Phenylalanyl tRNA synthetase (PheRS) substrate mimics: design, synthesis, molecular dynamics and antimicrobial evaluation†
Abstract
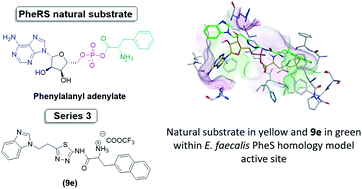
Antimicrobial resistance is a very challenging medical issue and identifying novel antimicrobial targets is one of the means to overcome this challenge. Phenylalanyl tRNA synthetase (PheRS) is a promising antimicrobial target owing to its unique structure and the possibility of selectivity in the design of inhibitors. Sixteen novel benzimidazole based compounds (5a–b), (6a–e), (7a–d), (9a–e) and three N,N-dimethyl-7-deazapurine based compounds (16a–c) were designed to mimic the natural substrate of PheRS, phenylalanyl adenylate (Phe-AMP), that was examined through flexible alignment. The compounds were successfully synthesised chemically in two schemes using 4 to 6-steps synthetic pathways, and evaluated against a panel of five microorganisms with the best activity observed against Enterococcus faecalis. To further investigate the designed compounds, a homology model of E. faecalis PheRS was generated, and PheRS-ligand complexes obtained through computational docking. The PheRS–ligand complexes were subjected to molecular dynamics simulations and computational binding affinity studies. As a conclusion, and using data from the computational studies compound 9e, containing the (2-naphthyl)-L-alanine and benzimidazole moieties, was identified as optimal with respect to occupancy of the active site and binding interactions within the phenylalanine and adenosine binding pockets.


 Please wait while we load your content...
Please wait while we load your content...