Tuned structure and DNA binding properties of metal complexes based on a new 4-acylpyrazolone derivative†
Abstract
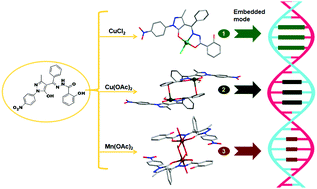
It is common knowledge that the spatial structure of substrates is the major influencing factor in DNA binding. To tune the binding affinity of DNA, a new 4-acylpyrazolone derivative ligand, (2-hydroxy-N′-((5-hydroxy-3-methyl-1-(4-nitrophenyl)-4,5-dihydro-1H-pyrazol-4-yl)(phenyl)methylene)benzohydrazide) (H2L) and its three complexes have been prepared and well characterized. Reaction of H2L with CuCl2 resulted in a mononuclear compound with tetra-coordinated quadrilateral plane, [Cu(HL)Cl] (1). When H2L was coordinated to Cu(OAc)2, a dinuclear Cu(II) compound with chemical formula of [Cu2L2(CH3OH)2]·CH3OH (2) was obtained, and the coordination geometry of Cu(II) is a square pyramid. Upon assembly of H2L with Mn(OAc)2, a quite different dinuclear compound with chemical composition of [Mn2L2(O CH3)2(H2O)2]·CH3OH (3) was afforded, where Mn(III) displayed distorted octahedral configurations. DNA binding studies were performed on H2L and its three complexes by means of electron absorption titration and EB–DNA competition experiments, and the results indicate they all bind DNA in an intercalation mode, and their binding affinity follows the order of 1 > 2 > 3 > H2L. In addition, time-dependent density functional theory (TD-DFT) calculations were performed for H2L and its three complexes to better clarify the electronic transitions in the UV-vis spectra.


 Please wait while we load your content...
Please wait while we load your content...