Open Access Article
Open Access ArticleAcidic in vitro selection of metal-specific deoxyribozymes
Pan
Jia
a,
Yangyang
Chang
*ab,
Shen
Li
b,
Wei
Xue
a,
Shusen
Xiao
a,
Qiang
Zhang
c,
Jiuxing
Li
ab,
Ying
Wang
 d,
Zijie
Zhang
*ab and
Meng
Liu
d,
Zijie
Zhang
*ab and
Meng
Liu
 *ab
*ab
aSchool of Environmental Science and Technology, Key Laboratory of Industrial Ecology and Environmental Engineering (Ministry of Education), Dalian University of Technology, Dalian POCT Laboratory, Dalian, 116024, China. E-mail: yychang@dlut.edu.cn; zhangzijie@dlut.edu.cn; mliu@dlut.edu.cn
bCentral Hospital of Dalian University of Technology, Dalian 116033, China
cSchool of Bioengineering, Key Laboratory of Bio-Intelligent Manufacturing (Ministry of Education), Dalian University of Technology, Dalian, 116024, China
dState Key Laboratory of Pollution Control and Resources Reuse, College of Environmental Science and Engineering, Tongji University, Shanghai, 200092, China
First published on 20th January 2026
Abstract
Deoxyribozymes (DNAzymes) are in vitro selected catalytic DNA molecules that recruit metal ions to function. However, nearly all previous DNAzymes generated through conventional selection methods exhibit poor metal selectivity. Here, we report an acidic in vitro selection strategy for isolating truly metal-specific DNAzymes. By using Ca2+ as the target in positive selection and a mixture of competing metal ions in counter-selection, and conducting the selections under acidic conditions to suppress metal hydrolysis, we have successfully selected an acidic RNA-cleaving DNAzyme, termed aRCD-Ca2, which is only activated by Ca2+ and shows no response to all other tested metal ions, including monovalent ions and chemically similar competing divalent ions (Mg2+, Cu2+, Zn2+, Co2+, Ni2+, Mn2+ and Pb2+). This represents the first acidic DNAzyme with exclusive metal selectivity. Moreover, aRCD-Ca2 exhibits fast catalytic activity, with a kobs of 0.026 min−1 toward Ca2+. A trans-acting aRCD-Ca2TCQ was also engineered from aRCD-Ca2 that enabled highly specific and sensitive monitoring of Ca2+ dynamics in HT22 cell lysosomes through a fluorescent probe. We envision that the described acidic in vitro selection strategy can be readily adapted to obtain more new DNAzymes with high specificity for other metal ions and advance the development of nucleic acid catalysts for a wide range of applications.
Introduction
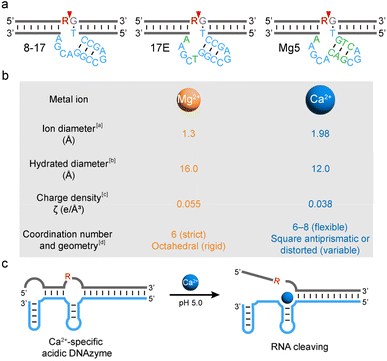
Deoxyribozymes (or DNAzymes) are single-stranded DNA molecules with catalytic activity.1,2 In contrast to naturally evolved protein enzymes and ribozymes, DNAzymes are isolated from large pools of random DNA sequences via in vitro selection, a process also referred to as systematic evolution of ligands by exponential enrichment (SELEX).3–6 Since their first report in 1994, numerous DNAzymes that can catalyze various chemical reactions have been identified by in vitro selection.7–10 One of the most extensively studied classes of DNAzymes is RNA-cleaving DNAzymes (RCDs), which catalyze RNA cleavages in a site-specific manner.11,12Because metal ions are commonly recruited as cofactors for structural folding and catalysis, RCDs have been widely applied to sensing or imaging of metal ions.13–15 However, nearly all RCDs obtained through conventional in vitro selection suffer from poor metal specificity, particularly difficult in distinguishing chemically similar competing ions, such as Ca2+ and Mg2+.16 A notable example is the 8–17 DNAzyme, which was first isolated in 1997 under simulated physiological conditions (i.e., 2 mM MgCl2/150 mM KCl, pH 7.5, 37 °C).17 Although derived with Mg2+, 8–17 exhibits activity with many other divalent ions, including Pb2+, Zn2+, Ca2+, Mn2+, and Co2+, among others.18,19 Recent structural studies have shown that these metals induce a similar tertiary fold of 8–17, demonstrating its non-specificity in metal binding.20 Interestingly, the 8–17 motif has been repeatedly rediscovered under different selection conditions, including variants such as 17E (isolated with Zn2+), Mg5 (isolated with histidine and Mg2+) and others.21–24 These variants share the catalytic core of 8–17 (Scheme 1a), demonstrating the difficulty in obtaining truly metal-specific DNAzymes via traditional in vitro selection. In fact, many other RCDs (e.g., 10–23) suffer from poor metal specificity.25,26
An overlooked factor in conventional in vitro selection is metal hydrolysis. In aqueous environments, metal-bound water molecules deprotonate and produce metal-bound hydroxyl groups.16 As a result, DNA is not fully exposed to free metal ions during selection under neutral conditions. Lowering pH can suppress metal hydrolysis, thus facilitating the interactions between DNA and free metal ions, potentially enhancing the isolation of specific DNAzymes for a given metal ion.
Recently, a few new DNAzymes have been selected under unconventional conditions to reveal their hidden abilities.27–31 We hypothesize that, rather than conventional selections near neutral pH, selecting DNAzymes under acidic conditions could yield highly metal-specific RCDs by suppressing metal hydrolysis in a low-pH environment. In addition, acidic in vitro selections offer several inherent advantages. First, RNA substrates exhibit higher chemical stability under acidic conditions. For instance, the rate constant for RNA degradation at pH 6 is approximately 10-fold lower than under otherwise similar conditions at pH 7.4, which could improve the chemical stability of RNA-cleaving DNAzymes.32 Second, acidic selection may avoid the recurrence of existing metal-nonspecific catalytic motifs such as 8–17, since DNA folding is strongly influenced by base protonation.33,34
To test this idea, we recruited the chemically similar Ca2+ and Mg2+ as the first case. Ca2+ and Mg2+ were chosen for three reasons: (1) they share similar properties. Both are Group 2 alkaline earth metal ions with a +2 charge and a preference for oxygen-containing ligands, making them difficult to distinguish, and many DNAzymes fail to differentiate between them;18,24 (2) Ca2+ and Mg2+ differ in ionic radius, charge density, and coordination dynamics (Scheme 1b).16,35–38 Ca2+ prefers flexible and fast-exchanging coordination (with 6–8 ligands), while Mg2+ exhibits rigid (mostly octahedral), slow-exchanging coordination and has a higher dehydration enthalpy due to its higher charge density;16,36,37 (3) a Ca2+-specific DNAzyme is valuable for monitoring Ca2+ dynamics in the acidic microenvironment of lysosomes in live cells.39 Currently, monitoring of lysosomal components remains challenging due to the small size, acidic pH, and compartmentalization of lysosomes.
In this work, we conducted the first attempt to use acidic in vitro selection to isolate metal-specific DNAzymes. We have successfully isolated a Ca2+-specific acidic RNA-cleaving DNAzyme, named aRCD-Ca2 (Scheme 1c), which exhibited effective catalytic activity with an observed rate constant (kobs) of 0.026 min−1 at pH 5.0 and demonstrated excellent selectivity over other competing metal ions (Mg2+, Cu2+, Zn2+, Co2+, Ni2+, Mn2+ and Pb2+). We further developed a trans-acting aRCD-Ca2T-based fluorescent probe for imaging of Ca2+ dynamics within lysosomes in HT22 hippocampal neuron cells with high sensitivity and stability, validating its practical intracellular sensing applications.
Results and discussion
Acidic in vitro selection of Ca2+-specific RNA-cleaving DNAzymes
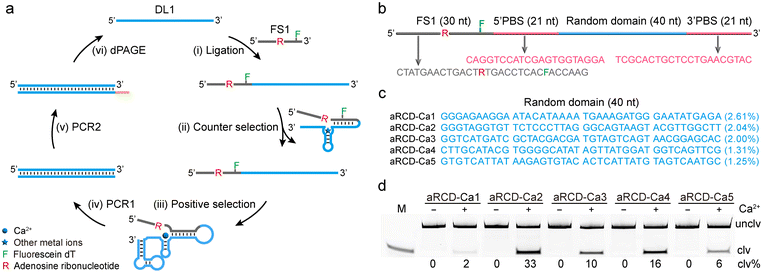
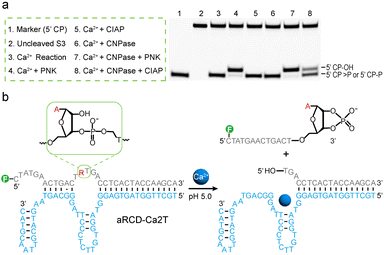
Before performing the acidic in vitro selection, we first examined the Ca2+-dependent cleavage activity of the canonical 8–17 DNAzyme and its variant 17E at pH 7.0 and 4.6. Both displayed high activity at pH 7.0 but were completely inactive at pH 4.6 (Fig. S1), thus laying the foundation for avoiding the recurrence of the 8–17 motif and generating completely new DNAzymes under acidic conditions.The key steps in the acidic in vitro selection of Ca2+-specific DNAzymes are illustrated in Fig. 1a. To isolate Ca2+-specific DNAzymes, we performed an acidic in vitro selection in a low-pH selection buffer (SB) containing 50 mM glycine at pH 4.6, without using other buffer salts to avoid contamination from other metal ions. Positive selection that uses Ca2+ as the target metal ion and counter selection that uses a mixture of competing divalent metal ions (Mg2+, Zn2+, Mn2+, Cu2+, and Pb2+) were both incorporated to promote the selection of DNAzymes with metal specificity.
The starting pool of ∼1014 DNA molecules was initially incubated with 5 mM Ca2+ in the acidic selection buffer at 37 °C for positive selection. DNA molecules that cleaved the substance (FS1) in the presence of Ca2+ were purified using 10% denaturing polyacrylamide gel electrophoresis (dPAGE) and amplified by two rounds of PCR to regenerate the library for the next selection cycle. To enrich for highly active DNAzymes, the incubation time was gradually reduced from 12 hours to 0.5 hours, and the Ca2+ concentration was decreased from 5 mM to 1 mM from round 1 (R1) to round 8 (R8). Counter selection steps were introduced in rounds 6–8 to enhance the metal specificity. Additional experimental details are provided in the SI.
As illustrated in Fig. 1b, an 82-nucleotide (nt) DNA library containing a 40-nt randomized region flanked by two 21-nt primer-binding sites (PBSs) for PCR amplification was used (oligonucleotide sequences are provided in Tables S1 and S2 of the SI). The library was ligated to a 30-nt substrate FS1, which was purposely engineered with a single ribonucleotide adenosine (rA) as the intended cleavage site (R). FS1 was labeled with fluorescein (F) at position 24 to enable fluorescent monitoring of DNAzyme cleavage during selection. Selection progress was monitored by quantifying the cleavage yield (clv%) to confirm the enrichment of Ca2+-specific DNAzymes (Fig. S2). After eight rounds of selection, the cleavage products were collected and sequenced. Many candidate sequences were identified. The five top sequences in the R8 pool are shown in Fig. 1c, and they occupied 9.21% of the total population of R8. The top five candidates were designated aRCD-Ca1 through aRCD-Ca5, where “aRCD-Ca” stands for acidic RNA-cleaving DNAzyme for Ca2+, and the number reflects their enrichment rank in R8. Due to sequence differences, aRCD-Ca1 to aRCD-Ca5 exhibit diverse features in their predicted secondary structures (Fig. S3).
Catalytic proficiency and metal specificity of aRCD-Ca2
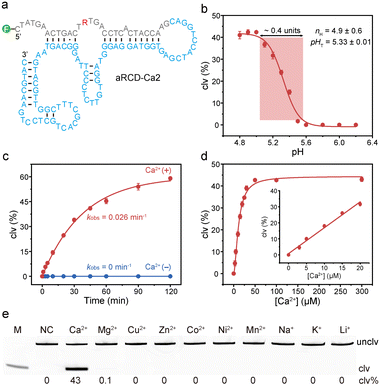
We first assessed the catalytic activity of the top five candidate sequences (aRCD-Ca1 to aRCD-Ca5). Each of them was ligated to 5′-fluorescein-labeled substrate FS2 for testing. After a 1-hour incubation with 0.5 mM Ca2+ in acidic selection buffer, the cleaved and uncleaved products were separated via dPAGE, and the clv% was calculated based on the fluorescence intensity (Fig. 1d). These DNA sequences exhibited varying activities. Notably, aRCD-Ca2 showed the highest activity, achieving a clv% of 33% within 1 hour.The predicted secondary structure of aRCD-Ca2 is shown in Fig. 2a. Since aRCD-Ca2 was selected under acidic conditions, we next examined its cleavage activity across a range of pH values (Fig. S4). The highest cleavage was achieved at pH 5.0 (clv%: 42%), close to the selection condition. Sigmoidal fitting of clv% versus pH revealed a transition midpoint (pHT) of 5.33 ± 0.01 (Fig. 2b). Moreover, aRCD-Ca2 displayed a narrow response range, defined as the pH interval in which it retains 10–90% of maximal activity, spanning only ∼0.4 pH units. The Hill coefficient (nH) was determined to be 4.9 ± 0.6, indicating strong cooperative binding of protons. This high degree of cooperativity enables an ultrasensitive pH response, where small pH shifts lead to pronounced changes in the cleavage activity. This property has been effectively exploited in previous studies to track drug-induced pH changes in cellular lysosomes.34 Notably, a gradual reduction in activity was observed between pH 5.0 and 5.6, suggesting that aRCD-Ca2 may use general acid catalysis, in which protonation is progressively suppressed as the pH increases.40
We next examined the cleavage kinetics of aRCD-Ca2 (Fig. S5). As shown in Fig. 2c, aRCD-Ca2 achieved a clv% of 59% within two hours of reaction in the presence of Ca2+ at pH 5.0 and 37 °C, with a kobs of 0.026 min−1. In contrast, no detectable cleavage was observed in the absence of Ca2+, confirming its dependency on Ca2+.
We quantitatively assessed the cleavage activity of aRCD-Ca2 across a range of Ca2+ concentrations (Fig. S6). As shown in Fig. 2d, aRCD-Ca2 exhibited a clear dose-dependent response to Ca2+ in the range of 0–300 µM, with a linear detection range between 3 and 20 µM (R2 = 0.991). We next evaluated its specificity for metal ions, a key property for DNAzymes in metal detection. The cleavage activity of aRCD-Ca2 was tested in the presence of competing metal ions, including 1 mM divalent cations (Mg2+, Cu2+, Zn2+, Co2+, Ni2+, and Mn2+) and 150 mM monovalent cations (Na+, K+, and Li+) (Fig. 2e). aRCD-Ca2 exhibits high catalytic activity exclusively in the presence of Ca2+, with no detectable activity toward other metal ions. Due to the low pKa of water coordinated to Pb2+, lead ions can efficiently hydrolyze RNA, and many DNAzymes exhibit higher activity with Pb2+ than with other metal ions.16 However, under equivalent metal ion concentrations, aRCD-Ca2 shows minimal cleavage with Pb2+ (clv%: 3%), while retaining high selectivity for Ca2+ (clv%: 45%, Fig. S7). As noted, the Ca2+/Pb2+ discrimination achieved by aRCD-Ca2 is much greater than that of the EtNa DNAzyme, which lacks the selectivity between Ca2+ and Pb2+.41
The success of aRCD-Ca2 suggests that acidic in vitro selection can facilitate the discovery of new DNAzymes with enhanced metal specificity. In contrast, DNAzymes isolated using conventional selection under neutral conditions often struggle to distinguish similar metal ions. For example, the classical 8–17 and 10–23 DNAzymes cannot differentiate between Ca2+ and Mg2+.16,17 Similarly, Mg5 and 17EV1 DNAzymes also fail to discriminate between Ca2+ and Mg2+.22,24 In addition to ionic size and charge density, Mg2+ and Ca2+ also differ in their hydration properties and coordination flexibility. Previous theoretical calculations have shown that Mg2+ forms a rigid six-coordinate hydration structure with a high dehydration and water-exchange energy barrier, whereas Ca2+ can adopt more flexible coordination numbers (6–8) with lower water-exchange barriers.42–47 Compared to Mg2+, Ca2+ more readily undergoes transient coordination rearrangement, which may allow it to interact more effectively with DNA. Acidic conditions could facilitate Ca2+ coordination reorganization, thereby enhancing the discrimination from Mg2+.
Sequence optimization of aRCD-Ca2 by nucleotide truncation
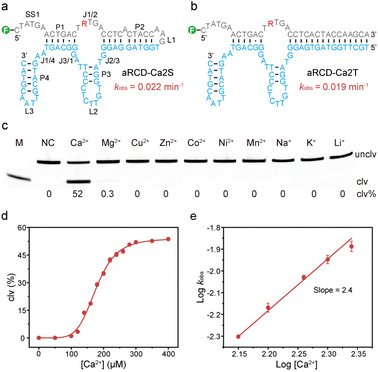
To investigate the functional importance of structural elements and improve the practicality of DNAzymes, we systematically truncated aRCD-Ca2. It was found that 32 nucleotides of aRCD-Ca2 could be removed without affecting its catalytic activity (Fig. S8). This leads to a minimized cis-acting version named aRCD-Ca2S (Fig. 3a). aRCD-Ca2S retains essential structural features, including a single-stranded region (SS1), four short duplexes (P1–P4), three hairpin loops (L1–L3), and four interhelical unpaired junctions (J1/2, J2/3, J3/1, and J1/4).We next evaluated the catalytic performance of aRCD-Ca2S. It exhibited a kobs of 0.022 min−1 and a clv% of 43% within 1 hour in 1× RB at pH 5.0 and 37 °C (Fig. S9), confirming that the truncation did not impair the catalytic efficiency of the parent aRCD-Ca2.
Based on the minimized cis-acting aRCD-Ca2S, we next engineered a trans-acting version by splitting the L1 loop of aRCD-Ca2S. The resulting construct, named aRCD-Ca2T, is shown in Fig. 3b. Unlike cis-acting RCDs, which require ligation to their RNA substrates, trans-acting RCDs bind substrates through Watson–Crick base pairing. We then evaluated the catalytic activity of aRCD-Ca2T by determining its cleavage kinetics. In the presence of Ca2+, aRCD-Ca2T exhibited a kobs of 0.019 min−1, comparable to the original aRCD-Ca2, and achieved a final cleavage yield of 53% within 1 hour (Fig. S10). This catalytic activity exceeds that of EtNa in catalyzing reactions with Ca2+.41 No cleavage activity was observed in the absence of Ca2+, confirming the metal-dependent properties of aRCD-Ca2T.
We further examined the metal specificity of aRCD-Ca2T (Fig. 3c). The results showed that aRCD-Ca2T retained high selectivity for Ca2+ over other competing metal ions. Similarly, its cleavage activity toward Pb2+ remained minimal (clv%: 3%, Fig. S11). We next tested the Ca2+-dependence of aRCD-Ca2T to understand its binding mechanism (Fig. 3d and S12). The clv% was measured with increasing Ca2+ concentrations. Curve fitting shows the sigmoidal behavior characteristic of a cooperative system, suggesting that aRCD-Ca2T requires multiple Ca2+ ions for efficient catalysis.
To quantitatively determine the number of Ca2+ ions involved in catalysis, we measured cleavage kinetics across a range of Ca2+ concentrations (Fig. S13a). The kinetics were fitted to determine kobs at each Ca2+ concentration (Fig. S13b). By fitting the data to the equation kobs = k[Ca2+]n,48 a double-logarithmic plot of kobsversus Ca2+ concentration (Fig. 3e) yielded a slope of 2.4, indicating that at least two Ca2+ ions are involved in the binding and catalytic process.
Characterization of the cleavage products and site selectivity of aRCD-Ca2T
Because deprotonation of the RNA 2′-hydroxyl is disfavored at low pH, the catalytic mechanism of aRCD-Ca2T under acidic conditions may differ from that at neutral pH. We therefore investigated whether RNA cleaving generates distinct terminal groups at low pH.29 To investigate the 3′-phosphate moieties generated on the 5′-cleavage product (5′ CP), the RNA-containing substrate was incubated with aRCD-Ca2T in the presence of Ca2+ at pH 5.0 for 1 hour. The resulting products were then treated with various phosphodiesterases for terminal group analysis, including polynucleotide kinase (PNK), calf intestinal alkaline phosphatase (CIAP), and 2′,3′-cyclic nucleotide 3′-phosphodiesterase (CNPase). These enzymes differ in catalytic specificity: PNK possesses both 2′,3′-cyclic nucleotide 3′-phosphodiesterase and 2′/3′-phosphatase activities; CIAP exhibits only 2′/3′-phosphatase activity; while CNPase hydrolyzes 2′,3′-cyclic phosphate to 2′-phosphate.PNK treatment reduced the gel mobility of 5′ CP, likely due to the removal of the negatively charged phosphate group (Fig. 4a, lane 4). In contrast, CIAP treatment had no effect on its migration (Fig. 4a, lane 5), and CNPase treatment similarly caused no change (Fig. 4a, lane 6). However, sequential treatment with CNPase followed by either PNK (Fig. 4a, lane 7) or CIAP (Fig. 4a, lane 8) resulted in a decreased mobility of 5′ CP.
These results suggest that aRCD-Ca2T-mediated RNA-cleavage produces a 5′-cleavage fragment bearing a 2′,3′-cyclic phosphate and a 3′-cleavage fragment with a 5′-OH (Fig. 4b), which is similar to the cleavage products generated by conventional RCDs. To confirm the identity of the cleaved product, electrospray ionization mass spectrometry (ESI-MS) analysis of the 5′-cleavage fragment was performed, confirming that the cleavage product contains a 2′,3′-cyclic phosphate (Fig. S14).
We investigated the cleavage-site selectivity of aRCD-Ca2T. The original rAT cleavage site in the substrate was replaced with rCT, rUT, rAC, or rGC, and the catalytic activity of aRCD-Ca2T toward these substrates was examined (Fig. S15). Replacement of the adenosine ribonucleotide (rA) at the cleavage site with cytidine ribonucleotide (rC) or uridine ribonucleotide (rU) resulted in nearly undetectable cleavage activity. Similarly, substitution of the rAT cleavage site with rAC or rGC abolished catalytic activity. These results demonstrate that aRCD-Ca2T is highly specific for the rAT cleavage site. Adenosine is known to undergo protonation under acidic conditions,40 which may contribute to the observed site selectivity of aRCD-Ca2T.
Visualizing lysosomal Ca2+ in HT22 hippocampal neuron cells
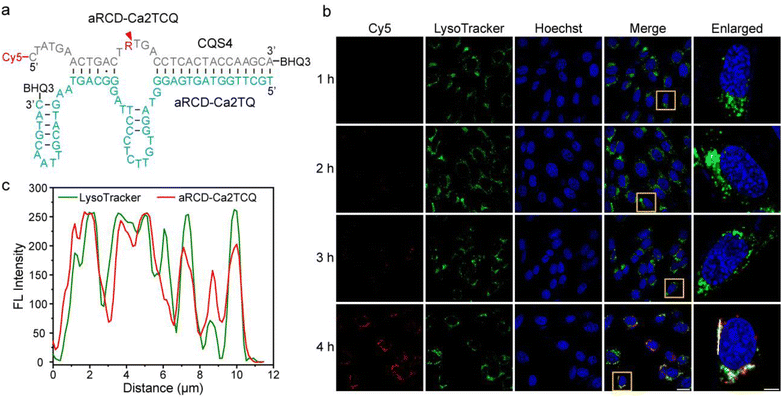
To convert aRCD-Ca2T into a fluorescent probe, we first employed a catalytic beacon design (named aRCD-Ca2TCQ), in which the RNA-containing substrate strand was labeled with a fluorophore (Cy5) at the 5′ end and a quencher (BHQ3) at the 3′ end of the enzyme strand. To further reduce background fluorescence, the substrate strand (CQS4) was additionally labeled with BHQ3 (Fig. 5a). In the presence of Ca2+, aRCD-Ca2T can catalyze the cleavage of the substrate strand, resulting in fluorescence generation. We first evaluated the biostability of aRCD-Ca2TCQ in cell transfection medium (Opti-MEM). dPAGE results indicated that the construct remained highly stable over an 8-hour incubation (Fig. S16). We also assessed its cellular compatibility in HT22 hippocampal neuron cells using a standard MTT assay, which showed no significant reduction in cell viability (Fig. S17).Given the excellent in vitro performance of aRCD-Ca2T, we next investigated its potential for sensing Ca2+ in living cells. Lysosomes have recently been recognized as acidic Ca2+ reservoirs, with luminal Ca2+ concentrations maintained at approximately 0.5 mM—nearly 5000 times higher than that of the cytosol (∼100 nM).49 Lysosomal Ca2+ plays a crucial role in regulating diverse cellular processes, including vesicle fusion, intracellular signaling, transcriptional regulation, lysosomal dynamics, autophagy, and exocytosis.50 However, the absence of Ca2+-specific DNAzymes that function exclusively under acidic conditions has hindered the accurate quantification of Ca2+ in acidic organelles. Therefore, we investigated the potential of aRCD-Ca2T for visualizing lysosomal Ca2+ fluctuations in living cells.
To evaluate lysosomal colocalization, HT22 hippocampal neuron cells were incubated with aRCD-Ca2TCQ for various durations. At 4 hours post-transfection, colocalization analysis between the Cy5 and LysoTracker channels yielded a Pearson correlation coefficient of 0.80, demonstrating its strong lysosomal localization (Fig. 5b). Fluorescence intensity profiling analysis along the white line in the merged image further confirmed the spatial overlap between these two signals (Fig. 5c). We observed a gradual increase in red fluorescence in the Cy5 channel over time, indicative of DNAzyme activation. In contrast, a mutant version (maRCD-Ca2TCQ), containing critical nucleotide substitutions in the J2/3 junction (GT replaced with AA) and the J3/1 junction (TAG replaced with ATT), showed no appreciable signal increase (Fig. S18). These results demonstrate that aRCD-Ca2TCQ is specifically activated by lysosomal Ca2+, enabling visualization of endogenous Ca2+ within lysosomes.
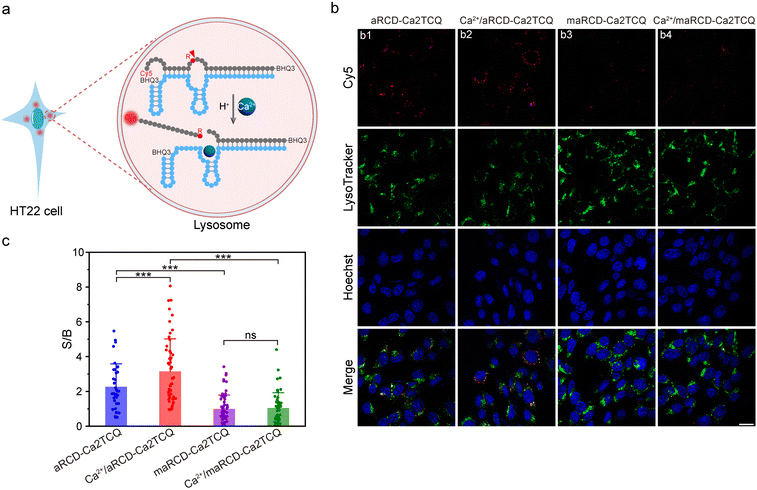
To verify the responsiveness of aRCD-Ca2TCQ to changes in lysosomal Ca2+ levels (Fig. 6a), we introduced exogenous Ca2+ and applied ionomycin to facilitate the equilibration of Ca2+ across cellular membranes.39,51 As shown in Fig. 6b, the fluorescence observed in the Cy5 channel was attributed to the activation of aRCD-Ca2TCQ by lysosomal Ca2+ (Image b1). Upon Ca2+ treatment, a marked increase in Cy5 fluorescence intensity was observed (Image b2), confirming that the probe is capable of detecting changes in lysosomal Ca2+ levels. In contrast, cells transfected with the mutant maRCD-Ca2TCQ displayed minimal fluorescence signals, regardless of Ca2+ influx (Images b3 and b4), thus highlighting the high specificity of aRCD-Ca2TCQ for Ca2+. Quantitative fluorescence analysis revealed that HT22 hippocampal neuron cells transfected with aRCD-Ca2TCQ exhibited 2.3-fold and 3.1-fold higher fluorescence intensities in the absence and presence of Ca2+ influx, respectively, compared to cells treated with the mutant probe (Fig. 6c). Together, these results demonstrate that aRCD-Ca2TCQ enables reliable monitoring of lysosomal Ca2+ fluctuations in living cells.
Conclusions
In summary, we have demonstrated an innovative approach for enhancing the metal specificity in DNAzyme evolution by harnessing acidic in vitro selection. While certain RCDs capable of recognizing Ca2+ have been reported, no Ca2+-specific DNAzymes have previously been directly isolated via in vitro selection under acidic conditions. Leveraging this strategy, we successfully isolated a Ca2+-specific aRCD from a random DNA library by performing positive selection in the presence of Ca2+ at low pH, followed by counter selection against competing metal ions (Mg2+, Zn2+, Mn2+, Cu2+, and Pb2+). The resulting aRCD-Ca2 exhibited robust activity and exceptional discrimination against other competing metals at pH 5.0. This strategy can be readily adapted to isolate new high-specific DNAzymes for other metal ions.Although the role of Ca2+ in the catalysis of aRCD-Ca2 remains to be fully elucidated, the reduced activity with increasing pH from 5.0 to 5.6 supports the notion that RNA cleavage by aRCD-Ca2 likely proceeds through general acid catalysis. The specific recognition principles and catalytic mechanisms still need to be clarified through high-resolution techniques, such as crystallography and nuclear magnetic resonance (NMR), to gain a deeper understanding of its molecular structures. We further engineered a trans-acting variant, aRCD-Ca2T, which displays strongly cooperative binding to Ca2+, a feature that likely underlies its high metal specificity. Based on this trans-acting design, we developed a fluorescent probe, aRCD-Ca2TCQ, for imaging endogenous lysosomal Ca2+ in HT22 hippocampal neuron cells, thereby offering a highly selective potential tool for studying Ca2+-mediated biological processes within lysosomes.
Together, these findings not only elucidate the potential of specific metal–DNA interactions in acidic media but also expand the toolkit for selective metal sensing in complex biological environments. Future work will focus on extending this platform to other biologically relevant metal ions and combining these DNAzymes into diagnostic and therapeutic applications.
Author contributions
P. J. was responsible for the conception and design of the study, conducted the experiments, analyzed the data, and drafted the manuscript. S. L., W. X., S. S. X., Q. Z., J. X. L., and Y. W. assisted in revising the manuscript. Y. Y. C., Z. J. Z., and M. L. contributed to the study design, provided guidance during the research, and assisted in manuscript editing. All authors read and approved the final manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
All the data supporting this study are included in the main text and the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5sc08656f.Acknowledgements
This work was supported by the National Key R&D Program (2023YFC3711100), National Natural Science Foundation of China (NSFC; 22425602), Dalian Science and Technology Innovation Fund (2023YGZD04), Dalian Science and Technology Talent Innovation Support Program (2024RJ001) and Program of Introducing Talents of Discipline to Universities (B25041).Notes and references
- B. Cuenoud and J. W. Szostak, Nature, 1995, 375, 611–614 CrossRef CAS PubMed.
- R. R. Breaker, Nat. Biotechnol., 1997, 15, 427–431 CrossRef CAS PubMed.
- R. R. Breaker and G. F. Joyce, Chem. Biol., 1994, 1, 223–229 CrossRef CAS PubMed.
- H. Gu, K. Furukawa, Z. Weinberg, D. F. Berenson and R. R. Breaker, J. Am. Chem. Soc., 2013, 135, 9121–9129 CrossRef CAS PubMed.
- Q. Hu, Z. Tong, A. Yalikong, L.-P. Ge, Q. Shi, X. Du, P. Wang, X.-Y. Liu, W. Zhan, X. Gao, D. Sun, T. Fu, D. Ye, C. Fan, J. Liu, Y.-S. Zhong, Y.-Z. Jiang and H. Gu, Nat. Chem., 2023, 16, 122–131 CrossRef PubMed.
- T. Chang, G. Li, D. Chang, R. Amini, X. Zhu, T. Zhao, J. Gu, Z. Li and Y. Li, Angew. Chem., Int. Ed., 2023, 62, e202310941 CrossRef CAS PubMed.
- J. Shi, Q. Zhang, Y. Wu, Y. Chang and M. Liu, Chem. Sci., 2024, 15, 13452–13458 RSC.
- X. Li, Y. Chang, Y. Wu and M. Liu, Chem. Sci., 2024, 15, 2996–3002 RSC.
- S. K. Silverman, Acc. Chem. Res., 2009, 42, 1521–1531 CrossRef CAS PubMed.
- S. K. Silverman, Angew. Chem., Int. Ed., 2010, 49, 7180–7201 CrossRef CAS PubMed.
- B. Dou, H. Shen, Z. Li, H. Cheng and P. Wang, Chem. Sci., 2025, 16, 3470–3478 RSC.
- D. Luo, X. Lin, Y. Zhao, J. Hu, F. Mo, G. Song, Z. Zou, F. Wang and X. Liu, Chem. Sci., 2022, 13, 5155–5163 RSC.
- R. J. Lake, Z. Yang, J. Zhang and Y. Lu, Acc. Chem. Res., 2019, 52, 3275–3286 CrossRef CAS PubMed.
- E. M. McConnell, I. Cozma, Q. Mou, J. D. Brennan, Y. Lu and Y. Li, Chem. Soc. Rev., 2021, 50, 8954–8994 RSC.
- C. Hong, Q. Wang, Y. Chen, Y. Gao, J. Shang, X. Weng, X. Liu and F. Wang, Chem. Sci., 2021, 12, 15339–15346 RSC.
- W. Zhou, R. Saran and J. Liu, Chem. Rev., 2017, 117, 8272–8325 CrossRef CAS PubMed.
- S. W. Santoro and G. F. Joyce, Proc. Natl. Acad. Sci. U. S. A., 1997, 94, 4262–4266 Search PubMed.
- A. K. Brown, J. Li, C. M. B. Pavot and Y. Lu, Biochemistry, 2003, 42, 7152–7161 Search PubMed.
- H. Liu, X. Yu, Y. Chen, J. Zhang, B. Wu, L. Zheng, P. Haruehanroengra, R. Wang, S. Li, J. Lin, J. Li, J. Sheng, Z. Huang, J. Ma and J. Gan, Nat. Commun., 2017, 8, 2006 Search PubMed.
- J. Wieruszewska, A. Pawłowicz, E. Połomska, K. Pasternak, Z. Gdaniec and W. Andrałojć, Nat. Commun., 2024, 15, 4218 Search PubMed.
- J. Li, W. Zheng, A. H. Kwon and Y. Lu, Nucleic Acids Res., 2000, 28, 481–488 CrossRef CAS PubMed.
- D. Faulhammer and M. Famulok, Angew. Chem., Int. Ed., 1996, 35, 2837–2841 CrossRef CAS.
- K. L. Y. Schlosser, Biochemistry, 2004, 43, 9695–9707 CrossRef CAS PubMed.
- W. Zhou, Y. Zhang, J. Ding and J. Liu, ACS Sens., 2016, 1, 600–606 CrossRef CAS.
- H. Rosenbach, J. Victor, M. Etzkorn, G. Steger, D. Riesner and I. Span, Molecules, 2020, 25, 3100 CrossRef CAS PubMed.
- J. Borggräfe, J. Victor, H. Rosenbach, A. Viegas, C. G. W. Gertzen, C. Wuebben, H. Kovacs, M. Gopalswamy, D. Riesner, G. Steger, O. Schiemann, H. Gohlke, I. Span and M. Etzkorn, Nature, 2022, 601, 144–149 CrossRef PubMed.
- Z. Liu, S. H. Mei, J. D. Brennan and Y. Li, J. Am. Chem. Soc., 2003, 125, 7539–7545 CrossRef CAS PubMed.
- S. A. Kandadai, Nucleic Acids Res., 2005, 33, 7164–7175 CrossRef CAS PubMed.
- S. A. Kandadai, W. W. K. Mok, M. M. Ali and Y. Li, Biochemistry, 2009, 48, 7383–7391 CrossRef CAS PubMed.
- Y. Shen, J. D. Brennan and Y. Li, Biochemistry, 2005, 44, 12066–12076 CrossRef CAS PubMed.
- Q. Zhou, G. Zhang, Y. Wu, Q. Zhang, Y. Liu, Y. Chang and M. Liu, J. Am. Chem. Soc., 2023, 145, 21370–21377 CrossRef CAS PubMed.
- Y. Li and R. R. Breaker, J. Am. Chem. Soc., 1999, 121, 5364–5372 CrossRef CAS.
- Y. Lee, R. Roy, S. Gu, S. B. Shetty, A. K. Rangadurai and H. M. Al-Hashimi, J. Am. Chem. Soc., 2025, 147, 19643–19666 CrossRef CAS PubMed.
- P. Jia, Q. Meng, Q. Zhang, R. Zhang, Z. Zhang, J. Li, X. Wang, Y. Liu, Y. Chang and M. Liu, Anal. Chem., 2025, 97, 9378–9385 CrossRef CAS PubMed.
- E. Koculi, C. Hyeon, D. Thirumalai and S. A. Woodson, J. Am. Chem. Soc., 2007, 129, 2676–2682 CrossRef CAS PubMed.
- T. Ikeda, M. Boero and K. Terakura, J. Chem. Phys., 2007, 127, 074503 CrossRef PubMed.
- I. Persson, Liquids, 2022, 2, 210–242 CrossRef CAS.
- Z. Sun, W. Zhang, M. Ji, R. Hartsock and K. J. Gaffney, J. Phys. Chem. B, 2013, 117, 12268–12275 CrossRef CAS PubMed.
- N. Narayanaswamy, K. Chakraborty, A. Saminathan, E. Zeichner, K. Leung, J. Devany and Y. Krishnan, Nat. Methods, 2018, 16, 95–102 CrossRef PubMed.
- L. Ma, S. Kartik, B. Liu and J. Liu, Nucleic Acids Res., 2019, 47, 8154–8162 CrossRef CAS PubMed.
- W. Zhou, R. Saran, P.-J. J. Huang, J. Ding and J. Liu, ChemBioChem, 2017, 18, 518–522 CrossRef CAS PubMed.
- Y. I. Neela, A. S. Mahadevi and G. N. Sastry, Struct. Chem., 2013, 24, 637–650 CrossRef CAS.
- D. Rutkowska-Zbik, M. Witko and L. Fiedor, J. Mol. Model., 2013, 19, 4661–4667 CrossRef CAS PubMed.
- J. Weston, The Chemistry of Organomagnesium Compounds, 2008, pp. 315–367 Search PubMed.
- S. F. Lincoln, D. T. Richens and A. G. Sykes, Compr. Coord. Chem. II, 2003, 1, 515–555 Search PubMed.
- M. Pavlov, P. E. M. Siegbahn and M. Sandström, J. Phys. Chem. A, 1998, 102, 219–228 CrossRef CAS.
- P.-C. Cheng, F.-S. Tseng, C.-T. Yeh, T.-G. Chang, C.-C. Kao, C.-H. Lin, W.-R. Liu, J.-S. Chen and V. Zima, CrystEngComm, 2012, 14, 6812–6822 RSC.
- P.-J. J. Huang, D. de Rochambeau, H. F. Sleiman and J. Liu, Angew. Chem., Int. Ed., 2020, 59, 3573–3577 CrossRef CAS PubMed.
- J. Yang, Z. Zhao, M. Gu, X. Feng and H. Xu, Protein Cell, 2018, 10, 8–19 CrossRef PubMed.
- C. Settembre and R. M. Perera, Nat. Rev. Mol. Cell Biol., 2023, 25, 223–245 CrossRef PubMed.
- Y. Kuronuma, Y. Shindo, R. Kumada, A. Sakama, D. Citterio, K. Oka and Y. Hiruta, ACS Chem. Neurosci., 2025, 16, 649–658 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |