Open Access Article
Open Access ArticleEngineered bacteria-enabled biosensing: integration with artificial intelligence for enhanced diagnostic precision
Qiuke
Jing
,
Hongmei
Liu
 * and
Decheng
Wu
* and
Decheng
Wu
 *
*
Guangdong Provincial Key Laboratory of Advanced Biomaterials, Department of Biomedical Engineering, Southern University of Science and Technology, Shenzhen 518055, China. E-mail: liuhm@sustech.edu.cn; wudc@sustech.edu.cn
First published on 4th March 2026
Abstract
Engineered bacteria—microbial strains endowed with bespoke functionalities via genetic or protein engineering—represent a cornerstone of synthetic biology. These living diagnostics have shown immense potential across diverse sensing applications, including disease diagnostics, environmental surveillance, and food safety assessment. Recent advances in artificial intelligence (AI) have further propelled the field by enabling the efficient analysis of complex biological datasets, identification of nuanced signal patterns, and rational optimization of genetic circuits. In this review, we first summarize the fundamental design principles underpinning engineered bacterial biosensors, encompassing chassis selection, genetic circuit construction, and signal transduction modules. We then explore emerging roles of AI in component discovery, regulatory circuit refinement, and predictive modeling of system behavior. Representative case studies highlight the translational potential of these intelligent biosensors in real-world monitoring scenarios. Finally, we discuss key challenges—including biosafety, long-term stability, and regulatory hurdles—and propose future directions for the development and clinical implementation of AI-augmented engineered bacterial sensing platforms.
1 Introduction
Accurate and timely detection is critical for public health protection, environmental preservation, and sustainable development. However, as real-world sensing scenarios become more complex—from multiplexed environmental monitoring to rapid, decentralized disease diagnostics—conventional analytical platforms are increasingly constrained by their cost, technical complexity, and limited deployability. For instance, detecting trace pollutants such as heavy metals and volatile organic compounds (VOCs) in environmental matrices remains challenging due to their low abundance and heterogeneous distribution.1,2 In food safety, the prompt identification of microbial pathogens and chemical residues is essential for preventing outbreaks,3 while in clinical settings, early diagnosis is directly linked to improved outcomes and reduced healthcare burdens.4–7 Accurate and timely detection represents a cornerstone of public health protection. Nevertheless, traditional analytical platforms are often hampered by high operational costs, limited throughput, and poor scalability. Detecting trace pollutants is particularly challenging because of their low environmental concentrations. While gold-standard techniques, including chromatography and mass spectrometry, deliver excellent accuracy and sensitivity, their reliance on costly instrumentation and prolonged analytical workflows restricts their applicability for routine, large-scale monitoring. Accordingly, there is a growing interest in developing affordable and scalable alternatives capable of enabling rapid, high-throughput screening, particularly in resource-constrained settings.8,9Engineered bacteria—microorganisms rewired through genetic and protein engineering—offer a promising solution to these limitations. Naturally equipped with diverse signal-sensing and adaptive capabilities, bacteria can be reprogrammed to function as living biosensors, capable of detecting specific chemical or physical cues and generating quantifiable outputs in real time.10,11 These systems are low-cost, self-replicating, and operable in complex environments, making them ideal for field-deployable applications in environmental monitoring, food safety, and medical diagnostics.12
Despite their promise, engineered bacterial biosensors face persistent challenges, including limited sensitivity, slow response kinetics, and suboptimal performance in complex biological matrices. Recently, artificial intelligence (AI) has emerged as a transformative enabler in biosensor engineering. AI techniques—ranging from deep learning to probabilistic modeling—enable high-throughput data analysis, optimization of genetic circuits, and predictive modeling of sensor performance.13 Integrating AI with engineered bacterial detection is expected to overcome the limitations of traditional methods in precision, speed, and adaptability. This fusion enables intelligent, rapid, high-throughput, and cost-effective detection, supporting applications in environmental monitoring, food safety, and personalized medicine.14 The convergence of synthetic biology and AI thus represents a new paradigm for intelligent, adaptive biosensing platforms. In this review, we first discuss core design principles of engineered bacterial biosensors, including chassis strain selection, genetic circuit architecture, and signal transduction mechanisms. We then explore how AI accelerates biosensor development through enhanced component discovery, circuit optimization, and predictive modeling. Representative case studies highlight translational applications in environmental and clinical settings. Finally, we outline current limitations and future opportunities for advancing AI-integrated bacterial biosensing toward clinical and field implementation.
2 Construction of engineered bacteria
Genetically engineered bacteria (GEBs) are microbial strains modified through synthetic biology and genetic engineering to express exogenous proteins, sense environmental cues, or execute programmed functions. The construction of GEBs for biosensing applications involves three key steps: (i) selection of appropriate chassis strains, (ii) integration of sensing and effector modules, and (iii) optimization of signal processing and output systems—including quorum sensing networks and logic circuits—for precise and spatiotemporal control of gene expression.15,162.1 Chassis strains selection
Chassis strain selection frequently varies across different detection scenarios (Table 1). Chassis strain selection forms the foundation of intelligent bacterial biosensor design. The host strain not only dictates the efficiency and stability of synthetic gene circuits but also determines the organism's adaptability, biosafety profile, and long-term viability in physiological or environmental settings.17 In AI-integrated biosensing systems, the inherent biological response dynamics of the chassis directly influence data fidelity, interpretability, and system responsiveness. Therefore, tailoring the chassis strain to the intended application scenario is a critical prerequisite for high-performance engineered biosensor platforms.| Species of strains | Advantages | Disadvantages | Applications | Selection criteria |
|---|---|---|---|---|
| Escherichia coli | Easy genetic engineering; strong ability to express | LPS; inclusion bodies; low stress tolerance | Host intestinal diagnosis; in vitro sensing platform | Select for rapid prototyping and high-yield signal output in sterile IVD or controlled laboratory settings |
| Lactobacillus spp. | GRAS certification; high safety | Slow growth; limited genetic tools | Inflammatory bowel disease; probiotic delivery systems | Choose when human biocompatibility and safety (GRAS) are the primary requirements for long-term oral administration |
| Bacillus subtilis | Good stability; easy for dry transportation | Protease secretion; sporulation control | Portable biological detectors; paper-based diagnostic devices | Select if the biosensor requires long-term shelf-life at room temperature or deployment via paper-based analytical devices |
| Acinetobacter baylyi | Strong natural drug resistance; multi-drug resistance; high natural competence | Fewer standardized promoters and inducible systems; strictly aerobic; potential crosstalk between endogenous and synthetic pathways | Tumor DNA | Prefer for detecting horizontal gene transfer or environmental/tumor DNA via natural transformation mechanisms |
| Attenuated Salmonella (e.g., S. typhimurium strains) | Excellent stability | Risk of virulence reversion; residual endotoxicity (LPS); strict containment regulations | Soil pollution monitoring; detecting pollutants in food (e.g., pesticide residues) | Use for sensing in complex food and soil matrices |
| Bacteroides spp. | Strong natural colonization ability; respond to endogenous signals | Genetic toolkit underdevelopment; difficult manipulation; slower growth; difficult to scale up in standard industrial fermenters | Long term in vivo diagnosis; chronic disease monitoring | Select if the target environment is the anaerobic human colon and long-term stable colonization (>1 week) is required |
| Vibrio natriegens | Fast growth rate; strong salt/pollution tolerance; non-pathogenic | High metabolic demand; elevated oxygenation/cooling costs; corrosive growth media; bioreactor damage risk; replication-induced instability; accelerated plasmid loss | Detection of organic pollutants; detection of pollutants in high salt wastewater | Choose if sample salinity exceeds 1.5% or if a rapid “time-to-result” (<1 hour) is critical for the diagnostic workflow |
| Pseudomonas putida | Good stability; strong ability to resist environmental stress | Metabolic rigidity; flux redirection difficulty; the formed biofilm may clog the equipment; fewer orthogonal genetic parts (promoters/ribosome binding sites) available compared to E. coli | Heavy metal detection; organic pollutant detection; metabolite detection | Prefer for monitoring industrial effluents or environments with high chemical toxicity and fluctuating osmotic pressures |
For environmental applications. Several fast-growing and environmentally resilient bacteria have emerged as preferred chassis for environmental biosensing. Vibrio natriegens, one of the fastest-growing bacterial species known, exhibits high gene-editing efficiency and robust salt tolerance, making it ideal for marine or high-salinity condition.18,19 Engineered variants such as VCOD-2 have demonstrated exceptional competence for exogenous DNA integration, with transformation efficiencies surpassing those of most natural microorganisms.20Pseudomonas putida, a metabolically versatile Gram-negative bacterium, thrives under a wide range of environmental stresses including fluctuations in temperature, pH, organic solvents, and oxidative conditions. Its model strain KT2440, recognized as a safe host organism by the U.S. NIH Recombinant DNA Advisory Committee, has been widely adopted in pollutant detection and metabolic engineering.21 KT2440 has been engineered to degrade persistent environmental toxins such as aromatic hydrocarbons, polycyclic aromatic hydrocarbons, and pesticides.22
For biomedical application. For clinical and diagnostic contexts, Escherichia coli and Lactobacillus species are the most frequently used chassis due to their well-characterized genomes, high genetic tractability, and generally recognized safety.23E. coli remains the prototypical host for in vitro biosensors, thanks to its rapid growth and extensive synthetic biology toolbox. Among clinical strains, E. coli Nissle 1917—a probiotic with a longstanding safety record—is gaining prominence in the development of in vivo biosensors, particularly for gastrointestinal diagnostics due to its ability to colonize the human gut.24Bacillus species also represent promising chassis, especially for applications requiring environmental persistence. Their spore-forming capability provides resilience in extreme or fluctuating conditions, such as those found in the digestive tract. Additionally, non-pathogenic human commensals are being explored as application-specific hosts. For instance, selected skin microbiota strains are under investigation as chassis for dermatological biosensors, owing to their native residence on human skin and their ability to avoid immune clearance.
2.2 Sensing modules
Sensing modules constitute the core functional units of engineered bacterial biosensors, enabling the conversion of pathological or environmental cues—such as heavy metal ions, pollutants, disease-associated metabolites, inflammatory markers, or pathogen-derived molecules—into quantifiable biological outputs. Recent advancements in synthetic biology and molecular engineering have facilitated the development of modular, programmable, and highly specific sensing architectures.25In engineered microbes, sensing modules are responsible for detecting target analytes and initiating downstream responses. Upon recognition of the target, the sensing element triggers signal transduction via mechanisms including ligand-induced conformational changes, allosteric dimerization, conditional protein stabilization, or enzymatic cascade amplification.26 These mechanisms ensure the fidelity and responsiveness of biosensing under physiological or environmental conditions.
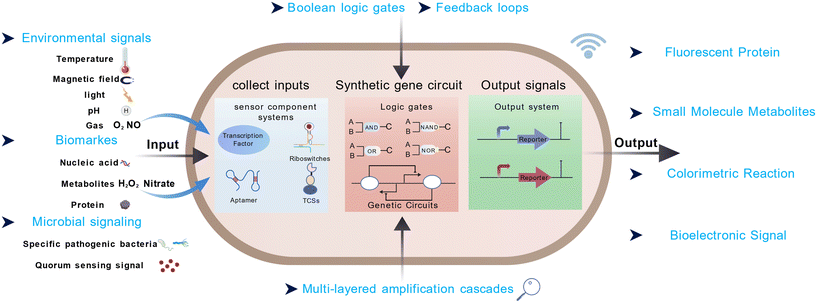
A functional sensing module typically comprises three components: (i) an input recognition element, which determines the specificity and sensitivity of the system; (ii) a signal transduction mechanism, which processes the input into an intracellular cue; and (iii) an output response module, which generates a measurable signal such as fluorescence, color change, or metabolite production (Fig. 1). The performance of the sensing module—especially its sensitivity, dynamic range, and background noise—is critically dependent on the design of the input recognition element. These recognition components can be drawn from natural sensor domains (e.g., transcription factors, two-component systems, or riboswitches) or designed de novo using computational protein engineering and machine learning-guided screening.
The modularity of synthetic sensing systems allows for flexible reconfiguration, enabling bacteria to respond to a wide range of targets with high selectivity. Furthermore, AI-driven strategies are increasingly being applied to enhance component selection, model allosteric behavior, and optimize signal transduction cascades, thereby accelerating the development of next-generation biosensors capable of real-time, multiplexed detection in complex environments.
| Type of sensing element | Typical target | Sensitivity (LOD) | Response time | Advantages | Disadvantages | Application |
|---|---|---|---|---|---|---|
| Transcription factors (TFs) | Heavy metals; antibiotics; intracellular metabolites | High | Medium (30–120 min) | Easy to engineer/modify; tunable dynamic range | Requires cell penetration; leaky expression | General type; intracellular small molecule |
| Two-component systems (TCSs) | Peptides; environmental factors (pH/temperature); extracellular toxins | Extremely high | Relatively fast (20–60 min) | Extracellular sensing; inherent signal amplification | Difficulties in expressing membrane proteins; and cross-talk is prone to occur between systems | Extracellular macromolecules; environmental monitoring |
| Riboswitches | Small molecules (purines; coenzymes; amino acids) | Medium | Fast (<30 min) | Fastest response speed; extremely low metabolic burden | Difficult to detect extremely low concentrations; complex structure design | Real-time monitoring; metabolic flux |
Transcription factors. Transcription factors are endogenous proteins that regulate gene expression in response to small molecules, metabolites, or redox signals.27,28 They are among the most widely used input modules in engineered bacterial systems due to their modularity and adaptability. Natural transcription factors can be repurposed or engineered to recognize non-native ligands, thereby expanding the sensing repertoire. For instance, the transcription factor LldR specifically binds to lactate and is commonly employed for sensing hypoxic tumor microenvironments characterized by elevated lactate levels.29 Mimee et al. utilized engineered bacteria expressing bile acid-responsive transcription factors to monitor gastrointestinal physiology, enabling noninvasive diagnostics of intestinal disorders.30 Other classic examples include MerR and ArsR, which recognize mercury (Hg2+) and arsenic (As3+), respectively. Upon metal ion binding, these transcription factors activate promoter regions that drive reporter gene expression, enabling heavy metal quantification in environmental samples. Additionally, the quorum sensing regulator LuxR, responsive to N-acyl homoserine lactones, has been leveraged in environmental biosensors for detecting petroleum-based pollutants. When coupled with the LuxI operon, this system enables construction of luminescent bacterial reporters for pollutant detection in aquatic ecosystems.
Nucleic acid aptamers. Aptamers are short, single-stranded DNA or RNA sequences capable of binding target molecules—including proteins, metabolites, and nucleic acids—with high affinity and specificity.31 Their synthetic accessibility, biocompatibility, and design flexibility make them attractive components for programmable biosensing. Recent efforts have incorporated aptamers into engineered bacteria to enable the detection of complex biological targets, such as cytokines and tumor biomarkers.32 For example, engineered microbial platforms equipped with cancer-specific aptamers have demonstrated enhanced sensitivity and selectivity in tumor detection. Moreover, strategies that couple extracellular aptamer-based recognition with intracellular signaling circuits have enabled real-time monitoring of immunological markers. Such systems offer a promising avenue for developing in vivo biosensors capable of operating in dynamic disease microenvironments.
Riboswitches. Riboswitches are structured RNA elements that regulate gene expression in response to direct binding of small molecules or ions, typically without requiring intermediary proteins.33,34 Upon ligand binding, riboswitches undergo conformational changes that affect transcription initiation or translation efficiency. These elements offer rapid signal response and minimal background noise, making them suitable for real-time detection of metabolic or ionic changes. Common applications of riboswitches include detection of magnesium ions (Mg2+) and fluoride (F−).35 Recent studies demonstrate that engineered riboswitches can precisely regulate target genes based on exogenous metabolite concentration gradients, revealing substantial application potential in precision medicine and biosensing.36 Their RNA-based nature also makes them amenable to integration with CRISPR-based gene regulation and RNA-sensing platforms.
Two-component systems (TCS). TCS are widespread signal transduction modules in bacteria that enable adaptive responses to a broad range of extracellular and periplasmic stimuli.26,37,38 A canonical TCS consists of a sensor histidine kinase that detects the external signal and a response regulator that modulates gene expression in response.
In contrast to their one-component counterparts, two-component systems (TCS) allow microorganisms to monitor a diverse spectrum of environmental cues, ranging from nutrient concentrations (e.g., amino acids, carbon sources) and the presence of chemical stressors (e.g., antibiotics) to changes in physicochemical conditions such as pH and osmotic pressure.39,40 Engineered TCS circuits have been designed to detect inflammatory cues such as nitric oxide and nitrate, or pathogen-associated molecular patterns such as lipopolysaccharide (LPS).41 Their robust dynamic range, high signal fidelity, and compatibility with complex biological environments make them ideal candidates for in vivo biosensing applications, including inflammation and infection monitoring.
In the context of in vivo disease diagnostics, the target signals—such as aberrant metabolites, inflammation-associated molecules, or tumor-derived biomarkers—are often present at low concentrations and accompanied by substantial background noise.42 These characteristics pose considerable challenges for sensitivity, specificity, and false-positive avoidance in microbial biosensing. To address these limitations, engineered bacterial systems increasingly rely on two key strategies: signal amplification and logic-based signal processing.43
Signal amplification strategies. Signal amplification plays a pivotal role in enhancing weak or transient stimuli into robust, detectable outputs. Several amplification architectures have been developed: (1) cascaded amplification: by arranging signal transduction elements in a hierarchical manner, weak input signals are progressively strengthened through multiple tiers, culminating in a significant phenotypic or reporter response.44 (2) Positive feedback loops (PFLs): upon initial activation, the system amplifies its own input, enabling exponential response propagation. For instance, Sayut et al. demonstrated the use of synthetic PFLs to boost low-level promoter activity, significantly increasing output signal strength.45 (3) Autoinduction systems: these rely on quorum sensing-like strategies, wherein an initially stimulated bacterium produces and releases autoinducers or signal molecules, triggering a synchronized response in neighboring cells.46 This collective behavior not only enhances detection sensitivity but also enables population-level signal integration. These amplification strategies can be customized or combined based on the specific requirements of target analytes, signal dynamics, and the background complexity of the sensing environment.
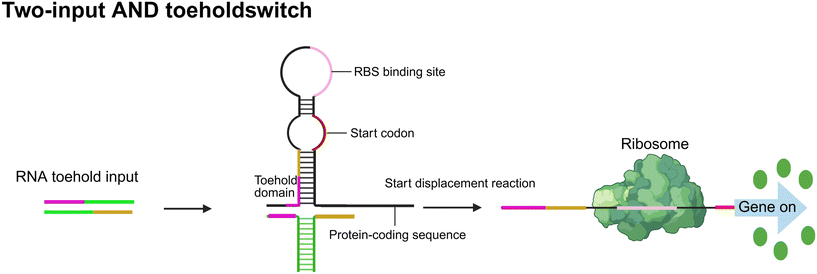
Logic gate-based signal processing. In addition to amplification, logic gate designs are frequently integrated into bacterial circuits to enhance decision-making precision and reduce false-positive responses.47–49 Boolean logic operations such as AND, OR, NOT, and NOR enable bacteria to respond only when signal combinations meet predefined thresholds or contextual constraints: (1) AND gates: trigger outputs only when multiple input conditions are met simultaneously—e.g., co-detection of lactate and hypoxia in tumor microenvironments ensures tumor-specific activation. (2) OR gates: permit response initiation upon recognition of any input signal, offering sensitivity across a broader range of pathophysiological states. (3) NOT/NOR gates: introduce inverse logic, suppressing reporter expression in the presence of inhibitory cues or absence of critical ligands, thereby improving context-selective detection. For example, a two-input AND gate toehold switch design not only enhances signal specificity but also effectively reduces background leakage. This system employs an engineered RNA structure that forms an inhibitory stem-loop, thereby sequestering the ribosome binding site (RBS) and blocking downstream gene translation. Only upon recognition of both specific RNA inputs does the structure unfold, exposing the RBS and initiating protein expression, ensuring that the downstream response is activated only when all input conditions are met50 (Fig. 2).
| Output module | Mechanism | Sensitivity | Kinetics | Robustness | Application scenarios | Limitations |
|---|---|---|---|---|---|---|
| Fluorescent/luminescent | Expression of reporter proteins (GFP; mCherry; bioluminescence) | High | Real-time monitoring possible | Proteins are generally stable; but signal requires external excitation | Laboratory settings; flow cytometry; and in situ imaging | Typically requires external equipment; less suitable for naked-eye field diagnosis |
| Colorimetric | Enzymatic conversion of substrate to visible color (β-galactosidase) | Medium | Dependent on enzymatic turnover rates | Needs optimization | Point-of-care (POC) and low-resource settings | Non-quantitative (often binary yes/no) |
| Small-molecule secretion | Metabolic production of diffusive compounds (indoles; short-chain fatty acids; aldehydes) | High | Faster kinetics than protein-based reporters | Excellent stability and diffusivity | Complex biological samples (e.g., wound exudates, stool) | Requires specific sensor integration |
| Bioelectronic | Extracellular electron transfer (EET) to electrodes | High | Real-time response | Long-term stability | Continuous, remote, and non-invasive monitoring (digital health) | Complex fabrication; requires managing the bio-material interface |
Fluorescent and luminescent outputs. Fluorescent proteins, such as green fluorescent protein (GFP) and mCherry, remain among the most widely used reporters for real-time visualization of bacterial activation. These proteins provide robust, non-invasive readouts and are easily monitored via fluorescence microscopy or flow cytometry.52 In advanced applications, engineered probiotic strains equipped with fluorescent reporters have been integrated with photodetector arrays to achieve in situ signal monitoring. For instance, intelligent bacterial systems sensitive to pro-inflammatory cytokines (e.g., TNF-α) can activate fluorescent or bioluminescent output upon exposure to disease markers.53 An illustrative example is the i-ROBOT system developed by Ye et al., in which fluorescent reporter circuits embedded in engineered E. coli enable noninvasive diagnosis of inflammatory bowel disease (IBD) in mice. Signal intensity in fecal samples correlates with inflammation severity, enabling real-time disease quantification.54
Colorimetric outputs. Colorimetric reactions offer intuitive, equipment-free readouts based on visible color changes triggered by enzymatic activity or chromogenic substrates. One widely adopted example is the expression of β-galactosidase, which catalyzes the conversion of X-gal into a blue precipitate.55 These outputs are ideal for point-of-care and low-resource settings due to their low cost, ease of use, and instrument-free operation. Ongoing efforts aim to enhance the stability and signal-to-noise ratio of colorimetric reporters, enabling broader deployment in portable diagnostic formats.
Small-molecule secretion. Engineered bacteria can also be programmed to produce small-molecule metabolites (e.g., indoles, short-chain fatty acids, or aldehydes) in response to specific stimuli. These compounds exhibit excellent stability and diffusivity, and can be detected using portable analytical devices such as electrochemical sensors.56 Compared to protein-based reporters, small-molecule outputs allow faster kinetics and reduced background interference, making them suitable for real-time detection in complex matrices such as stool or wound exudates.
Bioelectronic readouts. The convergence of synthetic biology and bioelectronics has enabled the development of electrogenic bacterial biosensors, which transduce molecular recognition events into measurable electrical currents. This is typically achieved by engineering extracellular electron transfer (EET) pathways that connect metabolic responses to electrodes.57,58 For example, E. coli engineered to sense nitrate can generate detectable current changes upon target recognition. This principle was applied in an implantable bioelectronic system developed by Mimee et al., which wirelessly transmitted diagnostic signals from the intestinal lumen to external receivers, enabling continuous and non-invasive monitoring of IBD.59 Compared with traditional fluorescent or colorimetric reporters, bioelectronic outputs offer higher temporal resolution, real-time response, remote readout capability, and seamless integration with digital health platforms.60
Future work in this domain is focused on improving long-term stability, minimizing host immune interference, and optimizing the interface between microbial circuits and conductive materials (e.g., carbon nanotubes, flexible electrodes). AI-driven signal processing will further refine noise filtering, pattern recognition, and early anomaly detection in bioelectronic systems.
2.3 AI-driven intelligent design and optimization
AI is emerging as a transformative force in synthetic biology, enabling the rational design, optimization, and deployment of engineered microbial systems across diverse applications, including disease diagnostics, environmental monitoring, and biosensing. Traditional trial-and-error approaches are increasingly being supplanted by AI-driven methodologies, which can accelerate workflows, improve predictive accuracy, and support adaptive decision-making. Recent advances in deep learning, graph neural networks, and evolutionary algorithms have equipped researchers with powerful computational tools that enable the systematic exploration and optimization of microbial chassis, genetic circuits, and sensing systems (Table 4).| Model | Application domain | Model architecture | Prediction accuracy | Data requirements | Computational cost | Strengths | Limitations |
|---|---|---|---|---|---|---|---|
| DeepPromoter65 | Promoter strength prediction | CNN + LSTM | Accuracy up to ∼98–99% on human and mouse promoter datasets | ∼60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 labeled promoter sequences (EPDnew); fixed-length (∼300 bp) DNA sequences 000 labeled promoter sequences (EPDnew); fixed-length (∼300 bp) DNA sequences |
Moderate training cost; single-GPU training feasible; fast inference | High accuracy with relatively small datasets; captures hierarchical sequence features | Limited generalization beyond trained species; relies on labeled promoter datasets |
| AlphaFold2 (ref. 80) | Protein structure prediction | Transformer-based attention networks with Evoformer | Near-experimental accuracy (median backbone RMSD ∼0.96 Å in CASP14) | >100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 PDB structures; millions of protein sequences from UniRef; BFD; MGnify 000 PDB structures; millions of protein sequences from UniRef; BFD; MGnify |
Extremely high; TPU-based training (∼1 week); GPU inference minutes–hours per protein | Unprecedented structural accuracy; broad proteome coverage | High computational and data demands; inference cost scales cubically with sequence length |
| RoseTTAFold81 | Protein structure prediction | 3-Track neural network (sequence; distance; coordinate) | High but lower than AlphaFold2 | PDB + multiple sequence alignments (MSAs) | High but lower than AlphaFold2 | Faster inference; lower hardware requirements | Slightly reduced accuracy |
| ESMFold82 | Protein structure prediction | Large language model (protein LLM) | Moderate–high; lower than AlphaFold2 | Hundreds of millions of protein sequences (no MSAs required) | Moderate; faster inference than AlphaFold2 | MSA-free; scalable to large datasets | Reduced accuracy for complex folds |
| DeepCRISPR67,83 | gRNA design and efficiency prediction | Hybrid CNN-autoencoder | High in human/mouse contexts; moderate in non-mammalian systems | Limited experimentally validated gRNA datasets | High training cost; moderate inference cost (GPU-dependent) | Rapid design and screening | Limited species generality; requires epigenetic data; high data dependency |
Crucially, multi-omics integration has transitioned from descriptive profiling to the predictive discovery of novel biological entities that were previously obscured by data complexity. For instance, in the context of chassis selection, machine learning frameworks applied to metagenomic data have successfully pinpointed specific probiotic strains, such as distinct Lactobacillus reuteri isolates associated with healthy gut metrics, validating their potential as robust vectors for gastrointestinal biosensing.61 Regarding genetic components, explainable AI models (e.g., XGBoost) have mined diverse fungal genomes to identify four previously uncharacterized xylose transporter genes, demonstrating the capacity to expand the library of functional parts available for sensing and metabolic engineering.62 Furthermore, deep learning has revealed characteristic tRNA sequence motifs that accurately distinguish probiotic from pathogenic bacteria, providing a novel, sequence-based metric for evaluating the biosafety of potential biosensor chassis.63 These findings illustrate how AI-driven screening can uncover previously overlooked chassis candidates and functional parts beyond conventional choices. These approaches enable rational chassis selection based on ecological compatibility, safety, and engineering flexibility.
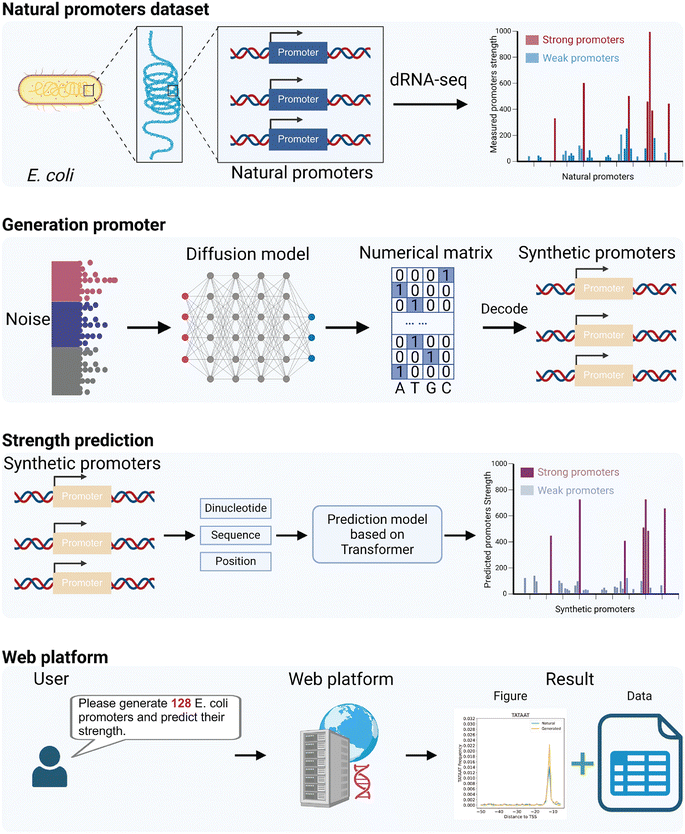 | ||
| Fig. 3 The computational framework for promoter generation and strength prediction.66 | ||
Structure-based protein engineering has also been revolutionized by deep learning models such as AlphaFold, which accurately predict the three-dimensional conformations of sensor proteins, including their ligand-binding domains, conformational stability, and membrane integration.68,69 This structural insight guides the engineering of input modules with improved sensitivity and selectivity. Beyond component-level design, generative adversarial networks (GANs) and reinforcement learning algorithms have enabled the construction of synthetic circuits capable of executing Boolean logic operations (e.g., AND, NOR, NAND), supporting multi-signal integration and programmable microbial behaviours.70 Microfluidic testing of these AI-designed circuits verified that the engineered bacterial populations executed logic operations with digital-like precision, matching the theoretical truth tables generated by the computational models.
AI-guided pathway design tools have also facilitated the optimization of metabolic flux, as exemplified by an E. coli strain engineered for 3-phenylpropanol production with significantly improved yield.73 Genome-scale kinetic models integrated with multi-agent reinforcement learning can iteratively refine enzyme expression profiles without predefined stoichiometric constraints.74 These capabilities support predictive modeling of complex, multi-input genetic systems and enable real-time control of biosynthetic processes.
A major strength of AI-powered platforms lies in their ability to detect subtle phenotypic signatures and complex interactions within high-dimensional datasets. These insights support robust predictions of disease markers, microbial activity states, or biosensor performance. In oncology, AI systems have been used to integrate microbial sensing outputs with clinical imaging data to predict tumour invasiveness and therapeutic response.78 Importantly, these systems exhibit continuous learning capabilities. As additional data are acquired, model parameters are dynamically refined, enhancing diagnostic precision and adaptability. Integration with expert knowledge systems further improves interpretability and user trust, facilitating deployment in clinical or environmental settings.79
By fusing data-driven inference, real-time feedback, and predictive modeling, AI-enabled analytics platforms represent a critical infrastructure for the deployment of intelligent microbial biosensors at scale. This convergence of synthetic biology and artificial intelligence is poised to redefine diagnostic and therapeutic paradigms.
| AI role | Functional objective | AI/ML algorithms | Representative application & outcome |
|---|---|---|---|
| Design optimization | Generation of high-affinity receptors and orthogonal genetic parts | GANs; transformer models (e.g., AlphaFold) | In silico design of specific promoters resulted in a 10-fold increase in sensor sensitivity |
| Performance prediction | Simulation of circuit behavior and host burden before wet-lab assembly | LSTM; CNN; kinetic modeling | Prediction of leakage rates in genetic circuits with >90% accuracy; reducing screening workload |
| Real-time analytics | Decoding complex/noisy output signals into binary diagnostic decisions | SVM, random forest; ANN | Smartphone app processing of colorimetric bacterial signals to classify patient samples with clinical-grade precision |
At the design optimization level, AI-driven generative and structure-prediction models shift biosensor development from empirical screening toward rational and proactive engineering. For example, generative adversarial networks (GANs) and other generative frameworks can propose novel genetic parts with predefined performance constraints, while structure-based deep learning tools such as AlphaFold enable the atomic-level prediction of receptor–ligand interactions, guiding the optimization of sensing proteins and orthogonal regulatory elements before experimental validation.84
At the performance prediction stage, sequence-to-function deep learning models, including convolutional neural networks (CNNs) and long short-term memory (LSTM) architectures, are employed to quantitatively map genetic sequence features to functional outputs. These models enable in silico evaluation of circuit dynamics, signal-to-noise characteristics, and metabolic burden, thereby eliminating low-performing or unstable designs prior to costly build–test cycles and improving overall engineering efficiency.85
Finally, in the real-time data analytics phase, AI serves as the computational engine that translates complex biological signals into actionable diagnostic outcomes. Machine learning classifiers such as support vector machines (SVMs) and random forest algorithms, when coupled with portable or point-of-care readout devices, can robustly decode multiplexed fluorescence, luminescence, or colorimetric outputs under noisy conditions and convert them into accurate, binary diagnostic decisions in real time.86
Collectively, these three functional layers clarify the concrete role of AI in engineered bacterial biosensing, demonstrating how AI systematically enhances design rationality, predictive accuracy, and diagnostic robustness across the entire biosensor lifecycle.
3 Applications of engineered bacteria in multi-scenario detection
3.1 Environmental monitoring
Amidst escalating concerns over environmental pollution, the demand for rapid, sensitive, and cost-effective monitoring tools continues to rise. Engineered bacteria, empowered by synthetic biology, have emerged as promising “living biosensors” for real-time, in situ environmental surveillance. By integrating pollutant-responsive genetic circuits, signal amplification modules, and robust microbial chassis, these systems offer high specificity and sensitivity toward diverse environmental contaminants—including heavy metals, organic compounds, and nutrient overloads. Of particular interest is the recent development of microbial bioelectronic sensors, which transduce biochemical recognition events into electrical outputs, enabling precise and continuous monitoring of target analytes (Table 6).11,87–89 Recent advances have further integrated AI-driven analysis with these living sensors, allowing automated interpretation of complex biosensor outputs and predictive environmental surveillance.90| Application domain | Target analyte | Biosensing mechanism | Operating matrix | Limit of detection (LOD)/range | Response time | Validation level |
|---|---|---|---|---|---|---|
| Environmental92 | Mercury (Hg2+) | E. coli (MerR + fluorescence/electrochem) | Water/buffer | Nanomolar sensitivity | Rapid (<1 h) | In vitro/lab |
| Environmental136 | Phenol | Pseudomonas (conductance-based) | Marine water | 2 μM | Real-time | Field (simulated) |
| Food safety106 | Methyl parathion | S. paucimobilis (enzyme inhibition) | Buffer/food | 0.1–1 μg mL−1 (linear range) | N/A | In vitro |
| Food safety107 | Histamine | P. aeruginosa (transcription factor + fluor.) | Seafood/wine | 0.39 ppm | 90 min | Real food samples |
| Disease (cancer)121,122 | Tumor volatiles/lactate | E. coli Nissle 1917 (metabolic conversion) | Urine | Qualitative presence/100% specificity | N/A | In vivo (mouse) |
| Disease (IBD)137,138 | Thiosulfate | E. coli Nissle 1917 (memory/base editing) | Gut lumen/feces | Qualitative event recording | Real-time (recording) | In vivo (mouse) |
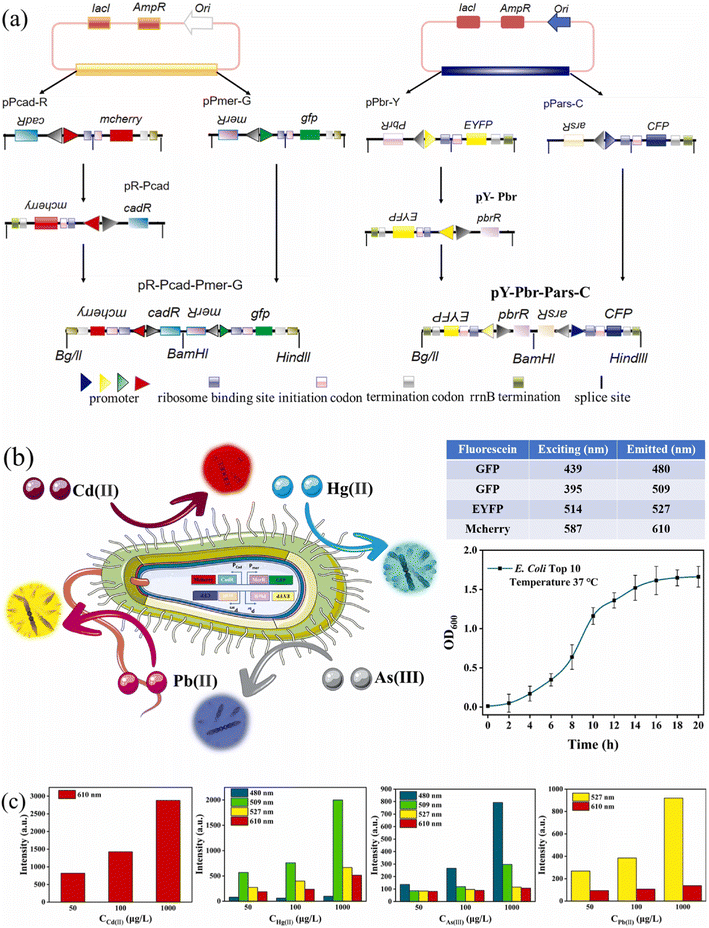 | ||
Fig. 4 (a) Schematic diagram of the construction process of carriers pR-Pcad-Pmer-G (left) and pY-Pbr-Pars-C (right), respectively. (b) The TOP10/pR-Pcad-Pmer-G+pY-Pbr-Pars-C, which integrates multiple promoters and reporter genes, can simultaneously sense stimuli from Cd(II), Hg(II), As(III), and Pb(II) and emit red, green, blue, and yellow fluorescence under corresponding wavelength excitation stimuli. (c) The fluorescence intensity bar graph of TOP10/pR-Pcad-Pmer-G+pY-Pbr-Pars-C cultured in different concentrations (50, 100, 1000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) μg L−1) of Cd(II), Hg(II), As(III), and Pb(II) solutions for 6 μg L−1) of Cd(II), Hg(II), As(III), and Pb(II) solutions for 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) h, respectively.92 h, respectively.92 | ||
To improve environmental robustness and field deployability, bacterial sensors have been immobilized in hydrogels, cellulose membranes, and embedded into microfluidic platforms—enabling the development of portable or wearable biosensing devices for real-time monitoring of heavy metals in aquatic environments.93
3.2 Applications of engineered bacteria in food safety monitoring
Building upon the sensing principles established for environmental monitoring, food safety applications face the unique diagnostic challenge of functioning within complex organic matrices. Biosensing platforms based on engineered bacteria integrate molecular recognition modules with genetic circuits to transduce contaminant detection into visible colorimetric, fluorescent, or electrochemical signals. These systems offer operational simplicity, scalability, and adaptability for on-site applications.Consistent with strategies employed in environmental toxicity monitoring, luminescent bacterial systems have been adapted for pesticide detection through matrix immobilization and optical readout.102,103 For example, Photobacterium leiognathi immobilized in alginate beads enables quantitative assessment of pesticide-induced luciferase inhibition, as demonstrated for 2,4-dichlorophenoxyacetic acid by Ranjan et al.104 Similarly, biohybrid platforms integrating pesticide-degrading bacteria with nanomaterial scaffolds parallel developments in foodborne contaminant sensing. Mishra et al. reported a microplate-based optical sensor combining Sphingomonas paucimobilis with polyethyleneimine-functionalized silica nanoparticles, achieving a linear detection range of 0.1–1 μg mL−1 for methyl parathion and long-term operational stability exceeding six months.105,106
Notably, recent pesticide biosensing systems increasingly adopt AI-enabled signal interpretation pipelines that mirror those described in other application domains. Smartphone-based platforms coupled with deep learning-assisted image analysis standardize optical signal acquisition across heterogeneous sample matrices, while machine learning classifiers such as support vector machines facilitate discrimination between structurally similar pesticide compounds (e.g., methyl parathion versus chlorpyrifos).102 Rather than introducing fundamentally new sensing mechanisms, these AI-driven approaches enhance robustness, specificity, and deployability, reinforcing a convergent trend toward unified, intelligent bacterial biosensing architectures across environmental, food and agricultural monitoring contexts.
3.3 Assessment of soil and crop environments
Crop infections during both cultivation and post-harvest storage pose a major threat to global food security. Diseases such as potato soft rot—caused primarily by Dickeya and Pectobacterium species—can progress rapidly under storage conditions due to the secretion of cell wall-degrading enzymes. Early-stage infections are often asymptomatic and therefore difficult to detect, resulting in missed opportunities for timely intervention and leading to substantial postharvest losses.110,111 Conventional pathogen detection approaches—including morphological characterization, PCR, and ELISA—offer high specificity but are hindered by long processing times, complex sample preparation, and destructive analysis, rendering them impractical for real-time or on-site diagnostics.112–114 Furthermore, these methods generally lack the sensitivity required to detect pathogens during the latent infection phase.Pathogens emit specific VOC profiles as metabolic byproducts during infection, reflecting microbial activity even before visible symptoms appear. Continuous monitoring of VOC fluctuations in the crop microenvironment offers the potential for non-destructive, real-time detection of latent infections. Recent advances in synthetic biology have enabled the engineering of living biosensors—typically bioluminescent bacterial strains—that respond selectively to pathogen-associated VOCs.115 These bacterial systems are programmed with high-affinity olfactory modules and transcriptional circuits that activate luciferase or other reporters upon exposure to trace amounts of infection-related volatiles.
To facilitate practical deployment, engineered bacteria can be immobilized on microfluidic chips or embedded within porous matrices placed near stored crops. Upon detecting VOC levels exceeding predefined thresholds, the bacteria emit bioluminescent signals that can be recorded using portable imaging systems, enabling early diagnosis and rapid mitigation.
Collectively, these emerging biosensing platforms offer a scalable, low-cost, and highly adaptable solution for mitigating postharvest losses and improving crop protection strategies.
3.4 Engineered bacteria for disease detection
Recent advances in synthetic biology have ushered in a new generation of programmable bacterial diagnostics capable of sensing, processing, and responding to disease-associated biomarkers. Engineered bacteria now serve as living diagnostics in diverse clinical contexts, ranging from infectious disease and cancer to gastrointestinal and metabolic disorders. These biosensors leverage modular genetic circuits, quorum-sensing systems, CRISPR-based logic gates, and integration with wearable or ingestible devices to enable real-time, minimally invasive, and context-aware diagnostics in vivo.116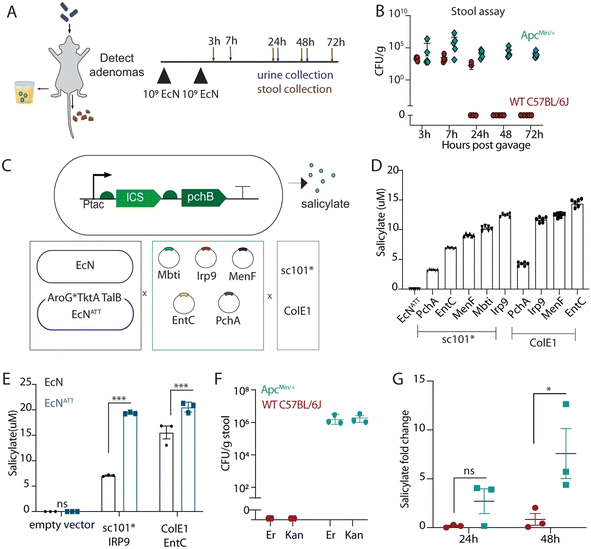 | ||
| Fig. 5 Engineering tumor-colonizing E. coli Nissle 1917 for detection and treatment of colorectal neoplasia. A. Oral delivery and fecal persistence. Schematic of EcN probiotic detection via fecal CFU or urinary metabolite quantification. B. Time-course of EcN recovery from stool pellets in 15-week-old ApcMin/+ mice following oral dosing (n = 5). C. Design of the ICS-pchB salicylate biosynthesis pathway. D and E. LC-MS quantification of salicylate from EcNATT variants, showing enhanced production compared to wild-type EcN (n = 3–6 ; ****p < 0.0001, Two-way ANOVA). Values normalized to D4-salicylate. F. Recovery of total and plasmid-retaining EcNATT from stool 48 h post-dosing (n = 3). G. Urinary salicylate quantification via LC-MS in WT vs. ApcMin/+ mice at 24 and 48 h, demonstrating differential diagnostic potential (n = 3; *p = 0.0228).122 | ||
Beyond metabolic markers, bacteria have been engineered to detect oncogenic mutations. The CRISPR-based CATCH system, for instance, utilizes Acinetobacter baylyi to acquire circulating mutant DNA (e.g. KRAS G12D) and activates kanamycin resistance only upon successful homologous recombination. This allows highly specific mutation detection in vitro (cell lines, organoids) and in vivo (mouse rectal models).5
To address the limitations of conventional imaging, which often fails to detect micrometastases, tumor-tropic Salmonella strains expressing ZsGreen fluorescent protein have been developed. These bacteria preferentially colonize microlesions as small as 0.043 mm3—offering detection sensitivity ∼2600-fold higher than current radiographic methods.123 Additionally, dual-responsive biosensors targeting tumor-specific lactate and hypoxia have been designed to provide spatiotemporal maps of tumor progression via fluorescence imaging.124,125
These systems exploit inflammatory biomarkers such as nitrate, thiosulfate, and hydrogen peroxide. Riglar et al. engineered E. coli with the NarX/NarL system to detect nitrate, enabling real-time in vivo inflammation monitoring.128 Building on this, the i-ROBOT platform developed by Ye and Zhou integrates molecular sensing, logic gating, and therapeutic response in a single EcN chassis. Upon sensing thiosulfate, i-ROBOT initiates base editing to create heritable genomic records and activates colorimetric output while releasing the anti-inflammatory protein AvCystatin. This platform demonstrated diagnostic and therapeutic efficacy in colitis mouse models129 (Fig. 6). By coupling these biological recorders with external AI analysis of fecal biomarkers, clinicians can reconstruct a temporal map of gut inflammation, enabling personalized treatment adjustments.
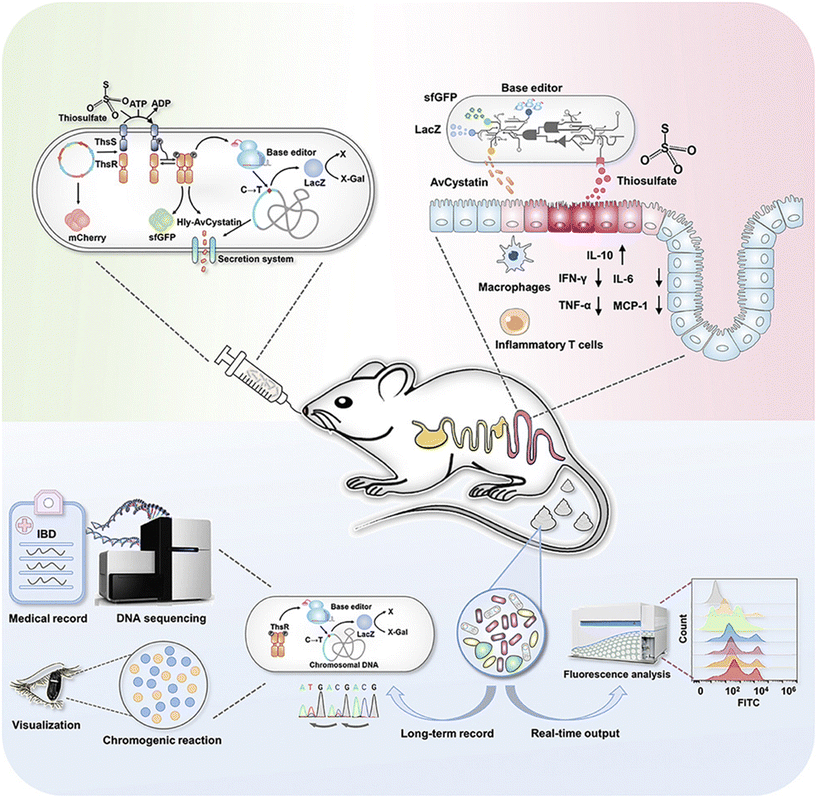 | ||
| Fig. 6 Biomarker-responsive engineered probiotic diagnoses, records, and ameliorates inflammatory bowel disease in mice.129 | ||
To further enhance diagnostic capability, researchers have developed ingestible biosensor devices incorporating engineered bacteria and microelectronics. For instance, commensal E. coli expressing hemoglobin-responsive luciferase were encapsulated in a swallowable device capable of detecting gastrointestinal bleeding in large animal models.30 These “bacterio-electronic” pills transmit real-time luminescence data to external receivers, where machine learning algorithms filter gastrointestinal noise to accurately identify bleeding events. Similarly, Liu et al. embedded bioluminescent heme-responsive bacteria into magnetic hydrogels to enable spatially controllable intestinal diagnostics using external magnets and real-time imaging.130
3.5 Artificial intelligence-driven engineering of bacterial biosensors
The integration of AI into whole-cell bacterial biosensing marks a transition from passive signal readout to data-driven and adaptive perception. In this framework, AI functions as the computational layer linking biological signal generation with quantitative interpretation and rational design. Machine learning algorithms, including artificial neural networks and support vector machines, enable the decoding of complex, non-linear outputs from multiplexed bacterial sensors and effectively alleviate signal crosstalk caused by analyte cross-reactivity, a persistent limitation in whole-cell diagnostics.139 Beyond signal analysis, AI increasingly supports biosensor engineering through data-driven optimization of genetic circuits.Embedded within design–build–test–learn workflows, predictive and generative models facilitate the selection of promoter architectures and regulatory logic, reducing experimental search space and accelerating strain development, particularly for multi-input and high-complexity circuits.140 AI-empowered bacterial biosensors offer several functional advantages. Deep learning-based denoising and feature extraction improve diagnostic precision in heterogeneous matrices such as soil, food, and the gastrointestinal tract, thereby lowering false-positive rates. Moreover, AI enables a shift from binary detection toward quantitative and predictive diagnostics by integrating temporal signal dynamics to infer exposure risk, concentration trends, or system trajectories, including food spoilage and disease progression.141 AI-driven automation further enhances high-throughput screening and evolutionary optimization of large mutant libraries, supporting rapid adaptation to emerging analytes.
Despite these advances, AI integration introduces new challenges. Model robustness remains dependent on training data quality and contextual diversity, limiting generalizability across environments and hosts. In addition, the limited interpretability of complex models—particularly deep neural networks—poses obstacles for regulatory approval and clinical adoption, highlighting the need for explainable AI, standardized datasets, and rigorous validation pipelines tailored to living diagnostics.
4 Current challenges and mitigation strategies for engineered bacteria in detection
Despite significant advances and promising applications across environmental, agricultural, and clinical domains, the practical implementation of engineered bacterial diagnostics continues to face multiple challenges. Key barriers include biosafety concerns, genetic and functional instability, regulatory ambiguity, and context-dependent sensitivity. Importantly, these challenges become more pronounced when engineered bacteria are coupled with AI-enabled diagnostic pipelines, as real-world deployment requires not only biological safety but also algorithmic robustness, interpretability, and regulatory compliance. Addressing these hurdles is therefore essential for enabling the safe, reliable, and scalable translation of living, AI-assisted biosensor technologies.4.1 Biosafety and biosecurity concerns
The deliberate release or clinical use of genetically engineered microorganisms raises critical questions regarding ecological safety and human health. Even in well-characterized systems, the potential for horizontal gene transfer (HGT), unintended ecological impacts, and evolution of harmful traits poses considerable risks.142Of particular concern is the possibility that engineered strains may acquire undesirable traits, such as increased persistence, virulence, or antibiotic resistance, through spontaneous mutation or genetic exchange. For instance, biosensors incorporating antibiotic- or metal-responsive regulators could inadvertently accelerate the environmental spread of resistance determinants if insufficiently contained.143 Such risks are further amplified in complex and dynamic settings, including the human microbiome or open environmental systems, where selective pressures are difficult to predict or control.
From a translational perspective, biosafety risks are not only theoretical but have been experimentally observed to vary across deployment contexts. Engineered bacteria may disseminate synthetic constructs—such as reporter genes or resistance elements—to native microbial communities via HGT, perturbing environmental gene pools or conferring fitness advantages to opportunistic strains. If improperly controlled, engineered bacteria may acquire harmful functions or undesirable antibiotic resistance, potentially leading to microbiome disruption or pathogenicity.143 For example, strains engineered with antibiotic or metal-resistance regulators could inadvertently accelerate the spread of resistance genes in the environment, threatening public health. Additionally, spontaneous mutations under selective pressure may result in the acquisition of undesirable phenotypes, such as increased virulence or persistence. In vivo studies have demonstrated that selective pressures in the gut, including bile acids, immune stress, and microbial competition, can accelerate genetic rearrangements, underscoring the need for context-aware containment strategies prior to clinical translation.
To mitigate these risks, synthetic biologists have developed advanced containment strategies. These include the engineering of auxotrophic strains that depend on synthetic nutrients absent in natural environments, and programmable self-destruction circuits that trigger lysis upon task completion.144 CRISPR-based systems have also been employed to achieve precise, task-dependent genome editing or to facilitate post-delivery clearance by host immunity.145 A particularly promising class of containment tools involves genetic “kill switches”, which activate bacterial suicide pathways in response to predefined stimuli—such as temperature shifts, nutrient depletion, or host-derived signals.146 These kill switches provide multilayered biosafety assurance and can be tuned to environmental or clinical conditions.147 In parallel, efforts are underway to establish standardized frameworks for biosafety evaluation, encompassing immune compatibility, genetic mobility, and long-term ecological impact.
4.2 Genetic stability and functional robustness
For a living diagnostic to be a viable product, it must survive not just the host environment, but also the supply chain. Maintaining genetic stability and functional performance over time is arguably the most significant barrier to commercialization (retained integration/encapsulation points, added manufacturing/storage context). Evolutionary instability: in vivo environments exert dynamic selective pressures (e.g., inflammation, competition) that accelerate evolutionary drift. “Genetic breaking” of sensor circuits leads to signal degradation (loss-of-function) or constitutive activation (false positives), causing “data drift” that renders pre-trained AI models obsolete. Strategies such as bi-stable toggle switches and redundant genetic encoding are being employed to stabilize memory and output.148–150These challenges are especially critical for AI-enabled diagnostics, as algorithmic prediction accuracy is highly sensitive to signal drift and longitudinal inconsistency. To counteract these issues, researchers are exploring several complementary strategies: (1) genomic integration of synthetic circuits reduces reliance on plasmids and enhances heritability. (2) Redundant circuit architectures (e.g., multi-copy integration) improve resilience to mutation and loss. (3) Resource-aware feedback loops modulate gene expression to balance performance and metabolic load. (4) Microencapsulation using hydrogels or biocompatible polymers protects engineered cells from host immunity, gastric acid, and shear stress while preserving viability and function. (5) Ecological support strategies, including co-culture with synergistic microbiota or expression of surface adhesion proteins, help establish stable niches within host or environmental contexts.142 (6) Manufacturing and shelf-life: a critical, often overlooked aspect of translation is the physical form of the diagnostic. Liquid cultures are impractical for widespread distribution. Recent efforts have focused on lyophilization (freeze-drying) and sporulation technologies to create “dormant-to-active” sensors that can be stored at room temperature and reactivated upon use. Ensuring that engineered gene circuits retain functionality after rehydration is a key engineering objective. Microencapsulation serves a dual purpose here: protecting the cells from gastric acid or environmental shear during deployment, and providing a defined micro-niche that stabilizes the population against competitive exclusion by native microbes. Recent in vivo studies demonstrate that encapsulated or chromosomally integrated bacterial sensors can maintain stable signal outputs over weeks, enabling reliable time-series data acquisition required for AI-based trend analysis and predictive modeling. When applied in combination, these strategies significantly improve the robustness, lifespan, and translational feasibility of engineered bacterial diagnostics in both ex vivo and in vivo settings.
4.3 Regulatory and ethical considerations
The transition of engineered bacterial diagnostics from laboratory prototypes to clinical and field-ready platforms is hindered by regulatory uncertainty and unresolved ethical questions. Despite their potential as precision medicine tools, these systems operate at the intersection of synthetic biology, data science, and healthcare—domains where existing regulatory frameworks remain fragmented or insufficient.143 From a regulatory readiness standpoint, living diagnostics are increasingly evaluated under frameworks developed for live biotherapeutic products. In the United States, the FDA currently classifies engineered bacterial systems intended for in vivo use as biological products, requiring Investigational New Drug submissions, Good Manufacturing Practice compliance, and validated assays for identity, purity, potency, and genetic stability. However, explicit guidance for diagnostic-only living organisms—particularly those integrated with AI-based data interpretation—remains underdeveloped. Similar regulatory gaps exist within European Medicines Agency and National Medical Products Administration frameworks, complicating international translation.In vivo diagnostic platforms that collect host-derived physiological or molecular data raise pressing concerns regarding data privacy, security, and ownership. These concerns are amplified in AI-enabled systems that rely on cloud-based computation or continuous data aggregation. Current protocols often lack robust mechanisms for anonymization, secure storage, or responsible data sharing—particularly in systems linked to cloud computing or AI-enabled analysis. Moreover, there is no global consensus on clinical trial design, quality control benchmarks, or approval pathways for microbial diagnostic tools.
To navigate these issues, several actions are essential: (1) establish standardized clinical guidelines for engineered bacterial diagnostics, including preclinical models, trial endpoints, and safety evaluation criteria. (2) Develop regulatory frameworks tailored to living biosensors, informed by precedents from gene and cell therapy. (3) Implement data governance protocols, such as tiered access control, cryptographic encryption, and real-time consent management. (4) Explore privacy-preserving technologies, including federated learning and blockchain, to enable secure, cross-institutional data sharing.
Proactive engagement with regulatory agencies, coupled with transparent risk–benefit assessment and explainable AI design, will be critical for the ethical and equitable deployment of AI-enabled living diagnostics in real-world healthcare and environmental monitoring.
Outlook
The convergence of synthetic biology, microbial engineering, and artificial intelligence is redefining the landscape of biosensing technologies. Engineered bacteria—rationally programmed to detect specific biomarkers, respond to dynamic microenvironments, and produce quantifiable outputs—are emerging as powerful tools for next-generation diagnostics. By integrating modular genetic circuits with adaptive sensing mechanisms, these living systems offer spatiotemporal resolution, real-time analysis, and seamless compatibility with both in vitro and in vivo platforms.The incorporation of artificial intelligence further elevates the capabilities of bacterial biosensors. Machine learning and neural network-based models are increasingly employed to decode complex biological signals, optimize circuit performance, and predict system behavior under multifactorial conditions. Looking ahead, AI-assisted microbial systems may acquire context-aware “biological intelligence”—enabling them to autonomously sense, interpret, and respond to intricate physiological and pathological cues. Such developments will support the realization of fully autonomous sense–analyze–respond closed-loop platforms.
Future iterations of these systems are expected to be tightly integrated with wearable devices, ingestible or implantable diagnostics, and cloud-connected healthcare infrastructures, giving rise to intelligent “living-data” networks. These networks will enable dynamic, continuous, and personalized monitoring of chronic diseases, environmental stressors, and health trajectories. Coordinated bacterial consortia, supported by AI-driven control architectures, may also facilitate distributed sensing and adaptive therapeutic interventions across tissue compartments or ecological niches.
As the field transitions from conceptual demonstrations to translational development, challenges remain—including biosafety, genetic stability, standardization of production pipelines, and alignment with regulatory frameworks. However, sustained advances in chassis design, synthetic circuit optimization, and biocontainment strategies, coupled with increased cross-disciplinary collaboration, are accelerating the path toward clinical and industrial deployment. Ultimately, engineered bacterial diagnostics are poised to become a cornerstone of next-generation biotechnologies—bridging synthetic biology, smart medicine, and digital health. Their broad applicability across environmental monitoring, food safety, agriculture, and precision healthcare holds promise for significant societal and economic impact, ushering in a new era of intelligent, responsive, and personalized biosensing.
Author contributions
Decheng Wu: writing – review & editing, supervision. Hongmei Liu: writing – review & editing, supervision. Qiuke Jing: investigation, writing – original draft.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This work was supported by National Key R&D Program of China, the Ministry of Science and Technology of China (2020YFA0908900), Shenzhen Science and Technology Program (JCYJ20220530114409020 and 20231115134555001), Guangdong Basic and Applied Basic Research Foundation (2025A1515012076) and Guangdong Provincial Key Laboratory of Advanced Biomaterials (2022B1212010003).References
- Y. Li, et al., Advances in environmental pollutant detection techniques: Enhancing public health monitoring and risk assessment, Environ. Int., 2025, 197, 109365, DOI:10.1016/j.envint.2025.109365.
- J. Chang, G. Zhou, E. R. Christensen, R. Heideman and J. Chen, Graphene-based sensors for detection of heavy metals in water: a review, Anal. Bioanal. Chem., 2014, 406, 3957–3975, DOI:10.1007/s00216-014-7804-x.
- P. Kumar, D. K. Mahato, M. Kamle, T. K. Mohanta and S. G. Kang, Aflatoxins: A global concern for food safety, human health and their management, Front. Microbiol., 2016, 7, 2170, DOI:10.3389/fmicb.2016.02170.
- M. B. Amrofell, A. G. Rottinghaus and T. S. Moon, Engineering microbial diagnostics and therapeutics with smart control, Curr. Opin. Biotechnol., 2020, 66, 11–17, DOI:10.1016/j.copbio.2020.05.006.
- R. M. Cooper, et al., Engineered bacteria detect tumor DNA, Science, 2023, 381, 682–686, DOI:10.1126/science.adf3974.
- I. Tanniche and B. Behkam, Engineered live bacteria as disease detection and diagnosis tools, J. Biol. Eng., 2023, 17, 65, DOI:10.1186/s13036-023-00379-z.
- K. Jin, Y. Huang, H. L. Che and Y. H. Wu, Engineered bacteria for disease diagnosis and treatment using synthetic biology, Microb. Biotechnol., 2025, 18, 1, DOI:10.1111/1751-7915.70080.
- X. Tan, J. H. Letendre, J. J. Collins and W. W. Wong, Synthetic biology in the clinic: engineering vaccines, diagnostics, and therapeutics, Cell, 2021, 184, 881–898, DOI:10.1016/j.cell.2021.01.017.
- P. H. Tsou, et al., Exploring volatile organic compounds in breath for high-accuracy prediction of lung cancer, Cancers, 2021, 13, 1431, DOI:10.3390/cancers13061431.
- I. Del Valle, et al., Translating new synthetic biology advances for biosensingeinto the earth and environmental sciences, Front. Microbiol., 2020, 11, 618373, DOI:10.3389/fmicb.2020.618373.
- S. A. Ali, D. Mittal and G. Kaur, In-situ monitoring of xenobiotics using genetically engineered whole-cell-based microbial biosensors: recent advances and outlook, World J. Microbiol. Biotechnol., 2021, 37, 81, DOI:10.1007/s11274-021-03024-3.
- T. J. Ford and P. A. Silver, Synthetic biology expands chemical control of microorganisms, Curr. Opin. Chem. Biol., 2015, 28, 20–28, DOI:10.1016/j.cbpa.2015.05.012.
- J. Zhou, J. Dong, H. Hou, L. Huang and J. Li, High-throughput microfluidic systems accelerated by artificial intelligence for biomedical applications, Lab Chip, 2024, 24, 1307–1326, 10.1039/d3lc01012k.
- Y. P. Shelke, A. K. Badge and N. J. Bankar, Applications of artificial intelligence in microbial diagnosis, Cureus, 2023, 15, e49366, DOI:10.7759/cureus.49366.
- S. Fooladi, N. Rabiee and S. Iravani, Genetically engineered bacteria: a new frontier in targeted drug delivery, J. Mater. Chem. B, 2023, 11, 10072–10087, 10.1039/d3tb01805a.
- R. Xie, et al., Living therapeutics: Precision diagnosis and therapy with engineered bacteria, Biomaterials, 2025, 321, 123342, DOI:10.1016/j.biomaterials.2025.123342.
- S. Sridhar, C. M. Ajo-Franklin and C. A. Masiello, A Framework for the systematic selection of biosensor chassis for environmental synthetic biology, ACS Synth. Biol., 2022, 11, 2909–2916, DOI:10.1021/acssynbio.2c00079.
- F. Wu, Y. Liang, Y. Zhang, Y. Huo and Q. Wang, Construction of seamless genome editing system for fast-growing vibrio natriegens, Shengwu Gongcheng Xuebao, 2020, 36, 2387–2397, DOI:10.13345/j.cjb.200157.
- J. Xu, S. Yang and L. Yang, Vibrio natriegens as a host for rapid biotechnology, Trends Biotechnol., 2022, 40, 381–384, DOI:10.1016/j.tibtech.2021.10.007.
- M. T. Weinstock, E. D. Hesek, C. M. Wilson and D. G. Gibson, Vibrio natriegens as a fast-growing host for molecular biology, Nat. Methods, 2016, 13, 849–851, DOI:10.1038/nmeth.3970.
- M. Fan, S. Tan, W. Wang and X. Zhang, Improvement in salt tolerance ability of pseudomonas putida KT2440, Biology, 2024, 13(6), 404, DOI:10.3390/biology13060404.
- E. T. Mohamed, et al., Adaptive laboratory evolution of pseudomonas putida KT2440 improves p-coumaric and ferulic acid catabolism and tolerance, Metab. Eng. Commun., 2020, 11, e00143, DOI:10.1016/j.mec.2020.e00143.
- Y. T. Liu, J. Feng, H. C. Pan, X. W. Zhang and Y. L. Zhang, Genetically engineered bacterium: Principles, practices, and prospects, Front. Microbiol., 2022, 13, 997587, DOI:10.3389/fmicb.2022.997587.
- C. Beimfohr, A Review of research Conducted with probiotic E. coli marketed as symbioflor, Int. J. Bacteriol., 2016, 2016, 3535621, DOI:10.1155/2016/3535621.
- Y. Lai, N. Hayashi and T. K. Lu, Engineering the human gut commensal Bacteroides thetaiotaomicron with synthetic biology, Curr. Opin. Chem. Biol., 2022, 70, 102178, DOI:10.1016/j.cbpa.2022.102178.
- C. A. Miller, J. M. L. Ho and M. R. Bennett, Strategies for improving small-molecule biosensors in bacteria, Biosensors, 2022, 12, 64, DOI:10.3390/bios12020064.
- D. Roncarati, V. Scarlato and A. Vannini, Targeting of regulators as a promising approach in the search for novel antimicrobial agents, Microorganisms, 2022, 10, 185, DOI:10.3390/microorganisms10010185.
- C. Mejía-Almonte, et al., Redefining fundamental concepts of transcription initiation in bacteria, Nat. Rev. Genet., 2020, 21, 699–714, DOI:10.1038/s41576-020-0254-8.
- D. Xiao, et al., A D,L-lactate biosensor based on allosteric transcription factor LldR and amplified luminescent proximity homogeneous assay, Biosens. Bioelectron., 2022, 211, 114378, DOI:10.1016/j.bios.2022.114378.
- M. Mimee, et al., An ingestible bacterial-electronic system to monitor gastrointestinal health, Science, 2018, 360, 915–918, DOI:10.1126/science.aas9315.
- W. J. Wang, et al., A brief review of aptamer-based biosensors in recent years, Biosensors, 2025, 15, 120, DOI:10.3390/bios15020120.
- T. Mo, et al., Aptamer-based biosensors and application in tumor theranostics, Cancer Sci., 2022, 113, 7–16, DOI:10.1111/cas.15194.
- M. McKeague, R. S. Wong and C. D. Smolke, Opportunities in the design and application of RNA for gene expression control, Nucleic Acids Res., 2016, 44, 2987–2999, DOI:10.1093/nar/gkw151.
- C. Berens, F. Groher and B. Suess, RNA aptamers as genetic control devices: the potential of riboswitches as synthetic elements for regulating gene expression, Biotechnol. J., 2015, 10, 246–257, DOI:10.1002/biot.201300498.
- R. R. Breaker, Riboswitches and the RNA world, Cold Spring Harbor Perspect. Biol., 2012, 4(2), a003566, DOI:10.1101/cshperspect.a003566.
- Y. F. Wu, et al., High content design of riboswitch biosensors: All-around rational module-by-module design, Biosens. Bioelectron., 2023, 220, 114887, DOI:10.1016/j.bios.2022.114887.
- R. B. Bourret and R. E. Silversmith, Two-component signal transduction, Curr. Opin. Microbiol., 2010, 13, 113–115, DOI:10.1016/j.mib.2010.02.003.
- J. T. Lazar and J. J. Tabor, Bacterial two-component systems as sensors for synthetic biology applications, Curr. Opin. Syst. Biol., 2021, 28, 100398, DOI:10.1016/j.coisb.2021.100398.
- S. Ravikumar, M. G. Baylon, S. J. Park and J. I. Choi, Engineered microbial biosensors based on bacterial two-component systems as synthetic biotechnology platforms in bioremediation and biorefinery, Microb. Cell Fact., 2017, 16, 62, DOI:10.1186/s12934-017-0675-z.
- M. T. Laub and M. Goulian, Specificity in two-component signal transduction pathways, Annu. Rev. Genet., 2007, 41, 121–145, DOI:10.1146/annurev.genet.41.042007.170548.
- E. A. Groisman, A. Duprey and J. Choi, How the PhoP/PhoQ system controls virulence and Mg(2+) homeostasis: lessons in signal transduction, pathogenesis, physiology, and evolution, Microbiol. Mol. Biol. Rev., 2021, 85, e0017620, DOI:10.1128/mmbr.00176-20.
- M. Lebovich, M. A. Lora, J. Gracia-David and L. B. Andrews, Genetic circuits for feedback control of gamma-aminobutyric acid biosynthesis in probiotic escherichia coli Nissle 1917, Metabolites, 2024, 14, 44, DOI:10.3390/metabo14010044.
- C. M. Robinson, N. E. Short and D. T. Riglar, Achieving spatially precise diagnosis and therapy in the mammalian gut using synthetic microbial gene circuits, Front. Bioeng. Biotechnol., 2022, 10, 959441, DOI:10.3389/fbioe.2022.959441.
- X. Y. Wan, et al., Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals, Nat. Chem. Biol., 2019, 15, 540–548, DOI:10.1038/s41589-019-0244-3.
- D. J. Sayut, Y. Niu and L. H. Sun, Construction and engineering of positive feedback loops, ACS Chem. Biol., 2006, 1, 692–696, DOI:10.1021/cb6004245.
- S. K. Kim, et al., A genetically encoded biosensor for monitoring isoprene production in engineered escherichia coli, ACS Synth. Biol., 2018, 7, 2379–2390, DOI:10.1021/acssynbio.8b00164.
- B. C. Stanton, et al., Genomic mining of prokaryotic repressors for orthogonal logic gates, Nat. Chem. Biol., 2014, 10, 99–105, DOI:10.1038/nchembio.1411.
- H. S. Kang, D. Park and J. Kim, Logical regulation of endogenous gene expression using programmable, multi-input processing CRISPR guide RNAs, Nucleic Acids Res., 2024, 52, 8595–8608, DOI:10.1093/nar/gkae549.
- S. J. Chen, X. L. Chen, H. F. Su, M. Z. Guo and H. L. Liu, Advances in synthetic-biology-based whole-cell biosensors: principles, genetic modules, and applications in food safety, Int. J. Mol. Sci., 2023, 24, 7989, DOI:10.3390/ijms24097989.
- D. T. Riglar and P. A. Silver, Engineering bacteria for diagnostic and therapeutic applications, Nat. Rev. Microbiol., 2018, 16, 214–225, DOI:10.1038/nrmicro.2017.172.
- T. H. T. Chau, D. H. A. Mai, D. N. Pham, H. T. Q. Le and E. Y. Lee, Developments of riboswitches and toehold switches for molecular detection-biosensing and molecular diagnostics, Int. J. Mol. Sci., 2020, 21, 3192, DOI:10.3390/ijms21093192.
- L. Liu, W. Huang and J. D. Huang, Synthetic circuits that process multiple light and chemical signal inputs, BMC Syst. Biol., 2017, 11, 5, DOI:10.1186/s12918-016-0384-y.
- M. E. Inda-Webb, et al., Sub-1.4 cm3 capsule for detecting labile inflammatory biomarkers in situ, Nature, 2023, 620, 386–392, DOI:10.1038/s41586-023-06369-x.
- Z. P. Zou, Y. Du, T. T. Fang, Y. Zhou and B. C. Ye, Biomarker-responsive engineered probiotic diagnoses, records, and ameliorates inflammatory bowel disease in mice, Cell Host Microbe, 2023, 31(2), 199-212.e5, DOI:10.1016/j.chom.2022.12.004.
- D. H. Juers, B. W. Matthews and R. E. Huber, LacZ β-galactosidase: structure and function of an enzyme of historical and molecular biological importance, Protein Sci., 2012, 21, 1792–1807, DOI:10.1002/pro.2165.
- Q. Z. Wang, et al., Developing clostridia as cell factories for short- and medium-chain ester production, Front. Bioeng. Biotechnol., 2021, 9, 661694, DOI:10.3389/fbioe.2021.661694.
- J. Zhu, B. Wang, Y. Zhang, T. Wei and T. Gao, Living electrochemical biosensing: Engineered electroactive bacteria for biosensor development and the emerging trends, Biosens. Bioelectron., 2023, 237, 115480, DOI:10.1016/j.bios.2023.115480.
- F. Simonte, G. Sturm, J. Gescher and K. Sturm-Richter, Extracellular electron transfer and biosensors, Adv. Biochem. Eng./Biotechnol., 2019, 167, 15–38, DOI:10.1007/10_2017_34.
- M. Mimee, et al., An ingestible bacterial-electronic system to monitor gastrointestinal health, Science, 2018, 360, 915–918, DOI:10.1126/science.aas9315.
- V. A. Pavlov, S. S. Chavan and K. J. Tracey, Bioelectronic medicine: From preclinical studies on the inflammatory reflex to new approaches in disease diagnosis and treatment, Cold Spring Harbor Perspect. Med., 2020, 10, a034140, DOI:10.1101/cshperspect.a034140.
- S. Wu, et al., metaProbiotics: a tool for mining probiotic from metagenomic binning data based on a language model, Briefings Bioinf., 2024, 25(2), bbae085, DOI:10.1093/bib/bbae085.
- M. B. Fiamenghi, et al., Machine learning and comparative genomics approaches for the discovery of xylose transporters in yeast, Biotechnol. Biofuels Bioprod., 2022, 15, 57, DOI:10.1186/s13068-022-02153-7.
- C. M. Bergamini, et al., Machine learning algorithms highlight tRNA information content and chargaff's second parity rule score as important features in discriminating probiotics from non-probiotics, Biology, 2022, 11(7), 1024, DOI:10.3390/biology11071024.
- P. Zhang, et al., Deep flanking sequence engineering for efficient promoter design using DeepSEED, Nat. Commun., 2023, 14, 6309, DOI:10.1038/s41467-023-41899-y.
- M. Oubounyt, Z. Louadi, H. Tayara and K. T. Chong, DeePromoter: Robust promoter predictor using deep learning, Front. Genet., 2019, 10, 286, DOI:10.3389/fgene.2019.00286.
- X. Lei, X. Wang, G. Chen, C. Liang, Q. Li, H. Jiang and W. Xiong, Combining diffusion and transformer models for enhanced promoter synthesis and strength prediction in deep learning, mSystems, 2025, 10, e0018325, DOI:10.1128/msystems.00183-25.
- J. G. Doench, et al., Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9, Nat. Biotechnol., 2016, 34, 184–191, DOI:10.1038/nbt.3437.
- A. G. Genc and L. J. McGuffin, Beyond AlphaFold2: The impact of AI for the further improvement of protein structure prediction, Methods Mol. Biol., 2025, 2867, 121–139, DOI:10.1007/978-1-0716-4196-5_7.
- G. W. Wei, Protein structure prediction beyond AlphaFold, Nat. Mach. Intell., 2019, 1, 336–337, DOI:10.1038/s42256-019-0086-4.
- A. Racovita and A. Jaramillo, Reinforcement learning in synthetic gene circuits, Biochem. Soc. Trans., 2020, 48, 1637–1643, DOI:10.1042/bst20200008.
- Y. Matsuo, et al., Deep learning, reinforcement learning, and world models, Neural Netw., 2022, 152, 267–275, DOI:10.1016/j.neunet.2022.03.037.
- Z. P. Zou, et al., Self-Adjusting engineered probiotic for targeted tumor colonization and local therapeutics delivery, Adv. Sci., 2025, e06486, DOI:10.1002/advs.202406486.
- Z. Liu, X. Zhang, D. Lei, B. Qiao and G. R. Zhao, Metabolic engineering of escherichia coli for de novo production of 3-phenylpropanol via retrobiosynthesis approach, Microb. Cell Fact., 2021, 20, 121, DOI:10.1186/s12934-021-01615-1.
- M. Sabzevari, S. Szedmak, M. Penttilä, P. Jouhten and J. Rousu, Strain design optimization using reinforcement learning, PLoS Comput. Biol., 2022, 18, e1010177, DOI:10.1371/journal.pcbi.1010177.
- Y. Cheng, et al., Machine learning for metabolic pathway optimization: A review, Comput. Struct. Biotechnol. J., 2023, 21, 2381–2393, DOI:10.1016/j.csbj.2023.03.045.
- K. Rai, Y. Wang, R. W. O'Connell, A. B. Patel and C. J. Bashor, Using machine learning to enhance and accelerate synthetic biology, Curr. Opin. Biomed. Eng., 2024, 31, 100553, DOI:10.1016/j.cobme.2024.100553.
- D. Zhang, F. Xu, F. Wang, L. Le and L. Pu, Synthetic biology and artificial intelligence in crop improvement, Plant Commun., 2025, 6, 101220, DOI:10.1016/j.xplc.2024.101220.
- M. P. McRae, K. S. Rajsri, T. M. Alcorn and J. T. McDevitt, Smart diagnostics: combining artificial intelligence and in vitro diagnostics, Sensors, 2022, 22, 6355, DOI:10.3390/s22176355.
- M. Mirbabaie, S. Stieglitz and N. R. J. Frick, Artificial intelligence in disease diagnostics: A critical review and classification on the current state of research guiding future direction, Health Technol., 2021, 11, 693–731, DOI:10.1007/s12553-021-00555-5.
- J. Jumper, et al., Highly accurate protein structure prediction with AlphaFold, Nature, 2021, 596, 583–589, DOI:10.1038/s41586-021-03819-2.
- M. Baek, et al., Accurate prediction of protein structures and interactions using a three-track neural network, Science, 2021, 373, 871–876, DOI:10.1126/science.abj8754.
- Z. Lin, et al., Evolutionary-scale prediction of atomic-level protein structure with a language model, Science, 2023, 379, 1123–1130, DOI:10.1126/science.ade2574.
- G. Chuai, et al., DeepCRISPR: optimized CRISPR guide RNA design by deep learning, Genome Biol., 2018, 19, 80, DOI:10.1186/s13059-018-1459-4.
- Y. Wang, et al., Synthetic promoter design in Escherichia coli based on a deep generative network, Nucleic Acids Res., 2020, 48, 6403–6412, DOI:10.1093/nar/gkaa325.
- J. A. Valeri, et al., Sequence-to-function deep learning frameworks for engineered riboregulators, Nat. Commun., 2020, 11, 5058, DOI:10.1038/s41467-020-18676-2.
- N. Hashemi Goradel, et al., Biosensors for the detection of environmental and urban pollutions, J. Cell. Biochem., 2018, 119, 207–212, DOI:10.1002/jcb.26030.
- N. Hashemi Goradel, et al., Biosensors for the detection of environmental and urban pollutions, J. Cell. Biochem., 2018, 119, 207–212, DOI:10.1002/jcb.26030.
- B. Wang, M. Barahona and M. Buck, A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals, Biosens. Bioelectron., 2013, 40, 368–376, DOI:10.1016/j.bios.2012.08.011.
- Q. Gui, T. Lawson, S. Shan, L. Yan and Y. Liu, The application of whole cell-based biosensors for use in environmental analysis and in medical diagnostics, Sensors, 2017, 17(7), 1623, DOI:10.3390/s17071623.
- A. Doshi, et al., Engineered bacterial swarm patterns as spatial records of environmental inputs, Nat. Chem. Biol., 2023, 19, 878–886, DOI:10.1038/s41589-023-01325-2.
- C. Liu, et al., Engineering whole-cell microbial biosensors: Design principles and applications in monitoring and treatment of heavy metals and organic pollutants, Biotechnol. Adv., 2022, 60, 108019, DOI:10.1016/j.biotechadv.2022.108019.
- H. Che, et al., Multi-targets detection via combination of multi-stimulus-response engineered bacteria and hydrogel-based separation platform, J. Hazard. Mater., 2024, 478, 135578, DOI:10.1016/j.jhazmat.2024.135578.
- C. Liu, et al., Engineering whole-cell microbial biosensors: Design principles and applications in monitoring and treatment of heavy metals and organic pollutants, Biotechnol. Adv., 2022, 60, 108019, DOI:10.1016/j.biotechadv.2022.108019.
- C. M. Reddy, et al., Composition and fate of gas and oil released to the water column during the Deepwater Horizon oil spill, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 20229–20234, DOI:10.1073/pnas.1101242108.
- B. Q. Huynh, et al., Public health impacts of an imminent Red Sea oil spill, Nat. Sustain., 2021, 4, 1084–1091, DOI:10.1038/s41893-021-00774-8.
- F. Dubocq, B. B. Bæringsdóttir, T. Wang and A. Kärrman, Comparison of extraction and clean-up methods for comprehensive screening of organic micropollutants in fish using gas chromatography coupled to high-resolution mass spectrometry, Chemosphere, 2022, 286, 131743, DOI:10.1016/j.chemosphere.2021.131743.
- N. Kolahchi, et al., Direct detection of phenol using a new bacterial strain-based conductometric biosensor, J. Environ. Chem. Eng., 2018, 6, 478–484, DOI:10.1016/j.jece.2017.12.023.
- C. I. L. Justino, A. C. Duarte and T. A. P. Rocha-Santos, Recent progress in riosensors for environmental monitoring: A review, Sensors, 2017, 17, 2918, DOI:10.3390/s17122918.
- T. T. Wang, et al., Enhanced assessment of water quality for both nitrate and nitrite using engineered E. coli with para-aminobenzoic acid biosynthesis, Anal. Chim. Acta, 2025, 1338, 343580, DOI:10.1016/j.aca.2024.343580.
- N. Verma and A. Bhardwaj, Biosensor technology for pesticides--a review, Appl. Biochem. Biotechnol., 2015, 175, 3093–3119, DOI:10.1007/s12010-015-1489-2.
- M. Kim, et al., Chemostat-like microfluidic platform for highly sensitive detection of heavy metal ions using microbial biosensors, Biosens. Bioelectron., 2015, 65, 257–264, DOI:10.1016/j.bios.2014.10.028.
- Q. Li, P. Wang, R. Chen, W. Li and Y. J. Wu, Construction of genetically engineered bacteria that degrades organophosphorus pesticide residues and can be easily detected by the fluorescence, Environ. Technol., 2014, 35, 556–561, DOI:10.1080/09593330.2013.837936.
- S. Parvez, C. Venkataraman and S. Mukherji, A review on advantages of implementing luminescence inhibition test (Vibrio fischeri) for acute toxicity prediction of chemicals, Environ. Int., 2006, 32, 265–268, DOI:10.1016/j.envint.2005.08.022.
- R. Ranjan, N. K. Rastogi and M. S. Thakur, Development of immobilized biophotonic beads consisting of Photobacterium leiognathi for the detection of heavy metals and pesticide, J. Hazard. Mater., 2012, 225–226, 114–123, DOI:10.1016/j.jhazmat.2012.04.076.
- J. Kumar and S. F. D'Souza, An optical microbial biosensor for detection of methyl parathion using Sphingomonas sp. immobilized on microplate as a reusable biocomponent, Biosens. Bioelectron., 2010, 26, 1292–1296, DOI:10.1016/j.bios.2010.07.016.
- A. Mishra, J. Kumar and J. S. Melo, An optical microplate biosensor for the detection of methyl parathion pesticide using a biohybrid of Sphingomonas sp. cells-silica nanoparticles, Biosens. Bioelectron., 2017, 87, 332–338, DOI:10.1016/j.bios.2016.08.048.
- A. Bhatt, S. Jain and N. K. Navani, Rapid, sensitive, and specific microbial whole-cell biosensor for the detection of histamine: A potential food toxin, J. Agric. Food Chem., 2024, 72, 27466–27478, DOI:10.1021/acs.jafc.4c06315.
- J. Ma, et al., Accurate and non-destructive monitoring of mold contamination in foodstuffs based on whole-cell biosensor array coupling with machine-learning prediction models, J. Hazard. Mater., 2023, 449, 131030, DOI:10.1016/j.jhazmat.2023.131030.
- J. Ma, et al., Accurate and non-destructive monitoring of mold contamination in foodstuffs based on whole-cell biosensor array coupling with machine-learning prediction models, J. Hazard. Mater., 2023, 449, 131030, DOI:10.1016/j.jhazmat.2023.131030.
- V. Felipe, et al., First report of Pectobacterium brasiliense causing blackleg and soft rot of potato in Argentina, Plant Dis., 2024 DOI:10.1094/pdis-03-24-0558-pdn.
- X. Ma, A. Schloop, B. Swingle and K. L. Perry, Pectobacterium and dickeya responsible for potato blackleg disease in new york state in 2016, Plant Dis., 2018, 102, 1834–1840, DOI:10.1094/pdis-10-17-1595-re.
- M. B. Suárez, et al., New pCR-based assay for the identification of pectobacterium carotovorum causing potato soft rot, Plant Dis., 2022, 106, 676–684, DOI:10.1094/pdis-08-21-1676-re.
- R. Czajkowski, et al., Detection, identification and differentiation of pectobacterium and dickeya species causing potato blackleg and tuber soft rot: a review, Ann. Appl. Biol., 2015, 166, 18–38, DOI:10.1111/aab.12166.
- Y. Fang and R. P. Ramasamy, Current and prospective methods for plant disease detection, Biosensors, 2015, 5, 537–561, DOI:10.3390/bios5030537.
- N. Zanger and E. Eltzov, Optimizing whole-cell biosensors for the early detection of crop infections: A proof-of-concept study, Biosensors, 2025, 15(5), 300, DOI:10.3390/bios15050300.
- M. Liu, W. Yang, W. Zhu and D. Yu, Innovative applications and research advances of bacterial biosensors in medicine, Front. Microbiol., 2025, 16, 1507491, DOI:10.3389/fmicb.2025.1507491.
- S. Tuljapurkar, Infectious diseases of humans—Dynamics and control, Science, 1991, 254, 591–592, DOI:10.1126/science.254.5031.591.
- R. Rafique, et al., Transmission of infectious vibrio cholerae through drinking water among the household contacts of cholera patients (CHoBI7 trial), Front. Microbiol., 2016, 7, 1635, DOI:10.3389/fmicb.2016.01635.
- Z. Zhou, et al., Engineering probiotics as living diagnostics and therapeutics for improving human health, Microb. Cell Fact., 2020, 19, 56, DOI:10.1186/s12934-020-01318-z.
- M. B. Holowko, H. J. Wang, P. Jayaraman and C. L. Poh, Biosensing vibrio cholerae with genetically engineered escherichia coli, ACS Synth. Biol., 2016, 5, 1275–1283, DOI:10.1021/acssynbio.6b00079.
- T. Danino, et al., Programmable probiotics for detection of cancer in urine, Sci. Transl. Med., 2015, 7, 289ra84, DOI:10.1126/scitranslmed.aaa3519.
- C. R. Gurbatri, et al., Engineering tumor-colonizing E. coli Nissle 1917 for detection and treatment of colorectal neoplasia, Nat. Commun., 2024, 15, 646, DOI:10.1038/s41467-024-44776-4.
- J. T. Panteli, B. A. Forkus, N. Van Dessel and N. S. Forbes, Genetically modified bacteria as a tool to detect microscopic solid tumor masses with triggered release of a recombinant biomarker, Integr. Biol., 2015, 7, 423–434, 10.1039/c5ib00047e.
- T. V. Plavec, et al., Targeting of fluorescent lactococcus lactis to colorectal cancer cells through surface display of tumour-antigen binding proteins, Microb. Biotechnol., 2021, 14, 2227–2240, DOI:10.1111/1751-7915.13907.
- Z. M. Geng, et al., Aptamer-assisted tumor localization of bacteria for enhanced biotherapy, Nat. Commun., 2021, 12, 6584, DOI:10.1038/s41467-021-26956-8.
- S. W. de Melo and J. A. Di Palma, The role of capsule endoscopy in evaluating inflammatory bowel disease, Gastroenterol. Clin. N. Am., 2012, 41, 315–323, DOI:10.1016/j.gtc.2012.01.005.
- Z. P. Zou, et al., Genetically engineered bacteria as inflammatory bowel disease therapeutics, Eng. Microbiol., 2024, 4, 100167, DOI:10.1016/j.engmic.2024.100167.
- D. T. Riglar, et al., Engineered bacteria can function in the mammalian gut long-term as live diagnostics of inflammation, Nat. Biotechnol., 2017, 35, 653–658, DOI:10.1038/nbt.3879.
- Z. P. Zou, Y. Du, T. T. Fang, Y. Zhou and B. C. Ye, Biomarker-responsive engineered probiotic diagnoses, records, and ameliorates inflammatory bowel disease in mice, Cell Host Microbe, 2023, 31, 199–211, DOI:10.1016/j.chom.2022.12.004.
- X. Liu, et al., Magnetic living hydrogels for intestinal localization, retention, and diagnosis, Adv. Funct. Mater., 2021, 31, 2010918, DOI:10.1002/adfm.202010918.
- A. Courbet, D. Endy, E. Renard, F. Molina and J. Bonnet, Detection of pathological biomarkers in human clinical samples via amplifying genetic switches and logic gates, Sci. Transl. Med., 2015, 7, 289ra283, DOI:10.1126/scitranslmed.aaa3601.
- F. J. van Spronsen, et al., Phenylketonuria, Nat. Rev. Dis. Primers, 2021, 7, 36, DOI:10.1038/s41572-021-00267-0.
- P. Dongiovanni, et al., Salivary biomarkers: novel noninvasive tools to diagnose chronic inflammation, Int. J. Oral Sci., 2023, 15, 27, DOI:10.1038/s41368-023-00231-6.
- J. Shi, et al., Active biointegrated living electronics for managing inflammation, Science, 2024, 384, 1023–1030, DOI:10.1126/science.adl1102.
- J. L. Cummings, et al., Biomarker-guided decision making in clinical drug development for neurodegenerative disorders, Nat. Rev. Drug Discovery, 2025, 24(8), 589–609, DOI:10.1038/s41573-025-01165-w.
- N. Kolahchi, M. Braiek, G. Ebrahimipour, S. O. Ranaei-Siadat, F. Lagarde and N. Jaffrezic-Renault, Direct detection of phenol using a new bacterial strain-based conductometric biosensor, J. Environ. Chem. Eng., 2018, 6, 478–484, DOI:10.1016/j.jece.2017.12.023.
- D. Zhu, et al., Microbial biosensor for sensing and treatment of intestinal inflammation, Adv. Sci., 2025, 12, e2504364, DOI:10.1002/advs.202504364.
- M. T. Buss, L. Zhu, J. H. Kwon, J. J. Tabor and M. G. Shapiro, Probiotic acoustic biosensors for noninvasive imaging of gut inflammation, Nat. Commun., 2025, 16, 7931, DOI:10.1038/s41467-025-62569-1.
- R. J. S. Banicod, et al., Integration of Artificial Intelligence in Biosensors for Enhanced Detection of Foodborne Pathogens, Biosensors, 2025, 15(10), 690, DOI:10.3390/bios15100690.
- C. E. Lawson, et al., Machine learning for metabolic engineering: A review, Metab. Eng., 2021, 63, 34–60, DOI:10.1016/j.ymben.2020.10.005.
- A. Coutant, et al., Closed-loop cycles of experiment design, execution, and learning accelerate systems biology model development in yeast, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 18142–18147, DOI:10.1073/pnas.1900548116.
- J. Li, H. Zhao, L. Zheng and W. An, Advances in synthetic biology and biosafety governance, Front. Bioeng. Biotechnol., 2021, 9, 598087, DOI:10.3389/fbioe.2021.598087.
- A. Lerner, C. Benzvi and A. Vojdani, The potential harmful effects of genetically engineered microorganisms (GEMs) on the intestinal microbiome and public health, Microorganisms, 2024, 12, 238, DOI:10.3390/microorganisms12020238.
- C. M. Whitford, et al., Auxotrophy to xeno-DNA: an exploration of combinatorial mechanisms for a high-fidelity biosafety system for synthetic biology applications, J. Biol. Eng., 2018, 12, 13, DOI:10.1186/s13036-018-0105-8.
- D. R. George, et al., A bumpy road ahead for genetic biocontainment, Nat. Commun., 2024, 15, 650, DOI:10.1038/s41467-023-44531-1.
- C. T. Chan, J. W. Lee, D. E. Cameron, C. J. Bashor and J. J. Collins, 'Deadman' and 'Passcode' microbial kill switches for bacterial containment, Nat. Chem. Biol., 2016, 12, 82–86, DOI:10.1038/nchembio.1979.
- F. Stirling, et al., Rational design of evolutionarily stable microbial kill switches, Mol. Cell, 2017, 68, 686–697.e3, DOI:10.1016/j.molcel.2017.10.033.
- F. Du, et al., Optimizing multicopy chromosomal integration for stable high-performing strains, Nat. Chem. Biol., 2024, 20, 1670–1679, DOI:10.1038/s41589-024-01650-0.
- A. Chen, et al., Application of plasmid stabilization systems for heterologous protein expression in Escherichia coli, Mol. Biol. Rep., 2024, 51, 939, DOI:10.1007/s11033-024-09881-z.
- A. Egli, J. Schrenzel and G. Greub, Digital microbiology, Clin. Microbiol. Infect., 2020, 26, 1324–1331, DOI:10.1016/j.cmi.2020.06.023.
| This journal is © The Royal Society of Chemistry 2026 |