Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Going full circle: dynamic covalent enzyme immobilisation via visually trackable boronate esters
Glenn
Bojanov
 ,
Juliette
Swit
and
Francesca
Paradisi
,
Juliette
Swit
and
Francesca
Paradisi
 *
*
Department of Chemistry, Biochemistry and Pharmaceutical Sciences, University of Bern, Freiestrasse 3, 3012 Bern, Switzerland. E-mail: francesca.paradisi@unibe.ch
First published on 3rd March 2026
Abstract
Enzyme immobilisation on solid supports enables biocatalyst recycling but generates significant waste due to single-use resins that are discarded when enzyme activity declines. Here we report a reversible immobilisation strategy based on boronate ester formation between alizarin-functionalised enzymes and boronic acid-modified supports. Alizarin-methyliminodiacetic acid (alizarin-IDA) serves dual roles as both a pH-responsive binding handle and a visual reporter, enabling real-time colourimetric tracking of enzyme loading (red solution → orange resin), immobilisation completeness, and pH-triggered release. A universal labelling protocol was established and successfully applied to four structurally diverse enzymes retaining 77–95% of native activity. All alizarin-labelled enzymes achieved >90% immobilisation yield on different supports, were extensively reusable, and could be removed by acidic treatment with full regeneration of the supports. The load–use–cleave sequence was repeated five times without loss of binding capacity, enabling more than 50 catalytic cycles per support across multiple enzyme lifecycles (5 regeneration cycles × 10+ reactions each) with identical performance.
Introduction
Biocatalysis delivers high chemo-, regio-, and stereoselectivity under mild conditions and is widely deployed across fine chemicals, pharmaceuticals, food, and fuels.1–3 In practice, these processes often depend on immobilised enzymes for recovery and process integration, yet implementation remains materially linear: carriers, which constitute ∼90–99% of the immobilised catalyst mass, are discarded when the enzyme deactivates.4–6 Lipase B from Candida antarctica (CALB) has been one of the first immobilised lipases via non-covalent adsorption on an acrylic polymeric resin. Commercialised as Novozym 435 by Novonesis, it is likely the most used biocatalyst in academia and industry.7 To prevent enzyme leaching and maximise stability, irreversible covalent bonds are favoured in industry,8–10 this however creates a dilemma: the same chemistry that enables robust catalysis prevents carrier regeneration. From a circular economy perspective, extending the lifetime of the support, not only of the enzyme, is a direct but underused lever to reduce cost and environmental footprint.11–13 The rationale for immobilisation over the use of soluble enzymes has been extensively discussed in terms of ease of separation (downstream processing), operational stability, and reactor integration.14,15 Aside from non-covalent chemistries which easily enable enzyme desorption, dynamic covalent chemistries are less explored in this field but are well established in bioconjugation for drug-delivery where the payload must be released under controlled but mild conditions.16,17 Imine, oxime, and hydrazone chemistries enable reversible bioconjugation, but their hydrolytic lability under typical reaction conditions would not make them suitable for enzyme immobilisation in biocatalytic processes.18,19 Disulfide exchange has demonstrated promise for reversible enzyme attachment, but introduces redox sensitivity that complicates processes involving reducing agents or oxidizing substrates.20–22 Boronate-ester formation between boronic acids and diols offers a compelling alternative. The reaction proceeds rapidly and reversibly in water: catechols exhibit high affinity at pH 7–9 and are released upon acidification or in the presence of competing polyols.23–26Boronate-affinity chromatography leverages this principle for the purification and separation of glycoproteins bearing native cis-diol carbohydrates.27–29 Several studies have applied this chemistry to protein immobilisation for sensing, but remain limited to naturally glycosylated proteins.30,31 One study broadened the scope beyond glycoproteins by successfully immobilising penicillin G acylase from E. coli, a non-glycosylated enzyme, on phenylboronate-activated supports.32 Multipoint attachment yielded stabilisation in organic solvents, and reversible desorption with mannitol was demonstrated, though support regeneration for repeated biocatalytic use was not explored. Beyond protein applications, the pH-responsive reversibility of boronate esters has been well-established in drug delivery systems,33–35 demonstrating pH-triggered release under mild conditions.
Despite these precedents, no method exists for biocatalysis that: (i) installs boronate-reactive handles on any enzyme regardless of native glycosylation, (ii) enables visual tracking of immobilisation and release, and (iii) permits true carrier regeneration across multiple enzyme lifecycles. Here we combine function and readout in a single molecular handle to enable cradle-to-cradle enzyme immobilisation. Phenylboronic-acid groups are installed on the carrier (chemically robust, batch-prepared, storable) and enzymes are labelled through surface lysine amino acids with alizarin-3-methyliminodiacetic acid (alizarin-IDA), which supplies both a catechol for boronate-ester formation and a vivid, pH-sensitive chromophore.36–39 The resulting red (free) → orange (bound) → yellow (released) colour sequence provides direct visual verification of each process step.24,36 This approach aims to establish a general immobilisation strategy compatible with diverse enzymes classes, standard support materials, and packed-bed operation, offering a practical route towards support regeneration in industrial biocatalysis.40–44
Results and discussion
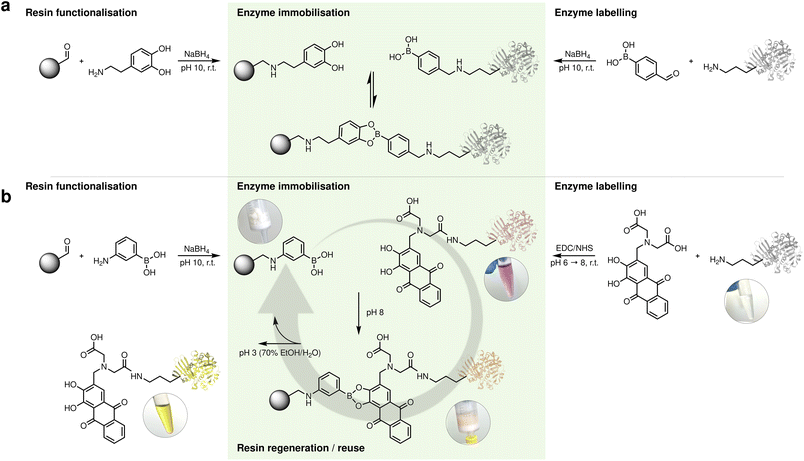
To evaluate our immobilisation strategies, we selected a panel of five structurally diverse enzymes representing different fold families and oligomeric states: Mycobacterium smegmatis acyltransferase (MsAcT, octamer), Candida boidinii formate dehydrogenase (CbFDH, dimer), Streptomyces pristinaespiralisL-lysine cyclodeaminase (SpLCD, tetramer), Thermomyces stellatus (R)-amine transaminase (TsRTA, dimer), and commercial porcine trypsin (monomer). Our initial screening explored the derivatisation of enzymes surface lysines with boronic acid in order to introduce the acidic binding handle while a set of resins were modified with dopamine to provide the catechol moiety for the reversible covalent interaction (Fig. 1, case a). Two structurally distinct enzymes were modified with 4-formylphenylboronic acid (FPBA): MsAcT, a benchmark transesterification catalyst, and CbFDH, essential for NAD+ regeneration. The number of exposed surface lysines on each enzyme was estimated from the available crystal structures (28 in MsAcT and 38 in CbFDH). FPBA was then reacted with the enzymes via reductive amination in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio with respect to the total surface lysines. MsAcT maintained native–level activity (229 U mg−1), while CbFDH exhibited a modest increase (3.01 vs. 2.68 U mg−1). In parallel, two different commercially available resins were functionalised with dopamine to introduce the catechol binding partner. Agarose resin required epichlorohydrin activation followed by acid ring-opening and periodate oxidation to generate aldehydes as previously described.47 EP400 epoxy resin was directly converted via acid ring-opening and periodate oxidation.48 Dopamine (5-fold molar excess) was then conjugated to both glyoxyl resins through reductive amination, introducing the catechol binding partner.
1 molar ratio with respect to the total surface lysines. MsAcT maintained native–level activity (229 U mg−1), while CbFDH exhibited a modest increase (3.01 vs. 2.68 U mg−1). In parallel, two different commercially available resins were functionalised with dopamine to introduce the catechol binding partner. Agarose resin required epichlorohydrin activation followed by acid ring-opening and periodate oxidation to generate aldehydes as previously described.47 EP400 epoxy resin was directly converted via acid ring-opening and periodate oxidation.48 Dopamine (5-fold molar excess) was then conjugated to both glyoxyl resins through reductive amination, introducing the catechol binding partner.
 | ||
| Fig. 1 Reaction scheme reversible boronic ester formation and cleavage. (Case a) 1 = enzyme and 2 = resin. (Case b) 1 = resin and 2 = enzyme. | ||
An overview of the setup is shown in Fig. 2a. Simple incubation of the FPBA-modified enzymes with the dopamine-functionalised resins for 2 hours showed high immobilisation yields (>99%) on both agarose and methacrylate support. However, the recovered activity (RA) was strongly influenced by the FPBA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) lysine ratio used during enzyme functionalisation. MsAcT retained 11–14% activity on methacrylate and up to 38% on agarose, substantially lower than the 73% reported for traditional multipoint covalent immobilisation on agarose.49 Reducing the FPBA
lysine ratio used during enzyme functionalisation. MsAcT retained 11–14% activity on methacrylate and up to 38% on agarose, substantially lower than the 73% reported for traditional multipoint covalent immobilisation on agarose.49 Reducing the FPBA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) lysine ratio to 0.7
lysine ratio to 0.7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 improved activity retention showed a similar trend, with RA improving from 30 to 44% on methacrylate as the ratio decreased from 0.5
1 improved activity retention showed a similar trend, with RA improving from 30 to 44% on methacrylate as the ratio decreased from 0.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 0.2
1 to 0.2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. These results demonstrated a clear correlation between labelling density and activity retention, suggesting that excessive surface modification disrupts the native enzyme structure, while moderate functionalisation preserves catalytic function.
1. These results demonstrated a clear correlation between labelling density and activity retention, suggesting that excessive surface modification disrupts the native enzyme structure, while moderate functionalisation preserves catalytic function.
Despite the success of our initial protocols, critical limitations emerged with the dopamine-based system. Progressive oxidation of dopamine residues caused resin darkening and compromised binding efficiency, which is particularly problematic for batch-prepared resins requiring long-term storage (Fig. S1). Furthermore, enzyme cleavage proved difficult to monitor: extensive screening of conditions (pH 2.5–4, competing diols, Ca2+) yielded no detectable protein in supernatants by Bradford assay, possibly due to protein precipitation under desorption conditions. While through SDS-PAGE analysis of urea-treated resins, proteins cleavage could be at least confirmed, the quantification remained challenging (Fig. S2).
In an effort to simplify the process and enhance chemical stability, we inverted the location of the chemical handles: the stable boronic acid moiety was to be placed on the resin (suitable for batch functionalisation and long-term storage) while the oxidation-prone catechol would be used to derivatise the enzyme, where it would be freshly installed on demand (Fig. 1, case b). This inversion would not only eliminate resin degradation over time but also avoid catechol instability during cleavage, ensuring reliable regeneration and consistent performance across cycles.
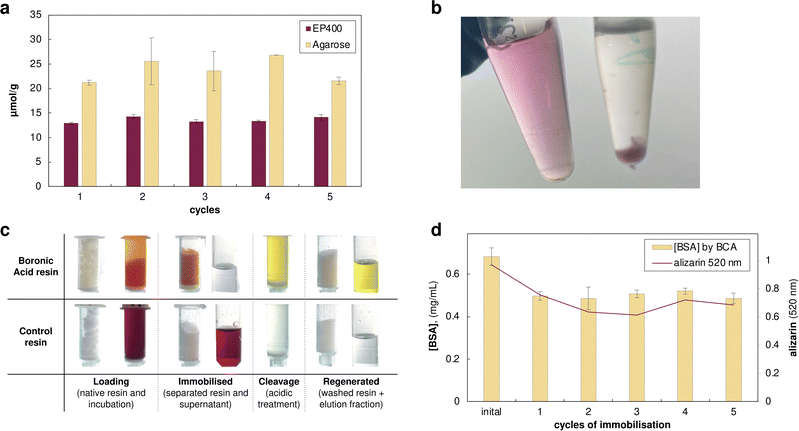
Following the design inversion, we first prepared boronic acid-functionalised EP400 methacrylate and agarose supports. To validate the functionalisation, we used alizarin (1,2-dihydroxyanthraquinone), an established colourimetric probe for boronate ester formation. Quantitative binding experiments revealed functional capacities of 13.6 ± 0.6 µmol alizarin per gram for EP400 resins and 23.7 ± 2.0 µmol g−1 for agarose supports, corresponding to the boronic acid groups on the support available for enzyme immobilisation (Fig. 3a). Control experiments with non-functionalised resins showed negligible alizarin binding (<0.5 µmol g−1), confirming the specificity of the boronate–alizarin interaction.
To our delight, the alizarin quantification experiments offered a striking colour variation. The deep red alizarin solution turned bright orange immediately upon contact with boronic acid resins, with the resin beads developing intense colouration within seconds, while the surrounding solution rapidly lost its colour (Video S1). After separation, the resins retained the orange colour while supernatants remained colourless. Control native resins showed no colour change, with any slight red hue quickly removed after washing with water. Acidic treatment (70% ethanol/30% water with HCl, pH ∼3) released alizarin quantitatively as a yellow solution, returning the boronic functionalised resin to its original white appearance. The resins maintained full binding capacity over five sequential quantification cycles, demonstrating the robustness of the boronic acid functionalisation (Fig. 3a).
Crucially, in this experiment alizarin served as a small-molecule mimic of the dopamine-functionalised enzymes, demonstrating that catechol–boronate ester formation and cleavage were indeed rapid and efficient, even though no enzyme was present in these initial validation experiments. This distinct colour sequence—red (free) → orange (bound) → yellow (acidic released)—provided unprecedented visual tracking of each process step without analytical instrumentation. This clear visual confirmation of complete catechol release demonstrated effective cleavage, resolving the detection challenges encountered with the dopamine system, where low protein recovery had complicated assessment. This observation suggested that alizarin-based molecules could serve dual roles as both binding handles and integrated visual indicators for process monitoring well beyond their use as a diagnostic probe. Its chemical stability compared to dopamine, combined with the visual clarity demonstrated in our validation experiments, prompted us to explore alizarin derivatives as the enzyme-labelling moiety itself. We identified alizarin-3-methyliminodiacetic acid (alizarin-IDA), which retains the catechol functionality while bearing a carboxylic acid handle suitable for EDC/NHS coupling to surface lysines (Fig. 2b). To evaluate this concept, we first tested alizarin-IDA conjugation to bovine serum albumin (BSA) as a model protein. Following EDC/NHS activation and coupling at pH 8, the reaction mixture was purified using PD-10 desalting columns to remove excess unreacted dye. The eluted BSA fraction displayed a distinct pink-red colouration, again a striking visual change from the typical colourless protein solution. UV-vis spectroscopy showed an absorption maximum at 520 nm, demonstrating alizarin conjugation. To confirm covalent nature of this attachment, we boiled the labelled BSA solution. Upon thermal denaturation, the protein precipitated, leaving a colourless supernatant while the protein pellet remained vividly red (Fig. 3b). The reversibility and visual tracking capabilities were systematically evaluated through five complete immobilisation–cleavage cycles (Fig. 3d). When alizarin-labelled BSA was incubated with one of the boronic acid-functionalised resin (agarose was selected for its higher binding capacity) at typical enzyme loading conditions (5 mg protein per g resin), we observed immediate and quantitative binding: the resin turned bright orange while the supernatant became colourless, with both Pierce™ BCA Protein Assay (Thermo Scientific, Cat. No. 23225) and 520 nm absorbance confirming nearly complete protein immobilisation. Control resins showed little protein binding, verifying the specificity of the boronate–catechol interaction. The covalent boronate ester formation was confirmed through standard thermal challenge experiments. While control experiments with unlabelled BSA showed complete protein release upon boiling, alizarin-BSA remained bound to the resin, which retained its orange colour. Both the flow-through collected after initial loading and the supernatant recovered after boiling the loaded resin showed negligible protein content (<5% of loaded amount) and minimal alizarin absorbance, demonstrating stable attachment under harsh conditions. Only upon acidic treatment did the protein release occur, accompanied by the expected colour transition, with the orange resin turning to white and the elution fraction yellow. Complete protein removal was confirmed when both BCA measurement and alizarin absorbance at 520 nm returned to baseline after washing with 3 resin volumes (fractions were pooled and neutralised before measurement). Subsequent neutralisation and reloading with fresh alizarin-BSA restored the orange resin colouration, confirming the reversibility and reusability of the system for at least 5 cycles (Fig. 3d).
Having established alizarin-IDA as a functional enzyme label, we sought to develop a standard functionalisation protocol applicable across diverse enzyme classes available either in our laboratory or commercially. Initial optimisation studies revealed that reaction parameters (pH, time, temperature, and alizarin-to-lysine ratios) could be fine-tuned for individual enzymes to maximize both labelling efficiency and activity retention. However, the practical implementation of this technology demanded a simpler approach: a single, robust protocol that would work sufficiently well across a broad spectrum of enzymes without requiring case-by-case optimisation.
The challenge lies in the inherent structural diversity of enzymes. Our analysis of several structurally distinct enzymes using both SASA calculations and CapiPy-assisted surface mapping (a software tool designed to rationalise immobilisation strategies) revealed dramatic variations in lysine accessibility.50 As shown in Table 1 the number of surface-exposed lysines (SASA ≥ 80 Å2) ranged from just 2 in SpLCD to 38 in CbFDH. Moreover, the reactivity of these lysines, which is influenced by local microenvironments, pKa variations, and dynamic protein motions, proved difficult to predict in silico. This challenge is well-recognised in enzyme immobilisation, where case-by-case empirical optimisation typically remains necessary. We therefore sought to establish a general protocol that could accommodate such structural diversity and serve as a practical starting point for further optimisation. Through experimental optimisation, we established that effective labelling occurred within a range of 1–10 alizarin molecules per reactive surface lysine. To ensure broad applicability, we designed a protocol using a 100-fold molar excess of alizarin-IDA relative to total enzyme concentration (equivalent to a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 enzyme
100 enzyme![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) dye ratio and at least 6 times the amount of lysines). This substantial excess ensures that even enzymes with numerous surface lysines achieve adequate labelling within the optimal range, while the mild conditions preserve enzymatic activity. We validated this strategy across a panel of structurally diverse enzymes: MsAcT (octameric acyltransferase), CbFDH (dimeric formate dehydrogenase), SpLCD (tetrameric cyclodeaminase), TsRTA (dimeric transaminase), and commercial trypsin from porcine pancreas (lyophilised powder). These enzymes span different fold families and oligomeric states (monomer to octamer), ranging from 25 to 187 kDa in their assembled forms, representing a broad cross-section of industrial biocatalysts. All enzymes displayed the characteristic pink-red colouration upon labelling, with absorption maxima at 520 nm confirming successful alizarin conjugation. Quantification of the Degree of Labelling (DoL) revealed modification levels ranging from ∼5 alizarin moieties for TsRTA and trypsin to ∼38 for CbFDH, in line with the abundance of surface-accessible lysines (Table 1). These values rationalise the observed differences in enzyme performance: while MsAcT proved remarkably robust despite ∼15 modifications, the saturation of available lysines in CbFDH (∼38 labels) likely restricted conformational flexibility, contributing to its lower recovered activity (43%, Table 2). Conversely, the low activity of trypsin is likely attributed to autolysis during the process rather than specific labelling interference. Critically, enzymatic activity was preserved across the panel. With the exception of trypsin, the enzymes retained 77–95% of their native activity.
dye ratio and at least 6 times the amount of lysines). This substantial excess ensures that even enzymes with numerous surface lysines achieve adequate labelling within the optimal range, while the mild conditions preserve enzymatic activity. We validated this strategy across a panel of structurally diverse enzymes: MsAcT (octameric acyltransferase), CbFDH (dimeric formate dehydrogenase), SpLCD (tetrameric cyclodeaminase), TsRTA (dimeric transaminase), and commercial trypsin from porcine pancreas (lyophilised powder). These enzymes span different fold families and oligomeric states (monomer to octamer), ranging from 25 to 187 kDa in their assembled forms, representing a broad cross-section of industrial biocatalysts. All enzymes displayed the characteristic pink-red colouration upon labelling, with absorption maxima at 520 nm confirming successful alizarin conjugation. Quantification of the Degree of Labelling (DoL) revealed modification levels ranging from ∼5 alizarin moieties for TsRTA and trypsin to ∼38 for CbFDH, in line with the abundance of surface-accessible lysines (Table 1). These values rationalise the observed differences in enzyme performance: while MsAcT proved remarkably robust despite ∼15 modifications, the saturation of available lysines in CbFDH (∼38 labels) likely restricted conformational flexibility, contributing to its lower recovered activity (43%, Table 2). Conversely, the low activity of trypsin is likely attributed to autolysis during the process rather than specific labelling interference. Critically, enzymatic activity was preserved across the panel. With the exception of trypsin, the enzymes retained 77–95% of their native activity.
| Enzyme | Quart. structure | MW (kDa) | Exposed Lys (calc.) | Reaction ratio (dye: Lys) | [Enzyme], mg mL−1 | Experimental DoL (mol mol−1) |
|---|---|---|---|---|---|---|
| SpLCD | Tetramer | 149.5 | 2 | 50 | 2.0 | 6.9 |
| CbFDH | Dimer | 161.9 | 38 | 6 | 0.1 | 38.4 |
| TsRTA | Dimer | 82.9 | 24 | 8 | 4.0 | 4.8 |
| MsAcT | Octamer | 187.6 | 28 | 28 | 1.6 | 15.3 |
| Trypsin | Monomer | 25.0 | 6 | 16 | 1.2 | 5.1 |
| Enzyme | Post-labelling | Immobilisation | Post-immobilisation | ||
|---|---|---|---|---|---|
| Activity | Resin | Loading (mg g−1) | IY | RA | |
| a Activity is intended as the retained after alizarin-IDA labelling and purification, relative to native enzyme. RA: recovered activity of immobilised enzyme relative to native free enzyme. b SpLCD activity could not be accurately quantified due to interference between alizarin and the FMOC derivatisation required for product detection by HPLC assay; conversion was clearly observed but not accurately quantifiable. | |||||
| SpLCD | Tracesb | Agarose | 5 | 99% | n.d. |
| CbFDH | 77% | Agarose | 5 | 92% | 43% |
| TsRTA | 92% | Agarose | 5 | 91% | 72% |
| EP400 | 10 | 93% | 48% | ||
| MsAcT | 95% | Agarose | 5 | 94% | 93% |
| Trypsin | 57% | Agarose | 5 | 78% | 22% |
With our functionalised enzyme panel in hand, we evaluated their immobilisation performance on the boronic acid-modified agarose resin at a typical loading of 5 mg enzyme per g support. All enzymes showed rapid binding behaviour similar to BSA, achieving >90% immobilisation efficiency within 2 h as determined by both colourimetric (at 520 nm) and Bradford analysis of residual supernatants (Table 2). Control experiments with unlabelled enzymes showed minimal (<10%) non-specific adsorption, typical for agarose supports.51 The immobilised enzymes retained catalytic activity across the panel, with RAs ranging from 22% (trypsin) to 93% (MsAcT). Notably, MsAcT and TsRTA showed particularly high activity retention, making them suitable candidates for further characterisation. For TsRTA, we extended the boronate ester immobilisation to EP400 methacrylate resin and compared it with the previously optimised covalent epoxy immobilisation performed at higher enzyme loadings (10 mg g−1) on the same support.45,46 When compared with the optimised conditions, the boronate ester immobilisation on EP400 achieved 93% IY with 48% RA (12.9 ± 2.2 U g−1), closely reproducing the performance of the epoxy methacrylate reference (>99% IY and 53% RA 13.7 ± 1.8 U g−1). This demonstrates that the reversible boronate ester linkage delivers comparable catalytic performance to irreversible covalent attachment, while enabling support regeneration.
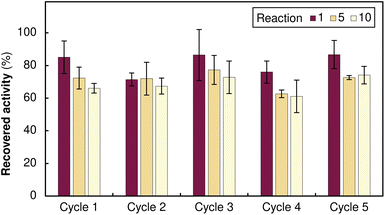
To gain further insight into the binding mechanism, we monitored the immobilisation kinetics using TsRTA as a model system. The process proved extremely rapid, with ∼50% of the protein immobilised within the first minute and >75% binding achieved within 15 minutes (Fig. S6). This rapid binding confirms that boronate ester formation is kinetically efficient, minimising the time required for resin loading. Eventually, to assess the full potential of our system, we performed multiple loading, cleaving, and reloading experiments using MsAcT as a representative biocatalyst. Each immobilisation cycle consisted of: (i) enzyme loading, (ii) 10 repeated reaction cycles, and (iii) enzyme removal and resin regeneration. The immobilised MsAcT maintained stable activity throughout 10 consecutive catalytic cycles, with no significant loss. Following this, the enzyme was removed by acidic wash, the orange-to-white transition confirming complete stripping, and fresh alizarin-functionalised MsAcT was loaded onto the regenerated resin. This complete sequence was systematically repeated 5 times, with identical performance of both the enzyme and the resin. The consistent activity across all immobilisation cycles confirms true cradle-to-cradle operation: each resin support enables 50+ reaction cycles (5 enzyme loadings × 10+ uses each) before any potential disposal (Fig. 4).
Finally, we assessed the pH stability of the boronate–catechol linkage under operationally relevant conditions (Fig. S4A). Alizarin-loaded boronic acid agarose resin was incubated between pH 3–10 for 2 h to assess spontaneous hydrolysis. The complex remained stable across a wide pH range (3–8), with only moderate release at pH 9–10 (6–17%) (Fig. S4A). This alkaline instability likely arises from hydroxide-mediated mechanisms, distinct from the protonation-driven dissociation observed at low pH; however, we selected acidic cleavage for regeneration as it offered superior speed and completeness. Beyond pH, we evaluated stability against common biocatalytic additives (Fig. S4B). Incubation with high ionic strength buffers (300 mM NaCl), reducing agents (1 mM DTT), competing polyols (10% glycerol), or substrates (formate) resulted in negligible protein leaching (<6%). These results confirm that the boronate ester bond is sufficiently stable for biocatalytic operation across the pH range (6–8), where most enzymes maintain high activity,52 while still permitting controlled cleavage at very low pH.
Conclusions
Enzyme immobilisation has long faced a fundamental dilemma: the irreversible covalent bonds that enable robust industrial biocatalysis simultaneously prevent carrier regeneration, generating substantial waste when activity declines. Here we address this through boronate ester chemistry combined with alizarin-IDA, a bifunctional molecule that serves as both a pH-responsive binding handle and as an integrated chromophoric reporter. By incorporating alizarin-IDA directly on the enzyme surface we established a system that enables: (1) direct confirmation of successful enzyme labelling via the distinct red colour of functionalised enzyme solutions; (2) visual tracking of IY during resin loading (red → orange shift; transparent supernatant); (3) immediate, reagent-free confirmation of pH-triggered release during cleavage (orange → yellow elution; resin turns colourless): (4) operational transparency across multiple loading-use-cleave cycles without the need for analytical instrumentation.We established a universal labelling protocol effective across five structurally diverse enzymes (25–187 kDa, monomer to octamer). All alizarin-functionalised enzymes retained 77–95% native activity and achieved >90% immobilisation yields with typical enzyme loadings. Commercial trypsin and SpLCD were the weakest performers which is possibly linked to the limited lysines available for reactivity. Critically, for TsRTA we further showed, that the chemistry extends to methacrylate supports and that the performance matched traditional covalent epoxy immobilisation at higher loadings (10 mg g−1). TsRTA retained 48% activity via boronate esters versus 53% via irreversible epoxy chemistry, whilst enabling complete support regeneration.
Multiple cycling with MsAcT could be carried out for at least five complete load–use–cleave sequences, enabling 50+ reaction cycles per support with no loss of performance. This represents a step-change in support utilisation: rather than discarding 90–99% of catalyst mass after a single enzyme lifetime, carriers now serve multiple lifecycles.
In combination with its wide operational pH range, the modular, aqueous chemistry provides broad compatibility with diverse enzymes, buffer systems and process configurations (i.e. packed-bed operation and standard process equipment). Looking forward, this dynamic binding opens opportunities beyond traditional support regeneration. The specific pH-triggered release enables enzyme immobilisation on high-value materials, such as electrodes for biosensors, optical fibres for process monitoring, or specialised coatings on medical devices, where recovery and reuse of both the enzyme and the substrate become economically justified. The visual tracking and reversibility are well-suited for automated flow systems with in-line regeneration and real-time process monitoring, whilst 3D-printed scaffolds and microfluidic devices could exploit the on-demand loading capability for spatially programmed biocatalysis. This work provides a practical route to address material circularity in industrial biocatalysis, demonstrating that catalytic performance and sustainability are compatible goals, and that reversibility expands rather than limits the scope of enzyme immobilisation.
Author contributions
G. B. conceived and developed all experiments on the alizarin-IDA immobilisation strategy, analysed data, and wrote the manuscript. J. S. performed the initial dopamine-based immobilisation experiments. F. P. conceptualised and supervised the project, contributed to experimental design and data interpretation, and co-wrote the manuscript. All authors have read and approved the final version.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5sc08585c.Acknowledgements
This publication was created as part of NCCR Catalysis (grant number 225147), a National Centre of Competence in Research funded by the Swiss National Science Foundation, with further support from the SNSF individual grant (200021_232199).Notes and references
- R. A. Sheldon and J. M. Woodley, Chem. Rev., 2018, 118, 801–838 CrossRef CAS PubMed.
- A. Basso and S. Serban, Mol. Catal., 2019, 479, 110607 CAS.
- Y. R. Maghraby, R. M. El-Shabasy, A. H. Ibrahim and H. M. E.-S. Azzazy, ACS Omega, 2023, 8, 5184–5196 CrossRef CAS PubMed.
- J. Zdarta, A. S. Meyer, T. Jesionowski and M. Pinelo, Catalysts, 2018, 8, 92 CrossRef.
- R. A. Sheldon, A. Basso and D. Brady, Chem. Soc. Rev., 2021, 50, 5850–5862 RSC.
- R. A. Sheldon and S. van Pelt, Chem. Soc. Rev., 2013, 42, 6223–6235 RSC.
- C. Ortiz, M. L. Ferreira, O. Barbosa, J. C. S. dos Santos, R. C. Rodrigues, Á. Berenguer-Murcia, L. E. Briand and R. Fernandez-Lafuente, Catal. Sci. Technol., 2019, 9, 2380–2420 RSC.
- P. Zucca and E. Sanjust, Molecules, 2014, 19, 14139–14194 CrossRef PubMed.
- T. Prabhakar, J. Giaretta, R. Zulli, R. J. Rath, S. Farajikhah, S. Talebian and F. Dehghani, Chem. Eng. J., 2025, 503, 158054 CrossRef CAS.
- R. C. Rodrigues, Á. Berenguer-Murcia, D. Carballares, R. Morellon-Sterling and R. Fernandez-Lafuente, Biotechnol. Adv., 2021, 52, 107821 CrossRef CAS PubMed.
- R. A. Sheldon, Philos. Trans. R. Soc., A, 2020, 378, 20190274 CrossRef CAS PubMed.
- M. Becker, A. Ziemińska-Stolarska, D. Markowska, S. Lütz and K. Rosenthal, ChemSusChem, 2023, 16, e202201629 CrossRef CAS PubMed.
- R. E. Ruscoe and S. C. Cosgrove, Curr. Opin. Green Sustainable Chem., 2024, 49, 100954 CrossRef CAS.
- A. A. Homaei, R. Sariri, F. Vianello and R. Stevanato, J. Chem. Biol., 2013, 6, 185–205 CrossRef PubMed.
- M. Nur Royhaila, N. H. C. Marzuki and R. A. Wahab, Biotechnol. Biotechnol. Equip., 2015, 29, 205–220 CrossRef PubMed.
- F. Lu, H. Zhang, W. Pan, N. Li and B. Tang, Chem. Commun., 2021, 57, 7067–7082 RSC.
- S. Ulrich, Acc. Chem. Res., 2019, 52, 510–519 CrossRef CAS PubMed.
- A. Dirksen and P. E. Dawson, Bioconjugate Chem., 2008, 19, 2543–2548 CrossRef CAS PubMed.
- D. K. Kölmel and E. T. Kool, Chem. Rev., 2017, 117, 10358–10376 CrossRef PubMed.
- K. Ovsejevi, C. Manta and F. Batista-Viera, Methods Mol. Biol., 2013, 1051, 89–116 CrossRef CAS PubMed.
- J. M. Bolivar, J. M. Woodley and R. Fernandez-Lafuente, Chem. Soc. Rev., 2022, 51, 6251–6290 RSC.
- A. I. Benítez-Mateos and F. Paradisi, ChemSusChem, 2022, 15, e202102030 CrossRef PubMed.
- G. Springsteen and B. Wang, Tetrahedron, 2002, 58, 5291–5300 CrossRef CAS.
- W. L. A. Brooks, C. C. Deng and B. S. Sumerlin, ACS Omega, 2018, 3, 17863–17870 CrossRef CAS PubMed.
- A. Stubelius, S. Lee and A. Almutairi, Acc. Chem. Res., 2019, 52, 3108–3119 CrossRef CAS PubMed.
- S. D. Bull, M. G. Davidson, J. M. H. van den Elsen, J. S. Fossey, A. T. A. Jenkins, Y.-B. Jiang, Y. Kubo, F. Marken, K. Sakurai, J. Zhao and T. D. James, Acc. Chem. Res., 2013, 46, 312–326 CrossRef CAS PubMed.
- D. Li, Y. Chen and Z. Liu, Chem. Soc. Rev., 2015, 44, 8097–8123 RSC.
- Z. Liu and H. He, Acc. Chem. Res., 2017, 50, 2185–2193 CrossRef CAS PubMed.
- Y. Chen, A. Huang, Y. Zhang and Z. Bie, Anal. Chim. Acta, 2019, 1076, 1–17 CrossRef CAS PubMed.
- X. C. Liu and W. H. Scouten, J. Mol. Recognit., 1996, 9, 462–467 Search PubMed.
- X. Wang, N. Xia and L. Liu, Int. J. Mol. Sci., 2013, 14, 20890–20912 CrossRef PubMed.
- R. Torres, B. Pessela, M. Fuentes, R. Munilla, C. Mateo, R. Fernández-Lafuente and J. M. Guisán, J. Biotechnol., 2005, 120, 396–401 CrossRef CAS PubMed.
- Y. Dong, W. Wang, O. Veiseh, E. A. Appel, K. Xue, M. J. Webber, B. C. Tang, X.-W. Yang, G. C. Weir, R. Langer and D. G. Anderson, Langmuir, 2016, 32, 8743–8747 Search PubMed.
- S. Morariu, Polymers, 2023, 15, 582 CrossRef CAS PubMed.
- B. Yang, Y. Lv, J.-Y. Zhu, Y.-T. Han, H.-Z. Jia, W.-H. Chen, J. Feng, X.-Z. Zhang and R.-X. Zhuo, Acta Biomater., 2014, 10, 3686–3695 CrossRef CAS PubMed.
- F. Duval, T. A. van Beek and H. Zuilhof, Synlett, 2012, 23, 1751–1754 Search PubMed.
- G. Springsteen and B. Wang, Chem. Commun., 2001, 1608–1609 RSC.
- Y. Suzuki, T. Sugaya, S. Iwatsuki, M. Inamo, H. D. Takagi, A. Odani and K. Ishihara, ChemistrySelect, 2017, 2, 2956–2964 CrossRef CAS.
- R. Nishiyabu, Y. Kubo, T. D. James and J. S. Fossey, Chem. Commun., 2011, 47, 1106–1123 Search PubMed.
- Z. Tang, Y. Oku and T. Matsuda, Org. Process Res. Dev., 2024, 28, 1308–1326 CrossRef CAS.
- M. Michaud, G. Nonglaton and Z. Anxionnaz-Minvielle, Chembiochem, 2024, 25, e202400086 Search PubMed.
- C. L. F. Regueiro, D. R. Padrosa and F. Paradisi, C. R. Chim., 2025, 28, 349–359 Search PubMed.
- M. L. Contente, S. Farris, L. Tamborini, F. Molinari and F. Paradisi, Green Chem., 2019, 21, 3263–3266 RSC.
- V. Marchini, A. I. Benítez-Mateos, D. Roura Padrosa and F. Paradisi, Front. Catal., 2021, 1 DOI:10.3389/fctls.2021.790461.
- P. Díaz-Kruik, D. Roura Padrosa, E. Hegarty, H. Lehmann, R. Snajdrova and F. Paradisi, Org. Process Res. Dev., 2024, 28, 2683–2691 Search PubMed.
- K. Stalder, A. I. Benítez-Mateos and F. Paradisi, ChemCatChem, 2024, 16, e202301671 Search PubMed.
- P. Zucca, R. Fernandez-Lafuente and E. Sanjust, Molecules, 2016, 21, 1577 CrossRef PubMed.
- N. Rueda, J. C. S. dos Santos, R. Torres, C. Ortiz, O. Barbosa and R. Fernandez-Lafuente, RSC Adv., 2015, 5, 11212–11222 Search PubMed.
- P. Cannazza, S. Donzella, A. Pellis and M. L. Contente, Biotechnol. Adv., 2022, 59, 107985 CrossRef CAS PubMed.
- D. Roura Padrosa, V. Marchini and F. Paradisi, Bioinformatics, 2021, 37, 2761–2762 Search PubMed.
- C. Mateo, J. M. Palomo, G. Fernandez-Lorente, J. M. Guisan and R. Fernandez-Lafuente, Enzyme Microb. Technol., 2007, 40, 1451–1463 CrossRef CAS.
- Z. Zhang, Z. Zhao, K. Huang and Z. Liang, Appl. Microbiol. Biotechnol., 2023, 107, 6163–6178 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |