Gibco), and phosphate buffered saline (PBS, Gibco). Three separate measurements including 13 runs per measurement were conducted for both size and zeta potential at room temperature. The final particle concentration was 4 μg mL−1. Additionally, the mean particle size of each particle after 24 hours was quantified via bioimaging analysis further described below.
2.2 Routine cell culture
Oncorhynchus mykiss (rainbow trout) gill epithelial cells (RTgill-W1, ATCC CRL-2523) were routinely cultured in 75 cm2 cell culture flasks (Millipore Sigma) in a phenol red Leibovitz's L-15 medium (ATCC) and supplemented with 1% penicillin–streptomycin (Gibco, ThermoFisher Scientific) and 10% FBS (Gibco). The flasks were incubated at 19 °C with the growth medium replaced twice weekly. Following 85% confluency, the cells were passaged and harvested for experiments. The growth medium was aspirated, and the cells were rinsed with PBS (ThermoFisher) and were dispersed using 0.25% trypsin–0.53 mM EDTA solution (ATCC).2.3 Cell viability
Following the NP preparation previously mentioned, 20 μg mL−1 solutions were prepared in a phenol red- and FBS-free L-15 growth medium by using particle solutions from the 10 mg mL−1 PS-NH2, and 1 mg mL−1 PS-COOH and PS-Plain manufacturer vials. This was followed immediately by a series of serial dilutions in 15 mL conical tubes to obtain desired concentrations. NP solutions were vortexed in between each dilution step to maintain dispersion in the growth medium. Exposures were conducted in Cellvis 96-well black polystyrene bottom cell culture treated microplates (P96-1.5H-N, Cellvis). To decrease the formation of a protein corona, NP exposures were conducted in a phenol red- and FBS-free L-15 growth medium.Cell viability was measured using an alamarBlue™ (AB) cell viability reagent (DAL1025, Invitrogen) which measures the reduction of resazurin to highly fluorescent red resorufin metabolized by viable cells. Cells were seeded at 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well in 96-well (P96-1.5H-N, Cellvis) plates for 24–28 hours before treatment. Following 85% confluence, the cells were rinsed with PBS and replaced with 200 μL of PS NP solution in a phenol red- and FBS-free L-15 growth medium for each nanoparticle type at each concentration (0, 2, 4, 6, 8, 10 μg ml−1) for 24 hours total. The cells exposed to the phenol red- and FBS-free L-15 growth medium only were used as the negative control. At the 20 hour mark, 20 μL of AB was added to each well, including blanks containing the NPs only, for 4 h at 19 °C. The relative fluorescence was read at 560/590 nm using a microplate with a monochrometer (Synergy H4 Hybrid Reader, Bio Tek). Viability was normalized to the non-treated control cells. Prior to the routine use of the test method above, 3,4-dichloroaniline (3,4-DCA) was used to demonstrate the sensitivity of the RTgill-W1 cell line to AB according to the OECD Test No. 249: Fish Cell Line Acute Toxicity (ESI† Fig. S1).36
000 cells per well in 96-well (P96-1.5H-N, Cellvis) plates for 24–28 hours before treatment. Following 85% confluence, the cells were rinsed with PBS and replaced with 200 μL of PS NP solution in a phenol red- and FBS-free L-15 growth medium for each nanoparticle type at each concentration (0, 2, 4, 6, 8, 10 μg ml−1) for 24 hours total. The cells exposed to the phenol red- and FBS-free L-15 growth medium only were used as the negative control. At the 20 hour mark, 20 μL of AB was added to each well, including blanks containing the NPs only, for 4 h at 19 °C. The relative fluorescence was read at 560/590 nm using a microplate with a monochrometer (Synergy H4 Hybrid Reader, Bio Tek). Viability was normalized to the non-treated control cells. Prior to the routine use of the test method above, 3,4-dichloroaniline (3,4-DCA) was used to demonstrate the sensitivity of the RTgill-W1 cell line to AB according to the OECD Test No. 249: Fish Cell Line Acute Toxicity (ESI† Fig. S1).36
2.4 Cytological imaging preparation
Exposures were conducted in PhenoPlate 96-well plates composed of black optically clear cyclic olefin plastic designed for adherent cell cultures and high-content screening systems (PhenoPlate 6055300, Revvity). Cells were seeded using the protocol referenced in the Cell viability section. A 12 μg mL−1 solution was prepared with 10 mg mL−1 PS-NH2, 1 mg mL−1 PS-COOH and PS-Plain manufacturer vials immediately after sonication in a phenol red- and FBS-free L-15 growth medium for each nanoparticle type at each concentration (0, 2, 4, 6 μg ml−1) followed immediately by a series of serial dilutions in 15 mL conical tubes. NP solutions were vortexed in between each dilution step to maintain dispersion in the growth medium.A cell painting assay37 was modified for a 96-well plate and a reduced number of channels for automated fluorescence microscopy on an ImageXpress Micro XLS Widefield High Content Screening System (Molecular Devices, California, USA) using fluorescence cytochemistry parameters in Table 1. First, live-cell staining was conducted with MitoTracker Deep Red in a complete culture medium, followed by fixation with 4% paraformaldehyde. The cells were permeabilized with 0.1% Triton X-100 and were washed prior to staining with 1% bovine serum albumin in PBS (1×) solution containing Hoechst-33342 and rhodamine phalloidin. The wells were washed and stored in PBS. The plates were covered in aluminum foil and placed at 4 °C until image acquisition. The detailed protocol is shown in ESI† Table S1.
| DAPI | GFP | Texas Red | Cy5 |
|---|---|---|---|
| 377/447 | 472/520 | 562/624 | 628/692 |
| Hoechst 33342 | PS NPs | Rhodamine phalloidin | MitoTracker Deep Red |
2.5 Image screening
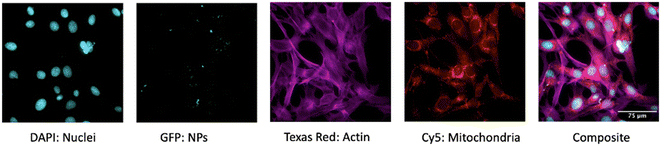
Images were acquired using an ImageXpress Micro XLS Widefield High Content Screening System at 60× magnification using four different channels (Fig. 1). During the acquisition setup, channel-specific Z-offsets, exposure times, and laser autofocus settings were optimized for the plate-type, objective, and cell line. Images included 9 sites per well (n = 5) to obtain 36 2D projection images for each well. Similarly, the acquisition set up was followed for the Z-series. Under the Z-series tab, based on the # of steps or z-planes chosen, the step size (2 μm) was automatically calculated by the MetaXpress Software and used as recommended.2.6 Bio-image analysis and phenotypic profiling
Cell Profiler 4.2.6 (ref. 38) was used to characterize qualitative morphological responses to quantifiable information. A pipeline was developed to automatically conduct measurements and was optimized to obtain multiple features at the resolution of a single cell (count, size, intensity, granularity, etc.) to quantify cellular phenotypic changes with the use of per-well profiles to analyze patterns across treatments. A data analysis pipeline was created to obtain per-well profiles to extract morphological image features from established methods.37 Parameters were also adjusted to overcome nanoparticle interference by using randomly selected wells and for deconvolution-based image sharpening. In addition to the steps described, PS NPs were created as a primary object, in which both were masked and related to only the regions of the cell (secondary object) on the pipeline. Cell data were exported to a local SQLite database file and were extracted using the RSQLite package in R.39 Phenotypic response data were analyzed using the methods described by Ostovich and Klaper, 2024.402.7 Caspase-3/7 assay
Following the NP preparation previously mentioned, a 12 μg mL−1 solution of PS-NH2 was prepared in a phenol red- and FBS-free L-15 growth medium by using a particle solution from the 10 mg mL−1 manufacturer vial. This was followed immediately by a series of serial dilutions in 15 mL conical tubes to obtain desired concentrations (0, 2, 3, 4, 6 μg ml−1). NP solutions were vortexed in between each dilution step to maintain dispersion in the growth medium. Exposures were conducted in Cellvis 96-well black polystyrene bottom cell culture treated microplates (P96-1.5H-N, Cellvis).Caspase proteins were measured using a Red CellEvent™ caspase-3/7 detection reagent (C10431, Invitrogen), a fluorogenic assay that cleaves activated caspases enabling the dye to bind to DNA producing a red fluorescent dye. Cells were seeded at 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well in 96-well (P96-1.5H-N, Cellvis) plates for 24–28 hours before treatment. Following 85% confluence, the cells were rinsed with PBS and replaced with 100 μL of NH2-PS NP solution in a phenol red- and FBS-free L-15 growth medium for each nanoparticle type at each concentration for 24 hours total. The cells exposed to the phenol red- and FBS-free L-15 growth medium only were used as the negative control. At the 24 hour mark, 10 μL of dye was added to each well, including blanks containing the NPs only, for 60 min at 19 °C. Fluorescence was captured in the Texas Red channel via automated fluorescence microscopy on the ImageXpress Micro XLS Widefield High Content Screening System (Molecular Devices, California, USA) immediately following the addition of the dye. Fluorescence was quantified via a developed pipeline with CellProfiler. Prior to the routine use of the test method above, 2 μM of camptothecin was used to demonstrate the sensitivity of the RTgill-W1 cell line to the caspase-3/7 dye. 0.1% DMSO and the phenol red- and FBS-free L-15 growth medium were used as negative controls (ESI† Fig. S2).
000 cells per well in 96-well (P96-1.5H-N, Cellvis) plates for 24–28 hours before treatment. Following 85% confluence, the cells were rinsed with PBS and replaced with 100 μL of NH2-PS NP solution in a phenol red- and FBS-free L-15 growth medium for each nanoparticle type at each concentration for 24 hours total. The cells exposed to the phenol red- and FBS-free L-15 growth medium only were used as the negative control. At the 24 hour mark, 10 μL of dye was added to each well, including blanks containing the NPs only, for 60 min at 19 °C. Fluorescence was captured in the Texas Red channel via automated fluorescence microscopy on the ImageXpress Micro XLS Widefield High Content Screening System (Molecular Devices, California, USA) immediately following the addition of the dye. Fluorescence was quantified via a developed pipeline with CellProfiler. Prior to the routine use of the test method above, 2 μM of camptothecin was used to demonstrate the sensitivity of the RTgill-W1 cell line to the caspase-3/7 dye. 0.1% DMSO and the phenol red- and FBS-free L-15 growth medium were used as negative controls (ESI† Fig. S2).
2.8 Quantifying colocalization
Analysis of co-localized PS NPs within the mitochondria, nuclei, and actin were determine through z-stack imaging calculations per well by using the Just Another Co-localization Plugin (JACoP) in Fiji.41 The most common methods used for obtaining colocalization measurements in the literature are predominantly qualitative to prove the existence of colocalization or subjectively through manual clusters in two-dimensional histograms.42,43 Although two probes may be subjectively identified to be colocalized when they are merged to a combined color, measurements tend to be highly prone to random error and bias since these techniques are typically based on visual estimation leading to inconsistent and irreproducible results.44 To overcome this issue, a threshold mask is created to include all areas of biological interest across the z-stacks by choosing a pixel cutoff. Only the pixels within that region will be measured excluding any artifacts generated by background and noise.45 An example of a thresholded image is available in ESI† Fig. S3 from Fiji.41 Measurements can be displayed as a biologically meaningful set of coefficients that are the proportion of each colocalized problem known as the Manders overlap coefficient (MOC) as opposed to the Pearson correlation coefficient invariant to intensity scales. The MOC allows one to compare the fraction of channel 1 with channel 2 and the fraction of channel 2 with channel 1.42,46 Thresholded Manders' overlap coefficients were calculated and are commonly used as co-localization indicators between two selected images. The thresholded Manders' coefficient (tM2) recorded represents the ratio of overlapping pixels in the GFP channel (channel 2) with the corresponding Cy5, Texas Red, or DAPI channels (channels 1, 3, and 4, respectively). Orthogonal projections were generated in Fiji.412.9 Statistical analyses
All statistical analyses were performed using R software.47,48 After confirming the normality using the Shapiro–Wilk test for each particle type, a one-way ANOVA with Tukey's post hoc test was performed for multiple comparisons. A non-parametric test was conducted on the data not normally distributed using Kruskal–Wallis' test followed by Dunn's test with Bonferroni adjustments. For cell viability, the EC50 was statistically calculated with a three-parameter logistic model using the drc package in R.49 For colocalization, a one-way ANOVA followed by a post hoc Tukey test was used to obtain differences per particle type (p-value > 0.05). A confidence interval of 95% was used to determine significant differences.3. Results and discussion
3.1 Polystyrene nanoparticle characterization
When NPs are introduced into media, they have the possibility of interacting with the various media components which can influence their physicochemical properties prior to making any biological contact.50,51 Because NP colloidal stability can be altered by high salt content,52 DLS was used to characterize the zeta potential and size of the PS NPs in phenol red- and FBS-free L-15 growth media, PBS, and UltraPure™ distilled water.As seen in ESI† Fig. S4a, in L-15 media, PS-COOH exhibited negatively charged values between −14 mV and −18 mV, whereas PS-NH2 displayed a positively charged value of +7 mV. The zeta potentials were significantly different across all three particle types. This was expected as the PS NPs are functionalized with negatively charged carboxyl groups and positively charged amino groups, respectively. It is also important to note that PS-Plain, although conventionally considered to be uncharged, exhibited a slightly negative charge (−2 mV to −4 mV) derived from fragments of the initiator used in the polymerization reaction as reported by the manufacturer (CD Bioparticles). However, these zeta potential values suggest that the PS NPs are not highly stable in L-15 media.
Significant differences were observed for each particle type between their sizes in water versus L-15 as shown in ESI† Fig. S4b. The cell culture medium strongly impacts particle size. All particles showed an increase in size while still maintaining their size at nanoscale. These results were in congruent with those of previous characterization of PS NPs in varying complex culture media regarding size.53–55 However, after 24 hours of exposure, PS NP agglomerates can be observed under the ImageXpress Micro XLS Widefield High Content Screening System (Molecular Devices, California, USA) presented in ESI† Fig. S5, suggesting that the observed toxicity, or lack thereof, was caused by the agglomerates as opposed to single NPs. Additionally, the size of the particles post 24 hour treatment was quantified by bio-imaging analysis software presented in ESI† Fig. S6. This emphasizes that the nanoparticles are not stable in solution, and the increase in agglomeration and sedimentation could also be due to the salt in the L-15 phenol red- and FBS-free medium.
3.2 PS-NH2 NPs induce phenotypic response at sublethal concentrations
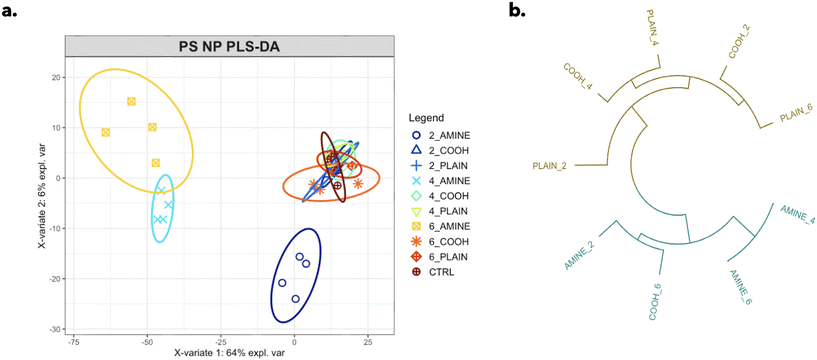
A total of 1326 different features were profiled in an unbiased robust manner in RTgill-W1 cells for each particle type. A PLS-DA (partial least squares-discriminant analysis) model was used to identify significant changes in the complex phenotypic profile across each particle type separating their responses by concentration with respect to the control (Fig. 4a). Variable importance in projection (VIP) scores measure the level of importance each variable contributes to the separation of response groups; here, scores >1 were considered the most important. Out of the top 10 VIP scores, 8 of the phenotypic markers were mitochondrial descriptors as shown in ESI† Table S2. This work emphasizes the relevance and applicability of image-based profiling techniques in assessing the environmental impacts of plastic NPs in aquatic organisms.
3.3 Differential sublethal and lethal effects of PS-NH2 NPs on cellular morphology
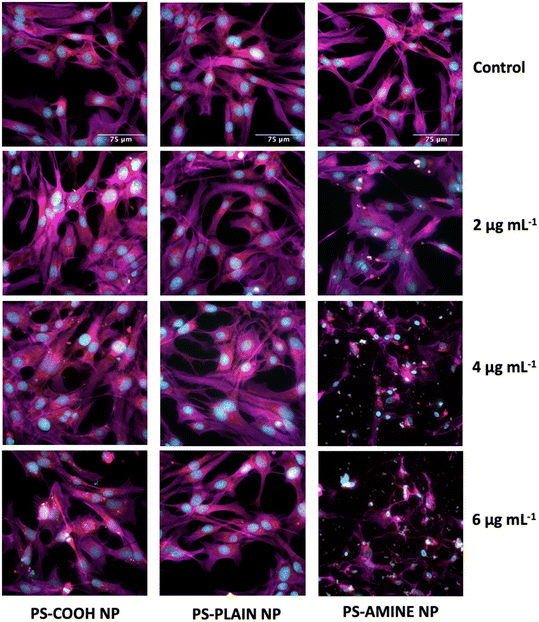
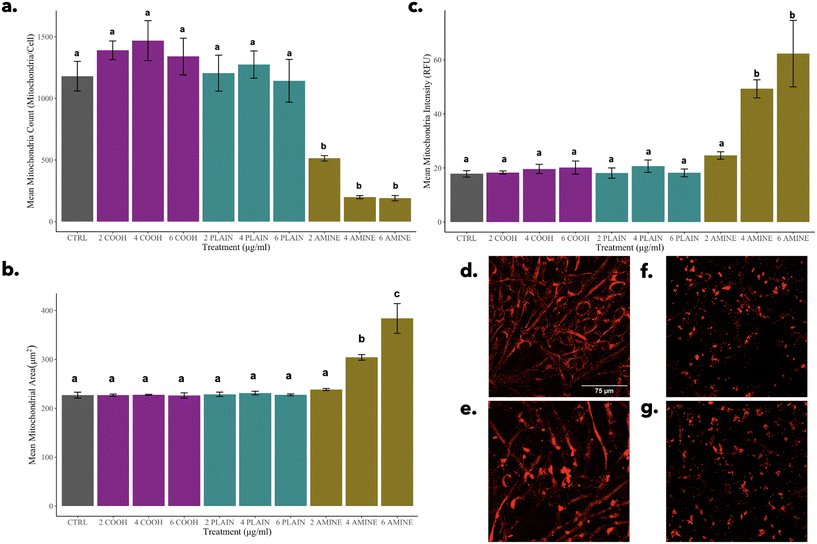
The effects of PS-NH2 NPs on cellular morphology were examined at both sublethal and lethal concentrations with a focus on mitochondrial, cytoskeletal, and nuclear responses. Data suggest that NP treatments at sublethal concentrations may have triggered an early cellular stress response without leading to the onset of apoptosis. However, observations described for NP treatments at lethal concentrations align with morphological hallmarks of apoptosis. The data highlight the complex cellular responses that are dose dependent.On the other hand, a significant decrease in mitochondria can be observed alongside the significant increases in both size and intensity for 4 and 6 μg mL−1 of PS-NH2 NPs (Fig. 5f and g). These three morphological features can be indicative of mitochondrial fusion or mitophagy.60 NH2-PS NPs have been shown to promote possible fusion due to a significant increase in gene expression for the mfn2 (mitofusion-2 protein) gene in HUVEC cells.57 Similarly, this phenomenon can also be explained by mitophagy which is the degradation of dysfunctional mitochondria through the fusion of the lysosomes.61 Overall, it is possible that the mitochondria may play a major role in the mechanism governing this morphological change at the sublethal concentration of 2 μg mL−1 of PS-NH2 NPs observed in the PLS-DA. This should be studied further to assess the possible mitochondrial pathways induced apart from the mechanism of cell death reported.57,62,63
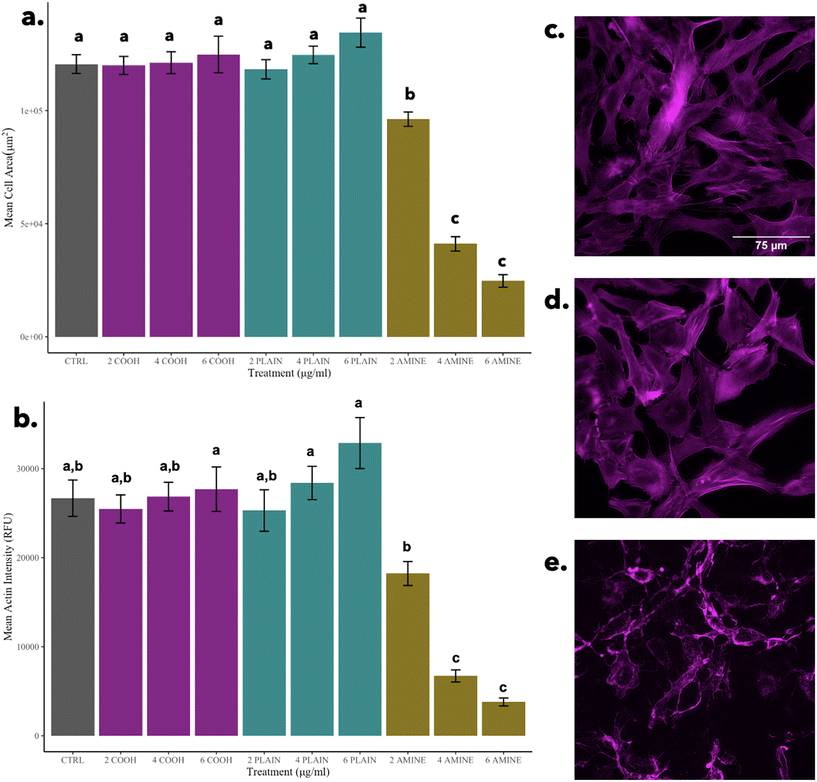
In general, cells decrease their size to conserve energy; however, the hypothetical possibility of nanoparticle–actin interactions should not be ignored. For example, the induction of untargeted cell-signaling pathways can be induced when a cell meets a nanoparticle which can result in non-specific degradation. In drug delivery models, nanomaterials have been designed to interact with enzymes to improve target efficiency and reduce unwanted cell signaling by decreasing protease activity.64 In general, nanoparticle–actin interactions need to be further studied at sublethal doses while monitoring protease activity in vertebrate cells in response to PS NPs. On the other hand, in Fig. 6a and b, when compared to the cells treated with 2 μg mL−1 of PS-NH2 NPs, there are significant decreases in cell size and actin intensity as seen in Fig. 6e, indicative of a morphological hallmark of apoptosis possibly governed by the mechanism of cell death elucidated by Wang et al., 2013.62 Alongside mitophagy, mitochondrial fusion, and nuclear condensation, the shrinkage of the cell is another morphological indicator of apoptosis.60
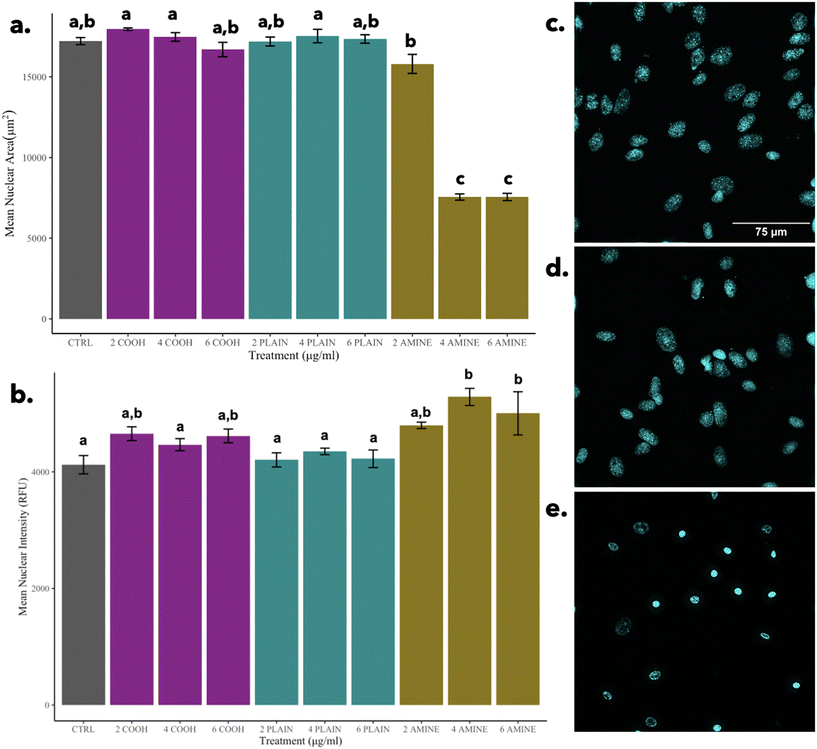
In contrast, it can be observed that in comparison to cells treated with 2 μg mL−1 of PS-NH2 NPs, there is a significant decrease in nuclear area but no significant differences in nuclear intensity as seen in Fig. 7e. The significant decrease in nuclear area is indicative of the morphological hallmark of apoptosis and possibly governed by the mechanism of cell death for concentrations of 4 and 6 μg mL−1 of PS-NH2 NPs, elucidated by Wang et al., 2013.62 This finding reinforces that the indication of apoptosis is nuclear condensation as observed in other particles.65 Although cells can undergo secondary necrosis after apoptosis, it is important to note that the morphological indicator of non-apoptotic cell death via necrosis is the swelling and rupture of the nuclei which are not observed.
3.4 Nanoparticles colocalize with cellular compartments
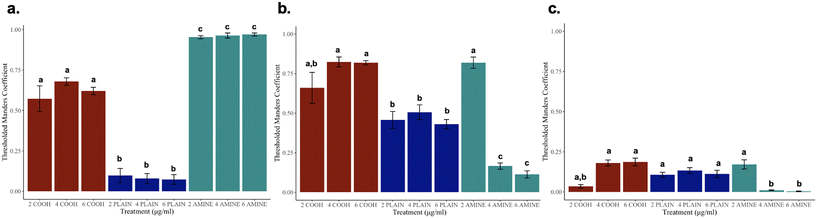
To resolve whether NPs are internalized, Z-stacks of RTgill-W1 cells were obtained to examine colocalization with various cellular components. Orthogonal projections can be found in ESI† Fig. S8. In Fig. 8a–c, thresholded Manders coefficients (tM2) and their corresponding concentrations are displayed per particle. Fig. 8a presents the GFP channel (tM2) colocalized with the Cy5 channel (tM1) across all three particle types visually depicted in ESI† Fig. S8a–c. These observations suggest that unlike the plain particle, nano-mito interactions in functionalized NPs may play a larger role in treated cells even though the colocalization measurements were significantly different across all the particle types. These observations suggest that the NPs have been trafficked into the cell.Interestingly, Fig. 8b presents the GFP channel (tM2) colocalized with the TexasRed channel (tM1) across all three particle types, visually depicted in ESI† Fig. S8d–f. At the sublethal concentration of 2 μg mL−1 of PS-NH2 NPs, there is more colocalization present in amine-functionalized NPs before the cytoskeleton becomes disorganized at the lethal concentrations. It should also be noted that the colocalization measurements for cells treated with PS-COOH NPs were not different to that of the 2 μg mL−1 of PS-NH2 NPs which are in congruent with the results based on Euclidean distance. This was not a surprise as it has been elucidated in the literature that –COOH enhances uptake due to the negative charge alongside the presence of extracellular amino acids, such as cysteine which can mediate NP internalization via the mTOR signaling pathway.66,67 These observations suggest that there are possible nano-actin interactions across the negative and positive PS NPs that need to be studied further. Colocalization in Fig. 8c presents the GFP channel (tM2) colocalized with the DAPI channel (tM1) across all three particle types, visually depicted in ESI† Fig. S8g–i. These observations suggest that the NPs may not directly interact with the nucleus despite the decrease in PS-NH2 NP uptake and nuclear condensation previously observed. Interestingly, there was little to no colocalization of the nuclei and cytoskeleton at the lethal concentrations further supporting the possibility of nano-mito interactions and possible mitochondrial-induced nanoparticle cell signaling pathways, resulting in the previously discussed morphological responses. It is important to note that although we did find colocalization, it does not identify molecular interactions but whether where they are localized within a certain volume.44,68
4. Conclusion
The results of this present study show the intricate cellular responses of PS NPs to RTgill-W1 gill cells at sublethal and lethal concentrations through the comprehensive analysis of phenotypic profiling and high-content imaging analysis of mitochondria, nuclear and cytoskeletal features. Image-based profiling highlighted the phenotypic differences induced by positively charged PS-NH2 NPs compared to negatively charged and uncharged counterparts. Additionally, we observed alterations to the mitochondria at the sublethal concentration of 2 μg mL−1 of PS-NH2 NPs suggesting the role of mitophagy and the involvement of apoptosis and fusion at higher concentrations. This suggests that mitochondria may play a vital role in the sublethal cellular responses to positively charged nanoparticles. While we did not observe nuclear alterations at the sublethal concentration of 2 μg mL−1 of PS-NH2 NPs, the alterations at higher concentrations suggest apoptosis. We also observed the cytoskeletal shrinkage associated with cells treated with 2 μg mL−1 of PS-NH2 NPs, indicating a pre-apoptotic stress response.Thus, it is imperative to understand the mechanisms governing the observed phenotypic changes, especially those associated with mitochondrial dynamics, nuclear condensation, and cytoskeletal disorganization. The findings of the paper not only provide mechanistic insights into the possible mode of action governing the phenotypic response at sublethal concentrations of positively charged nanoparticles, but also highlight a potential unified mechanism that may be nanoparticle specific to the mitochondria; however, pristine polystyrene nanoparticles do not reflect most nanoplastics found in the environment due to existing varying shapes, sizes, compositions, and surface chemistries. Thus, it is important to study these impacts with other primary- or secondary-plastics. Further research into the underlying mechanisms will contribute to our understanding of nanoparticle toxicity in aquatic vertebrates guiding both the policy and the sustainable design of nanoparticles.
Data availability
Data for this article including CellProfiler pipelines and cellular images are available at Mendeley Data DOI: https://doi.org/10.17632/8kb3twbbf6.1.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work was supported by the National Science Foundation under Grant No. CHE-2001611, the NSF Center for Sustainable Nanotechnology (CSN), and the Mick A. Naulin Foundation Research Award. The CSN is a part of the Centers for Chemical Innovation Program.References
- L. M. Rios, C. Moore and P. R. Jones, Persistent organic pollutants carried by synthetic polymers in the ocean environment, Mar. Pollut. Bull., 2007, 54(8), 1230–1237 CrossRef CAS PubMed.
- M. Wagner, C. Scherer, D. Alvarez-Muñoz, N. Brennholt, X. Bourrain and S. Buchinger, et al., Microplastics in freshwater ecosystems: what we know and what we need to know, Environ. Sci. Eur., 2014, 26(1), 1–9, DOI:10.1186/s12302-014-0012-7.
- S. Wagner and T. Reemtsma, Things we know and don't know about nanoplastic in the environment, Nat. Nanotechnol., 2019, 14(4), 300–301 CrossRef CAS PubMed.
- E. Kelpsiene, M. T. Ekvall, M. Lundqvist, O. Torstensson, J. Hua and T. Cedervall, Review of ecotoxicological studies of widely used polystyrene nanoparticles, Environ. Sci.: Processes Impacts, 2022, 24(1), 8–16 RSC , available from: https://pubs.rsc.org/en/content/articlehtml/2022/em/d1em00375e.
- Z. Chen, X. Shi, J. Zhang, L. Wu, W. Wei and B. J. Ni, Nanoplastics are significantly different from microplastics in urban waters, Water Res.: X, 2023, 19, 100169 CAS.
- N. M. Schaeublin, L. K. Braydich-Stolle, A. M. Schrand, J. M. Miller, J. Hutchison and J. J. Schlager, et al., Surface charge of gold nanoparticles mediates mechanism of toxicity, Nanoscale, 2011, 3(2), 410–420 RSC , available from: https://pubs.rsc.org/en/content/articlehtml/2011/nr/c0nr00478b.
- C. He, Y. Hu, L. Yin, C. Tang and C. Yin, Effects of particle size and surface charge on cellular uptake and biodistribution of polymeric nanoparticles, Biomaterials, 2010, 31(13), 3657–3666 CrossRef CAS PubMed.
- W. Wei, Y. Li, M. Lee, N. Andrikopoulos, S. Lin and C. Chen, et al., Anionic nanoplastic exposure induces endothelial leakiness, Nat. Commun., 2022, 13(1), 1–14 Search PubMed , available from: https://www.nature.com/articles/s41467-022-32532-5.
- X. Jin, H. Yu, Z. Zhang, T. Cui, Q. Wu and X. Liu, et al., Surface charge-dependent mitochondrial response to similar intracellular nanoparticle contents at sublethal dosages, Part. Fibre Toxicol., 2021, 18(1), 1–15 CrossRef PubMed , available from: https://particleandfibretoxicology.biomedcentral.com/articles/10.1186/s12989-021-00429-8.
- M. Teng, X. Zhao, F. Wu, C. Wang, C. Wang and J. C. White, et al., Charge-specific adverse effects of polystyrene nanoplastics on zebrafish (Danio rerio) development and behavior, Environ. Int., 2022, 163, 107154 CrossRef CAS PubMed.
- X. D. Sun, X. Z. Yuan, Y. Jia, L. J. Feng, F. P. Zhu and S. S. Dong, et al., Differentially charged nanoplastics demonstrate distinct accumulation in Arabidopsis thaliana, Nat. Nanotechnol., 2020, 15(9), 755–760 CrossRef CAS PubMed.
- E. McKeel, H. I. Kim, S. J. Jeon, J. P. Giraldo and R. Klaper, The effect of nanoparticle surface charge on freshwater algae growth, reproduction, and lipid production, Environ. Sci.: Nano, 2023, 11(2), 657–666 RSC.
- Z. V. Feng, I. L. Gunsolus, T. A. Qiu, K. R. Hurley, L. H. Nyberg and H. Frew, et al., Impacts of gold nanoparticle charge and ligand type on surface binding and toxicity to Gram-negative and Gram-positive bacteria, Chem. Sci., 2015, 6(9), 5186–5196 RSC , available from: https://pubs.rsc.org/en/content/articlehtml/2015/sc/c5sc00792e.
- E. Ostovich, A. Henke, C. Green, R. Hamers and R. Klaper, Predicting the phytotoxic mechanism of action of LiCoO2 nanomaterials using a novel multiplexed algal cytological imaging (MACI) assay and machine learning, Environ. Sci.:Nano, 2024, 11(2), 507–517 RSC.
- X. Ma, R. Hartmann, D. Jimenez De Aberasturi, F. Yang, S. J. H. Soenen and B. B. Manshian, et al., Colloidal Gold Nanoparticles Induce Changes in Cellular and Subcellular Morphology, ACS Nano, 2017, 11(8), 7807–7820, DOI:10.1021/acsnano.7b01760.
- E. Eisenbarth, P. Linez, V. Biehl, D. Velten, J. Breme and H. F. Hildebrand, Cell orientation and cytoskeleton organisation on ground titanium surfaces, Biomol. Eng., 2002, 19(2–6), 233–237 CrossRef CAS PubMed.
- S. J. H. Soenen, N. Nuytten, S. F. De Meyer, S. C. De Smedt and M. De Cuyper, High intracellular iron oxide nanoparticle concentrations affect cellular cytoskeleton and focal adhesion kinase-mediated signaling, Small, 2010, 6(7), 832–842 CrossRef CAS PubMed.
- D. Wu, Y. Ma, Y. Cao and T. Zhang, Mitochondrial toxicity of nanomaterials, Sci. Total Environ., 2020, 702, 134994 CrossRef CAS PubMed.
- C. L. Browning, A. Green, E. P. Gray, R. Hurt and A. B. Kane, Manganese Dioxide Nanosheets Induce Mitochondrial Toxicity in Fish Gill Epithelial Cells, Nanotoxicology, 2021, 15(3), 400 CrossRef CAS PubMed.
- S. Lin, H. Zhang, C. Wang, X. L. Su, Y. Song and P. Wu, et al., Metabolomics Reveal Nanoplastic-Induced Mitochondrial Damage in Human Liver and Lung Cells, Environ. Sci. Technol., 2022, 56(17), 12483–12493, DOI:10.1021/acs.est.2c03980.
- Y. Pan, A. Leifert, D. Ruau, S. Neuss, J. Bornemann and G. Schmid, et al., Gold nanoparticles of diameter 1.4 nm trigger necrosis by oxidative stress and mitochondrial damage, Small, 2009, 5(18), 2067–2076 CrossRef CAS PubMed.
- S. E. Lee, Y. Yi, S. Moon, H. Yoon and Y. S. Park, Impact of Micro- and Nanoplastics on Mitochondria, Metabolites, 2022, 12(10), 897 CrossRef CAS PubMed.
- R. J. Mailloux, J. Treberg, C. Grayson, L. B. Agellon and H. Sies, Mitochondrial function and phenotype are defined by bioenergetics, Nat. Metab., 2023, 5(10), 1641 CrossRef PubMed , available from: https://www.nature.com/articles/s42255-023-00885-w.
- Y. L. Wu, N. Putcha, K. W. Ng, D. T. Leong, C. T. Lim and S. C. J. Loo, et al., Biophysical responses upon the interaction of nanomaterials with cellular interfaces, Acc. Chem. Res., 2013, 46(3), 782–791, DOI:10.1021/ar300046u.
- J. S. Zheng, S. Y. Zheng, Y. B. Zhang, B. Yu, W. Zheng and F. Yang, et al., Sialic acid surface decoration enhances cellular uptake and apoptosis-inducing activity of selenium nanoparticles, Colloids Surf., B, 2011, 83(1), 183–187 CrossRef CAS PubMed.
- L. T. Allen, E. J. P. Fox, I. Blute, Z. D. Kelly, Y. Rochev and A. K. Keenan, et al., Interaction of soft condensed materials with living cells: Phenotype/transcriptome correlations for the hydrophobic effect, Proc. Natl. Acad. Sci. U. S. A., 2003, 100(11), 6331–6336, DOI:10.1073/pnas.1031426100.
- D. A. Fletcher and R. D. Mullins, Cell mechanics and the cytoskeleton, Nature, 2010, 463(7280), 485 CrossRef CAS PubMed.
- X. Qin, H. Zhang, X. Xing, P. Wang, J. Yan and D. Liu, et al., Robust strategies in nuclear-targeted cancer therapy based on functional nanomaterials, Mater. Des., 2022, 221, 110999 CrossRef CAS.
- I. Tomé, V. Francisco, H. Fernandes and L. Ferreira, High-throughput screening of nanoparticles in drug delivery, APL Bioeng., 2021, 5(3), 31511 CrossRef PubMed.
- S. Lin, K. Schorpp, I. Rothenaigner and K. Hadian, Image-based high-content screening in drug discovery, Drug Discovery Today, 2020, 25(8), 1348–1361 CrossRef CAS PubMed.
- C. Willis, J. Nyffeler and J. Harrill, Phenotypic Profiling of Reference Chemicals across Biologically Diverse Cell Types Using the Cell Painting Assay, SLAS Discovery, 2020, 25(7), 755–769, DOI:10.1177/2472555220928004.
- J. C. Caicedo, S. Singh and A. E. Carpenter, Applications in image-based profiling of perturbations, Curr. Opin. Biotechnol., 2016, 39, 134–142 CrossRef CAS PubMed.
- A. Kizhedath, S. Wilkinson and J. Glassey, Applicability of Traditional In Vitro Toxicity Tests for Assessing Adverse Effects of Monoclonal Antibodies: A Case Study of Rituximab and Trastuzumab, Antibodies, 2018, 7(3), 30 CrossRef CAS PubMed.
- H. Kimura, T. Sawada, S. Oshima, K. Kozawa, T. Ishioka and M. Kato, Toxicity and roles of reactive oxygen species, Curr. Drug Targets: Inflammation Allergy, 2005, 4(4), 489–495 CAS , available from: https://pubmed.ncbi.nlm.nih.gov/16101527/.
- R. S. Thomas, T. Bahadori, T. J. Buckley, J. Cowden, C. Deisenroth and K. L. Dionisio, et al., The Next Generation Blueprint of Computational Toxicology at the U.S. Environmental Protection Agency, Toxicol. Sci., 2019, 169(2), 317–332 CrossRef CAS PubMed , available from: https://pubmed.ncbi.nlm.nih.gov/30835285/.
- Test No. 249: Fish Cell Line Acute Toxicity - The RTgill-W1 cell line assay, 2021 Jun 18 [cited 2023 Nov 21], available from: https://read.oecd-ilibrary.org/environment/test-no-249-fish-cell-line-acute-toxicity-the-rtgill-w1-cell-line-assay_c66d5190-en.
- M. A. Bray, S. Singh, H. Han, C. T. Davis, B. Borgeson and C. Hartland, et al., Cell Painting, a high-content image-based assay for morphological profiling using multiplexed fluorescent dyes, Nat. Protoc., 2016, 11(9), 1757–1774 CrossRef CAS PubMed , available from: https://www.nature.com/articles/nprot.2016.105.
- D. R. Stirling, et al., CellProfiler4: Improvements in speed, utility, and usability, BMC Bioinf., 2021, 22, 1–11 CrossRef PubMed.
- K. Muller, H. Wickham, D. A. James and S. Falcon, RSQLite: SQLite Interface for R, 2023 Search PubMed.
- E. Ostovich and R. Klaper, Using a Novel Multiplexed Algal Cytological Imaging (MACI) Assay and Machine Learning as a Way to Characterize Complex Phenotypes in Plant-Type Organisms, Environ. Sci. Technol., 2024, 58(11), 4894–4903 CrossRef CAS PubMed.
- J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair and T. Pietzsch, et al., Fiji: An open-source platform for biological-image analysis, Nat. Methods, 2012, 9(7), 676–682 CrossRef CAS PubMed.
- M. EMM, F. J. Verbeek and J. A. Aten, Measurement of co-localization of objects in dual-colour confocal images, J. Microsc., 1993, 169(3), 375–382, DOI:10.1111/j.1365-2818.1993.tb03313.x.
- D. Demandolx and J. Davoust, Multicolour analysis and local image correlation in confocal microscopy, J. Microsc., 1997, 185(1), 21–36, DOI:10.1046/j.1365-2818.1997.1470704.x.
- S. V. Costes, D. Daelemans, E. H. Cho, Z. Dobbin, G. Pavlakis and S. Lockett, Automatic and Quantitative Measurement of Protein-Protein Colocalization in Live Cells, Biophys. J., 2004, 86(6), 3993 CrossRef CAS PubMed.
- A. P. French, S. Mills, R. Swarup, M. J. Bennett and T. P. Pridmore, Colocalization of fluorescent markers in confocal microscope images of plant cells, Nat. Protoc., 2008, 3(4), 619–628 CrossRef CAS PubMed , available from: https://www.nature.com/articles/nprot.2008.31.
- E. M. Manders, J. Stap, G. J. Brakenhoff, R. Van Driel and J. A. Aten, Dynamics of three-dimensional replication patterns during the s-phase, analysed by double labelling of dna and confocal microscopy, J. Cell Sci., 1992, 103(3), 857–862, DOI:10.1242/jcs.103.3.857.
- RStudio Team, RStudio: Integrated Development Environment for R, 2020 Search PubMed.
- R Core Team, R: A language and Environment for Statistical Computing, 2019 Search PubMed.
- J. C. Strebig, Package ‘drc’ Title Analysis of Dose-Response Curves, 2022 Search PubMed.
- X. Jiang, S. Weise, M. Hafner, C. Röcker, F. Zhang, W. J. Parak and G. U. Nienhaus, Quantitative analysis of the protein corona on FePt nanoparticles formed by transferrin binding, J. R. Soc. Interface, 2010, 7(Suppl 1), S5–S13 CAS.
- E. C. Cho, Q. Zhang and Y. Xia, The effect of sedimentation and diffusion on cellular uptake of gold nanoparticles, Nat. Nanotechnol., 2011, 6(6), 385–391 CrossRef CAS PubMed , available from: https://pubmed.ncbi.nlm.nih.gov/21516092/.
- T. L. Moore, L. Rodriguez-Lorenzo, V. Hirsch, S. Balog, D. Urban and C. Jud, et al., Nanoparticle colloidal stability in cell culture media and impact on cellular interactions, Chem. Soc. Rev., 2015, 44(17), 6287–6305 RSC , available from: https://pubs.rsc.org/en/content/articlehtml/2015/cs/c4cs00487f.
- T. Xia, M. Kovochich, M. Liong, J. I. Zink and A. E. Nel, Cationic polystyrene nanosphere toxicity depends on cell-specific endocytic and mitochondrial injury pathways, ACS Nano, 2008, 2(1), 85–96 CrossRef CAS PubMed.
- C. González-fernández, F. G. Díaz Baños, M. Á. Esteban and A. Cuesta, Functionalized nanoplastics (Nps) increase the toxicity of metals in fish cell lines, Int. J. Mol. Sci., 2021, 22(13), 7141 CrossRef PubMed.
- A. Katsumiti, M. P. Losada-Carrillo, M. Barros and M. P. Cajaraville, Polystyrene nanoplastics and microplastics can act as Trojan horse carriers of benzo(a)pyrene to mussel hemocytes in vitro, Sci. Rep., 2021, 11(1), 1–17 CrossRef PubMed , available from: https://www.nature.com/articles/s41598-021-01938-4.
- X. Shi, X. Wang, R. Huang, C. Tang, C. Hu and P. Ning, et al., Cytotoxicity and Genotoxicity of Polystyrene Micro- and Nanoplastics with Different Size and Surface Modification in A549 Cells, Int. J. Nanomed., 2022, 17, 4509 CrossRef CAS PubMed.
- Y. Fu, M. Fan, L. Xu, H. Wang, Q. Hu and Y. Jin, Amino-Functionalized Polystyrene Nano-Plastics Induce Mitochondria Damage in Human Umbilical Vein Endothelial Cells, Toxics, 2022, 10(5), 215 CrossRef CAS PubMed , available from: https://www.mdpi.com/2305-6304/10/5/215/htm.
- A. Nemmar, M. F. Hoylaerts, P. H. M. Hoet, J. Vermylen and B. Nemery, Size effect of intratracheally instilled particles on pulmonary inflammation and vascular thrombosis, Toxicol. Appl. Pharmacol., 2003, 186(1), 38–45 CrossRef CAS PubMed.
- H. Zhang, H. Cheng, Y. Wang, Z. Duan, W. Cui and Y. Shi, et al., Influence of Functional Group Modification on the Toxicity of Nanoplastics, Front. Mar. Sci., 2022, 8, 800782 CrossRef.
- A. Saraste and K. Pulkki, Morphologic and biochemical hallmarks of apoptosis, Cardiovasc. Res., 2000, 45(3), 528–537, DOI:10.1016/S0008-6363(99)00384-3.
- D. A. Chistiakov, I. A. Sobenin, V. V. Revin, A. N. Orekhov and Y. V. Bobryshev, Mitochondrial Aging and Age-Related Dysfunction of Mitochondria, BioMed Res. Int., 2014, 2014, 238463 Search PubMed.
- M. Wang, S. Bexiga, P. Anguissola, P. Boya and J. Simpson, Time resolved study of cell death mechanisms induced by amine-modified polystyrene nanoparticles, Nanoscale, 2013, 5, 10868–10876 RSC.
- J. Zhang, G. Hu, H. Guo, W. Yang, X. Li, Y. Ni, M. He, P. Ding and Y. Yu, Amino modifications exacerbate the developmental abnormalities of polystyrene microplastics via mitochondria-mediated apoptosis pathway in zebrafish larvae, Sci. Total Environ., 2025, 958, 178031 CrossRef CAS PubMed.
- C. F. Anderson and H. Cui, Protease-Sensitive Nanomaterials for Cancer Therapeutics and Imaging, Ind. Eng. Chem. Res., 2017, 56(20), 5761 CrossRef CAS PubMed.
- D. Chanda, W. Dudefoi, J. Anadu and M. Minghetti, Evaluation of the effect of silver and silver nanoparticles on the function of selenoproteins using an in-vitro model of the fish intestine: The cell line RTgutGC, Ecotoxicol. Environ. Saf., 2021, 211, 111930 CrossRef CAS PubMed.
- P. Foroozandeh and A. A. Aziz, Insight into Cellular Uptake and Intracellular Trafficking of Nanoparticles, Nanoscale Res. Lett., 2018, 13(1), 1–12, DOI:10.1186/s11671-018-2728-6.
- Y. Wei, T. Tang and H. B. Pang, Cellular internalization of bystander nanomaterial induced by TAT-nanoparticles and regulated by extracellular cysteine, Nat. Commun., 2019, 10(1), 1–11 CrossRef PubMed , available from: https://www.nature.com/articles/s41467-019-11631-w.
- K. W. Dunn, M. M. Kamocka and J. H. McDonald, A practical guide to evaluating colocalization in biological microscopy, Am. J. Physiol., 2011, 300(4), C723 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4en01149j |
| This journal is © The Royal Society of Chemistry 2025 |