 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
DP5 without DFT: uncertainty-calibrated graph neural net accelerates structure confirmation via NMR
Ruslan
Kotlyarov
 *,
Alexander
Howarth
and
Jonathan M.
Goodman
*,
Alexander
Howarth
and
Jonathan M.
Goodman
 *
*
Yusuf Hamied Department of Chemistry, Lensfield Road, Cambridge CB2 1EW, UK. E-mail: jmg11@cam.ac.uk
First published on 5th March 2026
Abstract
The evaluation and assignment of candidate structures to NMR spectra can be facilitated by the DP4 method, which assumes that one of the candidate structures is correct, and the DP5 method, which calculates the probability of a correct assignment for each candidate individually. Both of these methods require DFT calculations and thus a significant amount of computer resources. In this paper we present DP5q, a new version of DP5, which uses a graph convolutional neural network and quantile regression to replace the DFT-based algorithm. This dramatically increases the speed of the calculation at the cost of a modest decrease in accuracy. We demonstrate the efficacy of this rapid calculation both on a test set of thousands of molecules and also on cases selected for the difficulty of assigning the structure.
1 Introduction
Nuclear magnetic resonance spectroscopy is an important technique for characterisation of small molecules. It requires very little material for successful spectrum collection. The sample is easy to prepare, and the technique is non-destructive, i.e., the product may be recovered intact due to the very low energy of electromagnetic radiation employed in the method.Structure confirmation is relatively easy to accomplish when an experimental spectrum of a pure substance has already been recorded under the same conditions, such as temperature, solvent, and acidity. However, when a new compound is made, there are no reference spectra to compare.
The assignment of candidate structures to NMR spectra can be a challenging task, particularly when multiple structures have similar spectra. This has led to many examples of misassignment that have later been corrected.1–23 This issue is becoming more acute with increased automation, miniaturisation, and parallelisation of chemical synthesis. The amount of data generated within a single experimental campaign grows beyond individual chemists' capacity to process it. A fast, automated way to confirm a structure from a spectrum would address this growing challenge.
Our previous studies in the area of computer-aided structure elucidation led to CP3 (ref. 24) and DP4 (ref. 25) methods, which select the structure from the best-matching structure from the user-provided list, DP4-AI,26–28 which automates the process, and, most recently, DP5 (ref. 29), which determines the confidence with which a single structure may be assigned to a single spectrum. All of these methods require DFT analyses and the resources needed for these calculations are the rate determining steps for the process. In this work, we present a new version of DP5 analysis, DP5q, which obviates the need for DFT calculations while maintaining the accuracy to a high level. This dramatically improves the throughput for the DP5 process.
2 Neural net design
2.1 Learning explicit distribution parameters
Neural nets have already been used to help with structure elucidation.30–33 Several models can predict NMR spectra.34,35 IMPRESSION34 featurises molecular geometries using FCHL representations36 and uses kernel ridge regression to predict both the shift value and its variance. Graph-convolutional neural nets have been used to generate new learned features and predict molecular and atomic properties together with the associated uncertainties.35,37Estimation of uncertainty for chemical problems is well-precedented and is used in several models that can predict NMR spectra.34,35 IMPRESSION,34 which was developed by Gerrard et al., featurises molecular geometries using FCHL representations36 and uses kernel ridge regression to predict both the shift value and its variance. Work by Jonas and Kuhn,35 on the other hand, used a graph-convolutional neural net to generate new learned features and predicted shift and its uncertainty using two separate heads. Both approaches assume normal distributions for their predictions.
2.2 Quantile regression
In our model, we have used quantile regression for our uncertainty estimation. Instead of predicting explicit parameters of a probability distribution (PDF) as an output, it predicts the values of inverse cumulative distribution of a chemical shift for any specified set of quantiles (Fig. 1). | ||
| Fig. 1 Quantile regression as an uncertainty estimation method is easy to incorporate into existing deep learning model architectures for chemical shift prediction. | ||
The approach is easy to integrate into the architecture of CASCADE,38 a graph neural network: instead of a single output, the final layer of the model can be reconfigured to produce multiple outputs corresponding to each specified quantile. For example, we can configure and train the model to predict the median, the first quartile, and the seventh decile of the probability distribution for a chemical shift. Realising that each additional quantile requires significantly fewer trainable parameters compared with the total number of the parameters within the model, we have evaluated chemical shift predictions for all percentiles from the 1st to the 99th, as the 0th and the 100th percentiles would simply correspond to minimum and maximum shifts observed in the training dataset. This allows us to balance effective interpolation of the cumulative distribution function for accurate analysis and the number of trainable parameters in the model.
2.3 Comparison with prior approaches
This work builds upon our prior efforts in uncertainty quantification for NMR shift predictions. Previously, we relied extensively on DFT calculations. For example, the DP4 approach25,28 uses errors between scaled DFT-predicted and unscaled experimental shifts and compares them to the previously benchmarked normal error distributions for correct structure assignments. These errors are converted into atomic probabilities, which are then multiplied together, and the sum of molecular probabilities for all candidate structures is normalised to unity. The structure with the highest DP4 score is selected as the most probable structure. This approach assumes that one of the proposed structures is correct and does not account for any structural features.DP5 analysis29 goes further and constructs a dataset with tens of thousands of FCHL representations36 of atomic environments and their associated DFT errors. For an observed atomic environment, a probability distribution is constructed based on similar atomic environments to generate the probability of observing such an error. These atomic probabilities are then combined into the absolute molecular probability of correct structure assignment.
Our new neural net-based approach removes the need for DFT calculations altogether. Instead, the model constructs the probability distribution for chemical shifts directly. The speed of quantile regression-based DP5q analysis no longer depends on the size of the reference dataset, which allows us to arbitrarily scale up the size of the training data without increasing the computational cost.
3 Training
3.1 Loss function
The proposed loss function for the quantile regression is made of two parts:39 the prediction loss for all modelled quantiles and a regularisation term.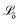 (eqn (1)) for each quantile prediction ŷτ using quantiles specified at the point of model creation (eqn (2)). In the limit of a low δ parameter (here, set to 1 × 10−4 for good numerical convergence during training), modified Huber loss reduces to mean absolute error and our result for a single quantile τ becomes indistinguishable from pinball loss.
(eqn (1)) for each quantile prediction ŷτ using quantiles specified at the point of model creation (eqn (2)). In the limit of a low δ parameter (here, set to 1 × 10−4 for good numerical convergence during training), modified Huber loss reduces to mean absolute error and our result for a single quantile τ becomes indistinguishable from pinball loss.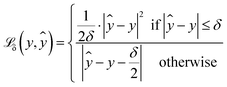 | (1) |
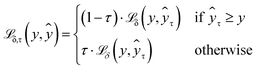 | (2) |
An example of the proposed loss function for a single quantile τ is shown in Fig. 2. For all quantiles, the correct prediction has no associated loss. However, the parameter τ is used to weight the loss for over- and under-predictions. For example, if τ is set to 0.75, the loss for under-prediction is three times larger than the loss for over-prediction. Conversely, if τ is set to 0.5, the penalty for over- and under-prediction is equal and guides the model to predict the median of the distribution.
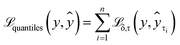 | (3) |
By minimising the sum of losses 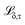 for each quantile τ (eqn (3)), we hope that predictions for all quantiles will both accurately reflect the probability distribution and approach the true value y, thus preventing the model from overestimating its uncertainty. In this study, we have used n = 99 quantiles, corresponding to percentiles from 1 to 99, i.e., τ = (0.01, 0.02, …, 0.99).
for each quantile τ (eqn (3)), we hope that predictions for all quantiles will both accurately reflect the probability distribution and approach the true value y, thus preventing the model from overestimating its uncertainty. In this study, we have used n = 99 quantiles, corresponding to percentiles from 1 to 99, i.e., τ = (0.01, 0.02, …, 0.99).
We can enforce the non-decreasing character by adding the second loss term 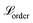 (eqn (4)), which penalises higher quantiles having lower values, i.e., preserves ranking order and monotonicity. With a larger number of prediction quantiles, it should be easier to maintain the desirable effects of the proposed loss function. Here, ε is a small positive number set to 1 × 10−6 to ensure a positive difference between two consecutive quantiles.
(eqn (4)), which penalises higher quantiles having lower values, i.e., preserves ranking order and monotonicity. With a larger number of prediction quantiles, it should be easier to maintain the desirable effects of the proposed loss function. Here, ε is a small positive number set to 1 × 10−6 to ensure a positive difference between two consecutive quantiles.
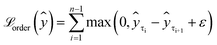 | (4) |
We, therefore, adapted the sum of the two components as the final loss function for the model (eqn (5)).
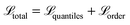 | (5) |
3.2 Dataset preparation
The training dataset has been curated from NMRShiftDB40 data (Fig. 3). Since the goal is to analyse carbon NMR spectra, all entries without the carbon NMR spectrum were discarded. To aid training of the graph neural net, all molecules containing multiple fragments (e.g., salts) were removed as well. Molecules containing atoms other than H, B, C, N, O, F, P, S, Cl, Br, and I were discarded, as rare atom types hinder effective training. Possible duplicates were treated as independent entries.For each curated molecule, a 3D conformer was generated using ETKDG, as implemented in RDKit.41 The structures had their geometries optimised using the MMFF force field. Shielding tensors were calculated using DFT (mPW1PW91/6-311g(d)) in line with our previous studies28 without further geometry optimisation and then converted into chemical shifts using reference shielding tensor values for tetramethylsilane. After the calculations, any molecule with a large difference between predicted and observed shifts (50 ppm) was deemed to be an erroneous interpretation or assignment of the spectrum and discarded, leaving 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 349 records for 22
349 records for 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 248 unique molecules and their associated experimental chemical shifts.
248 unique molecules and their associated experimental chemical shifts.
3.3 Training procedure
The model was then trained using the procedure described by Guan38 for 1200 epochs, an initial learning rate of 5 × 10−4, and reduction of the learning rate by 4% every 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 training steps using the Adam optimiser. Early stopping after 10 epochs without improvement in validation loss was used to prevent overfitting.
000 training steps using the Adam optimiser. Early stopping after 10 epochs without improvement in validation loss was used to prevent overfitting.
4 Transforming model uncertainty into structure confidence
Accurate uncertainty quantification is important for our use case. However, it is not pursued for its own sake, but rather as a step towards structure confirmation and to generate the probability distribution we will then integrate for an error score Pi as defined in eqn (6) (orange area in Fig. 4). Here, the integration limits are set between the predicted median minus absolute error ŷ−|ŷ−y| and predicted median plus mean absolute error ŷ+|ŷ−y|. The advantage of quantile regression in this case is that the model outputs the quantiles of the cumulative distribution directly without the need to integrate the probability distribution.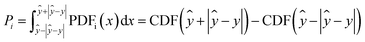 | (6) |
 | ||
| Fig. 4 Calculation of the error score for the subsequent DP5 analysis using the experimental shift value and the predicted probability distribution. | ||
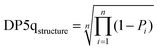
The DP5q score for atom i is defined in eqn (7) (blue area in Fig. 4) and the molecular DP5 score for n atoms would be their geometric average (eqn (8)). A high DP5q score therefore means that the observed shift is consistent with the chemical environment in the molecule.
| DP5qi = 1 − Pi | (7) |
 | (8) |
5 Testing
5.1 Impact of uncertainty quantification on shift prediction accuracy
To see if the model has been trained correctly, its median predictions were compared to experimental data (Fig. 5). The accuracy against experimental data deteriorated somewhat, with MAE increasing from 1.76 to 2.19 ppm. The increase in mean absolute error could mean an overfit to training data. However, the overfit must have been mitigated by the use of a validation set and early stopping. Another explanation would be a shift of the objective. Since the median is only one of 99 quantiles to be predicted, it now has a lower contribution to the loss function than in the single-output scenario, where mean absolute error was employed for a single prediction. Other quantiles are as important as the median in our setup, since we now need to accurately predict the probability distribution for chemical shifts.5.2 DP5q is more sensitive than mean absolute error analysis
When deploying DP5q analysis in practice, we need to demonstrate that the scores we obtain carry practical meaning, i.e., a high score means that the structure is likely to be correct, and a low score means that the structure is likely to be incorrect. We can readily generate the distribution of DP5q scores for the correctly assigned molecule, but how could we generate the scores for incorrect assignments? Some studies generate ‘incorrect’ structures artificially by altering the molecular structures already present in the dataset.42,43 However, we typically have no information about the experimental spectra of these structures and therefore cannot compute DP5q scores for them.In combinatorial studies,29 we obtain ‘incorrect’ data without generating new molecules by taking advantage of existing databases. We notice that the database is a set of molecules with interpreted NMR spectra. Since there is a one-to-one match between a molecule and a spectrum and vice versa, all molecules but one in a data set will be an incorrect interpretation of a given spectrum. Therefore, using a dataset of size N provides us with N ‘correct’ spectrum–structure pairs and at most N2 − N ‘incorrect’ spectrum–structure pairs. For each pair, we compute mean absolute error and the DP5q score using eqn (8).
Within each group, we then assigned spectra to right and wrong structures and checked if the scores for correct structures are visibly different from the scores for incorrect structures. The utility of the DP5q score is to discriminate between right and wrong proposals.
We have taken the test set of 5000 molecules the NMR predicting model (Exp5k dataset from ref. 38) has not seen and assigned a spectrum to all of the appropriate structure-spectrum pairs using the algorithm developed by Lewis et al.44 Here, we have obtained 5000 correct assignments and 5614 incorrect assignments where the incorrect proposal had the same molecular formula.
Their distributions of mean absolute errors and DP5q scores are shown in Fig. 6. While the distributions of mean absolute errors for correct and incorrect proposals overlap significantly, the distributions of DP5q scores are much more distinct, with scores for incorrect proposals concentrated around zero. This demonstrates that the DP5q score is a useful way to select correct proposals.
Two summary metrics can be combined together to assess confidence in the structure interpretation (Fig. 7). We group the neighbouring data points into bins and calculate the ratio of correct interpretations to the total number of interpretations within each bin. This ratio may be interpreted as the probability of a spectrum being interpreted correctly given the combination of mean absolute error and DP5 score.
Here, we clearly observe a steep decline of the DP5q score with increasing mean absolute error. This is reasonable to expect as correct interpretations of the spectra must match the data closely. We then see that beyond a mean absolute error of 4 ppm, the DP5 score almost does not vary with increasing error. This is expected, as the structures with high errors are near certain to be incorrect.
The graph also highlights a clear separation between correct and incorrect proposals. Correct proposals tend to cluster at higher DP5q scores and lower mean absolute errors, while incorrect proposals are more dispersed and skewed towards lower DP5 scores. This separation demonstrates the robustness of the DP5 metric in distinguishing correct structures from incorrect ones.
The combination of two metrics also helps us isolate high-confidence proposals with greater care. For example, proposals with low DP5 scores combined with low mean absolute errors are considered more likely than proposals with the same low DP5 scores combined with high mean absolute errors.
Additionally, the steep decline in DP5 scores for small increases in mean absolute error indicates that the method is highly sensitive to small deviations in predicted spectra. This sensitivity is advantageous for identifying subtle differences between closely related structures, making the DP5 approach particularly useful for challenging cases of structure elucidation.
5.3 Challenging structural revision case studies
In these studies, we have demonstrated the effectiveness of the new DP5q approach as a summary metric. These offer a bird’s-eye, abstract view of its performance. To gain the confidence of laboratory-based chemists, we consider concrete examples, taken from the structure revision studies (see Fig. 8). Molecules S12, S23, S34, S45, S56, S6–S87, S98, S109, S1110, S1211, and S1312 were used in our previous DFT-based DP5 study.29 Molecules S1413, S1514, S1615, S1716, S1817, S1918, S2019, S2120, S2221, S2322, and S2423 are new.Here, a structure for a molecule was proposed based on multiple methods, including several NMR experiments, mass, UV-vis spectroscopy, and infrared spectroscopy. However, subsequent attempts to re-create the proposed structure did not succeed, with recorded spectra not matching that of the authentic sample. New structures are then proposed and confirmed by de novo chemical synthesis or by comparison with spectra of known natural products.
We took 24 case studies and conducted DP5q analysis using single conformers, optimised at the force-field level. For 22 cases, the score for the correct proposal was higher than the score for the incorrect proposals (Fig. 9). This result is comparable to that of 23 examples where the mean absolute error of a revised structure is lower than the mean absolute error of the original structure. We note that this is a very challenging test set: someone has already analysed the spectra in detail and published a conclusion that was later found to be incorrect. For more common cases, by distinguishing automatically between large numbers of diverse structures the results are likely to be even more reliable.
 | ||
| Fig. 9 DFT-free DP5q for structure reassignment case studies. In 22 cases out of 24, the correct structure had a higher score than the initial proposal. | ||
This is a key result: we achieve the DFT performance at the neural network cost. What previously took CPU months is now achievable in seconds. The only example that the model could not tackle was S13. Therefore, it merits a closer look (Fig. 10).
 | ||
| Fig. 10 For S13, the initial and revised proposals have drastically different molecular weights and formulae. | ||
Here, we see that the original proposed structure was a symmetric binaphthyl, while the revised structure lacks any symmetry elements. Furthermore, the structures have different numbers of distinct NMR environments, 11 and 13 respectively. Both proposals, however, have four non-labile proton environments and four non-quaternary carbon environments. If the spectra have a low signal-to-noise ratio, the other, less intense, signals may be indistinguishable from the baseline. This shows that our approach depends on correct interpretation of the NMR spectra: if signals are lost into noise, an accurate analysis may not be possible. The molecular masses of the two proposals are quite different as well. This highlights the importance of using several structural determination techniques at once, so that spurious proposals based on a single modality are rejected by an orthogonal method and good proposals are corroborated. Addressing this issue is a topic of on-going research.
Using the ETKDGv3 algorithm within RDKit as a quick conformational search tool improves the accuracy of our analysis, with higher scores predicted for the correct compound in all 24 cases (Fig. 11). On the other hand, a simple comparison of mean absolute errors returned the correct result 23 times out of 24. This highlights the importance of the representative conformer ensemble in accurate prediction of NMR shifts, even when DFT spectrum calculation has been eliminated from the workflow.
The combination of DP5q scores and mean absolute errors (MAE) of the carbon spectra (Fig. 12) shows that they are linked to some extent, as higher scores tend to correspond to lower errors. The distribution of correct and incorrect proposals resembles the one observed in the combinatorial studies (Fig. 7). There is a small overlap between DP5q scores of correct and incorrect proposals, but the correct proposals still tend to have a lower error and higher DP5q score. This is a good result and shows that the DP5q score is an effective method even for structurally complex molecules, such as natural products.
5.4 Relative stereochemistry elucidation
Another worthwhile and very challenging test of the method would be to apply it to an established test set used to determine performance of DP4-AI.28 It consists of 42 molecules with their interpreted carbon spectra (Fig. 13). The dataset is made of natural products and synthetic intermediates to represent a broad range of chemical space. The differences between the diastereomers are rather small, which makes it a robust test of DP5 analysis. A good result for this challenging dataset would demonstrate the power of the method. We have already demonstrated the power of DP5q for less challenging datasets (Fig. 7). | ||
| Fig. 13 Molecules used in the DP4-AI test set. AT3, TS3A, TS4 and NL1A had no corresponding carbon spectra and were, therefore, excluded from analysis. | ||
We consider three scenarios with varying degrees of sophistication. In the first scenario, we use a single conformer, optimised at the force-field level. This is the fastest way to run DP5 analysis. In the second scenario, we use multiple conformers generated using low-mode conformational search, with geometries optimised at the DFT-level and energies recalculated using single-point DFT calculations. This is the most sophisticated way to run DP5q analysis, and its previous use for stereochemistry determination28 has shown it to be the most accurate. In the third scenario, we use multiple conformers generated using the ETKDGv3 algorithm41 as provided with RDKit, further optimised at the force-field level of theory.
To visualise the resulting DP5q scores, we divide them by the number of possible diastereomers. For example, if the molecule has 8 diastereomers, the plotted DP5q scores will be divided by 8. This helps us both indicate the relative magnitudes and absolute values for DP5 probabilities.
Here, stereochemistry was determined correctly for 18 examples out of 42 (Fig. 15), with a 0.13% chance of obtaining a result this good at random. This is an encouraging improvement, which is consistent with the trend where more sophisticated treatment improves the overall accuracy.
This approach may take a few minutes instead of a fraction of a second and helps us determine correct stereochemistry in 25 out of 42 cases, based on 1D13C NMR data only, using no 1H NMR or multidimensional spectroscopy (Fig. 16). This result has a 2.9 × 10−6% probability of occurring by chance. The method meets and exceeds purpose-developed DP4 analysis at a similar level of theory,29 with 19 out of 42 cases identified correctly, the DFT-based DP5 approach, where the correct diastereomer has been found in 16 cases out of 42,29 and the CASCADE mean absolute error comparison, where the lowest-error diastereomer was the correct one 21 times out of 42. Therefore, we recommend this for the comparison of stereoisomers and other very similar structures.
6 Conclusions
A new approach has been developed for estimating the confidence of interpretation of an NMR spectrum. It is fully integrated into an existing DP5 suite, accelerating structural elucidation by several orders of magnitude by eliminating costly DFT calculations. Our approach enhances the DP5 methodology by incorporating a 3D neural network capable of predicting NMR chemical shifts along with their uncertainties. This advancement allows for rapid and accurate assessment of molecular structures based on NMR data.DP5q analysis performs admirably on the very complex structural revision case studies. Inclusion of quick conformational search results in scoring the correct proposal higher in 24 cases out of 24. DP5q analysis is also helpful for determining stereochemistry of a diverse range of molecules, with the correct diastereomer selected in 25 cases out of 42, which exceeds performance of DP4 analysis (15 examples correct) using a similar level of theory.28 Such performance makes DFT-free DP5q ideal for high-throughput NMR data curation. To aid such workflows, we recommend using ETKDGv3 conformer generation for best results and selecting a threshold of 0.2 for the DP5q score to reliably discard incorrect structure proposals.
This work sets a strong foundation for incorporating more sophisticated spectroscopy methods, including J-value analysis, 2D NMR spectra interpretation, and processing the mixtures. In addition to that, DP5q can be used as a scoring function for the plethora of molecular optimisation methods available, potentially enabling automated structure revision.
Author contributions
Prof J. M. Goodman conceptualised and supervised the project. R. Kotlyarov implemented the machine learning model and evaluated its performance. A. Howarth curated the datasets for model training and evaluation, ran the DFT calculations, and performed preliminary research.Conflicts of interest
There are no conflicts to declare.Data availability
The code for DP5 structure confirmation software can be found at https://github.com/ruslankotl/DP5.The NMRdb dataset used in this study is available at https://github.com/ruslankotl/DP5.
Supplementary information (SI): data on the acceleration of DP5q analysis over DP5, a TMAP plot showing how predicted uncertainty varies with the chemical environment, and detailed outputs of DP5 analyses. See DOI: https://doi.org/10.1039/d5sc06988b.
Acknowledgements
R.K. thanks Recursion (formerly Exscientia) and EPSRC Centre for Doctoral Training – SynTech (EP/S024220/1) for funding and his wife for support. The authors thank Benji Rowlands for visualisations of the DP5q results.Notes and references
- K. C. Nicolaou and S. A. Snyder, Angew. Chem., Int. Ed., 2005, 44, 1012–1044 CrossRef CAS PubMed.
- G. Buchi, W. D. MacLeod and J. Padilla, J. Am. Chem. Soc., 1964, 86, 4438–4444 CrossRef.
- A. V. Kalinin and V. Snieckus, Tetrahedron Lett., 1998, 39, 4999–5002 CrossRef CAS.
- G. Bringmann, J. Schlauer, H. Rischer, M. Wohlfarth, J. Mühlbacher, A. Buske, A. Porzel, J. Schmidt and G. Adam, Tetrahedron, 2000, 56, 3691–3695 CrossRef CAS.
- H. Takikawa, K. Isono, M. Sasaki and F. A. Macías, Tetrahedron Lett., 2003, 44, 7023–7025 CrossRef CAS.
- P. Wipf and A. D. Kerekes, J. Nat. Prod., 2003, 66, 716–718 CrossRef CAS PubMed.
- A. G. Kutateladze, E. H. Krenske and C. M. Williams, Angew. Chem., 2019, 131, 7181–7186 CrossRef.
- B. S. Dyson, J. W. Burton, T.-i. Sohn, B. Kim, H. Bae and D. Kim, J. Am. Chem. Soc., 2012, 134, 11781–11790 CrossRef CAS PubMed.
- M. W. Lodewyk, C. Soldi, P. B. Jones, M. M. Olmstead, J. Rita, J. T. Shaw and D. J. Tantillo, J. Am. Chem. Soc., 2012, 134, 18550–18553 CrossRef CAS PubMed.
- B. Verbraeken, J. Hullaert, J. van Guyse, K. Van Hecke, J. Winne and R. Hoogenboom, J. Am. Chem. Soc., 2018, 140, 17404–17408 CrossRef CAS PubMed.
- A. G. Kutateladze, J. Org. Chem., 2016, 81, 8659–8661 CrossRef CAS.
- E. E. Podlesny and M. C. Kozlowski, J. Nat. Prod., 2012, 75, 1125–1129 CrossRef CAS PubMed.
- S. O. Simonetti, E. L. Larghi, A. B. J. Bracca and T. S. Kaufman, Org. Biomol. Chem., 2012, 10, 4124 RSC.
- M. Tori, K. Nakashima, M. Toyota and Y. Asakawa, Tetrahedron Lett., 1993, 34, 3751–3752 CrossRef CAS.
- X. Zhang, Z. Li, H. Yong and Z. Xie, Org. Chem. Front., 2018, 5, 3469–3475 RSC.
- J. A. May, R. K. Zeidan and B. M. Stoltz, Tetrahedron Lett., 2003, 44, 1203–1205 CrossRef CAS.
- I. Cornella and T. R. Kelly, J. Org. Chem., 2004, 69, 2191–2193 CrossRef CAS PubMed.
- S. Venkateswarlu, G. K. Panchagnula, M. B. Guraiah and G. V. Subbaraju, Tetrahedron, 2005, 61, 3013–3017 CrossRef CAS.
- M. Kita, R. Miwa, T. Widianti, Y. Ozaki, S. Aoyama, K. Yamada and D. Uemura, Tetrahedron Lett., 2007, 48, 8628–8631 CrossRef CAS.
- F. Saito, K. Kuramochi, A. Nakazaki, Y. Mizushina, F. Sugawara and S. Kobayashi, Eur. J. Org Chem., 2006, 2006, 4796–4799 CrossRef.
- G. Schlingmann, T. Taniguchi, H. He, R. Bigelis, H. Y. Yang, F. E. Koehn, G. T. Carter and N. Berova, J. Nat. Prod., 2007, 70, 1180–1187 CrossRef CAS.
- Y. H. Ye, H. L. Zhu, Y. C. Song, J. Y. Liu and R. X. Tan, J. Nat. Prod., 2005, 68, 1106–1108 CrossRef CAS.
- F. Dhoro, J. Parkin-Gibbs, M. McIldowie, B. W. Skelton and M. J. Piggott, J. Nat. Prod., 2018, 81, 1658–1665 CrossRef CAS PubMed.
- S. G. Smith and J. M. Goodman, J. Org. Chem., 2009, 74, 4597–4607 CrossRef CAS.
- S. G. Smith and J. M. Goodman, J. Am. Chem. Soc., 2010, 132, 12946–12959 CrossRef CAS PubMed.
- K. Ermanis, K. E. B. Parkes, T. Agback and J. M. Goodman, Org. Biomol. Chem., 2016, 14, 3943–3949 RSC.
- K. Ermanis, K. E. B. Parkes, T. Agback and J. M. Goodman, Org. Biomol. Chem., 2017, 15, 8998–9007 RSC.
- A. Howarth, K. Ermanis and J. M. Goodman, Chem. Sci., 2020, 11, 4351–4359 RSC.
- A. Howarth and J. M. Goodman, Chem. Sci., 2022, 13, 3507–3518 RSC.
- M. Alberts, F. Zipoli and A. C. Vaucher, Learning the Language of NMR: Structure Elucidation from NMR Spectra Using Transformer Models, 2023.
- M. Alberts, O. Schilter, F. Zipoli, N. Hartrampf and T. Laino, Unraveling Molecular Structure: A Multimodal Spectroscopic Dataset for Chemistry, 2024.
- O. Schilter, M. Alberts, F. Zipoli, A. Vaucher, P. Schwaller and T. Laino, NeurIPS 2023 AI for Science Workshop, 2023.
- M. Priessner, R. J. Lewis, I. Lemurell, M. J. Johansson, J. Goodman, J. P. Janet and A. Tomberg, Angew. Chem., Int. Ed., 2026, 65, e17611 CrossRef CAS PubMed.
- W. Gerrard, L. A. Bratholm, M. J. Packer, A. J. Mulholland, D. R. Glowacki and C. P. Butts, Chem. Sci., 2020, 11, 508–515 RSC.
- E. Jonas and S. Kuhn, J. Cheminf., 2019, 11, 50 Search PubMed.
- A. S. Christensen, L. A. Bratholm, F. A. Faber and O. A. von Lilienfeld, J. Chem. Phys., 2020, 152, 044107 CrossRef CAS PubMed.
- S. Jiang, S. Qin, R. C. V. Lehn, P. Balaprakash and V. M. Zavala, Digital Discovery, 2024, 3, 1534–1553 RSC.
- Y. Guan, S. V. S. Sowndarya, L. C. Gallegos, P. C. S. John and R. S. Paton, Chem. Sci., 2021, 12, 12012–12026 RSC.
- Deep Quantile Regression in Keras, https://www.kaggle.com/code/abiolatti/deep-quantile-regression-in-keras/notebook.
- S. Kuhn and N. E. Schlörer, Magn. Reson. Chem., 2015, 53, 582–589 CrossRef CAS PubMed.
- S. Wang, J. Witek, G. A. Landrum and S. Riniker, J. Chem. Inf. Model., 2020, 60, 2044–2058 CrossRef CAS PubMed.
- M. M. Zanardi and A. M. Sarotti, J. Org. Chem., 2015, 80, 9371–9378 CrossRef CAS PubMed.
- M. Priessner, R. J. Lewis, M. J. Johansson, J. M. Goodman, J. P. Janet and A. Tomberg, J. Chem. Inf. Model., 2024, 64, 3180–3191 CrossRef CAS PubMed.
- R. Lewis, B. Rowlands, L. Jonsson, J. M. Goodman, P. Howe, W. Czechtizky and T. Leek, Towards Automatically Verifying Chemical Structures: The Powerful Combination of 1H NMR and IR Spectroscopy, 2024, doi: DOI:10.21203/rs.3.rs-4719113/v1.
| This journal is © The Royal Society of Chemistry 2026 |











