Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Fully protected pyrophosphates via phosphorobromidates for the synthesis of biopolymers
Sven
Wijngaarden
,
Femke L. A. M.
van der Heijden
,
Koen J.
Rijpkema
,
Bob
van Puffelen
,
Nico J.
Meeuwenoord
,
Danicia
Ramcharan
,
Gijsbert A.
van der Marel
,
Jeroen D. C.
Codée
 ,
Herman S.
Overkleeft
,
Herman S.
Overkleeft
 and
Dmitri V.
Filippov
and
Dmitri V.
Filippov
 *
*
Leiden Institute of Chemistry, Leiden University, Einsteinweg 55, 2333 CC, Leiden, The Netherlands. E-mail: filippov@chem.leidenuniv.nl
First published on 9th March 2026
Abstract
Natural oligomers containing phosphodiester linkages, such as DNA and RNA oligomers, are readily accessible through the phosphoramidite P(III) approach. However, the repetitive introduction of pyrophosphates into an oligomeric backbone, as found in poly-ADP-ribose, remains challenging. The most effective method to date relies on P(III)–P(V) coupling, in which a phosphate monoester is linked with a phosphoramidite, facilitating chain elongation of the biopolymer following the oxidation of the P(III)–P(V) intermediate. This anionic and nucleophilic intermediate limits the chain length that can be achieved. To overcome this limitation, we designed a methodology based on P(V)–P(V) chemistry, where a phosphodiester is coupled to a protected and activated P(V) donor. To this end, the Atherton–Todd oxidation of H-phosphonates is utilised to generate an electrophilic halophosphate, which is then coupled with a phosphodiester to yield a fully protected pyrophosphate species. The chemistry was optimised to balance the reactivity and stability of the coupling partners and the product by introducing the o-chloro-p-nitrophenyl ethyl protecting group for the phosphodiester and pyrophosphates. The full protection of the pyrophosphate moiety during oligomer synthesis eliminates the issues associated with the nucleophilicity and charge of the deprotected pyrophosphate and enables the solid-phase synthesis of pyrophosphate-based ADP-ribose oligomers of unprecedented length.
Introduction
To date, phosphodiester-based biopolymers such as DNA and RNA are readily available due to advancements in oligonucleotide chemistry over the past few decades.1 The chemical synthesis of oligonucleotides has provided powerful molecular tools, such as guide strands for CRISPR/Cas9 gene editing2 and has enabled many new technologies like gene synthesis3 and antisense mRNA suppression.4 The use of phosphoramidites, which are P(III) reagents, form the basis of the remarkably efficient synthetic methods employed in DNA and RNA fragment synthesis.Adenosine-di-phosphate ribose (ADPr) is a polymer composed of ribosyl adenosines linked through pyrophosphate bonds, and is installed as a post-translational modification on many proteins to fulfill a myriad of biological functions.5–9 Well-defined ADPr fragments and synthetic ADPr peptides/proteins have been instrumental in unravelling ADPr biology.10,11 However, the generation of longer ADPr chains has been a major challenge.12,13 Pyrophosphates are intrinsically labile, as the P(V)–O–P(V) linkage is very susceptible to cleavage. As a result, the synthesis of pyrophosphate-containing biomolecules is far less developed as compared to their phosphodiester-based counterparts. The longest ADPr chain synthesized to date, was assembled using a solid-phase synthesis strategy in which resin-bound phosphate monoesters were reacted with a P(III) phosphoramidite.14 This P(III)–P(V) methodology has enabled the synthesis of various biomolecules and their derivatives as well as an oligo-ADPr fragment of five repeating units in length.15–18 However, the latter synthesis produced substantial amounts of truncated fragments, even when excess coupling reagents and multiple coupling cycles were employed. These shorter fragments likely originate from undesired reactivity associated with the unprotected pyrophosphate during chain elongation.19–21 This reduction in synthesis quality for longer ADPr oligomers, together with the nucleophilic and anionic character of the unprotected pyrophosphate during chain elongation, motivated the development of an alternative to the P(III)–P(V) coupling strategy.
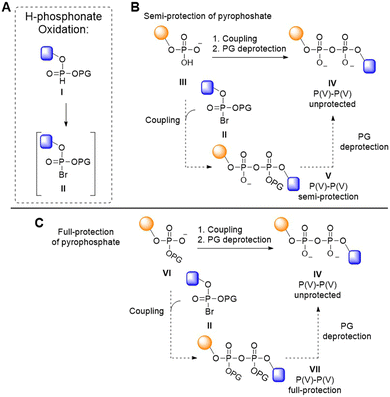
Pyrophosphates can be introduced through the coupling of a phosphomonoester with an activated P(V) species, such as a halophosphate (Fig. 1). Halophosphates are readily obtained by the Atherton–Todd oxidation of the corresponding H-phosphonate precursors (Fig. 1A).22 They are commonly used in situ, while they are generally too unstable for standard isolation procedures.23 The Atherton–Todd reaction entails oxidation of an H-phosphonate (I) by reaction with a halogenating agent in the presence of a base to provide the electrophilic halogenated P(V) species (II).24 This reaction was originally developed to produce phosphoramides, but changing the nucleophilic reaction partner from an amine to a phosphate has delivered methodology to construct diphosphate bonds.22 Using this approach, the group of Meier has created di- and triphosphate linkages.25–27 Likewise, Hergenrother and colleagues employed an in situ generated halophosphate (II) to couple a phosphomonoester (III) in their solution-phase synthesis of di-ADPr, yielding a partially protected pyrophosphotriester (V) that, upon deprotection, furnished the desired diphosphate (Fig. 1B, IV).28,29 These partially protected intermediates are difficult to handle in solution, and no method based on this approach has been able to access constructs longer than a dimer.
We reasoned that full protection of the pyrophosphate species as a tetraester during the chain elongation would negate its nucleophilicity (Fig. 1C, VII), which could potentially benefit the synthesis of longer oligo-ADPr and related biopolymers, in line with contemporary oligonucleotide synthesis, in which the phosphates are protected as phosphotriesters. Because phosphodiesters do not react productively with phosphoramidites30 we set out to develop a P(V)–P(V) method for repetitive pyrophosphate formation where the coupling proceeds via attack of a phosphate diester (VI) on an electrophilic P(V) phosphorohalidate (II), obtained via an Atherton–Todd (AT) reaction (Fig. 1).22 This approach requires careful handling of the anhydride-type intermediates formed during chain elongation, as these species are highly susceptible to nucleophilic substitution leading to monophosphate byproducts instead of the desired pyrophosphate.31–34 Consequently, stringent demands are placed on the protecting groups used to mask both the phosphate that is to be elongated as well as the intermediate pyrophosphate that is formed. Here, we describe the successful development of such an approach, which was enabled by the use of the little-used o-chloro-p-nitrophenyl ethyl protecting group35 for the phosphodiester and pyrophosphotetraester. This protecting group strikes the right balance between pyrophosphate stability, ease of halophosphate formation and removal at the end of the synthesis. We demonstrate the applicability of the methodology in the solid phase assembly of ADPr oligomers up to the decamer level, representing the longest well-defined ADPr chains synthesized to date.
Results and discussion
Development of P(V)–P(V) method using oligo-TDP as the model system
To develop the P(V)–P(V) pyrophosphate coupling methodology and protecting group strategy, we first explored the usability of halophosphates (II, Fig. 1) for the synthesis of pyrophosphate-linked oligonucleotides (oligo-TDP), that can be assembled from relatively simple H-phosphonates.Four 4,4-di-methoxytrityl (DMTr) protected thymidine H-phosphonates were synthesized, each featuring a distinct protecting group on the H-phosphonate functionality: an acid-labile tert-butyl (tBu) ester or one of the three increasingly base-labile esters para-nitrophenylethyl (NPE), 2-chloro-4-nitrophenylethyl (ClNPE) or 2-cyanoethyl (CNE) (1–4, Fig. 2). Base labile pyrophosphate protecting groups that can be cleaved via an elimination reaction have previously been shown to be effective in the P(III)–P(V) methodology.14 The NPE- and ClNPE groups, which can be cleaved by a strong base such as 1,8-diazabicyclo(5.4.0)undec-7-ene (DBU),35,36 were envisioned to be compatible with triethylamine (TEA), which is the base commonly used for the Atherton–Todd reaction. While CNE-protected phosphates are susceptible to cleavage by TEA, the milder base N-methylmorpholine (NMM) was employed instead.37
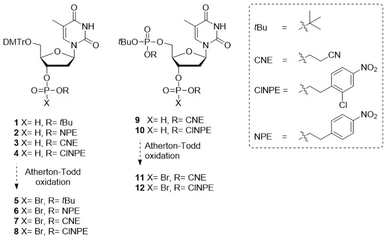 | ||
| Fig. 2 H-phosphonate building blocks required for solid-phase synthesis of oligo-TDP fragments via the P(V)–P(V) strategy. | ||
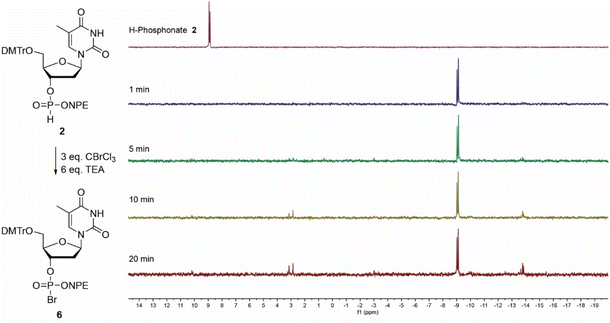
First, H-phosphonates 1–4 were assessed for their reactivity in the Atherton–Todd reaction using trichlorobromomethane (CBrCl3) as the halogenating agent, generating the phosphorobromidates (5–8), the formation of which was monitored by 31P-NMR.38 We explored two different bases in the reaction: TEA (pKaH 10.8) and the milder base NMM (pKaH 7.4).
The oxidation of NPE-protected H-phosphonate 2 (9 ppm) into phosphorobromidate 6 (−9 ppm) using TEA as base, proceeded uneventfully and the conversion was finished within one minute (Fig. 3). After twenty minutes, some hydrolysis to the phosphodiester (3 ppm) and formation of the symmetrical pyrophosphate (−13 ppm) could be detected.39 Oxidation of the more base labile ClNPE-protected H-phosphonate 4 was explored using either NMM or TEA as base. In an experiment with NMM no conversion to halogenated phosphate 8 (−9 ppm) could be detected after 2 minutes and the symmetrical pyrophosphotetraester (−14 ppm) was observed as the sole product after twenty minutes (Fig. S1). In contrast, the Atherton–Todd oxidation using TEA showed rapid conversion (<1 minute) into the bromidate 8 (−9 ppm), which was fully hydrolyzed and converted into the symmetrical pyrophosphate (−14 ppm) within three minutes (Fig. S2). No cleavage of the ClNPE-group could be detected under these conditions. The NMM-assisted oxidation of CNE-protected H-phosphonate 3 was significantly slower and complete conversion into the corresponding phosphorobromidate 7 was observed after approximately twenty minutes (Fig. S3). Here, slow hydrolysis, accompanied by the formation of the protected symmetrical pyrophosphate (−14 ppm) was also observed. The conversion of the tBu-protected H-phosphonate 1 into the corresponding phosphorobromidate 5 (−15 ppm) proceeded the slowest and took approximately 30 minutes (Fig. S4).
The halophosphates (5–8) featuring the four different protective groups were next evaluated as electrophilic agents in the preparation of fully protected pyrophosphates. To this end, the solid-phase syntheses of oligo-TDP dimer 21 and trimer 26 was undertaken (Table 1). After the manual construction of the pyrophosphate linkages on resin, they were deprotected, the product was cleaved from the solid support and the coupling quality assessed via LCMS analysis.
| # | Pyrophosphate protecting group R | Product | Base | Halogenating agent | Coupling time | R cleavage conditions | Notes | Yieldc |
|---|---|---|---|---|---|---|---|---|
| a The scale of test syntheses was 2 µmol. General conditions during coupling reactions: 3 eq. of H-phosphonate 1–4, 6 eq. CBrCl3 and 9 eq. of base. The concentration of immobilized phosphodiester during coupling was 11 mM. tBu was deprotected using 2 cycles of 10% v/v TFA in DCM. After tBu deprotection, a washing step with the base was performed. Protecting groups depicted in blue are removed under basic conditions and those in red under acidic conditions. Cleavage from the solid support depicted in green was achieved under alkaline conditions. Solid-phase syntheses towards oligo-TDP compounds 21 and 26 and optimization of the protecting group strategy. b Double coupling procedure. c Estimated yield based on integration of UV-signal at 268 nm in the LC-MS traces. | ||||||||
| 1 | tBu | Dimer 21 | TEA | CBrCl3 | 30 min | 20% TFA | — | 77% |
| 2 | NPE | Dimer 21 | TEA | CBrCl3 | 10 min | 15% DBU | Incomplete deprotection of NPE | 53% |
| 3 | CNE | Dimer 21 | NMM | CBrCl3 | 10 min | 5% DBU | — | 89% |
| 4 | CNE | Trimer 26 | NMM | CBrCl3 | 10 min | 5% DBU | 19% dimer 21c | 66% |
| 5 | ClNPE | Dimer 21 | NMM | CBrCl3 | 10 min | 20% DBU | — | 98% |
| 6 | ClNPE | Trimer 26 | NMM | CBrCl3 | 10 min | 20% DBU | 25% dimer 21c + other side products | 68% |
| 7 | ClNPE | Dimer 21 | TEA | CBrCl3 | 1 min | 20% DBU | — | 93% |
| 8 | ClNPE | Trimer 26 | TEA | CBrCl3 | 1 min | 20% DBU | 10% dimer 21c | 83% |
| 9 | ClNPE | Trimer 26 | TEA | NCS | 1 min | 20% DBU | 27% dimer 21c | 65% |
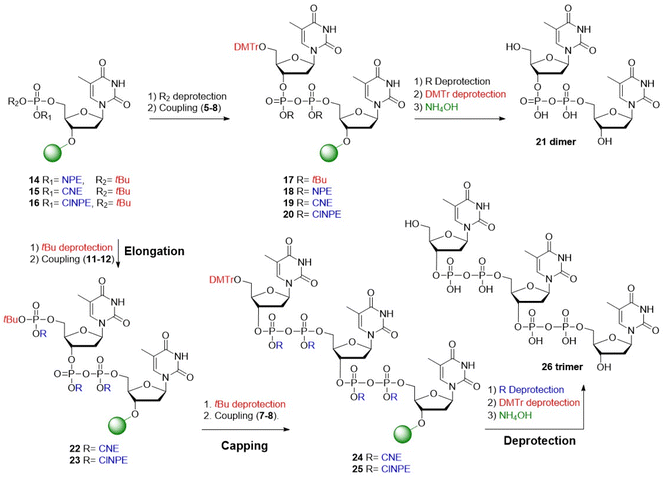
The first approach explored the use of base-labile temporary phosphate protection in combination with acid-labile “permanent” pyrophosphate protection. Solid-phase synthesis commenced with the preparation of immobilized phosphotriester 16. The corresponding resin-bound tBu-protected phosphodiester was obtained after cleavage of the ClNPE ester, which was condensed with tBu-protected P(V) bromidate 5 to yield the acid-sensitive pyrophosphotetraester 17. Subsequent acidolysis of the tBu pyrophosphate esters and cleavage from the solid support resulted in dimer 21 in suboptimal yield (Table 1, entry 1 and Fig. S5A).
The second approach employed acid-labile temporary protection together with base-labile “permanent” pyrophosphate protection. Acidolysis of the tBu ester in immobilized phosphotriester 14, was followed by a wash with base to provide the phosphodiester as an organic salt, enabling coupling to phosphorobromidate 6 using TEA as a base. Deprotection of the pyrophosphotetraester with DBU, followed by cleavage of the oligomer from the solid support afforded dimer 21 (Table 1, entry 2 and Fig. S5B). Unfortunately, LCMS analysis of the reaction mixture revealed incomplete removal of the NPE-ester in pyrophosphotetraester 18. Repetition of the NPE-cleavage procedure or increasing the concentration of DBU did not improve the yield of the synthesis, and the NPE protecting group was therefore not pursued further.
Next, the CNE-group was evaluated in the construction of dimer 21. Using the coupling cycle described above, the oligo-TDP dimer 21 was obtained in good purity (Table 1, entry 3 and Fig. S5C). Encouraged by these results, the synthesis of trimer 26 was undertaken. Cleavage of the tBu-ester in 15 generated the corresponding phosphodiester, which was coupled with phosphorobromidate 11 to afford the immobilized pyrophosphate 22, featuring a protected phosphate for the next elongation. Subsequent removal of the tBu ester in 22 and coupling with the chain terminating bromidate 7 yielded the fully protected trimer 24. Next, deprotection of the CNE-phosphate esters, removal of the primary DMT and cleavage of the oligomer from the solid support yielded trimer 26. LCMS analysis revealed formation of the trimer in 68%, but also substantial amount of dimer 21 (∼20%, Table 1, entry 4 and Fig. S5D).
Lastly, the ClNPE ester was investigated following the same strategy. Elimination of the ClNPE pyrophosphate esters using DBU and subsequent cleavage from the solid support afforded dimer 21, again in good purity with minimal side products (Table 1, entry 5 and Fig. S5E). Using these coupling and deprotection conditions, trimer 26 was produced as a mixture with dimer 21 along with other byproducts originating from incomplete elongation and termination (Table 1, entry 6 and Fig. S5F). Based on the Atherton–Todd activation studies described above, we hypothesized that switching the base from NMM to TEA would accelerate halophosphate formation and thereby improve the overall efficiency of the synthesis.
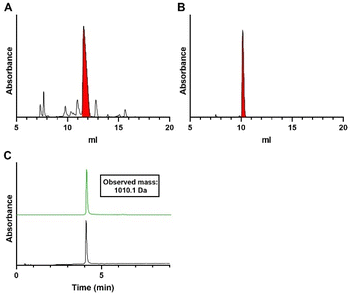
Indeed, the synthesis of dimer 21 using H-phosphonate 4, CBrCl3 and TEA for the coupling reaction, delivered 21 in excellent purity (Table 1, entry 7 and Fig. S5G). The ensuing solid-phase synthesis of trimer 26via sequential coupling of phosphorobromidates 12 and 8 delivered the target trimer 26 in 83% with significantly less dimer being formed (∼10%, Table 1, entry 8 and Fig. S5H), representing a significant improvement over the previous attempts. Substitution of CBrCl3 with the commonly used halogenating reagent N-chlorosuccinimide (NCS) did not enhance the outcome and instead gave a lower yield of trimer 26 (Table 1, entry 9 and Fig. S5I). Using the optimized conditions of Table 1, entry 8, a preparative synthesis of the trimer 26 was undertaken (Fig. 4A). The pure oligo-TDP trimer could be isolated by ion exchange chromatography (IEX), followed by size exclusion chromatography, in 30% yield (Fig. 4B and C).
Utilizing the P(V)–P(V) methodology for synthesis of ADPr-oligomers
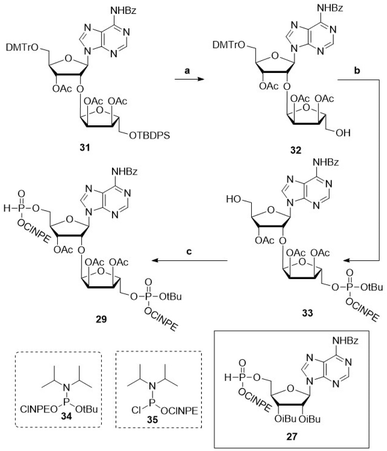
With conditions in hand for the solid-phase construction of the pyrophosphate linkages via fully protected pyrophosphates, we next set out to explore this chemistry for the synthesis of oligo-ADPr fragments. To this end, appropriately protected elongating and terminating H-phosphonate building blocks were designed based on the protecting group strategy developed above (27 and 29, Scheme 1). The central elongating building block 29 (Scheme 1) was prepared starting from orthogonally protected, ribosylated adenosine 31.15 Cleavage of the silyl ether using tetra-n-butylammoniumfluoride (TBAF), produced primary alcohol 32 in 80% yield, which was phosphitylated using the asymmetric tBu/ClNPE protected phosphoramidite 34 giving the phosphite triester. This P(III) species was oxidized by tBuOOH into the corresponding phosphotriester, after which acidolysis of the DMTr ether gave alcohol 33 in 90% yield over three steps. The H-phosphonate was then installed on the 5′-hydroxyl of compound 33via phosphitylation with chlorophosphoramidite 35 followed by pyridine·HCl mediated hydrolysis of the resulting phosphoramidite, to furnish the advanced phosphoryl H-phosphonate adenosine diriboside 29 in 74% yield over two steps.Atherton–Todd oxidation of the synthesized H-phosphonates showed comparable reaction kinetics as compared to the oxidation experiments described for oligo-TDP building block 4 (Fig. S6 and S7). A preparative solution-phase synthesis of mono-ADPr 66 afforded the product in 11% yield after purification, demonstrating the applicability of this approach in solution-phase settings (Scheme S7A). Attempts to isolate the corresponding pyrophosphotetraester 48 were unsuccessful, consistent with the pronounced lability of this species. Likewise, an attempted synthesis of dimer 42 proved abortive as no full-length product was detected (Scheme S7B).
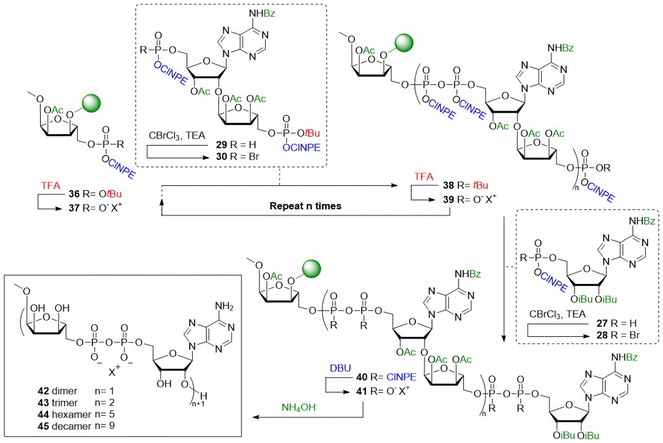
With the required building blocks in hand, the solid phase assembly of the ADPr oligomers was undertaken (Scheme 2). Starting from resin bound triester 36, the elongation cycle began with acidolysis of the tBu ester, followed by a wash with NMM to yield phosphodiester 37, which was then coupled with phosphorobromidate 30, generated in situ via the Atherton–Todd oxidation of H-phosphonate 29. This coupling procedure was repeated once to improve conversion, delivering immobilized pyrophosphotetraester 40 (n = 1). Repeating this cycle 2 times allowed for the synthesis of an oligo-ADPr trimer (38, n = 2). Termination of the oligomers involved removal of the tBu ester and coupling with phosphorobromidate 28, to produce the protected full-length ADPr-oligomers 40.
The pyrophosphates in the oligo-ADPr chain were then deprotected by first removal of the ClNPE esters using DBU, giving the pyrophosphodiesters 41, followed by global deprotection with concomitant cleavage from the solid support using aqueous ammonia to deliver the desired poly-ADPr fragments. The so-generated oligomers were analyzed by LCMS and purified by ion exchange chromatography (IEX) and size-exclusion chromatography. As shown in Fig. 5A the assembly of dimer proceeded uneventfully to deliver 42, which was isolated in a 27% yield based on resin loading. Analysis of the product mixture, generated in the synthesis of trimer 43, revealed the presence of significant amounts of dimer 42, which could originate from incomplete deprotection of the tBu phosphate ester. We therefore adapted the synthetic protocol by using an increased concentration of TFA for the elimination of the tBu ester, resulting in trimer 43 in a significantly improved yield (30% isolated yield, Fig. 5B).
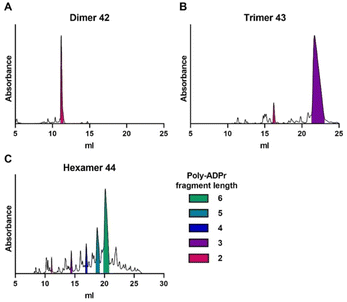 | ||
| Fig. 5 Analytical anion exchange chromatograms of crude poly-ADPr (A) dimer 42, (B) trimer 43 and (C) hexamer 44, obtained via the solid-phase synthetic strategy depicted in Scheme 2. The area under the curve of the individual poly-ADPr fragments are colored as depicted in the legend. UV absorbance is measured at 260 nm. | ||
With the successful preparation of trimer 43, the synthesis of fragments longer than a pentamer were attempted, which were previously inaccessible.16 The synthesis of ADP-ribose hexamer 44 proceeded well, delivering the target compound as the main product (Fig. 5C), although deletion sequences were also present in the mixture. Nonetheless, the hexamer could be effectively isolated to give the poly-ADPr fragment in 8.8% overall yield.
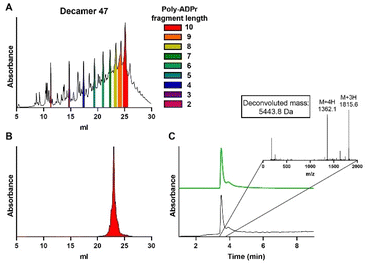
Finally, the synthesis of decamer 45 was undertaken. Repetition of the elongation cycle nine times, followed by the termination cycle, furnished ADPr decamer 45. Analysis of the crude reaction mixture (Fig. 6A) showed the successful synthesis of the target decamer 45. Isolation by ion exchange chromatography afforded decamer 45 in 4.8% yield (Fig. 6A and B), completing the synthesis of the longest ADPr oligomer to date.
During the synthesis of decamer 45, truncated fragments ranging from the nonamer to the dimer were detected. Several processes could account for these species, including incomplete coupling, insufficient removal of the temporary protecting group, or phosphoanhydride scission during chain elongation. Trace amounts of water during coupling can consume phosphorobromidate, leading to incomplete coupling. Indeed, performing the solid-phase synthesis without rigorously dried reagents resulted in no detectable full-length product (Fig. S8, top vs. middle). Incomplete deprotection of the tBu group could provide another possible mechanism for the observed deletion products. If the tBu group is not fully removed at the end of an elongating cycle, the resulting protected intermediate cannot undergo the subsequent coupling step. In the next cycle, however, this unreacted intermediate, now lacking the monomer that should have been added, can re-enter the coupling chemistry and propagate through the remaining cycles, ultimately generating the truncation products. This was already observed during an exploratory synthesis of hexamer 44, in which 10% TFA yielded small amounts of full-length product. Increasing the TFA concentration to 20% improved deprotection efficiency (Fig. S8, middle vs. bottom). Phosphoanhydride scission during synthesis is also conceivable, given the intrinsic sensitivity of the pyrophosphotetraester linkage. Although such fragments were less apparent in shorter oligomers, cumulative low-frequency side reactions over multiple coupling cycles could reasonably cause the pronounced truncation pattern seen in the decamer.
Conclusions
In this work, a solid-phase synthesis protocol to generate well-defined ADPr oligomers was developed, based on halophosphate P(V)–P(V) coupling methodology. The approach relies on the generation of fully protected pyrophosphotetraesters as intermediates during chain elongation, obtained by condensation of a phosphodiester with a phosphorobromidate, generated in situ via Atherton–Todd oxidation of the corresponding H-phosphonate. Crucial to the success of the approach has been the development of a protecting group scheme that hinges on the protection of the pyrophosphate with the ClNPE group, that balances the right stability and reactivity during the phosphorobromidate generation, the condensation reaction and can be cleaved under mild conditions that do not jeopardize the integrity of the labile (protected) pyrophosphate moieties. The presented solid-phase method successfully produced ADPr oligomers of unprecedented length, delivering the longest ADPr oligomer, assembled to date. The developed methodology is a valuable addition to the current toolbox for the synthesis of well defined poly-ADP-ribose chains and poly-ADPr proteins and peptides, as well as other biopolymers containing sugar-pyrophosphate chains. The generated ADPr oligomers will be valuable tools to probe interactions with ADPr processing enzymes and receptors.Author contributions
S. W.: investigation, methodology, conceptualisation, writing – original draft; F. L. A. M. v. d. H.: investigation, resources, writing – review & editing; K. J. R.: investigation, resources, writing – review & editing; B. v. P.: investigation, resources, writing – review & editing; N. J. M.: investigation, resources; D. R.: investigation, resources; G. A. v. d. M.: conceptualisation, writing – original draft, writing – review & editing; J. D. C. C.: supervision, writing – review & editing; H. S. O.: supervision, funding acquisition, writing – review & editing: D. V. F.: conceptualisation, supervision, funding acquisition, writing – original draft, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). The authors have cited two additional references within the SI.40,41 Supplementary information: figures, synthetic procedures and raw NMR spectra. See DOI: https://doi.org/10.1039/d6sc01119e.Acknowledgements
The authors thank the Dutch Research Council (NWO; grant number OCENW.KLEIN.497) for supporting the research presented in this paper.References
- C. B. Reese, Org. Biomol. Chem., 2005, 3, 3851 RSC
.
- Q. Chen, Y. Zhang and H. Yin, Adv. Drug Delivery Rev., 2021, 168, 246 CrossRef CAS PubMed
.
- M. J. Czar, J. C. Anderson, J. S. Bader and J. Peccoud, Trends Biotechnol., 2009, 27, 63 CrossRef CAS PubMed
.
- K. A. Lennox and M. A. Behlke, Gene Ther., 2011, 18, 1111 CrossRef CAS PubMed
.
- A. R. Fehr, S. A. Singh, C. M. Kerr, S. Mukai, H. Higashi and M. Aikawa, Genes Dev., 2020, 34, 341 CrossRef CAS PubMed
.
- M. J. Suskiewicz, E. Prokhorova, J. G. M. Rack and I. Ahel, Cell, 2023, 186, 4475 CrossRef CAS PubMed
.
- F. Dantzer and R. Santoro, FEBS J., 2013, 280, 3508 CrossRef CAS PubMed
.
- R. Gupte, Z. Liu and W. L. Kraus, Genes Dev., 2017, 31, 101 CrossRef CAS PubMed
.
- M. M. Rosado, E. Bennici, F. Novelli and C. Pioli, Immunology, 2013, 139, 428 Search PubMed
.
- A. K. L. Leung, Trends Cell Biol., 2020, 30, 370 Search PubMed
.
- H. Minnee, J. D. C. Codee and D. V. Filippov, ChemBioChem, 2024, 25, e202400440 Search PubMed
.
- Q. Liu, G. A. van der Marel and D. V. Filippov, Org. Biomol. Chem., 2019, 17, 5460 Search PubMed
.
- G. J. van der Heden van Noort, ACS Omega, 2020, 5, 1743 CrossRef CAS PubMed
.
- H. Gold, P. van Delft, N. Meeuwenoord, J. D. C. Codee, D. V. Filippov, G. Eggink, H. S. Overkleeft and G. A. van der Marel, J. Org. Chem., 2008, 73, 9458 CrossRef CAS PubMed
.
- H. A. V. Kistemaker, L. N. Lameijer, N. J. Meeuwenoord, H. S. Overkleeft, G. A. van der Marel and D. V. Filippov, Angew. Chem., Int. Ed., 2015, 54, 4915 Search PubMed
.
- Q. Liu, G. Knobloch, J. Voorneveld, N. J. Meeuwenoord, H. S. Overkleeft, G. A. van der Marel, A. G. Ladurner and D. V. Filippov, Chem. Sci., 2021, 12, 12468 RSC
.
- K. W. Kliza, Q. Liu, L. W. M. Roosenboom, P. W. T. C. Jansen, D. V. Filippov and M. Vermeulen, Mol. Cell, 2021, 81, 4552 CrossRef CAS PubMed
.
- H. A. V. Kistemaker, N. J. Meeuwenoord, H. S. Overkleeft, G. A. Van Der Marel and D. V. Filippov, Eur. J. Org Chem., 2015, 6084 CrossRef CAS
.
- G. Weimann and H. G. Khorana, J. Am. Chem. Soc., 1962, 84, 4329 CrossRef CAS
.
- Q. Sun, S. Liu, J. Sun, S. Gong, Q. Xiao and L. Shen, Tetrahedron Lett., 2013, 54, 3842 CrossRef CAS
.
- D. G. Knorre, A. V Lebedev, A. S. Levina, A. I. Rezvukhin and V. F. Zarytova, Tetrahedron, 1974, 30, 3073 CrossRef CAS
.
- F. R. Atherton, H. T. Openshaw and A. R. Todd, J. Chem. Soc., 1945, 660 RSC
.
- A. Kraszewski and J. Stawinski, Pure Appl. Chem., 2007, 79, 2217 CrossRef CAS
.
- S. S. Le Corre, M. Berchel, H. Couthon-Gourvès, J. P. Haelters and P. A. Jaffrès, Beilstein J. Org. Chem., 2014, 10, 1166 CrossRef PubMed
.
- N. Jeschik, T. Schneider and C. Meier, Eur. J. Org Chem., 2020, 6776 CrossRef CAS
.
- X. Jia, D. Schols and C. Meier, J. Med. Chem., 2023, 66, 12163 CrossRef CAS PubMed
.
- S. Weising, S. Weber, D. Schols and C. Meier, J. Med. Chem., 2022, 65, 12163 CrossRef CAS PubMed
.
- M. J. Lambrecht, M. Brichacek, E. Barkauskaite, A. Ariza, I. Ahel and P. J. Hergenrother, J. Am. Chem. Soc., 2015, 137, 3558 CrossRef CAS PubMed
.
- B. S. Drown, T. Shirai, J. G. M. Rack, I. Ahel and P. J. Hergenrother, Cell Chem. Biol., 2018, 25, 1562 CrossRef CAS PubMed
.
- A. P. Guzaev and M. Manoharan, J. Org. Chem., 2001, 66, 1798 CrossRef CAS PubMed
.
- E. Sonveaux, Bioorg. Chem., 1986, 14, 274 CrossRef CAS
.
- E. M. Ivanova, L. . M. Khalimskaya, V. P. Romanenko and V. F. Zarytova, Tetrahedron Lett., 1982, 23, 5447 CAS
.
- S. Chandrasegaran, A. Murakami and L. Kan, J. Org. Chem., 1984, 49, 4951 Search PubMed
.
- G. W. Kenner, A. R. Todd and F. J. Weymouth, J. Chem. Soc., 1952, 3675 RSC
.
- E. Uhlmann and W. Pfleiderer, Helv. Chim. Acta, 1981, 64, 1688 CrossRef CAS
.
- W. Pfleiderer, F. Himmelsbach, R. Charubala, H. Schirmeister, A. Beiter, B. Schulz and T. Trichtinger, Nucleosides Nucleotides, 1985, 4, 81 CrossRef CAS
.
- H. M. Hsiung, Tetrahedron Lett., 1982, 23, 5119 Search PubMed
.
- F. R. Atherton and A. R. Todd, J. Chem. Soc., 1947, 674 Search PubMed
.
- J. Nilsson, A. Kraszewski and J. Stawinski, J. Chem. Soc., Perkin Trans. 2, 2001, 2263 Search PubMed
.
- C. A. A. Claesen, R. P. A. M. Segers and G. I. Tesser, Recl. Trav. Chim. Pays-Bas, 1985, 104, 119–122 CrossRef CAS
.
- G. J. Van Der Heden Van Noort, H. S. Overkleeft, G. A. Van Der Marel and D. V. Filippov, J. Org. Chem., 2010, 75, 5733–5736 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |