Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Computationally driven discovery of SARS-CoV-2 Mpro inhibitors: from design to experimental validation†‡
Léa
El Khoury§
 a,
Zhifeng
Jing§
a,
Zhifeng
Jing§
 a,
Alberto
Cuzzolin
b,
Alessandro
Deplano
c,
Daniele
Loco
a,
Boris
Sattarov
a,
Florent
Hédin
a,
Alberto
Cuzzolin
b,
Alessandro
Deplano
c,
Daniele
Loco
a,
Boris
Sattarov
a,
Florent
Hédin
 a,
Sebastian
Wendeborn
d,
Chris
Ho
a,
Dina
El Ahdab
a,
Sebastian
Wendeborn
d,
Chris
Ho
a,
Dina
El Ahdab
 n,
Theo
Jaffrelot Inizan
n,
Mattia
Sturlese
g,
Alice
Sosic
e,
Martina
Volpiana
e,
Angela
Lugato
e,
Marco
Barone
e,
Barbara
Gatto
e,
Maria Ludovica
Macchia
e,
Massimo
Bellanda
f,
Roberto
Battistutta
f,
Cristiano
Salata
n,
Theo
Jaffrelot Inizan
n,
Mattia
Sturlese
g,
Alice
Sosic
e,
Martina
Volpiana
e,
Angela
Lugato
e,
Marco
Barone
e,
Barbara
Gatto
e,
Maria Ludovica
Macchia
e,
Massimo
Bellanda
f,
Roberto
Battistutta
f,
Cristiano
Salata
 h,
Ivan
Kondratov
i,
Rustam
Iminov
i,
Andrii
Khairulin
i,
Yaroslav
Mykhalonok
i,
Anton
Pochepko
i,
Volodymyr
Chashka-Ratushnyi
i,
Iaroslava
Kos
i,
Stefano
Moro
h,
Ivan
Kondratov
i,
Rustam
Iminov
i,
Andrii
Khairulin
i,
Yaroslav
Mykhalonok
i,
Anton
Pochepko
i,
Volodymyr
Chashka-Ratushnyi
i,
Iaroslava
Kos
i,
Stefano
Moro
 g,
Matthieu
Montes
j,
Pengyu
Ren
k,
Jay W.
Ponder
g,
Matthieu
Montes
j,
Pengyu
Ren
k,
Jay W.
Ponder
 lm,
Louis
Lagardère
n,
Jean-Philip
Piquemal
lm,
Louis
Lagardère
n,
Jean-Philip
Piquemal
 *no and
Davide
Sabbadin
*a
*no and
Davide
Sabbadin
*a
aQubit Pharmaceuticals, Incubateur Paris Biotech Santé, 24 Rue du Faubourg Saint Jacques, 75014 Paris, France. E-mail: davide@qubit-pharmaceuticals.com
bChiesi Farmaceutici S.p.A, Nuovo Centro Ricerche, Largo Belloli 11a, 43122, Parma, Italy
cPharmacelera, Torre R, 4a planta, Despatx A05, Parc Cientific de Barcelona, Baldiri Reixac 8, 08028 Barcelona, Spain
dUniversity of Applied Sciences and Arts Northwestern Switzerland, School of LifeSciences, Hofackerstrasse 30, CH-4132 Muttenz, Switzerland
eDepartment of Pharmaceutical and Pharmacological Sciences, University of Padova, via Marzolo 5, 35131, Padova, Italy
fDepartment of Chemistry, University of Padova, via Marzolo 1, 35131, Padova, Italy
gMolecular Modeling Section, Department of Pharmaceutical and Pharmacological Sciences, University of Padua, via F. Marzolo 5, 35131, Padova, Italy
hDepartment of Molecular Medicine, University of Padua, via Gabelli 63, 35121, Padova, Italy
iEnamine Ltd, 78 Chervonotkats'ka Str., Kyiv 02094, Ukraine
jLaboratoire GBCM, EA7528, Conservatoire National des Arts et Métiers, Hesam Université, 2 Rue Conte, 75003 Paris, France
kUniversity of Texas at Austin, Department of Biomedical Engineering, TX 78712, USA
lDepartment of Chemistry, Washington University in Saint Louis, MO 63130, USA
mDepartment of Biochemistry and Molecular Biophysics, Washington University School of Medicine, MO 63110, USA
nSorbonne Université, Laboratoire de Chimie Théorique, UMR 7616 CNRS, 75005, Paris, France. E-mail: jean-philip.piquemal@sorbonne-universite.fr
oInstitut Universitaire de France, 75005, Paris, France
First published on 10th February 2022
Abstract
We report a fast-track computationally driven discovery of new SARS-CoV-2 main protease (Mpro) inhibitors whose potency ranges from mM for the initial non-covalent ligands to sub-μM for the final covalent compound (IC50 = 830 ± 50 nM). The project extensively relied on high-resolution all-atom molecular dynamics simulations and absolute binding free energy calculations performed using the polarizable AMOEBA force field. The study is complemented by extensive adaptive sampling simulations that are used to rationalize the different ligand binding poses through the explicit reconstruction of the ligand–protein conformation space. Machine learning predictions are also performed to predict selected compound properties. While simulations extensively use high performance computing to strongly reduce the time-to-solution, they were systematically coupled to nuclear magnetic resonance experiments to drive synthesis and for in vitro characterization of compounds. Such a study highlights the power of in silico strategies that rely on structure-based approaches for drug design and allows the protein conformational multiplicity problem to be addressed. The proposed fluorinated tetrahydroquinolines open routes for further optimization of Mpro inhibitors towards low nM affinities.
1. Introduction
Since December 2019, the COVID-19 global pandemic has put the entire world on edge.1,2 The disease is due to a coronavirus (CoV) called SARS-CoV-2 (severe acute respiratory syndrome, SARS) that has triggered the start of an unprecedented research effort.3–5 While the vaccination strategy6 has been particularly successful with the rise of mRNA techniques, additional programs have been launched to obtain antivirals able to reduce the impact of COVID-19 on ill patients. Despite these efforts, few potential treatments are presently available with the exception of Paxlovid, a nirmatrelvir/ritonavir combo proposed by Pfizer.7 Due to the persistence of the pandemic, it remains essential to propose new antiviral drugs. A possible strategy consists in designing small molecules to interact with one of the main proteins of the SARS-CoV-2 virus, thus blocking its activity. Among the potential targets, the main protease protein, denoted as Mpro or 3CLpro, is a primary choice8 as it has no human homolog and it is well conserved among coronaviruses,9 especially in terms of the structure of its active site, catalytic dyad, and dimer interface. Furthermore, Mpro is required to release viral proteins for particle assembly, and is thus essential to the virus replication cycle.Developing a new drug targeting the viral Mpro is challenging as it requires extensive resources and the success rate is notoriously low.10 Relying on in silico driven rational design could accelerate the process. In fact, it diminishes the cost by reducing the need for synthetic iterations while also providing an interpretation of the interactions occurring between the target protein and potential inhibitors.
It is important to note that theoretical modeling of Mpro is challenging as the protein exhibits high structural flexibility11–13 leading to high conformational complexity. Mpro is also involved in a variety of complex protein–ligand–solvent interaction networks.12,13 These challenges can be tackled using a high-resolution modeling approach12,13 going beyond rigid docking procedures (see ref. 14 for a detailed discussion of the difficulties of docking approaches in predicting the native binding modes of small molecules within Mpro).
Many studies have been devoted to the design of new Mpro inhibitors3,5,15–25 through joint computational and experimental approaches. In particular, a recent study by the Jorgensen group highlighted the usefulness of relative binding free energy (RBFE) computations as part of the drug design process.26
In this paper, we present a computationally driven discovery and binding mode rationalization of new SARS-CoV-2 Mpro inhibitors. In doing so, we build on our previous high-resolution Mpro molecular dynamics studies.12,13 Here, we explore more deeply some specific subpockets of the substrate binding site of the protease using absolute binding free energy (ABFE) calculations and adaptive sampling grounded on extensive molecular dynamics simulations with high-resolution polarizable force fields (PFFs). Using the GPU-accelerated module27 (GPU = Graphics Processing Unit) of the Tinker-HP molecular dynamics package28 coupled to the AMOEBA PFF,29–32 it has been shown that simulations can reach the required level of accuracy and μs timescales needed to explore the structural rearrangement and interactions profile of this flexible protein.12,13 More precisely, the modeling of Mpro necessitates the ability to evaluate at high resolution various types of key interactions including hydrogen bonds, salt bridges, π–π stacking, and specific solvation effects. Long timescales are required to achieve sufficient sampling. This is now possible by using the large number of graphics processing units (GPUs) that are presently available on supercomputers and high-performance cloud computing platforms. In this study, we combine our computationally driven strategy, using absolute binding free energy computations33–37 and unsupervised adaptive sampling,12,13 with machine learning-assisted property predictions, while conducting extensive characterization experiments including nuclear magnetic resonance (NMR), mass spectrometry (MS), and FRET-based assays to evaluate the activity of the newly designed compounds.
In the following, we introduce our design strategy, which led to non-covalent and covalent inhibitors of Mpro (ESI-Fig. 1‡). Then, we describe how an interplay between experiments and molecular simulations allowed the discovery of a final compound (QUB-00006-Int-07) with a high affinity to the protease (IC50 = 830 ± 50 nM).
2. Computational details
2.1. Systems preparation
The protease dimer structure (PDB code: 7L11) was used for all the MD simulations and it was prepared at physiological pH (pH = 7). This structure has a higher resolution (1.80 Å) than the PDB structure (PDB code: 6LU7) used in our previous work12 (resolution of 2.16 Å). Both structures are of the holo state in complex with covalent inhibitors, and the rotamers of the key residues at the catalytic site (Cys145, His41, His162, His163, and His172) are virtually identical. The protonation states of His residues were assigned based on previous work,38 where His41 and His80 are protonated at the delta carbon atom and all other His residues are epsilon-protonated, which is favorable for substrate binding.38 This is different from our previous work where His64 and His80 are protonated at the delta carbon atom and all other histidines are epsilon-protonated.12 All water molecules were retained except for those that might collide with the ligands.2.2. Simulation protocols
All-atom simulations were performed using Qubit Pharmaceuticals' Atlas platform which enables the use of any type of High-Performance Computing (HPC) system including cloud supercomputing infrastructures. Among its possibilities, Atlas has the ability to efficiently handle polarizable force field molecular dynamics simulations using a custom version of the multi-GPU module27 of the Tinker-HP molecular dynamics package,28,39 to perform docking runs using either Autodock-Vina40 or Autodock-GPU,41 and to enable machine learning predictions of molecular properties.Since molecular docking could suggest reasonable potential binding modes, but does not always rank the most likely binding mode as the best docked pose,14,48 we visually inspected the generated docked poses and chose an ensemble of binding poses with different binding orientations that we used to run MD and ABFE calculations in order to explore the binding mode of QUB-00006-Int-01, as described in the Results and discussion section.
2.3. Quantitative structure–property relationship (QSPR) modeling: predicting solubility using machine learning
Qubit Pharmaceuticals' Atlas internal machine learning-based QSPR module was used to predict the water solubility (log S, S measured in mol L−1) and octanol/water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P). To build a water solubility QSPR predictor, the AqSolDB dataset52 was used as a training set. To predict octanol/water partition coefficients (log
P). To build a water solubility QSPR predictor, the AqSolDB dataset52 was used as a training set. To predict octanol/water partition coefficients (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), the dataset from EPA's OPERA53 was used as a training set.
P), the dataset from EPA's OPERA53 was used as a training set.
Selected datasets were preprocessed and standardized to some extent by authors of the corresponding publications. However, the need for additional processing was identified when doing exploratory data analysis. We discarded compounds with less than two carbon atoms and kept molecules with molecular weight between 50 and 750 daltons. Additional rules of fragment standardization developed at Qubit Pharmaceuticals were applied.
3. Experimental protocol
3.1. Recombinant expression of SARS-CoV-2 Mpro in E. coli
The plasmid pGEX-6P-1 encoding SARS-CoV-2 Mpro56 was a generous gift from Prof. Rolf Hilgenfeld, University of Lübeck, Lübeck, Germany. Protein expression and purification were adapted from Zhang et al.56 The expression plasmid was transformed into E. coli strain BL21 (DE3) and then pre-cultured in YT medium at 37 °C (100 μg mL−1 ampicillin) overnight. The pre-culture was used to inoculate fresh YT medium supplied with an antibiotic and the cells were grown at 37 °C to an OD600 of 0.6–0.8 before induction of overexpression with 0.5 mM isopropyl-D-thiogalactoside (IPTG). After 5 h at 37 °C, cells were harvested by centrifugation (5000g, 4 °C, 15 min) and frozen. The pellets were resuspended in buffer A (20 mM Tris, 150 mM NaCl, pH 7.8) supplemented with lysozyme, DNase I and PMSF for the lysis. The lysate was clarified by centrifugation at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g at 4 °C for 1 h and loaded onto a HisTrap HP column (GE Healthcare) equilibrated with 98% buffer A/2% buffer B (20 mM Tris, 150 mM NaCl, 500 mM imidazole, pH 7.8). The column was washed with 95% buffer A/5% buffer B and then His-tagged Mpro was eluted with a linear gradient of imidazole ranging from 25 mM to 500 mM. Pooled fractions containing the target protein were subjected to buffer exchange with buffer A using a HiPrep 26/10 desalting column (GE Healthcare). Next, PreScission protease was added to remove the C-terminal His tag (20 μg of PreScission protease per mg of target protein) at 12 °C overnight. Protein solution was loaded onto a HisTrap HP column connected to a GSTrap FF column (GE Healthcare) equilibrated in buffer A to remove the GST-tagged PreScission protease, the His-tag, and the uncleaved protein. Mpro was finally purified with a Superdex 75 prep-grade 16/60 (GE Healthcare) SEC column equilibrated with buffer C (20 mM Tris, 150 mM NaCl, 1 mM EDTA, 1 mM DTT, pH 7.8). Fractions containing the target protein at high purity were pooled, concentrated at 25 mg mL−1 and flash-frozen in liquid nitrogen for storage in small aliquots at −80 °C.
000g at 4 °C for 1 h and loaded onto a HisTrap HP column (GE Healthcare) equilibrated with 98% buffer A/2% buffer B (20 mM Tris, 150 mM NaCl, 500 mM imidazole, pH 7.8). The column was washed with 95% buffer A/5% buffer B and then His-tagged Mpro was eluted with a linear gradient of imidazole ranging from 25 mM to 500 mM. Pooled fractions containing the target protein were subjected to buffer exchange with buffer A using a HiPrep 26/10 desalting column (GE Healthcare). Next, PreScission protease was added to remove the C-terminal His tag (20 μg of PreScission protease per mg of target protein) at 12 °C overnight. Protein solution was loaded onto a HisTrap HP column connected to a GSTrap FF column (GE Healthcare) equilibrated in buffer A to remove the GST-tagged PreScission protease, the His-tag, and the uncleaved protein. Mpro was finally purified with a Superdex 75 prep-grade 16/60 (GE Healthcare) SEC column equilibrated with buffer C (20 mM Tris, 150 mM NaCl, 1 mM EDTA, 1 mM DTT, pH 7.8). Fractions containing the target protein at high purity were pooled, concentrated at 25 mg mL−1 and flash-frozen in liquid nitrogen for storage in small aliquots at −80 °C.
3.2. Protein characterization and enzymatic activity
The molecular mass of the recombinant SARS-CoV-2 Mpro was determined by direct infusion electrospray ionization mass spectrometry (ESI-MS) on a Xevo G2-XS QTOF mass spectrometer (Waters). Samples were diluted in 50% acetonitrile with 0.1% formic acid to achieve a final 1 μM concentration of protein. The detected species displayed a mass of 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 796.64 Da, which matches very closely the value of 33
796.64 Da, which matches very closely the value of 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 796.81 Da calculated from the theoretical full-length protein sequence (residues 1–306). To characterize the enzymatic activity of our recombinant Mpro, we adopted a FRET-based assay using the fluorogenic substrate 5-FAM-AVLQ′SGFRK(DABCYL)K (ProteoGenix) harbouring the cleavage site of SARS-CoV-2 Mpro (′ indicates the cleavage site). The fluorescence of the intact peptide is very low since the fluorophore 5-FAM and the quencher Dabcyl are in close proximity. When the substrate is cleaved by the protease, the fluorophore and the quencher are separated, increasing the fluorescence signal. Freshly unfrozen recombinant SARS-CoV-2 Mpro was used in our assays. The assay was performed by mixing 0.05 μM Mpro with different concentrations of substrate (1–128 μM) in the reaction buffer (20 mM Tris–HCl, 100 mM NaCl, 1 mM EDTA and 1 mM DTT, pH 7.3) in the final volume of 100 μL. Fluorescence intensity (Ex = 485 nm/Em = 535 nm) was monitored at 37 °C with a Victor3 microplate reader (PerkinElmer) for 50 min. A calibration curve was created by measuring multiple concentrations (from 0.001 to 5 μM) of free fluorescein in a final volume of 100 μL reaction buffer. Initial velocities were determined from the linear section of the curve, and the corresponding relative fluorescence units per unit of time (ΔRFU/s) were converted to the amount of the cleaved substrate per unit of time (μM s−1) by fitting to the calibration curve of free fluorescein. Inner-filter effect corrections were applied for the kinetic measurements according to ref. 57. The catalytic efficiency kcat/km resulted in 4819 ± 399 s−1 M−1, in line with literature data.56,58
796.81 Da calculated from the theoretical full-length protein sequence (residues 1–306). To characterize the enzymatic activity of our recombinant Mpro, we adopted a FRET-based assay using the fluorogenic substrate 5-FAM-AVLQ′SGFRK(DABCYL)K (ProteoGenix) harbouring the cleavage site of SARS-CoV-2 Mpro (′ indicates the cleavage site). The fluorescence of the intact peptide is very low since the fluorophore 5-FAM and the quencher Dabcyl are in close proximity. When the substrate is cleaved by the protease, the fluorophore and the quencher are separated, increasing the fluorescence signal. Freshly unfrozen recombinant SARS-CoV-2 Mpro was used in our assays. The assay was performed by mixing 0.05 μM Mpro with different concentrations of substrate (1–128 μM) in the reaction buffer (20 mM Tris–HCl, 100 mM NaCl, 1 mM EDTA and 1 mM DTT, pH 7.3) in the final volume of 100 μL. Fluorescence intensity (Ex = 485 nm/Em = 535 nm) was monitored at 37 °C with a Victor3 microplate reader (PerkinElmer) for 50 min. A calibration curve was created by measuring multiple concentrations (from 0.001 to 5 μM) of free fluorescein in a final volume of 100 μL reaction buffer. Initial velocities were determined from the linear section of the curve, and the corresponding relative fluorescence units per unit of time (ΔRFU/s) were converted to the amount of the cleaved substrate per unit of time (μM s−1) by fitting to the calibration curve of free fluorescein. Inner-filter effect corrections were applied for the kinetic measurements according to ref. 57. The catalytic efficiency kcat/km resulted in 4819 ± 399 s−1 M−1, in line with literature data.56,58
3.3. Nuclear magnetic resonance
All the NMR screening experiments were performed with a Bruker Neo 600 MHz spectrometer, equipped with a nitrogen cooled 5 mm Prodigy CryoProbe at 298 K. The ligand binding was monitored by WaterLOGSY (wLogsy)59 and Saturation Transfer Difference (STD)60 experiments in the presence and in the absence of the protein. Samples contained 10 μM Mpro and 100 μM to 2 mM ligand dissolved in 150 mM NaCl, 20 mM phosphate, 5% D2O, and 4% DMSO-d6 (pH = 7.3). WaterLOGSY experiments were performed with a 180° inversion pulse applied to the water signal at 4.7 ppm using a Gaussian-shaped selective pulse of 5 ms. Each WaterLOGSY spectrum was acquired with 320 scans, a mixing time of 1.5 s and a relaxation delay of 4.5 s. STD experiments were performed with 256 scans. Selective saturation of the protein at 0.4 ppm frequency was carried out by a 2 s pulse train (60 Gaussian pulses of 50 ms separated by 1 ms intervals) included in the relaxation delay and a 30 ms spin-lock was used to reduce the broad background protein signal. The estimation of the KD was achieved by a STD titration according to a previously reported procedure and fitting the curves using OriginPro 2018 (OriginPro version 2018 developed by OriginLab Corporation, Northampton, MA, USA). The water suppression was achieved by the excitation sculpting pulse scheme.3.4. Screening of potential Mpro inhibitors and hits validation
A FRET-based assay employed to test the enzymatic activity of the recombinant SARS-CoV-2 Mpro was used to evaluate the ability of the compounds to inhibit its activity in vitro. In fact, inhibition of Mpro by the tested compounds results in a decrease of the fluorescence signal compared to the Mpro activity in the absence of an inhibitor. A preliminary screening was first performed at a single compound concentration to rapidly identify the ability of the compounds to inhibit Mpro activity and to rank them according to their inhibitory activity. The protein was diluted in the reaction buffer (20 mM Tris–HCl, 100 mM NaCl, 1 mM EDTA and 1 mM DTT, pH 7.3) and pipetted into a 96-well plate to a final protein concentration of 0.02 μM in a final volume of 100 μL. Each compound at the final concentration of 100 μM was incubated with Mpro for 20 minutes at room temperature. After incubation, the peptide substrate (5 μM final) was added to initiate the reaction which was monitored for 50 min at 37 °C. The final DMSO amount was 3.75%. Two controls were prepared for each experiment: the peptide substrate in the absence of Mpro (0% Mpro activity, hence minimal fluorescence intensity detected) and the reaction mixture in the absence of the compounds (100% Mpro activity, therefore maximal fluorescence intensity detected). Following the preliminary screening, the most active compounds (hits) were tested at increasing concentrations (0.25, 0.5, 1, 5, 25, 50, 100, and 150 μM) to determine the dose–response curves and calculate IC50 values fitted using GraphPad Prism 5 software. Each experiment was performed in triplicate and the results were used to calculate an average and a standard deviation.3.5. Binding studies by mass spectrometry
Samples were prepared by mixing appropriate volumes of Mpro (10 μM final) with each compound in the reaction buffer (20 mM Tris–HCl, 100 mM NaCl, 1 mM EDTA and 1 mM DTT, pH 7.3). The final mixtures had a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 or 10
1 or 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 compound
1 compound![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) protein molar ratio. Samples were incubated at room temperature for 20 min before analysis. Control experiments were performed on 10 μM solutions of Mpro in the absence of the compounds. Mass spectrometric analyses were carried out in positive ion mode by ESI-MS under denaturing conditions, i.e. water/acetonitrile 50
protein molar ratio. Samples were incubated at room temperature for 20 min before analysis. Control experiments were performed on 10 μM solutions of Mpro in the absence of the compounds. Mass spectrometric analyses were carried out in positive ion mode by ESI-MS under denaturing conditions, i.e. water/acetonitrile 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 with 0.1% formic acid on a Q-Tof Xevo G2S (Waters, Manchester, UK). Data were processed using MassLynx V4.1 software.
50 with 0.1% formic acid on a Q-Tof Xevo G2S (Waters, Manchester, UK). Data were processed using MassLynx V4.1 software.
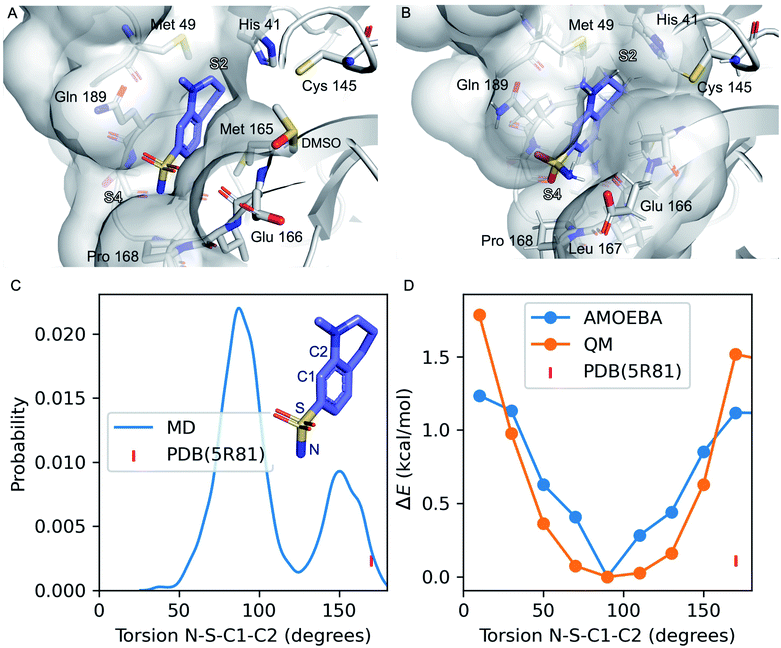 | ||
| Fig. 1 Refinement of the co-crystal structure of x0195 and Mpro using MD simulations. (A) An unusual conformation of x0195 (in purple) located in the binding pocket formed by His41, Met49, Glu166, Gln189, and Pro168 and their surroundings (PDB code: 5R81), and (B) the relaxed structure of x0195 (in purple), obtained after the equilibration step, interacting with the amino acid residues of the substrate binding site. Mpro is shown in light grey. (C) Torsion angle distribution for the sulfonamide group during 20 ns of MD simulations (in blue) performed on the Mpro dimer in complex with x0195; the torsion angle of the sulfonamide group in the co-crystal structure is shown in pink. (D) Torsion energy scan calculated by AMOEBA (in blue) and QM (in orange); the torsion angle of the sulfonamide group in the co-crystal structure is shown in pink. QM level = ωB97x-D/6-31g*.65–67 | ||
3.6. Synthesis
The detailed synthetic protocol used to prepare all molecules can be found in the ESI.‡4. Results and discussion
Several diverse fragments binding the viral Mpro have been identified by high-throughput crystallographic screening of this protease. Among the screened fragments, x0195 (PDB ID: 5R81 (ref. 61) – Fig. 1A) shows one of the highest binding affinities62 and therefore provides a reasonable starting point for fragment-based design of novel Mpro inhibitors.The crystal structure shows that x0195 is located within the Mpro substrate binding pocket, at the interface of the two subpockets S2 and S4 as described by Cannalire et al.63 S4 is a solvent exposed subpocket that is partially composed of a flexible loop delimited by Gln189 and Gln192, while S2 is defined by the side chain residues of Phe140, Asn142, His163, Glu166, and His172, and the backbone atoms of Phe140 and Leu141.
In the co-crystal structure corresponding to Mpro in complex with x0195 (see Fig. 1A), the aromatic portion of the molecule is located between the side chains of Gln189 and Met 165, while the unsaturated region of the tetrahydroquinoline scaffold establishes a hydrophobic interaction with the side chains of His41 and Met49. The N-methyl group attached to the tetrahydroquinoline core is solvent exposed, while the sulfonamide moiety is in contact with Pro168 and Glu166. In particular, the aromatic ring of the small molecule is bisecting the SO2 unit and the polar sulfonamide nitrogen (–NH2) is reaching the boundaries of the hydrophobic part of the binding pocket composed of the alkyl chain of Pro168.
After comparing the available X-ray structural information with previously conducted studies on small molecule conformational preferences derived from crystal structure data,64 we noticed that x0195 was modeled in a high energy conformation and that an unusual high-energy (i.e. repulsive) contact occurs between the sulfonamide oxygen and the carbonyl oxygen of the Glu166 backbone. Additionally, the tetrahydroquinoline scaffold was not fully exploring S2 subpocket boundaries. As reported by Cannalire et al.63 and Zhang et al.,8 the volume of the S2 subpocket in SARS-CoV Mpro is very similar to that of the MERS-CoV homologue. However, the volume of S2 in SARS-CoV Mpro (252 Å3) is significantly larger than in other CoV homologues of the α-genus, such as the HCoV-NL63 Mpro (45 Å3).8,63 Therefore, exploiting this knowledge might be key to designing specific inhibitors of CoV Mpro.
In order to refine the available X-ray structural model and to gather more structural insights (e.g. protein flexibility and binding pocket rearrangements12,13) to guide the design of better binders of the subpocket S2, we ran all-atom molecular dynamics simulations using the AMOEBA polarizable force field29–32 on Mpro (PDB code: 7L11) in complex with x0195 (PDB code: 5R81).
Our simulations show that the unusual high-energy contacts between the sulfonamide oxygen and the carbonyl oxygen of the Glu166 backbone no longer occurred. Also, regarding the electronic structure, we noticed that the p orbitals of the aromatic carbon C1 bisect (e.g. are parallel to) the SO2 angle, compared with a 90° value for the same angle as reported in the crystal structure (see Fig. 1). Moreover, the NH2 of the sulfonamide group is engaged in favorable polar interactions with the Gln189 side chain and the solvent.
Then, we performed absolute binding free energy calculations on the refined protein–ligand structure. Our results show that x0195 binds to the protein with a binding free energy of −2.83 kcal mol−1 at 283 K, which is comparable to the experimental binding energy (−3.59 ± 0.1 kcal mol−1, see Table 1).
| Compound | Computed ΔG | Experimental ΔG |
|---|---|---|
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006(R) 006(R) |
−2.73 ± 0.34 | N.A. |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006(S) 006(S) |
−2.72 ± 0.22 | |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01(R) 006-Int-01(R) |
−4.30 ± 0.35 | −3.71 ± 0.2 |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01(S) 006-Int-01(S) |
−4.45 ± 0.29 | |
| x0195 | −2.83 ± 0.66 | −3.59 ± 0.1 |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-07 006-Int-07 |
−5.37 ± 0.23 | Covalent binder |
We obtained the experimental binding free energy by converting the experimental Kd (1.7 mM ± 0.2) provided in the literature62 using the Gibbs free energy equation and the experimental temperature used in the binding assays (283 K). The agreement of the computed free energy prediction with the experimental results is reasonable. Further analysis of MD simulations suggests that the tetrahydroquinoline scaffold of x0195 is sub-optimally occupying the binding pocket.
We put in place design strategies to modify the chemical moieties of x0195 and potentially increase its binding affinity. Here, we introduce the design of a new molecule, namely QUB-00006 (Fig. 2), where we added two fluorines and a methyl group on the tetrahydroquinoline core of x0195. Also, we substituted the sulfonamide group on the aromatic ring of the molecule by a methanethiol. Fluorination at position 3 of the tetrahydroquinoline core could increase ligand occupancy with no disruption of the water network surrounding the binding pocket,12,13 while methylation at position 4 seemed an interesting modification to increase the potential interactions of the ligand with binding pocket residues. QUB-00006 was generated based on the structure and position of x0195 in the co-crystal (5R81), and then placed in the receptor structure (Mpro dimer with the PDB code: 71LL); next, the Mpro–QUB-00006 complex was equilibrated using MD simulations (see ESI Section 1‡ for the detailed protocol), followed by free energy calculations. To explore the potential of our computational platform in designing new binders with no or few experimental data such as ligand–Mpro co-crystal structures, we leveraged all-atom molecular dynamics simulations on QUB-00006 complexed with Mpro. The aim of this approach is to gather insights on the binding conformation of the newly in silico designed ligand, assess pocket fitness, and evaluate its binding affinity using ABFE calculations.
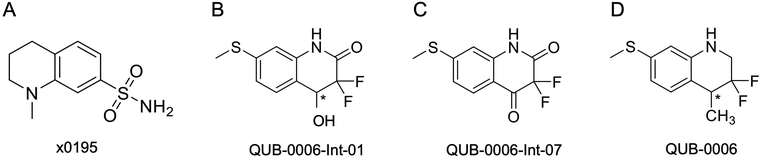 | ||
| Fig. 2 2D structures of (A) x0195, (B) QUB-00006-Int-01, (C) QUB-00006-Int-07, and (D) QUB-00006. The asterisk represents a chiral center. | ||
The initial molecular conformation is mostly anchored at the binding pocket, with the α,α-difluoro-methyl group attached to the tetrahydroquinoline core fully occupying the buried part of the S2 subpocket, which is composed of the side chains of Met49 and His41, while the sulfonamide moiety extends to S4 (Leu167 and Pro168). We note that methylation at position 4 of the tetrahydroquinoline core introduces a chiral center, however no significant differences in terms of pocket occupancy between the R and S enantiomers were observed.
The computed absolute binding free energies for QUB-00006(R) and QUB-00006(S) are −2.73 ± 0.34 kcal mol−1 and −2.72 ± 0.22 kcal mol−1, respectively (Table 1). These results suggest that the designed fluorinated fragment is a binder at the Mpro S2 subpocket and could represent a starting point for structure-based design of novel Mpro inhibitors.
The identified binding mode is defined by several favorable intermolecular interactions occurring between the newly designed ligand and the Mpro binding pocket: (i) the sulfur group of QUB-00006(R) interacts with the oxygen of the carbonyl belonging to the backbone of Glu166 with a distance of 3.3 Å, (ii) the α,α-difluoro moiety points towards His41, and (iii) the sulfur of Met49 establishes a favorable interaction with one of the two fluorines of the substrate (distance 3.3 Å). In fact, the sulfur–oxygen contact observed in our simulations is in agreement with the findings of a study conducted by Iwaoka et al.,68 where they found that a total of 1200 and 626 fragments from the Cambridge Structural Database (CSD) and Protein Data Bank (PDB), respectively, have close intermolecular S–O contacts (with a distance of 3.52 Å or less). Another study analyzing the protein structures deposited in the Protein Data Bank reports 1133 interactions between His and halogen atoms found in 3833 PDB entries with one or more halogenated ligands co-crystallized with a protein.69 Moreover, the strong S–F interaction identified during the simulations is in good agreement with experimentally observed distances for fluorine–sulfur contacts in crystal structures (2.8–3.4 Å).70 It is worth noting that such interactions involving sulfur and halogen atoms are usually better captured with polarizable models than with their classical counterparts.71–73
QUB-00006 was then synthesized following the path in Fig. 4 in order to validate in vitro the simulation outcomes.
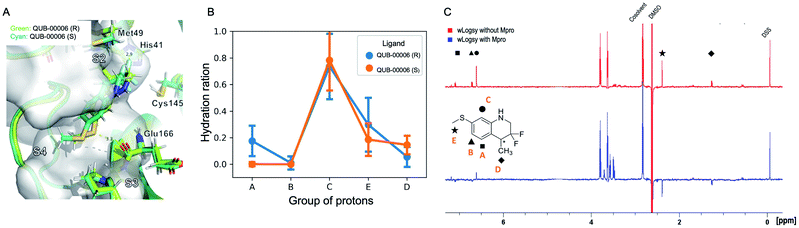
The ligand orientation in the MD simulations and the computed hydration ratio of the different atoms of QUB-00006 during ABFE simulations suggest that proton C is solvent exposed, while the protons of the methyl thioether group (group E) and the methyl group at position 4 of the tetrahydroquinoline core (group D) are buried (Fig. 3B).
Those findings strongly correlate with the NMR characterization of QUB-00006 obtained via WaterLOGSY experiments. In fact, WaterLOGSY epitope mapping confirms that QUB-00006 binds to the protein binding pocket. We leveraged the experimental approach to better identify the region of the ligand in contact with the protein. In Fig. 3C, the proton signals arising from the two methyl groups (D and E) in the presence of Mpro show a change in the sign suggesting that these protons are in close contact with the protein. Similarly, the aromatic protons A and B undergo a sign inversion. In contrast, the aromatic proton C is not significantly perturbed, which suggests that this position is solvent exposed. The binding mode suggested by NMR is in agreement with the MD-derived hydration ratios confirming the predictive power of our MD-based approach to characterize the binding mode of novel ligands at the experimental level of accuracy (Fig. 3B and C).
Although we were able to gather structural information about the binding mode of QUB-00006 using a WaterLOGSY assay, we could not measure its experimental binding affinity via STD NMR due to solubility challenges.
Several synthetic steps were performed in order to obtain QUB-00006, as detailed in Fig. 4. Through this synthetic scheme, we obtained different intermediates characterized by a better solubility profile (Table 2). Interestingly, the hydroxyquinolinone QUB-00006-Int-01 displayed the best solubility profile of all the synthetic intermediates, making it a strong candidate for in vitro evaluation.
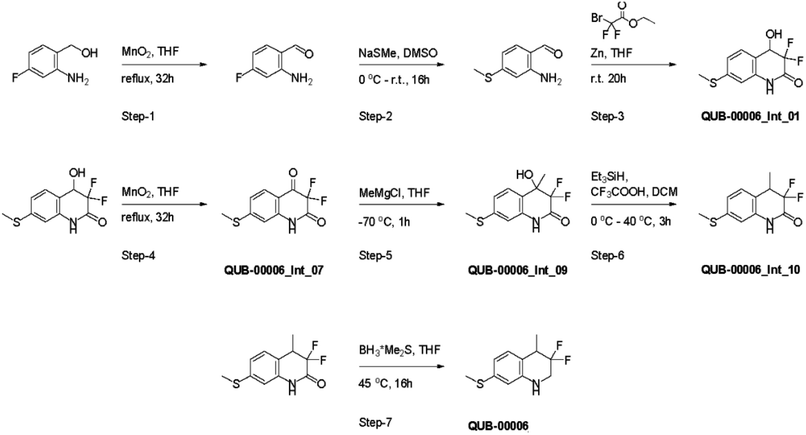 | ||
| Fig. 4 Synthesis path of 3,3-difluoro-4-methyl-7-(methylsulfanyl)-1,2,3,4-tetrahydroquinoline named QUB-00006. | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S is the predicted solubility of the different compounds, log
S is the predicted solubility of the different compounds, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P represents the differential solubility, and the Tanimoto coefficient reflects the similarity of the selected compounds relative to x0195
P represents the differential solubility, and the Tanimoto coefficient reflects the similarity of the selected compounds relative to x0195
| MW (Da) | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S S |
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
Tanimoto (MACCS) | |
|---|---|---|---|---|
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006 006 |
229.07 | −3.99 | 3.56 | 0.391 |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-07 006-Int-07 |
243.02 | −3.73 | 1.96 | 0.371 |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01 006-Int-01 |
245.03 | −2.73 | 1.66 | 0.338 |
| x0195 | 226.08 | −1.94 | 0.56 | 1 |
Before conducting NMR STD experiments to determine the dissociation constant (Kd) of the more polar QUB-00006-Int-01 compound, we decided to predict its binding conformation at the binding pocket and compute the respective absolute binding free energy. Modification of the molecular scaffolds, especially in fragment-like molecules, might affect the binding mode74 compared to a reference structure (e.g. x0195 as per PDB ID:5R81).
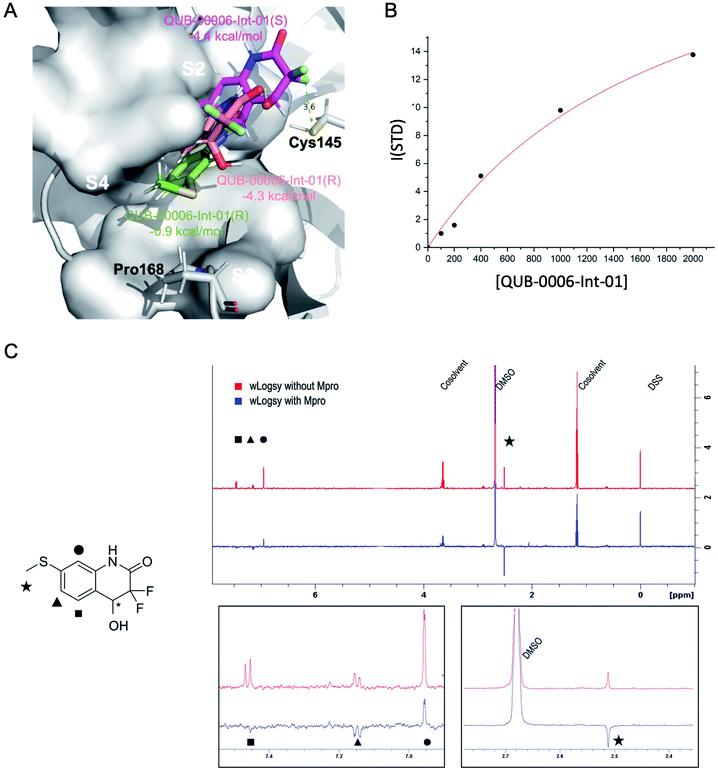
We used a combination of docking, MD and ABFE calculations to explore the putative binding mode of QUB-00006-Int-01. Those calculations identified two dominant binding modes for QUB-00006-Int-01(R) and QUB-00006-Int-01(S) (Fig. 5A) with computed binding free energies of −4.4 and −4.3 kcal mol−1, respectively. Then, we estimated the binding affinity of QUB-00006-Int-01 towards Mpro by a STD NMR titration and we found a dissociation constant in the low millimolar range, with an estimated Kd of 1.9 ± 0.6 mM (−3.71 ± 0.2 kcal mol−1), which agrees reasonably well with our binding free energy calculations (Table 1). As shown in Fig. 5A, both enantiomers bind to the S2 and S4 subpockets with the thioether group being fully buried in subpocket S2, which correlates with WaterLOGSY experiments (Fig. 5C). Additionally, QUB-00006-Int-01(R) and QUB-00006-Int-01(S) fill up a binding pocket space that is different from the one occupied by QUB-00006. On the other hand, starting with a QUB-00006-like binding mode, we ran an additional absolute binding free energy calculation on an Mpro–QUB-00006-Int-01(R) complex and obtained a binding free energy of −0.9 kcal mol−1. These results suggest that QUB-00006-Int-01 and QUB-00006 might have different dominant binding conformations (see Fig. 3A and 5A).
Since a fragment-like molecule could have multiple binding modes and the ligand conformation is unlikely to be fully sampled during 20 ns of binding free energy simulations, we used unsupervised adaptive sampling (AS) to further explore the conformational space of QUB-00006-Int-01. AS can be used here as an interpretative tool able to gather structural insights on the various potential Mpro–ligand interactions (see the ESI‡ for details). The AS trajectories were clustered using average-linkage hierarchical clustering algorithms and the top ten largest clusters were chosen for analysis. These clusters have comparable populations (the smallest clusters have 3–4 times smaller populations or 0.3 kcal mol−1 higher free energy than the largest clusters, see Table 3), indicating the coexistence of multiple binding modes.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01(R) and (S). ΔΔG (kcal mol−1) is the relative free energy at 298 K. The relative binding free energies reported for QUB-00
006-Int-01(R) and (S). ΔΔG (kcal mol−1) is the relative free energy at 298 K. The relative binding free energies reported for QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01(R) and (S) are calculated using the respective cluster 1 as a reference ligand
006-Int-01(R) and (S) are calculated using the respective cluster 1 as a reference ligand
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01(R) 006-Int-01(R) |
QUB-00![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 006-Int-01(S) 006-Int-01(S) |
||||
|---|---|---|---|---|---|
| Cluster | Fraction | ΔΔG | Cluster | Fraction | ΔΔG |
| 1 | 0.101 | 0 | 1 | 0.103 | 0 |
| 2 | 0.083 | 0.05 | 2 | 0.093 | 0.03 |
| 3 | 0.067 | 0.11 | 3 | 0.088 | 0.04 |
| 4 | 0.053 | 0.17 | 4 | 0.065 | 0.12 |
| 5 | 0.042 | 0.23 | 5 | 0.059 | 0.14 |
| 6 | 0.035 | 0.27 | 6 | 0.054 | 0.17 |
| 7 | 0.034 | 0.28 | 7 | 0.049 | 0.19 |
| 8 | 0.033 | 0.29 | 8 | 0.039 | 0.25 |
| 9 | 0.032 | 0.30 | 9 | 0.033 | 0.29 |
| 10 | 0.032 | 0.30 | 10 | 0.031 | 0.31 |
More precisely, starting from these clusters, absolute binding free energies would yield results within 0.3 kcal mol−1 of what was previously obtained. The simulations of QUB-00006-Int-01(R) and QUB-00006-Int-01(S) converged to similar ensembles containing several possible binding modes. Clusters 3, 5, and 6 of QUB-00006-Int-01(R) and cluster 4 of QUB-00006-Int-01(S) (ESI-Fig. 2‡) correspond to the respective dominant binding modes predicted by ABFE simulations (Fig. 5A). For both enantiomers, the most conserved interactions are the hydrophobic contacts between C9 (methyl thioether) and Gln189, and between C5 (proton B) and His41, Arg188, and Gln189.
Overall, our computational findings on QUB-00006-Int-01 confirm that the structural approach we introduce in this work using a sequence of MD-based techniques (classical MD simulations, adaptive sampling, and absolute binding free energy calculations) is able to capture potential binding orientations of fragment-like compounds in the binding pocket of a protein, and to accurately predict their binding free energies.
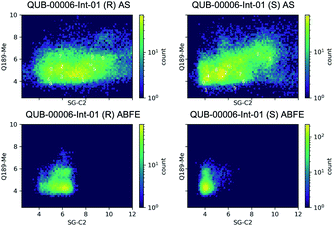
Then, we analyzed the clustered QUB-00006-Int-01 binding conformations from the adaptive sampling simulations plotted as a function of the distance between the methyl thioether group in QUB-00006-Int-01 and the beta carbon of Gln189, and the distance between C2 (carbon connected to the hydroxyl group) and the sulfur atom (SG) of the catalytic side chain of the Cys145 residue (Fig. 6). We noticed that the distance of C2–SG in the most populated cluster generated by the AS simulations is around 4 Å. To reinforce our analysis, we leveraged another unsupervised reduction of dimension technique: TICA (time-lagged independent component analysis),75 which aims at finding the slow collective variables of the data, and applied it to QUB-00006-Int-01(R). We then used the k-means clustering method on the data projected on this space and built a Hidden Markov State Model (HMSM).76 Three clusters emerged, whose characteristics also show the coexistence of several binding modes of QUB-00006-Int-01(R), one of which corresponds to a distance between C2 and SG below 4 Å. Detailed results can be found in the ESI.‡
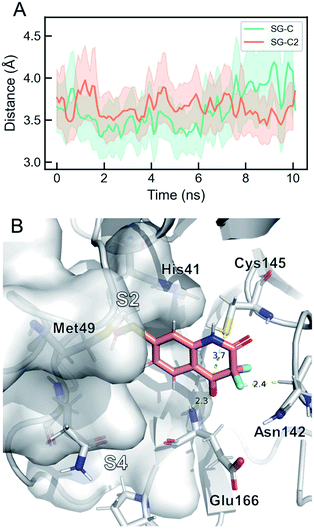
Targeting Cys145 with covalent warheads has been used by several researchers to discover novel potent inhibitors of Mpro.38,63,77 As a matter of fact, a simple chemical modification to QUB-00006-Int-01 would lead to QUB-00006-Int-07 bearing an α,α-difluoro-keto moiety, which is prone to a nucleophilic attack by the vicinal R-SH of Cys145. In order to enable the latter, QUB-00006-Int-07 would need to access the Mpro substrate pocket and adopt a stable binding conformation prior to the covalent binding. Thus, we conducted absolute binding free energy simulations on the Mpro–QUB-00006-Int-07 complex, which confirmed a favorable binding energy of QUB-00006-Int-07 to the Mpro substrate pocket (−5.37 ± 0.23 kcal mol−1). As reported in Fig. 7, compound QUB-00006-Int-07 is bound to the S2 and S4 subpockets with the thioether group being fully buried in subpocket S2 and the α,α-difluoro-keto moiety facing Cys145. More precisely, the average distance between SG and the C is 3.65 angstroms (±0.33) and the average distance between the SG and C2 is 3.61 angstroms (±0.43) as can be seen in Fig. 7.
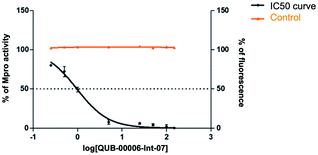
Our computational findings motivated us to test the compound with a FRET-based proteolytic assay. This assay should detect potent functional binders to the viral Mpro. Being a fluorogenic assay, compounds with fluorescence quenching properties can suppress the fluorescence signal generated by the protease activity. To eliminate false positive results, we conducted a preliminary counter screen and verified that the tested compound possesses negligible fluorescence quenching effects. Subsequently, to assess the potential inhibitory activity of the compound against SARS-CoV-2 Mpro, increasing concentrations of QUB-00006-Int-07 (0.25–150 μM) were incubated with 20 nM Mpro before the addition of 5 μM FRET substrate. As shown in Fig. 8, QUB-00006-Int-07 inhibited Mpro with 50% inhibitory concentration (IC50 value of 830 ± 50 nM), thus resulting in a fairly potent inhibitor of the Mpro enzymatic activity. The binding of QUB-00006-Int-07 to Mpro was confirmed by electrospray ionization (ESI) mass spectrometry.
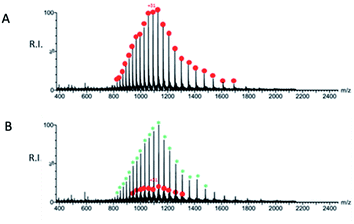
A preliminary determination of the initial protein showed an experimental mass of 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 796.40 Da, which matches very closely the expected value of 33
796.40 Da, which matches very closely the expected value of 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 796.64 Da calculated from the sequence (Fig. 9A). The sample obtained after incubation of QUB-00006-Int-07 with Mpro (compound
796.64 Da calculated from the sequence (Fig. 9A). The sample obtained after incubation of QUB-00006-Int-07 with Mpro (compound![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) protein ratio = 10
protein ratio = 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) was analyzed by ESI-MS under denaturing conditions, and a representative spectrum is provided in Fig. 9B. In addition to the signals corresponding to multiple charge states of the initial protein (red dots), we identified the distribution of signals corresponding to the Mpro modified by the presence of the compound (green asterisks) which is therefore covalently linked to the protein given the non-native conditions of the experiment. The nature of the adduct and the molecular mechanism of binding are under investigation and will be the subject of further studies.
1) was analyzed by ESI-MS under denaturing conditions, and a representative spectrum is provided in Fig. 9B. In addition to the signals corresponding to multiple charge states of the initial protein (red dots), we identified the distribution of signals corresponding to the Mpro modified by the presence of the compound (green asterisks) which is therefore covalently linked to the protein given the non-native conditions of the experiment. The nature of the adduct and the molecular mechanism of binding are under investigation and will be the subject of further studies.
Finally, in this work, the introduction of multiple modifications (e.g. gem-difluoro, thioether, hydroxyl and methyl groups) to the tetrahydroquinoline scaffold of x0195, and the design and synthesis of novel molecular scaffolds, enabled the exploration of binding pocket boundaries and provided additional information related to druggability of the S2 subpocket. Other molecules were produced over the course of this research but, due to their weaker activity, their detailed analysis is not provided here. Their list can be found in the ESI.‡ These compounds were either designed computationally without leading to improved affinities or were synthesis intermediates. All resulting molecules were submitted to biological testing, but none of them were found to be as potent as QUB-00006-Int-07 nor presented a strongly druggable profile, compared to the previously discussed compounds.
5. Conclusion and perspectives
We presented a computationally driven discovery of a new set of non-covalent and covalent inhibitors of Mpro that have been further characterized experimentally. The best compound, QUB-00006-Int-07, has been found to be a covalent binder that resulted in a potent inhibition of the Mpro enzymatic activity (IC50 = 830 ± 50 nM). The results of the innovative scaffold design described here were obtained within three months via a fast-track project that took place in the summer of 2021. It involved a small consortium of theoreticians, organic chemists and drug designers, and demonstrated the effectiveness of a computation-guided synthetic strategy. Indeed, GPU-accelerated high-performance computing platforms can now provide access to high-resolution molecular dynamics simulations, which are able to predict detailed protein conformational maps and provide accurate absolute binding free energy results. Such computations can be further rationalized by means of adaptive sampling simulations, an approach which is able to decipher multiple binding modes. Coupled to NMR, in vitro experiments and machine learning, such high-resolution predictions yield structural insights regarding the design of new active compounds, while offering an atomic level understanding of binding affinities.Beyond this preliminary proof of concept study, the next research steps will be devoted to the QM/MM modeling78,79 of the warhead reaction mechanism38,77,80 leading to the covalent binding of QUB-00006-Int-07, and to optimization of active compounds with the goal of reaching low nanomolar activity.
Data availability
All data have been provided in the main text and ESI.†Author contributions
L. E. K., Z. J., D. L., T. J. I., D. E. A., B. S., and L. L. performed the simulations. L. L., T. J. I., D. S., and J.-P. P. contributed new code. M. S., M. B., A. S., and M. V. performed NMR and mass spectroscopy experiments. I. K., R. I., A. K., Y. M., A. P., and V. C.-R. performed the synthesis. L. L., B. S., P. R., J. W. P., D. S., and J.-P. P. contributed new methodology. L. L., B. S., F. H., P. R., J. W. P., and J.-P. P. contributed analytical tools. All authors analyzed the data. L. E. K., Z. J., L. L., M. S., A. S., D. S. and J.-P. P. wrote the paper. D. S. and J.-P. P. designed the research.Conflicts of interest
P. R., M. M., L. L., J. W. P. and J.-P. P. are co-founders and shareholders of Qubit Pharmaceuticals.Acknowledgements
This work has received funding from the European Research Council (ERC) under the European Union's Horizon 2020 research and innovation program (grant agreement No. 810367), project EMC2 (JPP). Computations have been made possible thanks to special COVID-19 grants from: (i) Amazon Web Services (AWS) allowing access to the AWS Cloud supercomputing platform (JPP; Qubit Pharmaceuticals); (ii) GENCI on the Jean Zay supercomputer (IDRIS, Orsay, France) through projects AP010712339 and AD011012316 (Qubit Pharmaceuticals); GENCI allocation no. A0070707671 (JPP). The work on experimental validation was supported by funding from the CARIPARO Foundation (“Progetti di ricerca sul COVID-19” No. 55812 to BG) and from the Department of Chemical Sciences (project P-DiSC 01BIRD2018-UNIPD to MB). DS thanks Denis Klapishevskiy for discussions, and Prof. Carsten Bolm and his team for providing the synthetic scheme for scaffolds related to the x0195 fragment (https://bolm.oc.rwth-aachen.de/content/outreach).References
- P. Zhou, X.-L. Yang, X.-G. Wang, B. Hu, L. Zhang, W. Zhang, H.-R. Si, Y. Zhu, B. Li and C.-L. Huang, et al. , nature, 2020, 579, 270–273 CrossRef CAS PubMed.
- F. Wu, S. Zhao, B. Yu, Y.-M. Chen, W. Wang, Z.-G. Song, Y. Hu, Z.-W. Tao, J.-H. Tian and Y.-Y. Pei, et al. , Nature, 2020, 579, 265–269 CrossRef CAS PubMed.
- E. N. Muratov, R. Amaro, C. H. Andrade, N. Brown, S. Ekins, D. Fourches, O. Isayev, D. Kozakov, J. L. Medina-Franco, K. M. Merz, T. I. Oprea, V. Poroikov, G. Schneider, M. H. Todd, A. Varnek, D. A. Winkler, A. V. Zakharov, A. Cherkasov and A. Tropsha, Chem. Soc. Rev., 2021, 50, 9121–9151 RSC.
- F. von Delft, M. Calmiano, J. Chodera, E. Griffen, A. Lee, N. London, T. Matviuk, B. Perry, M. Robinson and A. von Delft, A white-knuckle ride of open COVID drug discovery, 2021 Search PubMed.
- J. Breidenbach, C. Lemke, T. Pillaiyar, L. Schäkel, G. Al Hamwi, M. Diett, R. Gedschold, N. Geiger, V. Lopez, S. Mirza, V. Namasivayam, A. C. Schiedel, K. Sylvester, D. Thimm, C. Vielmuth, L. Phuong Vu, M. Zyulina, J. Bodem, M. Gütschow and C. E. Müller, Angew. Chem., Int. Ed., 2021, 60, 10423–10429 CrossRef CAS PubMed.
- F. Krammer, Nature, 2020, 586, 516–527 CrossRef CAS PubMed.
- H. Ledford, et al. , Nature, 2021, 599, 358–359 CrossRef CAS PubMed.
- L. Zhang, D. Lin, Y. Kusov, Y. Nian, Q. Ma, J. Wang, A. von Brunn, P. Leyssen, K. Lanko, J. Neyts, A. de Wilde, E. J. Snijder, H. Liu and R. Hilgenfeld, J. Med. Chem., 2020, 63, 4562–4578 CrossRef CAS PubMed.
- H. Yang, W. Xie, X. Xue, K. Yang, J. Ma, W. Liang, Q. Zhao, Z. Zhou, D. Pei and J. Ziebuhr, et al. , PLoS Biol., 2005, 3, e324 CrossRef PubMed.
- Z. Cournia, B. Allen and W. Sherman, J. Chem. Inf. Model., 2017, 57, 2911–2937 CrossRef CAS PubMed.
- D. W. Kneller, G. Phillips, H. M. O'Neill, R. Jedrzejczak, L. Stols, P. Langan, A. Joachimiak, L. Coates and A. Kovalevsky, Nat. Commun., 2020, 11, 1–6 Search PubMed.
- T. Jaffrelot Inizan, F. Célerse, O. Adjoua, D. El Ahdab, L.-H. Jolly, C. Liu, P. Ren, M. Montes, N. Lagarde, L. Lagardère, P. Monmarché and J.-P. Piquemal, Chem. Sci., 2021, 12, 4889–4907 RSC.
- D. El Ahdab, L. Lagardère, T. J. Inizan, F. Célerse, C. Liu, O. Adjoua, L.-H. Jolly, N. Gresh, Z. Hobaika, P. Ren, R. G. Maroun and J.-P. Piquemal, J. Phys. Chem. Lett., 2021, 12, 6218–6226 CrossRef CAS PubMed.
- S. Zev, K. Raz, R. Schwartz, R. Tarabeh, P. K. Gupta and D. T. Major, J. Chem. Inf. Model., 2021, 61, 2957–2966 CrossRef CAS PubMed.
- Z. Li, X. Li, Y.-Y. Huang, Y. Wu, R. Liu, L. Zhou, Y. Lin, D. Wu, L. Zhang, H. Liu, X. Xu, K. Yu, Y. Zhang, J. Cui, C.-G. Zhan, X. Wang and H.-B. Luo, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 27381–27387 CrossRef CAS PubMed.
- H. T. H. Chan, M. A. Moesser, R. K. Walters, T. R. Malla, R. M. Twidale, T. John, H. M. Deeks, T. Johnston-Wood, V. Mikhailov, R. B. Sessions, W. Dawson, E. Salah, P. Lukacik, C. Strain-Damerell, C. D. Owen, T. Nakajima, K. Świderek, A. Lodola, V. Moliner, D. R. Glowacki, J. Spencer, M. A. Walsh, C. J. Schofield, L. Genovese, D. K. Shoemark, A. J. Mulholland, F. Duarte and G. M. Morris, Chem. Sci., 2021, 12, 13686–13703 RSC.
- A. Morris, W. McCorkindale, T. C. M. Consortium, N. Drayman, J. D. Chodera, S. Tay, N. London and A. A. Lee, Chem. Commun., 2021, 57, 5909–5912 RSC.
- D. Shcherbakov, D. Baev, M. Kalinin, A. Dalinger, V. Chirkova, S. Belenkaya, A. Khvostov, D. Krut’ko, A. Medved’ko, E. Volosnikova, E. Sharlaeva, D. Shanshin, T. Tolstikova, O. Yarovaya, R. Maksyutov, N. Salakhutdinov and S. Vatsadze, ACS Med. Chem. Lett., 2021, 13(1), 140–147 CrossRef PubMed.
- E. Glaab, G. B. Manoharan and D. Abankwa, J. Chem. Inf. Model., 2021, 61, 4082–4096 CrossRef CAS PubMed.
- Z. Jin, X. Du, Y. Xu, Y. Deng, M. Liu, Y. Zhao, B. Zhang, X. Li, L. Zhang, C. Peng, Y. Duan, J. Yu, L. Wang, K. Yang, F. Liu, R. Jiang, X. Yang, T. You, X. Liu, X. Yang, F. Bai, H. Liu, X. Liu, L. W. Guddat, W. Xu, G. Xiao, C. Qin, Z. Shi, H. Jiang, Z. Rao and H. Yang, Nature, 2020, 582, 289–293 CrossRef CAS PubMed.
- J. Gossen, S. Albani, A. Hanke, B. P. Joseph, C. Bergh, M. Kuzikov, E. Costanzi, C. Manelfi, P. Storici, P. Gribbon, A. R. Beccari, C. Talarico, F. Spyrakis, E. Lindahl, A. Zaliani, P. Carloni, R. C. Wade, F. Musiani, D. B. Kokh and G. Rossetti, ACS Pharmacol. Transl. Sci., 2021, 4, 1079–1095 CrossRef CAS PubMed.
- A. Manandhar, V. Srinivasulu, M. Hamad, H. Tarazi, H. Omar, D. J. Colussi, J. Gordon, W. Childers, M. L. Klein, T. H. Al-Tel, M. Abou-Gharbia and K. M. Elokely, J. Chem. Inf. Model., 2021, 61, 4745–4757 CrossRef CAS PubMed.
- G. Amendola, R. Ettari, S. Previti, C. Di Chio, A. Messere, S. Di Maro, S. J. Hammerschmidt, C. Zimmer, R. A. Zimmermann, T. Schirmeister, M. Zappalà and S. Cosconati, J. Chem. Inf. Model., 2021, 61, 2062–2073 CrossRef CAS PubMed.
- M. M. Ghahremanpour, J. Tirado-Rives, M. Deshmukh, J. A. Ippolito, C.-H. Zhang, I. Cabeza de Vaca, M.-E. Liosi, K. S. Anderson and W. L. Jorgensen, ACS Med. Chem. Lett., 2020, 11, 2526–2533 CrossRef CAS PubMed.
- C.-H. Zhang, K. A. Spasov, R. A. Reilly, K. Hollander, E. A. Stone, J. A. Ippolito, M.-E. Liosi, M. G. Deshmukh, J. Tirado-Rives, S. Zhang, Z. Liang, S. J. Miller, F. Isaacs, B. D. Lindenbach, K. S. Anderson and W. L. Jorgensen, ACS Med. Chem. Lett., 2021, 12, 1325–1332 CrossRef CAS PubMed.
- C.-H. Zhang, E. A. Stone, M. Deshmukh, J. A. Ippolito, M. M. Ghahremanpour, J. Tirado-Rives, K. A. Spasov, S. Zhang, Y. Takeo, S. N. Kudalkar, Z. Liang, F. Isaacs, B. Lindenbach, S. J. Miller, K. S. Anderson and W. L. Jorgensen, ACS Cent. Sci., 2021, 7, 467–475 CrossRef CAS PubMed.
- O. Adjoua, L. Lagardère, L.-H. Jolly, A. Durocher, T. Very, I. Dupays, Z. Wang, T. J. Inizan, F. Célerse, P. Ren, J. W. Ponder and J.-P. Piquemal, et al. , J. Chem. Theory Comput., 2021, 17, 2034–2053 CrossRef CAS PubMed.
- L. Lagardère, L.-H. Jolly, F. Lipparini, F. Aviat, B. Stamm, Z. F. Jing, M. Harger, H. Torabifard, G. A. Cisneros, M. J. Schnieders, N. Gresh, Y. Maday, P. Y. Ren, J. W. Ponder and J.-P. Piquemal, Chem. Sci., 2018, 9, 956–972 RSC.
- P. Ren and J. W. Ponder, J. Phys. Chem. B, 2003, 107, 5933–5947 CrossRef CAS.
- J. W. Ponder, C. Wu, P. Ren, V. S. Pande, J. D. Chodera, M. J. Schnieders, I. Haque, D. L. Mobley, D. S. Lambrecht and R. A. DiStasio Jr, et al. , J. Phys. Chem. B, 2010, 114, 2549–2564 CrossRef CAS PubMed.
- Y. Shi, Z. Xia, J. Zhang, R. Best, C. Wu, J. W. Ponder and P. Ren, J. Chem. Theory Comput., 2013, 9, 4046–4063 CrossRef CAS PubMed.
- C. Zhang, C. Lu, Z. Jing, C. Wu, J.-P. Piquemal, J. W. Ponder and P. Ren, J. Chem. Theory Comput., 2018, 14, 2084–2108 CrossRef CAS PubMed.
- D. Jiao, P. A. Golubkov, T. A. Darden and P. Ren, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 6290–6295 CrossRef CAS PubMed.
- Y. Shi, C. Z. Zhu, S. F. Martin and P. Ren, J. Phys. Chem. B, 2012, 116, 1716–1727 CrossRef CAS PubMed.
- R. Qi, Z. Jing, C. Liu, J.-P. Piquemal, K. N. Dalby and P. Ren, J. Phys. Chem. B, 2018, 122, 6371–6376 CrossRef CAS PubMed.
- Z. Jing, J. A. Rackers, L. R. Pratt, C. Liu, S. B. Rempe and P. Ren, Chem. Sci., 2021, 12, 8920–8930 RSC.
- Y. Shi, M. L. Laury, Z. Wang and J. W. Ponder, J. Comput.-Aided Mol. Des., 2021, 35, 79–93 CrossRef CAS PubMed.
- K. Arafet, N. Serrano-Aparicio, A. Lodola, A. J. Mulholland, F. V. González, K. Świderek and V. Moliner, Chem. Sci., 2021, 12, 1433–1444 RSC.
- L.-H. Jolly, A. Duran, L. Lagardère, J. W. Ponder, P. Ren and J.-P. Piquemal, Living Journal of Computational Molecular Science, 2019, 1, 10409 Search PubMed.
- O. Trott and A. J. Olson, J. Comput. Chem., 2010, 31, 455–461 CAS.
- D. Santos-Martins, L. Solis-Vasquez, A. F. Tillack, M. F. Sanner, A. Koch and S. Forli, J. Chem. Theory Comput., 2021, 17, 1060–1073 CrossRef CAS PubMed.
- J. A. Rackers, Z. Wang, C. Lu, M. L. Laury, L. Lagardère, M. J. Schnieders, J.-P. Piquemal, P. Ren and J. W. Ponder, J. Chem. Theory Comput., 2018, 14, 5273–5289 CrossRef CAS PubMed.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, J. Chem. Phys., 1995, 103, 8577–8593 CrossRef CAS.
- L. Lagardère, F. Lipparini, E. Polack, B. Stamm, E. Cancès, M. Schnieders, P. Ren, Y. Maday and J.-P. Piquemal, J. Chem. Theory Comput., 2015, 11, 2589–2599 CrossRef PubMed.
- L. Lagardère, F. Aviat and J.-P. Piquemal, J. Phys. Chem. Lett., 2019, 10, 2593–2599 CrossRef PubMed.
- L. Lagardère, F. Lipparini, E. Polack, B. Stamm, E. Cancès, M. Schnieders, P. Ren, Y. Maday and J.-P. Piquemal, J. Chem. Theory Comput., 2015, 11, 2589–2599 CrossRef PubMed.
- G. M. Morris, R. Huey, W. Lindstrom, M. F. Sanner, R. K. Belew, D. S. Goodsell and A. J. Olson, J. Comput. Chem., 2009, 30, 2785–2791 CrossRef CAS PubMed.
- G. L. Warren, C. W. Andrews, A.-M. Capelli, B. Clarke, J. LaLonde, M. H. Lambert, M. Lindvall, N. Nevins, S. F. Semus, S. Senger, G. Tedesco, I. D. Wall, J. M. Woolven, C. E. Peishoff and M. S. Head, J. Med. Chem., 2006, 49, 5912–5931 CrossRef CAS PubMed.
- R. T. McGibbon, K. A. Beauchamp, M. P. Harrigan, C. Klein, J. M. Swails, C. X. Hernández, C. R. Schwantes, L.-P. Wang, T. J. Lane and V. S. Pande, Biophys. J., 2015, 109, 1528–1532 CrossRef CAS PubMed.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss and V. Dubourg, et al. , Journal of Machine Learning Research, 2011, 12, 2825–2830 Search PubMed.
- C. H. Bennett, J. Comput. Phys., 1976, 22, 245–268 CrossRef.
- M. C. Sorkun, A. Khetan and S. Er, Sci. Data, 2019, 6, 143 CrossRef PubMed.
- K. Mansouri, C. M. Grulke, R. S. Judson and A. J. Williams, J. Cheminf., 2018, 10, 10 Search PubMed.
- D. Bajusz, A. Rácz and K. Héberger, J. Cheminf., 2015, 7, 1–13 CAS.
- H. L. Morgan, J. Chem. Doc., 1965, 5, 107–113 CrossRef CAS.
- L. Zhang, D. Lin, X. Sun, U. Curth, C. Drosten, L. Sauerhering, S. Becker, K. Rox and R. Hilgenfeld, Science, 2020, 368, 409–412 CrossRef CAS PubMed.
- Y. Liu, W. Kati, C.-M. Chen, R. Tripathi, A. Molla and W. Kohlbrenner, Anal. Biochem., 1999, 267, 331–335 CrossRef CAS PubMed.
- C. Ma, M. D. Sacco, B. Hurst, J. A. Townsend, Y. Hu, T. Szeto, X. Zhang, B. Tarbet, M. T. Marty, Y. Chen and J. Wang, Cell Res., 2020, 30, 678–692 CrossRef CAS PubMed.
- C. Dalvit, G. Fogliatto, A. Stewart, M. Veronesi and B. Stockman, J. Biomol. NMR, 2001, 21, 349–359 CrossRef CAS PubMed.
- A. Mier, I. Maffucci, F. Merlier, E. Prost, V. Montagna, G. U. Ruiz-Esparza, J. V. Bonventre, P. K. Dhal, B. Tse Sum Bui, P. Sakhaii and K. Haupt, Angew. Chem., Int. Ed., 2021, 60, 20849–20857 CrossRef CAS PubMed.
- A. Douangamath, D. Fearon, P. Gehrtz, T. Krojer, P. Lukacik, C. D. Owen, E. Resnick, C. Strain-Damerell, A. Aimon, P. Ábrányi Balogh and others, Nat. Commun., 2020, 11, 1–11 Search PubMed.
- A. L. Kantsadi, E. Cattermole, M.-T. Matsoukas, G. A. Spyroulias and I. Vakonakis, J. Biomol. NMR, 2021, 75, 167–178 CrossRef CAS PubMed.
- R. Cannalire, C. Cerchia, A. R. Beccari, F. S. Di Leva and V. Summa, J. Med. Chem., 2020 DOI:10.1021/acs.jmedchem.0c01140.
- K. A. Brameld, B. Kuhn, D. C. Reuter and M. Stahl, J. Chem. Inf. Model., 2008, 48, 1–24 CrossRef CAS PubMed.
- J.-D. Chai and M. Head-Gordon, Phys. Chem. Chem. Phys., 2008, 10, 6615–6620 RSC.
- V. A. Rassolov, J. A. Pople, M. A. Ratner and T. L. Windus, J. Chem. Phys., 1998, 109, 1223–1229 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian ∼16 Revision C.01, Gaussian Inc, Wallingford CT, 2016 Search PubMed.
- M. Iwaoka, S. Takemoto and S. Tomoda, J. Am. Chem. Soc., 2002, 124, 10613–10620 CrossRef CAS PubMed.
- S. Kortagere, S. Ekins and W. J. Welsh, J. Mol. Graphics Modell., 2008, 27, 170–177 CrossRef CAS PubMed.
- M. R. Bauer, R. N. Jones, M. G. J. Baud, R. Wilcken, F. M. Boeckler, A. R. Fersht, A. C. Joerger and J. Spencer, ACS Chem. Biol., 2016, 11, 2265–2274 CrossRef CAS PubMed.
- K. El Hage, J.-P. Piquemal, Z. Hobaika, R. G. Maroun and N. Gresh, J. Comput. Chem., 2013, 34, 1125–1135 CrossRef CAS PubMed.
- X. Mu, Q. Wang, L.-P. Wang, S. D. Fried, J.-P. Piquemal, K. N. Dalby and P. Ren, J. Phys. Chem. B, 2014, 118, 6456–6465 CrossRef CAS PubMed.
- J. Melcr and J.-P. Piquemal, Front. Mol. Biosci., 2019, 6, 143 CrossRef CAS PubMed.
- C. P. Mpamhanga, D. Spinks, L. B. Tulloch, E. J. Shanks, D. A. Robinson, I. T. Collie, A. H. Fairlamb, P. G. Wyatt, J. A. Frearson, W. N. Hunter, I. H. Gilbert and R. Brenk, J. Med. Chem., 2009, 52, 4454–4465 CrossRef CAS PubMed.
- G. Pérez-Hernández, F. Paul, T. Giorgino, G. De Fabritiis and F. Noé, J. Chem. Phys., 2013, 139, 07B604_1 CrossRef PubMed.
- M. K. Scherer, B. Trendelkamp-Schroer, F. Paul, G. Pérez-Hernández, M. Hoffmann, N. Plattner, C. Wehmeyer, J.-H. Prinz and F. Noé, J. Chem. Theory Comput., 2015, 11, 5525–5542 CrossRef CAS PubMed.
- S. Martí, K. Arafet, A. Lodola, A. J. Mulholland, K. Świderek and V. Moliner, ACS Catal., 2022, 12, 698–708 CrossRef PubMed.
- D. Loco, L. Lagardère, S. Caprasecca, F. Lipparini, B. Mennucci and J.-P. Piquemal, J. Chem. Theory Comput., 2017, 13, 4025–4033 CrossRef CAS PubMed.
- D. Loco, L. Lagardère, G. A. Cisneros, G. Scalmani, M. Frisch, F. Lipparini, B. Mennucci and J.-P. Piquemal, Chem. Sci., 2019, 10, 7200–7211 RSC.
- K. Świderek and V. Moliner, Chem. Sci., 2020, 11, 10626–10630 RSC.
Footnotes |
| † Qubit Pharmaceuticals and Sorbonne Université have submitted a preliminary patent application on the compounds reported in this study. |
| ‡ Electronic supplementary information (ESI) available. See DOI: 10.1039/d1sc05892d |
| § These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2022 |