Open Access Article
Open Access ArticleFoldamer-based K+ channels with ion selectivity surpassing the KcsA channel
Ze
Lin
 ,
Zhaocheng
Xu
,
Jun
Tian
,
Zhaocheng
Xu
,
Jun
Tian
 ,
Chenyang
Zhang
,
Chuqiao
Sheng
and
Zeyuan
Dong
,
Chenyang
Zhang
,
Chuqiao
Sheng
and
Zeyuan
Dong
 *
*
State Key Laboratory of Supramolecular Structure and Materials, College of Chemistry, Jilin University, Changchun 130012, China. E-mail: zdong@jlu.edu.cn
First published on 14th February 2026
Abstract
This study presents biomimetic supramolecular K+ channels based on pore-containing foldamers with pyridine and oxadiazole backbones, aimed at high ion selectivity and transport efficiency. Modification of helical peripheral chains controlled their distribution in aqueous and lipid phases. LUV experiments demonstrated that a partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) of ∼5.0 optimally balanced this distribution, significantly enhancing ion transport activity. The M1 channel with isopropyl side chains exhibited an EC50 of 1.1 nM, the highest known activity for synthetic K+ channels, supporting the role of side chains in enhancing transport efficiency, as seen in natural K+ carriers. Bilayer lipid membrane (BLM) experiments revealed a potassium-to-sodium permeability ratio (PK/Na) of 55.2 for the M1 channel under 1 M asymmetric salt conditions, setting a new record for bioinspired K+ channels. Under 2 M asymmetric salt conditions, PK/Na increased to 138, surpassing the selectivity of the KcsA channel (80). BLM tests showed that M1 and M2 channels, with identical scaffolds, had similar selectivity and transport rates, indicating that the scaffold structure governs ion selectivity, while side chains primarily modulate transport activity. These findings offer key insights for bioinspired channel design and underscore the potential of biomimetic K+ channels in treating channelopathies.
P) of ∼5.0 optimally balanced this distribution, significantly enhancing ion transport activity. The M1 channel with isopropyl side chains exhibited an EC50 of 1.1 nM, the highest known activity for synthetic K+ channels, supporting the role of side chains in enhancing transport efficiency, as seen in natural K+ carriers. Bilayer lipid membrane (BLM) experiments revealed a potassium-to-sodium permeability ratio (PK/Na) of 55.2 for the M1 channel under 1 M asymmetric salt conditions, setting a new record for bioinspired K+ channels. Under 2 M asymmetric salt conditions, PK/Na increased to 138, surpassing the selectivity of the KcsA channel (80). BLM tests showed that M1 and M2 channels, with identical scaffolds, had similar selectivity and transport rates, indicating that the scaffold structure governs ion selectivity, while side chains primarily modulate transport activity. These findings offer key insights for bioinspired channel design and underscore the potential of biomimetic K+ channels in treating channelopathies.
Introduction
Natural ion channels are pore-forming proteins located on cell membranes, providing selective and rapid pathways for transmembrane ion conduction and regulating the concentration of ions.1,2 They play crucial roles in electrical signal transmission, nervous system regulation, and cell growth and apoptosis.3,4 Potassium ion channels, in particular, exhibit exceptional selectivity and transport activity.5 For example, the KcsA channel shows a high selectivity for K+, as reflected by its permeability ratio (PK/Na) of 80,6 owing to its selective filter that coordinates effectively with K+ but less efficiently with Na+.7,8 Dysfunction in these channels can lead to diseases such as cystic fibrosis, myotonia, and long QT syndrome.9 However, the complex structure and instability of these natural ion channels pose significant limitations in practical applications, especially in drug development and disease treatment. This has driven the search for artificial ion channels as viable alternatives.10–12To address these challenges, researchers have developed artificial systems for transmembrane ion transport, ranging from pore-forming ion channels to macroscopic ion-selective membranes.13–20 Among the various designs inspired by nature, ion channels have been the primary focus, offering potential replacements for natural channels. Inspired by natural K+ channels, various artificial K+ channels have been designed using materials such as crown ethers,21 biomimetic multiblock amphiphiles,22 polypeptides,23 macrocyclic compounds,24 and inorganic materials.25 Despite the advances in artificial K+ channels, improvements are still needed in terms of selectivity and transport activity before they can match the performance of their natural counterparts.
Within the last few years, our group has developed a series of aromatic foldamer-based molecules that self-assemble on bilayer membranes to form supramolecular ion channels. It has been proven that modulating the backbone structure of these molecules can significantly affect the properties of the channels. Adjusting the pore size, number of binding sites, and charge distribution within the pore, for instance, can fine-tune the ion selectivity of the channels, creating a series of artificial channels that are highly selective for potassium, sodium, and lithium ions.26–29
Although significant progress has been made in the internal structural design of foldamer-based ion channels, a critical gap remains in optimizing their external architecture, particularly in replicating the membrane-embedding characteristics of natural ion channels. This area remains largely unexplored. To address this gap, we turn to natural ion transport proteins for guidance, as their design has been optimized for stability and function within lipid membranes. These proteins rely on specific amino acid side chains that are precisely arranged to interact with the hydrophobic lipid bilayer, enhancing their membrane embedding and ensuring channel stability. For instance, membrane proteins enhance anchoring by matching amino acids (such as Val, Leu, Ala, Ile) with hydrophobic residues with the lipid bilayer.30 Similarly, the gramicidin A (gA) dimer, a prototypical linear peptide channel, demonstrates how judicious incorporation of isopropyl and isobutyl side chains enhances membrane integration while maintaining dynamic channel assembly.
Building on these insights, we hypothesize that rational side-chain engineering can regulate the phase distribution balance in synthetic channels, thereby enhancing their transport activity. Therefore, we adopted a helical biomimetic channel based on pyridine and oxadiazole moieties, which is capable of highly selective K+ transport. The backbone structure is preserved, while side chains, commonly found in natural ion channels, are introduced to modulate the channel properties. By adjusting these side chains, we aim to investigate how their distribution influences channel activity and selectivity.
To validate the ion selectivity and conduction properties of the engineered channel, we employed BLM technology. At a salt concentration of 1 M, the selectivity of the channel M1 was measured at 53, which increased to 138 at 2 M—significantly surpassing the natural KcsA channel (80) under the identical conditions. This exceptional selectivity makes our artificial channel the highest-performing K+/Na+ selective channel in biomimetic supramolecular K+ channels measured using the BLM method. Furthermore, its ion conductance was remarkable, with a rate of 3 × 107 K+ s−1 at 100 mV, demonstrating its high transport efficiency.
Notably, by drawing inspiration from the structures of natural channels and carriers, we selected the isopropyl side chain as the peripheral group of the artificial channel. This optimization achieved a distribution balance between the aqueous phase and the phospholipid membrane, significantly enhancing the activity of channel M1 to as high as 1.1 nM. This represents the highest activity reported for artificial K+ channels to date. Our findings underscore the crucial role of balancing amphiphilicity between water and lipid phases in the design of artificial ion channels. This not only provides valuable insights for side chain selection, but also paves the way for future applications in drug delivery and disease treatment.
Results and discussion
Design and synthesis of foldamers M1–M5
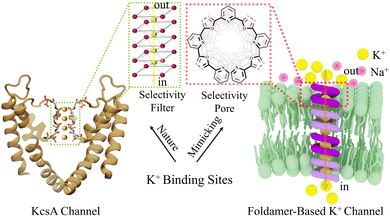
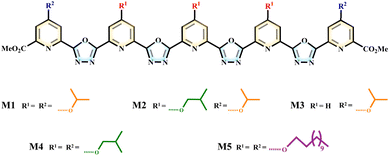
Natural K+ channels selectively transport K+ through specific ion recognition sites on the selectivity filter (SF). Inspired by this mechanism, we employed aromatic foldamers as scaffolds for artificial ion channels. These foldamers can self-assemble into pre-organized pore structures, which mimic the ion recognition sites of natural channels, enabling selective transmembrane transport of specific ions (Fig. 1).Learning from natural structures is an important direction for optimizing the properties of artificial systems. In this process, we found that short-chain side groups (such as isopropyl and isobutyl) are key structural features in natural K+ channels, playing a crucial role in their embedding in the membrane and maintaining stability. This idea led us to construct a series of foldamer-based channels with different side chains (Fig. 2 and S1). These foldamers (M1–M5), characterized by 1H-NMR, 13C-NMR, and MS (Fig. S2–S48), were synthesized using pyridine and oxadiazole as the skeletal framework. The side groups included are isopropyl, isobutyl, and dodecyl chains in varying combinations, designed as hydrophobic exterior anchors. Isopropyl and isobutyl represent typical short functional groups seen in natural K+ channels, while dodecyl is a longer alkyl chain that exhibits good lipophilicity.
Specifically, M1 features the shortest isopropyl side chain, M4 uses slightly longer isobutyl, M2 combines both isopropyl and isobutyl, and M3 has a partial isopropyl component. To explore the effect of highly lipophilic groups on channel activity, M5 includes dodecyl side chains. These side groups at the periphery of the helix confer amphiphilicity to the channel, enabling the helical molecules to diffuse in aqueous environments and assemble into transmembrane channels within phospholipid membranes. Meanwhile, the nitrogen and oxygen atoms in the helical skeleton form recognition sites within the channel pore, allowing the selective binding and recognition of K+.
It is worth noting that the molecular frameworks of M2 and M4 have been reported,31 but their transmembrane transport behaviours and systematic comparison with other side-chain-modified channels have not been investigated. Therefore, in this study, they are used as reference structures for in-depth exploration, providing important insights into the regulatory effects of different alkyl side chains on channel function.
To better understand the helical structure, we shortened the backbone and obtained the crystal structure of molecule M0 (Fig. S49). The results revealed that the conformation of M0 aligns with expectations: the N atom on the pyridine and the N atoms on the oxadiazole exhibit repulsion, causing the O atom on the oxadiazole to orient toward the pore interior. The overall structure adopts a folded conformation, with a pore size of approximately 3.6 Å. This simplified structure provides valuable insight into the molecular conformation, which serves as the foundation for designing the more complex M1–M5 foldamers.
Molecular self-assembly and ion response of foldamers M1 and M2
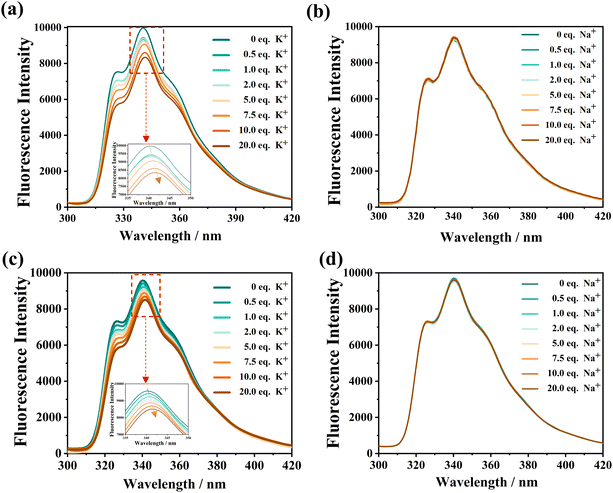
To further investigate the self-assembly behaviour in solution, concentration-dependent fluorescence titrations were performed. M1 and M2 exhibited fluorescence quenching at a concentration of 27 µM, suggesting that the foldamers undergo self-assembly at this concentration (Fig. S50). This observation strongly implies that the formation of supramolecular assemblies is governed by the skeletal structure of the foldamer, which is built from repeating pyridine and oxadiazole units.To determine the ion preference of the foldamers, fluorescence titration experiments were conducted on M1 and M2 with varying concentrations of K+ and Na+. The introduction of K+ caused a decrease in the fluorescence intensity of the M1 and M2 systems, accompanied by a slight red shift (Fig. 3).
The fluorescence intensity decrease and red shift became more pronounced as the K+ concentration increased. In contrast, the addition of Na+ had no noticeable effect on the fluorescence intensity of M1 and M2. These results suggest that M1 and M2 have a specific recognition and binding affinity for K+.
Liposome-based ion transmembrane transport
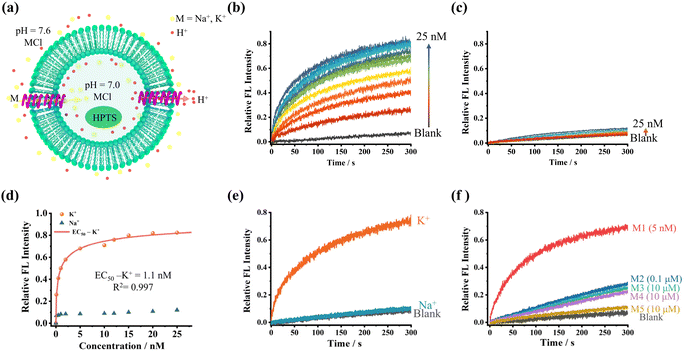
To investigate the transmembrane ion transport characteristics, a liposome-based kinetic test was conducted on channels M1 to M5. Suspensions of large unilamellar vesicles (LUVs, pH inside = 7.0, HEPES 10 mM) containing egg yolk L-α-phosphatidylcholine (EYPC) and encapsulated HPTS (1 mM) were prepared and added to HPTS buffer solution with pH = 7.8, creating a pH gradient inside and outside the vesicles. The translocation of cations into LUVs was assessed by continuously monitoring the change in fluorescence intensity of the embedded HPTS dye (Fig. 4a).Foldamers M1–M4 can all mediate transmembrane K+ transport. To identify the transport direction, we conduct three experiments (Fig. S51). The results indicate the H+/K+ antiport. The chloride ion indicator assay confirmed that the channels lack Cl− permeability (Fig. S51a and b), excluding anions from charge balance. Given that H+ efflux was observed in HPTS assays, the protonophore FCCP did not enhance transport efficiency (Fig. S51c and d), indicating that H+ efflux is not the rate-limiting step. Conversely, adding the potassium ionophore valinomycin significantly increased efficiency (Fig. S51e and f). These results demonstrate that the channels achieve H+/K+ antiport by mediating H+ efflux coupled with K+ influx.
M1–M5 transport K+ to varying degrees, whereas the transport of Na+ is very low, suggesting that these channels all have good K+ transport selectivity (Fig. 4b–d and S52–S55).
This selectivity difference is remarkably significant and surprising. Even at a concentration of 12 nM, the transport efficiency of M1 for K+ reaches 76%, while the transport of Na+ is almost negligible, showing no significant difference from the baseline level (Fig. 4e). To quantify the K+/Na+ ion selectivity (SK/Na) of the channel, we determined the selectivity of M1 to be 55.8 ± 5.2 by comparing the first-order rate constants of the potassium and sodium ion transport curves (Fig. S56–S59). This result represents a new milestone in the field of biomimetic supramolecular K+ channels, demonstrating significant selectivity. Notably, the selectivity of M2 to M4 is similar to that of M1, indicating that side-chain modifications do not significantly affect the channel selectivity (Fig. S60–S63).
After confirming the high selectivity of the foldamers for K+/Na+, the transport efficiency for other monovalent alkali metal ions was investigated (Fig. S64). The results showed the selectivity order: Rb+ > Cs+ > K+ > Na+ > Li+. This order is primarily controlled by the matching of the ion size with the foldamer cavity and the dehydration energy barrier: the larger Rb+ and Cs+ ions match well with the foldamer cavity (∼3.6 Å) and have lower dehydration energies, resulting in high permeability; while the smaller Li+ and Na+ ions are effectively filtered due to size mismatch and higher dehydration energy barriers. This characteristic is consistent with the behaviour of some natural potassium channels, which are permeable to K+ and its larger congeners.
Given the strict monovalent cation selectivity of M1, we further investigated its ability to exclude divalent ions, a key feature of natural K+ channels. Transmembrane transport tests of Ca2+ and Mg2+ were also performed on these channels. These channels have almost no transport function for Ca2+ or Mg2+, suggesting that they are specific K+ channels and have almost no transport function for other physiologically important cations in the human body (Fig. S65). This behaviour aligns with natural K+ channels, which have evolved to distinguish between K+, Ca2+, and Mg2+.
Building on its exceptional selectivity, M1 maintained efficient K+-selective transport in artificial membrane systems. This functional preservation prompted investigation of its biological compatibility, where co-localization assays demonstrated spontaneous aggregation on both HUVEC and leukocyte membranes (Fig. S66). High-resolution imaging further confirmed stable integration into native cellular membrane architectures while retaining robust binding affinity. These findings provide a foundation for further exploration in the field of biomedicine, including future functional validation in live cells through patch-clamp electrophysiology or K+-flux assays, to support the development of potential therapeutic applications. Moreover, although channels M1–M5 all exhibit good K+ selectivity, the activity of transporting K+ varies considerably. Channel activities decrease progressively from M1 to M5. M1 exhibits high K+ transport efficiency, achieving 70% at 5 nM, far outperforming M2 at 0.1 µM (28%) and M3 to M5 at 10 µM (25%, 20%, and 10%, respectively) (Fig. 4f). It is worth noting that the K+ transport efficiency of M1 is already near saturation at a concentration of 10 nM, whereas M2 requires a concentration of 30 µM to approach saturation, and M3 needs nearly 100 µM. To further quantify these differences, the EC50 (the effective concentration required to reach 50% activity) for each channel was determined using the Hill equation, Y = 1/(1 + (EC50/[C])n).32Table 1 summarizes the EC50 values for K+ transport, with M4 and M5 failing to reach 50% efficiency and thus no EC50 values are provided.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) of foldamers M1–M5
P) of foldamers M1–M5
| Foldamer | EC50 (nM) | Mol% relative to lipid | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pa Pa |
|---|---|---|---|
| a The lipid-water partition coefficients were obtained by computer simulation with the ALOGPS 2.1 program. b N.D.: not detectable due to the inability of M4 and M5 to transport K+ to 50% efficiency. [phospholipid] = 0.09 mM. | |||
| M1 | 1.1 | 1.2 × 10−3 | 5.0 |
| M2 | 3100 | 3.3 | 5.9 |
| M3 | 110![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
116.7 | 4.3 |
| M4 | N.Db | N.Db | 6.2 |
| M5 | N.Db | N.Db | 9.1 |
The EC50 values of M1, M2, and M3 increase gradually and vary greatly, implying that their transmembrane K+ transporting activities vary greatly (Table 1). The EC50 of M1, which has all isopropyl side chains, is 1.1 nM, 4.5 times more active than that of gA under the same conditions (5.0 nM),26 and is the most active K+ channel in the field of artificial K+ channels so far. This also suggests that M1 is among the most active and selective artificial K+ channels identified so far. The EC50 values for M2, which contains just isobutyl side chains, are 3.1 µM, while M3, which contains partial isopropyl side chains, has an EC50 of 110 µM. These values differ by 2800 and 105 times from the EC50 values for M1, respectively. The activity difference is significant, indicating that not only the type of channel side chain should be considered, but also the number of side chains is important. With an all-isobutyl side chain, M4 exhibits poor channel activity, whereas M5, with an all-dodecyl side chain, has nearly no K+ transfer capacity.
Amphiphilicity of channels
To understand the reasons behind these large variations in channel activity, we examined the entire mechanism of ion transport across the membrane. The process begins with the foldamer dissolving and diffusing in water, followed by self-assembly and embedding into the lipid membrane (Fig. S67). The efficiency of these steps depends on the balance between the channel distribution in the aqueous and phospholipid phases, which in turn determines activity (EC50 from liposome-based testing). The final stage involves ion transport through the channel aperture, which dictates ion selectivity and transport rate. The side chains of M1 to M5 are different, resulting in distinct dissolution and diffusion behaviours, which subsequently affect the membrane insertion process. In particular, the isopropyl side chain exhibits the best properties, as it easily dissolves and diffuses in water while remaining relatively stable in lipid membranes. In contrast, the solubility of the isobutyl and dodecyl side chains in water gradually decreases, leading to poor ion transport activity in M4 and almost no ion transport activity in M5.To further understand the impact of these side chains on channel activity, we examined the lipid-water partition coefficients (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) of the five foldamers to investigate their distribution in water and phospholipid membranes. Log
P) of the five foldamers to investigate their distribution in water and phospholipid membranes. Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P serves as a key indicator of lipophilicity, which plays a crucial role in its dissolution and permeability, which in turn determines its ability to insert into the membrane and ion channel activity.33 Typically, this value falls within a reasonable range to ensure effective membrane interaction. To assess this, the ALOGPS 2.1 program was used to predict the lipid-water distribution coefficients of M1 to M5 (Table 1).34 The log
P serves as a key indicator of lipophilicity, which plays a crucial role in its dissolution and permeability, which in turn determines its ability to insert into the membrane and ion channel activity.33 Typically, this value falls within a reasonable range to ensure effective membrane interaction. To assess this, the ALOGPS 2.1 program was used to predict the lipid-water distribution coefficients of M1 to M5 (Table 1).34 The log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values of these five molecules vary significantly, reflecting how the length and number of side chains greatly affect the hydrophilicity of foldamer-based channels. For instance, M3, with the shortest and fewest side chains, has a log
P values of these five molecules vary significantly, reflecting how the length and number of side chains greatly affect the hydrophilicity of foldamer-based channels. For instance, M3, with the shortest and fewest side chains, has a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of 4.3, while M5, with a dodecyl side chain, has a log
P of 4.3, while M5, with a dodecyl side chain, has a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of 9.1. However, they both exhibit poor channel transport activity. The low activity of M3 may be due to the strong hydrophilicity of its side chain, which restrains it from embedding into the membrane. On the other hand, the low activity of M5 can be attributed to the lipophilic nature of the side chain, which hinders solubilization and diffusion in water, limiting the ability to interact with the membrane.
P of 9.1. However, they both exhibit poor channel transport activity. The low activity of M3 may be due to the strong hydrophilicity of its side chain, which restrains it from embedding into the membrane. On the other hand, the low activity of M5 can be attributed to the lipophilic nature of the side chain, which hinders solubilization and diffusion in water, limiting the ability to interact with the membrane.
The log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values of M1 (5.0), M2 (5.9), and M4 (6.2) fall within a favourable range, indicating moderate hydrophilicity. This clear correlation between log
P values of M1 (5.0), M2 (5.9), and M4 (6.2) fall within a favourable range, indicating moderate hydrophilicity. This clear correlation between log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and ion transport activity reveals the primary role of the side chains. Their dominant function is to modulate the amphiphilicity of the foldamers, which in turn governs the critical step of membrane insertion efficiency. An optimal log
P and ion transport activity reveals the primary role of the side chains. Their dominant function is to modulate the amphiphilicity of the foldamers, which in turn governs the critical step of membrane insertion efficiency. An optimal log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ensures a balance between sufficient aqueous solubility for diffusion to the membrane and adequate lipophilicity for partitioning into and stabilizing within the lipid bilayer. This optimal balance maximizes the number of foldamer molecules that can successfully embed into the membrane and assemble into functional channels, thereby determining the macroscopic transport activity. Interestingly, the log
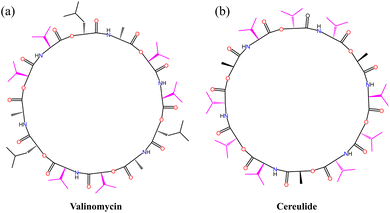
P ensures a balance between sufficient aqueous solubility for diffusion to the membrane and adequate lipophilicity for partitioning into and stabilizing within the lipid bilayer. This optimal balance maximizes the number of foldamer molecules that can successfully embed into the membrane and assemble into functional channels, thereby determining the macroscopic transport activity. Interestingly, the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of M1 aligns closely with that of the K+ carriers cereulide (5.1) and valinomycin (5.0), both predicted using the same method, and it demonstrates high transmembrane K+ transport activity. This suggests that the amphiphilicity of the side chains is crucial in designing foldamer-based biomimetic ion channels, underlining that a log
P of M1 aligns closely with that of the K+ carriers cereulide (5.1) and valinomycin (5.0), both predicted using the same method, and it demonstrates high transmembrane K+ transport activity. This suggests that the amphiphilicity of the side chains is crucial in designing foldamer-based biomimetic ion channels, underlining that a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value around 5.0 is particularly advantageous for efficient function. After screening, the isopropyl side chain was found to be particularly effective in enhancing K+ transport. Interestingly, the design of K+ channel M1 aligns closely with evolutionary strategies observed in nature. The use of isopropyl groups in M1 mirrors the side chain composition of natural K+ ion carriers like valinomycin and cereulide, which also feature isopropyl side chains (Fig. 5).35 This similarity suggests that the design of M1 unintentionally reflects an evolutionary optimization developed over time. The exceptional transport activity highlights the critical role of the isopropyl group in facilitating efficient K+ transport. This finding not only demonstrates a surprising alignment between artificial design and natural evolution but also provides valuable insight for the future development of synthetic ion channels. The successful incorporation of isopropyl side chains serves as a key reference for creating more efficient ion transport systems.
P value around 5.0 is particularly advantageous for efficient function. After screening, the isopropyl side chain was found to be particularly effective in enhancing K+ transport. Interestingly, the design of K+ channel M1 aligns closely with evolutionary strategies observed in nature. The use of isopropyl groups in M1 mirrors the side chain composition of natural K+ ion carriers like valinomycin and cereulide, which also feature isopropyl side chains (Fig. 5).35 This similarity suggests that the design of M1 unintentionally reflects an evolutionary optimization developed over time. The exceptional transport activity highlights the critical role of the isopropyl group in facilitating efficient K+ transport. This finding not only demonstrates a surprising alignment between artificial design and natural evolution but also provides valuable insight for the future development of synthetic ion channels. The successful incorporation of isopropyl side chains serves as a key reference for creating more efficient ion transport systems.
Electrophysiological properties of the channels
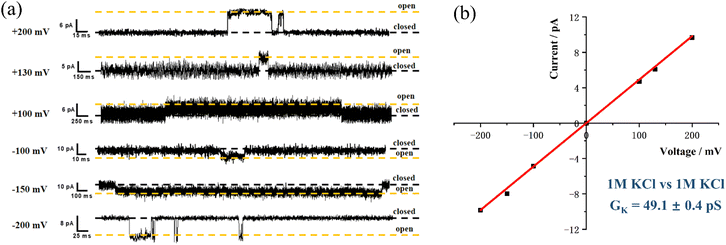
M2 is a high-speed K+ channel with an electrical conductance (G) of 48.3 pS, which corresponds to a transport rate of 3 × 107 K+ s−1 at 100 mV.31 Its conductance is notably higher than that of gA (G = 18 pS) under identical conditions,36 highlighting its efficiency in K+ transport.To investigate the effect of different side chains on the K+ conductivity of channels, the BLM experiment was carried out on M1 in the 1 M KCl environment. M1 displayed a regular square signal (Fig. 6), with a fitted K+ conductance of 49.1 ± 0.4 pS, similar to M2. This suggests that the structural framework of the channel, including interior binding sites, is crucial in determining the transport properties of the channel. This similarity in conductance aligns with the principle that natural channel conductance is primarily influenced by the channel pore size and the electronegativity of the vestibular opening.7,37 Since only the side chains were altered in this system while the aperture and electronegativity remained unchanged, it is hypothesized that these structural elements dominate the channel conductance.
The nearly identical single-channel conductance of M1 and M2 suggests that they form transmembrane pores with highly similar architectures, prompting an investigation into their assembly stoichiometry. We propose a tetrameric model, supported by the columnar supramolecular packing observed in the crystal structure of M0 (interplanar distance approximately 4.3 Å). In the functional molecules M1–M5, the hydrophobic side chains likely optimize this packing within the membrane, reducing the helical pitch to approximately 3.5–3.8 Å. A pore composed of four such molecules, each contributing about two helical turns, yields a total length of 2.8–3.0 nm. This dimension aligns well with the hydrophobic thickness of a lipid bilayer (approximately 2.6–2.8 nm), thereby supporting the observed stable, unitary currents. Alternative oligomeric states (e.g., trimer or pentamer) are less compatible with these physical constraints.
BLM recordings of M1 and M2 reveal clear open–close transitions, suggesting that gating behaviour may arise from the dynamic nature of their supramolecular assemblies. Channel closure could result from transient sliding or partial dissociation of the π–π stacked helical foldamers under the influence of transmembrane voltage. Additionally, the side chains might contribute to gating by sterically blocking the channel or destabilizing the overall assembly structure.
Further analysis under symmetrical salt concentration conditions in the BLM experiment revealed identical K+ transport speeds for both M1 and M2, suggesting that the ion transport rates through their pores are similar. However, during the LUVs test, M1 exhibited a K+ transport activity 2800 times higher than M2. This discrepancy is mainly attributed to variations in the membrane embedding process rather than differences in pore transport properties. Specifically, the amphiphilic effects of the side chains on the external surface of the pores likely enhance the transport activity of M1.
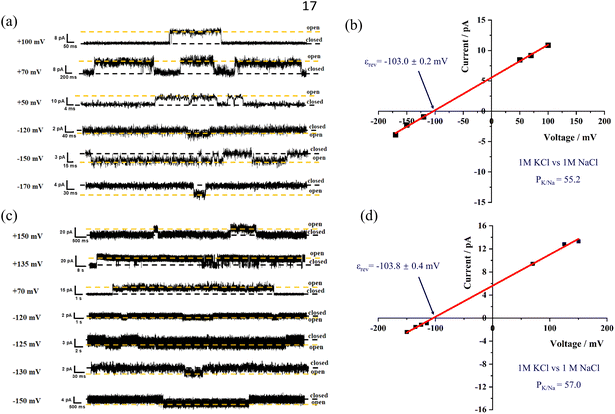
To accurately determine the K+/Na+ ion selectivity of the channel formed by this framework, an asymmetric BLM method was employed for measurement. M1 demonstrates excellent single-channel selectivity for K+ over Na+ under asymmetric conditions (Fig. 7a and b). The reversal potential was measured using the current–voltage (I–V) curve fitting method and was found to be −103.0 mV, indicating a clear preference for K+ over Na+. Based on this reversal potential and the Goldman-Hodgkin-Katz equation, the potassium-to-sodium permeation ratio (PK/Na) for M1 was calculated to be 55.2, the highest single-channel selectivity for K+ over Na+ reported among biomimetic K+ channels characterized by BLM.17,24,26 Given the high accuracy of this method and the fact that most recent reports on artificial K+ channels show selectivities below 50, the selectivity of this framework represents a significant improvement. This suggests that well-designed helical skeleton cavities can act as highly selective ion filters, allowing specific ions to pass through the similar to natural channels.
Under the same testing conditions, the potassium-sodium ion permeability selectivity of channel M2 was evaluated (Fig. 7c and d). The fitted reversal potential for M2 was −103.8 mV, which closely matches that of M1 under the same conditions. Additionally, the calculated PK/Na value for M2 was 57.0, which is comparable to that of M1. This result is in agreement with the potassium-sodium ion selectivity observed in LUV tests, and both M1 and M2 exhibited similar ion selectivity in both LUV and BLM asymmetric tests. These findings further highlight that for foldamer-based supramolecular channels, the scaffold structure of the channel determines its ion selectivity, rather than the external side chains.
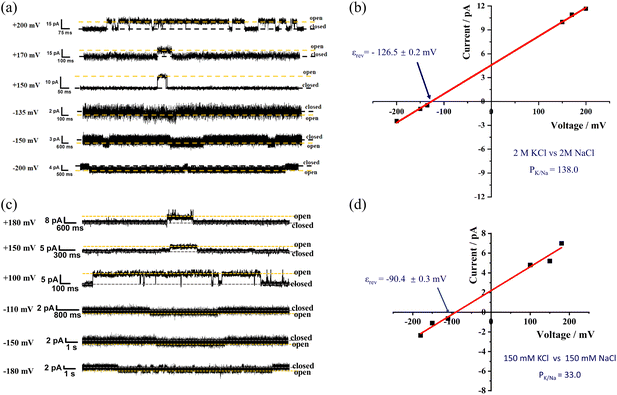
Because the ion selectivity of the channel can be more reliably determined under elevated salt concentrations, the salt concentration on both sides of the channel was increased to 2M to determine the PK/Na of the channel M1. Under the identical experimental system previously used to characterize the KcsA channel, at identical conditions that include the identical testing methods, salt solutions, and concentrations, the PK/Na value of M1 increased to 138.0, surpassing the selectivity of the natural KcsA channel (80) (Fig. 8a and b).6 This shows that the PK/Na of the channel can be enhanced by increasing the concentration of ions on both sides of the membrane. The enhanced selectivity under high salt conditions likely results from increased electrostatic screening. At lower ionic strength, fixed negative charges near the channel entrance generate a non-specific electric field that attracts both K+ and Na+, thereby obscuring the intrinsic selectivity of the pore. As ionic strength rises, these charges become effectively screened, weakening the external field. Consequently, the inherent selectivity of the pore, which is determined by its geometry and chemical coordination, becomes the dominant factor. This phenomenon is consistent with the higher selectivity exhibited by natural channels at higher ion concentrations, further proving the superiority of this channel framework in terms of ion selectivity.
The exceptional K+/Na+ selectivity of artificial channel M1—surpassing synthetic counterparts at 1 M and even exceeding natural KcsA channels at 2 M—prompted us to investigate its performance under physiologically relevant conditions. Since the fundamental physiological role of K+ channels is to generate and stabilize the resting membrane potential through high K+/Na+ selectivity, we assessed this key function by measuring the reversal potential under a standard physiological gradient (150 mM KCl inside/150 mM NaCl outside). The measured reversal potential was −90.4 mV (Fig. 8c and d), confirming the high selectivity of our artificial channel. This value closely approximates the theoretical K+ equilibrium potential (Ek = −90 mV), which is more negative than the resting membrane potential of natural cells (approximately −70 mV).2 This suggests that channel M1 can significantly influence transmembrane electrical signalling.
The PK/Na for M1 was calculated based on the reversal potential using the Goldman-Hodgkin-Katz equation, yielding a value of 33.0. This high PK/Na ratio underscores the exceptional K+ selectivity of M1 compared to Na+, further supporting that M1 maintains high K+/Na+ selectivity under physiological conditions. These findings suggest that the high selectivity of M1 could have valuable applications in biomedicine and bioelectronics, particularly in fields requiring precise ion transport regulation.
Conclusions
In conclusion, we have constructed biomimetic K+ channels by utilizing pyridine and oxadiazole as the structural backbones of foldamers. Inspired by the natural K+ transmembrane transport systems, we introduced various side chains into the foldamers to explore their effects on channel properties. The heteroatoms within the channel pore specifically interact with K+ ions, and LUV experiments demonstrate a potassium-to-sodium selectivity ratio of 55.8 for these channels.Our findings reveal that the channel activity is strongly influenced by the lipid-water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), which is modulated by the side chains. When the log
P), which is modulated by the side chains. When the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P is approximately 5.0, the channel achieves an optimal distribution between the aqueous and phospholipid phases, resulting in efficient transmembrane K+ transport. Notably, the inclusion of isopropyl side chains significantly enhances channel activity, reflecting the evolutionary characteristics of natural K+ carriers. Channels with isopropyl side chains exhibit exceptionally high activity (EC50 = 1.1 nM), making them the most active artificial K+ channels reported to date.
P is approximately 5.0, the channel achieves an optimal distribution between the aqueous and phospholipid phases, resulting in efficient transmembrane K+ transport. Notably, the inclusion of isopropyl side chains significantly enhances channel activity, reflecting the evolutionary characteristics of natural K+ carriers. Channels with isopropyl side chains exhibit exceptionally high activity (EC50 = 1.1 nM), making them the most active artificial K+ channels reported to date.
Additionally, BLM testing reveals that the channel's backbone structure governs its ion transport rate and selectivity. The channel with pyridine and oxadiazole sequences exhibits a K+ transport rate as high as 3 × 107 K+ s−1 at 100 mV. Under 1M conditions, the potassium-to-sodium permeability ratio (PK/Na) is 55.2, and when the ion concentration is increased to 2M, the PK/Na value rises to 138, setting a record for the highest selectivity biomimetic K+ channel achieved by this method. Moreover, this artificial channel maintains high selectivity under physiological concentration conditions, with a reversal potential of −90.4 mV. Thus, this study not only provides a highly active and selective K+ channel, offering potential strategies for the treatment of channel-related diseases, but also provides important theoretical guidance for the design of high-performance biomimetic channels.
Author contributions
Z. L. performed data curation, formal analysis, and drafted the original manuscript. Z. X. was responsible for cultivating the single crystals. J. T. and C. S. conducted data analysis and contributed to the manuscript draft. C. Z. and Z. D. provided supervision.Conflicts of interest
There are no conflicts to declare.Data availability
CCDC 2467629 contains the supplementary crystallographic data for this paper.38The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: synthetic details, crystal structure figures, and experimental details. See DOI: https://doi.org/10.1039/d5sc08786d.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 22471093).Notes and references
- E. Gouaux and R. MacKinnon, Science, 2005, 310, 1461–1465 CrossRef CAS PubMed.
- B. Hille, Ion channels of excitable membranes, Sinauer Associates, 2001 Search PubMed.
- L. A. Pardo, Physiology, 2004, 19, 285–292 CrossRef CAS PubMed.
- C. D. Bortner and J. A. Cidlowski, Philos. Trans. R. Soc., B, 2014, 369, 20130104 CrossRef PubMed.
- J. T. Littleton and B. Ganetzky, Neuron, 2000, 26, 35–43 CrossRef CAS PubMed.
- K. Mita, T. Sumikama, M. Iwamoto, Y. Matsuki, K. Shigemi and S. Oiki, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2017168118 CrossRef CAS PubMed.
- D. A. Doyle, J. M. Cabral, R. A. Pfuetzner, A. Kuo, J. M. Gulbis, S. L. Cohen, B. T. Chait and R. MacKinnon, Science, 1998, 280, 69–77 CrossRef CAS PubMed.
- Y. F. Zhou, J. H. Morais-Cabral, A. Kaufman and R. MacKinnon, Nature, 2001, 414, 43–48 CrossRef CAS PubMed.
- M. A. Zaydman, J. R. Silva and J. Cui, Chem. Rev., 2012, 112, 6319–6333 CrossRef CAS PubMed.
- S. Sun, Z. Xu, Z. Lin, W. Chen, Y. Zhang, M. Yan, S. Ren, Q. Liu, H. Zhu, B. Tian, J. Zhang, W. Zhang, S. Jiang, C. Sheng, J. Ge, F. Chen and Z. Dong, Acta Biomater., 2024, 181, 391–401 Search PubMed.
- J. A. Malla, R. M. Umesh, A. Vijay, A. Mukherjee, M. Lahiri and P. Talukdar, Chem. Sci., 2020, 11, 2420–2428 RSC.
- W. Huang, C. Jia and C. Ren, Chemmedchem, 2025, 20, e202400811 Search PubMed.
- S. H. Yu and L. B. Mao, ACS Cent. Sci., 2021, 7, 1467–1469 Search PubMed.
- M. Zhang, P. P. Zhu, P. Y. Xin, W. Si, Z. T. Li and J. L. Hou, Angew. Chem., Int. Ed., 2017, 56, 2999–3003 Search PubMed.
- J. A. Malla, R. M. Umesh, S. Yousf, S. Mane, S. Sharma, M. Lahiri and P. Talukdar, Angew. Chem., Int. Ed., 2020, 59, 7944–7952 CrossRef CAS PubMed.
- K. Sato, R. Sasaki, R. Matsuda, M. Nakagawa, T. Ekimoto, T. Yamane, M. Ikeguchi, K. V. Tabata, H. Noji and K. Kinbara, J. Am. Chem. Soc., 2022, 144, 11802–11809 Search PubMed.
- L. Jin, C. Sun, Z. Y. Li, J. Shen and H. Q. Zeng, Chem. Commun., 2023, 59, 3610–3613 RSC.
- T. F. Yan, S. D. Liu, C. Li, J. Y. Xu, S. J. Yu, T. T. Wang, H. C. Sun and J. Q. Liu, Angew. Chem., Int. Ed., 2022, 61, e202210214 Search PubMed.
- P. Xin, L. Xu, W. Dong, L. Mao, J. Guo, J. Bi, S. Zhang, Y. Pei and C. P. Chen, Angew. Chem., Int. Ed., 2023, 62, e202217859 CrossRef CAS PubMed.
- A. D. Peters, S. Borsley, F. della Sala, D. F. Cairns-Gibson, M. Leonidou, J. Clayden, G. F. S. Whitehead, I. J. Vitórica-Yrezábal, E. Takano, J. Burthem, S. L. Cockroft and S. J. Webb, Chem. Sci., 2020, 11, 7023–7030 RSC.
- M. Barboiu, Acc. Chem. Res., 2018, 51, 2711–2718 CrossRef CAS PubMed.
- K. Sato, T. Muraoka and K. Kinbara, Acc. Chem. Res., 2021, 54, 3700–3709 CrossRef CAS PubMed.
- G. Su, M. Zhang and W. Si, Angew. Chem., Int. Ed., 2016, 55, 14678–14682 CrossRef CAS PubMed.
- P. Xin, H. Kong, Y. Sun, L. Zhao, H. Fang, H. Zhu, T. Jiang, J. Guo, Q. Zhang, W. Dong and C. P. Chen, Angew. Chem., Int. Ed., 2019, 58, 2779–2784 CrossRef CAS PubMed.
- J. P. Li, L. H. Du, X. Kong, J. Z. Wu, D. N. Lu, L. Jiang and W. Guo, Natl. Sci. Rev., 2023, 10, nwad260 Search PubMed.
- S. W. Qi, C. Y. Zhang, H. Yu, J. Zhang, T. F. Yan, Z. Lin, B. Yang and Z. Y. Dong, J. Am. Chem. Soc., 2021, 143, 3284–3288 Search PubMed.
- L. Zhang, C. Zhang, X. Dong and Z. Dong, Angew. Chem., Int. Ed., 2023, 62, e202214194 Search PubMed.
- L. Zhang, J. Tian, Z. Lin and Z. Dong, J. Am. Chem. Soc., 2024, 146, 8500–8507 Search PubMed.
- J. Tian, L. Zhang, Z. Lin, S. Mao and Z. Dong, Chem. Sci., 2025, 16, 15111–15120 Search PubMed.
- G. Von Heijne, Nat. Rev. Mol. Cell Biol., 2006, 7, 909–918 Search PubMed.
- C. Lang, X. Deng, F. Yang, B. Yang, W. Wang, S. Qi, X. Zhang, C. Zhang, Z. Dong and J. Liu, Angew. Chem., Int. Ed., 2017, 56, 12668–12671 CrossRef CAS PubMed.
- A. Gilles and M. Barboiu, J. Am. Chem. Soc., 2016, 138, 426–432 Search PubMed.
- C. A. Lipinski, F. Lombardo, B. W. Dominy and P. J. Feeney, Adv. Drug Deliv. Rev., 2012, 64, 4–17 CrossRef.
- I. V. Tetko, J. Gasteiger, R. Todeschini, A. Mauri, D. Livingstone, P. Ertl, V. Palyulin, E. Radchenko, N. S. Zefirov, A. S. Makarenko, V. Y. Tanchuk and V. V. Prokopenko, J. Comput. Aided Mol. Des., 2005, 19, 453–463 CrossRef CAS PubMed.
- A. Makarasen, K. Yoza and M. Isobe, Chem.–Asian J., 2009, 4, 688–698 CrossRef CAS PubMed.
- N. Thompson, G. Thompson, C. D. Cole, M. Cotten, T. A. Cross and D. D. Busath, Biophys. J., 2001, 81, 1245–1254 CrossRef CAS PubMed.
- D. Naranjo, H. Moldenhauer, M. Pincuntureo and I. Diaz-Franulic, J. Gen. Physiol., 2016, 148, 277–291 Search PubMed.
- CCDC 2467629: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2ntry6.
| This journal is © The Royal Society of Chemistry 2026 |