Identification of new hit scaffolds by INPHARMA-guided virtual screening†
Abstract
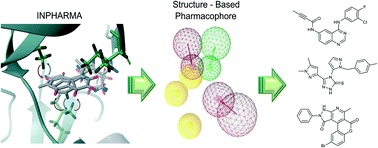
Structure-based drug design (SBDD) relies on the availability of high-quality structures that describe protein–ligand interactions. INPHARMA is an NMR-based method that allows the determination of ligand binding poses to accuracy higher than 2 Å. In this work, we demonstrate that INPHARMA can be used to find novel ligand scaffolds as inhibitors of a model system protein, the cyclin-dependent kinase (Cdk-2). The workflow is given as follows: first, we determine the binding poses to Cdk-2 of six low-affinity fragments and use them to derive a structure-based pharmacophore. Two of the ligands show an unexpected binding mode, which differs from the one observed in crystal structures of other kinases. Second, we use the INPHARMA-generated pharmacophore for virtual screening of the ZINC database; one of the hit compounds is found to bind Cdk-2 in the low μM range and shows selectivity for Cdk-2 against kinases of other families. Our results demonstrate that INPHARMA is an efficient structure-based tool in solution to evolve low-affinity fragments into hit compounds.

 Please wait while we load your content...
Please wait while we load your content...