Open Access Article
Open Access ArticleAmmonia monooxygenase: a work in progress
Thomas C.
Arndt
 ,
Alexander L.
Laughlin
and
Kyle M.
Lancaster
,
Alexander L.
Laughlin
and
Kyle M.
Lancaster
 *
*
Department of Chemistry and Chemical Biology Cornell University, Baker Laboratory, 162 Sciences Drive, Ithaca, NY 14853, USA. E-mail: kml236@cornell.edu
First published on 8th April 2026
Abstract
Ammonia monooxygenase (AMO) is a multimeric, multidomain integral membrane protein belonging to the copper membrane monooxygenase (CuMMO) family. It is responsible for the initiation of nitrification, the process by which certain bacteria and archaea oxidize ammonia (NH3) to nitrite (NO2−) as their primary metabolism. AMO is responsible for the selective conversion of NH3 to hydroxylamine (NH2OH) using the green oxidant dioxygen (O2). While the archaeal AMO and bacterial AMO both produce NH2OH, the structure and genetic underpinnings of this chemical behavior vary significantly. Further, there are many Cu-binding sites in AMO, and consequently there has been substantial discussion regarding the nature of the Cu site or sites responsible for substrate activation. Overall, AMO has largely eluded direct study because active protein has only recently been purified due to difficulties in cultivation of the native organism and recalcitrance of the protein to recombinant expression. This review is intended to serve as a primer to AMO, yielding the necessary context to understand its importance.
The nitrogen cycle, nitrification, and ammonia monooxygenase
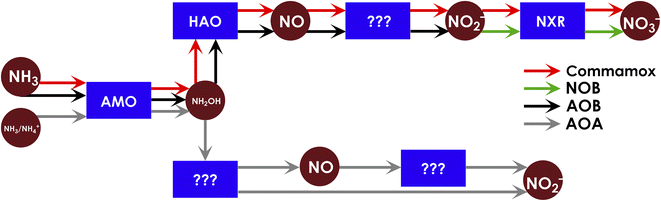
In every environment on Earth, there exist a multitude of organisms that are capable of fueling metabolism entirely upon inorganic nitrogen-based fuel.1 These microbes are responsible for the aerobic oxidation of ammonia (NH3) to nitrite (NO2−) and nitrate (NO3−), via primary metabolic reactions collectively termed nitrification (Fig. 1).2–4 Both ammonia oxidizing bacteria (AOB) and archaea (AOA) are capable of the oxidation of NH3 to NO2− (eqn (1)).| NH3 + O2 → NO2− + 3H+ + 2e− | (1) |
The final 2-electron oxidation to form NO3− is completed either by NO2− oxidizing bacteria (NOB)2–4 (eqn (2)), or by ‘comammox’ bacteria that can effect the complete 8-electron oxidation of NH3 to NO3−.5,6
| NO2− + H2O → NO3− + 2H+ + 2e− | (2) |
Given the precise enzymatic control required to form, transport, and use reactive nitrogen compounds such as hydroxylamine (NH2OH) and nitroxyl (HNO), an understanding of the fundamental enzymology and the associated mechanisms can inform our ability to selectively transform nitrogenous species. Yet, despite its importance, the enzymology of nitrification is understudied due to challenges associated with the growth of nitrifying organisms and the recalcitrance of several key nitrification enzymes to recombinant expression. Furthermore, there are also challenges with genetic systems and tools that lag far behind other microbial model systems.
Perfectly embodying these challenges is the poorly understood enzyme ammonia monooxygenase (AMO), which AOA, AOB, and comammox bacteria all use to hydroxylate NH3 to NH2OH as their common first primary metabolic step (eqn (3)).7,8
| NH3 + O2 + 2H+ + 2e− → NH2OH + H2O | (3) |
AMO belongs to the Cu membrane monooxygenase (CuMMO) family. This family includes the better-studied enzyme particulate membrane monooxygenase (pMMO), which carries out a similar oxidation reaction that is essential to the metabolism of methanotrophic bacteria: the conversion of methane (CH4) to methanol (CH3OH).9 Due to their similarity, much of what is known about AMO at the molecular level is drawn from inference to pMMO. Both AMO and pMMO catalyze product formation using as-yet unidentified oxidizing intermediates derived from diatomic oxygen (O2) and Cu.10,11 At present, it is not certain whether said intermediates are common to the two enzymes (vide infra). Both reactions formally require activation of particularly strong bonds: the C–H bond of CH4 with a bond-dissociation free energy (BDFE) of 105 kcal mol−1 and the N–H bond of NH3, with a BDFE of 107 kcal mol−1.12 While AMO and pMMO are highly homologous enzymes with overlapping substrate scopes, the precise structure and mechanism by which these enzymes react with their respective substrates may not be the same.
Selective and controlled functionalization of strong C–H and N–H bonds, such as those found in CH4 and NH3, would tremendously advance understanding and functionality of synthetic oxidation reactions. AMO and pMMO are both capable of CH4 oxidation, overcoming the significant C–H bond strength of CH4 to yield CH3OH.12 Such C–H functionalization is of interest, as CH3OH is of significant industrial value as a feedstock for synthesis of chemicals such as olefins, acetic acid, formaldehyde, and in fuel additives such as methyl tert-butyl ester or biodiesel.13 Many industrial systems exist for the synthesis of CH3OH, although most require high heat or pressure and frequently face overoxidation to undesired products.13 CuMMOs have demonstrated little potential for overoxidation, with pMMO specifically demonstrating stereoselective hydroxylation of substrate.14 Thus, these systems have inspired efforts to synthesize catalysts capable of hydroxylating C–H bonds using an environmentally friendly oxidant, O2. Discussion of such biomimetic synthetic Cu catalysts that attempt to replicate benign C–H activation chemistry of CuMMOs are beyond the scope of this review and documented elsewhere.15
An understanding of the active site architecture and mechanism of AMO would yield valuable insight into this challenging reactivity, but this information remains elusive. Significant challenge exists in purification of active AMO, which to date, has only been purified from large volumes of Nitrosomonas halophila.16 This can be attributed to the slow growth and low cell density of the native organism, a lack of success in recombinant expression platforms, and the inherent difficulty of membrane protein manipulation. Due to these confounding factors, the current understanding of AMO activity is based almost entirely on experiments using whole-cell or cell lysate experiments.
This review is purposed as a primer to the biochemistry of AMO. We have sought to offer sufficient biogeochemical context to underscore AMO's importance, and we have attempted to aggregate available reactivity data.
Nitrification: the basics
Bacteria were definitively identified as drivers of environmental NH3 oxidation upon the isolation of Nitrosomonas europaea in 1890 by Sergei Winogradsky.17 In subsequent years, it was determined by isotopic labeling studies that NH3 consumption in N. europaea accumulated NH2OH, implicating NH2OH as a key intermediate and the initial product of bacterial ammonia oxidation.18 By Arrhenius analysis of NO2− production by whole cell suspensions of N. europaea, it was determined that the apparent activation energy for the oxidation of ammonia is 1.1 kcal mol−1.19 Studies of ethylene oxidation by N. europaea also suggest that in substrate-saturated conditions, the ability to reduce AMO, as opposed to substrate oxidation, is rate-limiting.20| NH3 + 1.5 O2 → NO2− + H2O + H+ | (4) |
The catabolism of NH3 to NO2− is exergonic, with a ΔG°′ of −73.45 kcal per mol NH3 (eqn (4)), which is significantly less than the energy released by the oxidation of glucose (ΔG°′ = −685.94 kcal per mol glucose), even when normalizing for the number of oxidized C atoms (ΔG°′ = −114.24 kcal per mol glucose/C atom).21,22 Further restricting the energetic economy of ammonia oxidizing organisms is the fact that only a maximum of 32.7 kcal per mol NH3 are productively recovered from the electron transport chain, an approximately 53% yield, alongside recovery of less than 2 net electrons per oxidized NH3 molecule.23
| NH2OH → NO + 3H+ + 3e− | (5) |
The NH2OH produced by AMO is oxidized by hydroxylamine oxidoreductase (HAO), yielding the second obligatory AOB intermediate, nitric oxide (NO) (eqn (5)).24 This step is proposed to have an activation energy of 0.89 kcal mol−1, which is less than that of the oxidation of NH3.19 Oxidation of NH2OH to NO yields 3 electrons, which is a net 1 electron gain after budgeting for AMO catalysis.24 The chemical trajectory of the NO generated by AOB remains a matter of investigation. While the terminal product of NH3 oxidation by AOB is NO2− (and this is an intermediate formed by comammox bacteria), no enzyme has yet been identified for the oxidation of NO to NO2−. An enzyme is likely operative rather than abiotic aerobic oxidation of NO because only one of the two O atoms in the NO2− formed by AOB originates from O2 (via AMO). The second comes from H2O.25
In 2005, David Stahl and coworkers26 reported their discovery of the first AOA, Nitrosopumilus maritimus. Since then, AOA have been shown to comprise nearly 40% of the cells in the ocean and account for approximately 34% of oceanic N2O emission.27,28 Thus, despite their until-recent obscurity, these organisms constitute major participants in the biogeochemical nitrogen cycle. Moreover, AOA carry out highly efficient CO2 fixation and organic compound biosynthesis.25,28–34 Similar to AOB, AOA are also found in soil and are implicated in land-based N2O emissions. Within soil, it has been suggested that AOA may outnumber AOB by up to 3 orders of magnitude, although some soils show a more even distribution between these types of nitrifiers.35–38 Despite their higher abundance, the soil-based N2O emissions of AOA are lower than those of AOB.39 Generally, it is understood that the interplay between AOA and AOB in the soil is dynamic, and their relative contributions to nitrification depend on a variety of factors such as pH, temperature, and nitrogen availability.40,41
The AOA primary metabolism remains largely speculative, although it appears that both AOA and AOB initiate nitrification in the same manner. AOA encode an AMO (aAMO) that is a CuMMO, although it exhibits several distinct features from bacterial AMO (bAMO) (vide infra). This implies that NH2OH is a common intermediate to both AOA and AOB, which has been experimentally validated by Arp and co-workers.8 However, it is at this point that AOA and AOB diverge. AOA lack genes encoding HAO homologs as well as for the AOB c-heme electron transfer proteins.3 This has led to the suggestion that either aAMO produces a novel, non-NH2OH intermediate (although the Arp results dispute this), or that a unique, non-heme AOA enzyme serves a similar function to HAO.3,8 Studies demonstrate that in the absence of ammonium (NH4+), N. maritimus is capable of converting NH2OH to NO2−, and that this reaction is directly correlated with both the stoichiometric consumption of O2 and the biosynthesis of ATP.8 It was shown that the marine AOA N. maritimus possesses numerous blue-Cu proteins and multicopper oxidases in its genome; this has since been generalized to numerous other AOA.42 Thus, it is possible that post-AMO primary metabolism of AOA is Cu-dependent, as opposed to the Fe-dependent metabolism used by AOB.3 In recent work, it has been demonstrated that one of these multicopper oxidase proteins, Nmar_1354, is capable of oxygen-coupled consumption of NH2OH to produce HNO.43 This reactivity may be irrelevant to primary metabolism, however, as it shunts reducing equivalents from NH2OH directly to O2 and in doing so, bypasses the respiratory electron transport chain.
Bioenergetic studies predict that isolated AOA and AOB should grow with a theoretical maximum of 0.16 g biomass per g N, and a realistic maximum of 0.13 g biomass per g N after the demands of cellular maintenance are accounted for.23 This is complicated by the experimentally observed yields of biomass in pure and mixed cultures, which range from 0.04 to 0.45 g biomass per g N.23 Such variation implies that our understanding of how energy is harvested from NH3 in AOA and AOB is incomplete, and there is potential for symbiosis between AOA, AOB, and other microorganisms. Interdependence between multiple species would also explain why growth of isolated AOA/AOB is traditionally difficult, slow, and yields minimal biomass, yet wastewater treatment plants grow diverse species at relatively high density.44–50
Foundational studies of AMO
Early studies carried out by Hooper and Terry19 of biological NH3 oxidation by the AOB N. europaea led to the conclusion that intact membranes and metal ions are required. Certain short-chain alcohols were determined to act as specific inhibitors for the enzyme responsible for NH3 oxidation while allowing NH2OH oxidation to proceed, implicating NH2OH as an obligate nitrification intermediate.19 To assist in assignment of product stoichiometry, Hollocher and coworkers21 established a H+/O ratio for NH4+ oxidation by N. europaea of ca. 4, indicating that NH3 oxidation is initiated by a monooxygenase-type enzyme. Later on, Ensign and Arp11 demonstrated that inactive metal-free N. europaea extracts show restored NH3 oxidation activity (quantified via observation of NO2− production and O2 consumption) upon addition of Cu, further supporting the identity of the required metal ion as Cu. In subsequent years, NH3 oxidation was directly assigned to AMO by suicide inhibition with radiolabeled 14C acetylene, which inhibits NH2OH production and yields radiolabeled AMO.51 Moreover, treatment of N. europaea with 15NH4Cl led to the observation of 15NH2OH, showing that the nitrogen atom of NH2OH is derived from NH3.10,18 Similarly, exposure of N. europaea to 18O2 leads to the observation of NH218OH, clearly demonstrating that a monooxygenase enzyme is responsible for the incorporation of an O atom from O2 into NH2OH, with the other oxygen atom being lost as H2O.10 Further tying the oxidation of NH3 to the consumption of O2 is the correlated increase in O2 consumption of N. europaea cell suspensions upon introduction of NH3.52Comparative AMO genetics
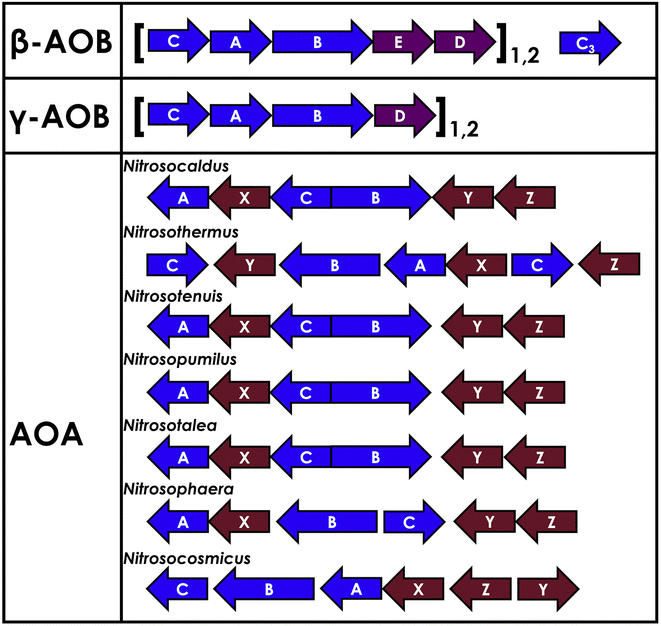
Both aAMO and bAMO are CuMMOs, albeit with kingdom-specific differences in structure and function. These are multi-subunit integral membrane proteins. NH2OH is observed as an intermediate common to both AOA and AOB, suggesting their AMOs share this product.8,53 It is particularly interesting that, despite similar reactivity, bAMOs are more closely related to pMMOs than aAMOs. The bAMOs share ∼70% amino acid sequence similarity to pMMOs, while aAMOs share ∼40% sequence similarity to pMMOs (Fig. 2).54,55 The operons responsible for encoding AMO in bacteria and archaea are shown in Fig. 3.56 The bAMO genes include, in order on their AMO-associated gene clusters, amoC, amoA, and amoB, which encode the respective amoC, amoA, and amoB subunits of the bAMO complex. At the end of betaproteobacterial amoCAB, two open reading frames, orf4 and orf5 (also referred to as amoE and amoD) are present. These are associated with the putative protein amoD, which bears high similarity to methanotrophic pmoD.57,58 PmoD has been characterized as a cupredoxin homodimer that houses an interfacial CuA site. PmoD has been implicated as essential to the Cu-dependent metabolism of methanotrophic bacteria.58 Betaproteobacteria such as N. europaea contain both amoD and the homologous amoE, plausibly due to a duplication event, and thus have an amoCABED gene cluster. Gammaproteobacteria such as Nitrosococcus oceani only possess amoD, thus presenting an amoCABD gene cluster.59 | ||
| Fig. 2 Sequence comparison of the individual subunits of aAMO from N. maritimus, bAMO from N. europaea, and pMMO from M. capsulatus str. Bath. Fully conserved regions are shown in yellow, identity between aAMO and bAMO is shown in cyan, identity between bAMO and pMMO is shown in grey, and identity between aAMO and pMMO is shown in magenta. The initial portion of the subunit A sequences are a signal sequence. Copyright 2018, Cell Press, reprinted with permission.55 | ||
In the gammaproteobacterium N. oceani, an amoR gene was discovered and determined to be transcriptionally linked with amoC.57,60 Attempts to search for peptide sequence homology to amoR have yielded little success, although some authors believe it to encode a small cytoplasmic protein (71 residues) containing 3 small helices (9, 4, and 9 residues at the N terminus) and one larger helix (19 residues at the C-terminus). This region of the gene is preceded by a Shine–Dalgarno sequence to promote transcription.57 Structurally, it has been proposed that amoR is similar to the N-terminal domain of frizzled related protein 3, which has been implicated in protein–protein interactions.57,61 Given the presence of 3 Cys residues in amoR and analogy to frizzled related protein 3, it may be possible that changes in the reduction potential of the environment (e.g. the cytoplasm) can enable formation of a disulfide bond between Cys37 and Cys71, locking the protein to a particular conformation to promote reactivity or protein–protein interaction.57 However, due to this gene only being present in N. oceani, it is unlikely that this gene is part of the conserved bAMO complex.56,57 Little further study has occurred.
Early studies of N. europaea bAMO revealed a 27 kDa protein expressed by the amoA gene and a 42 kDa peptide expressed from amoB, both of which are highly homologous with the corresponding pmoA and pmoB components of methanotrophic pMMO.62–65 Notably, amoA binds acetylene, which has been used as an irreversible inhibitor of AMO, as demonstrated by exposure to 14C acetylene yielding radiolabeled peptide with an apparent mass of 28 kDa (vide supra).51 In recent years, the primary interest in amoA has been as a biomarker of AOB and AOA. Detection of amoA transcripts in water and soil have led to developments in the phylogeny of AOA and AOB, discovery of new species, and an understanding of how these populations respond to environmental stimuli.66–72
In AOB, an upstream gene encoding a highly hydrophobic 27 kDa protein was found ahead of the previous known amoAB operons, which came to be called amoC.73 In N. europaea, there exist 2 copies of amoA and amoB, while in another AOB, Nitrosospira sp., 3 copies are present, with minimal mutation between the copies.74–77 The gene clusters amoCAB1 and amoCAB2 appear to be differentially regulated, and this has been invoked as a means for the cell to rapidly adjust to external stimuli.74 The amoC gene is unique, in that it is the only constituent of bAMO to exist as both part of the gene cluster and duplicated as an isolated gene, identified as amoC3.78 This divergent copy of amoC is primarily expressed when the cell is recovering from NH3 starvation, with the duration of starvation directly correlating to the magnitude of amoC3 expression.78,79 The amoC3 gene is also upregulated upon heat shock or oxidative damage, further implicating it as part of a stress response.80 While the divergent amoC3 unit bears significant deviation in sequence identity (67.5%) from the nearly identical amoC1 and amoC2 (99.6% sequence identity), it maintains 81.4% sequence similarity, suggesting relatively high homology (Fig. 4). As the product-forming active site of AMO may exist in the amoC subunit (vide infra), it is plausible to assume frequent oxidative damage may occur to the protomer, damaging it to an extent that requires replacement. This also suggests that in vitro, the subunits in the AMO complex are exchangeable, which has yet to be validated. The aAMO is the most distinct of the CuMMOs.56 Within the AOA genome, homologs of amoA, amoB, and amoC were found to be conserved across a variety of archaeal species.81 Closely associated with amoA is orf38, also known as amoX, initially identified in the soil AOA Nitrososphaera viennensis.37,56,81,82 Gene cluster analysis of N. viennensis by Schleper and co-workers56 has demonstrated the co-regulation of not just amoX, but also two additional genes amoY and amoZ with amoA, amoB, and amoC, leading to a more complicated archaeal AMO complex that is predicted to be a trimer of hexamers. These additional subunits (X, Y, Z) are relatively small, offering an explanation for their prior omission.56 These 3 small subunits are conserved across all AOA, are highly transcribed, and upon addition to amoA, amoB, and amoC, increase the number of transmembrane (TM) helixes per protomer from 10 to 14, increasing similarity to bAMO and pMMO (vide infra).56 Archaeal amoC is also abundant in the genome of viruses that infect Thaumarchaeota like AOA, although the precise interplay between AOA, amoC, and the associated viruses remains yet to be elucidated.83
AMO structure
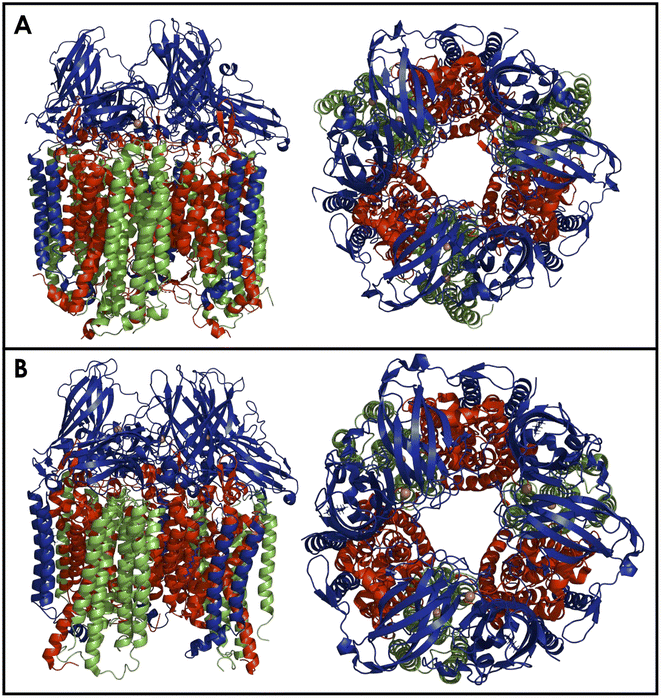
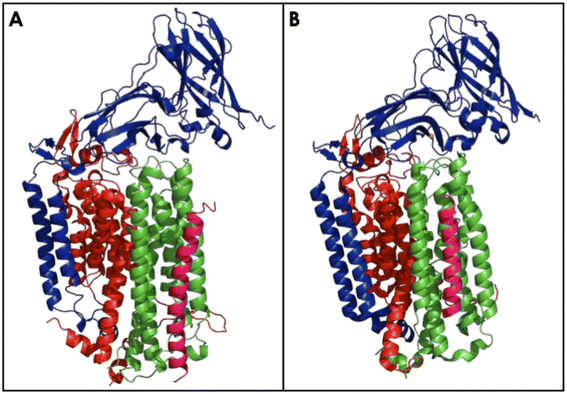
The bAMO is a trimer of heterotrimers, in the form of an α3β3γ3 arrangement of the amoA, amoB, and amoC subunits (Fig. 5).84 The structure of AMO in native N. europaea membranes has recently been solved by use of cryogenic electron microscopy (cryo-EM) to 2.8 Å resolution.84 Furthermore, a similar cryo-EM structure in native membranes has been reported from Nitrosospira briensis, to a resolution of 2.6 Å.85 Cryo-EM of the bAMO complex has also been reported to 2.36 Å from N. halophilia.16 This has allowed for comparison with pMMO, of which numerous examples have been structurally characterized by both X-ray crystallography and cryo-EM techniques under several conditions.84,86–94 Within the native membrane, both bAMO and pMMO form hexagonal arrays, and these arrays have been implicated as necessary for maximal pMMO activity (Fig. 6).84,93 Therefore, it can be inferred that the arrangement of the bAMO heterotrimers within the membrane likely impacts protein structure and function, potentially impacting the ability of endogenous electron donors or substrate to bind correctly. Mechanical pressure imposed by packing of bAMO/pMMO in these arrays may also be important to active site formation, e.g., through imposing structural proximity of multiple Cu binding sites (vide infra). To date, aAMO has not been structurally characterized, nor have hexagonal arrays of the protein in its native environment been observed. | ||
| Fig. 6 Cryo-EM of AMO from N. europaea, as 2-D class averages demonstrating hexagonal packing in the native membranes. Box size are 320 pixels (top) and 640 pixels (bottom). Copyright 2025, National Academy of Sciences. Portion of original figure published under a CC BY-NC-ND license.84 | ||
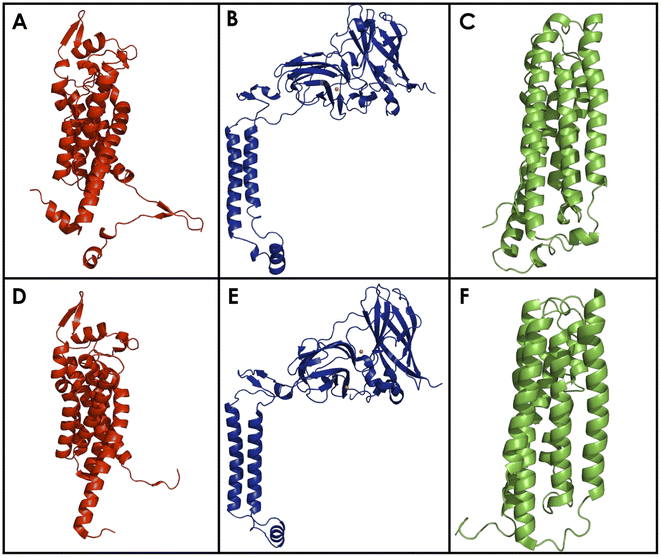
The amoA and amoC subunits of bAMO are largely composed of TM helices (Fig. 7). AmoA and amoC are both composed of 6 TM helices, and measure 32 kDa and 30 kDa respectively.84 The archaeal amoB subunit has only 2 TM helices, with the rest of the protein comprising a soluble cupredoxin-like domain, contrasting with the bacterial amoB (43 kDa) and pmoB, which consists of a pair of cupredoxin-like domains connected by the 2 TM domains.84 Many early structures of the pmoC subunit contained an unmodeled region on the interior pore of the protein, made of highly conserved residues. This region was later resolved in high resolution cryo-EM, demonstrating that this region of pmoC/amoC requires significant stabilization from native lipids in the interior pore of the protein to avoid disorder.84,90
Within the native structures of pMMOs from type-II alphaproteobacterial strains and bAMO, an additional helix of unknown origin is consistently observed in a groove along the external face of amoC (Fig. 8).16,84 In bAMO, the enigmatic extra helix is significantly longer than is observed in pMMO, and is oriented in the opposite direction.84 It has been associated with a previously unannotated 46 residue orf in the genome of the AOB N. europaea.84 Attempts to identify this helix from pMMO by mass spectrometry have failed, although it has been observed that lipids wedged between this helix and amoC interact strongly with a pair of conserved arginine residues.84,88–91 In the recent native-membrane structure of pMMO from M. sp. Rockwell, this helix has been associated with a 23 residue orf in the genome of this organism.84 It has been suggested that the Cu-containing protein, pmoD/amoD, may bind in this site due to the highly conserved nature of those proteins and their genetic proximity to the pMMO/AMO subunits.95 However, attempts to model or observe pmoD in this position have yielded no strong evidence of association. The most recent high-resolution cryo-EM structure of bAMO, from N. halophilia, contains this helix and identifies strong interactions with a lipid identified as phosphatidylmethanol.16 Note that this study refers to the enigmatic helix as amoD, and this is distinct from the pmoD/amoD protein which contains a binuclear CuA site and is proposed to be an electron-transfer partner protein.16 Furthermore, the gene responsible for the encoding of this small helical protein are only present in the lineage of AOB containing N. europaea and Nitrosomonas mobilis.16
The structure of bAMO from N. halophilia also shows an additional density: a small soluble protein comprised of 2 small helixes and 2 small loops.16 This protein has been termed amoE, conflicting with the previously proposed CuA-containing amoE protein.16
Other studies of pMMO have assigned electron density in this cleft to duroquinol, a synthetic mimic of native quinones, potentially implicating this position as the binding site for a biological reductant.90 Ultimately, this helix remains of unknown significance in both pMMO and bAMO, though it has been speculated that this helix could promote array formation or binding with partner proteins.
AOA are uniquely adapted to leverage the membrane-bound nature of AMO. Most archaea and some bacteria have a porous, proteinaceous layer around the outermost cell membrane, called the S-layer, creating a pseudoperiplasmic space between the cell membrane and the environment at large (vide infra). N. maritimus has an S-layer with hexameric pores lined with negatively charged residues, potentially creating a locally high concentration of NH4+ between the S-layer and the cell membrane, where AMO is found, allowing these organisms to sustain themselves in nutrient poor environments.96
The aAMO, while bearing low sequence homology to bAMO, comes to bear high structural similarity with bAMO upon the inclusion of the amoXYZ subunits, as determined by AlphaFold modeling studies (vide infra).56 To date, the only structurally characterized component of aAMO is the cupredoxin domain of the B subunit, from thermophilic Nitrosocaldus yellowstonensis (Fig. 9).97 This soluble amoB unit and pmoB yield low sequence similarity (∼20%), yet structurally, the cupredoxin domains are highly structurally homologous and nearly superimposable (Calpha RMSD = 1.5–1.6 Å).97 While structurally similar to pmoB, the isolated amoB subunit does not appear to oxidize CH4.97 This is remarkable, as in a previous study, using a similar soluble pmoB subunit from Methylococcus capsulatus (Bath), demonstrated a specific activity for the oxidation of CH4 of 203.1 ± 20.2 nmol CH3OH per µmoles per min which was confirmed to be dependent on Cu rather than Fe.98 It is possible that the oxidation of CH4 observed does not represent the biological mechanism of pMMO, and that the Cu site in the B subunit generates a reactive oxygen species that can then proceed to engage in chemistry with aqueous CH4. Studies have shown that oxidation of CH4 can occur by a reaction that is dependent on exogenous reductant duroquinol and O2 and this reaction is most likely not enzymatic.99,100 The reactivity of CH4 and duroquinol suggest that the absence of this reactivity in amoB is not entirely unexpected, although it is possible that the functionality of the archaeal amoB subunit is reliant on stabilization of the cupredoxin domain by another protein such as amoZ, as amoZ likely contains a small, soluble domain.56,97 Without a high resolution structure displaying the precise position and role of amoX, amoY, and amoZ in the aAMO complex, any proposal as to their role is highly speculative and future work regarding these additional subunits remains necessary.
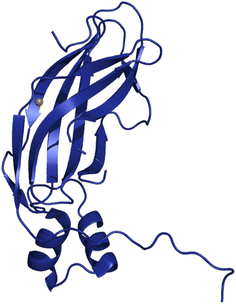 | ||
| Fig. 9 The soluble domain of archaeal amoB, from N. yellowstonii, as characterized to 1.80 Å by X-ray diffraction. PDB: 4O65. | ||
A general overview of the potential metal centers in AMO
Within pMMO and AMO, current evidence suggests at least two primary metal binding sites, one located within the B subunit (CuB), and the other(s) located within amoC (CuC or CuD), totaling approximately 2 to 3 Cu per protomer.95 Some groups have adopted interpretations assigning significantly more Cu to each protomer, such as trinuclear Cu clusters in amoA or Cu “sponges” within amoC that contains ca. 10 metal ions.92,101–105 Over the years, a variety of metal ions have been associated with the active sites of pMMO and AMO, not limited to but including Cu, Zn, and Fe. It is known that some methanotrophic bacteria can use soluble methane monooxygenase (sMMO), a diiron enzyme, to grow under Cu-depleted conditions.58 In parallel with the sMMO/pMMO pair, previous work has suggested the purification and isolation of a soluble form of AMO to be possible, containing approximately 9.5 mol of Cu, 4.0 mol of Fe, and 0.5 to 2.6 mol Zn per mol of N. europaea AMO complex.106,107 These 4 iron are proposed to be divided into 3 heme iron, in the form of a cytochrome c1, and 1 nonheme iron.106 An alternative explanation, however, is that Gilch had isolated partially functional amoA and amoB. Thus, the observed Cu would be attributable to amoB. Yet, the third subunit purified for this soluble form of AMO contained one heme per protomer, with distinct heme Q bands at 552 nm. One heme per trimer would yield 3 heme-bound iron, with a 4th Fe somewhere else in the protein. Thus, another constituent of this preparation could be cytochrome c552 or c554, both of which have shown potential to act as a biological reductant for AMO in accordance with NH3 oxidation activity.108 No further studies of “soluble AMO” have been reported.Other studies have also proposed an intermediate-spin (S = 3/2) Fe ion, capable of forming a ferrous nitrosyl, to be required for oxygen activation in AMO/pMMO.109–111 However, all of these proposals are at odds with the recent structural data supplied by Rosenzweig and co-workers, which does not show Fe or heme cofactors in AMO.84
Studies of membranes containing pMMO show that activity is abolished upon treatment with cyanide, but can be restored with addition of Cu ions, pinpointing the enzymatic active site to a membrane-affiliated protein.91 In the high resolution structures of both AMO and pMMO, there has not yet been a heme cofactor detected, and the electron density of the bound metal ions is consistently best modeled with Cu and/or Zn. Therefore, more recent work regarding CuMMOs has focused on the potentially catalytic Cu sites in amoB and amoC.
The non-conserved bis-His site
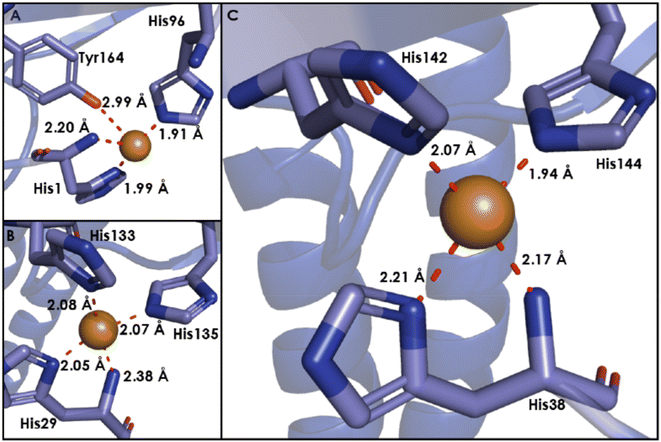
Within pmoB of pMMO, there exists a mononuclear Cu center ligated by 2 histidines, denoted as the bis-His site (Fig. 10). This site has been observed in both crystallographic and cryo-EM structures of M. capsulatus (Bath), yet it is not strictly conserved, as one of the His ligands is replaced by asparagine in some methanotrophs.86,88–92 This lack of conservation, alongside the absence of this site in AOA or AOB, suggests that this site plays little catalytic role, and Cu may be adventitious. In methanotrophs containing the bis-His site, it is proposed that in vivo, this site binds CuI, due to the nearly identical EPR spectra of pMMO in methanotrophs lacking this Cu ion.100,112 While some attempt to observe or characterize a potential bis-His site in AMO may be warranted, it is unlikely to be important to the catalytic mechanism.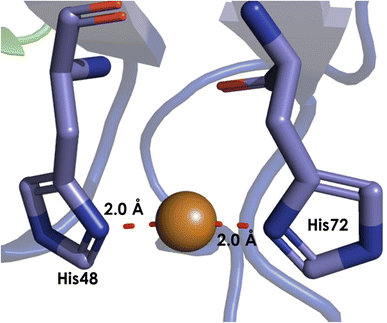 | ||
| Fig. 10 The bis-His site from M. capsulatus (Bath) pMMO (PDB: 7S4H), showing the 2 coordinated His residues, as resolved to 2.14 Å by cryo-EM. | ||
The conserved CuB site
Within amoB and pmoB, there exists a generally conserved Cu site, CuB, coordinated by 3 His residues (Fig. 11).84,113 This motif resembles the ‘His-brace’ structure found within lytic polysaccharide monooxygenases (LPMOs), which catalyze the cleavage of glycosidic bonds in polysaccharides.114,115 In resting LPMOs, a CuI ion is coordinated by the amino group and sidechain of an N-terminal histidine and the sidechain of an additional histidine residue. X-ray absorption near edge spectroscopy (XANES) has been used to assign the oxidation state of the Cu ion in CBM33, a polysaccharide oxidase.116 Comparison of a sample treated with sodium ascorbate and an as-isolate sample demonstrate that both samples show the rising Cu K-edge to be at 8982–8983 eV, indicating that the resting active site binds CuI.116 In CuMMOs, this motif is distinct, with the resting CuB site binding CuII and coordination by an additional His residue. While there has been previous discussion of a binuclear CuB site, recent studies using high-energy resolution fluorescence detected (HERFD) extended X-ray absorption fine structure (EXAFS) has determined this site to be mononuclear CuII.117 This determination was made by observation of a pre-edge feature at 8978 eV, which is assigned as the 1s to 3d transition of CuII.117 Given the similarity to the active site of LPMOs, it has been suggested that the CuB site may be the active site of CuMMOs. However, this conjecture suffers from the fact that some novel LPMO-like fungal proteins contain a His-brace motif, yet do not demonstrate oxidative enzymatic activity.118,119Attribution of activity to the CuB site has been challenging. Studies of a soluble form of pmoB have demonstrated that the CuB site is capable of oxidizing both CH4 and propylene, while a soluble archaeal amoB was not (vide supra).97,98 Furthermore, the His-brace motif B-site is not rigorously conserved, as methanotrophic Verrucomicrobial pMMOs contain residues such as glycine, proline, and methionine in place of histidine in their B-subunits.120,121 It is also important to note that in a recombinantly expressed CuMMO, mycobacterial hydrocarbon monooxygenase (HMO), mutation of the CuB site did not completely abolish enzymatic activity, ultimately supporting the conclusion that the active site resides in amoC.122,123
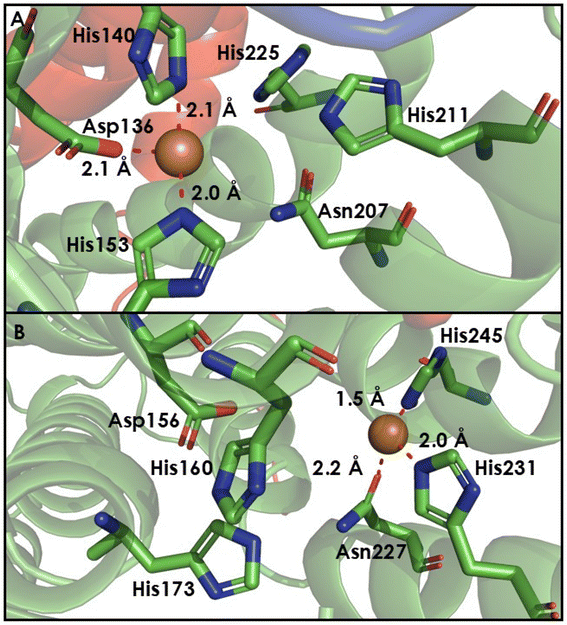
A tale of 2 Cu sites (CuC and CuD) in the amoC subunit
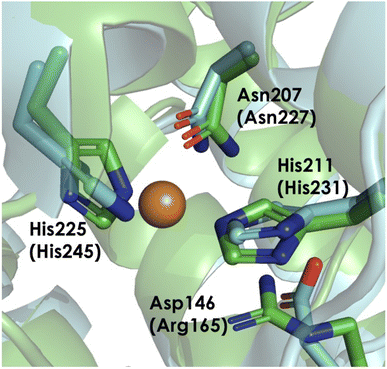
Mounting evidence suggests that the AMO active site is in the amoC subunit. More precisely, activity could occur at either the CuC or the recently proposed CuD site (Fig. 12). Reactivity may also involve both Cu ions, as recent work has demonstrated the simultaneous occupancy of the CuC and CuD sites in functional bAMO (Fig. 13).16,85 Meanwhile, pMMO structures are available whether either CuC or CuD are individually metalated. Both the CuC and CuD site are highly conserved, even in the aforementioned methanotrophic Verrucomicrobia that lack the ligands constituting the CuB site.120,121 Both potential Cu sites in the amoC subunit comprise 2 His residues along with Asp or Asn within CuC or CuD, respectively.84 In computational modeling studies by Ciurli and coworkers,124 it is proposed that an α-helix is formed by residues 211–251 in N. europaea amoC. Three of these helices, one from each component amoC, interact with one another inside the central barrel of the multimeric unit of AMO, helping to stabilize the CuC site in a solvent accessible cave. While these residues do form an α-helix, these residues were observed in the recent native membrane cryo-EM structures of pMMO and bAMO to have a structure more similar to the other TM helices, with no protrusion toward the central cavity of the multimer.84NO2− has been previously established as an inhibitor of AMO activity.125 In the presence of isotopically labeled 15N NO2−, the EPR and electron nuclear double resonance (ENDOR) spectroscopic signatures of the CuII at the CuC site of pMMO is perturbed (Fig. 14).98 This suggests NO2− inhibits NH3 oxidation by binding to an active site Cu. Studies of HMO have demonstrated that mutations at the CuB site do not abolish enzymatic activity, while mutation of any one of the three residues at the CuC center lead to a complete loss of function.122,123 Historically, it is clear that the C subunit is essential for oxidative activity, although the exact nature of the bound Cu has recently come into question.
 | ||
| Fig. 14 (A) X-band CW EPR of purified pMMO, before and after introduction of NaNO2. Perturbations to CuC are most apparent ∼3300 G. (B) Q-band Mims ENDOR of purified pMMO, before and after introduction of NaNO2. The full spectrum is displayed above, whereas the lower spectrum highlights the precise 15N frequency and associated hyperfine at improved signal to noise ratio. Copyright 2019, American Association for the Advancement of Sciences, reprinted with permission.100 | ||
It is known that short chain primary alcohols, such as CH3OH, ethanol, and tert-butanol are capable of acting as inhibitors of pMMO and AMO.19 This information has enabled a study using ENDOR spectroscopy in tandem with cryo-EM to identify the spectroscopic signature of trifluoroethanol (TFE) alongside its physical binding location, which was revealed to be the CuD site.94 In the ENDOR spectrum of TFE bound pMMO in native lipid bilayer nanodiscs, a 19F doublet is observed that is not present in uninhibited sample.94 TFE binds to the CuD metal ion through its hydroxyl group, leading to a 2.2 Å Cu–O distance and an average Cu–F distance of ∼5 Å, after accounting for the fact that the trifluoromethyl group demonstrates free rotation.94 As pMMO is also capable of butane oxidation, these experiments were repeated with 4,4,4-trifluorobutanol (TFB), in which a larger, yet less ordered density, attributed to TFB, was reported attached to the CuD site.94
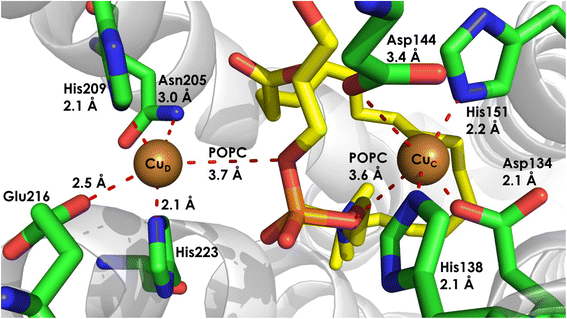
To further strengthen the requirement of CuD at the catalytic site responsible for hydroxylation of NH3, it has been observed by cryo-EM of pMMO that the CuD site is occupied in active pMMO, while a disordered CuD site and occupied CuC site are observed in inactive pMMO (Fig. 16).90 These studies directly correlate CuD occupancy to the activity of pMMO. Initial Cryo-EM data of bAMO demonstrates significantly lower density in the region surrounding the CuD site than the CuC site, suggesting that this site is particularly labile.84 More recent evidence has been gathered in support of co-occupancy, creating cooperative active site requiring both CuC and CuD may also be worth consideration, due to their close proximity.16,85 Such a site could be formed dynamically, with driving force supplied by pressure from the membrane. This offers a plausible explanation for the wildly different activities encountered between pMMO preparations in, e.g., detergent, native membranes, and lipid bilayer nanodiscs. Finally, it is important to note that a site amenable to NH3 substrate binding would likely require a coordinatively unsaturated Cu ion in a hydrophobic pocket, and the cryo-EM structures of pMMO, both with and without TFE, demonstrate a small hydrophobic cavity lined by phenylalanine residues.84,94
The most recent work regarding bAMO has demonstrated occupancy of both the CuC and CuD sites. In the structure of bAMO, in native membranes from N. briensis, the CuC/CuD site is observed to be in a diamagnetic resting state, most likely Cu(I)/Cu(I).85 The two Cu ions are separated by ∼8.0 Å, as a non-coordinating phospholipid (identified as 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine) rests between them.85 Other authors have reported an interatomic Cu distance of 7.9 Å, with an unidentified density residing between the Cu ions.16 Addition of this lipid and binuclear occupation of the site bends the helix containing the residues coordinating CuD towards the central pore.16,85 This transformation is assisted by the highly conserved Pro200 residue and yields a previously unobserved conformation of the potential active site.16,85 This kinked conformation, in which the helix from amoC bends towards the central pore, also enforces changes in helixes of amoA, creating a hydrophobic channel from the central pore to the binuclear CuC/CuD site.16 With these considerations, significant development has been made in demonstrating the fluid and dynamic nature of the amoC subunit and the associated proposed active site(s).
AlphaFold modeling of AMO
Given the limited structural data of AMO, computational methods of sequence analysis and protein folding become potential sources of insight. To investigate conservation at and near the CuC and CuD site, we have run sequence similarity networks (SSN) using EFI-EST, comparing amoC to 679 known unique sequences of other xmoC sequences.126,127 Analysis of the SSN shows high conservation of the ligands directly coordinating both CuC and CuD, yet the loop between the His residues of the CuD site is poorly conserved (Fig. 15). This has been presumed to be a region of protein with little-to-no secondary structure, and it is possible that variation in this structural region is responsible for the substrate specificity of various CuMMOs. Reiterating the potential for structural differences in this region between various CuMMO's, overlap of the cryo-EM structures of amoC and pmoC demonstrate that pmoC contains an arginine at the CuD site where amoC has an aspartate residue in this position (Fig. 16). To expand beyond the available structural data, AlphaFold 3 was used to create renditions of the folded bAMO trimer (αβγ) using the sequence from N. europaea ATCC 19718, in the presence of 2 Cu atoms.128 This computational structure is generally in good agreement with the cryo-EM data recently published. The model templated onto the cryo-EM structure (ipTM = 0.83, pTM = 0.85) demonstrates metalation of the CuB site and the CuD site, whereas the untemplated model (ipTM = 0.85, pTM = 0.87) shows metalation of the CuB and CuC site. In observation of the AlphaFold amoC subunit, a small dangling region of residues is observed at the N-terminus, which has not been observed in the recent cryo-EM structure. It is possible that this may be a misannotation or a previously unknown signal sequence, as it is 19 residues from start to the next Met residue. In preparation of these models, SignalP analysis of the amoB subunit also revealed a signal sequence that is cleaved in vivo. This portion of the sequence was omitted from the AlphaFold3 models. Given these results, it is apparent that the CuD site is becoming of increasing interest, and the forces that dictate substrate differentiation between CuMMO's may lie in the region surrounding the CuD site.AlphaFold 2.1 has also been used in previous work to demonstrate the effective homology between bAMO from Nitrosocaldus cavascurensis and the more distantly related aAMO from N viennensis upon inclusion of the additional amoX, amoY, and amoZ subunits (Fig. 17).56,129 AmoX, amoY, and amoZ all appear to play a role in anchoring aAMO into the membrane, alongside amoA, amoB, and amoC. AmoZ was also determined to contain a disulfide bond assisting in structural stability and a flexible loop that allows for positioning of a pair of helices in a soluble domain that interacts with the cupredoxin domain of amoB, potentially acting as a replacement for the cupredoxin domain ‘missing’ from aAMO.
AmoD: a unique CuA site
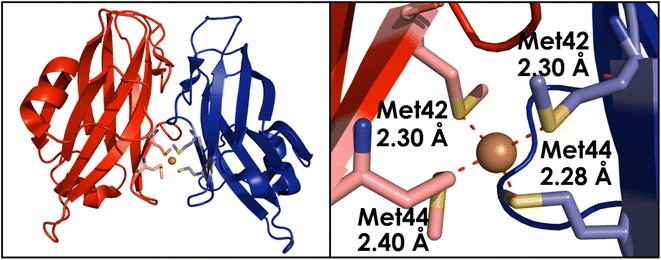
Unique to AOB and methanotrophs is the closely associated pmoD/amoD gene, which has yet to have an identified homolog in AOA.58 Betaproteobacteria contain amoE and amoD directly after the amoB/pmoB gene, while gammaproteobacteria only contain amoD, also located directly after the amoB/pmoB gene.59 Studies by Fisher58 and colleagues have demonstrated that disruption of pmoD results in a Cu-dependent growth defect. In methanotrophs lacking pmoD, growth was stunted in conditions that rely on pMMO for primary metabolism, although the cells grew as expected under conditions that take advantage of sMMO.58 This implicated the pmoD protein as particularly relevant to Cu-dependent metabolism in methanotrophic bacteria. Given the high similarity between pMMO and bAMO, it is possible that the role of amoD is similar to that of pmoD, although little work has been done regarding amoD to date.Sequence analysis of pmoD/amoD provide evidence for an N-terminal signal sequence, followed by a soluble domain and a C-terminal TM helix.58 The soluble domain of PmoD can be recombinantly expressed in E. coli, allowing for purification of a purple homodimeric protein containing a unique CuA site (Fig. 18).58 CuA sites are characterized by a delocalization of a single electron across a binuclear Cu site; similar sites containing a mixed valent Cu(+1.5)–Cu(+1.5) ligated by 2 µ2-S(Cys) and 2 planar N(His), alongside axial methionine and carbonyl backbone ligands, are found in cytochrome c oxidase (CcO) and nitrous oxide reductase (N2OR).130–136 The CuA site exists at the interface of the homodimer, with each monomer contributing 1 µ2-S(Cys) ligand to the catalytic core.137 In monomeric form, no spectroscopic signatures for the CuA site are observed, suggesting that amoD/pmoD must exist as a dimer in vivo. The Cu–Cu distance is particularly short, at 2.41 Å by EXAFS data, and this conclusion is supported by the observation that the near-IR transition associated with the Cu–Cu bond strength is blue shifted relative to those observed in N2OR and CcO, indicating a stronger bonding interaction in amoD/pmoD than other known CuA sites.137–139 The CuA site of pmoD is significantly more unstable in the presence of oxygen than other reported CuA sites, decomposing to a 2 [Cu+2] site on the scale of a few hours.
Sequence comparison by Musiani, Ciurli and coworkers124 of pmoD from Methylocystis sp. ATCC 49242 (Rockwell) and amoD/E from N. europaea demonstrate 38% and 28% sequence identity, respectively. These authors also note that the methionine residue responsible for axial ligation in pmoD is not conserved in amoD or amoE. Computational modeling of the CuA site in amoE yields a long Cu–Cu distance of 2.55, alongside a conformational change at the N-terminus that reduces the protein–protein interaction surface.124 While intuition may suggest that these amoD/pmoD proteins play a stabilizing role, act as a biological electron transport partner or a source of Cu ions, or some other role in relation to AMO, significantly more work is required in this area, as these proteins have only recently been purified.
Substrate scope of AMO
Beyond NH3, AMO is known to have a wide scope of substrates and inhibitors. Contained in the supplementary information is a compilation of all currently known substrates, inhibitors, and associated kinetic parameters, alongside relevant experimental information. Due to high homology between AMO and pMMO, one may intuit AMO is capable of CH4 oxidation. This was proven by early studies by Hyman and Wood140 demonstrated that N. europaea cell suspensions incubated under O2 are capable of oxidizing CH4 to CH3OH. These authors reported a Km for NH4+ of 1.2 mM and Ki for CH4 of 2 mM, suggesting that these may function more as alternative substrates for CuMMOs than inhibitors.140 It is worth noting that later studies by Jones and Morita141 demonstrated that both N. oceanus and N. europaea are capable of CH4 oxidation, with whole cell assays in the presence of 14CH4 showing both production of 14CO2 and incorporation of carbon from CH4 into cellular components. These whole cell assays also exhibit robust pH stability, with CH4 oxidation observed in significant quantity between pH of 5 and 9.141N. europaea has been shown to oxidize CH3OH to formaldehyde and formate following incubation with 13CH3OH.142 Addition of known NH3-oxidation inhibitors such as acetylene, or allylthiourea both prevent the oxidation of CH3OH, suggesting that AMO is the source of CH3OH oxidation.142 While such promiscuity may suggest AMO to be a rather nonspecific enzyme, ratios of kcat/Km for CH3OH and CH4 relative to NH3 are determined to be 0.008 and 0.004, respectively.141,142 Further, the enzyme is also capable of reaction with a wide variety of hydrocarbons.143 C1–C8 alkanes can be hydroxylated selectively at the 1 and 2 positions, with the ratio of 1-ol/2-ol products varying with increasing chain length.143 Work by Chan and coworkers14 has shown that, in pMMO, these hydroxylation events occur is a stereospecific manner, leading them to propose a side-on oxene insertion mechanism at a binuclear active site, suggesting that AMO may also be capable of stereospecific reactivity.
Whole cell N. europaea have also demonstrated the ability to hydroxylate benzene to phenol, phenol to hydroquinone, and cyclohexane to cyclohexanol. These reactions have not been demonstrated using methanotrophic pMMO.144 Further reactivity studies of AMO and benzene, ethylbenzene, and toluene show the formation of associated phenolic compounds alongside hydroxylation of the alkyl groups, leading to formation of benzyl alcohol, phenylethyl alcohol, and sec-phenylethyl alcohol.145 Aniline was converted to nitrobenzene, which is further oxidized to 3-nitrophenol, although it is worth noting that this transformation may actually be occurring through reactions mediated by HAO.145 Halobenzenes can also be oxidized in the para position, producing low yields of halo-phenols.145 The tolerance of N. europaea to halogenated organic compounds is rather diverse, with chloromethane, chloroethane, and 1,2-dichloroethane showing substantial conversion to formaldehyde, acetaldehyde, and chloroacetaldehyde respectively, with minimal inactivation of the AMO complex.146 Other substrates undergo biodegradation but form strong inhibitors as products, such as dichloromethane, which becomes CO, and 1,1,1-trichloroethane, which becomes 2,2,2-trichloroethanol.146 A common, seemingly essential feature of these substrates required for AMO reactivity is an abstractable H-atom. Compounds such as carbon tetrachloride or tetrachloroethylene do not show any reaction with or inhibition of AMO.146 Whole-cell activity has also demonstrated consumption of methyl bromide, suggesting that brominated and/or iodinated species may also function as alternative substrates for AMO.147 AMO is also capable of epoxide formation upon introduction to alkenes, showing activity with ethylene, propylene, 1-butene, and both cis and trans 2-butene, alongside styrene.143 Further extending the substrate scope of AMO are thioethers, which act as inhibitors of the enzyme but can also be converted to their corresponding sulfoxide.148 Examples include the oxidation of dimethylsulfide to dimethylsulfoxide (DMSO) and tetrahydrothiophene to its corresponding sulfoxide, alongside various other substrates and their associate products.148 Given the unclear nature of the mechanism of AMO and the lack of abstractable protons on the S atom, a different mechanism may be operative for thioether substrates.
Ammonia, ammonium, and the substrate specificity of CuMMOs
In the environment, AOA and AOB occupy ecosystems not limited to but including oceans, fresh water, soil, and geothermal geysers. This has led to unique adaptations in NH3 scavenging and the fundamental reactivity of AMO. Given the wide range of pH experienced in such environments and the ecologically accessible pKa of NH3 allowing for both it and the NH4+ ion (pKa = 9.25) to exist in solution, one must consider whether the substrate of AMO is NH3 or NH4+.149 Initial studies by Suzuki, Dular, and Kwok149 investigated the effect of pH on the KM of NH3 by tracking the rate of oxygen consumption by whole cell and cell-free extracts of N. europaea. Both pH and the concentration of NH3 impacted the rate of O2 consumption, with an increase in pH being associated with a decrease in KM (increased affinity) and no change in maximal velocity being observed.149 This has prompted the proposal that NH3 is the true biological substrate of AMO, although it is possible that both the substrate and mechanism may vary between AOA and AOB, due to the differences in structure indicated by the presence of amoX, amoY, and amoZ in aAMO. Early studies of N. europaea suggest a KM between 1.2 mM and 1.5 mM for NH4+, and after accounting for dissolution of the NH4+ ion, a KM of 50 µM was determined for NH3 alongside a Vmax of 1.68 ± 0.15 µmol N per min per mg protein−1.140,150A persistent assumption has been that AOA possess a higher substrate affinity than AOB or comammox bacteria, although recent work by Wagner and colleagues151 has called this into question. The NH3 and NH4+ affinity of a given species of AOA often vary widely, with examples including those of the acidophilic genus ‘Ca. nitrosotaleales’ demonstrating a particularly low affinity for NH4+ (KM(app)=3.41–11.23 µM) and a very strong affinity for NH3 (KM(app) of 0.6–2.8 nM), which is likely to assist in NH3 acquisition in acidic conditions.151 Generally, the substrate affinity for AOA is on the order of µM, although the complete range spans orders of magnitude, as evidenced by the particularly strong affinity observed in ‘Ca. nitrosotaleales’.151 While all AOA, aside from some Nitrosocosmicus species, alongside the comammox bacteria, Nitrospira inopinata, have a higher affinity than AOB, the tail end of the affinity range does overlap with those characteristic of AOB, such as N. europaea.151 In contrast to the high variability in affinity between various ammonia oxidizing species, the maximum substrate oxidation rate (Vmax) of all AOA span a narrow range of approximately 1 order of magnitude (Vmax = ∼4.27–54.68 µmol N per mg protein per h).151 This is similar to that observed for N. inopinata (Vmaxca. 12 µmol N per mg protein per h) and N. oceani ATCC 19707 (Vmax = ∼38 µmol N per mg protein per h), a marine AOB.151 The only species, to date, with a Vmax outside of this range is N. europaea, with a Vmax = 84 µmol N per mg protein per h.151 Whether this unusually fast rate is unique to N. europaea, shared with other closely related AOB, or due to adaptation to a lab environment over prolonged study is presently unknown.
As evidence suggests NH3 to be the biological substrate of AMO, it becomes particularly interesting to note that AOA have specialized adaptations to attract and transport NH4+, which may be related to niche separation between AOA and AOB.152 This has led to ambiguity in the identity of the native substrate of aAMO, as previously displayed in differences between bacterial and archaeal nitrification (Fig. 1) Archaeal cells possess a proteinaceous S-layer outside of the cell membrane where aAMO resides (Fig. 19).96,153,154 In the AOA N. maritimus, the S-layer is predicted to consist largely of repeating subunits of the proteins Nmar_1547 and Nmar_1201, which modeling studies have suggested may assist in creating a high local NH4+ concentration in the pseudoperiplasmic space between the S-layer and the cell membrane.79,155–157 It is known that the disruption of the archaeal S-layer leads to inhibition of NH4+ binding, and cryo-EM of the isolated S-layer exhibits unassigned density bound near negatively charged sites within pores.96 This evidence suggests that the lattice itself binds NH4+ for release into the pseudoperiplasmic space and subsequent consumption by AMO, although the S-layer has also been implicated in transport of Na+.96 While a negatively charged S-layer may explain how AOA are capable of acquiring NH4+ in nutrient-deficient environments, this may be incompatible with NH3 as the active substrate of aAMO.
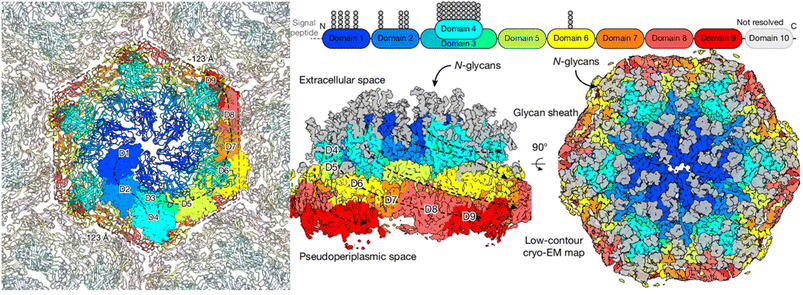 | ||
| Fig. 19 2.7 Å global resolution cryo-EM structure of isolated S-layer sheets, in vitro from N. maritimus (Left). Sharpened, annotated images of the cryo-EM map show the individual domains of the protein, alongside peripheral glycans shown in grey. The colors of each domain are demonstrated in the ribbon map above. Copyright 2024, Springer Nature, reprinted with permission.96 | ||
An aside to the substrate scope of pMMO
As with AMO, significant challenges exist in the isolation and purification of pMMO, yet some progress has been made. This has allowed for the determination that pMMO has a narrower substrate scope than AMO and is primarily capable of oxidizing C1–C5 straight-chain alkanes, terminal alkenes to stereoselective secondary alcohols or epoxides, respectively.158–160 It appears that pMMO, in contrast to AMO, is entirely incapable of oxidizing benzene, ethylbenzene, styrene, or cyclohexane.159The kinetic parameters for various pMMOs are on the same order of magnitude as those observed for AMO, with a KM(app) = 9.2–63 µM for CH4 and a KM(app) = 0.1 µM for O2.161,162 The Vmax of whole cell methanotrophs have also been reported to range from ∼82–300 nmol per mg total protein per min, which is again approximately what is observed for AMO.162 AMO and pMMO have a similar substrate scope and enzymatic parameters, yet it is probable that the active site of pMMO is more constrained than AMO, given the inability to react with larger species such as aromatics and cyclohexane. This difference in reactivity may also be due to differences in mechanism between the homologs.
Inhibitors of AMO
Acetylene and allylthiourea (a common Cu chelate) are two of the first discovered specific inhibitors for NH3 oxidation activity in N. europaea.19 While the precise mechanism of action is unknown, it is speculated that allylthiourea acts as a chelating agent, stripping Cu from the active site of the enzyme. The mechanism by which acetylene inactivates AMO is also not yet understood, but it is presumed to bind to amoA.51 Inhibition by acetylene follows irreversible first order kinetics, most likely with inactivation upon the first enzymatic turnover.51 While acetylene is the primary inhibitor used in many studies of whole-cell N. europaea, it has been observed that n-alkynes, (n = 2 to 10) are also capable of inhibition, with larger alkynes (n ≥ 6) being related to faster inhibition of NH3 oxidation.143 While AOB are quite easily inhibited by alkynes such as 1-octyne, AOA such as N. viennensis and Nitrososphaera gargensis do not demonstrate inhibition by long chain alkyls (n ≥ 5).163 This difference between AOA and AOB has allowed for selective inhibition of AOB and growth of monoculture AOA, while also suggesting structural or chemical differences in the enzymatic region where alkynes bind to inhibit NH3 oxidation activity.163 While resistant to long chain alkynes, aAMO is significantly more susceptible to inhibition by short chain alkynes (n ≤ 5). Inhibition studies using phenylacetylene have demonstrated alkynes to be a noncompetitive inhibitor in both AOA and AOB, but the kinetic characteristics between these two kingdoms are incongruent, suggesting that there may be differences in the binding site and substrate tolerance between aAMO and bAMO. Given that aAMO and bAMO appear to have distinct inhibition profiles, it is important to note that 1,5-hexadiyne can act as an inhibitor of ammonia oxidation in both AOA and AOB while also acting as a fluorescent probe upon in vivo reaction.164 This bifunctional fluorescent probe-inhibitor has demonstrated applicability across a variety of AOA, AOB, and comammox bacteria.Mechanism of AMO
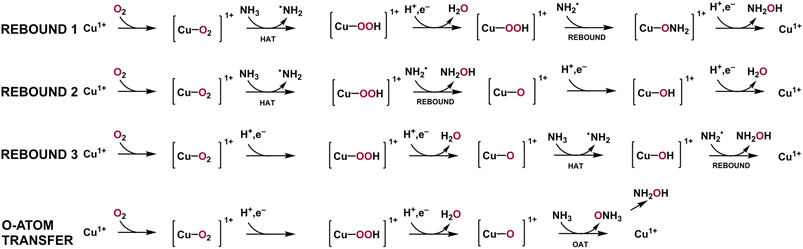
Due to the challenges of obtaining pure AMO, alongside potential differences from the mechanisms proposed for pMMO, the mechanism by which AMO forms NH2OH remains highly speculative. However, two plausible mechanisms serve as starting points for inquiry. One possibility is H-atom abstraction directly from NH3, followed by a radical rebound mechanism involving hydroxyl, in a mechanism resembling that of cytochrome P450.165–167 Such a mechanism would require elevated basicity of the oxygen atom that eventually is incorporated into NH2OH.168,169 Three potential rebound mechanisms are detailed in the scheme below (Scheme 1). The other mechanistic possibility is oxygen atom transfer directly to NH3, to form ammonia N-oxide, which spontaneously rearranges to form NH2OH. This rearrangement has been proposed to be thermodynamically favorable, with ΔG° = −3.0 kcal mol−1 determined by computation.170 Regardless of which mechanism is operational in vivo, both are heavily influenced by the nature of the Cu–O interaction and the electrophilicity of the oxygen-containing species.Given the multitude of Cu containing sites contained in AMO, their interplay may be complex. As the CuB site of AMO and pMMO is highly similar to the His-brace motif observed in LPMOs, a class of enzyme that demonstrates O2 activation at a mononuclear Cu site, it has been proposed that the CuB site may be capable of generation of reactive oxygen species.115,171 Computational studies of CuB from pMMO, using QM/MM methods by Lundgren and coworkers,171 propose that the CuB site can form a [CuO]+ species (best described as triplet CuII–O˙−) capable of H-atom abstraction from CH4. This would form a Cu-bound hydroxyl which may be capable of rebounding to H3C˙, yielding CH3OH. Such a process would begin in a CuII resting state and overcome 14.1 kcal mol−1 and 12.4 kcal mol−1 activation barriers for H-atom abstraction and hydroxyl rebound, respectively.171 In total, the proposed activation energy would be less than 17.2 kcal mol−1.171 While this gives credence to the possibility of H-atom abstraction and radical rebound at the true active site, this study also found that a competing pathway, in which CuB assists in the formation of hydrogen peroxide (H2O2) and perhydroxyl radical (HO2−), may be favored instead.171 Given the previous observation that soluble pmoB has demonstrated the ability to hydroxylate CH4, it is possible that the CH3OH observed was from the reaction of ROS generated by the CuB site and CH4.98 If only pMMO requires the generation of ROS followed by transfer to the catalytic CuC (or CuD) site, and AMO does not, this may also further implicate differences in the mechanisms of these homologs. Further evidence for cooperation between the CuB and CuC/CuD site has been proposed, in the form of a hydrogen bonding network spanning between CuB and CuC.16
Parallel ENDOR/cryo-EM studies by Rosenzweig and Hoffman94 implicate the CuC, CuD, or a potential binuclear CuC/CuD site as the site of CH3OH formation in pMMO. Given that both CuC and CuD exist in close proximity, containing 2 N-ligands and 1 O-ligand each, it is possible to infer that O2 (or a ROS from CuB) may bind at one (or both) of these Cu, which can subsequently engage in substrate hydroxylation.
Outside of the CuMMO family, a variety of other Cu monooxygenase enzymes capable of C–H hydroxylation exist, which may serve as potential inspiration for future mechanistic deductions in AMO.172–175 Two primary examples of enzymes that use Cu and O2 for C–H functionalization are the peptidylglycine α-amidating monooxygenase (PHM) subunit of peptidylglycine α-amidating monooxygenase (PAM) and dopamine β-monooxygenase (DβM).176–178 These species are proposed to hydroxylate secondary C–H bonds with a BDFE in excess of 90 kcal mol−1via use of a [Cu–O2˙−]+ intermediate as part of a radical rebound reaction.172 Given the recently demonstrated flexibility of PHM, which can change conformation to shorten the ∼14 Å Cu–Cu distance to 4–5 Å, a highly dynamic binuclear core may also be operational within AMO.179,180 It is possible that pressure from the membrane pushes the lipid separating CuC and CuD apart, allowing formation of the catalytically competent active site. Therefore, the possibility of a dynamic configuration and the precise role of this lipid require further investigation. Beyond these enzymatic examples, a bent [Cu2O]2+ core has been observed within Cu-ZSM-5, a zeolite capable of methane hydroxylation.181 While these intermediates (Fig. 20) have been proposed and largely accepted within other systems, there is still a significant amount of ongoing work in this area.
Recombinant expression of AMO
While AMO has been purified in active form, consistent and reliable recombinant access, and therefore mutagenesis, remains incomplete. One of the compounding barriers is the low proteomic abundance of AMO in native organisms such as N. europaea (∼5%, total between amoA, amoB, and amoC).182 In membrane fractions of N. viennensis, amoA, amoB, and amoC make up approximately 22% of total membrane protein content.56 Therefore, in a dense culture of a membrane-rich organism, large quantities of AMO may be achievable. This is diminished by difficulty in growing large quantities of AOB, helping native bAMO to remain elusive. Making aAMO even further inaccessible are the particularly slow growth rates and low cell densities of AOA.183 Even if these bottlenecks could be overcome, developing a complete mechanistic understanding of AMO would remain difficult without site-directed mutagenesis, which is unlikely to be achieved with native nitrifiers given the primary metabolic role of the target enzyme. Heterologous expression is complicated by the NH2OH produced by AMO, which is expected to be cytotoxic to any host organisms lacking means to process NH2OH.Thus, recombinant expression and access to site-directed mutations of AMO (and pMMO) have remained a challenge, despite the high value such an endeavor could yield. To date, the heterologous expression of complete CuMMO complexes has largely been restricted to the cloning of a fosmid bearing HMO into Mycobacterium smegmatis.122,123 These studies did not include the purification and isolation of recombinant HMO; focusing instead on observation of substrate oxidation by whole-cell suspensions.122,123 Cells containing the hmoCAB gene demonstrated the Cu-dependent ability to subsist on short-chain alkanes (C2–C4), alongside loss of this characteristic in media lacking Cu or upon addition of allylthiourea.122 A subsequent study of this system demonstrated mutagenesis at the residues coordinating CuB and CuC. In mutants where the proposed CuC site ligands had been individually removed (C-D139V, C-H143V, and C-D149A), no ethane or butane oxidation activity was observed, suggesting that the CuC site is essential for CuMMO activity. This is contrasted with a mutant at CuB, (B-H155V), which demonstrated significant hydrocarbon oxidase activity, albeit highly reduced activity relative to unmutated HMO. Mutations of nearby, noncoordinating residues, such as C-A151D, showed a decrease in activity that was markedly larger for butane than ethane, suggesting mutations in the secondary sphere may allow various CuMMOs to differentiate between their preferred substrate despite high sequence identity. Soluble fragments of pmoB from M. capsulatus (Bath) and amoB from N. yellowstonii have also been recombinantly expressed (vide supra).97,98 While a subunit-focused approach may yield valuable insight into the individual Cu centers of AMO, it also neglects the potential for dynamic interplay between the CuB, CuC, and CuD sites, alongside the potentially critical possibility of protein–protein or protein–membrane interactions.
To date, the only reported purification of active AMO was from expression of affinity-tagged AMO in N. halophilia, an ammonia oxidizing bacteria.16 To achieve this, a fusion protein was created, adding (3 × FLAG) tag to the C-terminus of amoA, enabling affinity purification of the complete protein complex.16 This purified AMO can be stabilized by the presence of detergent, although it is worth noting that the cryo-EM structures of isolated AMO do still show native lipids bound within the protein complex, suggesting their requirement for activity.16 The Cu-coordinating residues were identified as D136, H140, D146, H153 at CuC, and N207, H211, E218, and H225 for CuD, and preliminary mutagenesis studies have proceeded from this.16 To generate mutants, constructs of point-mutated amoC and FLAG-tagged amoA were generated and electroporated into N. halophilia.16 All attempts to generate D136A and H225A mutants did not yield viable cells, likely due to severe impacts on the ammonia-oxidation activity of the AMO protein and therefore, the central metabolism of the host.16 Mutations H140A, H153A, D146A, N207A, H211A, and E218A did yield both live cells and functional protein, after affinity purification.16 These mutants showed ∼30% lower ammonia oxidation activity than the wild type AMO and establishing these residues as a requirement for catalytic efficiency.16 Preliminary mutagenesis has taken place establishing the critical role of the D136 and H225 residues, potentially as targets for future mutagenesis work.
Another of the fundamental challenges regarding isolation of AMO is the hydrophobic nature of membrane proteins, which promotes aggregation and dissolution when removed from the lipid bilayer environment. Historically, this has been overcome by use of detergents, however, a variety of novel techniques, such as lipid bilayer nanodiscs, lipid bicelles, amphipols, and fluorinated surfactants have been employed in recent years, demonstrating wide-ranging success.184,185 As a final potential method of overcoming these challenges, it may be worthwhile to investigate the cell-free protein expression (CFPE) of AMO. Cell-free protein expression has been documented for a variety of membrane proteins to date.186,187 Notably, pMMO has been expressed and reconstituted into lipid nanodiscs using CFPE techniques.188 While this preparation did not yield pMMO capable of CH4 oxidation, possibly due to subtle misfoldings or a lack of hexagonal packing arrays, 2D class averages of negative stain micrographs demonstrate the formation of the heterotrimeric structure, demonstrating the potential for access to functional CuMMOs via CFPE in the future.188
Conclusion
While AMO has proved to be particularly challenging to study given the difficulties in native protein purification and recombinant expression, rigorous study has allowed for continuous improvement in our understanding. Advances in techniques in membrane protein crystallography and cryo-EM have allowed for the recent structure of AMO in native membranes, alongside a variety of structural data for comparison with pMMO. To date, the vast majority of studies of AMO have required whole-cell assays, given that AMO is particularly fragile outside of the native organism. Despite this, it has been observed that AMO has a broad substrate scope not observed in pMMO, and the differences between the elusive active site of these two homologs may yield great insight into nature's fundamental ability to selectively hydroxylate a given substrate. The differences between bAMO and aAMO are likely affiliated with the recently discovered amoX, amoY, and amoZ subunits, which are unique to archaea. These differences may also be cause for the difference in kinetic parameters regarding the oxidation of NH3 between AOA and AOB, giving insight into niche partition in nature. Given the poor energetic yields of NH3 as fuel for primary metabolism, understanding how obligatory chemolithoautotrophs survive on uncommon fuel can inform understanding of metabolism and global nutrient cycles. Given the significant challenges remaining in the field, collaboration across microbiologists, ecologists, biochemists, and other specialists involved in the study of nitrification processes will be essential to uncovering the mechanistic and biological importance of AMO.Many avenues exist to progress understanding of AMO. In the absence of significant quantities of protein acquired from native biomass, an approach based on the expression and subsequent combination of individual subunits is desirable. One possibility is individual expression, purification, and characterization of amoA, amoB, and amoC, followed by the subsequent combination of these units. Said combination could proceed via sequential addition of subunits to either detergent micelles or lipid nanodiscs. To assist in the solubility of these constructs, development of fusion proteins containing the subunit and a solubility tag may be required. A major advantage of this approach is that the functional protein would be amenable to site directed mutagenesis, enabling examination of the role of individual Cu sites.
If only the C subunit can be isolated, this may still provide viable information. The amoC subunit, assuming it contains Cu, can be exposed to O2 and NH3 or CH4. Rapid freeze quench techniques paired with spectroscopic methods such as resonance Raman, EXAFS, and EPR, may allow observation of some on-target intermediates, providing fundamental insight into the electronic structure of the active site and mechanism of AMO.
Beyond this, the structure of aAMO has yet to be experimentally revealed. A cryo-EM structure, preferably in native membranes and from an archetypical AOA such as N. maritimus, would be a significant breakthrough, allowing for direct comparison of the aAMO and bAMO quaternary structure. Currently, many questions exist surrounding aAMO. Does aAMO form hexagonal packing arrays like bAMO and pMMO? How do amoX, amoY, and amoZ interact with the canonical subunits? Is AMO metalated at CuC (like bAMO) or CuD (like pMMO)? These investigations may also begin to pave the way for understanding of the unique ammonia consumption kinetics observed across AOA. Furthermore, an understanding of the relation between various AMOs may paint a clearer picture of common intermediates and metabolic processes across a variety of niches in the nitrogen cycle.
Computational methods
AlphaFold 3 was used to generate templated and untemplated models of the AMO, using sequences from AMO found in N. europaea ATCC 19718. In the case of the templated model, cryo-EM data from AMO in native membranes was used.84 These structures excluded the predicted signal sequence on the amoB subunit. Additionally, two CuII ions were added to the model. The templated model yielded scores of ipTM = 0.83 and pTM = 0.85, while the untemplated model scored ipTM = 0.85 and pTM = 0.87. Files generated by AlphaFold, and the associated PYMOL structures, can be found in the supplementary information.Author contributions
TCA, ALL, and KML wrote and revised the article.Conflicts of interest
There are no conflicts of interest to declare.Abbreviations
| aAMO | Archaeal ammonia monooxygenase |
| AMO | Ammonia monooxygenase |
| AOA | Ammonia oxidizing archaea |
| AOB | Ammonia oxidizing bacteria |
| ATP | Adenosine triphosphate |
| bAMO | Bacterial ammonia monooxygenase |
| BDFE | Bond dissociation free energy |
| CcO | Cytochrome c oxidase |
| CFPE | Cell-free protein expression |
| CuMMO | Copper membrane monooxygenase |
| Cryo-EM | Cryogenic electron microscopy |
| DβM | Dopamine β-monooxygenase |
| DMSO | Dimethylsulfoxide |
| EDTA | Ethylenediamine tetraacetic acid |
| ENDOR | Electron-nuclear double resonance |
| EPR | Electron paramagnetic resonance |
| EXAFS | Extended X-ray absorption fine structure |
| LPMO | Lytic polysaccharide monooxygenase |
| HMO | Hydrocarbon monooxygenase |
| HERFD | High-energy resolution fluorescence detected |
| HNO | Nitroxyl |
| N2OR | Nitrous oxide reductase |
| HAO | Hydroxylamine oxidoreductase |
| NCBI | National center for biotechnology information |
| NH3 | Ammonia |
| NH4+ | Ammonium |
| NO | Nitric oxide |
| NOB | Nitrite oxidizing bacteria |
| PAM | Peptidylglycine α-amidating monooxygenase |
| PHM | Peptidylglycine α-hydroxylating monooxygenase |
| pMMO | Particulate methane monooxygenase |
| ROS | Reactive oxygen species |
| sMMO | Soluble methane monooxygenase |
| SSN | Sequence similarity networks |
| TFB | 4,4,4-Trifluorobutanol |
| TFE | Trifluoroethanol |
| TM | Transmembrane |
| XANES | X-ray absorption near edge spectroscopy |
Data availability
The AlphaFold 3 model of the N. europaea AMO trimer is available from the corresponding author upon request.Supplementary information: tables of AMO substrates and inhibitors including reaction conditions under which these substances were tested as well as the ammonia and ammonium affinities of several nitrifying microorganisms. See DOI: https://doi.org/10.1039/d6sc01048b.
Acknowledgements
This review is dedicated to Harry B. Gray on the occasion of his 90th birthday. AMO should be right up his alley: AMO is a Cu protein that likely generates a reactive metal–oxygen intermediate to drive a reaction important to energy transduction that most certainly involves finely tuned electron transfer pathways. Perhaps most in line with Harry Gray's scientific legacy, however, is one key point—AMO is a really, really hard problem. Harry would never consider a problem too hard to try to solve. Work on AMO in the Lancaster laboratory on nitrification has been generously supported by the US Department of Energy Office of Science under grant number DE-SC0021003 as well as NIGMS under grant R35GM124908. We thank Colby Gekko for a critical read of this manuscript.References
- J. I. Prosser and G. W. Nicol, Environ. Microbiol., 2008, 10, 2931–2941 CrossRef CAS PubMed.
- M. G. Klotz and L. Y. Stein, FEMS Microbiol. Lett., 2008, 278, 146–156 Search PubMed.
- C. B. Walker, J. R. de la Torre, M. G. Klotz, H. Urakawa, N. Pinel, D. J. Arp, C. Brochier-Armanet, P. S. G. Chain, P. P. Chan, A. Gollabgir, J. Hemp, M. Hügler, E. A. Karr, M. Könneke, M. Shin, T. J. Lawton, T. Lowe, W. Martens-Habbena, L. A. Sayavedra-Soto, D. Lang, S. M. Sievert, A. C. Rosenzweig, G. Manning and D. A. Stahl, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 8818–8823 CrossRef CAS PubMed.
- S. Lücker, M. Wagner, F. Maixner, E. Pelletier, H. Koch, B. Vacherie, T. Rattei, J. S. S. Damsté, E. Spieck, D. Le Paslier and H. Daims, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 13479–13484 CrossRef PubMed.
- M. A. H. J. van Kessel, D. R. Speth, M. Albertsen, P. H. Nielsen, H. J. M. Op den Camp, B. Kartal, M. S. M. Jetten and S. Lücker, Nature, 2015, 528, 555–559 CrossRef CAS PubMed.
- H. Daims, E. V. Lebedeva, P. Pjevac, P. Han, C. Herbold, M. Albertsen, N. Jehmlich, M. Palatinszky, J. Vierheilig, A. Bulaev, R. H. Kirkegaard, M. von Bergen, T. Rattei, B. Bendinger, P. H. Nielsen and M. Wagner, Nature, 2015, 528, 504–509 CrossRef CAS PubMed.
- K. D. Kits, C. J. Sedlacek, E. V. Lebedeva, P. Han, A. Bulaev, P. Pjevac, A. Daebeler, S. Romano, M. Albertsen, L. Y. Stein, H. Daims and M. Wagner, Nature, 2017, 549, 269–272 CrossRef CAS PubMed.
- N. Vajrala, W. Martens-Habbena, L. A. Sayavedra-Soto, A. Schauer, P. J. Bottomley, D. A. Stahl and D. J. Arp, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 1006–1011 CrossRef CAS PubMed.
- P. L. Tavormina, V. J. Orphan, M. G. Kalyuzhnaya, M. S. M. Jetten and M. G. Klotz, Environ. Microbiol. Rep., 2011, 3, 91–100 CrossRef CAS PubMed.
- T. C. Hollocher, M. E. Tate and D. J. Nicholas, J. Biol. Chem., 1981, 256, 10834–10836 CrossRef CAS PubMed.
- S. A. Ensign, M. R. Hyman and D. J. Arp, J. Bacteriol., 1993, 175, 1971–1980 CrossRef CAS PubMed.
- D. F. McMillen and D. M. Golden, Annu. Rev. Phys. Chem., 1982, 33, 493–532 Search PubMed.
- N. F. Dummer, D. J. Willock, Q. He, M. J. Howard, R. J. Lewis, G. Qi, S. H. Taylor, J. Xu, D. Bethell, C. J. Kiely and G. J. Hutchings, Chem. Rev., 2023, 123, 6359–6411 CrossRef CAS PubMed.
- S. S.-F. Yu, L.-Y. Wu, K. H.-C. Chen, W.-I. Luo, D.-S. Huang and S. I. Chan, J. Biol. Chem., 2003, 278, 40658–40669 Search PubMed.
- V. C.-C. Wang, S. Maji, P. P.-Y. Chen, H. K. Lee, S. S.-F. Yu and S. I. Chan, Chem. Rev., 2017, 117, 8574–8621 CrossRef CAS PubMed.
- X. Yang, Z. Li, T.-Q. Mao, C. Ma, G.-H. Chen, H.-P. Dong and S.-F. Sui, Nat. Commun., 2025, 17, 508 CrossRef PubMed.
- S. Winogradsky, Ann. Inst. Pasteur., 1890,(4), 213–231 Search PubMed.
- R. D. Dua, B. Bhandari and D. J. Nicholas, FEBS Lett., 1979, 106, 401–404 CrossRef CAS PubMed.
- A. B. Hooper and K. R. Terry, J. Bacteriol., 1973, 115, 480–485 CrossRef CAS PubMed.
- M. R. Hyman and P. M. Wood, Arch. Microbiol., 1984, 137, 155–158 CrossRef CAS.
- T. C. Hollocher, S. Kumar and D. J. Nicholas, J. Bacteriol., 1982, 149, 1013–1020 CrossRef CAS PubMed.
- Q. H. Tran and G. Unden, Eur. J. Biochem., 1998, 251, 538–543 CrossRef CAS PubMed.
- R. González-Cabaleiro, T. P. Curtis and I. D. Ofiţeru, Water Res., 2019, 154, 238–245 CrossRef PubMed.
- J. D. Caranto and K. M. Lancaster, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 8217–8222 CrossRef CAS PubMed.
- A. E. Santoro, C. Buchwald, M. R. McIlvin and K. L. Casciotti, Science, 2011, 333, 1282–1285 CrossRef CAS PubMed.
- M. Könneke, A. E. Bernhard, J. R. de la Torre, C. B. Walker, J. B. Waterbury and D. A. Stahl, Nature, 2005, 437, 543–546 CrossRef PubMed.
- M. B. Karner, E. F. DeLong and D. M. Karl, Nature, 2001, 409, 507–510 CrossRef CAS PubMed.
- S. Wang, J. Huang, Z. Wu, S. Li, X. Zhu, Y. Liu and G. Ji, Nat. Commun., 2025, 16, 3341 Search PubMed.
- X. S. Wan, L. Hou, S.-J. Kao, Y. Zhang, H.-X. Sheng, H. Shen, S. Tong, W. Qin and B. B. Ward, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2220697120 CrossRef CAS PubMed.
- E. T. Buitenhuis, P. Suntharalingam and C. Le Quéré, Biogeosciences, 2018, 15, 2161–2175 CrossRef CAS.
- A. Freing, D. W. R. Wallace and H. W. Bange, Philos. Trans. R. Soc. Lond. B Biol. Sci., 2012, 367, 1245–1255 CrossRef CAS PubMed.
- M. Könneke, D. M. Schubert, P. C. Brown, M. Hügler, S. Standfest, T. Schwander, L. Schada von Borzyskowski, T. J. Erb, D. A. Stahl and I. A. Berg, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 8239–8244 CrossRef PubMed.
- B. Bayer, R. L. Hansman, M. J. Bittner, B. E. Noriega-Ortega, J. Niggemann, T. Dittmar and G. J. Herndl, Environ. Microbiol., 2019, 21, 4062–4075 CrossRef CAS PubMed.
- A. K. Hawley, M. K. Nobu, J. J. Wright, W. E. Durno, C. Morgan-Lang, B. Sage, P. Schwientek, B. K. Swan, C. Rinke, M. Torres-Beltrán, K. Mewis, W.-T. Liu, R. Stepanauskas, T. Woyke and S. J. Hallam, Nat. Commun., 2017, 8, 1507 CrossRef PubMed.
- K. L. Adair and E. Schwartz, Microb. Ecol., 2008, 56, 420–426 CrossRef CAS PubMed.
- S. Leininger, T. Urich, M. Schloter, L. Schwark, J. Qi, G. W. Nicol, J. I. Prosser, S. C. Schuster and C. Schleper, Nature, 2006, 442, 806–809 CrossRef CAS PubMed.
- R. Bartossek, A. Spang, G. Weidler, A. Lanzen and C. Schleper, Front. Microbiol., 2012, 3, 208 CAS.
- A. E. Taylor, L. H. Zeglin, T. A. Wanzek, D. D. Myrold and P. J. Bottomley, ISME J., 2012, 6, 2024–2032 CrossRef CAS PubMed.
- L. Hink, C. Gubry-Rangin, G. W. Nicol and J. I. Prosser, ISME J., 2018, 12, 1084–1093 CrossRef CAS PubMed.
- A. E. Taylor, A. T. Giguere, C. M. Zoebelein, D. D. Myrold and P. J. Bottomley, ISME J., 2017, 11, 896–908 CrossRef CAS PubMed.
- H. J. Di, K. C. Cameron, J.-P. Shen, C. S. Winefield, M. O'Callaghan, S. Bowatte and J.-Z. He, FEMS Microbiol. Ecol., 2010, 72, 386–394 CrossRef CAS PubMed.
- M. Dreer, T. Pribasnig, L. H. Hodgskiss, Z.-H. Luo, F. Pozaric and C. Schleper, ISME J., 2025, 19, wraf182 CrossRef CAS PubMed.
- R. W. Voland, H. Wang, H. D. Abruña and K. M. Lancaster, Proc. Natl. Acad. Sci. U. S. A., 2025, 122, e2416971122 CrossRef CAS PubMed.
- G. F. Wells, H.-D. Park, C.-H. Yeung, B. Eggleston, C. A. Francis and C. S. Criddle, Environ. Microbiol., 2009, 11, 2310–2328 CrossRef CAS PubMed.
- T. Limpiyakorn, M. Fürhacker, R. Haberl, T. Chodanon, P. Srithep and P. Sonthiphand, Appl. Microbiol. Biotechnol., 2013, 97, 1425–1439 CrossRef CAS PubMed.
- Y. Bai, Q. Sun, D. Wen and X. Tang, FEMS Microbiol. Ecol., 2012, 80, 323–330 CrossRef CAS PubMed.
- T. Limpiyakorn, P. Sonthiphand, C. Rongsayamanont and C. Polprasert, Bioresour. Technol., 2011, 102, 3694–3701 CrossRef CAS PubMed.
- P. Srithep, P. Pornkulwat and T. Limpiyakorn, Environ. Sci. Pollut. Res., 2018, 25, 8676–8687 CrossRef CAS PubMed.
- J.-F. Gao, X. Luo, G.-X. Wu, T. Li and Y.-Z. Peng, Bioresour. Technol., 2013, 138, 285–296 CrossRef CAS PubMed.
- J. Gao, X. Luo, G. Wu, T. Li and Y. Peng, Appl. Microbiol. Biotechnol., 2014, 98, 3339–3354 CrossRef CAS PubMed.
- M. R. Hyman and P. M. Wood, Biochem. J., 1985, 227, 719–725 CrossRef CAS PubMed.
- O. Meyerhof, Pflueg. Arch. Eur. J. Physiol., 1917, 166, 240–280 CrossRef CAS.
- D. J. Arp, L. A. Sayavedra-Soto and N. G. Hommes, Arch. Microbiol., 2002, 178, 250–255 CrossRef CAS PubMed.
- D. A. Stahl and J. R. de la Torre, Annu. Rev. Microbiol., 2012, 66, 83–101 CrossRef CAS PubMed.
- K. M. Lancaster, J. D. Caranto, S. H. Majer and M. A. Smith, Joule, 2018, 2, 421–441 CrossRef CAS.
- L. H. Hodgskiss, M. Melcher, M. Kerou, W. Chen, R. I. Ponce-Toledo, S. N. Savvides, S. Wienkoop, M. Hartl and C. Schleper, ISME J., 2023, 17, 588–599 CrossRef CAS PubMed.
- A. F. El Sheikh, A. T. Poret-Peterson and M. G. Klotz, Appl. Environ. Microbiol., 2008, 74, 312–318 CrossRef CAS PubMed.
- O. S. Fisher, G. E. Kenney, M. O. Ross, S. Y. Ro, B. E. Lemma, S. Batelu, P. M. Thomas, V. C. Sosnowski, C. J. DeHart, N. L. Kelleher, T. L. Stemmler, B. M. Hoffman and A. C. Rosenzweig, Nat. Commun., 2018, 9, 4276 CrossRef PubMed.
- D. J. Arp, P. S. G. Chain and M. G. Klotz, Annu. Rev. Microbiol., 2007, 61, 503–528 CrossRef CAS PubMed.
- A. F. El Sheikh and M. G. Klotz, Environ. Microbiol., 2008, 10, 3026–3035 CrossRef CAS PubMed.
- C. E. Dann, J.-C. Hsieh, A. Rattner, D. Sharma, J. Nathans and D. J. Leahy, Nature, 2001, 412, 86–90 CrossRef CAS PubMed.
- H. McTavish, J. A. Fuchs and A. B. Hooper, J. Bacteriol., 1993, 175, 2436–2444 CrossRef CAS PubMed.
- D. J. Bergmann and A. B. Hooper, Biochem. Biophys. Res. Commun., 1994, 204, 759–762 CrossRef CAS PubMed.
- A. J. Holmes, A. Costello, M. E. Lidstrom and J. C. Murrell, FEMS Microbiol. Lett., 1995, 132, 203–208 CrossRef CAS PubMed.
- M. G. Klotz and J. M. Norton, Gene, 1995, 163, 159–160 CrossRef CAS PubMed.
- A. Burns, M. Kerou, D. Zak, C. Schleper, T. Urich and H. Wang, Appl. Soil Ecol., 2025, 214, 106395 Search PubMed.
- A. M. Duff, L.-M. Zhang and C. J. Smith, Sci. Rep., 2017, 7, 13200 CrossRef PubMed.
- J. Johnston, Z. Du and S. Behrens, Microbiol. Spectr., 2023, 11, e0257122 CrossRef PubMed.
- J. Li, D. B. Nedwell, J. Beddow, A. J. Dumbrell, A. McKew Boyd, E. L. Thorpe and W. Corinne, Appl. Environ. Microbiol., 2015, 81, 159–165 CrossRef PubMed.
- M. Li, H. He, T. Mi and Y. Zhen, Sci. Total Environ., 2022, 825, 153972 Search PubMed.
- M. Pester, T. Rattei, S. Flechl, A. Gröngröft, A. Richter, J. Overmann, B. Reinhold-Hurek, A. Loy and M. Wagner, Environ. Microbiol., 2012, 14, 525–539 CrossRef CAS PubMed.
- H. Wang, A. Bagnoud, R. Ponce-Toledo, M. Kerou, M. Weil, C. Schleper and T. Urich, mSystems, 2021, 6 DOI:10.1128/msystems.00546-21.
- M. G. Klotz, J. Alzerreca and J. M. Norton, FEMS Microbiol. Lett., 1997, 150, 65–73 CrossRef CAS PubMed.
- L. Y. Stein, L. A. Sayavedra-Soto, N. G. Hommes and D. J. Arp, FEMS Microbiol. Lett., 2000, 192, 163–168 CrossRef CAS PubMed.
- J. M. Norton, J. M. Low and M. G. Klotz, FEMS Microbiol. Lett., 1996, 139, 181–188 CrossRef CAS PubMed.
- N. G. Hommes, L. A. Sayavedra-Soto and D. J. Arp, J. Bacteriol., 1998, 180, 3353–3359 CrossRef CAS PubMed.
- N. G. Hommes, L. A. Sayavedra-Soto and D. J. Arp, J. Bacteriol., 2001, 183, 1096–1100 CrossRef CAS PubMed.
- P. M. Berube, R. Samudrala and D. A. Stahl, J. Bacteriol., 2007, 189, 3935–3944 CrossRef CAS PubMed.
- W. Qin, S. A. Amin, R. A. Lundeen, K. R. Heal, W. Martens-Habbena, S. Turkarslan, H. Urakawa, K. C. Costa, E. L. Hendrickson, T. Wang, D. A. Beck, S. M. Tiquia-Arashiro, F. Taub, A. D. Holmes, N. Vajrala, P. M. Berube, T. M. Lowe, J. W. Moffett, A. H. Devol, N. S. Baliga, D. J. Arp, L. A. Sayavedra-Soto, M. Hackett, E. V. Armbrust, A. E. Ingalls and D. A. Stahl, ISME J., 2018, 12, 508–519 CrossRef CAS PubMed.
- P. M. Berube and D. A. Stahl, J. Bacteriol., 2012, 194, 3448–3456 CrossRef CAS PubMed.
- M. Kerou, P. Offre, L. Valledor, S. S. Abby, M. Melcher, M. Nagler, W. Weckwerth and C. Schleper, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, E7937–E7946 CrossRef CAS PubMed.
- A. H. Treusch, S. Leininger, A. Kletzin, S. C. Schuster, H.-P. Klenk and C. Schleper, Environ. Microbiol., 2005, 7, 1985–1995 CrossRef CAS PubMed.
- N. A. Ahlgren, C. A. Fuchsman, G. Rocap and J. A. Fuhrman, ISME J., 2019, 13, 618–631 CrossRef CAS PubMed.
- F. J. Tucci and A. C. Rosenzweig, Proc. Natl. Acad. Sci. U. S. A., 2025, 122, e2417993121 CrossRef CAS PubMed.
- F. J. Tucci, M. B. Ho, A. A. B. Turner, L. Y. Stein, B. M. Hoffman and A. C. Rosenzweig, Chem. Sci., 2026, 17, 4470–4477 RSC.
- R. L. Lieberman and A. C. Rosenzweig, Nature, 2005, 434, 177–182 CrossRef CAS PubMed.
- S. Y. Ro, M. O. Ross, Y. W. Deng, S. Batelu, T. J. Lawton, J. D. Hurley, T. L. Stemmler, B. M. Hoffman and A. C. Rosenzweig, J. Biol. Chem., 2018, 293, 10457–10465 Search PubMed.
- S. M. Smith, S. Rawat, J. Telser, B. M. Hoffman, T. L. Stemmler and A. C. Rosenzweig, Biochemistry, 2011, 50, 10231–10240 CrossRef CAS PubMed.
- A. S. Hakemian, K. C. Kondapalli, J. Telser, B. M. Hoffman, T. L. Stemmler and A. C. Rosenzweig, Biochemistry, 2008, 47, 6793–6801 CrossRef CAS PubMed.
- C. W. Koo, F. J. Tucci, Y. He and A. C. Rosenzweig, Science, 2022, 375, 1287–1291 CrossRef CAS PubMed.
- S. Sirajuddin, D. Barupala, S. Helling, K. Marcus, T. L. Stemmler and A. C. Rosenzweig, J. Biol. Chem., 2014, 289, 21782–21794 CrossRef PubMed.
- W.-H. Chang, H.-H. Lin, I.-K. Tsai, S.-H. Huang, S.-C. Chung, I.-P. Tu, S. S.-F. Yu and S. I. Chan, J. Am. Chem. Soc., 2021, 143, 9922–9932 CrossRef CAS PubMed.
- Y. Zhu, C. W. Koo, C. K. Cassidy, M. C. Spink, T. Ni, L. C. Zanetti-Domingues, B. Bateman, M. L. Martin-Fernandez, J. Shen, Y. Sheng, Y. Song, Z. Yang, A. C. Rosenzweig and P. Zhang, Nat. Commun., 2022, 13, 5221 CrossRef CAS PubMed.
- F. J. Tucci, R. J. Jodts, B. M. Hoffman and A. C. Rosenzweig, Nat. Catal., 2023, 6, 1194–1204 CrossRef CAS PubMed.
- F. J. Tucci and A. C. Rosenzweig, Chem. Rev., 2024, 124, 1288–1320 CrossRef CAS PubMed.
- A. von Kügelgen, C. K. Cassidy, S. van Dorst, L. L. Pagani, C. Batters, Z. Ford, J. Löwe, V. Alva, P. J. Stansfeld and T. A. M. Bharat, Nature, 2024, 630, 230–236 CrossRef PubMed.
- T. J. Lawton, J. Ham, T. Sun and A. C. Rosenzweig, Proteins: Struct., Funct., Bioinf., 2014, 82, 2263–2267 CrossRef CAS PubMed.
- R. Balasubramanian, S. M. Smith, S. Rawat, L. A. Yatsunyk, T. L. Stemmler and A. C. Rosenzweig, Nature, 2010, 465, 115–119 CrossRef CAS PubMed.
- H. J. Kim, J. Huh, Y. W. Kwon, D. Park, Y. Yu, Y. E. Jang, B.-R. Lee, E. Jo, E. J. Lee, Y. Heo, W. Lee and J. Lee, Nat. Catal., 2019, 2, 342–353 CrossRef CAS.
- M. O. Ross, F. MacMillan, J. Wang, A. Nisthal, T. J. Lawton, B. D. Olafson, S. L. Mayo, A. C. Rosenzweig and B. M. Hoffman, Science, 2019, 364, 566–570 CrossRef CAS PubMed.
- S. S.-F. Yu, K. H.-C. Chen, M. Y.-H. Tseng, W. Yane-Shih, T. Chiu-Feng, C. Yu-Ju, H. Ded-Shih and S. I. Chan, J. Bacteriol., 2003, 185, 5915–5924 CrossRef CAS PubMed.
- S. I. Chan, V. C.-C. Wang, J. C.-H. Lai, S. S.-F. Yu, P. P.-Y. Chen, K. H.-C. Chen, C.-L. Chen and M. K. Chan, Angew Chem., Int. Ed. Engl., 2007, 46, 1992–1994 CrossRef CAS PubMed.
- S.-C. Hung, C.-L. Chen, K. H.-C. Chen, S. S.-F. Yu and S. I. Chan, J. Chin. Chem. Soc., 2004, 51, 1229–1244 CrossRef CAS.
- S. I. Chan, V. C.-C. Wang, P. P.-Y. Chen and S. S.-F. Yu, J. Chin. Chem. Soc., 2022, 69, 1147–1158 CrossRef CAS.
- S. S.-F. Yu, C.-Z. Ji, Y. P. Wu, T.-L. Lee, C.-H. Lai, S.-C. Lin, Z.-L. Yang, V. C.-C. Wang, K. H.-C. Chen and S. I. Chan, Biochemistry, 2007, 46, 13762–13774 CrossRef CAS PubMed.
- S. Gilch, O. Meyer and I. Schmidt, Biol. Chem., 2009, 390, 863–873 CrossRef CAS PubMed.
- S. Gilch, O. Meyer and I. Schmidt, BioMetals, 2010, 23, 613–622 CrossRef CAS PubMed.
- D. C. Tsang and I. Suzuki, Can. J. Biochem., 1982, 60, 1018–1024 CrossRef CAS PubMed.
- J. A. Zahn and A. A. DiSpirito, J. Bacteriol., 1996, 178, 1018–1029 CrossRef CAS PubMed.
- J. A. Zahn, D. M. Arciero, A. B. Hooper and A. A. DiSpirito, FEBS Lett., 1996, 397, 35–38 CrossRef CAS PubMed.
- M. Takeguchi, M. Ohashi and I. Okura, BioMetals, 1999, 12, 123–129 CrossRef CAS.
- R. J. Jodts, M. O. Ross, C. W. Koo, P. E. Doan, A. C. Rosenzweig and B. M. Hoffman, J. Am. Chem. Soc., 2021, 143, 15358–15368 CrossRef CAS PubMed.
- F. Sabbadin, S. Urresti, B. Henrissat, A. O. Avrova, L. R. J. Welsh, P. J. Lindley, M. Csukai, J. N. Squires, P. H. Walton, G. J. Davies, N. C. Bruce, S. C. Whisson and S. J. McQueen-Mason, Science, 2021, 373, 774–779 CrossRef CAS PubMed.
- J. Ø. Ipsen, M. Hallas-Møller, S. Brander, L. Lo Leggio and K. S. Johansen, Biochem. Soc. Trans., 2021, 49, 531–540 CrossRef CAS PubMed.
- P. H. Walton, G. J. Davies, D. E. Diaz and J. P. Franco-Cairo, FEBS Lett., 2023, 597, 485–494 CrossRef CAS PubMed.
- G. R. Hemsworth, E. J. Taylor, R. Q. Kim, R. C. Gregory, S. J. Lewis, J. P. Turkenburg, A. Parkin, G. J. Davies and P. H. Walton, J. Am. Chem. Soc., 2013, 135, 6069–6077 CrossRef CAS PubMed.
- G. E. Cutsail, M. O. Ross, A. C. Rosenzweig and S. DeBeer, Chem. Sci., 2021, 12, 6194–6209 RSC.
- A. Labourel, K. E. H. Frandsen, F. Zhang, N. Brouilly, S. Grisel, M. Haon, L. Ciano, D. Ropartz, M. Fanuel, F. Martin, D. Navarro, M.-N. Rosso, T. Tandrup, B. Bissaro, K. S. Johansen, A. Zerva, P. H. Walton, B. Henrissat, L. L. Leggio and J.-G. Berrin, Nat. Chem. Biol., 2020, 16, 345–350 CrossRef CAS PubMed.
- S. Garcia-Santamarina, C. Probst, R. A. Festa, C. Ding, A. D. Smith, S. E. Conklin, S. Brander, L. N. Kinch, N. V. Grishin, K. J. Franz, P. Riggs-Gelasco, L. Lo Leggio, K. S. Johansen and D. J. Thiele, Nat. Chem. Biol., 2020, 16, 337–344 CrossRef CAS PubMed.
- T. Kruse, C. M. Ratnadevi, H.-A. Erikstad and N.-K. Birkeland, BMC Genom., 2019, 20, 642 CrossRef PubMed.
- H. J. M. Op den Camp, T. Islam, M. B. Stott, H. R. Harhangi, A. Hynes, S. Schouten, M. S. M. Jetten, N.-K. Birkeland, A. Pol and P. F. Dunfield, Environ. Microbiol. Rep., 2009, 1, 293–306 CrossRef CAS PubMed.
- N. V. Coleman, N. B. Le, M. A. Ly, H. E. Ogawa, V. McCarl, N. L. Wilson and A. J. Holmes, ISME J., 2012, 6, 171–182 CrossRef CAS PubMed.
- E. F. Liew, D. Tong, N. V. Coleman and A. J. Holmes, Microbiology, 2014, 160, 1267–1277 CrossRef CAS PubMed.
- F. Musiani, V. Broll, E. Evangelisti and S. Ciurli, JBIC, J. Biol. Inorg. Chem., 2020, 25, 995–1007 CrossRef CAS PubMed.
- L. Y. Stein and D. J. Arp, Appl. Environ. Microbiol., 1998, 64, 4098–4102 CrossRef CAS PubMed.
- N. Oberg, R. Zallot and J. A. Gerlt, Comput. Resour. Mol. Biol., 2023, 435, 168018 CAS.
- R. Zallot, N. Oberg and J. A. Gerlt, Biochemistry, 2019, 58, 4169–4182 CrossRef CAS PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O'Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- P. M. H. Kroneck, W. A. Antholine, J. Riester and W. G. Zumft, FEBS Lett., 1989, 248, 212–213 CrossRef CAS PubMed.
- J. M. Charnock, A. Dreusch, H. Körner, F. Neese, J. Nelson, A. Kannt, H. Michel, C. D. Garner, P. M. H. Kroneck and W. G. Zumft, Eur. J. Biochem., 2000, 267, 1368–1381 CrossRef CAS PubMed.
- P. M. H. Kroneck, W. E. Antholine, D. H. W. Kastrau, G. Buse, G. C. M. Steffens and W. G. Zumft, FEBS Lett., 1990, 268, 274–276 CrossRef CAS.
- C. Greenwood, B. C. Hill, D. Barber, D. G. Eglinton and A. J. Thomson, Biochem. J., 1983, 215, 303–316 CrossRef CAS PubMed.
- S. I. Gorelsky, X. Xie, Y. Chen, J. A. Fee and E. I. Solomon, J. Am. Chem. Soc., 2006, 128, 16452–16453 CrossRef CAS PubMed.
- A. J. Leguto, M. A. Smith, M. N. Morgada, U. A. Zitare, D. H. Murgida, K. M. Lancaster and A. J. Vila, J. Am. Chem. Soc., 2019, 141, 1373–1381 CrossRef CAS.
- F. Neese, W. G. Zumft, W. E. Antholine and P. M. H. Kroneck, J. Am. Chem. Soc., 1996, 118, 8692–8699 CrossRef CAS.
- M. O. Ross, O. S. Fisher, M. N. Morgada, M. D. Krzyaniak, M. R. Wasielewski, A. J. Vila, B. M. Hoffman and A. C. Rosenzweig, J. Am. Chem. Soc., 2019, 141, 4678–4686 CrossRef CAS PubMed.
- M. Prudêncio, A. S. Pereira, P. Tavares, S. Besson, I. Cabrito, K. Brown, B. Samyn, B. Devreese, J. Van Beeumen, F. Rusnak, G. Fauque, J. J. G. Moura, M. Tegoni, C. Cambillau and I. Moura, Biochemistry, 2000, 39, 3899–3907 CrossRef PubMed.
- C. E. Slutter, D. Sanders, P. Wittung, B. G. Malmström, R. Aasa, J. H. Richards, H. B. Gray and J. A. Fee, Biochemistry, 1996, 35, 3387–3395 CrossRef CAS PubMed.
- M. R. Hyman and P. M. Wood, Biochem. J., 1983, 212, 31–37 CrossRef CAS PubMed.
- R. D. Jones and R. Y. Morita, Appl. Environ. Microbiol., 1983, 45, 401–410 CrossRef CAS.
- P. A. Voysey and P. M. Wood, Microbiology, 1987, 133, 283–290 CAS.
- M. R. Hyman, I. B. Murton and D. J. Arp, Appl. Environ. Microbiol., 1988, 54, 3187–3190 CrossRef CAS PubMed.
- M. R. Hyman, A. W. Sansome-Smith, J. H. Shears and P. M. Wood, Arch. Microbiol., 1985, 143, 302–306 CrossRef CAS.
- W. K. Keener and D. J. Arp, Appl. Environ. Microbiol., 1994, 60, 1914–1920 CrossRef CAS PubMed.
- M. E. Rasche, M. R. Hyman and D. J. Arp, Appl. Environ. Microbiol., 1991, 57, 2986–2994 CrossRef CAS PubMed.
- K. N. Duddleston, P. J. Bottomley, A. J. Porter and D. J. Arp, Appl. Environ. Microbiol., 2000, 66, 2726–2731 CrossRef CAS PubMed.
- L. Y. Juliette, M. R. Hyman and D. J. Arp, Appl. Environ. Microbiol., 1993, 59, 3718–3727 CrossRef CAS.
- I. Suzuki, U. Dular and S. C. Kwok, J. Bacteriol., 1974, 120, 556–558 CrossRef CAS PubMed.
- W. K. Keener and D. J. Arp, Appl. Environ. Microbiol., 1993, 59, 2501–2510 CrossRef CAS PubMed.
- M.-Y. Jung, C. J. Sedlacek, K. D. Kits, A. J. Mueller, S.-K. Rhee, L. Hink, G. W. Nicol, B. Bayer, L. Lehtovirta-Morley, C. Wright, J. R. de la Torre, C. W. Herbold, P. Pjevac, H. Daims and M. Wagner, ISME J., 2022, 16, 272–283 CrossRef CAS PubMed.
- W. Martens-Habbena, P. M. Berube, H. Urakawa, J. R. de la Torre and D. A. Stahl, Nature, 2009, 461, 976–979 CrossRef CAS PubMed.
- W. Qin, K. R. Heal, R. Ramdasi, J. N. Kobelt, W. Martens-Habbena, A. D. Bertagnolli, S. A. Amin, C. B. Walker, H. Urakawa, M. Könneke, A. H. Devol, J. W. Moffett, E. V. Armbrust, G. J. Jensen, A. E. Ingalls and D. A. Stahl, Int. J. Syst. Evol. Microbiol., 2017, 67, 5067–5079 CrossRef.
- S.-V. Albers and B. H. Meyer, Nat. Rev. Microbiol., 2011, 9, 414–426 CrossRef CAS PubMed.
- P.-N. Li, J. Herrmann, S. Wakatsuki and H. van den Bedem, J. Phys. Chem. B, 2019, 123, 10331–10342 CrossRef CAS PubMed.
- T. Nakagawa and D. A. Stahl, Appl. Environ. Microbiol., 2013, 79, 6911–6916 CrossRef CAS PubMed.
- P.-N. Li, J. Herrmann, B. B. Tolar, F. Poitevin, R. Ramdasi, J. R. Bargar, D. A. Stahl, G. J. Jensen, C. A. Francis, S. Wakatsuki and H. van den Bedem, ISME J., 2018, 12, 2389–2402 CrossRef CAS PubMed.
- D. D. S. Smith and H. Dalton, Eur. J. Biochem., 1989, 182, 667–671 CrossRef CAS PubMed.
- K. J. Burrows, A. Cornish, D. Scott and I. J. Higgins, Microbiology, 1984, 130, 3327–3333 Search PubMed.
- A. Miyaji, T. Miyoshi, K. Motokura and T. Baba, Biotechnol. Lett., 2011, 33, 2241–2246 CrossRef CAS PubMed.
- M. Baani and W. Liesack, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 10203–10208 CrossRef CAS PubMed.
- L. Sonny and J. D. Semrau, Appl. Environ. Microbiol., 1998, 64, 1106–1114 CrossRef PubMed.
- A. E. Taylor, K. Taylor, B. Tennigkeit, M. Palatinszky, M. Stieglmeier, D. D. Myrold, C. Schleper, M. Wagner and P. J. Bottomley, Appl. Environ. Microbiol., 2015, 81, 1942–1948 CrossRef CAS PubMed.
- D. Sakoula, A. Schatteman, P. Blom, M. S. M. Jetten, M. A. H. J. van Kessel, L. Lehtovirta-Morley and S. Lücker, ISME Commun., 2024, 4, ycae092 CrossRef PubMed.
- A. B. McQuarters, M. W. Wolf, A. P. Hunt and N. Lehnert, Angew Chem., Int. Ed. Engl., 2014, 53, 4750–4752 CrossRef CAS PubMed.
- J. T. Groves and G. A. McClusky, J. Am. Chem. Soc., 1976, 98, 859–861 CrossRef CAS.
- J. T. Groves, J. Chem. Educ., 1985, 62, 928 CrossRef CAS.
- M. T. Green, J. H. Dawson and H. B. Gray, Science, 2004, 304, 1653–1656 CrossRef CAS PubMed.
- J. J. Warren, T. A. Tronic and J. M. Mayer, Chem. Rev., 2010, 110, 6961–7001 CrossRef CAS PubMed.
- C. M. Silva, I. C. Dias and J. R. Pliego, Org. Biomol. Chem., 2015, 13, 6217–6224 RSC.
- K. J. M. Lundgren, L. Cao, M. Torbjörnsson, E. D. Hedegård and U. Ryde, Dalton Trans., 2025, 54, 3141–3156 RSC.
- J. Y. Lee and K. D. Karlin, Biocatal. Biotransform., 2015, 25, 184–193 CAS.
- E. I. Solomon, P. Chen, M. Metz, S.-K. Lee and A. E. Palmer, Angew. Chem., Int. Ed., 2001, 40, 4570–4590 CrossRef CAS.
- C. E. Elwell, N. L. Gagnon, B. D. Neisen, D. Dhar, A. D. Spaeth, G. M. Yee and W. B. Tolman, Chem. Rev., 2017, 117, 2059–2107 CrossRef CAS PubMed.
- S. Itoh, Biocatal. Biotransform., 2006, 10, 115–122 CAS.
- P. Chen and E. I. Solomon, J. Am. Chem. Soc., 2004, 126, 4991–5000 CrossRef CAS PubMed.
- J. P. Klinman, J. Biol. Chem., 2006, 281, 3013–3016 CrossRef CAS PubMed.
- M. C. Brenner and J. P. Klinman, Biochemistry, 1989, 28, 4664–4670 CrossRef CAS PubMed.
- R. J. Arias, E. F. Welch and N. J. Blackburn, Protein Sci., 2023, 32, e4615 CrossRef CAS PubMed.
- K. W. Rush, K. A. S. Eastman, E. F. Welch, V. Bandarian and N. J. Blackburn, J. Am. Chem. Soc., 2024, 146, 5074–5080 CrossRef CAS PubMed.
- J. S. Woertink, P. J. Smeets, M. H. Groothaert, M. A. Vance, B. F. Sels, R. A. Schoonheydt and E. I. Solomon, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 18908–18913 CrossRef CAS PubMed.
- J. K. Zorz, J. A. Kozlowski, L. Y. Stein, M. Strous and M. Kleiner, Front. Microbiol., 2018, 9, 938 CrossRef PubMed.
- E. French, J. A. Kozlowski, M. Mukherjee, G. Bullerjahn and A. Bollmann, Appl. Environ. Microbiol., 2012, 78, 5773–5780 CrossRef CAS PubMed.
- J.-L. Popot, Annu. Rev. Biochem., 2010, 79, 737–775 CrossRef CAS PubMed.
- U. H. N. Dürr, M. Gildenberg and A. Ramamoorthy, Chem. Rev., 2012, 112, 6054–6074 CrossRef PubMed.
- J.-L. Popot and D. M. Engelman, Biochemistry, 2016, 55, 5–18 CrossRef CAS PubMed.
- J.-L. Popot, Arch. Biochem. Biophys., 2014, 564, 314–326 CrossRef CAS PubMed.
- C. W. Koo, J. M. Hershewe, M. C. Jewett and A. C. Rosenzweig, ACS Synth. Biol., 2022, 11, 4009–4017 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |