Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
CBL ubiquitin ligase targets translation as a degrader E3
Alice T.
Wicks
 ab,
Lori
Buetow
ab,
Lori
Buetow
 a,
Toshiyasu
Suzuki
a,
Toshiyasu
Suzuki
 a,
Tobias
Schmidt
a,
Tobias
Schmidt
 a,
Sergio
Lilla
a,
Sergio
Lilla
 a,
Abigail
Macmillan-Jones
c,
Jennifer
Turney
d,
Andrea
Gohlke
a,
Abigail
Macmillan-Jones
c,
Jennifer
Turney
d,
Andrea
Gohlke
 d,
Martin
Bushell
d,
Martin
Bushell
 ab,
Andreas K.
Hock
c and
Danny T.
Huang
ab,
Andreas K.
Hock
c and
Danny T.
Huang
 *ab
*ab
aCancer Research UK Scotland Institute, Garscube Estate, Switchback Road, Glasgow, G61 1BD, UK. E-mail: d.huang@crukscotlandinstitute.ac.uk
bSchool of Cancer Sciences, University of Glasgow, Glasgow, G61 1QH, UK
cAstraZeneca, R&D Biopharmaceuticals, Discovery Sciences, Cellular Assay Development, The Discovery Centre, 1 Francis Crick Avenue, Cambridge, CB2 0AA, UK
dAstraZeneca, R&D Biopharmaceuticals, Discovery Sciences, Protein Sciences, Structure & Biophysics, The Discovery Centre, 1 Francis Crick Avenue, Cambridge, CB2 0AA, UK
First published on 26th December 2025
Abstract
Proteolysis Targeting Chimeras (PROTACs) are heterobifunctional molecules that recruit an ubiquitin ligase (E3) and a neo-substrate into a ternary complex, enabling selective protein degradation. Despite the presence of over 600 E3s, only a handful are utilised in PROTAC application, potentially limiting the number of druggable targets. Here, we investigate whether Casitas B-cell lymphoma (CBL) can be harnessed as a degrader E3 to promote ubiquitination and degradation of the eukaryotic translation initiation factor 4E (eIF4E). Using a selective CBL binding peptide, CBLock, we demonstrate that CBL facilitates the ubiquitination of CBLock-eIF4E fusion in cells and in in vitro reconstituted assays. We further developed peptidic PROTACs, termed eIFTerminators, by linking CBLock to an eIF4E-binding peptide. Among them, eIFTerminator4 rapidly eliminates endogenous eIF4E via both lysosomal and proteasomal pathways. Unexpectedly, eIFTerminator4 also caused a decrease in eIF4A and eIF4G levels, leading to a reduction in overall protein translation in cells. Our findings establish proof-of-concept that CBL can function as a degrader E3, expanding the arsenal of E3s available for targeted protein degradation in combating challenging drug targets.
Introduction
Proteins are constantly degraded and replenished to maintain homeostasis. Protein degradation, a process by which proteins are hydrolysed into amino acids, is primarily carried out by lysosomes or proteasomes. Many proteins destined for degradation are conjugated with ubiquitin (Ub) through a process known as ubiquitination, which involves the sequential action of Ub-activating enzyme (E1), Ub-conjugating enzyme (E2), and Ub-ligase (E3). The research community has harnessed this natural system to eliminate disease-causing proteins by inducing their degradation through a ubiquitin-mediated pathway. One such strategy involves PROTACs (PROteolysis TArgeting Chimeras), which are heterobifunctional molecules composed of two ligands: one binds the target protein, and the other binds an E3 Ub-ligase. This brings the E3 into proximity with the target, allowing it to catalyse the ubiquitination of the target protein, leading to its degradation.1Since the advent of a heterobifunctional chimeric compound recruiting an E3 to the substrate MetAP-2 in 2001,2 approximately 50 substrates have been degraded with this approach,3 and over 25 PROTACs are currently in clinical trials.4 These PROTACs primarily rely on ligands that recruit either Cereblon (CRBN) or VHL E3s to promote ubiquitination and degradation of neo-substrates.5 However, it has been shown that CRBN- or VHL-based PROTACs do not always result in efficient ubiquitination or selective degradation of the target protein.6,7 Furthermore, acquired resistance to CRBN- and VHL-based PROTACs have been reported.8–10 Given that the human genome encodes more than 600 E3s,11 it is plausible that previously untapped E3s could offer distinct structural plasticity and cooperativity that favour productive ubiquitination,12 enabling the degradation of otherwise difficult or undruggable targets and potentially improving potency. Interest in alternative E3s for PROTAC development has increased in recent years. To date, these include DCAF1,13 DCAF2,14 DCAF11,15 DCAF15,16 DCAF16,17 DDB1,18 FBXO22,19 GID4,20 IAP,21 KEAP1,22 KLHDC2,23 TRIM21,24 MDM2,25 SPOP,26 and ZYG11B.27
Casitas B-cell lymphoma (CBL) is an E3 best characterised for its role in signalling pathways involving non-receptor and receptor tyrosine kinases (RTKs). Several features make CBL a promising candidate as an alternative E3 in PROTAC applications. Firstly, CBL interacts with various substrates localised to the cytosol or plasma membrane,28 suggesting its potential to target a broad range of neo-substrates in these compartments. Secondly, CBL mediates Ub transfer that can lead to either lysosomal or proteasomal degradation,29–31 which may improve degradation efficiency by utilising both degradation pathways, unlike conventional PROTACs that typically rely solely on the proteasome. Thirdly, CBL is a monomeric RING E3 that becomes catalytically active upon phosphorylation of Tyr371 in its linker helix region (LHR).32,33 This distinguishes it from most RING E3s used in the PROTAC system, which generally belong to the multi-subunit Cullin RING ligase family.34–36 In addition, the modular domain architecture of CBL enables it to ubiquitinate substrates recruited via its N-terminal tyrosine kinase-binding domain (TKBD), proline-rich region or C-terminal region, suggesting its catalytic RING domain has ample spatial accessibility to carry out ubiquitination. Finally, CBL is ubiquitously expressed across most human tissues (The Human Protein Atlas, http://www.proteinatlas.org/),37 which potentially makes CBL a pan-cancer therapeutic.
Recently, we developed a peptide, CBLock, that binds selectively to the TKBD substrate-binding site of CBL with nanomolar binding affinity (Fig. 1A).38 In this study, we investigated whether CBL can be utilised as a degrader E3 by using CBLock as a handle to recruit neo-substrates to CBL. To demonstrate proof-of-concept, we targeted eIF4E, a key component of the eIF4F translation initiation complex, which also includes eIF4A and eIF4G, and plays a vital role in binding the 5′ cap of mRNA to facilitate translation.39 Overexpression of eIF4E has been linked to hallmarks of cancer such as uncontrolled cell growth and proliferation.40–43 Cancer-dependency analyses further show that many different cancer cell types rely heavily on eIF4E for survival (https://depmap.org/portal). Consistently, antisense oligonucleotide-mediated downregulation of eIF4E suppresses tumour growth in xenograft models.44 Moreover, studies in eIF4E haplo-insufficient mice revealed that a 50% reduction in eIF4E is compatible with normal development and global protein synthesis, yet is sufficient to block cellular transformation.45 Thus, there is an interest in targeting eIF4E for degradation by induced-proximity approaches. However, CRBN and VHL-recruiting PROTACs are so far unable to successfully degrade eIF4E in cells.46,47 To test if a previously unexplored CBL can be used as a degrader E3 in a PROTAC system, CBL-recruiting PROTACs, termed eIFTerminators, were generated by linking CBLock to the eIF4E-targeting sequence eIF4G1-D5S.48 Biochemical and cell-based analyses showed that eIFTerminators enable ternary complex formation and promote CBL-mediated eIF4E degradation as well as other components of eIF4F complex, including eIF4A and eIF4G, leading to a reduction in protein translation. These findings demonstrate the feasibility of using CBL as a degrader E3 for targeted protein degradation.
Results and discussion
Appending CBL-binding peptide to eIF4E facilitates its ubiquitination by CBL
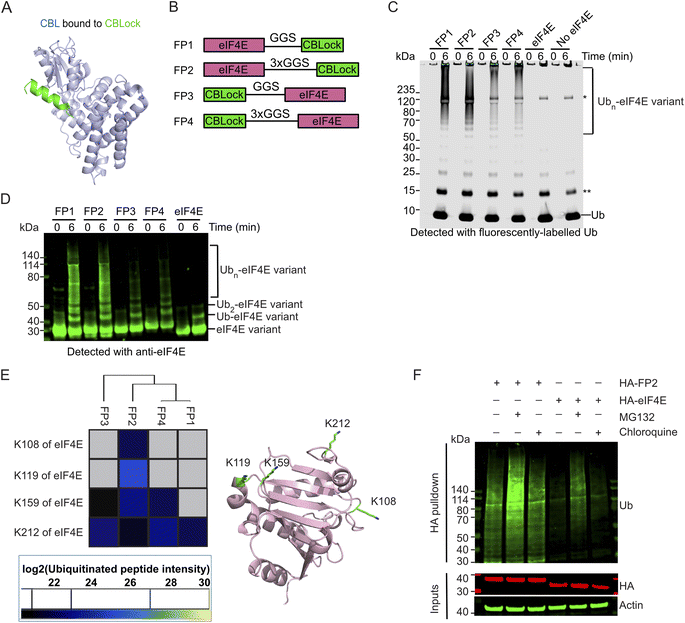
To test whether CBL could serve as a degrader E3, we examined its ability to ubiquitinate a neo-substrate in vitro upon induced proximity. As the 16-mer peptide CBLock occupies TKBD substrate-binding site where multiple substrates are known to bind and undergo ubiquitination (Fig. 1A; Ahmed et al., 2025; PDB: 9ERZ), we decided to use CBLock as a ligand to promote ubiquitination of a neo-substrate eIF4E by CBL. We fused a CBLock peptide sequence to either the N- or C-terminus of eIF4E, using short or long Gly-Gly-Ser (GGS) linkers (Fig. 1B) to facilitate CBL recruitment. The length of the GGS linkers was chosen based on a structural model of the N-terminal catalytic fragment of pTyr371-CBL bound to E2-Ub and CBLock, with eIF4E's globular body modelled near the E2-Ub active site and its N- or C-terminus proximal to CBLock's N- or C-terminus, respectively (SI Fig. 1A). These CBLock-eIF4E fusion constructs, referred to as FP1-FP4, were purified and then assessed for CBL-mediated ubiquitination.In vitro ubiquitination assays were performed using recombinant UBA1 (E1), UBE2D2 (E2), fluorescent-labelled Ub and the Tyr371-phosphorylated CBL fragment (residues 47-435, pTyr371-CBL), which represents the catalytically active portion of CBL as described previously.32 FP1-FP4 were readily ubiquitinated by pTyr371-CBL, as indicated by the appearance of Ub ladders detected with fluorescent-Ub or Coomassie staining, which were absent in control reactions with eIF4E lacking a CBLock peptide appendage as well as reactions without any form of eIF4E (Fig. 1C and SI Fig. 1B). In addition, when detected with an anti-eIF4E antibody, FP1-FP4 showed pronounced eIF4E ladders compared to lanes containing eIF4E lacking a CBLock fusion peptide (Fig. 1D). To investigate the modification sites on FP1-FP4 and the poly-Ub chain types, we subjected the smears above the 120 kDa region, indicated in black boxes, to mass spectrometry analysis (SI Fig. 1B). Mass spectrometry confirmed that Ub was the most abundant protein identified in these smears. Various Ub linkages were detected, with K11- and K48-linked Ub peptides being the most prevalent, consistent with the promiscuous activity of UBE2D2 (SI Fig. 1C). Ubiquitination was observed at four lysine residues of eIF4E: K108, K119, K159, and K212 (Fig. 1E). These lysine sites are distributed across one face of the eIF4E protein, with K212 ubiquitination consistently detected in FP1-FP4 (Fig. 1E). Notably, FP2 was the only construct ubiquitinated at all four lysine sites. In contrast, FP1, which shares the same eIF4E-CBLock orientation as FP2 but has a shorter GGS-linker, was only ubiquitinated at K212, and the K29-Ub linkage was absent. FP4, which retains FP2's GGS-linker but has the reversed eIF4E-CBLock orientation, was only ubiquitinated at K159 and K212, with the K27-, K29-, and K33-Ub linkages being absent. These findings suggested that the spatial arrangement of eIF4E relative to CBLock influences both the ubiquitination sites on eIF4E and the resulting Ub chain patterns.
Since FP2 exhibited a greater extent of ubiquitination sites on eIF4E in a reconstituted system, we next investigated whether endogenous CBL could facilitate ubiquitination of FP2 in cells. We generated stable cell lines that constitutively express either HA-eIF4E as a control or HA-FP2. The HA-tag was appended to the N-terminus of FP2, as this sequence lacks lysine residues that could undergo ubiquitination. Previous studies have shown that EGF stimulation activates CBL E3 ligase activity via phosphorylation of its Tyr371.32,49,50 To assess the phosphorylation status of CBL Tyr371, HEK293 cells expressing Myc-tag CBL were serum-starved overnight, treated with EGF, and harvested at different time points. CBL exhibited basal Tyr371 phosphorylation, which peaked at 5 min post-stimulation and returned to baseline within 2 h (SI Fig. 1D). Based on this, we treated HA-FP2 expressing cells with EGF after serum starvation to determine whether HA-FP2 undergoes CBL-mediated ubiquitination. HA-FP2 protein levels were detectable, and treatment with the proteasome inhibitor MG132 or lysosomal inhibitor chloroquine had little impact on its abundance. To directly assess ubiquitination, we performed HA pulldown under denatured conditions followed by anti-Ub immunoblotting. HA-FP2 was poly-ubiquitinated to a greater extent than HA-eIF4E (Fig. 1F). MG132 treatment intensified the poly-ubiquitinated bands, suggesting stabilisation, whereas chloroquine had a minimal effect. These findings demonstrate that CBL can induce poly-ubiquitination of a neo-substrate and promote its degradation when brought into proximity via a CBL-binding peptide fusion.
Designing peptidic PROTACs to recruit CBL for targeting eIF4E
Previous studies have identified a sub-micromolar 12-mer peptide binder of eIF4E, eIF4G1-D5S, using phage display technology.48 The crystal structure of eIF4G1-D5S bound to eIF4E revealed that it occupies the same dorsal surface of eIF4E as the eIF4G canonical helix (Fig. 2A).51 To recruit CBL to eIF4E's proximity, we designed peptidic PROTACs by linking CBLock to eIF4G1-D5S. We modelled the eIF4G1-D5S-eIF4E complex onto a structural model of the pTyr371-CBL-E2-Ub-CBLock complex by simultaneously positioning eIF4G1-D5S in proximity to CBLock and eIF4E near the E2-Ub active site (SI Fig. 2A and B). The model suggested that linking eIF4G1-D5S to either terminus of CBLock would support this arrangement. Based on this, we generated four variants incorporating either a short GGSG or a long GGSGGSGGSG linker, with CBLock positioned at either terminus of eIF4G1-D5S. These constructs were named eIFTerminator1-4 (Fig. 2B).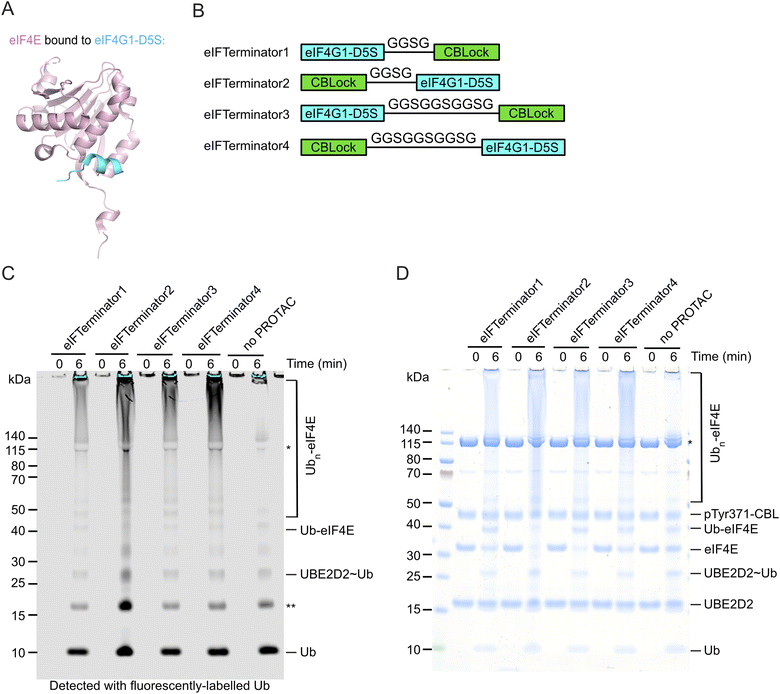 | ||
| Fig. 2 In vitro pTyr371-CBL-catalysed ubiquitination assay testing eIF4E ubiquitination with eIFTerminator1-4 (A) Crystal structure of eIF4E (pink) bound to eIF4G1-D5S peptide (cyan) (PDB ID: 4AZA).48 (B) Designs of eIF4E targeting PROTACs. The cartoons represent the peptide sequences of eIFTerminator1-4, which consist of eIF4G1-D5S and CBLock connected with a flexible GGSG or GGSGGSGGSG linker. (C) SDS-PAGE gel showing pTyr371-CBL-catalysed ubiquitination of eIF4E in the absence or presence of eIFTerminator1-4 detected with fluorescently-labelled Ub. Data are representative of three independent experiments. The single asterisk indicates the E1-Ub band and the double asterisk indicates a contaminant from the fluorescently-labelled Ub. (D) Coomassie-stained gel from C is shown. The single asterisk indicates the E1 band. | ||
To assess whether eIFTerminator1-4 could induce CBL-mediated eIF4E ubiquitination, we performed in vitro ubiquitination assays using recombinant UBA1, UBE2D2, fluorescent-labelled Ub, His-eIF4E, and pTyr371-CBL in the absence or presence of eIFTerminator1-4. All four PROTACs promoted poly-ubiquitination of eIF4E with the concomitant disappearance of the eIF4E band as compared to the control reaction lacking PROTAC (Fig. 2C and D). These findings suggest that eIFTerminator1-4 could bring CBL and eIF4E into proximity to facilitate CBL-mediated eIF4E ubiquitination.
eIFTerminator1-4 promote the formation of the CBL-eIF4E ternary complex
To investigate the binding kinetics and affinity of eIFTerminator1-4, surface plasmon resonance (SPR) analysis was performed. SPR measures changes in the refractive index near the sensor surface, where the analyte interacts with an immobilised ligand. This allows real-time monitoring of molecular interactions, providing quantitative binding affinity and kinetic parameters. First, we assessed whether eIFTerminator1-4 designs have any impact on CBL or eIF4E binding affinity. A three-fold dilution series of the eIFTerminator1-4, starting at 1 µM, was injected over immobilised CBL to assess the protein–protein interactions between eIFTerminator1-4 to CBL (SI Fig. 3A). The binding affinities of eIFTerminator1-4 were similar (KD: 42.9–64.6 nM; Table 1 and SI Fig. 3B) and comparable to that of CBLock (KD: 122 nM; Table 1 and SI Fig. 3B). Next, we examined the interactions between eIF4E and eIFTerminator1-4 by immobilising His-eIF4E on the SPR chip and injecting a dilution series of eIFTerminator1-4. The KD values ranged from 38.9 nM to 67.5 nM, and the binding affinity as well as kinetics were similar to that observed for the control ligand eIF4G1-D5S (Table 2 and SI Fig. 4A). These findings suggest that the design of eIFTerminator1-4 did not affect the binding affinity of the individual ligands for their respective binding partners, namely CBLock for CBL and eIF4G1-D5S for eIF4E.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding model was used to determine the KD (n = 3, mean ± SD, except CBLock is n = 2, mean ± SD)
1 binding model was used to determine the KD (n = 3, mean ± SD, except CBLock is n = 2, mean ± SD)
| Binary KD (nM) | |
|---|---|
| eIFTerminator1 | 64.6 ± 41.2 |
| eIFTerminator2 | 56.9 ± 25.4 |
| eIFTerminator3 | 42.9 ± 22.8 |
| eIFTerminator4 | 59.9 ± 27.0 |
| CBLock (positive ctrl) | 122 ± 4.2 |
| Binary KD (nM) | Ternary KD: CBL (nM) | |
|---|---|---|
| eIFTerminator1 | 54.0 ± 9.0 | 77.8 ± 9.7 |
| eIFTerminator2 | 67.5 ± 7.4 | 60.7 ± 22 |
| eIFTerminator3 | 39.5 ± 3.4 | 81.8 ± 8.2 |
| eIFTerminator4 | 60.6 ± 8.5 | 43.8 ± 6.0 |
| eIF4G1-D5S (positive ctrl) | 38.9 ± 6.5 | N/A |
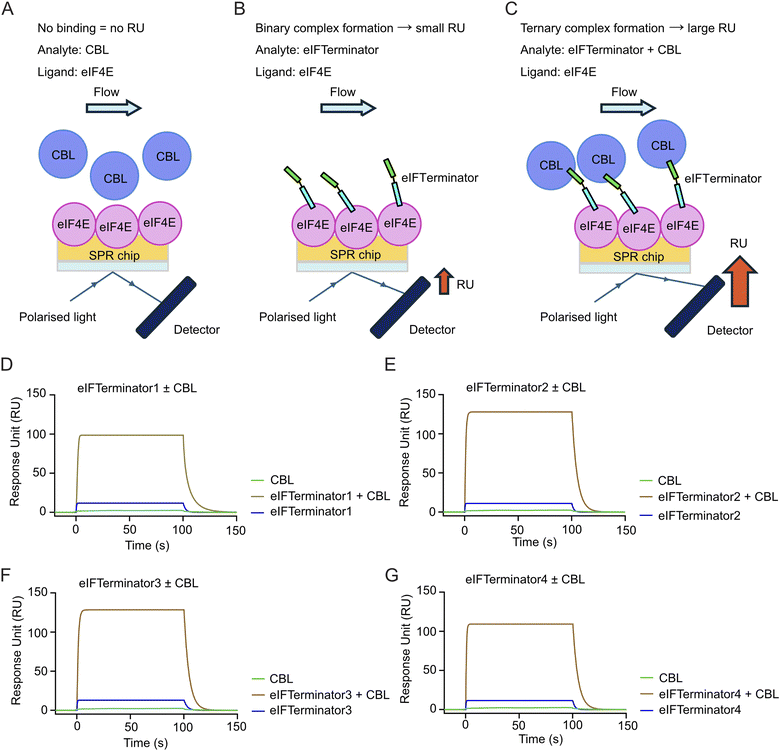
In SPR, the binding of a large analyte results in a greater change in refractive index, leading to a higher response unit (RU). By immobilising eIF4E on the SPR chip, we anticipated that binding of the eIFTerminator-CBL binary complex will cause a greater change in refractive index, leading to a larger RU compared to eIFTerminator alone if the ternary complex between eIF4E, eIFTerminator and CBL has assembled (Fig. 3A–C). To investigate the ternary complex formation, a saturating concentration of CBL (1 µM), which is over five-fold higher than the KD for the CBL-CBLock interaction, was included with the eIFTerminator1-4 analytes and flowed over immobilised eIF4E. Addition of CBL alone, in the absence of eIFTerminators, caused little change in RU, indicating that CBL did not interact with eIF4E (Fig. 3D–G). When eIFTerminator1-4 was injected alone, there was an increase of 10–15 RU. In contrast, in the presence of CBL, there was an increase of 100-120 RU (Fig. 3D–G), confirming ternary complex formation. In addition, since positive cooperativity in ternary complexes has been shown to influence degradation efficiency,12 we compared the eIFTeminator1-4 binding affinities for eIF4E in binary complexes to ternary complexes in the presence of 1 µM CBL. The KD values and binding kinetics were similar (Table 2 and SI Fig. 4B), suggesting no cooperative effect. Together, these findings demonstrate that eIFTerminators bind both CBL and eIF4E to form the ternary complex.
CP-eIFTerminator4 reduces the level of eIF4E in a dose-dependent manner
To examine whether eIFTerminator1-4 could induce eIF4E degradation in cells, we incorporated a 16-mer penetratin peptide sequence from the Drosophila Antennapedia homeodomain52 at their N-terminus to facilitate cell entry. Additionally, an N-terminal FITC label was included to monitor cell uptake. These modified peptidic PROTACs are referred to as cell-penetrating eIFTerminator1-4 (CP-eIFTerminator1-4). Among them, CP-eIFTerminator1, CP-eIFTerminator3, and CP-eIFTerminator4 readily dissolved in water or DMSO, whereas CP-eIFTerminator2 was insoluble and therefore not pursued further.A key consideration when characterising PROTACs in cells is cytotoxicity, as compounds that significantly reduce cell viability within a few hours may exert off-target cytotoxic effects rather than on-target degradation. To assess this, cell viability assays were performed after incubating HeLa cells with CP-eIFTerminators at concentrations ranging from 0.16–24.4 µM for 3 h and 8 h (SI Fig. 5A and B). The parental cell-penetrating peptides CP-CBLock and CP-eIF4G1-D5S were included as controls. The assay showed that CP-eIF4G1-D5S exhibited cytotoxicity at concentrations of 16.10 µM or higher, reducing viable cells to as low as 35% at 3 h and 25% at 8 h post-peptide treatment, whereas lower concentrations (0.16–16.10 µM) had no apparent effect on cell viability (SI Fig. 5A and B). In contrast, CP-CBLock and CP-eIFTerminator4 did not affect cell viability under the tested conditions. However, CP-eIFTerminator1 and CP-eIFTerminator3 showed mild cytotoxic effects at higher concentrations (16.10–24.4 µM). Although it remains unclear why the eIF4E warhead eIF4G1-D5S exhibited cytotoxicity at higher concentrations, it is intriguing that linking CBLock to its N-terminus in CP-eIFTerminator4 mitigates the cytotoxicity.
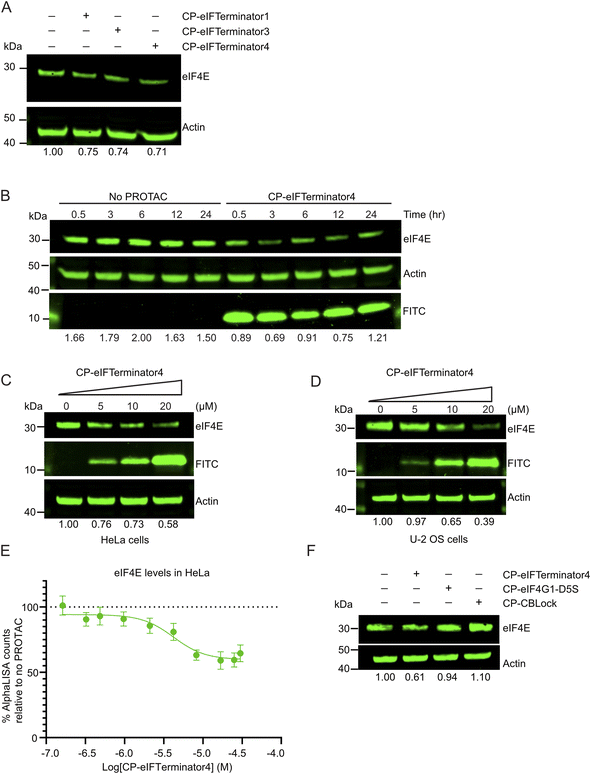
Based on these findings, we treated HeLa cells with 10 µM of CP-eIFTerminator1, CP-eIFTerminator3 or CP-eIFTerminator4 and assessed the effects on the endogenous levels of eIF4E. HeLa cells were chosen, due to their high EIF4E and CBL mRNA expression (The Human Protein Atlas, https://www.proteinatlas.org/). Treatment with CP-eIFTerminator1, CP-eIFTerminator3 or CP-eIFTerminator4 resulted in a similar extent of reduction in eIF4E protein levels, namely a reduction between 24.6% and 28.7%, compared to the no PROTAC treatment control (Fig. 4A). CP-eIFTerminator4 was selected for further testing, as it did not cause acute cytotoxicity at 3-h and 8-h post treatment (SI Fig. 5A and B), which enabled the assessment of its potency at higher concentrations.
Next, a time course experiment was performed to assess whether the reduction in eIF4E levels was sustained over time. HeLa cells were incubated with 20 µM CP-eIFTerminator4 and examined at selected time points over a 24-h period. Western blot analysis showed minimal changes in FITC levels over the 24-h period (Fig. 4B). CP-eIFTerminator4 readily entered cells as soon as 30 min and caused a reduction in eIF4E protein levels between 0.5–12 h. Although eIF4E levels gradually increased by 24 h, the level was still lower in CP-eIFTerminator4-treated cells compared to untreated cells. Since levels of eIF4E were most reduced at 3 h following CP-eIFTerminator4 treatment, we opted to further characterise this PROTAC at the 3-h time point.
Two cancer cell lines, HeLa and U-2 OS, were chosen based on their high mRNA expression of CBL and EIF4E (The Human Protein Atlas, https://www.proteinatlas.org/), in order to validate that CP-eIFTerminator4 exerts its effect in a concentration-dependent manner. HeLa cells were treated with varying concentrations of CP-eIFTerminator4 and harvested after 3 h for immunoblotting analysis. A concentration-dependent decrease in eIF4E levels was observed (Fig. 4C). Similarly, U-2 OS cells treated under comparable conditions showed a reduction in intracellular eIF4E levels as the concentration of CP-eIFTerminator4 was increased (Fig. 4D). Since the two cancer cell lines exhibited similar results, we selected HeLa cells for further characterisation.
Next, we examined the degradation efficiency of CP-eIFTerminator4 in HeLa cells using AlphaLISA. Cells were treated with a titration series of CP-eIFTerminator4 (0.49–30.07 µM) for 3 h, followed by AlphaLISA analysis to quantify intracellular eIF4E levels. CP-eIFTerminator4 showed a half-maximal degradation concentration (DC50) of approximately 4.66 µM and a maximum degradation (Dmax) of 39.4% (Fig. 4E). Since it was not possible to prepare eIF4E-knockout HeLa cells as a negative control, background subtraction was done using empty wells instead. For this reason, it was unclear if the true extent of degradation was under-represented by the measured Dmax. In comparison, quantification of eIF4E bands on immunoblots from four independent samples treated with 20 µM CP-eIFTerminator4 for 3 h yielded an average Dmax of 44.0% (SD ± 18.6%). The AlphaLISA and western blot data were similar, and the extent of degradation varied more across experiments in the latter case. Nonetheless, the lack of an eIF4E knock-out negative control is unlikely to significantly affect the determination of the DC50.
To determine whether the degradation of eIF4E by CP-eIFTerminator4 is driven by an induced-proximity mechanism rather than by the individual warheads, we tested CP-eIFTerminator4, CP-eIF4G1-D5S, and CP-CBLock separately in HeLa cells (Fig. 4F). Since CP-eIF4G1-D5S affected cell viability at concentrations above 16 µM while CP-CBLock and CP-eIFTerminator4 did not affect cell viability (SI Fig. 5A and B), all compounds were tested at 10 µM. Under this condition, only CP-eIFTerminator4 significantly reduced eIF4E (Fig. 4F). This confirms that CP-eIFTerminator4 functions as a PROTAC and that the individual warheads do not trigger eIF4E degradation.
Degradation of eIF4E by an artificial induced-proximity approach using a dTAG system has been shown to be feasible.53 However, degradation of endogenous eIF4E via a PROTAC approach using CRBN and VHL ligands has remained challenging; despite achieving cell entry and ternary complex formation, these PROTACs failed to induce eIF4E degradation.46,47 Our work presents the first case in which eIF4E is targeted for degradation in a PROTAC system.
CP-eIFTerminator4 recruits CBL and induces eIF4E degradation using two cellular degradation pathways
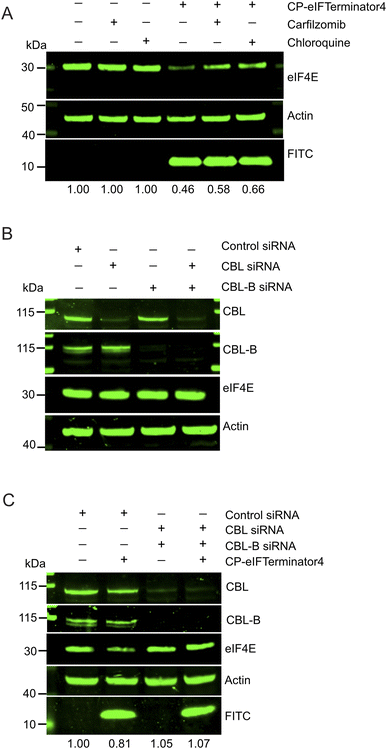
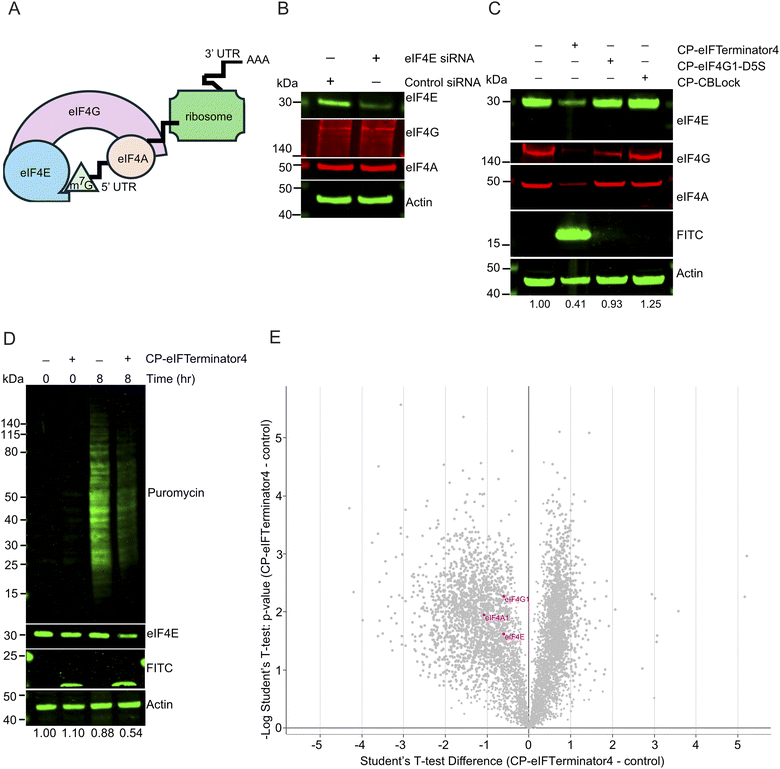
Since degradation was observed in the initial hours following CP-eIFTerminator4 treatment (Fig. 4B), the next step was to investigate the mechanism of degradation. To determine which degradation pathway is responsible for eIF4E reduction, HeLa cells were treated with 20 µM CP-eIFTerminator4 alongside the proteasome inhibitor carfilzomib or the lysosomal inhibitor chloroquine. Carfilzomib, an irreversible proteasomal inhibitor, was pre-incubated at a concentration of 100 nM, while chloroquine was used at a concentration of 50 µM for 4 h prior to adding CP-eIFTerminator4 for 3 h. Inhibition of protein degradation via either proteasome or lysosome led to eIF4E stabilisation (Fig. 5A).We previously showed that CBLock binds to both CBL and CBL-B, as both CBL proteins share identical sequences at the TKBD substrate-binding site targeted by CBLock.38 To determine whether eIF4E degradation by CP-eIFTerminator4 is mediated by the CBL proteins, we performed siRNA knockdown of CBL and CBL-B in HeLa cells. We confirmed that CBL siRNA and CBLB siRNA were able to deplete CBL or CBL-B, respectively, in HeLa cells (Fig. 5B). In addition, knockdown of CBL and/or CBL-B had no effect on eIF4E protein levels. As expected from a peptide that recruits CBL, co-treatment of CBL and CBLB siRNA inhibited CP-eIFTerminator4-mediated eIF4E degradation (Fig. 5C). These findings demonstrate that CBL proteins are required for CP-eIFTerminator4-mediated degradation of eIF4E and suggest that CBL and CBL-B can facilitate eIF4E degradation through both lysosomal and proteasomal pathways.
Unlike lysosome-targeting chimeras (LYTACs), which rely on lysosomal targeting receptors such as the cation-independent mannose-6-phosphate receptor or the asialoglycoprotein receptor,54 or autophagy-targeting chimeras (AUTACs), which induces autophagosome formation through S-guanylation,55 our study, to our knowledge, represents the first example of a PROTAC hijacking a cytoplasmic E3 to degrade a cytoplasmic neo-substrate via the lysosomal pathway. It remains unclear which types of Ub chains are assembled on the neo-substrate by CBL in cells to direct it toward either lysosomal or proteasomal degradation. However, the ability to engage both degradation pathways may offer a distinct advantage by increasing the likelihood of effective neo-substrate clearance. Interestingly, a recent study showed that the natural CBL substrate Lyn kinase was degraded by both CBL and CBL-B upon treatment with the Lyn-targeting inhibitor SI-3 (Scholes, Bertoni56). This degradation involved both lysosomal and proteasomal pathways. Together, these results suggest that CBL proteins can mediate substrate degradation through distinct mechanisms depending on cellular context and pharmacological cues.
CP-eIFTerminator4 disrupts the eIF4F complex and translation
Since the eukaryotic cap-dependent translation complex eIF4F consists of the RNA helicase eIF4A1, mRNA cap-binder eIF4E, and the scaffold protein eIF4G1 (Fig. 6A), we sought to assess whether the depletion of eIF4E also affects the protein levels of eIF4A1 and the scaffold eIF4G1 concomitantly. Consistent with previous reports,45,57EIF4E siRNA treatment in HeLa cells for 96 h did not perturb eIF4A1 or eIF4G1 levels (Fig. 6B). In contrast, 3 h of CP-eIFTerminator4 treatment in HeLa cells led to a reduction in eIF4A1, eIF4E, and eIF4G1 levels (Fig. 6C). The warheads, CP-eIF4G1-D5S or CP-CBLock, had minimal effects on eIF4A, eIF4E, and eIF4G levels (Fig. 6C). Combined, it appears that CP-eIFTerminator4 acts on the eIF4F complex via a unique mechanism that is distinct from siRNA-mediated knockdown of eIF4E, or occupancy-driven inhibition mediated by eIF4G1-D5S.To gain insight into the effect on translation as a result of depletion of the core eIF4F components, HeLa cells were pre-treated with or without CP-eIFTerminator4 for 7 h, followed by addition of puromycin for 1 h (Fig. 6D). Puromycin is a tyrosyl-tRNA mimetic that inhibits translation, but it is widely used to monitor the rate of protein synthesis.58,59 It binds to the ribosome during translation and is incorporated into the nascent polypeptide chain, causing premature termination and the release of puromycin-labelled peptides, which can be detected using an anti-puromycin antibody. Treatment with CP-eIFTerminator4 for 8 h was found to reduce polypeptide formation by 61.5% (SD ± 23.2, n = 3) (Fig. 6D).
As the puromycin incorporation assay suggested that translation is impaired in HeLa cells treated with CP-eIFTerminator4, we next performed a quantitative expression proteomics experiment to examine changes in the global proteome. Three independent replicates of HeLa cell treated with CP-eIFTerminator4 for 3 h were compared to untreated controls. Consistent with the puromycin incorporation data (Fig. 6D), CP-eIFTerminator4 treatment induced a widespread change in the proteome (Fig. 6E). The asymmetric volcano plot highlights a stronger trend towards downregulation, and many proteins were reduced to a greater extent than eIF4E. Proteomic analysis revealed that 1207 proteins were upregulated and 1779 were downregulated (SI Data 1). Notably, eIF4A1, eIF4E, and eIF4G1 showed a statistically significant reduction as a result of CP-eIFTerminator4 treatment. These findings suggest that perturbation of eIF4E and the eIF4F complex results in extensive downstream proteome remodelling.
Further KEGG and Reactome pathway analyses of the proteomics data revealed that enzymes involved in tRNA aminoacylation, the pentose phosphate pathway, and fructose and mannose metabolism were downregulated. In particular, 11 out of the 12 detected aminoacyl-tRNA synthetases were decreased, raising the possibility that their reduced levels may further compromise protein synthesis (SI Data 1).
Given the essential role of eIF4E in cap-dependent translation, our proteomics data do not allow a clear distinction between direct and indirect effects of CP-eIFTerminator4 on the proteome. It also remains to be determined how eIF4A and eIF4G are downregulated by CP-eIFTerminator4 treatment. Nevertheless, in light of previous studies such as the 30% reduction in [35S]methionine incorporation into the newly synthesised proteins despite an 80–90% decrease in eIF4E levels in rabbit reticulocytes,60 and the largely unchanged global translation in eIF4E haplo-sufficient mice,45 our results demonstrate that targeted degradation of eIF4E and other components of eIF4F complex by CP-eIFTerminator4 produces a substantial impact on translation.
Conclusion
Targeted protein degradation (TPD) via PROTACs has expanded rapidly over the past decade. While an increasing number of E3s have been exploited for PROTAC application, there remains a strong demand to diversify the E3 toolbox, particularly to address undruggable substrates, tissue-specific E3 expression limitations, and emerging drug resistance.In this study, we investigated the potential of the previously uncharacterised E3, CBL, as a degrader E3 to target neo-substrates for ubiquitination and degradation. We designed peptidic PROTACs, eIFTerminators, by linking CBLock to an eIF4E-binding peptide, eIF4G-D5S. We demonstrated that eIFTerminators promote ternary complex formation between CBL, eIFTerminator, and eIF4E, resulting in CBL-mediated eIF4E ubiquitination in vitro. In cells, eIFTerminator4 rapidly targets eIF4E for degradation via both lysosomal and proteasomal pathways in a manner dependent on CBL and CBL-B. Unexpectedly, eIFTerminator4 also led to the depletion of eIF4A and eIF4G, resulting in a marked reduction in global protein translation. Using a range of biochemical, biophysical, and cell-based assays, we have established CBL as a viable degrader E3 for TPD applications.
Unlike siRNA, which typically requires 24–72 h to show results, or CRISPR-Cas9, which involves months of intensive effort to knock out a gene, CP-eIFTerminator4 achieved intracellular eIF4E degradation in 30 min simply by adding the peptide to cell culture media with EGF. Furthermore, CP-eIFTermiantor4 did not acutely affect cell viability. Thus, our eIF4E PROTACs may be a useful chemical tool to study essential survival genes with minimal disruption of cellular systems.
Importantly, our CBL PROTAC was able to degrade an oncoprotein that is regarded undruggable with other E3s or inhibitors,46,47,53,61,62 highlighting its potential to expand the scope of PROTAC-based anti-cancer therapies. Future studies are required to elucidate the precise mechanism of action of CP-eIFTerminator4. Nonetheless, designing a PROTAC that targets the entire eIF4F complex for degradation may represent an interesting avenue for cancer therapy. Furthermore, a small molecule targeting the CBL's substrate-binding site, similar to eIFTerminator4, has recently been reported.63 In contrast to CBL ligands that bind the LHR site and inhibit CBL E3 activity,64,65 this compound does not interfere with CBL's E3 activity. Combined with previously described small-molecule inhibitors of eIF4E,53,62,66–68 it could enable the development of next-generation CBL-based eIF4E PROTACs with improved potency.
Author contributions
A. T. W, A. H., and D. T. H. conceived the project. A. T. W and D. T. H. wrote the manuscript. A. T. W, L. B, T. S. generated constructs and purified proteins. A. T. W. performed all experiments under the guidance and scientific input of L. B., T. S., T. S., A. M.-J., J. T, A. G, M. B, A. K. H. and D. T. H. S. L. performed and analysed mass spectrometry data. A. T. W., J. T., and A. G. analysed SPR data. All authors commented on the manuscript.Conflicts of interest
A. T. W., L. B., T. S., T. S., S. L., M. B., D. T. H. are current employees of the Cancer Research UK Scotland Institute. The above authors declare no competing interests. A. M.-J., J. T, A. G., and A. K. H. are employees of AstraZeneca and have stock ownership and/or stock options and commercial interests in the company.Data availability
Raw mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org/) via the PRIDE partner repository with the dataset identifiers PRIDE: PXD063494 and PXD071639. All data generated during this study are included in this article or are available from the corresponding author upon request. All raw gels, western blot images, AlphaLISA data, and SPR data generated during this study have been deposited to Mendeley (https://data.mendeley.com/preview/ptcdwrhj2x?a=b14d0213-7a88-4820-bd0c-2506dfc23dcc).Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5sc06141e.
Acknowledgements
We would like to thank Dr Feroj Ahmed for the scientific discussion and the late Dr Wei Zhang for his instrumental work on the CBLock peptide. We would also like to express our gratitude to Dr Catherine Winchester (Cancer Research UK Scotland Institute) for critically reviewing the manuscript. This work was supported by a BBSRC-CTP in collaboration with AstraZeneca (BB/V509413/1), CRUK Scotland Institute's Advanced Technology Facilities (A31287), and Cancer Research UK core funding to D. T. H. (A29256). M. B. and T. S. were supported by CRUK core funding (A29252) and BBSRC grants (BB/Y004248/1 and DRCQQR-Jun24/100002) awarded to M. B. T. S. was funded by Beatson Cancer Charity (24-25-079).References
- M. Bekes, D. R. Langley and C. M. Crews, PROTAC targeted protein degraders: the past is prologue, Nat. Rev. Drug Discov., 2022, 21(3), 181–200 CrossRef CAS PubMed.
- K. M. Sakamoto, K. B. Kim, A. Kumagai, F. Mercurio, C. M. Crews and R. J. Deshaies, Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation, Proc. Natl. Acad. Sci. U. S. A., 2001, 98(15), 8554–8559 CrossRef CAS PubMed.
- X. Li and Y. Song, Proteolysis-targeting chimera (PROTAC) for targeted protein degradation and cancer therapy, J. Hematol. Oncol., 2020, 13(1), 50 CrossRef CAS PubMed.
- S. Zeng, Y. Ye, H. Xia, J. Min, J. Xu and Z. Wang, et al., Current advances and development strategies of orally bioavailable PROTACs, Eur. J. Med. Chem., 2023, 261, 115793 CrossRef CAS PubMed.
- M. Zengerle, K. H. Chan and A. Ciulli, Selective Small Molecule Induced Degradation of the BET Bromodomain Protein BRD4, ACS Chem. Biol., 2015, 10(8), 1770–1777 CrossRef CAS PubMed.
- A. C. Lai, M. Toure, D. Hellerschmied, J. Salami, S. Jaime-Figueroa and E. Ko, et al., Modular PROTAC Design for the Degradation of Oncogenic BCR-ABL, Angew Chem. Int. Ed. Engl., 2016, 55(2), 807–810 CrossRef CAS PubMed.
- D. P. Bondeson, B. E. Smith, G. M. Burslem, A. D. Buhimschi, J. Hines and S. Jaime-Figueroa, et al., Lessons in PROTAC Design from Selective Degradation with a Promiscuous Warhead, Cell Chem. Biol., 2018, 25(1), 78–87 CrossRef CAS PubMed.
- R. Shirasaki, G. M. Matthews, S. Gandolfi, S. R. de Matos, D. L. Buckley and V. J. Raja, et al., Functional Genomics Identify Distinct and Overlapping Genes Mediating Resistance to Different Classes of Heterobifunctional Degraders of Oncoproteins, Cell Rep., 2021, 34(1), 108532 Search PubMed.
- L. Zhang, B. Riley-Gillis, P. Vijay and Y. Shen, Acquired Resistance to BET-PROTACs (Proteolysis-Targeting Chimeras) Caused by Genomic Alterations in Core Components of E3 Ligase Complexes, Mol. Cancer Ther., 2019, 18(7), 1302–1311 CrossRef CAS PubMed.
- P. Ottis, C. Palladino, P. Thienger, A. Britschgi, C. Heichinger and M. Berrera, et al., Cellular Resistance Mechanisms to Targeted Protein Degradation Converge Toward Impairment of the Engaged Ubiquitin Transfer Pathway, ACS Chem. Biol., 2019, 14(10), 2215–2223 CAS.
- W. Li, M. H. Bengtson, A. Ulbrich, A. Matsuda, V. A. Reddy and A. Orth, et al., Genome-wide and functional annotation of human E3 ubiquitin ligases identifies MULAN, a mitochondrial E3 that regulates the organelle's dynamics and signaling, PLoS One, 2008, 3(1), e1487 CrossRef PubMed.
- M. S. Gadd, A. Testa, X. Lucas, K. H. Chan, W. Chen and D. J. Lamont, et al., Structural basis of PROTAC cooperative recognition for selective protein degradation, Nat. Chem. Biol., 2017, 13(5), 514–521 CrossRef CAS PubMed.
- M. Schroder, M. Renatus, X. Liang, F. Meili, T. Zoller and S. Ferrand, et al., DCAF1-based PROTACs with activity against clinically validated targets overcoming intrinsic- and acquired-degrader resistance, Nat. Commun., 2024, 15(1), 275 Search PubMed.
- E. J. McMahon, A. G. Cioffi, P. R. Visperas, Y. Lin, M. Shaghafi and C. M. Daczkowski, et al., Structural basis for DCAF2 as a novel E3 ligase for PROTAC-mediated targeted protein degradation, Structure, 2025, 33(12), 8 CrossRef PubMed.
- G. Xue, J. Xie, M. Hinterndorfer, M. Cigler, L. Dotsch and H. Imrichova, et al., Discovery of a Drug-like, Natural Product-Inspired DCAF11 Ligand Chemotype, Nat. Commun., 2023, 14(1), 7908 CrossRef CAS PubMed.
- D. E. Bussiere, L. Xie, H. Srinivas, W. Shu, A. Burke and C. Be, et al., Structural basis of indisulam-mediated RBM39 recruitment to DCAF15 E3 ligase complex, Nat. Chem. Biol., 2020, 16(1), 15–23 CrossRef CAS PubMed.
- O. Hsia, M. Hinterndorfer, A. D. Cowan, K. Iso, T. Ishida and R. Sundaramoorthy, et al., Targeted protein degradation via intramolecular bivalent glues, Nature, 2024, 627(8002), 204–211 CrossRef CAS PubMed.
- M. Meyers, S. Cismoski, A. Panidapu, B. Chie-Leon and D. K. Nomura, Targeted Protein Degradation through Recruitment of the CUL4 Complex Adaptor Protein DDB1, ACS Chem. Biol., 2024, 19(1), 58–68 CrossRef CAS PubMed.
- C. Kagiou, J. A. Cisneros, J. Farnung, J. Liwocha, F. Offensperger and K. Dong, et al., Alkylamine-tethered molecules recruit FBXO22 for targeted protein degradation, Nat. Commun., 2024, 15(1), 5409 CrossRef CAS PubMed.
- Y. Li, K. Bao, J. Sun, R. Ge, Q. Zhang and B. Zhang, et al., Design of PROTACs utilizing the E3 ligase GID4 for targeted protein degradation, Nat. Struct. Mol. Biol., 2025, 32, 1825–1837 CrossRef CAS PubMed.
- N. Shibata, N. Miyamoto, K. Nagai, K. Shimokawa, T. Sameshima and N. Ohoka, et al., Development of protein degradation inducers of oncogenic BCR-ABL protein by conjugation of ABL kinase inhibitors and IAP ligands, Cancer Sci., 2017, 108(8), 1657–1666 CrossRef CAS PubMed.
- G. Du, J. Jiang, N. J. Henning, N. Safaee, E. Koide and R. P. Nowak, et al., Exploring the target scope of KEAP1 E3 ligase-based PROTACs, Cell Chem. Biol., 2022, 29(10), 1470–81e31 CrossRef CAS PubMed.
- C. M. Hickey, K. M. Digianantonio, K. Zimmermann, A. Harbin, C. Quinn and A. Patel, et al., Co-opting the E3 ligase KLHDC2 for targeted protein degradation by small molecules, Nat. Struct. Mol. Biol., 2024, 31(2), 311–322 CrossRef CAS PubMed.
- P. Lu, Y. Cheng, L. Xue, X. Ren, X. Xu and C. Chen, et al., Selective degradation of multimeric proteins by TRIM21-based molecular glue and PROTAC degraders, Cell, 2024, 187, 7126–7142 CrossRef CAS PubMed.
- J. Hines, S. Lartigue, H. Dong, Y. Qian and C. M. Crews, MDM2-Recruiting PROTAC Offers Superior, Synergistic Antiproliferative Activity via Simultaneous Degradation of BRD4 and Stabilization of p53, Cancer Res., 2019, 79(1), 251–262 CrossRef CAS PubMed.
- Z. Deng, J. Catlett, Y. Lee, Q. Wu, Z. Xu and L. Xie, et al., Harnessing the SPOP E3 Ubiquitin Ligase via a Bridged Proteolysis Targeting Chimera (PROTAC) Strategy for Targeted Protein Degradation, J. Med. Chem., 2025, 68(8), 8634–8647 CrossRef CAS PubMed.
- Z. Yang, M. Chen, R. Ge, P. Zhou, W. Pan and J. Song, et al., Identification of a non-inhibitory aptameric ligand to CRL2(ZYG11B) E3 ligase for targeted protein degradation, Nat. Commun., 2025, 16(1), 2494 CrossRef CAS PubMed.
- M. H. H. Schmidt and I. Dikic, The Cbl interactome and its functions, Nat. Rev. Mol. Cell Biol., 2005, 6(12), 907–918 CrossRef CAS PubMed.
- P. Castro-Sanchez, A. R. Teagle, S. Prade and R. Zamoyska, Modulation of TCR Signaling by Tyrosine Phosphatases: From Autoimmunity to Immunotherapy, Front. Cell Dev. Biol., 2020, 8, 608747 CrossRef PubMed.
- G. Levkowitz, H. Waterman, E. Zamir, Z. Kam, S. Oved and W. Y. Langdon, et al., c-Cbl/Sli-1 regulates endocytic sorting and ubiquitination of the epidermal growth factor receptor, Genes Dev., 1998, 12(23), 3663–3674 CrossRef CAS PubMed.
- M. Javadi, T. D. Richmond, K. Huang and D. L. Barber, CBL linker region and RING finger mutations lead to enhanced granulocyte-macrophage colony-stimulating factor (GM-CSF) signaling via elevated levels of JAK2 and LYN, J. Biol. Chem., 2013, 288(27), 19459–19470 CrossRef CAS PubMed.
- H. Dou, L. Buetow, A. Hock, G. J. Sibbet, K. H. Vousden and D. T. Huang, Structural basis for autoinhibition and phosphorylation-dependent activation of c-Cbl, Nat. Struct. Mol. Biol., 2012, 19(2), 184–192 CrossRef CAS PubMed.
- L. Buetow, G. Tria, S. F. Ahmed, A. Hock, H. Dou and G. J. Sibbet, et al., Casitas B-lineage lymphoma linker helix mutations found in myeloproliferative neoplasms affect conformation, BMC Biol., 2016, 14(1), 76 CrossRef PubMed.
- G. E. Winter, D. L. Buckley, J. Paulk, J. M. Roberts, A. Souza and S. Dhe-Paganon, et al., Drug development. Phthalimide conjugation as a strategy for in vivo target protein degradation, Science, 2015, 348(6241), 1376–1381 CrossRef CAS PubMed.
- D. P. Bondeson, A. Mares, I. E. Smith, E. Ko, S. Campos and A. H. Miah, et al., Catalytic in vivo protein knockdown by small-molecule PROTACs, Nat. Chem. Biol., 2015, 11(8), 611–617 CrossRef CAS PubMed.
- C. Galdeano, M. S. Gadd, P. Soares, S. Scaffidi, I. Van Molle and I. Birced, et al., Structure-guided design and optimization of small molecules targeting the protein–protein interaction between the von Hippel-Lindau (VHL) E3 ubiquitin ligase and the hypoxia inducible factor (HIF) alpha subunit with in vitro nanomolar affinities, J. Med. Chem., 2014, 57(20), 8657–8663 CrossRef CAS PubMed.
- Human Protein Atlas, The open access resource for human proteins, 2025, Available from: https://www.proteinatlas.org/ Search PubMed.
- S. F. Ahmed, J. Anand, W. Zhang, L. Buetow, L. Rishi and L. Mitchell, et al., Locking CBL TKBD in its native conformation presents a novel therapeutic opportunity in mutant CBL-dependent leukemia, Mol. Ther., 2025, 33, 3624–3643 CrossRef CAS PubMed.
- J. Pelletier, J. Graff, D. Ruggero and N. Sonenberg, Targeting the eIF4F translation initiation complex: a critical nexus for cancer development, Cancer Res., 2015, 75(2), 250–263 CrossRef CAS PubMed.
- S. Avdulov, S. Li, V. Michalek, D. Burrichter, M. Peterson and D. M. Perlman, et al., Activation of translation complex eIF4F is essential for the genesis and maintenance of the malignant phenotype in human mammary epithelial cells, Cancer Cell, 2004, 5(6), 553–563 CrossRef CAS PubMed.
- F. Shi, Y. Len, Y. Gong, R. Shi, X. Yang and D. Naren, et al., Ribavirin Inhibits the Activity of mTOR/eIF4E, ERK/Mnk1/eIF4E Signaling Pathway and Synergizes with Tyrosine Kinase Inhibitor Imatinib to Impair Bcr-Abl Mediated Proliferation and Apoptosis in Ph+ Leukemia, PLoS One, 2015, 10(8), e0136746 Search PubMed.
- D. Hanahan, Hallmarks of Cancer: New Dimensions, Cancer Discov., 2022, 12(1), 31–46 CrossRef CAS PubMed.
- J. Wan, F. Shi, Z. Xu and M. Zhao, Knockdown of eIF4E suppresses cell proliferation, invasion and enhances cisplatin cytotoxicity in human ovarian cancer cells, Int. J. Oncol., 2015, 47(6), 2217–2225 CrossRef CAS PubMed.
- J. R. Graff, B. W. Konicek, T. M. Vincent, R. L. Lynch, D. Monteith and S. N. Weir, et al., Therapeutic suppression of translation initiation factor eIF4E expression reduces tumor growth without toxicity, J. Clin. Invest., 2007, 117(9), 2638–2648 CrossRef CAS PubMed.
- M. L. Truitt, C. S. Conn, Z. Shi, X. Pang, T. Tokuyasu and A. M. Coady, et al., Differential Requirements for eIF4E Dose in Normal Development and Cancer, Cell, 2015, 162(1), 59–71 CrossRef CAS PubMed.
- T. Kaur, A. Menon and A. L. Garner, Synthesis of 7-benzylguanosine cap-analogue conjugates for eIF4E targeted degradation, Eur. J. Med. Chem., 2019, 166, 339–350 CrossRef CAS PubMed.
- P. D. Fischer, E. Papadopoulos, J. M. Dempersmier, Z. F. Wang, R. P. Nowak and K. A. Donovan, et al., A biphenyl inhibitor of eIF4E targeting an internal binding site enables the design of cell-permeable PROTAC-degraders, Eur. J. Med. Chem., 2021, 219, 113435 CrossRef CAS PubMed.
- W. Zhou, S. T. Quah, C. S. Verma, Y. Liu, D. P. Lane and C. J. Brown, Improved eIF4E binding peptides by phage display guided design: plasticity of interacting surfaces yield collective effects, PLoS One, 2012, 7(10), e47235 CrossRef CAS PubMed.
- C. K. Kassenbrock and S. M. Anderson, Regulation of ubiquitin protein ligase activity in c-Cbl by phosphorylation-induced conformational change and constitutive activation by tyrosine to glutamate point mutations, J. Biol. Chem., 2004, 279(27), 28017–28027 CrossRef CAS PubMed.
- Y. Zhang, A. Wolf-Yadlin, P. L. Ross, D. J. Pappin, J. Rush and D. A. Lauffenburger, et al., Time-resolved mass spectrometry of tyrosine phosphorylation sites in the epidermal growth factor receptor signaling network reveals dynamic modules, Mol. Cell. Proteomics, 2005, 4(9), 1240–1250 CrossRef CAS PubMed.
- S. Gruner, D. Peter, R. Weber, L. Wohlbold, M. Y. Chung and O. Weichenrieder, et al., The Structures of eIF4E-eIF4G Complexes Reveal an Extended Interface to Regulate Translation Initiation, Mol. Cell, 2016, 64(3), 467–479 CrossRef PubMed.
- S. Futaki, Membrane-permeable arginine-rich peptides and the translocation mechanisms, Adv. Drug Deliv. Rev., 2005, 57(4), 547–558 CrossRef CAS PubMed.
- S. Y. Sharp, M. Martella, S. D'Agostino, C. I. Milton, G. Ward and A. J. Woodhead, et al., Integrating fragment-based screening with targeted protein degradation and genetic rescue to explore eIF4E function, Nat. Commun., 2024, 15(1), 10037 CrossRef CAS PubMed.
- S. M. Banik, K. Pedram, S. Wisnovsky, G. Ahn, N. M. Riley and C. R. Bertozzi, Lysosome-targeting chimaeras for degradation of extracellular proteins, Nature, 2020, 584(7820), 291–297 CrossRef CAS PubMed.
- D. Takahashi, J. Moriyama, T. Nakamura, E. Miki, E. Takahashi and A. Sato, et al., AUTACs: Cargo-Specific Degraders Using Selective Autophagy, Mol. Cell, 2019, 76(5), 797–810 CrossRef CAS PubMed.
- N. S. Scholes, M. Bertoni, A. Comajuncosa-Creus, K. Kladnik, F. Frommelt and M. Hinterndorfer, et al., Inhibitor-induced supercharging of kinase turnover via endogenous proteolytic circuits, bioRxiv, 2024, 2024.07.10.602881 Search PubMed.
- A. Yanagiya, E. Suyama, H. Adachi, Y. V. Svitkin, P. Aza-Blanc and H. Imataka, et al., Translational homeostasis via the mRNA cap-binding protein, eIF4E, Mol. Cell, 2012, 46(6), 847–858 Search PubMed.
- D. Nathans, Puromycin Inhibition of Protein Synthesis: Incorporation of Puromycin into Peptide Chains, Proc. Natl. Acad. Sci. U. S. A., 1964, 51(4), 585–592 Search PubMed.
- E. K. Schmidt, G. Clavarino, M. Ceppi and P. Pierre, SUnSET, a nonradioactive method to monitor protein synthesis, Nat. Methods, 2009, 6(4), 275–277 CrossRef CAS PubMed.
- M. Rau, T. Ohlmann, S. J. Morley and V. M. Pain, A reevaluation of the cap-binding protein, eIF4E, as a rate-limiting factor for initiation of translation in reticulocyte lysate, J. Biol. Chem., 1996, 271(15), 8983–8990 CrossRef CAS PubMed.
- C. A. Lucchesi, J. Zhang, M. Gao, J. Shaw and X. Chen, Identification of a First-in-Class Small-Molecule Inhibitor of the EIF4E-RBM38 Complex That Enhances Wild-type TP53 Protein Translation for Tumor Growth Suppression, Mol. Cancer Ther., 2023, 22(6), 726–736 CrossRef CAS PubMed.
- N. J. Moerke, H. Aktas, H. Chen, S. Cantel, M. Y. Reibarkh and A. Fahmy, et al., Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G, Cell, 2007, 128(2), 257–267 CrossRef CAS PubMed.
- J. Liang, M. J. Lambrecht, T. L. Arenzana, S. Aubert-Nicol, L. Bao and F. Broccatelli, et al., Optimization of a Novel DEL Hit That Binds in the Cbl-b SH2 Domain and Blocks Substrate Binding, ACS Med. Chem. Lett., 2024, 15(6), 864–872 CrossRef CAS PubMed.
- A. M. Mfuh, J. A. Boerth, G. Bommakanti, C. Chan, A. J. Chinn and E. Code, et al., Discovery, Optimization, and Biological Evaluation of Arylpyridones as Cbl-b Inhibitors, J. Med. Chem., 2024, 67(2), 1500–1512 CrossRef CAS PubMed.
- T. R. Quinn, K. A. Giblin, C. Thomson, J. A. Boerth, G. Bommakanti and E. Braybrooke, et al., Accelerated Discovery of Carbamate Cbl-b Inhibitors Using Generative AI Models and Structure-Based Drug Design, J. Med. Chem., 2024, 67(16), 14210–14233 CrossRef CAS PubMed.
- R. Cencic, D. R. Hall, F. Robert, Y. Du, J. Min and L. Li, et al., Reversing chemoresistance by small molecule inhibition of the translation initiation complex eIF4F, Proc. Natl. Acad. Sci. U. S. A., 2011, 108(3), 1046–1051 CrossRef CAS PubMed.
- R. Cencic, M. Desforges, D. R. Hall, D. Kozakov, Y. Du and J. Min, et al., Blocking eIF4E-eIF4G interaction as a strategy to impair coronavirus replication, J. Virol., 2011, 85(13), 6381–6389 CrossRef CAS PubMed.
- X. Wan, T. Yang, A. Cuesta, X. Pang, T. E. Balius and J. J. Irwin, et al., Discovery of Lysine-Targeted eIF4E Inhibitors through Covalent Docking, J. Am. Chem. Soc., 2020, 142(11), 4960–4964 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |