Open Access Article
Open Access ArticleOptically switchable CLEAR probes enable rapid, biocompatible and high-efficiency fluorophore exchange for ultra-plex, high-resolution immunofluorescence imaging
Simanta
Kalita
a,
Pratibha
Kumari
a,
Arka
Som
a,
Srishti
Mandal
 b,
Resmi V.
Nair
a,
Saswata
Bandyopadhyay
a,
Sushma
Rao
c,
Sudha
Kumari
b and
Sarit S.
Agasti
b,
Resmi V.
Nair
a,
Saswata
Bandyopadhyay
a,
Sushma
Rao
c,
Sudha
Kumari
b and
Sarit S.
Agasti
 *a
*a
aNew Chemistry Unit and Chemistry & Physics of Materials Unit, School of Advanced Materials (SAMat), Jawaharlal Nehru Centre for Advanced Scientific Research (JNCASR), Bangalore, Karnataka 560064, India. E-mail: sagasti@jncasr.ac.in
bDepartment of Microbiology and Cell Biology, Indian Institute of Science (IISc), Bangalore, Karnataka 560012, India
cNeuroscience Unit, Jawaharlal Nehru Centre for Advanced Scientific Research (JNCASR), Bangalore, Karnataka 560064, India
First published on 25th February 2026
Abstract
Immunofluorescence imaging remains a foundational technique in biological research, enabling visualization of protein targets within native environments. However, the ability of conventional immunofluorescence approaches to fully capture biological complexity is limited by low multiplexing capacity, typically restricted to four or five targets due to spectral overlap among fluorophores. Recent advances, including DNA-labelled antibodies or chemical fluorescence ablation, aim to enhance multiplexing but face considerable challenges. DNA-based approaches require non-conventional reagents and are operationally complex, while chemical clearing methods are time-intensive and risk sample damage from harsh reagents. To address these limitations, we introduce CLEAR (Cleavable Light-Erased Antibody Reporter) probes—a simple, rapid, and broadly applicable platform for scalable multiplexing using conventionally conjugated antibodies. Through a highly optimized bottom-up molecular design, CLEAR probes deliver one of the fastest and most efficient signal-clearing strategies, achieving >98% fluorescence removal in under two minutes using a mild, non-toxic light dose. This allows their integration into iterative immunostaining workflows and, importantly, preserves sample and image quality across multiple staining cycles for supporting high-throughput, potentially unlimited multiplexing. Moreover, the excellent biocompatibility of this approach enables its extension from fixed to live-cell and tissue imaging. Its compatibility with diverse fluorophores makes it readily adaptable to advanced imaging modalities, including super-resolution microscopy. Finally, we demonstrate the synergistic potential of the CLEAR platform—where ultrafast, efficient, and biocompatible fluorophore clearance allows high-dimensional interrogation of coordinated cytoskeletal remodeling and organelle redistribution during dynamically regulated immunological synapse formation.
Introduction
Biological systems are fundamentally multi-scale, with networks of interacting biomolecules, subcellular structures, and cellular communities operating in concert to elicit emergent function at the cell, tissue, and organism scale. To interrogate these systems in sufficient depth—whether during development, homeostasis, or disease progression—researchers seek to map biomolecules across several orders of magnitude, from nanometer-scale subcellular distributions to micrometer-scale single cells and millimeter-to-centimeter-scale tissues.1,2 Considering that proteins are the primary functional molecules of the cell, mapping the spatial landscape of the proteome remains a pivotal goal for bringing transformative insights in this direction, thereby stimulating the recent interest in single-cell proteomics and spatial proteomics imaging technologies.2–7 To meet these demands, a versatile imaging platform capable of highly multiplexed protein mapping with high resolution and rapid throughput—while maintaining compatibility with delicate biological specimens—is essential; yet such technologies remain exceedingly scarce.8 In recent years, mass spectrometry (MS)-based imaging techniques (e.g., multiplexed ion beam imaging (MIBI) and imaging mass cytometry (IMC)) by employing metal-isotope-labeled antibodies have been developed to increase multiplexing capability.9–11 However, MS-based techniques suffer from limited resolution and rely on point-scanning of small fields (∼2–5 min per 50 µm × 50 µm), making them inherently slow for imaging large, centimetre-scale tissue sections. In addition, they require complex instrumentation and highly specialized personnel to manage the operational complexity.12 Moreover, the necessary vacuum stability of the samples hinders the utility of these techniques in profiling delicate samples and living specimens.Fluorescence microscopy, on the other hand, is a powerful and non-invasive imaging tool that has been widely adopted in biological research. It offers exceptional sensitivity and specificity for visualizing molecules in complex biological systems, including live specimens. With its high accessibility and high-throughput acquisition capability, it has been extensively adopted in biological research and integrated into pathological workflows for disease diagnostics, particularly through immunofluorescence imaging.13,14 In recent years, the emergence of super-resolution imaging techniques has further revolutionized fluorescence microscopy, making it possible to precisely map biomolecules at resolutions down to a few nanometres.3,5,15–17 At the same time, integrating these approaches with diffraction-limited modalities such as confocal microscopy extends imaging to larger tissue dimensions, thereby linking nanoscale molecular insights with tissue-level context. Despite its remarkable success in biological exploration and biomedical research, fluorescence microscopy faces a significant challenge in terms of its multiplexing capabilities—the ability to visualize a large number of distinct molecular species within a single sample. This bottleneck arises from spectral overlap between fluorophores, limiting the number of targets that can be reliably distinguished within a single sample to approximately 4–5. To address this challenge and enhance the multiplexing capability of fluorescence microscopy, innovative strategies have begun to emerge. One particularly promising approach is fluorescence erasing (or signal-clearing), which allows researchers to reuse the same optical detection window for different analytes.18 By employing iterative cycles of targeting, imaging, and signal-clearing, researchers can image additional molecular targets without interference from previously labeled fluorophores, thereby substantially increasing the number of species visualized within a single sample. In the earlier implementations, the signal-removal step relied on stripping the antibody labels or deactivating fluorophores under harsh chemical conditions,19–24 a process that risked altering tissue architecture and damaging biomolecules, thereby compromising protein localization, antigen recognition, and overall imaging accuracy in downstream data interpretation. In recent years, more refined strategies have emerged, notably DNA–antibody conjugate-based approaches and chemically triggered signal-removal methods, such as bioorthogonal reactions or disulfide-cleaving reducing agents, that enable controlled detachment of fluorophores from antibodies.4,5,25–35 In DNA-based strategies, DNA-barcoded antibodies are visualized through hybridization with complementary fluorescently tagged oligonucleotides, after which fluorescence signals are erased through mechanisms such as washing with duplex-destabilizing buffers or toehold-mediated strand-displacement reactions, among other approaches.5,36–38 Recent integrations of signal amplification strategies, including immuno-SABER, QD-SABER, CAD-HCR, and POSA, have further improved the sensitivity of these methods, enabling imaging of low-abundance proteins.4,38–40 Notably, the recently reported POSA strategy employs optical amplification using highly bright fluorescent π-conjugated polymers (FCPs), whose intrinsic brightness simplified the workflow while enabling rapid and sensitive multiplexed imaging through toehold-mediated strand displacement-based signal erasing.38 While these DNA-based approaches represent a powerful and rapidly evolving direction, DNA–antibody conjugates are non-conventional probe architectures that introduce additional synthetic challenges, sequence design requirements, and careful optimization of hybridization and strand-displacement conditions, thereby increasing experimental complexity and often requiring specialized expertise. Moreover, the incorporation of large, highly charged nucleic acid moieties into the antibody structure can compromise target recognition and significantly increase nonspecific interactions.41,42 On the other hand, chemically triggered signal-removal methods that enable detachment of fluorophores from antibodies are based on small-molecule fluorophore-conjugated antibodies and build upon established immunofluorescence workflows;31–34 however, they typically demand rigorous liquid handling and manipulation. Except for rare cases,31 signal removal often requires prolonged reaction times, and achieving consistently high removal efficiency can remain challenging, thereby limiting the number of achievable imaging cycles. Even in those more reactive cases, reports suggest the possibility of unexpected nonspecific proteome reactivity, which may potentially interfere with antibody binding during iterative staining rounds.43 In addition, the chemical trigger must be meticulously eliminated to avoid interfering with subsequent fluorophore labeling cycles—factors that collectively extend the overall cycle time and pose particular challenges when applied to delicate tissue samples or live-cell systems. To address these limitations, we were interested in developing a distinct solution that combines the operational simplicity of conventional small-molecule antibody–fluorophore conjugates with a rapid, efficient, biocompatible, and externally regulated signal-erasing mechanism.
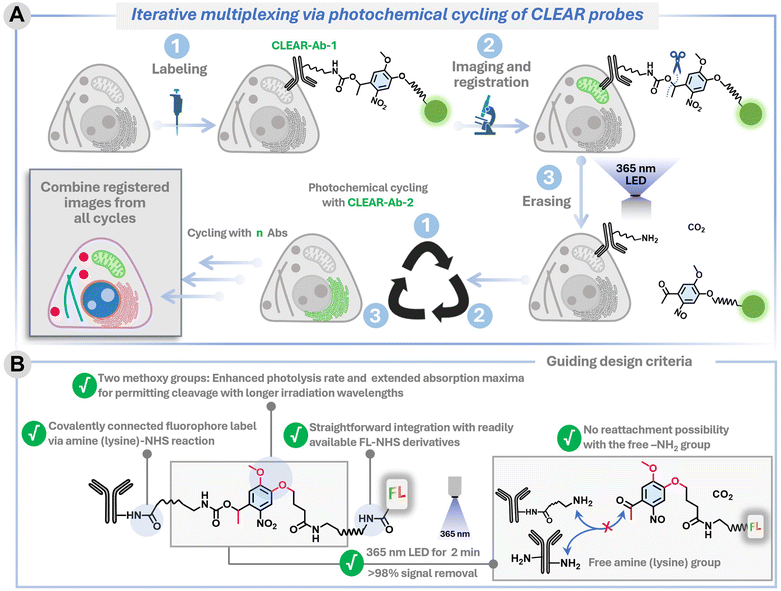
Light provides a gentle and minimally invasive alternative widely used in biology for stimulating reactions. Unlike chemical methods, light can be controlled remotely and toggled on and off with ease, simplifying workflows and eliminating the need for complex liquid handling steps. Early efforts to employ light for multiplexing involved photobleaching fluorophores to erase signals after each imaging cycle.44 However, this approach has several limitations. Fluorophores must be photobleached individually by illuminating each at its respective excitation wavelength. Moreover, since the microscope laser can only target a small area at a time (i.e., the field of view), this process is inherently slow, resulting in low throughput and limited area coverage. Photobleaching can be particularly time-consuming for photostable fluorophores, which are increasingly favoured in modern imaging applications. Additionally, the photobleaching process generates reactive oxygen species, which can compromise the integrity of both the sample and biomolecules.45,46 A more efficient and non-destructive alternative involves the use of photoinduced bond cleavage reactions, which can proceed under mild, non-toxic light doses, thereby minimizing photodamage and preserving sample integrity.47–50 Another key advantage of this approach is that a single type of bond-cleaving reaction can facilitate the simultaneous removal of multiple fluorophores, irrespective of their specific spectral properties. In addition, the incorporation of a precisely engineered photoresponsive moiety can permit efficient bond cleavage using low-intensity, broad-area illumination from handheld LED sources, thereby enabling rapid and high-throughput signal clearance with wide spatial coverage. Despite this promising potential, a broadly generalizable light-driven signal-clearing platform has not yet been realized, one that builds upon conventionally conjugated antibodies, operates rapidly and with high efficiency, remains compatible with diverse biological samples (including live cells), and is adaptable to virtually any fluorophore. To address this gap, we introduce a highly efficient and universally applicable signal-clearing platform, CLEAR (Cleavable Light-Erased Antibody Reporter) probes, which achieve >98% signal removal within two minutes under a mild, non-toxic light dose (Fig. 1A). When combined with iterative immunostaining, CLEAR probes establish a powerful multiplexing platform capable of profiling a large number of biological targets across diverse sample types. Critically, CLEAR probes provide a gentle means of near-quantitative signal erasure, which minimizes background accumulation across multiple staining cycles and preserves sample integrity—features essential for sustaining high signal-to-background and image quality during large-scale serial profiling. Moreover, the platform is readily extendable to live-cell imaging and compatible with state-of-the-art fluorophores for advanced imaging modalities such as dSTORM (direct stochastic optical reconstruction microscopy) super-resolution microscopy. As a proof of concept, we first visualized six distinct targets using a single fluorophore and subsequently expanded the workflow to accommodate three spectrally distinct fluorophores per cycle. Finally, we demonstrated the broader utility of this multiplexing platform by spatially mapping ten proteins to investigate cytoskeletal remodeling and organelle redistribution during immunological synapse formation.
Results and discussion
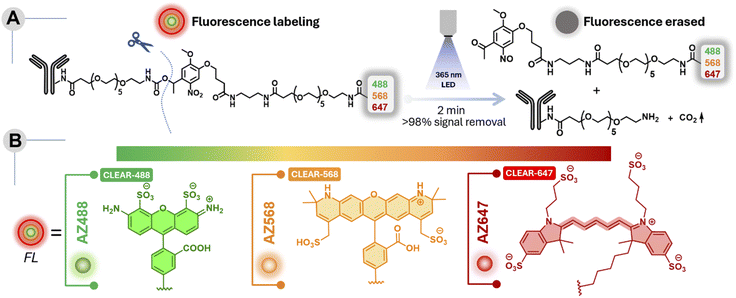
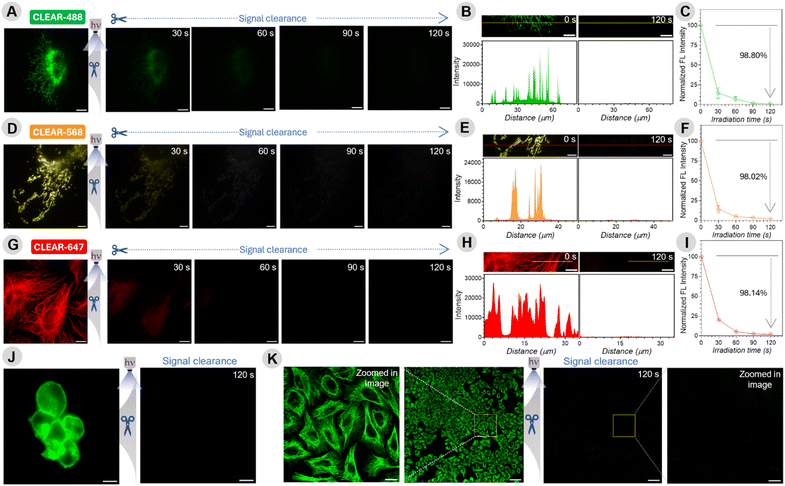
The development of this multiplexing platform was driven by several key design considerations aimed at overcoming the existing challenges and enhancing its potential for widespread adoption (Fig. 1). (1) It leverages iterative immunostaining with fluorophore–antibody conjugates that are covalently coupled—a robust and commercially viable approach that ensures stable and reproducible labeling, making it broadly adoptable in biological laboratories. While alternative fluorophore-coupling strategies, such as the use of DNA hybridization or secondary binders, are successful in many contexts,5,48 they often require extensive binding optimization, risk cross-reactivity if the same binder is used across different targets, and necessitate fresh preparation of constructs for each experiment, thereby precluding off-the-shelf availability. (2) Our antibody labeling method employs an amine-NHS ester reaction to conjugate engineered fluorophore tags to lysine residues—a widely used and robust approach that ensures stable attachment and prolongs reagent shelf life. This method is simple, versatile, and suitable for labeling small antibody quantities with minimal purification. In contrast, the other two-tag conjugation methods, such as DBCO-azide chemistry, require pre-conjugation separation steps, leading to potential antibody loss, increased batch-to-batch variability, and limited scalability. (3) Our multiplexing strategy is designed around a “plug-and-play” concept, allowing the direct integration of virtually any commercially available fluorophore without necessitating structural modifications or re-engineering of the dye. This flexibility enables researchers to select any off-the-shelf fluorophores tailored to meet specific experimental requirements—for instance, dyes optimized for super-resolution microscopy. (4) Our signal-clearance strategy is centered around the highly robust and extensively validated ortho-nitrobenzyl (ONB) photoresponsive moiety, recognized for its stability, ease of handling, and proven performance in diverse biological contexts. Through further molecular engineering of this photoresponsive moiety, we achieved highly efficient signal clearance within minutes at low light doses and non-toxic wavelengths (>350–405 nm), all while preserving the high signal-to-background ratio required for cyclic profiling and enabling use in live-cell applications (Fig. 1B). Another key innovation in our design is the prevention of fluorophore reattachment after cleavage (Fig. 1B)—a limitation of many conventional ONB derivatives that generate reactive aldehyde (–CHO) byproducts.51,52 By employing a ketone-generating variant instead, we effectively eliminated this reattachment issue, thereby ensuring traceless release and seamless performance across multiple imaging cycles. Together, these design considerations make our multiplexing platform both powerful and accessible, offering a streamlined approach that supports high-throughput and highly multiplexed fluorescence imaging, poised for facile integration into a wide range of biological exploration. SI Scheme S1 depicts the synthetic route employed to construct the CLEAR multiplexing probes. The overall process consists of four main steps: (1) synthesis of the photoresponsive bifunctional linker: we first synthesized an ONB-based bifunctional linker bearing an amine group on one terminus and a carboxylic acid on the other, connected by a polyethylene glycol (PEG) spacer. Incorporating PEG improves water solubility, reduces nonspecific interactions, and helps spatially separate the ONB moiety from the fluorophore to minimize fluorescence quenching. The ONB core was selected based on a few specific criteria, including chemical stability, compatibility with aqueous biological environments, and ease of structural modification. The ONB core is strategically designed with two methoxy groups, thereby enhancing the rate of photolysis and shifting the absorption maximum to enable photocleavage under longer-wavelength irradiation—important for fast and highly efficient photocleavage and also to extend its utility for live-cell applications.52 Furthermore, introducing a methyl substituent at the benzylic position yields the photolysis product in the form of a ketone rather than an aldehyde, thereby minimizing any risk of reattachment through imine formation with biomolecules (e.g., free lysine from antibody). (2) Fluorophore conjugation: the amine terminus of the photoresponsive linker was coupled with an N-hydroxysuccinimide (NHS)-activated fluorophore-PEG derivative. This step effectively installs the fluorescent label onto one side of the ONB linker. (3) NHS activation of the carboxylic acid terminus: the free carboxylic acid on the opposite end of the linker was converted to its NHS ester form. This activation step is essential for subsequent conjugation to biomolecules, as NHS esters rapidly and efficiently react with primary amines in lysine under mild conditions. (4) Antibody conjugation: finally, the NHS-activated photoresponsive fluorophore conjugate was reacted with antibodies against various target proteins, forming a stable amide bond between the antibody and the linker. The resulting constructs are photocleavable fluorophore–antibody probes whose fluorescence can be selectively erased upon light irradiation, enabling iterative labeling and multiplexed imaging. These probes were termed CLEAR (Cleavable Light-Erased Antibody Reporter) probes. The conjugation efficiency was evaluated by determining the degree of labeling (DOL) from UV-vis absorption spectra of the antibody conjugates, yielding approximately 1.4–2.7 fluorophores per antibody when five equivalents of NHS ester derivatives of the photocleavable linker–fluorophore conjugates were used (see Table S5, SI). This labeling range is comparable to that obtained using conventional dye–NHS ester conjugation and lies within the optimal range for fluorescence imaging. To enable simultaneous imaging of three molecular targets per imaging cycle, we incorporated three spectrally distinct Alexa Fluor dyes—each rendered photocleavable through conjugation to the ONB linker—and independently coupled them to antibodies against different protein targets. We modified a diverse set of targeting agents—including primary antibodies, secondary antibodies, and small molecules—to enable flexible and application-specific use of CLEAR probes. Fig. 2A shows the precise molecular connectivity between the fluorophore and the targeting ligand (here, an antibody) through the photocleavable linker, as well as the light-triggered cleavage reaction that releases the fluorophore from the antibody. Fig. 2B displays the library of spectrally distinct fluorophores integrated into the CLEAR probe platform. Comprehensive synthetic protocols and characterization details are described in the SI.To establish the chemical basis of efficient signal erasure, we first examined the intrinsic photochemical reaction pathway of the photocleavable group. A UV-vis spectroscopy study of a representative small-molecule analogue 1d incorporating the identical ONB photocleavable unit used in CLEAR probes revealed well-defined isosbestic points throughout 365 nm light irradiation, indicating a clean conversion between two dominant absorbing species consistent with a single-step bond-cleavage process (Fig. S1, SI). Importantly, identical spectral behaviour was observed in the presence of cell lysate, demonstrating that the photocleavage mechanism is preserved under biologically complex conditions without detectable secondary photoreactions (Fig. S1). Consistent with this observation, MALDI–MS analysis of irradiated CLEAR antibody conjugates revealed mass signatures corresponding to the expected cleaved fluorophore fragment (Fig. S2), further confirming a clean and well-defined photocleavage pathway in the conjugated system. We next evaluated whether the ONB-based photo-cleavable linker affects fluorophore performance due to potential nitrobenzene-induced quenching. Notably, fluorescence lifetime measurements of ONB-conjugated dyes (AZ488, AZ568, and AZ647) showed minimal impact (Fig. S3), indicating that PEG-based spatial separation effectively prevents significant quenching. We then subjected the CLEAR probes to a rigorous labelling–erasing assay targeting an intracellular protein, designed to quantitatively evaluate their erasing efficiency under biologically relevant conditions. Specifically, we immunostained the outer mitochondrial membrane protein TOM20 in U2OS cells using a conventional two-step immunolabeling protocol with CLEAR probes—employing a primary anti-TOM20 antibody followed by a CLEAR secondary antibody conjugate. The resulting TOM20 staining patterns exhibited the characteristic filamentous and reticulated morphology of mitochondria, closely resembling those obtained using conventional fluorophore–antibody conjugates (Fig. 3A). This demonstrated that CLEAR probes preserve the core antibody functionality and antigen accessibility following conjugation with the fluorophore-containing photocleavable tag—a critical requirement for ensuring specificity and reliability in CLEAR-based multiplexed immunostaining workflows. To evaluate the efficiency and kinetics of photocleavage in cells, we selected a defined region of interest and acquired a baseline fluorescence image before light exposure (0 s). The cells were then exposed to handheld 365 nm LED illumination (47.19 mW cm−2) in successive 30 s bursts, with fluorescence images captured from the same region of interest after each burst, until a cumulative irradiation time of 120 s was reached (Fig. 3A and B).
Fluorescence intensities from mitochondrial filaments were tracked over time, revealing that 98.80% of the initial signal from the AZ488-labeled CLEAR probe (CLEAR-488) was eliminated within just 120 s of cumulative light exposure (Fig. 3C). This near-quantitative erasing efficiency establishes a crucial basis for preventing background buildup and maintaining high image quality during extended serial profiling (>15 cycles).25 Importantly, similar performance was observed with other CLEAR probes: the AZ568-labeled conjugate (CLEAR-568) showed a 98.02% reduction in fluorescence intensity under the same conditions (Fig. 3D–F). To further evaluate the versatility and robustness of the CLEAR platform across diverse molecular targets and subcellular compartments, we extended our analysis to include additional proteins, such as tubulin, using the AZ647-labeled CLEAR probe (CLEAR-647, Fig. 3G). Consistent with other probes, CLEAR-647 achieved 98.14% fluorescence clearance within 120 seconds of light exposure (Fig. 3H and I). These findings validate the design of our photocleavable moiety and highlight its effectiveness in enabling highly efficient signal removal across multiple fluorophores within a short time frame under gentle, low-intensity light exposure. Additionally, control experiments with fluorophore-labeled secondary antibodies lacking the ONB photocleavable linker showed minimal signal reduction under identical illumination, confirming that the fluorescence loss in CLEAR probes is attributable to photocleavage rather than photobleaching (Fig. S4, S5). To demonstrate versatility beyond intracellular labeling, CLEAR probes were applied to label the extracellular domain of the EGFR in A431 cells. Immunostaining with CLEAR antibody conjugates showed efficient signal erasure upon 365 nm light exposure, validating the photocleavable design for clearing surface-bound targets (Fig. 3J). To evaluate spatial scalability—critical for high-throughput and tissue-scale imaging—we labeled α-tubulin in U2OS cells with CLEAR probes and irradiated a ∼1 mm2 region using a handheld 365 nm LED (64.20 mW cm−2) for 120 s. Confocal images confirmed uniform and efficient signal clearance across the entire field of view (Fig. 3K), validating the homogeneity of signal removal across large-scale dimensions, which is particularly useful for tissue-scale imaging. Another common concern in iterative multiplexing strategies is the possibility that repeated labeling–imaging–erasing cycles may lead to epitope degradation or compromise antigenicity, thus limiting the number of viable cycles. To address this, cells were exposed to 365 nm LED illumination for 20 sequential cycles of 1 minute each, followed by staining with CLEAR-488 antibodies against α-tubulin. Control cells were labeled in parallel with the same antibodies but without prior 365 nm light exposure. Fluorescence images of the stained microtubules were acquired for both irradiated and non-irradiated samples, and mean fluorescence intensities (MFIs) were compared. As shown in Fig. S6, the MFI—indicative of staining efficiency—remained virtually unchanged between the two conditions, indicating minimal epitope loss and confirming that antigenicity is well maintained even after multiple rounds of 365 nm light illumination and staining. Taken together, these results position the CLEAR probes as a highly efficient, versatile, and scalable solution for cyclic immunofluorescence imaging. Their compatibility with a wide range of fluorophores and protein targets, combined with their efficiency and mild operating conditions, offers a compelling strategy for high-throughput, highly multiplexed biological imaging.
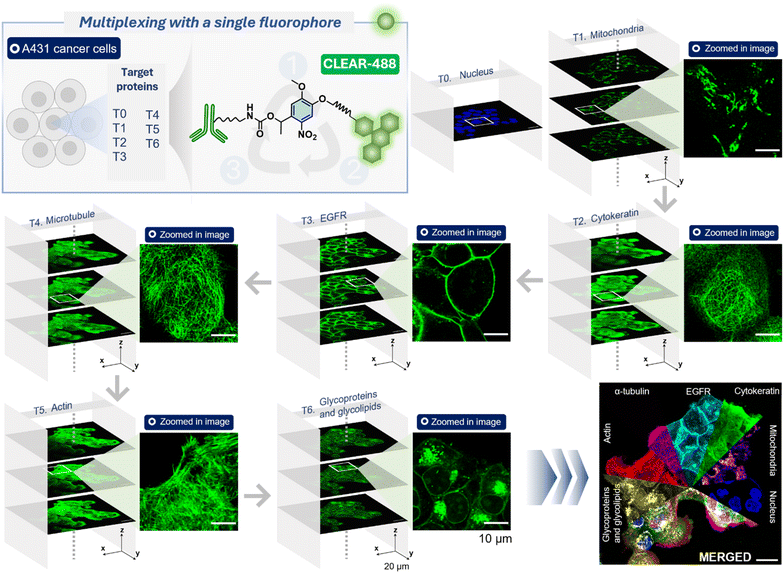
Having established the efficient signal-clearing performance of the CLEAR probes in single-target immunofluorescence, we next assessed their suitability for iterative, multitarget imaging—an essential requirement for high-content and spatially resolved biological studies. Specifically, we aimed to demonstrate that a single fluorophore (or an optical window) could be reused across multiple iterative cycles of labeling, imaging, and signal erasure to enable sequential visualization of multiple targets within the same sample. To evaluate this, we employed A431 epidermoid carcinoma cells and performed five consecutive rounds of cyclic immunofluorescence imaging, targeting six proteins comprising both intracellular and membrane-associated proteins. In the first cycle, the mitochondrial protein TOM20 was immunostained with a CLEAR-AZ488 probe, and a defined region of interest (ROI) was imaged using confocal laser scanning microscopy, with z-stacks acquired across multiple focal planes to capture the three-dimensional distribution of the target (Fig. 4). A magnified view of the ROI confirmed the characteristic distribution of the mitochondrial protein TOM20, confirming the specificity of the CLEAR-AZ488 probes towards its intended targets. After imaging, the fluorescent signal was erased by irradiating the sample with 365 nm LED light for 2 minutes, followed by a mild wash to remove the cleaved fluorophore fragments. In the subsequent cycles, cytokeratin18, EGFR, α-tubulin, and actin filaments were sequentially labelled and imaged using the respective CLEAR-AZ488 probes, with each cycle incorporating a light-induced cleavage step to eliminate the fluorescence signal before proceeding. A final sixth target, the glycoproteins—was visualized using the same detection channel by staining with FITC-conjugated wheat germ agglutinin (WGA-FITC). The nucleus stain served as a reference marker to maintain spatial registration across cycles. Notably, confocal images from each round of staining clearly revealed the characteristic subcellular distribution of each target (Fig. 4), demonstrating that CLEAR probes enable iterative multiplex imaging using the same fluorophore channel while preserving staining specificity across successive imaging cycles. Moreover, analysis of post–light exposure images showed no measurable buildup of residual fluorescence across cycles, indicating that photocleavage products do not accumulate or interfere with subsequent labelling or imaging steps (Fig. S7). After establishing the feasibility of multiplexed imaging using repeated cycles with a single fluorophore, we next aimed to demonstrate the potential of CLEAR probes for high-throughput multiplexing using spectrally distinct dye conjugates in each imaging round (CLEAR-488, CLEAR-568, and CLEAR-647). A unique advantage of the CLEAR platform is that all fluorophores—regardless of their spectral properties—can be cleaved simultaneously using a single, brief exposure to 365 nm light. This capability allowed the imaging of multiple targets per round, significantly accelerating multiplexing throughput by enabling multi-color imaging followed by synchronous fluorophore removal (Fig. 5). Together, these results establish CLEAR probes as a robust, scalable platform for high-throughput, cyclic immunofluorescence imaging. Their ability to enable rapid, repeated use of the same fluorophore channel to image multiple targets—with high specificity and minimal signal spillover between imaging rounds—makes CLEAR probes particularly well-suited for high-content spatial profiling in complex biological systems. Moreover, repeated gentle photocleavage cycles exerted minimal impact on cell morphology, underscoring the compatibility of CLEAR probes with sensitive biological samples. CLEAR conjugates also showed excellent stability during typical storage and under buffer conditions, retaining over 80% of attached fluorophores after approximately three years while preserving target recognition, imaging performance, and rapid signal erasure capability (see Fig. S8). This preserved chemical integrity and functional activity supports the practical and reproducible use of CLEAR probes in cyclic and multiplex imaging applications.
 | ||
| Fig. 5 Accelerated multiplexed imaging using CLEAR probes, enabling multi-target visualization per round through combined multicolor acquisition and synchronous fluorophore erasure. Scale bar: 20 µm. | ||
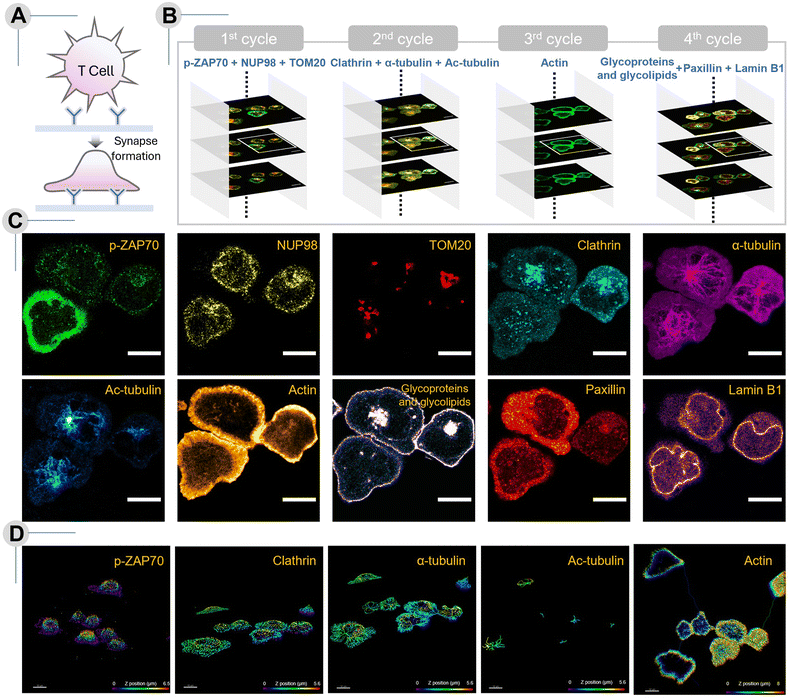
Building on the demonstrated scalability and efficiency of our CLEAR-based multiplexed imaging platform, we extended this approach to interrogate a highly dynamic cellular event in an immunological context. The immune synapse, a transient but highly organized structure formed between T cells and antigen-presenting surfaces, provides an ideal testbed for our high-content, spatially resolved imaging. Complex molecular interactions and rearrangements underlie immune synapse initiation, establishment, and maturation.53 During this process, subcellular localization and temporal dynamics of the signaling and cytoskeletal components together determine the overall signaling state of a single cell in a heterogeneous population. However, visually investigating the relationship between these different parameters requires concurrent imaging of several crucial effectors within that cell—a feat that has not been possible before. T cells, in particular, are highly challenging and delicate for high-content microscopy, and although multiplexing is highly sought after, imaging has typically been limited to just 3–4 markers.54 To overcome this limitation, we applied the CLEAR platform for high-dimensional imaging of ten molecular targets—including both immune synapse–associated and other subcellular proteins—in Jurkat T cells, generating a multiplexed, single-cell–level map of cells undergoing immune synapse formation (Fig. 6). In our system, synapse formation was induced by seeding Jurkat T cells onto coverslips coated with anti-CD3 antibody and His-ICAM, followed by fixation after 5 minutes. Here, the T-cell synaptic interface and closely apposed sub-synaptic zones are roughly up to 300 nm in depth from the coverslip. Thus, signalling molecules and cytoskeletal elements observed “at the synapse” indicate their specific recruitment and positioning at the synaptic interface with functional ramifications on T-cell synapse progression and T-cell activation. Ligation of the agonist antibody with the T-cell receptor (TCR) complex triggers a series of phosphorylation events that drive the overall immune response. One such early event is the recruitment and phosphorylation of adaptor protein ZAP70. The intensity of the phosphorylated-ZAP70 (p-ZAP70) signal near the interface provides a direct readout of proximal TCR signaling.55 Our imaging data revealed variation in p-ZAP70 intensity near the synaptic interface between cells, capturing the cell-intrinsic heterogeneous engagement of proximal TCR signaling among individual cells (see Fig. 6C and 6D—p-ZAP70). Such heterogeneity could also indicate differences in synapse maturity amongst the cells (synapse age ranging between 1–5 min), since Zap70 phosphorylation decays with synapse maturation. Cell 3, as indicated in Fig. S9, showed the highest signal at the interface, while others were lower. This indicated the relative maturity of Cell 1 and 2 synapses compared to Cell 3. To further infer the synapse age and maturation state, we examined cytoskeletal features captured in our dataset. The localization of the microtubule-organizing center (MTOC) reflects the maturity of the synapse, since MTOC repositions to the synapse only in its mature phase. In our data, MTOC, visualized via α-tubulin staining, frequently shifted toward the synaptic interface, as seen in the mature synapses of Cell 1 and 2 (Fig. 6C, 6D—α-tubulin, S13).56 Matching this heterogeneity in synapse age, we found acetylated microtubules enriched at the center of synapses in an age-dependent manner. Higher acetylation corresponds to a more stable microtubule network, and it was enhanced in cells with complete MTOC polarization, suggesting localized cytoskeletal stabilization following synaptic MTOC polarization during TCR engagement (Fig. 6C, 6D—acetylated MTs, S14).57 Other cytoskeletal elements, such as actin, play a critical role in T-cell spreading, creating a scaffold for organizing signaling clusters, and modulation of synapse symmetry, and critically determine the adaptive immune response.58,59 In Jurkat cells, the generation and contraction of the peripheral filamentous actin (f-actin) ring is a characteristic feature of the initial stages of synapse formation. This dynamic structure dictates adhesion strength in immunological synapses by engaging with integrins as well as generating an overall radial symmetry. In our imaging studies, Jurkat synapses fixed at the 5 minute mark consistently displayed prominent f-actin rings; however, we observed notable variability in ring morphology, continuity, and intensity across individual cells. As shown in Fig. S15, Cell 1 had the most distinct periphery with central clearing consistent with synapse maturation, Cell 2 had a slightly contracted ring, and Cell 3 still had actin networks visible centrally. These differences highlight the extent of cell-to-cell heterogeneity in early actin remodeling during immune synapse progression (Fig. 6C and 6D—actin).60 Adhesion complex organization was also captured through paxillin staining, indicating the formation of focal contacts at the cell–substrate interface, with a higher peripheral signal in Cell 1 indicating stable adhesion ring formation (Fig. 6C—Paxillin, S17).57 Synapse maturation, as evidenced by lower p-Zap70 levels, MTOC polarization, and actin distribution, was accompanied by increased central vesicular accumulation in Cell 1 and 2, as visualized by clathrin and WGA labeling (Fig. 6C and D —clathrin and WGA, S12, S16).61 Overlaying these classical markers, our multiplexed imaging revealed the spatial organization of nuclear and mitochondrial components during synapse formation. At this stage of synapse, the nucleus is typically large, occupying a majority of the cell, and juxtaposed to the synaptic interface as observed in the signal from nuclear envelope proteins NUP 98 and Lamin B1 across all cells (6C-NUP8, 6C-Lamin B1, S10, S18). Meanwhile, TOM20-labeled mitochondria showed the beginnings of redistribution toward the interface during synapse maturation. As mitochondria do not polarize to the synapse completely until 10–15 minutes of synaptic contact, these cells, being fixed at 5 minutes, do not have as distinct an accumulation of the TOM20 signal at the interface (Fig. 6C – TOM20, S11).62,63 Together, these findings demonstrate the power of CLEAR-based multiplexed imaging which has a powerful potential to uncover many coordinated subcellular rearrangements across signaling, cytoskeletal, membrane, and organellar processes in the same cell. By providing a systems-level view of spatial coordination during T-cell activation at high spatial resolution, our method offers a powerful tool for dissecting heterogeneity in immune responses at the single-cell level.
Super-resolution microscopy has become a cornerstone of modern biology, enabling the visualization of cellular structures and molecular organization at nanometer resolution. Among various methods, dSTORM is one of the most widely adopted techniques. Alexa Fluor 647 (Alexa647) is widely regarded as the gold-standard fluorophore for dSTORM imaging, owing to its superior photostability, high photon yield per switching event, and low duty cycle, which together enable optimal localization precision and image quality.64 However, the heavy reliance on a single fluorophore imposes a critical bottleneck by severely restricting the multiplexing capacity of dSTORM imaging. To overcome this limitation, we leveraged the rapid and near-quantitative fluorescence-clearing capability of our CLEAR probes, enabling iterative use of the same Alexa647 dye to perform sequential super-resolution imaging of multiple targets within a single sample (Fig. 7). We first immunostained the mitochondrial outer membrane protein TOM20 with a CLEAR-647-conjugated antibody and acquired super-resolved images. Following imaging, the CLEAR-647 signal was efficiently erased by irradiating the sample for two minutes with 365 nm light. We subsequently stained for vimentin, followed by α-tubulin, acquiring high-quality super-resolved images for each filamentous cellular structure in successive cycles. Sequential imaging cycles yielded sharp, distinct images without cross-contamination, validating the compatibility of CLEAR probes with high precision dSTORM workflows (Fig. 7). Overall, this strategy substantially enhances the multiplexing capability of dSTORM by allowing the sequential use of a single optimal fluorophore through controlled photocleavage and relabeling. The successful integration of CLEAR probes into dSTORM workflows highlights their versatility, robustness, and potential for advancing high-throughput, highly multiplexed super-resolution imaging.
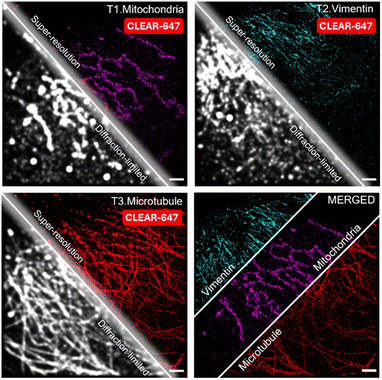 | ||
| Fig. 7 Multiplexed dSTORM super-resolution imaging using CLEAR-647 probes based on the gold-standard dye Alexa Fluor 647. Scale bar: 2 µm. | ||
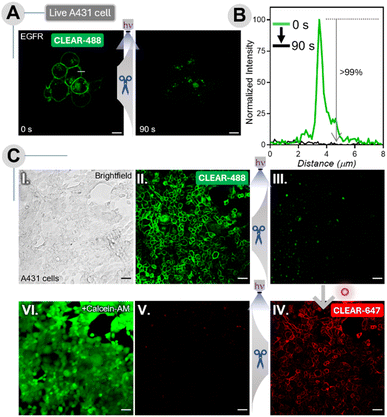
We next evaluated the live-cell compatibility of the CLEAR probes. Unlike chemical cleavage methods that require harsh reagents such as TCEP, DMF, NaOH, or H2O2—conditions that can compromise cell viability—CLEAR probes enable fluorescence erasure using only a mild dose of light, making them potentially more suitable for live-cell applications. To test this, we labeled cultured A431 cells with an EGFR-targeting CLEAR-488-antibody for 10 min, followed by live-cell imaging. Cells were then irradiated with 365 nm light (47.19 mW cm−2) for 30 s, and fluorescence images were captured from a region of interest to assess scission efficiency. This cycle of irradiation and imaging was repeated up to a total of 90 s of cumulative exposure. Notably, we observed that within 90 s, >99% of the membrane-localized EGFR signal was efficiently erased (Fig. 8A and B), demonstrating the high photocleavage efficiency of CLEAR probes under live-cell conditions. Before proceeding to live-cell multiplexing experiments, we evaluated the effect of light exposure under optimized irradiation conditions and found that the applied 365 nm illumination produced minimal phototoxic effects, as supported by Calcein-AM staining, cell viability measurements, and oxidative stress analyses (Fig. S19–S22). The generated photoproducts also showed negligible toxicity (Fig. S20 and S22). Cell viability remained high across increasing irradiation doses (Fig. S21), with a viability of >85% observed even after 540 s of cumulative exposure. Based on these results, six erasing cycles with 90 s of light exposure per cycle (corresponding to seven imaging cycles) were considered tolerable for live-cell imaging without significant deterioration in cellular health. Encouraged by these results, we moved forward to demonstrate live-cell multiplexed imaging capability using CLEAR probes. In this experiment, live A431 cells were first labeled with a CLEAR-488 primary antibody against EGFR and imaged (Fig. 8C-II). The fluorophore was then cleaved by irradiating the cells with 365 nm light for 90 s (Fig. 8C-III). For the second round, cells were incubated with unlabelled cetuximab for 10 minutes, followed by a 10 min incubation with a CLEAR-647 secondary antibody. The EGFR was imaged in the respective channel (Fig. 8C-IV), and the fluorophore was subsequently cleaved by another round of 365 nm light exposure (Fig. 8C–V). Finally, to assess the overall impact of two complete imaging cycles on cell viability, cells were stained with Calcein-AM, a non-fluorescent dye that becomes highly fluorescent upon esterase-mediated hydrolysis in viable cells. Bright cytosolic fluorescence was observed throughout the sample (Fig. 8C-VI), corroborating that the cycling of fluorophores did not adversely affect cell health, which was also supported by the cell-viability assay (Fig. S20). Together, these results demonstrate that CLEAR probes enable efficient, multiplexed live-cell imaging with minimal phototoxicity, highlighting their promise for spatiotemporal dynamic exploration and high-resolution biological studies.
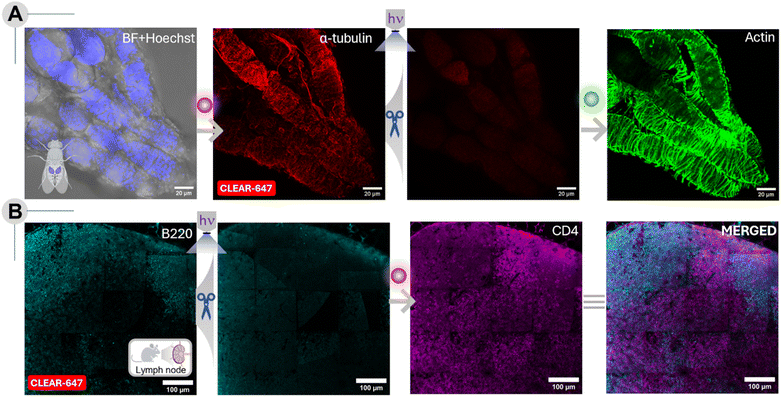
Multiplexed tissue imaging enables spatially resolved visualization of proteins within intact biological architectures, offering critical insights into the functional states of cells, their microenvironmental niches, and the complex molecular interplay underlying both physiological and pathological processes. Motivated by these needs, we evaluated the efficacy of our CLEAR probes in complex tissue environments using two distinct biological models: Drosophila melanogaster ovary tissue and mouse lymph node tissue. We first applied our method to Drosophila melanogaster ovary tissue to assess the ability of CLEAR probes to achieve efficient labeling and erasure in a complex system with multiple developmental stages. Sequential labeling was performed by targeting α-tubulin and actin filaments with CLEAR-647 and CLEAR-488, respectively. The resulting images revealed widespread and well-organized distributions of both cytoskeletal components throughout the tissue (Fig. 9A), highlighting the structural complexity and organization characteristics of ovarian tissue. Importantly, fluorescence erasure after the first labeling cycle was highly efficient, with minimal residual signal detected following photocleavage. This excellent clearing capability ensured that signals from prior rounds did not interfere with subsequent imaging, a critical requirement for accurate multiplexed analysis. Next, we evaluated the performance of CLEAR probes in mouse lymph node sections to test their adaptability to reveal complex mammalian tissue architecture. We sequentially labeled and imaged B220 (a B-cell marker) and CD4 (a T-cell marker) using CLEAR-647 over two imaging cycles, demonstrating the reuse of the same fluorophore through cyclic imaging (Fig. 9B). The staining patterns revealed distinct, stereotypic localization of B220+ and CD4+ cell populations, consistent with previous studies.65 Importantly, photocleavage of the CLEAR-647 fluorophore after the first staining cycle was highly efficient, leaving no detectable residual signal prior to the second labeling round. Together, these findings establish the applicability of CLEAR probes for reliable, consistent, multiplexed imaging across diverse biological and clinically relevant tissue models, extending their use beyond cultured cells into complex and heterogeneous tissue environments.
Conclusions
In summary, we introduce a versatile new platform for immunofluorescence imaging based on conventionally conjugated CLEAR probes that enables scalable and potentially unlimited multiplexed imaging. A defining strength of the CLEAR platform is its seamless integration into existing and widely adopted immunofluorescence workflows, along with broad compatibility across a diverse set of fluorophores, imaging modalities, and sample types. It supports techniques ranging from confocal microscopy for large µm-to mm-scale sample analysis to super-resolution methods for nanoscale visualization, while maintaining high signal fidelity in fixed cells, live cells, and complex tissue environments, making it ideally suited for high-content, spatially resolved proteomic investigations across diverse biological contexts. To realize such a platform, we designed CLEAR probes from the ground up using a highly optimized molecular architecture centered around a photo-cleavable moiety. This design enables one of the fastest and most efficient signal-clearing strategies reported to date, achieving >98% fluorescence removal in under two minutes using a mild, non-toxic light dose. This near-quantitative fluorescence erasure is critical for enabling high-degree multiplexing through serial profiling, as it prevents the accumulation of residual signals across imaging cycles which would otherwise lead to a rapid decline in signal-to-background ratios and severely limit the number of achievable imaging rounds in a multiplexed workflow. Built as a universal and modular platform, CLEAR probes readily accommodate a wide range of commercially available fluorophores and integrate seamlessly into iterative immunostaining protocols, enabling high-throughput, multiplexed imaging of diverse protein targets across a variety of biological samples, including intact and metabolically dynamic tissues. The excellent biocompatibility and mild activation conditions preserve both sample integrity and signal quality across multiple imaging cycles, also making the system highly suitable for live-cell imaging. In addition, its compatibility with super-resolution modalities further broadens its applicability. As a demonstration of the platform's capabilities, we employed CLEAR to spatially map ten distinct proteins and investigate the coordinated remodeling of cytoskeletal structures and organelle distribution during immunological synapse formation. Collectively, the CLEAR platform offers a simple, efficient, and broadly accessible strategy for high-content spatial proteomics, laying the foundation for comprehensive molecular mapping in both basic research and clinical applications.Ethical statement
All animal experiments were performed in accordance with institutional guidelines and were approved (approval No. SA004) by the Institutional Animal Ethics Committee (IAEC) of JNCASR.Author contributions
S. K. and S. S. A. conceived the project and designed the experiments. S. K., A. S., and S. B. designed and performed the synthesis, probe development, and photochemical characterization of the CLEAR probes. S. K., P. K., A. S., and R. V. N. performed the multiplexing and iterative imaging studies. S. M. and S. K. contributed to the immunological synapse studies. S. R. contributed to the tissue staining studies. All authors analyzed the data and interpreted the results. All authors contributed to the writing and editing of the manuscript. S. S. A. supervised the overall study.Conflicts of interest
There are no conflicts to declare.Data availability
All the relevant data of this study are available within the manuscript and its supplementary information (SI). Supplementary information: detailed experimental procedures, CLEAR probe synthesis and characterization, additional imaging data, control experiments, and supplementary figures. See DOI: https://doi.org/10.1039/d5sc08599c.Acknowledgements
This work was supported by a SERB Core Research Grant (CRG/2020/006183) and by the Indian Council of Medical Research (ICMR), Government of India, under Grant No. IIRPSG-2025-01-06141, awarded to S. S. A. R. V. N. gratefully acknowledges the Department of Science and Technology (DST), Government of India, for financial support through the INSPIRE Faculty Fellowship (DST/INSPIRE/04/2023/002548). The authors sincerely thank Prof. Sheeba Vasu (JNCASR) for valuable suggestions, expertise in Drosophila experiments, and critical reading of the manuscript. The authors also acknowledge Prof. Kushagra Bansal (JNCASR) and Prerana M. (JNCASR) for insightful discussions, helpful comments on the manuscript, and assistance with mouse lymph node experiments. Assistance from Anju A. K. with fluorescence lifetime measurements is gratefully acknowledged.References
- S. Jain, L. Pei, J. M. Spraggins, M. Angelo, J. P. Carson, N. Gehlenborg, F. Ginty, J. P. Gonçalves, J. S. Hagood, J. W. Hickey, N. L. Kelleher, L. C. Laurent, S. Lin, Y. Lin, H. Liu, A. Naba, E. S. Nakayasu, W.-J. Qian, A. Radtke, P. Robson, B. R. Stockwell, R. Van de Plas, I. S. Vlachos, M. Zhou, HuB. M. A. P. Consortium, K. Börner and M. P. Snyder, Nat. Cell Biol., 2023, 25, 1089–1100 CrossRef CAS PubMed.
- L. Liu, A. Chen, Y. Li, J. Mulder, H. Heyn and X. Xu, Cell, 2024, 187, 4488–4519 CrossRef CAS PubMed.
- F. Schueder, E. M. Unterauer, M. Ganji and R. Jungmann, Proteomics, 2020, 20, 1900368 CrossRef CAS PubMed.
- S. K. Saka, Y. Wang, J. Y. Kishi, A. Zhu, Y. Zeng, W. Xie, K. Kirli, C. Yapp, M. Cicconet, B. J. Beliveau, S. W. Lapan, S. Yin, M. Lin, E. S. Boyden, P. S. Kaeser, G. Pihan, G. M. Church and P. Yin, Nat. Biotechnol., 2019, 37, 1080–1090 CrossRef CAS PubMed.
- E. M. Unterauer, S. S. Boushehri, K. Jevdokimenko, L. A. Masullo, M. Ganji, S. Sograte-Idrissi, R. Kowalewski, S. Strauss, S. C. M. Reinhardt, A. Perovic, C. Marr, F. Opazo, E. F. Fornasiero and R. Jungmann, Cell, 2024, 187, 1785–1800.e16 CrossRef CAS PubMed.
- J. Ko, J. Oh, M. S. Ahmed, J. C. T. Carlson and R. Weissleder, Angew. Chem., Int. Ed., 2020, 59, 6839–6846 CrossRef CAS PubMed.
- N. R. Sundah, N. R. Y. Ho, G. S. Lim, A. Natalia, X. Ding, Y. Liu, J. E. eSeet, C. W. Chan, T. P. Loh and H. Shao, Nat. Biomed. Eng., 2019, 3, 684–694 CrossRef CAS PubMed.
- J. Eberwine, J. Kim, R. C. Anafi, S. Brem, M. Bucan, S. A. Fisher, M. S. Grady, A. E. Herr, D. Issadore, H. Jeong, H. Kim, D. Lee, S. Rubakhin, J. Y. Sul, J. V. Sweedler, J. A. Wolf, K. S. Zaret and J. Zou, Nat. Methods, 2023, 20, 331–335 CrossRef CAS PubMed.
- M. Angelo, S. C. Bendall, R. Finck, M. B. Hale, C. Hitzman, A. D. Borowsky, R. M. Levenson, J. B. Lowe, S. D. Liu, S. Zhao, Y. Natkunam and G. P. Nolan, Nat. Med., 2014, 20, 436–442 CrossRef CAS PubMed.
- S. C. Bendall, E. F. Simonds, P. Qiu, E. D. Amir, P. O. Krutzik, R. Finck, R. V. Bruggner, R. Melamed, A. Trejo, O. I. Ornatsky, R. S. Balderas, S. K. Plevritis, K. Sachs, D. Pe’er, S. D. Tanner and G. P. Nolan, Science, 2011, 332, 687–696 CrossRef CAS PubMed.
- X. K. Lun, K. Sheng, X. Yu, C. Y. Lam, G. Gowri, M. Serrata, Y. Zhai, H. Su, J. Luan, Y. Kim, D. E. Ingber, H. W. Jackson, M. B. Yaffe and P. Yin, Nat. Biotechnol., 2025, 43, 811–821 CrossRef CAS PubMed.
- A. R. Buchberger, K. DeLaney, J. Johnson and L. Li, Anal. Chem., 2018, 90, 240–265 CrossRef CAS PubMed.
- J. R. Lin, Y. A. Chen, D. Campton, J. Cooper, S. Coy, C. Yapp, J. B. Tefft, E. McCarty, K. L. Ligon, S. J. Rodig, S. Reese, T. George, S. Santagata and P. K. Sorger, Nat. Cancer, 2023, 4, 1036–1052 CrossRef CAS PubMed.
- R. Sasmal, A. Som, P. Kumari, R. V. Nair, S. Show, N. S. Barge, M. Pahwa, N. D. Saha, S. Rao, S. Vasu, R. Agarwal and S. S. Agasti, ACS Cent. Sci., 2024, 10, 1945–1959 CrossRef CAS PubMed.
- L. Schermelleh, A. Ferrand, T. Huser, C. Eggeling, M. Sauer, O. Biehlmaier and G. P. C. Drummen, Nat. Cell Biol., 2019, 21, 72–84 CrossRef CAS PubMed.
- B. Huang, M. Bates and X. Zhuang, Annu. Rev. Biochem., 2009, 78, 993–1016 CrossRef CAS PubMed.
- L. Möckl and W. E. Moerner, J. Am. Chem. Soc., 2020, 142, 17828–17844 CrossRef PubMed.
- J. W. Hickey, E. K. Neumann, A. J. Radtke, J. M. Camarillo, R. T. Beuschel, A. Albanese, E. McDonough, J. Hatler, A. E. Wiblin, J. Fisher, J. Croteau, E. C. Small, A. Sood, R. M. Caprioli, R. M. Angelo, G. P. Nolan, K. Chung, S. M. Hewitt, R. N. Germain, J. M. Spraggins, E. Lundberg, M. P. Snyder, N. L. Kelleher and S. K. Saka, Nat. Methods, 2022, 19, 284–295 CrossRef CAS PubMed.
- J. R. Lin, M. Fallahi-Sichani and P. K. Sorger, Nat. Commun., 2015, 6, 8390 CrossRef CAS PubMed.
- A. J. Ehrenberg, D. O. Morales, A. M. H. Piergies, S. H. Li, J. S. Tejedor, M. Mladinov, J. Mulder and L. T. Grinberg, J. Neurosci. Methods, 2020, 339, 108708 CrossRef CAS PubMed.
- M. M. Bolognesi, M. Manzoni, C. R. Scalia, S. Zannella, F. M. Bosisio, M. Faretta and G. Cattoretti, J. Histochem. Cytochem., 2017, 65, 431–444 CrossRef CAS PubMed.
- A. J. Radtke, E. Kandov, B. Lowekamp, E. Speranza, C. J. Chu, A. Gola, N. Thakur, R. Shih, L. Yao, Z. R. Yaniv, R. T. Beuschel, J. Kabat, J. Croteau, J. Davis, J. M. Hernandez and R. N. Germain, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 33455–33465 CrossRef CAS PubMed.
- D. L. Adams, R. K. Alpaugh, S. Tsai, C. M. Tang and S. Stefansson, Sci. Rep., 2016, 6, 33488 CrossRef CAS PubMed.
- M. J. Gerdes, C. J. Sevinsky, A. Sood, S. Adak, M. O. Bello, A. Bordwell, A. Can, A. Corwin, S. Dinn, R. J. Filkins, D. Hollman, V. Kamath, S. Kaanumalle, K. Kenny, M. Larsen, M. Lazare, Q. Li, C. Lowes, C. C. McCulloch, E. McDonough, M. C. Montalto, Z. Pang, J. Rittscher, A. Santamaria-Pang, B. D. Sarachan, M. L. Seel, A. Seppo, K. Shaikh, Y. Sui, J. Zhang and F. Ginty, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 11982–11987 CrossRef CAS PubMed.
- R. J. Giedt, D. Pathania, J. C. T. Carlson, P. J. McFarland, A. F. del Castillo, D. Juric and R. Weissleder, Nat. Commun., 2018, 9, 4550 Search PubMed.
- S. S. Agasti, Y. Wang, F. Schueder, A. Sukumar, R. Jungmann and P. Yin, Chem. Sci., 2017, 8, 3080–3091 RSC.
- Y. Goltsev, N. Samusik, J. Kennedy-Darling, S. Bhate, M. Hale, G. Vazquez, S. Black and G. P. Nolan, Cell, 2018, 174, 968–981.e15 Search PubMed.
- S.-M. Guo, R. Veneziano, S. Gordonov, L. Li, E. Danielson, K. Perez de Arce, D. Park, A. B. Kulesa, E.-C. Wamhoff, P. C. Blainey, E. S. Boyden, J. R. Cottrell and M. Bathe, Nat. Commun., 2019, 10, 4377 CrossRef PubMed.
- C. Wang, J. Zheng, Z. Xiong, W. Yin, Y. Zhao, H. Wu, S. Liang, L. Zhang, C. Yao, Z. Deng, Y. Liu, Q. Song, G. Zhou and B. Zou, Angew. Chem., Int. Ed., 2025, e202511629 CAS.
- F. Schueder, F. R. Molina, M. Su, Z. Marin, P. Kidd, J. E. Rothman, D. Toomre and J. Bewersdorf, Cell, 2024, 187, 1769–1784 CrossRef CAS PubMed.
- J. Ko, M. Wilkovitsch, J. Oh, R. H. Kohler, E. Bolli, M. J. Pittet, C. Vinegoni, D. B. Sykes, H. Mikula, R. Weissleder and J. C. T. Carlson, Nat. Biotechnol., 2022, 40, 1654–1662 CrossRef CAS PubMed.
- M. Mondal, R. Liao, L. Xiao, T. Eno and J. Guo, Angew. Chem., Int. Ed., 2017, 56, 2636–2639 CrossRef CAS PubMed.
- K. Tomimatsu, T. Fujii, R. Bise, K. Hosoda, Y. Taniguchi, H. Ochiai, H. Ohishi, K. Ando, R. Minami, K. Tanaka, T. Tachibana, S. Mori, A. Harada, K. Maehara, M. Nagasaki, S. Uchida, H. Kimura, M. Narita and Y. Ohkawa, Nat. Commun., 2024, 15, 3657 CrossRef CAS PubMed.
- M. Wilkovitsch, W. Kuba, P. Keppel, B. Sohr, A. Löffler, S. Kronister, A. F. D. Castillo, M. Goldeck, R. Dzijak, M. Rahm, M. Vrabel, D. Svatunek, J. C. T. Carlson and H. Mikula, Angew. Chem., Int. Ed., 2025, 64, e202411707 Search PubMed.
- M. Mondal, R. Liao, C. D. Nazaroff, A. D. Samuel and J. Guo, Chem. Sci., 2018, 9, 2909 RSC.
- Y. Wang, J. B. Woehrstein, N. Donoghue, M. Dai, M. S. Avendaño, R. C. J. Schackmann, J. J. Zoeller, S. S. H. Wang, P. W. Tillberg, D. Park, S. W. Lapan, E. S. Boyden, J. S. Brugge, P. S. Kaeser, G. M. Church, S. S. Agasti, R. Jungmann and P. Yin, Nano Lett., 2017, 17(10), 6131–6139 CrossRef CAS PubMed.
- F. Schueder, M. T. Strauss, D. Hoerl, J. Schnitzbauer, T. Schlichthaerle, S. Strauss, P. Yin, H. Harz, H. Leonhardt and R. Jungmann, Angew. Chem., Int. Ed., 2017, 56, 4052–4055 CrossRef CAS PubMed.
- W. Zhang, H. Jiang, L. Han, J. Liu, J. Wang, F. He and L. Tian, Nat. Commun., 2025, 16, 2300 CrossRef CAS PubMed.
- W. Zhou, Y. Han, B. J. Beliveau and X. Gao, Adv. Mater., 2020, 32, 1908410 CrossRef CAS PubMed.
- X. Liu, D. Mao, Y. Song, L. Zhu, A. N. Isak, C. Lu, G. Deng, F. Chen, F. Sun, Y. Yang, X. Zhu and W. Tan, Sci. Adv., 2022, 8, eabk0133 CrossRef CAS PubMed.
- H. J. Geertsema, G. Aimola, V. Fabricius, J. P. Fuerste, B. B. Kaufer and H. Ewers, Nat. Biotechnol., 2021, 39, 551–554 CrossRef CAS PubMed.
- V. Lehot, I. Kuhn, M. Nothisen, S. Erb, S. Kolodych, S. Cianférani, G. Chaubet and A. Wagner, Sci. Rep., 2021, 11, 5881 CrossRef CAS PubMed.
- J. Park, J. Hahm, J. Yim, H. Lee, H. M. Hwang, S. Lee, J. Y. Park, A. Velladurai, J. K. Gangasani, H. Cho, H. Park, M. Lee, J. Lee, H. Eom, W. J. Song, S. Lee, E. kim and J. Park, ACS Cent. Sci., 2025, 11, 878–889 CrossRef CAS PubMed.
- W. Schubert, B. Bonnekoh, A. J. Pommer, L. Philipsen, R. Böckelmann, Y. Malykh, H. Gollnick, M. Friedenberger, M. Bode and A. W. M. Dress, Nat. Biotechnol., 2006, 24, 1270–1278 CrossRef CAS PubMed.
- S. Wäldchen, J. Lehmann, T. Klein, S. van de Linde and M. Sauer, Sci. Rep., 2015, 5, 15348 CrossRef PubMed.
- C. A. Juan, J. M. Pérez de la Lastra, F. J. Plou and E. Pérez-Lebeña, Int. J. Mol. Sci., 2021, 22, 4642 CrossRef CAS PubMed.
- E. A. Halabi and R. Weissleder, J. Am. Chem. Soc., 2023, 145, 8455–8463 CrossRef CAS PubMed.
- N. Mougios, E. R. Cotroneo, N. Imse, J. Setzke, S. O. Rizzoli, N. A. Simeth, R. Tsukanov and F. Opazo, Nat. Commun., 2024, 15, 8771 Search PubMed.
- S. S. Agasti, M. Liong, V. M. Peterson, H. Lee and R. Weissleder, J. Am. Chem. Soc., 2012, 134, 18499–18502 CrossRef CAS PubMed.
- P. Klán, T. Šolomek, C. G. Bochet, A. Blanc, R. Givens, M. Rubina, V. Popik, A. Kostikov and J. Wirz, Chem. Rev., 2013, 113, 119–191 CrossRef PubMed.
- A. Patchornik, B. Amit and R. B. Woodward, J. Am. Chem. Soc., 1970, 92, 6333–6335 CrossRef CAS.
- C. P. Holmes, J. Org. Chem., 1997, 62, 2370–2380 Search PubMed.
- A. Grakoui, S. K. Bromley, C. Sumen, M. M. Davis, A. S. Shaw, P. M. Allen and M. L. Dustin, Science, 1999, 285, 221–227 CrossRef CAS PubMed.
- J. B. Huppa and M. M. Davis, Nat. Rev. Immunol., 2003, 3, 973–983 CrossRef CAS PubMed.
- J. Yi, L. Balagopalan, T. Nguyen, K. M. McIntire and L. E. Samelson, Nat. Commun., 2019, 10, 277 CrossRef PubMed.
- J. Yi, X. Wu, A. H. Chung, J. K. Chen, T. M. Kapoor and J. A. Hammer, Centrosome repositioning in T cells is biphasic and driven by microtubule end-on capture-shrinkage, J. Cell Biol., 2013, 202, 779–792 CrossRef CAS PubMed.
- J. M. Serrador, J. R. Cabrero, D. Sancho, M. Mittelbrunn, A. Urzainqui and F. Sánchez-Madrid, Immunity, 2004, 20, 417–428 CrossRef CAS PubMed.
- J. A. Hammer, J. C. Wang, M. Saeed and A. T. Pedrosa, Annu. Rev. Immunol., 2019, 37, 201–224 CrossRef CAS PubMed.
- L. Dupré, K. Boztug and L. Pfajfer, Front. Cell Dev. Biol., 2021, 9, 665519 CrossRef PubMed.
- A. Babich, S. Li, R. S. O'Connor, M. C. Milone, B. D. Freedman and J. K. Burkhardt, J. Cell Biol., 2012, 197, 775–787 CrossRef CAS PubMed.
- A. Kvalvaag, S. Valvo, P. F. Céspedes, D. G. Saliba, E. Kurz, K. Korobchevskaya and M. L. Dustin, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2211368120 CrossRef CAS PubMed.
- C. Ledderose, Y. Bao, M. Lidicky, J. Zipperle, L. Li, K. Strasser, N. I. Shapiro and W. G. Junger, J. Biol. Chem., 2014, 289, 25936–25945 CrossRef CAS PubMed.
- A. Quintana, C. Schwindling, A. S. Wenning, U. Becherer, J. Rettig, E. C. Schwarz and M. Hoth, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 14418–14423 CrossRef CAS PubMed.
- G. T. Dempsey, J. C. Vaughan, K. H. Chen, M. Bates and X. Zhuang, Nat. Methods, 2011, 8, 1027–1036 CrossRef CAS PubMed.
- G. Gasteiger, M. Ataide and W. Kastenmüller, Immunol. Rev., 2016, 271, 200 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |