Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Microbial cell lysis and nucleic acid extraction via nanofluidic channel
Krishna
Kant
a,
Jeongha
Yoo
b,
Steven
Amos
b,
Mason
Erkelens
b,
Craig
Priest
c,
Joe G.
Shapter
a and
Dusan
Losic
*b
aSchool of Chemical and Physical Sciences, Flinders University, Bedford Park, Adelaide 5042, Australia
bSchool of Chemical Engineering, The University of Adelaide, Adelaide, SA 5005, Australia. E-mail: Dusan.losic@adelaide.edu.au; Tel: +61 8 8013 4648
cIan Wark Research Institute, University of South Australia, Mawson Lakes, Adelaide 5095, Australia
First published on 4th March 2015
Abstract
This paper presents a microfluidic device with a nano-channel prepared by focused ion beam (FIB) milling for microbial cell lysis and nucleic acid extraction. The device prepared in a quartz slide combines two microchannels connected with one nanochannel in the middle with dimensions of 100 nm (width) and 250 nm (depth) and 1 μm (length). The capturing of a single or a few cells and their lysis for extraction of DNA in nanochannel were demonstrated. The purity and quantity of the extracted genetic material was tested using a UV spectrometer. The results revealed that the extraction methodology was successful and may be well-suited to integration in analytical lab-on-a-chip devices.
Introduction
Nanofluidic techniques enable extreme miniaturization of devices and enhanced analytical performance. These devices offer high sensitivity and reproducibility, low consumption of expensive, rare, or hazardous chemicals, and less energy on a single integrated device.1Coupled with advanced analytical techniques, they offer potential in applications from point of care to single cell analysis,1,2 and other examples where resources are limited.3,4 It was shown that incorporation of biological membranes and synthetic nanochannels in a device can create new opportunities for advanced analytical applications5,6 due to the increased surface area and reduced use of chemical reagents.7,8 Nanofluidic devices may also manipulate cell entities such as proteins, nucleic acids and cell organelles providing ultra-high sensitive analysis using ion source and electrochemical detection.9 Single-cell analysis can be achieved by integration of several biochemical steps into a micro total analysis system (μTAS) on a single chip.10,11 However, it is difficult to handle and manipulate single cells in microsystems, since very small volumes are involved in the analysis.12 The genome of any simple organism may contain thousands of base pairs and it is even more complex in higher species.13,14 Consequently, an integrated nanofluidic chip is a required approach to handling these tiny volumes of complex cell material without damage or loss. Ideally, all steps from cell lysis to analysis of the molecular cell contents, including DNA analysis, are needed on a single integrated chip.15–17 Furthermore, integration of several devices for parallel analysis for different molecules is an attractive prospect.
A number of sophisticated fabrication tools have been used to fabricate nanochannels in recent years.18 E-beam lithography followed by etching has been used to prepare nanochannels (and nanostructures within nanochannels19) with certain limitations of length and depth (depends on reactive ion etching (RIE) techniques). Nanoimprint lithography is capable of producing nanofluidic channels with dimensions of 10 nm in width.11,13,20,21
In this study, we first introduce the potential of cell lysis and extraction of nucleic acids in the nanofluidic device by trapping, lysing, and extracting microbial cells before eluting the genetic material. Moreover, we describe a suitable technique for the fabrication of nanochannels using focused ion beam (FIB) milling. The prepared devices offer an integrated micro- and nanofluidic system, with features that enable several functions performed during chemical analysis (sample preparation, fluid handling and injection, separation, and detection) and new potential for the development of advanced analytical chips.22,23
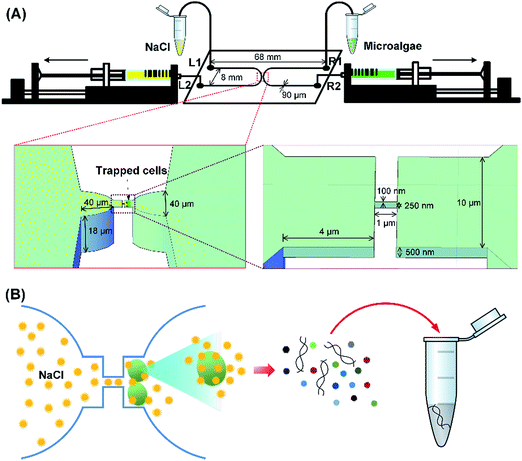
The schematic of the nanofluidic device is presented in Fig. 1. This scheme shows the flow of algae cells used as a model into the microchannels and diffusion of NaCl through the nanochannel for cell lysis. The collected raw material from cell lysis is analyzed using UV-vis spectrophotometer.
Experimental
Fabrication of microchannel and nanochannel
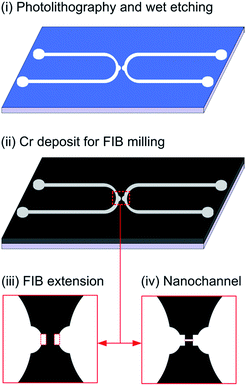
Microchannels were fabricated by using photolithography and wet chemical etching (Fig. 2(i)). Microfluidic chips were prepared in Quartz (Viosil, from Shin Etsu, Japan). It was first spin-coated with AZ 1518 photoresist (Delta 80, from SUSS MicroTec) and then baked for 1 min at 105 °C, the UV exposure was carried out at 340 nm by (EVG 620) Mask Aligner at 120 mJ cm−2. The exposed chip was developed using MIF 726 (Micro-chemicals). After patterning using photolithography, the wet chemical etching of the pattern was then achieved using hydrofluoric acid (HF) for 12 min to achieve 18 μm depth of the micro-channel. After that, the photoresist was removed by commercial AZ remover.The nanochannel was patterned by FIB milling using a Helios NanoLab 600 Dual Beam instrument (FEI Company) as schematically presented in Fig. 2(ii)–(iv). After photolithography the chip was sputtered with a 40 nm thick Cr layer to have good controls such as preventing from milling progressively deeper and protecting redeposition of material and structural deformations over the ion beam milling and imaging of channel.20 The ion beam energy for milling nanostructure is typically between 10 and 50 keV, with beam currents varying between 1 pA and 10 nA.24 100 nm width and 250 nm depth nanochannel was milled into a quartz substrate through a 40 nm thick Cr film using an ion beam probe with a beam current of 0.17 nA for 20 keV Ga+ ions at normal incidence on milling pattern of 100 nm width and 500 nm depth. Before milling the nanochannel to connect the two microchannels an extension in Fig. 2(iii) of 10 μm was milled to control the distance between the two big microchannels and provide a flat and smooth surface for final milling of nanochannel. The milled nanochannel in Fig. 2(iv) has Gaussian-like beam profile in the cross sections. Operating in this mode, the width of the nanochannel obtained for a given film thickness and ion beam current is highly reproducible. It is controlled by the user-defined width of the scan area. After milling, Cr was removed using ceric ammonium nitrate.
The access ports were drilled using abrasive powder blasting. The substrate and cover plate are cleaned using piranha solution. Their surfaces are activated in an oxygen plasma (Harrick Plasma, 18 W, 10 min), and then they are brought into contact, forming a reversible bond. The bonding is made permanent by heating the device to 1100 °C in a vacuum furnace and holding at this temperature for >10 h.
Cell injection and lysis into microfluidic channel
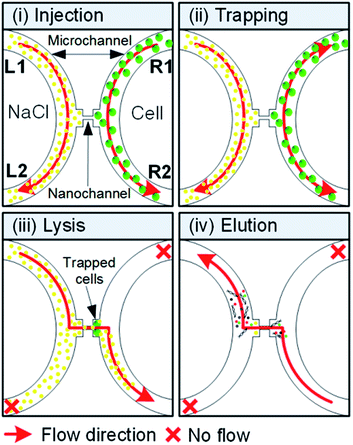
Chlorella vulgaris (ATCC30821) was obtained from the American Type Culture Collection (ATCC). It was grown at 25 °C with the cap loosened one half turn and maintained under a 14/10 h light–dark photoperiod and subcultured every 14–21 days. To quantify the C. vulgaris number, we measured the number of cells using a hemocytometer.The processes for cell injection, trapping, lysis, and elution in microchip are shown in Fig. 3 in brief. 1 × 105 cells per ml diluted with deionized (DI) water in Fig. 3(i) was injected into the R1 port of the microfluidic channel by using a syringe pump (Chemyx, Fusion-200) with flow rate of 3 μl min−1. Once the cells were flowing inside the microfluidic channel, 100 mM NaCl solution for osmotic stresses was injected and maintained with flow rate of 5 μl min−1 into the L1 port of the microchannel. In Fig. 3(ii), microalgae cells were trapped in the middle-extended part of the microchannel by changing the flow direction such as from R1 to R2 and from R2 to R1. By changing the direction from L1 to R2 for the flow of NaCl solution, it flowed across the nanochannel and reached to the other side of the microchannel where the cells were trapped. As soon as the NaCl reached to the other microchannel through the nanochannel, cell lysis was performed for 30 s and shown in Fig. 3(iii). After lysis, L1–L2 microchannel was washed by sterilized water with 5 μl min−1 flow rate. Then genetic materials of lysed alga cell were eluted by starting flow from R2 to L1 with 2 μl min−1 in Fig. 3(iv) and collected in an Eppendorf tube.
Instrument analysis
The microfluidic chip was positioned on an fluorescence microscope (BH 2-UMA, Olympus) to allow monitoring and image data acquisition of the detection of C. vulgaris images using Motic Images plus 2.0 software were taken by Moticam 2000 charge-coupled device (CCD) camera inside the microchannel. The cell movement and lysis was observed using a CCD camera at frame rate of 25 frames per second. The change of the fluorescence intensity of the cell after lysis was observed at fluorescence microscope by adopting a 460–490 nm excitation filter and a 515–550 nm emission filter with a mercury lamp.To determine the quantity and quality of the DNA extracted by the chip, a UV-vis spectrophotometer (Nanodrop, Thermo scientific, DE, USA) was used as a validation method. The quantity of DNA was displayed and purities of nucleic acids were monitored by 260 nm/280 nm absorbance ratio, which also gives the quality of the DNA extraction.25
Result and discussion
The functionality of the nanofluidic device was initially tested using a dye solution. Once flow was demonstrated, indicating that the nanochannel fabrication was successful, cell trapping, lysis, and extraction of genetic material were carried out, as described in the following sections.Nanochannel characterization
The SEM images of the fabricated microfluidic channel were taken before and after the FIB milling. In Fig. 4A two microchannels are shown before FIB milling. The distance between the two microchannels was approximately 18 μm depending on the etching time using HF. In this way, the length of the separation between the micro-channels can be controlled. In our case, we used FIB to extend the microchannel segments to adjust the length of the nanochannel and prepare a flat surface for FIB milling of the nanochannel (Fig. 4B). Once we have the extended microchannel using FIB, the distance between the microchannels is much smaller and we can easily mill a nanochannel with low energy (0.17 nA) ion beam in a very short time (2–3 min depending on the Cr layer thickness). The magnified view of the microchannel from Fig. 4B shows the milled extension of the microchannel in Fig. 4C and the image of cross section for a Gaussian-shaped nanochannel with dimensions of 100 nm width, 250 nm depth, and length of 1 μm shows in Fig. 4D. The length of nanochannel can be varied from few hundred nanometers to 20 μm. The width and depth of the fabricated nanochannel is dependent on the input parameters of the pattern and ion beam energy. A small variation in parameters can vary the dimension of the nanochannel; however it is also difficult to control the ion beam from shifting due to the charging effect from the sample surface. The use of low energy beam and recoating on the surface reduces this effect.After making the nanochannel on the chip it was cleaned by use of Cr etchant (ceric ammonium nitrate) and bonded under vacuum at 1100 °C. This quartz nanochannel chip was then used in the specially designed plastic holder with the inlet and outlet micro bore tubing (Fig. 4E). To confirm that nanochannel was working well and the chip was bonded perfectly we performed a small test with the flow of dye (red fruit dye) in the nano channel. In one microchannel red dye was flowed rate of 10 μl min−1 using a syringe pump and the other channel was filled with deionized (DI) water at same rate of flow. Once completely filled with the aqueous dye solution and DI water, the dye solution was pumped through the nanochannels and, after some time, the dye was observed at the left microchannel because the dye diffused by the concentration gradient via nanochannel in Fig. 4F. This demonstrated that the nanochannel was unblocked and able to transport liquid between the channels.
Cell flow and lysis
The algal cells were injected into the right microchannel (R1–R2) with a flow rate of 3 μl min−1. Once the microchannel was filled with the cells and maintained the flow of the cells, NaCl solution was flowed in the left microchannel (L1–L2) at a flow rate of 5 μl min−1. Once both microchannels were filled and flow was maintained continuously, we trapped a few cells in the small extended ‘beak’ of the right microchannel by changing the direction of the flow by the syringe pump. This process was monitored under fluorescence microscope to observe at trapping in the extended microchannel.C. vulgaris is a spherical microscopic cell with 2–10 μm diameter and has many structural elements similar to plants.26 Algae cells contain chlorophyll, which in are fluorescent under the green fluorescent light, so the algae cells were observed using fluorescence microscopy and shown in Fig. 5A. For trapping, cells were induced and trapped by the contraction/expansion in the microchannel beak by changing the flow direction at R1 and R2 ports. The trapped algae cells were fixed well into the microchannel beak and shown in Fig. 5B (fluorescence) and Fig. 5C (optical). Once we have the cells trapped in the microchannel, the injection of extra alga cells was stopped and the unwanted cells were removed from the microchannel by washing them with sterilized water. The cross flow of NaCl solution from L1 to R2 was started through nanochannel as schematically shown in Fig. 3(iii). The flow rate was maintained with 2 μl min−1 flow rate to keep the trapped cells in their place. An algae cell is large in size (more than 10 μm) and the low flow rate of NaCl solution does not move the cell. The osmotic stress in high concentration of NaCl induced the rupture of alga cell walls in solution following a sudden reduction in osmotic pressure. Lysis of the alga cells was thus triggered and the cell material was released. After the cell lysis is completed, the direction of flow was changed and cell materials along with genetic material (DNA, RNA) tend to flow through nanochannel towards the other side of microchannel. Due to the narrow size of nanochannel, some parts of lysed cell materials were separated on basis of their size in front of nanochannel. Finally the desired cell materials flowed along the microchannel via nanochannel and were collected in vials for further analysis.
After lysis of the cell, the flow of NaCl solution was stopped in left microchannel (L1–L2) and cross flow of sterilized water was started from microchannel R2 to L1. The cell organelles started moving towards microchannel (R1–R2), but due to small dimension of the nanochannel only small size molecules can pass through. In this way we separated the small molecules like DNA, RNA and protein along with some cell debris from the rest of the cell materials. The separated molecules are collected and used for further quantification of the material and after filtration through a 0.2 μm filter as discussed in the following section.
Quantitative analysis
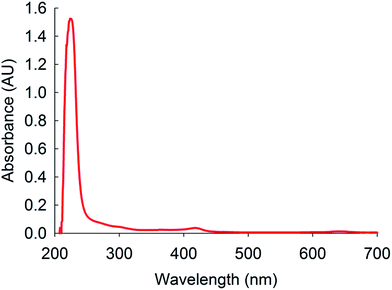
Absorbance and fluorescence spectroscopy for quantification of DNA and RNA are powerful tools in life sciences and therefore used in this work for sample analysis. The collected material was first measured and confirmed by UV-vis spectrometer (Shimadzu UV-visible, UV1601) for quantification of the genetic material. The spectrum was taken on the full range from 200–800 nm. A strong peak was observed at the range of 230–290 nm, which confirms the presence of molecules having double bonds in their chemical structure and are organic in nature (Fig. 6). The DNA, RNA, other protein and nucleotides contain double bonds in their chemical structure. The small peaks at 430 nm and 640 nm are due to the presence of some chlorophyll molecules, which is present only in trace amounts.The presence of DNA, RNA and protein in the collected cell material after lysis was quantified by Nano-Drop 1000 as a UV-vis spectrometer. During the measurement, the sample was analysed at 10 mm path length to report information about the available genetic materials.27 The ratio of absorbance at 260 nm and 280 nm was used to determine the composition of the nucleic acid. If the ratio is appreciably lower than expected, it may indicate the presence of contaminants which absorb at 230 nm. The ratio of absorbance at 260 nm and 280 nm is used to assess the purity of DNA and RNA.23 A ratio of ∼1.8 is generally accepted as “pure” for DNA; a ratio of ∼2.0 is generally accepted as “pure” for RNA. If the ratio is appreciably lower in either case in conventional methods, it may indicate the presence of protein, phenol or other contaminants that absorb strongly at or near 280 nm. It is important to note that the actual ratio will depend on the composition of the nucleic acid. If the ratio is appreciably lower than expected, it may indicate the presence of contaminants which absorb at 230 nm. In our study, otherwise, we absorb at 230 nm. In our study, otherwise, we did not worry about contamination to measure absorbance of DNA and RNA using UV-vis spectrometer because no chemicals except NaCl for lysis were used. After sample extraction from the nanofluidic device, an additional off-chip filtration was applied to purify genetic materials. The results of A260/280 were 1.84 and 1.97 for RNA and 1.72 and 1.8 for dsDNA before and after filtering, respectively. These ratios are accepted as a pure and the measured data were shown in Table 1. All data are averages from three replicate experiments and the concentration data indicate their standard deviations. After filtering, there was no change in the concentration of DNA. However, more than two thirds of the RNA has been lost possibly due to degration. Based on these results, the nanofluidic device shows easy and efficient lysis of cells and extraction of DNA without organic chemicals within short time.
| Type | Conc. (ng ml−1) | A 260 | A 280 | A 260/280 |
|---|---|---|---|---|
| Unfiltered | ||||
| RNA | 3.9 ± 0.6 | 0.098 | 0.053 | 1.84 |
| DNA | 3.1 ± 0.3 | 0.062 | 0.037 | 1.72 |
| Protein | 38.3 ± 6 | — | 0.038 | — |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Filtered | ||||
| RNA | 1.2 ± 0.2 | 0.03 | 0.016 | 1.97 |
| DNA | 2.8 ± 0.6 | 0.056 | 0.036 | 1.8 |
| Protein | 11.5 ± 3 | — | 0.01 | — |
Conclusions
We have successfully prepared a sealed microfluidic device with a nanofluidic channel (100 nm wide, 250 nm deep, and 1 μm long) in a quartz slide using FIB milling technique combined with wet-etching of microchannels The cross-section of the nanofluidic channel was Gaussian-shaped, which was demonstrated to be suitable for extracting cell materials from trapped cells isolated under flow. Cell lysis was carried out using NaCl solution and DNA molecules were extracted from algae cells used as models through the nanofluidic channels under positive pressure. In our experiments, sufficient DNA extraction was achieved to analyze the small sample with a Nano-Drop spectrophotometer. The ratio of A260/280 for the extracted material was 1.72 for DNA and 1.84 for RNA, consistent with expectations. These results, confirmed that this type of device with nanofluidic channel enables the separation of very small amounts of bio-samples.28 Possible outcomes of this study include integration of this kind of nanofluidic channel into a biochip, investigation of the migration of single DNA or protein molecules, or studies of their interactions.Acknowledgements
The authors acknowledge the financial support of Australian Research Council (FT110100711), and the University of Adelaide. Authors acknowledge support of South Australian Node of the Australian National Fabrication Facility (ANFF), for the start-up award to use their lithography and clean room facility and Australian Microscopy and Microanalysis Research Facility (AMMRF) for use their facilities at Adelaide Microscopy.Notes and references
- P. Abgrall and N. T. Nguyen, Anal. Chem., 2008, 80, 2326–2341 CrossRef CAS PubMed.
- H. Andersson and A. van den Berg, Curr. Opin. Biotechnol., 2004, 15, 44–49 CrossRef CAS PubMed.
- J. Hong, J. B. Edel and A. J. deMello, Drug Discovery Today, 2009, 14, 134–146 CrossRef CAS PubMed.
- D. M. Cannon, B. R. Flachsbart, M. A. Shannon, J. V. Sweedler and P. W. Bohn, Appl. Phys. Lett., 2004, 85, 1241–1243 CrossRef CAS.
- K. D. Dorfman, S. B. King, D. W. Olson, J. D. P. Thomas and D. R. Tree, Chem. Rev., 2012, 113, 2584–2667 CrossRef PubMed.
- K.-G. Wang, and H. Niu, in Micro and Nano Technologies in Bioanalysis, ed. R. S. Foote and J. W. Lee, Humana Press, 2009, p. 17 Search PubMed.
- L. D. Menard and J. M. Ramsey, Anal. Chem., 2012, 85, 1146–1153 CrossRef PubMed.
- K. Klepárník and F. Foret, Anal. Chim. Acta, 2013, 800, 12–21 CrossRef PubMed.
- J. O. Tegenfeldt, C. Prinz, H. Cao, R. L. Huang, R. H. Sustin, S. Y. Chou, E. C. Cox and J. C. Sturn, Anal. Bioanal. Chem., 2004, 378, 1678–1692 CrossRef CAS PubMed.
- H. Yin and D. Marshall, Curr. Opin. Biotechnol., 2012, 23, 110–119 CrossRef CAS PubMed.
- S. Le Gac and A. van den Berg, Trends Biotechnol., 2010, 28, 55–62 CrossRef CAS PubMed.
- A. Ros, W. Hellmich, J. Regtmeier, T. T. Duong and D. Anselmetti, Electrophoresis, 2006, 27, 2651–2658 CrossRef CAS PubMed.
- T. Maleki, S. Mohammadi and B. Ziaie, Nanotechnology, 2009, 20, 105302–105308 CrossRef CAS PubMed.
- H. Hoang, I. M. Segers-Nolten, J. W. Berenschot, M. J. de Boer, N. R. Tas, J. Haneveld and M. C. Elwenspoek, J. Micromech. Microeng., 2009, 19, 065017–065027 CrossRef.
- J. W. Hong and S. R. Quake, Nat. Biotech., 2003, 21, 1179–1183 CrossRef CAS PubMed.
- S. Lindström and H. Andersson-Svahn, Biochim. Biophys. Acta, 2011, 1810, 308–316 CrossRef PubMed.
- M. Mahalanabis, J. Do, H. ALMuayad, J. Y. Zhang and C. M. Klapperich, Biomed. Microdevices, 2010, 12, 353–359 CrossRef CAS PubMed.
- M. L. Kovarik and S. C. Jacobson, Anal. Chem., 2009, 81, 7133–7140 CrossRef CAS PubMed.
- K. Mawatari, S. Kubota, Y. Xu, C. Priest, R. Sedev, J. Ralston and T. Kitamori, Anal. Chem., 2012, 84, 10812–10816 CrossRef CAS PubMed.
- L. D. Menard and J. M. Ramsey, Nano Lett., 2011, 11, 512–517 CrossRef CAS PubMed.
- Q. He, S. Chen, Y. Su, Q. Fang and H. Chen, Anal. Chim. Acta, 2008, 628, 1–8 CrossRef CAS.
- L. J. Guo, Adv. Mater., 2007, 19, 495–513 CrossRef CAS.
- D. A. Triant and A. Whitehead, J. Hered., 2009, 100, 246–250 CrossRef CAS PubMed.
- K. Kant and D. Losic, in FIB Nanostructures, ed. Z. M. Wang, Springer, 2013, pp. 1–22 Search PubMed.
- J. A. Glasel, BioTechniques, 1995, 18, 62–63 CAS.
- C. Safi, B. Zebib, O. Merah, P. Pontalier and C. Vaca-Garcia, Renewable Sustainable Energy Rev., 2014, 35, 265–278 CrossRef.
- P. R. Desjardins and D. S. Conklin, Current Protocols in Molecular Biology, John Wiley & Sons, Inc., 2011, pp. A. 3J.1–A. 3J.16 Search PubMed.
- R. N. Zare and S. Kim, Annu. Rev. Biomed. Eng., 2010, 12, 187–201 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2015 |