Open Access Article
Open Access ArticleDeep profiling reveals coordinated remodeling of ganglioside metabolism in MCF-7 breast cancer cell line
Yichun
Wang†
a,
Gaoge
Sun†
 b,
Hang
Yin
b,
Hang
Yin
 *b and
Yu
Xia
*b and
Yu
Xia
 *a
*a
aMOE Key Laboratory of Bioorganic Phosphorus Chemistry & Chemical Biology, Department of Chemistry, Tsinghua University, Beijing 100084, China. E-mail: xiayu@mail.tsinghua.edu.cn
bState Key Laboratory of Membrane Biology, School of Pharmaceutical Sciences, Tsinghua-Peking Center for Life Sciences, Key Laboratory of Bioorganic Phosphorous Chemistry and Chemical Biology, Tsinghua University, Beijing 100084, China. E-mail: yin_hang@tsinghua.edu.cn
First published on 26th January 2026
Abstract
Gangliosides are vital cell membrane components whose metabolic dysregulation is implicated in various cancers. However, a systems-level understanding of their metabolism has been hindered by their structural complexity and low cellular abundance. Herein, we have developed a deep profiling workflow for gangliosides that integrates selective enrichment, liquid chromatography-ion mobility spectrometry, and isomer-resolved tandem mass spectrometry. This workflow enhances detection sensitivity 100-fold, enabling the identification of 391 ganglioside structures in a human breast adenocarcinoma cell line (MCF-7) at multiple structural levels. We further reveal coordinated remodeling of gangliosides in MCF-7 cancer cells, including shifts toward a-series glycans, increased incorporation of long-chain sphingosine bases, and altered C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location isomers. By integrating these lipidomic findings with targeted gene expression analysis and quantitative proteomics, we reconstruct a ganglioside biosynthetic network that delineates dysregulation across five key structural modules. This lipid-centric approach offers new insights into the metabolic reprogramming of gangliosides and holds potential for studying lipid metabolism in diverse diseases.
C location isomers. By integrating these lipidomic findings with targeted gene expression analysis and quantitative proteomics, we reconstruct a ganglioside biosynthetic network that delineates dysregulation across five key structural modules. This lipid-centric approach offers new insights into the metabolic reprogramming of gangliosides and holds potential for studying lipid metabolism in diverse diseases.
Introduction
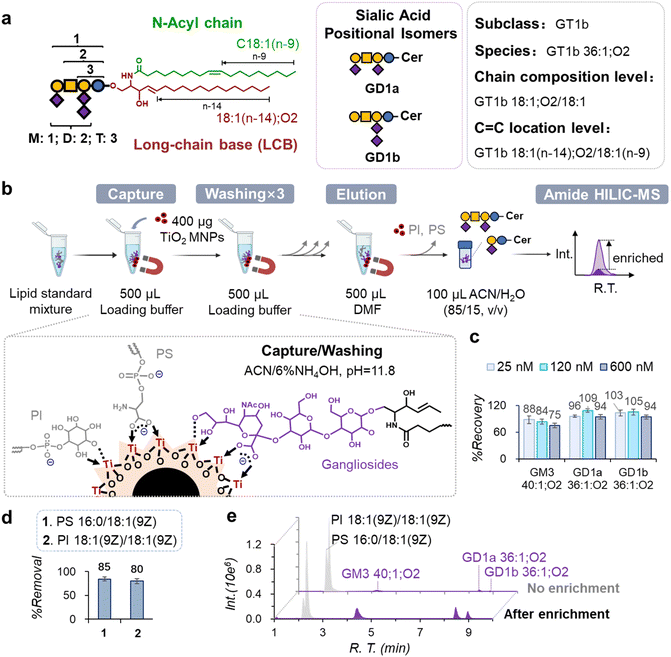
Gangliosides are a class of acidic glycosphingolipids, characterized by a sialoglycan headgroup linked to a ceramide moiety. Anchored on the outer leaflet of the plasma membrane, gangliosides are engaged in regulating diverse biological processes, including lipid raft formation,1 cell signaling,2 and cell–cell communication.3 Gangliosides are classified into subclasses according to the sequence of the neutral glycan core and the number, variants, and positions of sialic acids (Fig. 1a). Their structural diversity is further augmented by variations in the ceramide moiety, including differences in chain length, hydroxylation, and carbon–carbon double-bond (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) number and position. Cellular gangliosides are modulated by a complex metabolic network consisting of the synthesis, modification, and degradation of both glycan and ceramide moieties. Even minor perturbations within this network can drive tumor invasion,4 metastasis,5 and cancer stemness.6,7 For instance, MDA-MB-231 breast cancer cells exhibit overexpression of ST8SIA1, a GD3 synthase, leading to the accumulation of polysialylated gangliosides, including GD3 and GD2, which subsequently facilitate serum-independent cancer cell proliferation.8,9 GD2 also serves as a marker of CD44hiCD24lo breast cancer stem cells and as a therapeutic target,6,7 as GD2-CAR T cells effectively eliminate GD2+ tumors and prevent metastasis.10 These functions underscore gangliosides as key mediators in the “lipid rewiring” process characteristic of cancer.11,12
C) number and position. Cellular gangliosides are modulated by a complex metabolic network consisting of the synthesis, modification, and degradation of both glycan and ceramide moieties. Even minor perturbations within this network can drive tumor invasion,4 metastasis,5 and cancer stemness.6,7 For instance, MDA-MB-231 breast cancer cells exhibit overexpression of ST8SIA1, a GD3 synthase, leading to the accumulation of polysialylated gangliosides, including GD3 and GD2, which subsequently facilitate serum-independent cancer cell proliferation.8,9 GD2 also serves as a marker of CD44hiCD24lo breast cancer stem cells and as a therapeutic target,6,7 as GD2-CAR T cells effectively eliminate GD2+ tumors and prevent metastasis.10 These functions underscore gangliosides as key mediators in the “lipid rewiring” process characteristic of cancer.11,12
Despite these advances, the current understanding of the ganglioside metabolic network in cancer remains limited. The majority of research efforts have been focused on specific ganglioside subclasses or on particular glycosyltransferases, with the molecular structure of the ceramide moiety underexplored. Recent studies indicate that ceramide composition plays a regulatory role in modulating ganglioside function. For instance, the extent of ceramide unsaturation in GM1 affects its incorporation into membrane microdomains and modifies their signaling properties.13 Thus, detailed structural characterization of gangliosides is essential for elucidating their functional roles and mapping the metabolic regulatory network.
The inherent low abundance (0.1–5 mol% of total cellular lipids)14 and structural complexity of gangliosides present challenges for conventional lipidomic profiling methods,15–22 resulting in low molecular coverage and an inability to distinguish structural isomers. For example, Li et al. detected 147 ganglioside species in HeLa cells using reversed-phase liquid chromatography (RPLC)-tandem mass spectrometry (MS/MS).18 Hohenwallner et al. reported 33 gangliosides at the chain composition level from porcine brain using high-field asymmetric ion mobility spectrometry (FAIMS)-MS/MS.19 Vo et al. identified 159 species from human serum using RPLC-trapped ion mobility spectrometry (TIMS)-MS/MS.20 Nonetheless, these approaches do not capture deeper structural details, such as chain composition or C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C position, while RPLC- or ion mobility-based separations cannot distinguish sialic acid positional isomers like GD1a and GD1b (Fig. 1a).
C position, while RPLC- or ion mobility-based separations cannot distinguish sialic acid positional isomers like GD1a and GD1b (Fig. 1a).
A variety of isomer-resolved MS/MS techniques have been developed for profiling lipids at detailed structural levels.23,24 These include advanced ion activation methods, such as ozone-induced dissociation (OzID),25 ultraviolet photodissociation (UVPD),26 and electron impact excitation of ions from organics (EIEIO).27 Additionally, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C derivatization strategies coupled with MS/MS, such as the Paternò–Büchi (PB) reaction24,28 and aziridination,29–31 have shown considerable success. However, these advanced MS/MS methods have not been applied to ganglioside analysis.
C derivatization strategies coupled with MS/MS, such as the Paternò–Büchi (PB) reaction24,28 and aziridination,29–31 have shown considerable success. However, these advanced MS/MS methods have not been applied to ganglioside analysis.
Here, we report an integrated analytical workflow designed to overcome these limitations. Our approach builds upon a TiO2 magnetic nanoparticle (MNP)-based enrichment strategy previously established in our group for neutral glycosphingolipids,32,33 but tailored specifically for gangliosides. This new enrichment method is further integrated with a charge-tagging PB reaction and high-resolution amide hydrophilic interaction chromatography (HILIC)-trapped ion mobility spectrometry (TIMS). The workflow provides unprecedented profiling capability for gangliosides, enabling the identification of over 300 ganglioside structures at various structural levels in the human breast adenocarcinoma MCF-7 cell line, spanning a concentration range of three orders of magnitude. Furthermore, it facilitates the mapping of altered ganglioside metabolism in MCF-7 cells, encompassing enzymes involved in the biosynthesis of the five structural building blocks of gangliosides.
Methods
Shorthand notation of gangliosides
Gangliosides are annotated at multiple structural levels based on information from MS/MS experiments. The glycan moiety is abbreviated according to IUPAC nomenclature34 (e.g., GT1b). The ceramide portion is annotated following Lipid MAPS guidelines,35 reported at the species level (e.g., GT1b 36:1;O2), chain composition level (e.g., GT1b 18:1;O2/18:0;O), and the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C positional level using n-notation (e.g., GT1b 18:1(n-14);O2/24:1(n-9)). A detailed guide to this nomenclature system is provided in the SI.
C positional level using n-notation (e.g., GT1b 18:1(n-14);O2/24:1(n-9)). A detailed guide to this nomenclature system is provided in the SI.
Materials and chemicals
GM3 (bovine milk), SHexCer 18:1(Δ4);O2/24:1(15Z), PS 16:0/18:1(9Z), PI 18:1(9Z)/18:1(9Z), PG 16:0/18:1(9Z) and porcine brain total lipid extracts were purchased from Avanti Polar Lipids, Inc. (Alabaster, AL, USA). GD1a, GD1b mixtures (porcine brain) and GM1a 18:1;O2/16:0[D9] were purchased from Cayman Chemicals (Ann Arbor, Michigan, USA). Other chemicals were commercially sourced. Detailed information on the materials is provided in the SI. No unexpected or unusually high safety hazards were encountered.Extraction of gangliosides from cells
Gangliosides were extracted using a modified Folch method.15 Approximately 5.5 × 106 cells with 35 pmol of GM1a 18:1;O2/16:0[D9] were homogenized in 3 mL of CHCl3/MeOH (2:1, v/v) by ultrasonication for 30 min. After adding 600 µL of deionized water, the mixture was shaken at 3000 rpm for 10 min and centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 7 min. The upper layer was transferred to a new tube. The lower phase was re-extracted with 675 µL of MeOH and 600 µL of H2O following the same steps. The combined upper layers were dried under nitrogen.
000 rpm for 7 min. The upper layer was transferred to a new tube. The lower phase was re-extracted with 675 µL of MeOH and 600 µL of H2O following the same steps. The combined upper layers were dried under nitrogen.
Selective enrichment of gangliosides by TiO2 MNPs
Lipid standards (GM3 40:1;O2, GD1a 36:1;O2, GD1b 36:1;O2 at 1 µM; PI 18:1(9Z)/18:1(9Z) and PS 16:0/18:1(9Z) at 25 µM) were enriched using 400 µg of TiO2 MNPs. For gangliosides from cell extracts, lipids extracted from ∼5.5 × 106 cells were loaded onto 800 µg of TiO2 MNPs in methanol, dried, and incubated with loading buffer (ACN/NH4OH, 94:6, v/v). After 40 min of agitation, TiO2 MNPs were magnetically separated and washed four times with 500 µL of ACN/NH4OH (94:6, v/v). The gangliosides were eluted twice with 500 µL of DMF under 40 min of agitation, dried under nitrogen, filtered through a hydrophobic PTFE membrane and reconstituted in 100 µL of ACN/H2O (85:15, v/v) for analysis.Offline PB derivatization
Enriched gangliosides from 5.5 × 106 cells and 10 mM of 2-ccetylpyridine (2-AcPy) were dissolved in 100 µL of ACN/H2O (50:50, v/v). The solution was purged with nitrogen for 10 min to remove oxygen, then injected into a flow microreactor and UV-irradiated (254 nm) for 20 s. The reaction solution was collected for subsequent HILIC-MS/MS analysis without further purification. Further details are provided in the SI.Other experimental protocols
Detailed information on the TiO2 MNP synthesis, cell culture, RNA extraction, qRT-PCR, and label-free proteomics protocols are provided in the SI.Amide HILIC-TIMS-MS/MS
A Waters ACQUITY UPLC I-Class system (Waters, Milford, MA, USA) was used to separate the gangliosides on an ACQUITY Premier BEH Amide column (150 mm × 2.1 mm, 1.7 µm; Waters, Milford, MA, USA). HILIC-TIMS-MS/MS detection was performed on a hybrid trapped ion mobility-quadrupole time-of-flight mass spectrometer (timsTOF, Bruker Daltonics, Bremen, Germany) equipped with an ESI source. Data acquisition was carried out using otofControl software (version 5.0, Bruker Daltonics, Bremen, Germany). Detailed information on the amide HILIC separation and MS parameters is provided in the SI.Data Analysis
Data processing was conducted using Data Analysis 5.0 software (Bruker Daltonics, Bremen, Germany). Identification of the gangliosides was performed manually, utilizing retention time (R.T), accurate m/z, ion mobility (1/K0), and fragmentation rules in negative, positive ion mode MS/MS, and PB-MS/MS.Results and discussion
Selective enrichment of gangliosides by TiO2 MNPs
Previously, our group had developed an enrichment protocol employing TiO2 MNPs for neutral glycosphingolipids.32,33 The principle of enrichment is based on the multidentate coordination between Ti(IV) and the cis-diol groups of neutral glycosphingolipids under basic pH conditions, which are not favorable for PC and PE. However, we found that directly adapting this protocol resulted in poor ganglioside recovery (∼16%). This is likely caused by the higher hydrophilicity, negative charge, and increased steric hindrance associated with the complex glycan headgroups of gangliosides, compared to neutral glycosphingolipids. Using the modified Folch lipid extraction protocol,15 neutral glycosphingolipids remain in the organic phase with PC and PE as the major interfering phospholipids. As a comparison, gangliosides partition into the aqueous phase along with relatively polar phospholipids, such as phosphatidylinositol (PI) and phosphatidylserine (PS), owing to their strong anionic property and high polarity. In order to establish an optimized enrichment protocol, a mixture of lipid standards was used, mimicking a ganglioside extract from cells.36 The mixture contained 1 µM each of GM3 40:1;O2, GD1a 36:1;O2, and GD1b 36:1;O2, as well as 25 µM each of PI 18:1(9Z)/18:1(9Z) and PS 16:0/18:1(9Z). As the GM3, GD1a, and GD1b standards were provided as mixtures of different ceramide chain lengths, the concentrations of GM3 40:1;O2, GD1a 36:1;O2, and GD1b 36:1;O2 were only approximate. By using a basic pH loading buffer (6% aqueous NH3/94% ACN, pH = 11.8), gangliosides,37–41 PS,38,39 and PI42 are co-captured by 400 µg of TiO2 MNPs. Given that the aliphatic hydroxyl groups on glycans exhibit pKa values in the range 15 to 18,43 they engage in interactions with Ti(IV) through bidentate chelating mechanisms, as depicted in Fig. 1b. Conversely, the carboxylic acid moieties in gangliosides and the headgroup of PS are presumably deprotonated, facilitating their interaction with Ti(IV) via binuclear bidentate coordination. If the previously developed washing buffer is applied, substantial losses of gangliosides, as well as PS and PI, occur due to the high MeOH content (96% MeOH/4% aqueous, 20 mM NH4HCO3, pH = 10.2). One possible explanation is that the proton-donating ability of MeOH destabilizes lipid adhesion to TiO2 MNPs by weakening electrostatic and hydrogen-bonding interactions, while its high polarity facilitates lipid dissolution. Thus, optimized washing and elution conditions are needed to achieve selective enrichment of gangliosides over PS and PI. Considering the polarity and functional group differences between gangliosides and PS/PI, a-series of organic solvents were screened to select a suitable washing buffer (Fig. S1). We found that 6% aqueous NH3/94% ACN solvent was suitable for both loading and washing, as it did not elute gangliosides. For elution, dimethylformamide (DMF) provided the best performance (Fig. 1c). While the exact mechanism is unclear, it is likely that competitive coordination between DMF and Ti(IV)44 destabilizes interactions between gangliosides and TiO2, while the high polarity of DMF promotes the preferential solvation of the more polar gangliosides compared to PI and PS.We further evaluated the performance of the optimized enrichment protocol. Using 400 µg of TiO2 MNPs, we achieved 75–109% recovery for the ganglioside standards across three initial concentrations (25 nM, 120 nM, and 600 nM) (Fig. 1c), consistent across different ceramide chain lengths (C34–C42) and degrees of unsaturation (1–2) (Fig. S2). Over 80% of PS 16:0/18:1(9Z) and PI 18:1(9Z)/18:1(9Z) were successfully removed (Fig. 1d). This optimized enrichment protocol enabled a 10-fold increase in the maximal amount of gangliosides per injection, from 0.5 pmol to 5 pmol, for GM3 40:1;O2 (Fig. 1e).
Profiling of gangliosides at multiple structural levels in MCF-7 cells
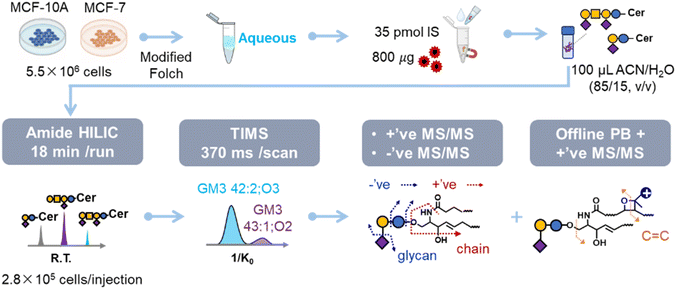
To profile gangliosides at multiple structural levels, we developed an integrated workflow combining TiO2 MNP-based selective enrichment, a charge-tagging PB reaction, and amide HILIC-TIMS-MS/MS (Scheme 1). Considering the important role that gangliosides play in breast cancer, we use MCF-7 cells as a model to evaluate the profiling capability of this workflow. Gangliosides were extracted from 5.5 × 106 MCF-7 cells using a modified Folch method with GM1a 18:1;O2/16:0[D9] (35 pmol) added as internal standard for normalization across samples, followed by enrichment using 800 µg of TiO2 MNPs. As a result, the maximum lipid amount for each injection was increased from 3.5 × 104 cells to 2.8 × 105 cells, leading to an eightfold increase (Fig. S3). The enriched gangliosides were analyzed using the following pipelines: (1) profiling on the subclass and species levels by HILIC-TIMS-MS/MS (precursor ions: [M − H]−, [M − 2H]2−); (2) profiling on the ceramide chain composition level by HILIC-MS/MS (precursor ions: [M + H]+, [M + NH4]+, [M + 2H]2+); (3) profiling on the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location level by performing offline charge-tagging PB derivatization followed by HILIC-MS/MS (precursor ions: [PBM + H]+, [PBM + 2H]2+).
C location level by performing offline charge-tagging PB derivatization followed by HILIC-MS/MS (precursor ions: [PBM + H]+, [PBM + 2H]2+).
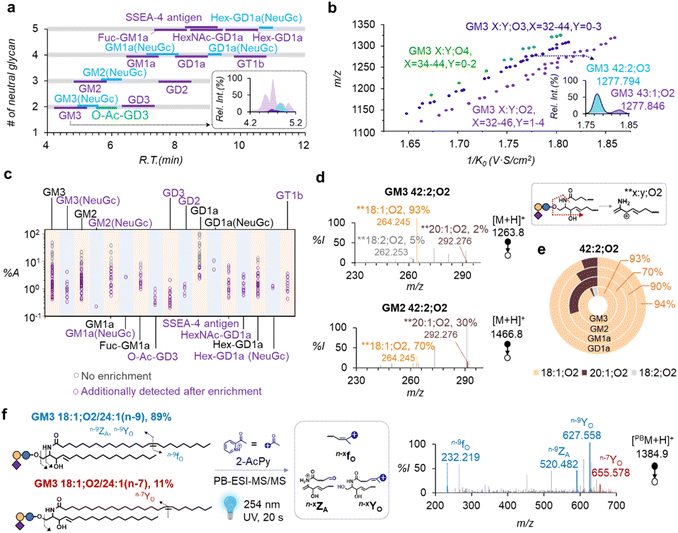
Amide HILIC was selected for its superior separation of ganglioside subclasses and its ability to resolve sialic acid positional isomers (e.g., GD1a/GD1b) compared with RPLC.18,20 It also yielded improved peak shapes over conventional diol-HILIC, attributed to stronger amide–glycan interactions (Fig. S4). Fig. 2a displays the retention behavior of 17 ganglioside subclasses from MCF-7 cells. In general, the retention time increased with the number of neutral glycans (2–6) and the number of sialic acid residues (1–3). The gangliosides containing NeuGc variants eluted ∼0.5 min later whereas NeuAc2 eluted ∼1 min earlier than the corresponding NeuAc counterparts. Several novel subclasses containing five neutral glycans were observed, including SSEA-4 antigen, HexNAc-GD1a, Hex-GD1a, and Fuc-GM1a. For each subclass, species with varying ceramide compositions co-eluted in a 1 min window, allowing limited chromatographic resolution of ceramide heterogeneity.
We integrated TIMS as an orthogonal separation method after amide HILIC to enhance the separation of co-eluting gangliosides based on the ceramide structure, i.e., X:Y;On, where X represents the number of carbons, Y indicates the degree of unsaturation, and On corresponds to the degree of hydroxylation. Taking GM3 as an example, the 73 species within this subclass occupy unique m/z vs. ion mobility, according to the number of hydroxylations (n = 2–4) in the ceramide moiety (Fig. 2b). Additionally, the ion mobility values (1/K0) increased monotonically with the total carbon number in ceramide (X = 33–46) and decreased with degree of unsaturation (Y = 0–4) or the number of hydroxylations (n = 2–4). Two isobaric species, i.e., GM3 42:2;O3 ([M − H]−, m/z 1277.794, 1/K0 = 1.775 V·S cm−2) and GM3 43:1;O2 ([M − H]−, m/z 1277.846, 1/K0 = 1.827 V·S cm−2), were resolved by TIMS, but were not fully separated by amide HILIC (inset of Fig. 2a). We also discovered previously unreported ceramide types, such as X:Y;O3 and X:Y;O4, in MCF-7 cells. Using this approach, we identified 17 subclasses and 319 species of gangliosides in MCF-7 cells, far exceeding the numbers without enrichment (7 subclasses and 76 species) (Fig. 2c). TiO2 MNP-based selective enrichment significantly boosts the detection of low-abundance species, expanding the dynamic range by two orders of magnitude. These low-abundance species also exhibit notable diversity in sialic acid types (NeuAc, NeuGc, NeuAc2), ceramide types (X:Y;O2, X:Y;O3, X:Y;O4), and degrees of unsaturation (0–4) (Fig. S5).
In order to characterize N-acyl and LCB composition in the ceramide moiety, we performed MS/MS on individual ganglioside species in positive ion mode. Tandem mass spectrometry employing collision-induced dissociation (MS2 CID) of the protonated molecular ions produces characteristic LCB fragment ions, viz. **x:y;O2 or *x:0;O3. Here, * signifies the loss of a water molecule; x and y represent the number of carbon atoms and the degree of unsaturation in the LCB moiety, respectively; and O2/3 indicates the degree of hydroxylation. TIMS was not activated in this process to reduce the degree of ion fragmentation before CID. This is because prolonged ion accumulation in TIMS results in elevated ion densities, inducing space charge effects that lead to ion losses, activation, and fragmentation.45 TIMS was not activated in this process to prevent space charge effects, which can cause ion losses and fragmentation that can degrade analytical utility. Fig. 2d shows the zoomed-in MS/MS spectra of GM3 42:2;O2 ([M + H]+, m/z 1263.8) and GM2 42:2;O2 ([M + H]+, m/z 1466.8). The high-quality data clearly suggest that GM3 42:2;O2 contains three chain composition isomers based on the detection of **18:1;O2, **18:2;O2, and **20:1;O2, whereas GM2 42:2;O2 has two (**18:1;O2 and **20:1;O2). The relative composition of each chain composition isomer was calculated by normalizing the corresponding LCB fragment to the sum of all LCB fragment ions in the same spectrum. We found that GM3 18:1;O2/24:1 accounted for 93% of GM3 42:2;O2 with minor contributions from GM3 18:2;O2/24:0 (5%) and GM3 20:1;O2/22:1 (2%). In contrast, GM2 18:1;O2/24:1 was most abundant (70%), followed by GM2 20:1;O2/22:1 (30%). Analysis from four subclasses, all consisting of the same ceramide moiety 42:2;O2 revealed that 18:1;O2 was the dominant LCB across all subclasses (70–94%). However, the compositions of minor components differed: GM3 contained higher 18:2;O2 (5%), whereas GM2, GM1a, and GD1a contained elevated 20:1;O2. Notably, 20:1;O2 reached 30% in GM2, compared with 2–9% in other subclasses. These findings highlight distinct chain composition patterns across subclasses (Fig. 2e).
We successfully resolved detailed chain composition information for 153 out of 319 species, yielding 220 distinct structures at the chain composition level. They can be categorized into seven types of LCBs, including four types of sphingosine (SPH, x:1;O2, x = 16–19), two types of sphingadiene (SPD, x:2;O2, x = 18, 20), and one type of phytosphingosine (PHS, 18:0;O3, Fig. S6a). Compared to the two commonly reported 18:1;O2 and 20:1;O2 in gangliosides,17 the gangliosides constituting the other LCBs are all of relatively low abundance, which can only be characterized by MS/MS after enrichment. N-acyl chains also exhibit diversity regarding chain lengths (C14–C26), degrees of unsaturation (0–3), and number of hydroxylations (0–1). Representative MS/MS spectra for hydroxylated fatty acids (hFA) are provided in Fig. S6b and c.
We found that more than 1/3 of gangliosides contained at least one C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond besides the C4–C5 C
C bond besides the C4–C5 C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C in sphingosine. Consequently, we conducted offline charge-tagging PB derivatization (Fig. S7) and performed targeted MS2 CID on the PB products in positive ion mode. We selected 2-acetylpyridine (2-AcPy) as the PB reagent, owing to its relatively high reaction yield, ability to enhance ionization efficiency of derivatized gangliosides in positive ion mode, and mitigation of in-source fragmentation. Because the C4–C5 C
C in sphingosine. Consequently, we conducted offline charge-tagging PB derivatization (Fig. S7) and performed targeted MS2 CID on the PB products in positive ion mode. We selected 2-acetylpyridine (2-AcPy) as the PB reagent, owing to its relatively high reaction yield, ability to enhance ionization efficiency of derivatized gangliosides in positive ion mode, and mitigation of in-source fragmentation. Because the C4–C5 C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C in the LCB reacts poorly in the PB reaction,32 PB-MS/MS produces mainly diagnostic ions of C
C in the LCB reacts poorly in the PB reaction,32 PB-MS/MS produces mainly diagnostic ions of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds other than at this location. In general, three types of C
C bonds other than at this location. In general, three types of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C diagnostic ions can be detected: n-xfO, n-xZA, and n-xYO. The generic structures of the diagnostic ions are shown in Fig. 2f. The superscript (n-x) denotes C
C diagnostic ions can be detected: n-xfO, n-xZA, and n-xYO. The generic structures of the diagnostic ions are shown in Fig. 2f. The superscript (n-x) denotes C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C position counting from the methyl end; the subscripts “A” and “O” indicate an aldehyde or olefin functional group at the C
C position counting from the methyl end; the subscripts “A” and “O” indicate an aldehyde or olefin functional group at the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond cleavage site, respectively. Fig. 2f shows the PB-MS2 CID spectrum of GM3 18:1;O2/24:1 ([PBM + H]+, m/z 1384.9) detected from MCF-7 cells. Diagnostic ions specific to n-9 (n-9fO, n-9ZA, n-9YO) and n-7 (n-7YO) C
C bond cleavage site, respectively. Fig. 2f shows the PB-MS2 CID spectrum of GM3 18:1;O2/24:1 ([PBM + H]+, m/z 1384.9) detected from MCF-7 cells. Diagnostic ions specific to n-9 (n-9fO, n-9ZA, n-9YO) and n-7 (n-7YO) C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C locations are of relatively high abundance. The relative compositions of n-9 and n-7 isomers were calculated as 89% and 11%, respectively, based on the relative abundances of YO ions. Using this approach, we identified 19 ganglioside structures at the C
C locations are of relatively high abundance. The relative compositions of n-9 and n-7 isomers were calculated as 89% and 11%, respectively, based on the relative abundances of YO ions. Using this approach, we identified 19 ganglioside structures at the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location level in MCF-7 cells, revealing the C
C location level in MCF-7 cells, revealing the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C positions in gangliosides for the first time. Monounsaturated N-acyls (C22–24) commonly consist of n-7, n-8, and n-9 isomers, while C24:2 is mainly contributed by an n–(6,9) C
C positions in gangliosides for the first time. Monounsaturated N-acyls (C22–24) commonly consist of n-7, n-8, and n-9 isomers, while C24:2 is mainly contributed by an n–(6,9) C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond configuration. Additionally, the SPD is determined as 18:2;O2(n-4,14). These structures have previously been reported in neutral glycosphingolipids from brain tissue, suggesting a potential homology of their fatty acid compositions.32
C bond configuration. Additionally, the SPD is determined as 18:2;O2(n-4,14). These structures have previously been reported in neutral glycosphingolipids from brain tissue, suggesting a potential homology of their fatty acid compositions.32
Our workflow presents the most extensive profiling of cellular gangliosides to date, documenting 391 distinct structures of gangliosides, encompassing 17 subclasses and 319 species, with 220 structures having defined chain compositions and 19 with specified C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C locations. Our findings not only surpass previous cellular ganglioside coverage, which reported up to ∼150 species but also extend structural resolution beyond the species level.18,21 Experimental details, including retention time, m/z, 1/K0, and relative abundance of identified ganglioside structures in MCF-7 cells, are provided in Tables S1–S3.
C locations. Our findings not only surpass previous cellular ganglioside coverage, which reported up to ∼150 species but also extend structural resolution beyond the species level.18,21 Experimental details, including retention time, m/z, 1/K0, and relative abundance of identified ganglioside structures in MCF-7 cells, are provided in Tables S1–S3.
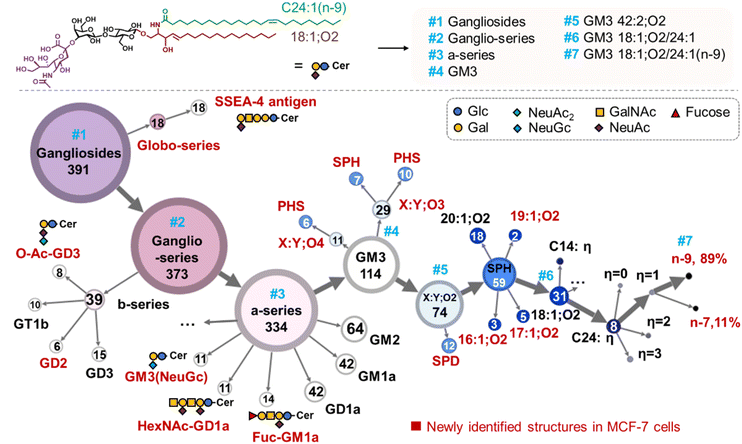
The rich molecular information allows us to delineate a top-down structural atlas of the gangliosides in MCF-7 cells in seven layers (Fig. 3). The atlas provides a visualization of the connections and divergences amongst individual structure layers, effectively reflecting the metabolic pathways of gangliosides. For instance, the second structural layer (#2) is clustered into ganglio- and globo-series according to the identity of the neutral glycan core. Starting from the ganglio-series, it further diverges into 0- and a-series (layer #3), defined by the number of sialo groups on the innermost galactose. The a-series contains 12 subclasses (layer #4) according to the numbers of neutral glycans and sialic acids, such as GM3, GM2, GM1a, and GD1a Fuc-GD1a and GM3(NeuGc). Each subclass is further categorized by the number of hydroxylations in the ceramide moiety (layer #5), such as GM3 X:Y;On (n = 2, 3, 4). Within GM3 X:Y;O2, the structures are grouped based on the types of LCB, viz. SPH, PHS, and SPD, and further characterized by chain length (e.g., 16:1;O2) (layer #6). On this same structural layer, the N-acyl is identified by carbon number and degrees of unsaturation (η), e.g. C24:1. Gangliosides consisting of additional C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C besides the sphingosine C4–C5 C
C besides the sphingosine C4–C5 C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, can be characterized by the location (layer #7). For instance, GM3 18:1;O2/24:1 is composed by two C
C, can be characterized by the location (layer #7). For instance, GM3 18:1;O2/24:1 is composed by two C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location isomers of C24:1(n-7) (11%) and C24:1(n-9) (89%).
C location isomers of C24:1(n-7) (11%) and C24:1(n-9) (89%).
Coordinated remodeling of ganglioside metabolism in MCF-7 breast cancer cells
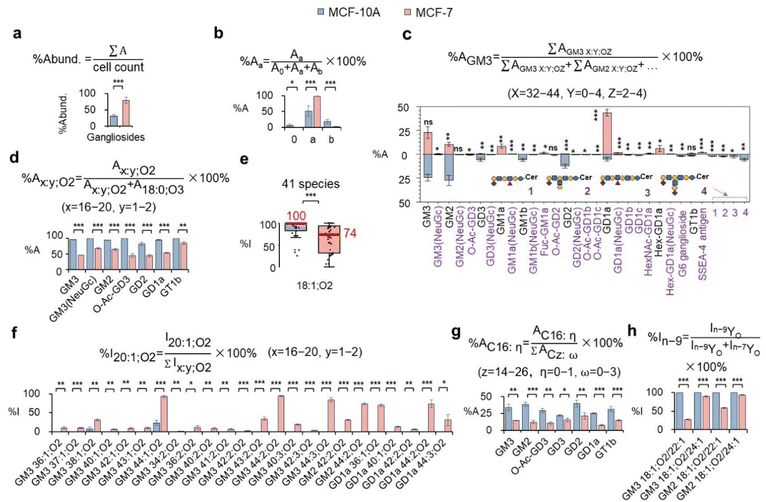
Aiming to map the metabolic changes of gangliosides at multiple structural levels in the MCF-7 breast cancer cell line, we used a human mammary epithelial cell line (MCF-10A) as a control, with three biological replicates analyzed for each cell line. From the MCF-10A cells, we identified 215 ganglioside species across 24 subclasses, with 122 and 30 structures having defined chain composition and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location (data provided in Tables S4–S6). Compared with MCF-10A, the MCF-7 cancer cells exhibited ∼3-fold more gangliosides (ion abundance/cell) (Fig. 4a). This observation is consistent with previous reports that breast cancer cells often exhibit elevated levels of sialylated glycoconjugates, including gangliosides.46 Due to the absence of commercially available ganglioside standards, the reported abundance of total gangliosides, series and subclasses does not represent actual molar ratios. For relative quantification, we employed integrated EIC peak areas for species-based analyses, including total gangliosides, series, and subclasses, as these species can be resolved by LC or TIMS. Given that chain composition and C
C location (data provided in Tables S4–S6). Compared with MCF-10A, the MCF-7 cancer cells exhibited ∼3-fold more gangliosides (ion abundance/cell) (Fig. 4a). This observation is consistent with previous reports that breast cancer cells often exhibit elevated levels of sialylated glycoconjugates, including gangliosides.46 Due to the absence of commercially available ganglioside standards, the reported abundance of total gangliosides, series and subclasses does not represent actual molar ratios. For relative quantification, we employed integrated EIC peak areas for species-based analyses, including total gangliosides, series, and subclasses, as these species can be resolved by LC or TIMS. Given that chain composition and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location isomers cannot be resolved by LC or TIMS, diagnostic fragment ion abundances were employed for the relative quantitation of these isomeric levels. By taking a deeper look at the ganglio-series, the a-series shows a prominent increase, with relative composition boosted from 52% to 98% (Fig. 4b), while the relative compositions of 0- and b-series decreased to 0% and 2%, respectively.
C location isomers cannot be resolved by LC or TIMS, diagnostic fragment ion abundances were employed for the relative quantitation of these isomeric levels. By taking a deeper look at the ganglio-series, the a-series shows a prominent increase, with relative composition boosted from 52% to 98% (Fig. 4b), while the relative compositions of 0- and b-series decreased to 0% and 2%, respectively.
Benefiting from selective enrichment, we profiled 31 distinct ganglioside subclasses across the two cell lines, including 21 novel ones highlighted in a purple font in Fig. 4c (MS/MS data provided in Fig. S8). Notably, 27 subclasses showed significant changes (p < 0.05) between cancer and normal breast cancer cells, based on the analysis of three biological replicates per cell line using a two-tailed t-test. The cancer cells showed significantly elevated GD1a (44% vs. 5%), GM1a (9% vs. 0%), Hex-GD1a (6% vs. 0%), SSEA-4 antigen (0% vs. 1.9%), and NeuGc-containing subclasses relative to the control (Fig. S9). In contrast, GD3 (0.6% vs. 6%) and GD2 (0.3% vs. 12%) were reduced; neolacto-series gangliosides, including G6 ganglioside and the four newly reported subclasses (1–4 in Fig. 4c), were below the limit of detection in MCF-7 cells. Our results differ from previous reports of elevated GD3 and GD2 in breast cancer,8,47 likely due to subtype- and model-specific heterogeneity. Elevated ganglioside subclasses in MCF-7 have been implicated in cancer-related functions. For instance, GM1a localizes to lipid rafts, where it interacts with TrkA receptors, stabilizes EGFR, and enhances PI3K/Akt signaling to support cell survival under stress and nutrient deprivation.48 GD1a enhances EGFR and PDGFR phosphorylation and recycling, sustaining proliferative signaling and cell cycle progression.49 SSEA-4 antigen acts as a cancer stem cell marker, promoting self-renewal, drug resistance, and invasiveness.50 Moreover, while both ganglio- and neolacto-series gangliosides were detectable in MCF-10A, MCF-7 cells exhibited only ganglio- and globo-series gangliosides, indicating a cancer-associated shift in the neutral glycan core composition.
MCF-7 cancer cells also exhibited significant alterations in ceramide chain compositions compared to MCF-10A. Gangliosides in both lines were composed of SPH (x:1;O2), SPD (x:2;O2), and PHS (x:0;O3). As SPH and SPD often coexisted in the same species, their combined composition (%Ax:y;O2, y = 1, 2) was calculated for each subclass. We found a significant decrease in the combined SPH and SPD, from nearly 100% to 44–86%, in seven out of ten subclasses commonly detected in both cell lines (Fig. 4d). A further look into the composition of SPH type revealed that %18:1;O2 decreased in 41 species of commonly detected ganglioside species with defined chain compositions in both cell lines (Fig. 4e). Conversely, an elevated %20:1;O2 was consistently observed in 24/24 ganglioside species commonly detected in both cell lines (Fig. 4f). Minor LCBs, including 17:1;O2 and 19:1;O2 also exhibited elevated levels in MCF-7 cells (Fig. S10). In terms of N-acyl composition, MCF-7 showed reduced C16: η (η = 0–1) content across seven subclasses (Fig. 4g). A detailed comparison of ganglioside structures at chain composition level is presented in Fig. S11. At the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location level, C22:1 and C24:1 N-acyls both contain n-7 and n-9 C
C location level, C22:1 and C24:1 N-acyls both contain n-7 and n-9 C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location isomers. Notably, the relative compositions of n-9 isomers all show a significant decrease in MCF-7 cells relative to the control (Fig. 4h), consistent with prior studies associating reduced n-9 isomers in phospholipids with increased breast cancer aggressiveness.51 These findings suggest that ganglioside remodeling in MCF-7 cells is a highly coordinated process, entailing rewiring of all five structural modules, such as the glycan headgroup, LCB, N-acyl, and C
C location isomers. Notably, the relative compositions of n-9 isomers all show a significant decrease in MCF-7 cells relative to the control (Fig. 4h), consistent with prior studies associating reduced n-9 isomers in phospholipids with increased breast cancer aggressiveness.51 These findings suggest that ganglioside remodeling in MCF-7 cells is a highly coordinated process, entailing rewiring of all five structural modules, such as the glycan headgroup, LCB, N-acyl, and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location.
C location.
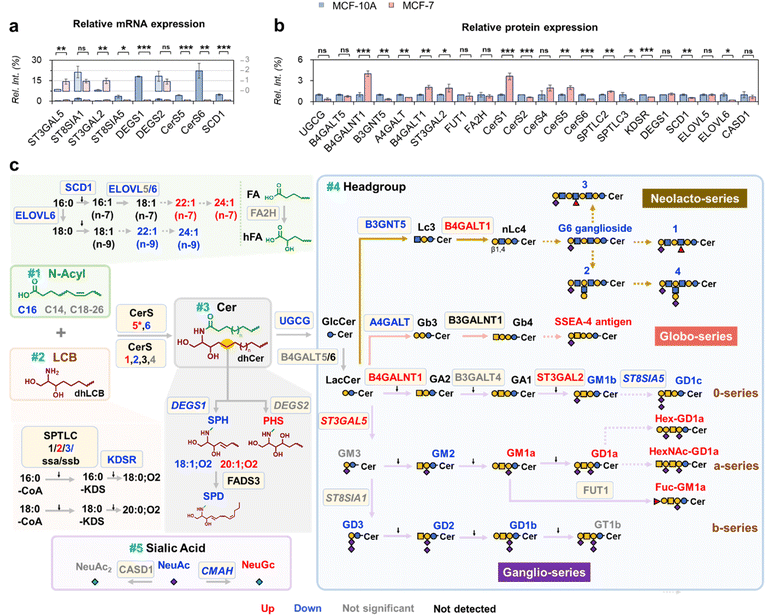
To dissect the regulatory logic behind this, we integrated the lipidomic data with targeted gene expression analysis and label-free quantitative proteomics. First, we performed quantitative reverse transcription PCR (qRT-PCR) to quantify nine enzymes participating in ganglioside biosynthesis. These enzymes were selected because their corresponding lipid products showed significant alterations in MCF-7 cells (Fig. 5a). The primer information used for qPCR is provided in Table S7. Consistent with elevated a-series in MCF-7 cells, ST3GAL5, which adds sialic acid to internal galactose to produce an a-series ganglioside, is significantly upregulated in MCF-7 cancer cells. ST8SIA1 is an enzyme responsible for converting a-series to b-series gangliosides via α2,8-sialylation. Interestingly, ST8SIA1 shows no significant change, leading toward a-series accumulation. ST3GAL2, which modifies GA1, GM1a, and GD1b to yield GM1b, GD1a, and GT1b, respectively, is also upregulated. Conversely, ST8SIA5, converting GM1b, GD1a, and GT1b into more complex gangliosides, is downregulated. The collective modulations of ST3GAL2 and ST8SIA5 expression align with observed GD1a accumulation. CMAH is responsible for converting NeuAc to NeuGc (Fig. S12). Despite a decrease in CMAH expression, NeuGc-containing gangliosides show an increased amount, possibly via exogenous NeuGc uptake or sialic acid pathway reprogramming.52 Ceramide-related enzymes are also altered from the transcription level. DEGS1, CerS5 (with C14:0 and C16:0 specificity), and CerS6 (with C16:0 specificity) are all downregulated, while DEGS2 remains unchanged. These coordinated changes are consistent with reduced %AC16: η (η = 0–1) and %Ax:y;O2 (x = 16−20, y = 1−2) in gangliosides from the cancer cell line. SCD1, which desaturates C16:0 or C18:0 to generate monounsaturated fatty acids (MUFAs), is also downregulated. However, its role in n-9 isomer reduction is unclear, as it contributes to both n-7 and n-9 MUFA synthesis.
To confirm enzymatic changes in protein level, we performed label-free quantitative proteomics (three replicates per cell line). Out of 39 enzymes involved in ganglioside biosynthesis (Table S8), 22 were detected in both cell lines, with abundances ranging from 0.005% to 0.7% relative to the most abundant protein (GTSE1, Fig. S13). The relative abundances of these enzymes are shown in Fig. 5b. Only 8 glycosyltransferases were detected, likely due to their membrane association, underscoring the limitations of current proteomic methods in detecting low abundance or membrane-bound enzymes. ST3GAL2, CerS6, and SCD1, showed consistent changes at transcription and protein expression levels, suggesting transcriptional regulation. In contrast, DEGS1 and CerS5 showed discordant patterns. That is, DEGS1 mRNA was downregulated in MCF-7 with protein expression unchanged, while CerS5 mRNA was decreased, accompanied by increased protein expression. These discrepancies may indicate post-transcriptional or post-translational regulation affecting enzyme abundance independently of mRNA levels. Several alterations in ganglioside metabolism cannot be explained by the enzymatic changes observed at the mRNA or protein levels. For instance, the biosynthetic origins of uncommon subclasses, such as G6 ganglioside, Hex-GD1a, HexNAc-GD1a, and subclasses 1 to 4, remain unclear. Likewise, the selective enzymatic mechanisms responsible for the elevated 20:1;O2 LCB and n-7 isomers in C22:1 and C24:1 species in MCF-7 cells are not yet understood. Nonetheless, the profiling data provide valuable clues for identifying previously unrecognized enzymatic activities.
Building on both lipidomics and proteomics data, we propose a lipid-centric framework for reconstructing the altered ganglioside metabolic network. The approach begins with lipidomic profiling to map changes across multiple structural levels of gangliosides, followed by targeted analysis of key biosynthetic enzymes at both transcript and protein levels. This leads to the construction of a five-module biosynthetic network (Fig. 5c), including N-acyl, LCB, ceramide, sialic acid, and glycan. This network provides the most comprehensive view of ganglioside remodeling in breast cancer cells to date, which cannot be obtained from existing databases, such as the Cancer Cell Line Encyclopedia (CCLE).
Conclusions
We developed a highly sensitive analytical workflow for the molecular profiling of gangliosides by integrating selective enrichment, amide HILIC-TIMS-MS/MS, and charge-tagging PB derivatization. The dynamic range for ganglioside detection from MCF-7 cells is extended by two orders of magnitude compared to conventional methods, leading to the construction of a structural atlas of 391 ganglioside molecules across subclass, species, chain composition, and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond positions. This workflow reveals numerous novel structures, including new subclasses (GM3(NeuGc), SSEA-4 antigens, etc.), highly hydroxylated ceramides (X:Y;O3/O4), unusual LCBs (17:1;O2, 19:1;O2, 18:0;O3), hydroxylated fatty acids, and precise C
C bond positions. This workflow reveals numerous novel structures, including new subclasses (GM3(NeuGc), SSEA-4 antigens, etc.), highly hydroxylated ceramides (X:Y;O3/O4), unusual LCBs (17:1;O2, 19:1;O2, 18:0;O3), hydroxylated fatty acids, and precise C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond locations. The deep profiling capability allows us to map the remodeling of five key building blocks of gangliosides in MCF-7 breast cancer cells relative to a normal control, such as altered glycan types, shifts in LCB and fatty acyl compositions, and changes in C
C bond locations. The deep profiling capability allows us to map the remodeling of five key building blocks of gangliosides in MCF-7 breast cancer cells relative to a normal control, such as altered glycan types, shifts in LCB and fatty acyl compositions, and changes in C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location isomers. These previously unrecognized changes in ganglioside metabolism may influence membrane properties and signaling in cancer cells.
C location isomers. These previously unrecognized changes in ganglioside metabolism may influence membrane properties and signaling in cancer cells.
The integration of targeted transcriptomic and untargeted proteomic analyses reveals that these lipid changes are underpinned by coordinated, yet sometimes uncoupled, regulation of biosynthetic enzymes. ST3GAL2, CerS6, and SCD1 show consistent trends across mRNA and protein levels, whereas DEGS1 and CERS5 exhibited discordant patterns, possibly reflecting complex regulatory layers. Using this lipid-centric approach, we construct a hypothesis-generating model of ganglioside metabolism, which contains five key structural modules: N-acyl, LCB, ceramide, sialic acid, and glycan. By acquiring the profiles of structurally resolved gangliosides, the lipid-centric approach delineates mechanistic links between lipid remodeling and enzyme regulation, expanding knowledge beyond the scope of current databases such as CCLE. Furthermore, our method overcomes the limited coverage of membrane-bound and low-abundance enzymes inherent in untargeted proteomics approaches by performing targeted enzyme discovery based on differential lipids identified via deep profiling. Although some ganglioside remodeling remains unexplained with existing knowledge, this type of information provides important experimental clues for the discovery of previously uncharacterized enzymatic functions in ganglioside biosynthesis. It is also important to note that analyzing additional breast cancer subtypes, such as triple-negative models, will be crucial for generating a more comprehensive and subtype-specific map of altered ganglioside metabolism in breast cancer.
The limitations of our study should also be acknowledged. Due to the scarcity of commercially available ganglioside standards, quantification is performed in a semi-quantitative fashion. Future efforts incorporating more diverse internal standards or employing parallel quantitation will be essential to achieving more accurate profiling data. The proposed biosynthetic connections are correlative and will require functional validation (ex., genetic perturbation) in biological and clinical models to confirm causality and pathophysiological relevance in breast cancer. Moreover, the potential of alternative enrichment materials and protocols, such as Zr4+-immobilized solid-phase extraction columns,53,54 can be explored.
Author contributions
Y. X. and Y. W. designed the experiments. Y. W. performed the experiments and analyzed the data. G. S. conducted the qRT-PCR experiment. Y. X., Y. W. co-wrote the paper. Y. X., Y. W., G. S., and H. Y. discussed the results and commented on the paper.Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting this article, including experimental procedures, characterization data, and spectra, are available in the supplementary information (SI). Supplementary information: additional experimental details and methods; optimization of the TiO2 MNPs enrichment procedure; characterized MS/MS spectra for ganglioside identification at multiple structural levels; and comprehensive profiles of gangliosides, mRNA expression, and protein expression of related enzymes in MCF-10A versus MCF-7 cell lines (PDF) tables of identified gangliosides at species, chain composition and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C location level from MCF-10A and MCF-7 cells, gene-specific primers, and enzymes involved in ganglioside biosynthesis (XLSX). See DOI: https://doi.org/10.1039/d5sc09445c.
C location level from MCF-10A and MCF-7 cells, gene-specific primers, and enzymes involved in ganglioside biosynthesis (XLSX). See DOI: https://doi.org/10.1039/d5sc09445c.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grants 22225404, 22137004, and 82430109), the National Key R&D Program of China (2023YFA0913902), and the Noncommunicable Chronic Diseases–National Science and Technology Major Project (2023ZD0501500). We thank the Protein Chemistry and Proteomics Platform at Tsinghua University for conducting label-free quantitative proteomics experiments and Dr Zidan Wang for her assistance in synthesizing TiO2 MNPs.References
- E. Sezgin, I. Levental, S. Mayor and C. Eggeling, The mystery of membrane organization: composition, regulation and roles of lipid rafts, Nat. Rev. Mol. Cell Biol., 2017, 18, 361–374 Search PubMed.
- V. Corradi, E. Mendez-Villuendas, H. I. Ingólfsson, R.-X. Gu, I. Siuda, M. N. Melo, A. Moussatova, L. J. DeGagné, B. I. Sejdiu, G. Singh, T. A. Wassenaar, K. Delgado Magnero, S. J. Marrink and D. P. Tieleman, Lipid–Protein Interactions Are Unique Fingerprints for Membrane Proteins, ACS Cent. Sci., 2018, 4, 709–717 CrossRef CAS PubMed.
- C. S. Delaveris, C. L. Wang, N. M. Riley, S. Li, R. U. Kulkarni and C. R. Bertozzi, Microglia Mediate Contact-Independent Neuronal Network Remodeling via Secreted Neuraminidase-3 Associated with Extracellular Vesicles, ACS Cent. Sci., 2023, 9, 2108–2114 CrossRef CAS PubMed.
- K. Hamamura, K. Furukawa, T. Hayashi, T. Hattori, J. Nakano, H. Nakashima, T. Okuda, H. Mizutani, H. Hattori, M. Ueda, T. Urano, K. O. Lloyd and K. Furukawa, Ganglioside GD3 promotes cell growth and invasion through p130Cas and paxillin in malignant melanoma cells, Proc. Natl. Acad. Sci. U.S.A., 2005, 102, 11041–11046 Search PubMed.
- C. W. Mount, R. G. Majzner, S. Sundaresh, E. P. Arnold, M. Kadapakkam, S. Haile, L. Labanieh, E. Hulleman, P. J. Woo, S. P. Rietberg, H. Vogel, M. Monje and C. L. Mackall, Potent antitumor efficacy of anti-GD2 CAR T cells in H3-K27M(+) diffuse midline gliomas, Nat. Med., 2018, 24, 572–579 Search PubMed.
- V. L. Battula, Y. Shi, K. W. Evans, R. Y. Wang, E. L. Spaeth, R. O. Jacamo, R. Guerra, A. A. Sahin, F. C. Marini, G. Hortobagyi, S. A. Mani and M. Andreeff, Ganglioside GD2 identifies breast cancer stem cells and promotes tumorigenesis, J. Clin. Invest., 2012, 122, 2066–2078 Search PubMed.
- Y. J. Liang, Y. Ding, S. B. Levery, M. Lobaton, K. Handa and S. I. Hakomori, Differential expression profiles of glycosphingolipids in human breast cancer stem cells vs. cancer non-stem cells, Proc. Natl. Acad. Sci. U.S.A., 2013, 110, 4968–4973 CrossRef CAS PubMed.
- A. Cazet, J. Lefebvre, E. Adriaenssens, S. Julien, M. Bobowski, A. Grigoriadis, A. Tutt, D. Tulasne, X. Le Bourhis and P. Delannoy, GD3 synthase expression enhances proliferation and tumor growth of MDA-MB-231 breast cancer cells through c-Met activation, Mol. Cancer Res., 2010, 8, 1526–1535 CrossRef CAS PubMed.
- A. Cazet, M. Bobowski, Y. Rombouts, J. Lefebvre, A. Steenackers, I. Popa, Y. Guérardel, X. Le Bourhis, D. Tulasne and P. Delannoy, The ganglioside GD2 induces the constitutive activation of c-Met in MDA-MB-231 breast cancer cells expressing the GD3 synthase, Glycobiology, 2012, 22, 806–816 Search PubMed.
- K. Wang, Y. Chen, S. Ahn, M. Zheng, E. Landoni, G. Dotti, B. Savoldo and Z. Han, GD2-specific CAR T cells encapsulated in an injectable hydrogel control retinoblastoma and preserve vision, Nat. Cancer., 2020, 1, 990–997 Search PubMed.
- S. K. Tan, H. Y. Hougen, J. R. Merchan, M. L. Gonzalgo and S. M. Welford, Fatty acid metabolism reprogramming in ccRCC: mechanisms and potential targets, Nat. Rev. Urol., 2023, 20, 48–60 CrossRef CAS PubMed.
- L. A. Broadfield, A. A. Pane, A. Talebi, J. V. Swinnen and S. M. Fendt, Lipid metabolism in cancer: New perspectives and emerging mechanisms, Dev. Cell, 2021, 56, 1363–1393 Search PubMed.
- S. Arumugam, S. Schmieder, W. Pezeshkian, U. Becken, C. Wunder, D. Chinnapen, J. H. Ipsen, A. K. Kenworthy, W. Lencer, S. Mayor and L. Johannes, Ceramide structure dictates glycosphingolipid nanodomain assembly and function, Nat. Commun., 2021, 12, 3675 CrossRef CAS PubMed.
- E. N. Schmidt, X. Y. Guo, D. T. Bui, J. Jung, J. S. Klassen and M. S. Macauley, Dissecting the abilities of murine Siglecs to interact with gangliosides, J. Biol. Chem., 2024, 300, 107482 CrossRef CAS PubMed.
- R. Hajek, R. Jirasko, M. Lisa, E. Cifkova and M. Holcapek, Hydrophilic interaction liquid chromatography-mass spectrometry characterization of gangliosides in biological samples, Anal. Chem., 2017, 89, 12425–12432 CrossRef CAS PubMed.
- M. Sarbu, A. C. Robu, R. M. Ghiulai, Z. Vukelic, D. E. Clemmer and A. D. Zamfir, Electrospray ionization ion mobility mass spectrometry of human brain gangliosides, Anal. Chem., 2016, 88, 5166–5178 Search PubMed.
- R. L. Schindler, A. Oloumi, J. Tena, M. R. S. Alvarez, Y. Liu, S. Grijaldo, M. Barboza, L. W. Jin, A. M. Zivkovic and C. B. Lebrilla, Profiling intact glycosphingolipids with automated structural annotation and quantitation from human samples with nanoflow liquid chromatography mass spectrometry, Anal. Chem., 2024, 96, 5951–5959 Search PubMed.
- H. Li, R. Xu, L. Yang, H. Luan, S. Chen, L. Chen, Z. Cai and R. Tian, Combinatory data-independent acquisition and parallel reaction monitoring method for deep profiling of gangliosides, Anal. Chem., 2020, 92, 10830–10838 Search PubMed.
- K. Hohenwallner, L. M. Lamp, L. Peng, M. Nuske, J. Hartler, G. E. Reid and E. Rampler, FAIMS shotgun lipidomics for enhanced class- and charge-state separation complemented by automated ganglioside annotation, Anal. Chem., 2024, 96, 12296–12307 CAS.
- H. G. Vo, G. Gonzalez-Escamilla, D. Mirzac, L. Rotaru, D. Herz, S. Groppa and L. Bindila, Extended coverage of human serum glycosphingolipidome by 4D-RP-LC TIMS-PASEF unravels association with Parkinson's disease, Nat. Commun., 2025, 16, 4567 CrossRef CAS PubMed.
- J. Lee, H. Hwang, S. Kim, J. Hwang, J. Yoon, D. Yin, S. I. Choi, Y. H. Kim, Y. S. Kim and H. J. An, Comprehensive profiling of surface gangliosides extracted from various cell lines by LC-MS/MS, Cells, 2019, 8, 1323 Search PubMed.
- T. Hu, Z. Jia and J. L. Zhang, Strategy for comprehensive profiling and identification of acidic glycosphingolipids using ultra-high-performance liquid chromatography coupled with quadrupole time-of-flight mass spectrometry, Anal. Chem., 2017, 89, 7808–7816 CrossRef CAS PubMed.
- J. Luo, Q. Wan and S. Chen, Chemistry-driven mass spectrometry for structural lipidomics at the C=C bond isomer level, Chinese. Chem. Lett., 2025, 36, 109836 CrossRef CAS.
- X. Zhao, H. Shi, Q. Lin and Y. Xia, Review: Deep lipidomic profiling enabled by the Patern'o-Büchi reaction - Pinpointing C=C locations and beyond, Anal. Chim. Acta, 2025, 1376, 344456 CrossRef CAS PubMed.
- D. L. Marshall, H. T. Pham, M. Bhujel, J. S. Chin, J. Y. Yew, K. Mori, T. W. Mitchell and S. J. Blanksby, Sequential collision- and ozone-induced dissociation enables assignment of relative acyl chain position in triacylglycerols, Anal. Chem., 2016, 88, 2685–2692 CrossRef CAS PubMed.
- P. E. Williams, D. R. Klein, S. M. Greer and J. S. Brodbelt, Pinpointing double bond and sn-positions in glycerophospholipids via hybrid 193 nm ultraviolet photodissociation (UVPD) mass spectrometry, J. Am. Chem. Soc., 2017, 139, 15681–15690 CrossRef CAS PubMed.
- J. L. Campbell and T. Baba, Near-complete structural characterization of phosphatidylcholines using electron impact excitation of ions from organics, Anal. Chem., 2015, 87, 5837–5845 CrossRef CAS PubMed.
- G. Feng, M. Gao, L. Wang, J. Chen, M. Hou, Q. Wan, Y. Lin, G. Xu, X. Qi and S. Chen, Dual-resolving of positional and geometric isomers of C=C bonds via bifunctional photocycloaddition-photoisomerization reaction system, Nat. Commun., 2022, 13, 2652 CrossRef CAS PubMed.
- B. Zhang, Y. Wang, B. W. Zhou, J. Cheng, Q. Xu, L. Zhang, T. Q. Sun, J. Zhang and Y. L. Guo, Chloramine-T-enabled mass spectrometric analysis of C=C isomers of unsaturated fatty acids and phosphatidylcholines in human thyroids, Anal. Chem., 2022, 94, 6216–6224 Search PubMed.
- T. Yang, S. Tang, S. T. Kuo, D. Freitas, M. Edwards, H. Wang, Y. Sun and X. Yan, Lipid mass tags via aziridination for probing unsaturated lipid isomers and accurate relative quantification, Angew. Chem., Int. Ed., 2022, 61, e202207098 CrossRef CAS PubMed.
- G. Feng, M. Gao, H. Chen, Z. Zhang, J. Chen, Y. Tong, P. Wu, R. Fu, Y. Lin and S. Chen, Stable-isotope N-Me aziridination enables accurate quantitative C=C isomeric lipidomics, Anal. Chem., 2024, 96, 2524–2533 Search PubMed.
- Z. Wang, D. Zhang, J. Wu, W. Zhang and Y. Xia, Illuminating the dark space of neutral glycosphingolipidome by selective enrichment and profiling at multi-structural levels, Nat. Commun., 2024, 15, 5627 CrossRef CAS PubMed.
- Z. Wang and Y. Xia, Selective enrichment via TiO2 magnetic nanoparticles enables deep profiling of circulating neutral glycosphingolipids, Anal. Chem., 2024, 96, 16955–16963 Search PubMed.
- M. A. Chester, IUPAC-IUB Joint Commission on Biochemical Nomenclature (JCBN). Nomenclature of glycolipids--recommendations 1997, Eur. J. Biochem., 1998, 257, 293–298 Search PubMed.
- G. Liebisch, E. Fahy, J. Aoki, E. A. Dennis, T. Durand, C. S. Ejsing, M. Fedorova, I. Feussner, W. J. Griffiths, H. Kofeler, A. H. Merrill Jr., R. C. Murphy, V. B. O'Donnell, O. Oskolkova, S. Subramaniam, M. J. O. Wakelam and F. Spener, Update on LIPID MAPS classification, nomenclature, and shorthand notation for MS-derived lipid structures, J. Lipid Res., 2020, 61, 1539–1555 Search PubMed.
- A. Prinetti, V. Chigorno, G. Tettamanti and S. Sonnino, Sphingolipid-enriched membrane domains from rat cerebellar granule cells differentiated in culture: A compositional study, J. Biol. Chem., 2000, 275, 11658–11665 Search PubMed.
- M. R. Larsen, S. S. Jensen, L. A. Jakobsen and N. H. H. Heegaard, Exploring the sialiome using titanium dioxide chromatography and mass spectrometry, Mol. Cell. Proteomics, 2007, 6, 1778–1787 CrossRef CAS PubMed.
- J. Liu, F. Zhang, S. Dou, M. Zhu, L. Ding and Y. Yang, Adsorption of serine at the anatase TiO2/water interface: A combined ATR-FTIR and DFT study, Sci. Total Environ., 2022, 807, 150839 CrossRef CAS PubMed.
- I. B. Ustunol, N. I. Gonzalez-Pech and V. H. Grassian, pH-dependent adsorption of alpha-amino acids, lysine, glutamic acid, serine and glycine, on TiO2 nanoparticle surfaces, J. Colloid Interface Sci., 2019, 554, 362–375 CrossRef CAS PubMed.
- Z. Huang, Q. Wu, H. Lu, Y. Wang and Z. Zhang, Separation of glycolipids/sphingolipids from glycerophospholipids on TiO2 coating in aprotic solvent for rapid comprehensive lipidomic analysis with liquid microjunction surface sampling-mass spectrometry, Anal. Chem., 2020, 92, 11250–11259 CrossRef CAS PubMed.
- L. Xu, H. Shi, T. Liang, J. Feng, Y. Jin, Y. Ke and X. Liang, Selective separation of flavonoid glycosides in Dalbergia odorifera by matrix solid-phase dispersion using titania, J. Sep. Sci., 2011, 34, 1347–1354 CrossRef CAS PubMed.
- H. Takeda, M. Takeuchi, M. Hasegawa, J. Miyamoto and H. Tsugawa, A procedure for solid-phase extractions using metal-oxide-coated silica column in lipidomics, Anal. Chem., 2024, 96, 17065–17070 CrossRef CAS PubMed.
- S. Feng, C. Bagia and G. Mpourmpakis, Determination of proton affinities and acidity constants of sugars, J. Phys. Chem. A, 2013, 117, 5211–5219 Search PubMed.
- J.-L. Lin, Y.-C. Lin, B.-C. Lin, P.-C. Lai, T.-E. Chien, S.-H. Li and Y.-F. Lin, Adsorption and reactions on TiO2: Comparison of N,N-dimethylformamide and dimethylamine, J. Phys. Chem. C, 2014, 118, 20291–20297 CrossRef CAS.
- P. Kwantwi-Barima, S. V. B. Garimella, I. K. Attah, X. Zheng, Y. M. Ibrahim and R. D. Smith, Accumulation of large ion populations with high ion densities and effects due to space charge in traveling wave-based structures for lossless ion manipulations (SLIM) IMS-MS, J. Am. Soc. Mass Spectrom., 2024, 35, 365–377 CrossRef CAS PubMed.
- S. S. Pinho and C. A. Reis, Glycosylation in cancer: mechanisms and clinical implications, Nat. Rev. Cancer, 2015, 15, 540–555 CrossRef CAS PubMed.
- C. Shao, V. Anand, M. Andreeff and V. L. Battula, Ganglioside GD2: a novel therapeutic target in triple-negative breast cancer, Ann. N. Y. Acad. Sci., 2022, 1508, 35–53 Search PubMed.
- Z. Guo, Ganglioside GM1 and the central nervous system, Int. J. Mol. Sci., 2023, 24, 9558 CrossRef CAS PubMed.
- M. Takahashi, Y. Hasegawa, K. Maeda, M. Kitano and N. Taniguchi, Role of glycosyltransferases in carcinogenesis; growth factor signaling and EMT/MET programs, Glycoconj. J., 2022, 39, 167–176 CrossRef CAS PubMed.
- N. Y. Chen, C. W. Lin, T. Y. Lai, C. Y. Wu, P. C. Liao, T. L. Hsu and C. H. Wong, Increased expression of SSEA-4 on TKI-resistant non-small cell lung cancer with EGFR-T790M mutation, Proc. Natl. Acad. Sci. U.S.A., 2024, 121, e2313397121 CrossRef CAS PubMed.
- W. Cao, S. Cheng, J. Yang, J. Feng, W. Zhang, Z. Li, Q. Chen, Y. Xia, Z. Ouyang and X. Ma, Large-scale lipid analysis with C=C location and sn-position isomer resolving power, Nat. Commun., 2020, 11, 375 CrossRef CAS PubMed.
- J. Heinzelbecker, M. Fauskanger, I. Jonson, U. Krengel, G. A. Loset, L. Munthe and A. Tveita, Chimeric antigen receptor T cells targeting the GM3(Neu5Gc) ganglioside, Front. Immunol., 2024, 15, 1331345 CrossRef CAS PubMed.
- C. Wang, M. Wang and X. Han, Comprehensive and quantitative analysis of lysophospholipid molecular species present in obese mouse liver by shotgun lipidomics, Anal. Chem., 2015, 87, 4879–4887 CrossRef CAS PubMed.
- H. Nagasawa, S. Miyazaki and M. Kyogashima, Simple separation of glycosphingolipids in the lower phase of a Folch's partition from crude lipid fractions using zirconium dioxide, Glycoconj. J., 2022, 39, 789–795 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |