Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Allosteric ligand–aptamer complexes orchestrate supramolecular or transient catalytic, transcription and fibrinogenesis processes
Diva
Froim
 a,
Hadar
Amartely
a,
Hadar
Amartely
 b,
Jiantong
Dong
b,
Jiantong
Dong
 *a,
Eli
Pikarsky
*a,
Eli
Pikarsky
 c and
Itamar
Willner
c and
Itamar
Willner
 *a
*a
aInstitute of Chemistry, The Hebrew University of Jerusalem, Jerusalem 91904, Israel. E-mail: jtdong2025@nankai.edu.cn; itamar.willner@mail.huji.ac.il
bWolfson Centre for Applied Structural Biology, The Hebrew University of Jerusalem, Jerusalem 91904, Israel
cThe Lautenberg Center for Immunology and Cancer Research, IMRIC, The Hebrew University of Jerusalem, Jerusalem 91120, Israel
First published on 2nd February 2026
Abstract
Allosteric regulation, the modulation of biological macromolecular function through binding of molecules at distant sites distinct from the active site, is a fundamental principle in biology that governs enzyme activity, signaling, and gene expression. In this work, we present allosteric ligand/aptamer complexes, coupled to biocatalytic reaction modules composed of enzymes, DNAzymes, or transcription machineries, regulating the catalytic and transient functions of these frameworks. This principle is exemplified by the assembly of ligand/aptamer subunits supramolecular complexes that allosterically stabilize the Mg2+-dependent DNAzyme, allowing its ribonucleobase cleavage activity, promoting the formation of transcription templates that yield RNA products, and modulating the assembly of thrombin aptamer subunits that inhibit thrombin-induced coagulation. Specifically, melamine (Mel)/aptamer subunits complexes allosterically stabilize the assembly of Mg2+-dependent DNAzyme strands for substrate cleavage, the formation of thrombin aptamer subunits that inhibit the conversion of fibrinogen to fibrin, and the stabilization of a transcription template encoding the Malachite Green (MG) RNA aptamer. Furthermore, coupling an enzyme that depletes the ligand/aptamer complex, which allosterically stabilizes the biocatalytic reaction module, demonstrates the dissipative and transient operation of the catalytic system. This concept is illustrated by the adenosine (Ade)/aptamer subunits supramolecular complex, which stabilizes thrombin aptamer subunits to inhibit thrombin-induced fibrinogenesis, and promotes the formation of an active transcription template for RNA synthesis. In the presence of adenosine deaminase (ADA), Ade is transformed into inosine, which lacks affinity for the aptamer subunits, thereby degrading the Ade/aptamer assemblies and depleting the allosteric complexes. The temporal disassembly of these allosteric stabilizing complexes leads to the transient inhibition of thrombin-induced coagulation or to the transient operation of a transcription machinery.
Introduction
Allosteric regulation, the modulation of the activity of macromolecules through binding of molecules at distant sites, is a fundamental principle governing biological processes. The communication between spatially separated molecular domains, provides the dynamic flexibility that underlies metabolism, signaling, and gene expression.1,2 Jacques Monod famously referred to allostery as the “second secret of life”, emphasizing its central role in the self-organization and adaptability of biological systems.3 Introduction of the allostery concept into synthetic nucleic acid structures might add an important dimension to the functional properties of the biopolymer and expand its application in synthetic biology. The base sequence comprising nucleic acids encodes functional information into the structure of the biopolymer. Sequence guided functions of DNA include sequence-specific recognition and binding of biomolecules or low molecular-weight ligands (aptamers),4–6 sequence dictated displacement of duplex nucleic acids or protein/nucleic acid complexes by auxiliary strands,7–9 and sequence regulated catalytic properties in the presence of auxiliary cofactors (DNAzymes or ribozymes), e.g., metal-ion or amino acid-dependent DNAzymes10–13 and hemin/G-quadruplex14 DNAzymes. Moreover, the sequences comprising DNA duplex frameworks dictate selective reactivity patterns towards auxiliary enzymes such as endonucleases15,16 or nickases.17 In addition, auxiliary enzymes such as DNA or RNA polymerases and added deoxyribonucleotide triphosphates (dNTPs) or ribonucleotide triphosphates (NTPs) as fuels, catalyze in the presence of nucleic acid templates dictated polymerization and displacement of DNA or RNA products.18,19 This arsenal of recognition and catalytic functions embedded in oligonucleotides provides a versatile “tool-box” for the rapidly developing area of DNA nanotechnology.20 Over the years, the functional information embedded in nucleic acids has been implemented to develop DNA switches,21,22 machines23,24 and two- and three-dimensional DNA nanostructures.25,26 In addition, dynamically reconfigured nanostructures,27 programmed logic gate circuits,28 dynamic reconfigurable DNA networks,29 dissipative circuits,30,31 and switchable transcription machineries32,33 were demonstrated. Moreover, chemical modifications of aptamers with light responsive34 or redox active units35 led to switchable binding properties of aptamers. Conjugation of aptamers to DNAzyme catalytic units yielded hybrid structures, “nucleoaptazymes”, emulating native enzymes by providing cooperative substrate binding sites in spatial proximity to the active site in the conjugated structure.36,37 Diverse applications of aptamers were demonstrated, including their use as sensing38,39 and imaging materials,40 engineering of stimuli-responsive drug-carriers, and their targeting to specific cell receptors.41 Also, aptamers were used as therapeutic agents through selective binding to proteins and their inhibition, e.g., association to VEGF (inhibiting angiogenesis)42,43 or thrombin (inhibiting fibrinogenesis).44 Similarly, DNAzymes have found broad applications as amplifiers of sensing events, using in vitro or in vivo assays45 and catalyzing diverse chemical transformations, such as oxidation of NADH46 or dopamine.47 Also, DNAzymes were employed as synthetic catalysts for gene therapy48 and for the generation of reactive oxygen species for chemo dynamic cancer therapy.49Here we wish to report on the conjugation of ligand/aptamer supramolecular complexes to DNAzyme subunits, anti-protein (thrombin) aptamer subunits and transcription templates, resulting in the allosteric operation of a DNAzyme, inhibition of fibrinogenesis and RNA transcription. Melamine (Mel) or adenosine (Ade) act as ligands assembling ligand/aptamer supramolecular complexes that allosterically guide the respective catalytic, fibrinogenic and transcription circuits. By coupling adenosine deaminase (ADA) to the Ade aptamer allosterically stabilized fibrinogenesis and transcription circuits, transient, dissipative, operation of the frameworks is achieved. Beyond the expansion of the functionalities of stimuli-responsive nucleic acid circuits, the significance of the ligand/aptamer complex, allosterically-driven, catalytic circuits is reflected by: (i) their possible application for sensing (e.g., Mel); (ii) the temporal, dose-controlled, therapeutic applications of the circuits (e.g., inhibition of thrombin-induced fibrinogenesis); and (iii) the spatiotemporal control-over transcription machineries by synthetic ligand/aptamer complexes emulating functions of native transcription factors. It should be noted that in a previous study50 we reported on the allosteric adenosine/aptamer subunits activation of the Mg2+-ion-dependent DNAzyme and its ADA-directed transient catalytic modulation in liposome protocells. The present study demonstrates, however, the versatility of the allosteric activation of diverse supramolecular and transient catalytic transformations such as fibrinogenesis or transcription machineries, using different ligand/aptamer subunits complexes as allosteric stabilizing units.
Results and discussion
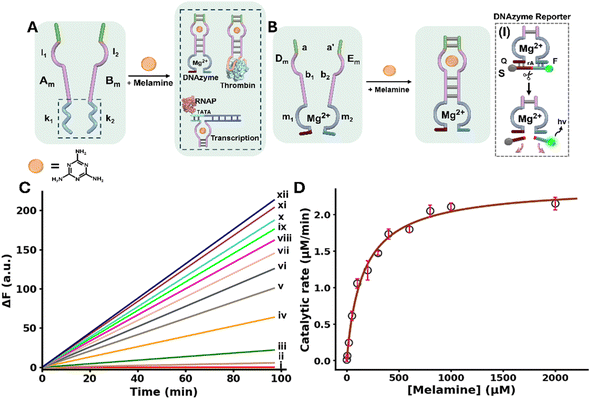
The general concept to allosterically operate catalytic circuits consisting of DNAzymes, anti-protein (thrombin) aptamer subunits or transcription machineries by ligand/aptamer complexes is exemplified in Fig. 1A using melamine (Mel)/aptamer complexes.51,52 The system consists of two strands Am and Bm that include engineered sub-domains l1 and l2 corresponding to aptamer subunits conjugated to the catalytic subunits k1 and k2. The catalytic subunits include the pre-engineered sequences to operate a DNAzyme, bind thrombin or, transcribe RNA sequences. While the strands Am and Bm have partial complementarity, it is insufficient to form a stable interstrand complex. However, binding of the ligand (Mel) to the aptamer subunits results in an interstrand supramolecular complex cooperatively stabilized by the ligand/aptamer complex and the complementarity associated with the two strands. Furthermore, the spatial proximity between the tethers k1 and k2 is pre-engineered to evolve the catalytic function in the supramolecular assembly. That is, the ligand-induced formation of ligand/aptamer supramolecular complex allosterically stabilizes and activates the catalytic function of the spatially confined k1 and k2 subunits.Fig. 1B schematically depicts the Mel/aptamer subunits complex induced allosteric stabilization of a supramolecular complex activating the function of a Mg2+-ion-dependent DNAzyme. The reaction circuit includes two strands Dm and Em which include subunits b1 and b2 of the split Mel aptamer, and subunits m1 and m2 corresponding to the arm/loop sequences of the DNAzyme. The complementary sequence domains a and a′ were added to strands Dm and Em. In the presence of Mel, the supramolecular complex cooperatively stabilized by the Mel/aptamer subunits complex and the interbridging duplex a/a′ is formed, resulting in the spatially confined DNAzyme configuration composed of m1 and m2. The allosteric stabilization of the functional Mg2+-ion-dependent DNAzyme is then probed by the cleavage of the fluorophore/quencher modified substrate S by the DNAzyme, panel I (F = FAM; Q = BHQ1). For the specific design of the structural units activating the allosteric Mel-induced operation of the DNAzyme see Page S9 in the SI.
Fig. 1C, depicts the rate of cleavage of the F/Q-modified substrate by the allosterically, Mel-stabilized DNAzyme framework in the presence of variable Mel concentrations. While no cleavage of the substrate proceeds in the absence of Mel, consistent with the lack of communication between strands Dm and Em, curve i, the supramolecular DNAzyme structure is activated in the presence of Mel, curves ii–xii. As the concentration of Mel increases, the rate of cleavage is enhanced. Fig. 1D, displays the rates of cleavage of the substrate strand in the presence of variable Mel concentrations. A saturation curve is observed consistent with the saturated formation of the Mel/aptamer subunits complex from which the Vmax = 2.41 ± 0.07 µmol min−1 and K0.5 = 162 ± 4.8 µM for the allosterically stabilized supramolecular DNAzyme were evaluated. The detection limit of the DNAzyme was calculated to be 867 ± 26 nM using the three-sigma method. Isothermal titration calorimetry (ITC) experiments supported the Mel/aptamer subunits formation of the supramolecular DNAzyme structure revealing a Kd of 0.890 ± 0.123 µM corresponding to the complex between Mel and the two subunits Dm and Em, Fig. S2. Beyond demonstrating the Mel/aptamer induced allosteric activation of the Mg2+-ion-dependent DNAzyme, the system presents as an amplified Mel sensing platform. Mel sensing is of importance as Mel has been used as an illegal additive in food products causing severe health problems in infants.53 Different analytical methods including mass-spectrometry,54 high performance liquid chromatography55 and CRISPR/Cas14a sensing platforms56 were developed to detect Mel residues in food products. The amplified Mel-dependent operation of the DNAzyme allows the analysis of Mel with a detection limit corresponding to 0.1 ppm, that is lower than the detection threshold defined by the FDA for infant formula (1.0 ppm).57
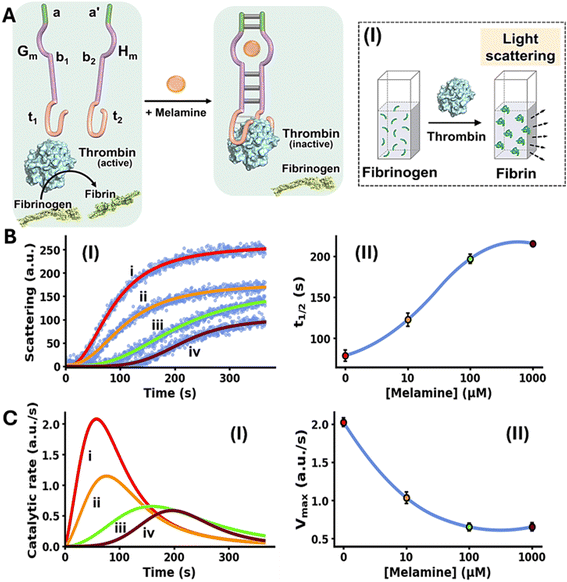
The allosteric Mel/aptamer subunits complex stimulating the catalytic functions of a conjugated DNA framework were further demonstrated with the Mel-mediated inhibition of fibrinogen to fibrin coagulation. Thrombin is a key physiological regulator of the blood clotting mechanism.58 While it plays a key role in hemostatic clotting of vascular injuries, its balanced dose activity is crucial to prevent blood clots and thrombosis.59,60 Diverse anti-coagulant therapeutic agents, controlling thrombin activity are known.61 Within these efforts, anti-thrombin aptamers that bind to thrombin were isolated and their inhibition of thrombin was implemented to design anti-coagulant agents.62,63 The dose-controlled, inhibition of thrombin catalytic functions by aptamers, acting as anti-coagulation agents, could thus be a significant advance in controlling blood clotting (thrombosis). Accordingly, the allosteric activation of the blood clotting inhibitory capacity of the anti-thrombin aptamer framework using an auxiliary ligand, e.g. Mel, could be an interesting path to follow. This concept is exemplified in Fig. 2A with the design of a Mel/aptamer subunits allosteric circuit for the controlled inhibition of thrombin's coagulation function.
The system consists of two strands Gm and Hm composed of the subsequences b1 and b2 corresponding to the Mel aptamer subunits, extended by four base complementary tethers, a and a′, cooperatively enhancing the stability of the Mel/aptamer subunits complex. The Mel aptamer subunits Gm and Hm are further extended by the tethers t1 and t2 that correspond to the anti-thrombin aptamer subunits. Despite the partial complementarity of strands Gm and Hm and the binding affinity of thrombin to the subunits t1 and t2, the strands Gm and Hm lack binding affinity to allow the formation of the Gm/Hm/thrombin complex, and thus, the inhibition of thrombin-induced fibrinogenesis (coagulation of fibrinogen to fibrin) is prohibited. The rate of thrombin-induced coagulation in the absence of the strands Gm/Hm yet in the presence of Mel, is reflected by the temporal light-scattering of the system, show similar temporal light-scattering intensity changes to free thrombin in the absence of ligands, Fig. S4. The allosteric Mel/aptamer subunits stabilization of the thrombin/anti-thrombin subunits results in the Mel-induced, controlled inhibition of the thrombin-induced coagulation of fibrinogen to fibrin. The rate of fibrinogenesis is followed by the temporal light-scattering features associated with the coagulation of fibrinogen to fibrin, in the presence of variable Mel concentrations, Fig. 2A, panel I. Fig. 2B, panel I, depicts the time-dependent light-scattering intensity changes associated with the thrombin-induced coagulation of fibrinogen to fibrin in the presence of strands Gm and Hm, in the absence of Mel, curve (i), and in the presence of variable concentrations of Mel, curves (ii)–(iv). While in the absence of Mel rapid fibrinogenesis is observed, the addition of Mel induces the formation of interstrand Mel/Gm/Hm and the coagulation of fibrinogen to fibrin is suppressed, thus as the concentration of Mel increases the degree of inhibition of fibrinogenesis is higher. The temporal light-scattering curves probing the allosteric inhibition efficacy of fibrinogenesis, in the presence of variable concentrations of Mel were quantitatively evaluated using two parameters.64,65 One parameter, t1/2, is the time interval corresponding to the light-scattering intensity reaching 50% of the saturation value at variable concentrations of Mel. Fig. 2B, panel II, depicts the relation of t1/2 to the concentrations of Mel inducing allosterically the inhibition of thrombin. As the concentration of Mel increases the t1/2 light-scattering intensity values are higher reflecting an enhanced thrombin inhibition capability of the circuit. A second parameter evaluating the Mel-induced inhibition of thrombin is the maximum coagulation rates of fibrinogen to fibrin (Vmax) that are derived from the temporal light-scattering curves in the presence of variable Mel concentrations, first order time-dependent derivative curves are shown in Fig. 2C, panel I. The Vmax values of the circuit, characterizing the inhibition efficacy induced by Mel, derived from the temporal light-scattering curves shown in Fig. 2B, panel I, are displayed in Fig. 2C, panel II. While a high Vmax in the absence of Mel is observed, reflecting low thrombin inhibition, the Vmax values decrease as the concentration of Mel increases, demonstrating the enhanced efficiency of Mel-induced inhibition of coagulation of fibrinogen to fibrin. The results displayed in Fig. 2 introduce a new paradigm for controlling thrombin-induced coagulation by employing an auxiliary ligand (Mel) that allosterically regulates the dose-controlled formation of the thrombin/anti-thrombin aptamer subunits affinity complex that inhibits the coagulation process.
In the next step, the allosteric Mel-induced activation of a transcription machinery was examined. Regulation of RNA transcription controls many biological processes ranging from cell cycle progression66 and maintenance of intracellular metabolism to cellular differentiation.67 The transcription apparatus demonstrates dynamic adaptive features, primarily modulated by transcription factors.68–71 Beyond the key functions of the native transcription machinery in maintenance of living organisms, misregulation of transcription programs by dysfunctional transcription factors is the origin of various diseases including cancer, viral infection, neurological disorders, autoimmune pathologies and diabetes.72,73 Development of biomimetic synthetic transcription circuits is important not only to emulate the native apparatus by artificial model systems, but it could provide versatile therapeutic applications. For example, the controlled programmed in vivo synthesis of pre-engineered RNA could be a valuable source of therapeutic agents, e.g. aptamers, siRNAs, and ribozymes. Indeed, recent research efforts demonstrated the modulation of transcription machineries by topological nucleic acid barriers conjugated to transcription templates, such as G-quadruplexes or DNA triplexes modeling native transcription factors' functions.64 In the forthcoming section we introduce the allosteric Mel-induced operation of a transcription circuit as a biomimetic model system emulating the functions of transcription factors.
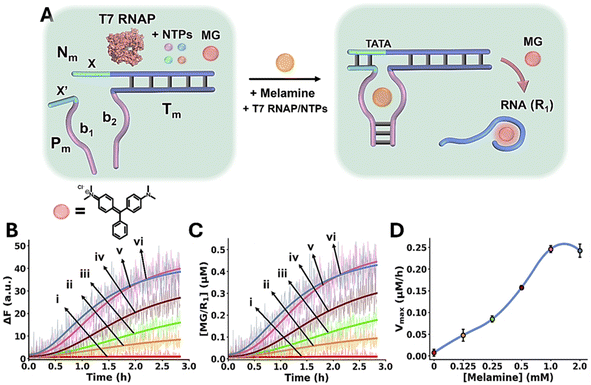
The Mel/aptamer subunits complex triggered activation of the transcription machinery is schematically displayed in Fig. 3A. The inactive reaction module consists of the template strands Nm/Tm containing an incomplete T7 RNA polymerase (RNAP) promoter, the strand Pm, Malachite Green (MG) and T7 RNAP/ribonucleotide triphosphates (NTPs) mixture. The strands Tm and Pm include tethers b1 and b2 corresponding to the Mel aptamer subunits, where b1 is extended by the sequence x′ that is complementary to domain x in the template Nm/Tm. While x′ contains the sequence to complete the promoter region that activates the Nm/Tm transcription machinery, the stability of the complementary duplex x/x′ is, however, insufficient to activate the transcription machinery. In the presence of Mel, the cooperative formation of the Mel/aptamer subunits complex, and the duplex x/x′ form an energetically stabilized, promoter-activated, transcription template enabling the activation of the transcription machinery, resulting in the T7 catalyzed RNAP/NTPs transcription of the RNA product, R1. The template Nm/Tm is pre-engineered to yield the MG RNA aptamer as the transcription product. The resulting fluorescent MG/RNA aptamer complex (λex = 632 nm; λem = 650 nm) provides, then, an optical readout signal for the temporal operation of the transcription machinery. Fig. 3B depicts the time-dependent fluorescence changes caused by the production of the MG RNA aptamer product, generated in the absence of Mel, curve i, and in the presence of variable concentrations of Mel, curves ii–vi. Using an appropriate calibration curve, relating the fluorescence intensity of MG/RNA aptamer to its concentration, Fig. S5, the rates of MG/RNA aptamer formation as a function of Mel concentrations were evaluated as depicted in Fig. 3C. Peak rates (Vmax) of the transcription template derived from Fig. 3C are displayed in Fig. 3D, as Mel concentration increases, Vmax increases.
The systems discussed so far demonstrated the allosteric ligand (Mel)-induced activation of catalytic processes involving a DNAzyme, thrombin-induced fibrinogenesis or a transcription machinery. Many of the catalytic processes in nature are, however, temporally modulated, leading to dissipative, transient and out-of-equilibrium operation.74 The ligand/aptamer complex allosterically regulating catalytic processes introduced a mechanism controlling the “dose” of the catalytic transformation. The coupling of a catalyst modulating temporally and transiently the allosteric mechanism could introduce an additional dimension to the “dose” regulated control over catalytic processes. The principle of dissipative out-of-equilibrium systems involves the design of reaction circuits that are activated by an auxiliary energy-fueled input (chemical fuel, light, electrical or magnetic stimuli) that generate an intermediate out-of-equilibrium state. The system includes, however, an internal mechanism depleting the auxiliary energy-fueled input resulting in the degradation of the intermediate state, into waste products while recovering the parent circuit. This leads to the temporal, transient, formation and depletion of the intermediate state. Substantial recent research efforts addressed the use of nucleic acid-based frameworks as functional reaction modules to design transient DNA circuits.75,76 Different triggers including nucleic acid fuel strands or light coupled to enzymes or DNAzymes were employed to trigger the temporal transitions of DNA frameworks into intermediate states that are temporally depleted by the catalysts to the parent reaction modules, thereby establishing transiently operating DNA circuits.77,78 Diverse applications of transient operating circuits were demonstrated, including transient operating biocatalytic cascades,79 transient DNA-based load release systems,80 or transient nucleic acid guided aggregation/de-aggregation of metal nanoparticles or semiconductor quantum dots.81 The allosteric ligand/aptamer stabilization of catalytic frameworks, and the availability of enzymes degrading the ligands, suggests that coupling of enzymes to the allosterically ligand/aptamer stabilized catalytic frameworks could provide a versatile mechanism to engineer dynamic, transient allosterically ligand-stabilized reaction modules. For example, diverse ligand/aptamer complexes can be coupled with ligand degrading enzymes (e.g., adenosine/adenosine deaminase, acetylcholine/acetylcholinesterase, uric acid/uricase) resulting in the separation of the complexes, as schematically shown in Fig. S6.
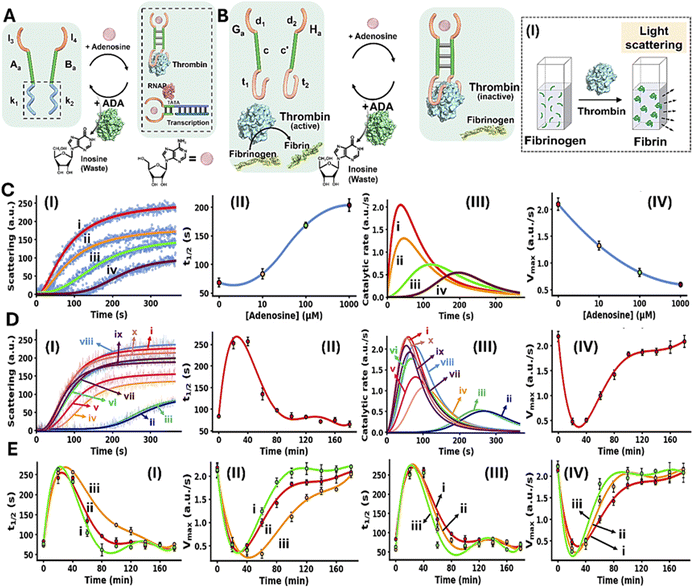
The conjugation of an enzyme to an allosteric ligand-induced catalytic transformation leading to transient allosterically-driven catalytic process is schematically presented in Fig. 4A, using the adenosine (Ade)/aptamer subunits complex and adenosine deaminase (ADA) as regulators controlling allosteric catalytic processes such as fibrinogenesis or transcription (for the design principles of the allosteric Ade/aptamer subunits inhibition of thrombin see Fig. S3 and accompanying discussion Page S11 in the SI). The two strands Aa and Ba coupled with ADA, act as the reaction module. The strands Aa and Ba include the Ade aptamer subunits l3 and l4 conjugated to strands k1 and k2 that encode the nucleic acid sequences comprising the catalytic frameworks. Fueling the system with Ade results in the formation of Ade/aptamer subunits supramolecular complex allosterically stabilizing the catalytic frameworks consisting of the thrombin/anti-thrombin aptamer subunits complex inhibiting fibrinogenesis, or the active transcription machinery as intermediate products. The ADA present in the system concomitantly transforms Ade to inosine, that lacks affinity towards the aptamer subunits. Separation of the Ade/Ade aptamer complex recovers the parent reaction circuit, in which the catalytic transformations are prohibited. This leads to the transient Ade/ADA allosteric operation of fibrinogenesis or transcription processes. In the forthcoming section, the transient Ade/ADA operation of transcription and fibrinogenesis machineries will be addressed.
Fig. 4B depicts schematically the reaction circuit used for the allosteric transient inhibition of the thrombin-induced coagulation of fibrinogen to fibrin in the presence of Ade and ADA. The system consists of two strands Ga and Ha as the DNA functional framework and ADA as the auxiliary catalyst. The strands Ga and Ha include the Ade aptamer subunits d1 and d2 conjugated to the complementary sequences c and c′ and extended by the anti-thrombin aptamer subunits t1 and t2. While in the absence of Ade, the base complementarity of c and c′ is insufficient to form a stable Ga/Ha duplex structure that binds to thrombin and inhibits the thrombin-induced fibrinogenesis. The addition of Ade leads to the formation of the Ade/aptamer subunits complex, cooperatively stabilized by the c/c′ duplex, leading to the assembly of the supramolecular Ga/Ha complex that stabilizes the thrombin aptamer subunits allowing the association of the interstrand Ade/Ga/Ha supramolecular thrombin aptamer subunits framework to thrombin. Binding of the interstrand Ade/Ga/Ha to thrombin allosterically inhibits thrombin-induced fibrinogenesis. The ADA coupled to the reaction circuit, concomitantly transforms Ade to inosine, leading to the separation of the Ga/Ha units from thrombin, thereby recovering the free thrombin, exhibiting non-inhibited coagulation rate. That is, the system reveals transient allosteric Ade-induced inhibition of thrombin-induced coagulation of fibrinogen to fibrin. The degree of inhibition is controlled by the concentration of Ade that regulates the allosteric formation of the Ade/aptamer subunits complex. While the ADA concentration dictates the rate of recovery of the parent module, showing non-inhibited fibrinogenesis. That is, the transient allosteric inhibition of thrombin is regulated by two parameters; the concentration of Ade that triggers the fibrinogenesis inhibition and the concentration of ADA degrading Ade thereby regulating the temporal depletion of the inhibition phenomenon. The temporal and transient allosteric inhibition of thrombin, is then, followed by the temporal light-scattering features associated with the coagulation of fibrinogen to fibrin, in the presence of variable Ade and ADA concentrations, Fig. 4B, panel I. The rate of thrombin-induced coagulation in the absence of the strands Ga/Ha, yet in the presence of Ade, show similar temporal light-scattering intensity changes to free thrombin in the absence of ligands, Fig. S4.
Fig. 4C, panel I, shows the temporal light-scattering intensities associated with the fibrinogenesis in the presence of different concentrations of Ade, yet in the absence of ADA: (i) 0 µM (ii) 10 µM (iii) 100 µM (iv) 1000 µM. As the concentration of Ade increases, the allosteric inhibition of the thrombin-induced fibrinogenesis increases as reflected by a prolonged initial lag and lower saturation values of temporal light-scattering curves. The analysis of the temporal light-scattering curves in panel I, in terms of t1/2, temporal catalytic rates and Vmax values are presented in Fig. 4C panels II-IV. Fig. 4D, panel I shows the temporal light-scattering curves associated with the thrombin-induced fibrinogenesis at time-intervals of the system's operation using Ade = 1.5 mM; ADA = 0.045 U ml−1. While the control of the system, curve (i), in the absence of Ade demonstrates non-inhibited fibrinogenesis, addition of Ade leads to effective inhibition of fibrinogenesis, curve (ii), the system reveals, however, a continuous temporal change in the rate of fibrinogenesis reflected by the decrease in the inhibition effect in the system and ultimately shows the recovery of the non-inhibited fibrinogenesis behavior of the system, curves (ii)–(x). Analysis of the temporal light-scattering curves shown in Fig. 4D panel I in terms of t1/2, temporal catalytic rates and Vmax are summarized in Fig. 4D, panels II–IV. The result demonstrates the transient allosteric Ade-induced inhibition of thrombin by Ade/ADA in the Ga/Ha reaction circuit. Fig. 4E depicts the effects of different Ade and ADA concentrations on the transient allosteric inhibition of thrombin-induced fibrinogenesis, as reflected by the t1/2 and Vmax values derived from the temporal light-scattering intensities in the respective Ga/Ha operating system. As the concentration of Ade increases, at fixed ADA concentration 0.045 U ml−1, the time-interval of the transient recovery of the allosterically inhibited thrombin-induced fibrinogenesis is prolonged, Fig. 4E, panels I and II. In addition, as the ADA concentration increases, at a fixed Ade concentration 1.5 mM, the time-interval of the transient recovery of the non-inhibited thrombin-induced fibrinogenesis is shortened, Fig. 4E, panels III and IV. These results are consistent with the allosteric transient inhibition of the thrombin-induced coagulation of fibrinogen to fibrin regulated by the Ade/ADA-Ga/Ha reaction circuit.
An additional process demonstrating the transient allosteric activation of a biocatalytic process included the Ade/ADA allosteric transient activation of a transcription machinery. Beyond transcription factor-modulated transcription machineries and accompanying regulated gene expression, promoter control elements regulate the dynamic interactions of transcription factors with the transcription machineries. These include for example, enhancer,82 silencer83 or switching elements,84 leading to temporal dynamic modulation of transcription and gene expression. For example, by coupling two mutually repressing transcription factor pathways, biomimetic oscillatory85 or bistable86 active gene expression were reported. Also, transient transcription machineries regulated by auxiliary enzymes87 or DNAzymes88 were reported. The allosteric ligand/aptamer transient activation of a transcription process is, to the best of our knowledge, unprecedented.
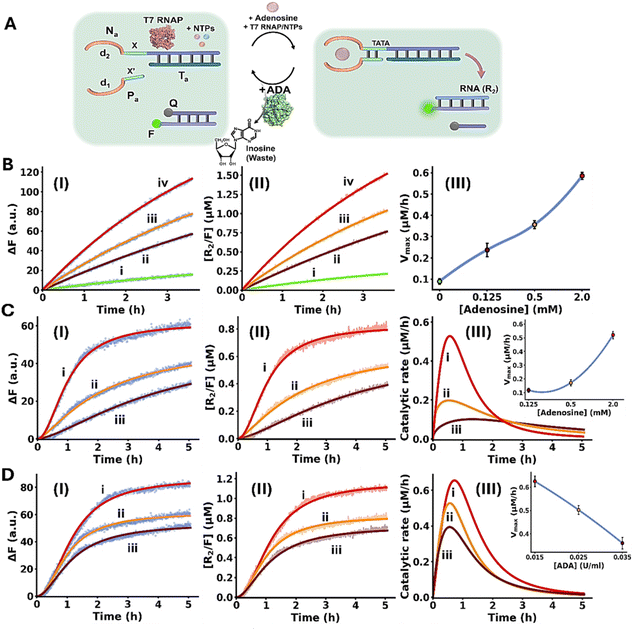
Fig. 5A depicts schematically the transient Ade/ADA allosteric operation of a transcription machinery. The reaction module consists of the strands Na/Ta that exist as an inactive transcription template lacking a full T7 promoter sequence. The strand Pa contains the sequence x′ that hybridizes to the sequence x in strand Na and completes the T7 promoter, however despite the complementarity of the domains x/x′ they are pre-engineered to form a non-stable five-base duplex. To assist the binding of Pa to the transcription template, Ade aptamer subunits, d1 and d2, were conjugated to strands Pa and Na. Added Ade cooperatively stabilize the formation of the active transcription template by the cooperative formation of the Ade/aptamer subunits complex and the x/x′ duplex completing the promoter domain in the transcription template. Formation of the intact Ade-stabilized transcription template activates, then, the T7 RNAP/NTPs transcription machinery transcribing the RNA product, R2. Since the affinity of the Ade aptamer towards other adenine-containing ligands is well established,89 ATP was excluded from the reaction circuit. The transcription template is pre-engineered to displace, by the transcribed RNA, R2, the auxiliary transducing F/Q DNA duplex that is comprised of a fluorophore-modified (FAM) strand and a quencher-modified (BHQ1) strand, where the fluorescent signal is effectively quenched, generating the fluorescent F/R2 duplex and the free quencher-modified strand Q. Thus, the time-dependent fluorescence changes in the system reflect the temporal performance of the transcription process induced by Ade. The ADA integrated in the circuit, temporally transforms Ade to inosine resulting in the depletion of Ade, leading to the separation of the strand Pa from the transcription template, that transiently recovers the parent reaction circuit. Thus, the ADA present in the system induces the transient dissipative evolution of the fluorescent intermediate F/R2 duplex in the system. The temporal fluorescence changes caused by the displacement induced by the RNA product reflect, then the dynamically modulated transcription occurring in the system. Fig. 5B, panels I–III, depict the time-dependent fluorescence changes generated by the reaction module in the presence of different concentrations of Ade, yet in the absence of ADA: (i) 0 mM (ii) 0.125 mM (iii) 0.5 mM (iv) 2.0 mM. While in the absence of Ade minimal transcription of R2 occurs (reflected by the lack of separation of the F/Q-transducing duplex), curve i, the time-dependent fluorescence changes are intensified as the concentration of Ade increases, curves ii–iv. Using an appropriate calibration curve relating the fluorescence of the released FAM-labeled DNA strand as a function of R2 concentration, Fig. S7, the temporal transcribed RNA product R2 in the presence of different Ade concentrations is displayed in Fig. 5B, panel II. Temporal first order maximum catalytic rates (Vmax) of the transcription template derived from panel II are displayed in panel III. The results confirm that Ade stabilizes allosterically the assembly of the promoter/transcription template Pa/Na/Ta transcribing the product R2. Fig. 5C depicts the temporal fluorescence changes generated by the transcription machinery, in the presence of variable Ade concentrations and a constant concentration of ADA = 0.025 U ml−1. The temporal fluorescence changes, reflecting the rate of R2 production reveal a non-linear behavior tending to reach a saturation value. The temporal fluorescence changes and the resulting saturation levels are controlled by the concentration of Ade. As the concentration of Ade increases, the intensities of the fluorescence changes are higher, consistent with the increased concentration of the Ade/aptamer subunits stabilized active transcription template. The non-linear temporal fluorescence intensity changes support that an accompanying mechanism slowing down the transcription process exists in the system, consistent with the ADA-induced depletion of the transcription machinery. Using a calibration curve relating the fluorescence intensities to the R2 RNA concentrations, Fig. S7, the temporal concentrations of the transcribed R2 were evaluated, Fig. 5C, panel II. The temporal catalytic rates (first order derivatives) of the transcription machinery, in the presence of different Ade concentrations, are displayed in panel III. Dissipative, transient catalytic rates revealing peak rates (Vmax) controlled by concentrations of Ade are observed (inset, panel III). Fig. 5D, panel I depicts the temporal fluorescence changes associated with R2 transcription, in the presence of variable ADA concentrations, and a fixed concentration of Ade = 2.0 mM. As the concentration of ADA increases, the maximal fluorescence level is lower. Fig. 5D, panel II shows the temporal concentration changes of the transcribed R2, and panel III displays the temporal catalytic rates of the transcription machinery. As the concentration of ADA increases Vmax lowers and the dissipative depletion of the transcription process is faster (inset, panel III). These results are consistent with the faster depletion of the allosteric Ade/aptamer subunits stabilized transcription template, as the concentration of ADA increases.
Conclusions
The study introduced allosteric ligand/aptamer complexes as functional structures orchestrating dynamic or transient catalytic DNA-based frameworks. These included the melamine (Mel)/aptamer subunits allosteric activation of the Mg2+-ion-dependent DNAzyme, the allosteric inhibition of thrombin and the allosteric operation of a transcription machinery. Moreover, by coupling the allosteric ligand/aptamer complex induced stabilization of the catalytic transformation to an auxiliary enzyme depleting the ligand, the transient, dissipative, operation of catalytic transformations was demonstrated. This was exemplified by the allosteric adenosine (Ade)/aptamer complex control over fibrinogenesis or transcription machineries and the transformation of these processes into dissipative and transient pathways in the presence of adenosine deaminase (ADA). Beyond mimicking native processes by synthetic circuits, such as transcription factor guided transcription machineries, the systems might be used for amplified sensing and biomedical applications, such as dose-controlled fibrinogenesis (blood coagulation) or a biomarker-induced synthesis of RNA inhibiting aptamers. The significance of the study rests, however, on the potential generalization of the allosteric stabilization concept of DNAzyme or transcription therapeutic circuits by other ligands, and particularly designing applications of such frameworks. Besides the use of Mel or Ade as allosteric activators of DNAzymes, protein biomarkers (e.g., VEGF43) or specific miRNAs could promote the allosteric biomarker-driven cleavage of mRNAs leading to autonomous gene therapy (for example, inducing cancer cell apoptosis by specific miRNAs).96 Also, biomarker triggered transcription machineries could be extended to miRNA or protein-stabilized transcription templates synthesizing pre-designed aptamers with therapeutic inhibiting functions.97 Furthermore, the transient operation of allosterically-driven DNA circuits could be extended to miRNA driven allosteric frameworks, using RNase H as the catalytic agent or by implementing other ligand/enzyme couples such as xanthine90/xanthine oxidase or uric acid91/uricase (applicable for Gout-related inflammation) or acetylcholine92/acetylcholinesterase (for the temporal treatment of neural diseases). Moreover, at present, all systems were operated in homogeneous solutions. Integration of the circuits within liposomes could yield functional circuit-loaded synthetic cells-protocells,93–95 and fusion between the liposomes and native cells could provide versatile means to deliver the loads into the cells thereby signaling cell functions by artificial circuits.50Author contributions
I. Willner and J. Dong designed the experiments and supervised the research. D. Froim carried out the experiments and conducted corresponding data analysis. H. Amartely contributed to the modules' characterization experiments. E. Pikarsky provided valuable insights to the topic of research. All authors reviewed and approved the final manuscript.Conflicts of interest
The authors declare no competing financial interests.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: reagents, instrumentation, experimental methods, calibration curves and ITC data. See DOI: https://doi.org/10.1039/d5sc09098a.Acknowledgements
The research was supported by the Israel Science Foundation (ISF) Precision Medicine Program, Grant No. 1696/20.References
- R. Nussinov, Introduction to Protein Ensembles and Allostery, Chem. Rev., 2016, 116, 6263–6266 Search PubMed
.
- J. Liu and R. Nussinov, Allostery: An overview of its history, concepts, methods, and applications, PLoS Comput. Biol., 2016, 12, e1004966 CrossRef PubMed
.
-
J. Monod, Chance and Necessity: Essay on the Natural Philosophy of Modern Biology, Vintage Books, New York, 1972 Search PubMed
.
- S. E. Osborne and A. D. Ellington, Nucleic acid selection and the challenge of combinatorial chemistry, Chem. Rev., 1997, 97, 349–370 CrossRef CAS PubMed
.
- M. Famulok and G. Mayer, Aptamer modules as sensors and detectors, Acc. Chem. Res., 2011, 44, 1349–1358 CrossRef CAS
.
- J. Lee, G. Stovall and A. Ellington, Aptamer therapeutics advance, Curr. Opin. Chem. Biol., 2006, 10, 282–289 Search PubMed
.
- F. C. Simmel, B. Yurke and H. R. Singh, Principles and applications of nucleic acid strand displacement reactions, Chem. Rev., 2019, 119, 6326–6369 CrossRef CAS PubMed
.
- D. Y. Zhang and G. Seelig, Dynamic DNA nanotechnology using strand-displacement reactions, Nat. Chem., 2011, 3, 103–113 CrossRef CAS PubMed
.
- D. Y. Zhang, A. J. Turberfield, B. Yurke and E. Winfree, Engineering Entropy-Driven reactions and networks catalyzed by DNA, Science, 2007, 318, 1121–1125 CrossRef CAS PubMed
.
- G. F. Joyce, Directed evolution of nucleic acid enzymes, Annu. Rev. Biochem., 2004, 73, 791–836 CrossRef CAS PubMed
.
- R. R. Breaker and G. F. Joyce, A DNA enzyme that cleaves RNA, Chem. Biol., 1994, 1, 223–229 Search PubMed
.
- S. K. Silverman, In vitro selection, characterization, and application of deoxyribozymes that cleave RNA, Nucleic Acids Res., 2005, 33, 6151–6163 CrossRef CAS PubMed
.
- S. V. Park, J.-S. Yang, H. Jo, B. Kang, S. S. Oh and G. Y. Jung, Catalytic RNA, ribozyme, and its applications in synthetic biology, Biotechnol. Adv., 2019, 37, 107452 Search PubMed
.
- P. Travascio, Y. Li and D. Sen, DNA-enhanced peroxidase activity of a DNA aptamer-hemin complex, Chem. Biol., 1998, 5, 505–517 Search PubMed
.
- A. F. Fagbemi, B. Orelli and O. D. Schärer, Regulation of endonuclease activity in human nucleotide excision repair, DNA Repair, 2011, 10, 722–729 Search PubMed
.
- P. Gao, H. Yang, K. R. Rajashankar, Z. Huang and D. J. Patel, Type V CRISPR-Cas Cpf1 endonuclease employs a unique mechanism for crRNA-mediated target DNA recognition, Cell Res., 2016, 26, 901–913 Search PubMed
.
- J. Bath, S. J. Green and A. J. Turberfield, A Free-Running DNA motor powered by a nicking enzyme, Angew. Chem., Int. Ed., 2005, 44, 4358–4361 Search PubMed
.
- Z. Li, J. Wang and I. Willner, Autoinhibited transient, gated, and cascaded dynamic transcription of RNAs, Sci. Adv., 2022, 8, eabq5947 Search PubMed
.
- Y. Weizmann, M. K. Beissenhirtz, Z. Cheglakov, R. Nowarski, M. Kotler and I. Willner, A virus spotlighted by an autonomous DNA machine, Angew. Chem., Int. Ed., 2006, 45, 7384–7388 CrossRef CAS PubMed
.
- N. C. Seeman and H. F. Sleiman, DNA nanotechnology, Nat. Rev. Mater., 2017, 3, 17068 CrossRef
.
- F. Wang, X. Liu and I. Willner, DNA switches: From principles to applications, Angew. Chem., Int. Ed., 2014, 54, 1098–1129 Search PubMed
.
- S. G. Harroun, C. Prévost-Tremblay, D. Lauzon, A. Desrosiers, X. Wang, L. Pedro and A. Vallée-Bélisle, Programmable DNA switches and their applications, Nanoscale, 2018, 10, 4607–4641 Search PubMed
.
- X. Liu, C.-H. Lu and I. Willner, Switchable reconfiguration of nucleic acid nanostructures by Stimuli-Responsive DNA machines, Acc. Chem.
Res., 2014, 47, 1673–1680 CrossRef CAS PubMed
.
- J. Bath and A. J. Turberfield, DNA nanomachines, Nat. Nanotechnol., 2007, 2, 275–284 CrossRef CAS
.
- F. Hong, F. Zhang, Y. Liu and H. Yan, DNA Origami: scaffolds for creating higher order structures, Chem. Rev., 2017, 117, 12584–12640 CrossRef CAS PubMed
.
- T. Tørring, N. V. Voigt, J. Nangreave, H. Yan and K. V. Gothelf, DNA origami: a quantum leap for self-assembly of complex structures, Chem. Soc. Rev., 2011, 40, 5636 Search PubMed
.
- J. Li, A. A. Green, H. Yan and C. Fan, Engineering nucleic acid structures for programmable molecular circuitry and intracellular biocomputation, Nat. Chem., 2017, 9, 1056–1067 Search PubMed
.
- G. Seelig, D. Soloveichik, D. Y. Zhang and E. Winfree, Enzyme-Free Nucleic Acid Logic Circuits, Science, 2006, 314, 1585–1588 CrossRef CAS PubMed
.
- L. Yue, S. Wang, Z. Zhou and I. Willner, Nucleic acid based constitutional Dynamic networks: From basic principles to applications, J. Am. Chem. Soc., 2020, 142, 21577–21594 Search PubMed
.
- E. Del Grosso, E. Franco, L. J. Prins and F. Ricci, Dissipative DNA nanotechnology, Nat. Chem., 2022, 14, 600–613 CrossRef CAS
.
- Z. Li, J. Wang and I. Willner, Transient Out-of-Equilibrium nucleic Acid-Based dissipative networks and their applications, Adv. Funct. Mater., 2022, 32, 2200799 Search PubMed
.
- K. Jiao, B. Zhu, L. Guo, H. Zhou, F. Wang, X. Zhang, J. Shi, Q. Li, L. Wang, J. Li and C. Fan, Programming switchable transcription of topologically constrained DNA, J. Am. Chem. Soc., 2020, 142, 10739–10746 CrossRef CAS PubMed
.
- P. Zhang, A. Fischer, Y. Ouyang, Y. S. Sohn, R. Nechushtai, J. Zhang, H. Tian, C. Fan and I. Willner, Topologically switchable and gated transcription machinery, Chem. Sci., 2022, 13, 10555–10565 RSC
.
- S. Xie, L. Qiu, L. Cui, H. Liu, Y. Sun, H. Liang, D. Ding, L. He, H. Liu, J. Zhang, Z. Chen, X. Zhang and W. Tan, Reversible and quantitative photoregulation of target proteins, Chem, 2017, 3, 1021–1035 CAS
.
- Y. Biniuri, G.-F. Luo, M. Fadeev, V. Wulf and I. Willner, Redox-Switchable binding properties of the ATP–Aptamer, J. Am. Chem. Soc., 2019, 141, 15567–15576 CrossRef CAS
.
- E. Golub, H. B. Albada, W.-C. Liao, Y. Biniuri and I. Willner, Nucleoapzymes: Hemin/G-Quadruplex DNAzyme–Aptamer Binding Site Conjugates with Superior Enzyme-like Catalytic Functions, J. Am. Chem. Soc., 2015, 138, 164–172 CrossRef PubMed
.
- Y. Ouyang, P. Zhang and I. Willner, Dynamic catalysis guided by nucleic acid networks and DNA nanostructures, Bioconjugate Chem., 2022, 34, 51–69 CrossRef PubMed
.
- Z. Huang, D. Wang, Q. Zhang, Y. Zhang, R. Peng and W. Tan, Leveraging Aptamer-Based DNA nanotechnology for bioanalysis and cancer therapeutics, Acc. Mater. Res., 2024, 5, 438–452 CrossRef CAS
.
- I. Willner and M. Zayats, Electronic Aptamer-Based sensors, Angew. Chem., Int. Ed., 2007, 46, 6408–6418 CrossRef CAS PubMed
.
- H.-M. Meng, H. Liu, H. Kuai, R. Peng, L. Mo and X.-B. Zhang, Aptamer-integrated DNA nanostructures for biosensing, bioimaging and cancer therapy, Chem. Soc. Rev., 2016, 45, 2583–2602 RSC
.
- C. Ji, J. Wei, L. Zhang, X. Hou, J. Tan, Q. Yuan and W. Tan, Aptamer–Protein interactions: From regulation to biomolecular detection, Chem. Rev., 2023, 123, 12471–12506 CrossRef CAS PubMed
.
- W.-H. Chen, G.-F. Luo, M. Vázquez-González, R. Cazelles, Y. S. Sohn, R. Nechushtai, Y. Mandel and I. Willner, Glucose-Responsive Metal–Organic-Framework nanoparticles act as “Smart” Sense-and-Treat carriers, ACS Nano, 2018, 12, 7538–7545 CrossRef CAS PubMed
.
- E. W. M. Ng, D. T. Shima, P. Calias, E. T. Cunningham, D. R. Guyer and A. P. Adamis, Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease, Nat. Rev. Drug Discovery, 2006, 5, 123–132 CrossRef CAS PubMed
.
- B. Deng, Y. Lin, C. Wang, F. Li, Z. Wang, H. Zhang, X.-F. Li and X. C. Le, Aptamer binding assays for proteins: The thrombin example—A review, Anal. Chim. Acta, 2014, 837, 1–15 Search PubMed
.
- J. Achenbach, W. Chiuman, R. Cruz and Y. Li, DNAZymes: From creation in vitro to application in vivo, Curr. Pharm. Biotechnol., 2004, 5, 321–336 CAS
.
- E. Golub, R. Freeman and I. Willner, A Hemin/G-Quadruplex acts as an NADH oxidase and NADH peroxidase mimicking DNAzyme, Angew. Chem., Int. Ed., 2011, 50, 11710–11714 CrossRef CAS
.
- E. Golub, H. B. Albada, W.-C. Liao, Y. Biniuri and I. Willner, Nucleoapzymes: Hemin/G-Quadruplex DNAzyme–Aptamer Binding Site Conjugates with Superior Enzyme-like Catalytic Functions, J. Am. Chem. Soc., 2015, 138, 164–172 Search PubMed
.
- N. Lin, Y. Ouyang, Y. Qin, O. Karmi, Y. S. Sohn, S. Liu, R. Nechushtai, Y. Zhang, I. Willner and Z. Zhou, Spatially localized Entropy-Driven evolution of nucleic Acid-Based constitutional dynamic networks for intracellular imaging and spatiotemporal programmable gene therapy, J. Am. Chem. Soc., 2024, 146, 20685–20699 Search PubMed
.
- Z. Tang, Y. Liu, M. He and W. Bu, Chemodynamic therapy: tumour Microenvironment-Mediated Fenton and Fenton-like reactions, Angew. Chem., Int. Ed., 2018, 58, 946–956 Search PubMed
.
- Y. Ouyang, Y. S. Sohn, X. Chen, R. Nechushtai, E. Pikarsky, F. Xia, F. Huang and I. Willner, Adenosine-Triggered dynamic and transient Aptamer-Based networks integrated in liposome protocell assemblies, J. Am. Chem. Soc., 2025, 147, 19282–19295 CrossRef CAS PubMed
.
- C. Gu, T. Lan, H. Shi and Y. Lu, Portable detection of melamine in milk using a personal glucose meter based on an in vitro selected Structure-Switching Aptamer, Anal. Chem., 2015, 87, 7676–7682 Search PubMed
.
- C. Gu, Y. Xiang, H. Guo and H. Shi, Label-free fluorescence detection of melamine with a truncated aptamer, Analyst, 2016, 141, 4511–4517 RSC
.
- Q. Li, P. Song and J. Wen, Melamine and food safety: a 10-year review, Curr. Opin. Food Sci., 2019, 30, 79–84 CrossRef
.
- E. D. C. Domingo, A. A. Tireli, C. A. Nunes, A. V. Batista, M. C. Guerreiro and S. M. Pinto, Rapid extraction of melamine in powdered milk for direct electrospray ionization tandem mass spectrometry analysis, Talanta, 2014, 132, 535–540 CrossRef PubMed
.
- A. M. R. Mondal, A. Desmarchelier, E. Konings, R. Acheson-Shalom and T. Delatour, Liquid Chromatography−Tandem Mass Spectrometry (LC−MS/MS) method extension to quantify simultaneously melamine and cyanuric acid in egg powder and soy protein in addition to milk products, J. Agric. Food Chem., 2010, 58, 11574–11579 CrossRef
.
- P. Ma, H. Guo, Y. Zhang and Z. Wang, A novel CRISPR/Cas14a1-Exo III aptasensor for melamine detection coupled with systematically studied binding mechanism of truncated aptamer, Sens. Actuators, B, 2022, 374, 132847 CrossRef
.
- U.S. Food and Drug Administration, Chemical Contaminants, Archived from the original on January 11, 2017, https://wayback.archive-it.org/7993/20170111174251/http://www.fda.gov/Food/FoodborneIllnessContaminants/ChemicalContaminants/ucm164520.htm, accessed October 2025.
- J. C. Chapin and K. A. Hajjar, Fibrinolysis and the control of blood coagulation, Blood Rev., 2014, 29, 17–24 Search PubMed
.
- R. I. Litvinov and J. W. Weisel, Blood clot contraction: Mechanisms, pathophysiology, and disease, Res. Pract. Thromb. Haemostasis, 2023, 7, 100023 CrossRef CAS PubMed
.
- A. S. Wolberg, Fibrinogen and fibrin: synthesis, structure, and function in health and disease, J. Thromb. Haemostasis, 2023, 21, 3005–3015 Search PubMed
.
- M. Di Nisio, S. Middeldorp and H. R. Büller, Direct thrombin inhibitors, N. Engl. J. Med., 2005, 353, 1028–1040 CrossRef CAS PubMed
.
- S. Zhao, R. Tian, J. Wu, S. Liu, Y. Wang, M. Wen, Y. Shang, Q. Liu, Y. Li, Y. Guo, Z. Wang, T. Wang, Y. Zhao, H. Zhao, H. Cao, Y. Su, J. Sun, Q. Jiang and B. Ding, A DNA origami-based aptamer nanoarray for potent and reversible anticoagulation in hemodialysis, Nat. Commun., 2021, 12, 358 CrossRef CAS PubMed
.
- C. Riccardi, E. Napolitano, C. Platella, D. Musumeci and D. Montesarchio, G-quadruplex-based aptamers targeting human thrombin: Discovery, chemical modifications and antithrombotic effects, Pharmacol. Ther., 2020, 217, 107649 Search PubMed
.
- J. Dong and I. Willner, Transient transcription machineries modulate dynamic functions of G-Quadruplexes: temporal regulation of biocatalytic circuits, gene replication and transcription, Angew. Chem., Int. Ed., 2023, 62, e202307898 CrossRef CAS PubMed
.
- Y. Ouyang, J. Dong and I. Willner, Dynamic DNA Networks-Guided directional and orthogonal transient biocatalytic cascades, J. Am. Chem. Soc., 2023, 145, 22135–22149 CrossRef CAS PubMed
.
- I. Simon, J. Barnett, N. Hannett, C. T. Harbison, N. J. Rinaldi, T. L. Volkert, J. J. Wyrick, J. Zeitlinger, D. K. Gifford, T. S. Jaakkola and R. A. Young, Serial regulation of transcriptional regulators in the yeast cell cycle, Cell, 2001, 106, 697–708 CrossRef CAS PubMed
.
- D. Accili and K. C. Arden, FOXOS at the crossroads of cellular metabolism, differentiation, and transformation, Cell, 2004, 117, 421–426 Search PubMed
.
- G. L. Hager, J. G. McNally and T. Misteli, Transcription dynamics, Mol. Cell, 2009, 35, 741–753 CrossRef CAS PubMed
.
- T. E. Sztal and D. Y. R. Stainier, Transcriptional adaptation: a mechanism underlying genetic robustness, Development, 2020, 147, dev186452 CrossRef CAS PubMed
.
- J. M. Vaquerizas, S. K. Kummerfeld, S. A. Teichmann and N. M. Luscombe, A census of human transcription factors: function, expression and evolution, Nat. Rev. Genet., 2009, 10, 252–263 CrossRef CAS PubMed
.
- T. L. Lenstra, J. Rodriguez, H. Chen and D. R. Larson, Transcription dynamics in living cells, Annu. Rev. Biophys., 2016, 45, 25–47 Search PubMed
.
- T. I. Lee and R. A. Young, Transcriptional regulation and its misregulation in disease, Cell, 2013, 152, 1237–1251 CrossRef CAS PubMed
.
- S. A. Ament, J. R. Pearl, J. P. Cantle, R. M. Bragg, P. J. Skene, S. R. Coffey, D. E. Bergey, V. C. Wheeler, M. E. MacDonald, N. S. Baliga, J. Rosinski, L. E. Hood, J. B. Carroll and N. D. Price, Transcriptional regulatory networks underlying gene expression changes in Huntington's
disease, Mol. Syst. Biol., 2018, 14, e7435 CrossRef PubMed
.
- G. Ragazzon and L. J. Prins, Energy consumption in chemical fuel-driven self-assembly, Nat. Nanotechnol., 2018, 13, 882–889 CrossRef CAS PubMed
.
- R. P. Goodman, M. Heilemann, S. Doose, C. M. Erben, A. N. Kapanidis and A. J. Turberfield, Reconfigurable, braced, three-dimensional DNA nanostructures, Nat. Nanotechnol., 2008, 3, 93–96 CrossRef CAS PubMed
.
- J. Wang, Z. Li and I. Willner, Dynamic reconfigurable DNA nanostructures, networks and materials, Angew. Chem., Int. Ed., 2023, 62, e202215332 CrossRef CAS PubMed
.
- Z. Zhou, Y. Ouyang, J. Wang and I. Willner, Dissipative gated and cascaded DNA networks, J. Am. Chem. Soc., 2021, 143, 5071–5079 Search PubMed
.
- J. Wang, Z. Li, Z. Zhou, Y. Ouyang, J. Zhang, X. Ma, H. Tian and I. Willner, DNAzyme- and light-induced dissipative and gated DNA networks, Chem. Sci., 2021, 12, 11204–11212 Search PubMed
.
- C. Wang, L. Yue and I. Willner, Controlling biocatalytic cascades with enzyme–DNA dynamic networks, Nat. Catal., 2020, 3, 941–950 Search PubMed
.
- E. Del Grosso, G. Ragazzon, L. J. Prins and F. Ricci, Fuel-Responsive allosteric DNA-Based aptamers for the transient release of ATP and cocaine, Angew. Chem., Int. Ed., 2019, 58, 5582–5586 CrossRef CAS PubMed
.
- Y. Ouyang, P. Zhang, H. Manis-Levy, Y. Paltiel and I. Willner, Transient dissipative optical properties of aggregated AU nanoparticles, CDSE/ZNS quantum dots, and supramolecular nucleic Acid-Stabilized AG nanoclusters, J. Am. Chem. Soc., 2021, 143, 17622–17632 Search PubMed
.
- F. Spitz and E. E. M. Furlong, Transcription factors: from enhancer binding to developmental control, Nat. Rev. Genet., 2012, 13, 613–626 CrossRef CAS PubMed
.
- H. M. Petrykowska, C. M. Vockley and L. Elnitski, Detection and characterization of silencers and enhancer-blockers in the greater CFTR locus, Genome Res., 2008, 18, 1238–1246 CrossRef CAS PubMed
.
- A. Travers, Transcriptional switches: the role of mass action, Phys. Life Rev., 2004, 1, 57–69 CrossRef
.
- M. Weitz, J. Kim, K. Kapsner, E. Winfree, E. Franco and F. C. Simmel, Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator, Nat. Chem., 2014, 6, 295–302 CrossRef CAS PubMed
.
- S. W. Schaffter and R. Schulman, Building in vitro transcriptional regulatory networks by successively integrating multiple functional circuit modules, Nat. Chem., 2019, 11, 829–838 CrossRef CAS PubMed
.
- M. Sun, J. Deng and A. Walther, Communication and Cross-Regulation between Chemically Fueled Sender and Receiver Reaction Networks, Angew. Chem., Int. Ed., 2022, 62, e202214499 CrossRef PubMed
.
- Z. Li, J. Wang and I. Willner, Alternate strategies to induce dynamically modulated transient transcription machineries, ACS Nano, 2023, 17, 18266–18279 CrossRef CAS PubMed
.
- D. E. Huizenga and J. W. Szostak, A DNA aptamer that binds adenosine and ATP, Biochemistry, 1995, 34, 656–665 CrossRef CAS PubMed
.
- Y. Ding, L. Gu, X. Wang, Z. Zhang, H. Zhang and J. Liu, Affinity-Guided coevolution of aptamers for guanine, xanthine, hypoxanthine, and adenine, ACS Chem. Biol., 2024, 19, 208–216 CrossRef CAS PubMed
.
- Y. Liu and J. Liu, Selection of DNA aptamers for sensing uric acid in simulated tears, Anal. Sens., 2022, 2, e202200010 CAS
.
- J. G. Bruno, M. P. Carrillo, T. Phillips and B. King, Development of DNA aptamers for cytochemical detection of acetylcholine, In Vitro Cell. Dev. Biol. Anim., 2008, 44, 63–72 CrossRef CAS PubMed
.
- B. C. Buddingh and J. C. M. Van Hest, Artificial Cells: Synthetic Compartments with Life-like Functionality and Adaptivity, Acc. Chem. Res., 2017, 50, 769–777 CrossRef CAS PubMed
.
- W. Jiang, Z. Wu, Z. Gao, M. Wan, M. Zhou, C. Mao and J. Shen, Artificial cells: past, present and future, ACS Nano, 2022, 16, 15705–15733 CrossRef CAS PubMed
.
- T. Trantidou, M. Friddin, Y. Elani, N. J. Brooks, R. V. Law, J. M. Seddon and O. Ces, Engineering compartmentalized biomimetic micro- and nanocontainers, ACS Nano, 2017, 11, 6549–6565 CrossRef CAS PubMed
.
- P. Zhang, L. Yue, M. Vázquez-González, Z. Zhou, W.-H. Chen, Y. S. Sohn, R. Nechushtai and I. Willner, MicroRNA-Guided Selective Release of Loads from Micro-/Nanocarriers Using Auxiliary Constitutional Dynamic Networks, ACS Nano, 2020, 14, 1482–1491 CrossRef CAS PubMed
.
- N. Que-Gewirth and B. Sullenger, Gene therapy progress and prospects: RNA aptamers, Gene Ther., 2007, 14, 283–291 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |