Open Access Article
Open Access ArticleDesign, synthesis, biological evaluation, and molecular dynamics study of asymmetric N-isopropyl-4-piperidone diarylpentanoids as potential anticancer and anti-inflammatory agents
Huy-Khoa Tranab,
Truc-Vy Maiab,
Van-Dung Le ab,
Tran-Nguyen Minh-An
ab,
Tran-Nguyen Minh-An c,
Dinh-Tri Maiab,
Quoc-Tuan Tran
c,
Dinh-Tri Maiab,
Quoc-Tuan Tran d,
Thi Quyen Vue,
Thanh-Danh Nguyen
d,
Thi Quyen Vue,
Thanh-Danh Nguyen ab and
Chi-Hien Dang
ab and
Chi-Hien Dang *ab
*ab
aInstitute of Advanced Technology, Vietnam Academy of Science and Technology, 1B, TL29 An Phu Dong Ward, Ho Chi Minh City, Vietnam. E-mail: dangchihien@gmail.com
bGraduate University of Science and Technology, Vietnam Academy of Science and Technology, 18 Hoang Quoc Viet, Nghia Do Ward, Hanoi 11000, Vietnam
cFaculty of Chemical Engineering (FCE), Industrial University of Ho Chi Minh City (IUH), Ho Chi Minh City, Vietnam
dDeparment of Biochemistry, Faculty of Biology-Biotechnology, Ho Chi Minh University of Science, Ho Chi Minh City, Vietnam
eFaculty of Applied Technology, Van Lang School of Technology, Van Lang University, Ho Chi Minh City, Vietnam
First published on 1st May 2026
Abstract
A series of symmetric and asymmetric diarylpentanoids based on N-isopropyl-4-piperidone were rationally designed and synthesized through a Claisen–Schmidt condensation reaction. This was done to investigate their potential as anticancer and anti-inflammatory agents. The synthesized compounds were evaluated for cytotoxic activity against the MCF-7 breast cancer cell line, as well as for nitric oxide inhibitory activity in LPS-stimulated RAW264.7 macrophages. Structure-activity relationship analysis revealed that the presence of electron-donating substituents and asymmetric aryl substitutions significantly enhanced biological activity. Among the synthesized derivatives, compound KB2d exhibited the strongest anticancer activity, with an IC50 value of 1.54 µM, while compound KB2c demonstrated promising anti-inflammatory activity, with an IC50 value of 2.51 µM. To understand the molecular basis of these activities, molecular docking simulations and 200 ns molecular dynamics were conducted on aromatase and PDE4D protein targets. The leading compound, KB2d, showed stable binding behavior, low ligand RMSD oscillations, and favorable binding free energy (ΔG_bind ≈ −33 kcal mol−1) as calculated using the MM-PBSA method. These findings suggest that asymmetric N-isopropyl-4-piperidone diarylpentanoids represent promising structural frameworks for the further development of anticancer and anti-inflammatory agents.
1. Introduction
Curcumin (diferuloylmethane) is a well-known natural polyphenol isolated from the rhizome of turmeric (Curcuma longa L). For decades, this compound has attracted considerable scientific interest due to its broad spectrum of biological activity, including antibacterial, anti-inflammatory, antioxidant effects, and especially its remarkable anticancer potential.1,2 Studies have demonstrated that curcumin can interact with many different molecular targets, thereby regulating key signaling pathways involved in inflammation, oxidative stress, cell proliferation, and programmed cell death (apoptosis) in cancer cells. This is clearly a very attractive pharmacological premise.However, despite its promising biological properties, the transition of curcumin from the laboratory to clinical application still faces significant obstacles. The main reasons stem from unfavorable pharmacokinetic characteristics such as low water solubility, rapid metabolism, and degradation in the body, leading to very limited systemic bioavailability.3–5 Structurally, these limitations are closely related to the instability of the central β-diketone group. The presence of the active methylene group along with the keto–enol tautomerism makes the molecule susceptible to chemical and biological degradation even under normal physiological conditions.6,7
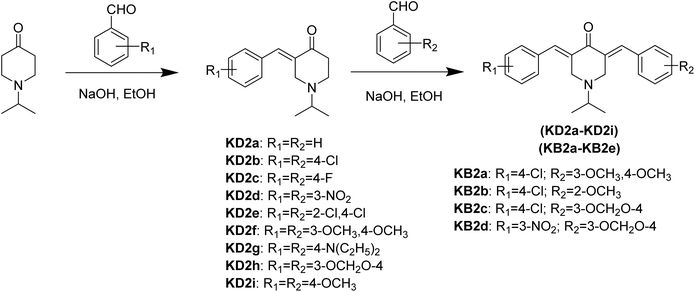
To overcome these barriers, many structural modification strategies have been developed to improve the stability and bioavailability of curcumin analogs.4,8 One approach considered particularly effective is the complete replacement of the β-diketone group with a more stable monoketone framework, thereby forming curcuminoid monoketone (MAC) groups, also known as diarylpentanoids.9,10 There is growing evidence that MAC derivatives exhibit better chemical stability and metabolism than natural curcumin, while maintaining or even enhancing important biological activities such as anticancer, anti-inflammatory, and antimicrobial.9,11,12 Notably, the integration of heterocyclic frameworks such as piperidin-4-one into diarylpentanoid structures (Fig. 1) has been shown to increase configuration rigidity, modulate electron distribution, and significantly improve selective toxicity against various cancer cell lines.13,14
Although symmetric MACs have been fairly extensively studied due to their simple synthesis procedures and often high reaction yields, the directed design of asymmetric MACs remains relatively underexploited.10,15,16 Meanwhile, the varying changes in aryl substituents at the C3 and C5 positions can create a delicate “push–pull” electronic system, allowing for controlled tuning of physicochemical properties such as lipophilicity, polarity, and adaptability when binding to biological targets.17,18 This structural asymmetry is expected to enhance molecular recognition and promote more complex ligand–target interactions. In fact, Zhang et al. showed that the asymmetric diarylpentanoid WZ26 (Fig. 1) can target the enzyme thioredoxin reductase 1, increasing levels of reactive oxygen species (ROS) and activating the JNK signaling pathway in colon cancer cells.19 Similarly, Jonathan et al. reported a MAC derivative carrying the N-piperidin-4-one framework, designated MBMP (Fig. 1), exhibiting potent antimicrobial activity against both Gram-positive and Gram-negative bacteria such as Bacillus subtilis, Staphylococcus aureus, Escherichia coli, and Pseudomonas aeruginosa, even outperforming ampicillin in some experiments.20 These results demonstrate the significant potential of asymmetric design strategies.
Parallel to the development of synthetic pharmaceutical chemistry, modern computational methods are playing an increasingly important role in drug discovery research. Molecular coupling allows for rapid prediction of potential ligand–protein interactions, but accuracy is sometimes limited due to the use of static receptor structures and the incomplete description of complex solvent effects.21,22 Therefore, long-time scale molecular dynamics (MD) simulations, combined with computational binding free energy methods such as MM-GBSA or MM-PBSA, have become powerful tools for evaluating binding stability, conformational adaptability, and binding affinity in environments closer to physiological conditions.23,24 Integrating these computational techniques with experimental biological data can provide deeper mechanistic insights and support the rational optimization of the lead substance.
Based on the above, this study was conducted to design and synthesize a library of diarylpentanoids based on the N-isopropyl-4-piperidone framework, including both symmetric derivatives (KD chains) and asymmetric analogs (KB chains). The synthesized compounds were evaluated for in vitro antimicrobial activity, cytotoxicity against the MCF-7 breast cancer cell line, and the ability to inhibit nitric oxide production as an indicator of anti-inflammatory potential. In addition, comprehensive computational studies including molecular coupling, MD 200 ns simulation, binding free energy calculation, and in silico ADMET prediction were performed to correlate computational results with experimental data. Through this, the study aims to provide a more comprehensive understanding of the structure-activity relationship of curcuminoid derivatives at the molecular level.
2. Materials and methods
2.1. Materials
1-Isopropyl-4-piperidinone, aromatic aldehydes, n-hexane, and ethanol were supplied by Acros Organics (Geel, Belgium). Lipopolysaccharide (LPS) from E. coli, sodium nitrite, sulfanilamide, N-(1-naphthyl)ethylenediamine dihydrochloride, and dimethyl sulfoxide (DMSO) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Dulbecco's Modified Eagle's Medium (DMEM) and fetal bovine serum (FBS) were obtained from Life Technologies Inc. (Gaithersburg, MD, USA). All other chemicals and reagents were of analytical grade and were used without further purification.The cell lines utilized in this study are as follows: MCF-7: a human breast carcinoma cell line provided by Professor Dr J. M. Pezzuto (Long Island University, USA) and Professor Dr Jeanette Maier (University of Milan, Italy); RAW 264.7: A murine macrophage cell line provided by Professor Dr Domenico Delfino (University of Perugia, Italy).
2.2. Synthesis
2.2.1.1 3,5-Di((E)-benzylidene)-1-isopropylpiperidin-4-one (KD2a). Yellow powder. Yield 93.6%. 1H NMR (600 MHz, CDCl3) δ 7.79 (s, 2H, Hβ và Hβ′), 7.44–7.39 (m, 8H, H–Ar), 7.38–7.34 (m, 2H, H–Ar), 3.88 (s, 4H, H8 và H8′), 2.93 (hept, J = 6.5 Hz, 1H, H9), 1.05 (d, J = 6.5 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 188.00, 135.90, 135.48, 134.09, 130.35, 128.87, 128.57, 53.57, 50.54, 18.52. FTIR (KBr, νmax cm−1): 3442, 2862, 1670, 1580, 1580, 1028, 762. ESI-MS (m/z): 318.1859 [M + H]+ calcd for C22H24NO ([M + H]+ = 318.1858).
2.2.1.2 3,5-Bis((E)-4-chlorobenzylidene)-1-isopropylpiperidin-4-one (KD2b). Yellow powder. Yield 94.2%. 1H NMR (600 MHz, CDCl3) δ 7.71 (s, 2H, Hβ và Hβ′), 7.41–7.38 (m, 4H, H–Ar), 7.35–7.30 (m, 4H, H–Ar), 3.82 (s, 4H, H8 và H8′), 2.93 (hept, J = 6.5 Hz, 1H, H9), 1.05 (d, J = 6.6 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 187.5, 135.0, 134.7, 134.4, 133.8, 131.5, 128.9, 53.6, 50.4, 18.5. FTIR (KBr, νmax cm−1): 3431, 3033, 2873, 1671, 1586, 1563, 1036, 702. ESI-MS (m/z): 386.1073 [M + H]+ calcd for C22H22Cl2NO ([M + H]+ = 386.1078).
2.2.1.3 3,5-Bis((E)-4-fluorobenzylidene)-1-isopropylpiperidin-4-one (KD2c). Yellow powder. Yield 94.1%. 1H NMR (600 MHz, CDCl3) δ 7.74 (s, 2H, Hβ và Hβ′), 7.42–7.36 (m, 4H, H–Ar), 7.15–7.09 (m, 4H, H–Ar), 3.83 (s, 4H, H8 và H8′), 2.94 (hept, J = 6.5 Hz, 1H, H9), 1.06 (d, J = 6.5 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 187.7, 162.9, 134.8, 133.7, 132.2, 131.6, 115.8, 53.63, 50.4, 18.5. FTIR (KBr, νmax cm−1): 3450, 2875, 1675, 1600, 1508, 1030, 759. ESI-MS (m/z): 354.1671 [M + H]+ calcd for C22H22F2NO ([M + H]+ = 354.1669).
2.2.1.4 1-Isopropyl-3,5-bis((E)-3-nitrobenzylidene)piperidin-4-one (KD2d). Yellow powder. Yield 92.8%. 1H NMR (600 MHz, CDCl3) δ 8.28–8.22 (m, 4H, H–Ar), 7.79 (d, J = 1.8 Hz, 2H, Hβ và Hβ′), 7.72 (m, 2H, H–Ar), 7.64 (m, 2H, H–Ar), 3.88 (s, 4H, H8 và H8′), 2.96 (hept, J = 6.5 Hz, 1H, H9), 1.26 (s, 0H), 1.07 (d, J = 6.6 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 187.1, 148.5, 136.8, 136.1, 135.9, 133.3, 129.8, 124.4, 123.6, 53.9, 50.2, 18.4. FTIR (KBr, νmax cm−1): 3062, 2872, 1678, 1597, 1528, 1039, 714. ESI-MS (m/z): 408.1559 [M + H]+ calcd for C22H22N3O5 ([M + H]+ = 408.1559).
2.2.1.5 3,5-Bis((E)-2,4-dichlorobenzylidene)-1-isopropylpiperidin-4-one (KD2e). Yellow powder. Yield 92.7%. 1H NMR (600 MHz, CDCl3) δ 7.88 (s, 2H, Hβ và Hβ′), 7.49 (m, 2H, H–Ar), 7.30 (m, 2H, H–Ar), 7.18 (m, 2H, H–Ar), 3.69 (s, 4H, H8 và H8′), 2.86 (hept, J = 6.5 Hz, 1H, H9), 0.97 (d, J = 6.5 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 186.9, 135.9, 135.5, 135.2, 132.4, 132.3, 130.9, 129.9, 126.9, 53.1, 50.1, 18.5. FTIR (KBr, νmax cm−1): 3434, 3074, 2928, 1671, 1589, 1549, 1033, 734. ESI-MS (m/z): 454.0300 [M + H]+ calcd for C22H20Cl4NO ([M + H]+ = 454.0299).
2.2.1.6 3,5-Bis((E)-3,4-dimethoxybenzylidene)-1-isopropylpiperidin-4-one (KD2f). Yellow powder. Yield 94.8%. 1H NMR (600 MHz, CDCl3) δ 7.75 (s, 2H, Hβ và Hβ′), 7.02 (m, 2H, H–Ar), 6.96 (m, 2H, H–Ar), 6.93 (m, 2H, H–Ar), 3.95–3.87 (m, 12H, –OCH3 và H8 và H8′), 2.95 (hept, J = 6.5 Hz, 1H, H9), 1.08 (d, J = 6.5, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 187.7, 149.9, 148.8, 135.9, 132.4, 128.5, 123.8, 113.8, 111.1, 55.9, 55.9, 53.4, 50.7, 18.8. FTIR (KBr, νmax cm−1): 3442, 3012, 2928, 1596, 1514, 1022, 740. ESI-MS (m/z): 438.2277 [M + H]+ calcd for C26H32NO5 ([M + H]+ = 438.2280).
2.2.1.7 3,5-Bis((E)-4-(diethylamino)benzylidene)-1-isopropylpiperidin-4-one (KD2g). Yellow powder. Yield 94.3%. 1H NMR (600 MHz, CDCl3) δ 7.73 (s, 2H, Hβ và Hβ′), 7.36–7.31 (m, 4H, H–Ar), 6.70–6.65 (m, 4H, H–Ar), 3.91 (s, 4H, H12 và H12′), 3.40 (q, J = 7.1 Hz, 8H, H7, H7′, H9 và H9′), 2.97 (hept, J = 6.5 Hz, 1H, H13), 1.20 (t, J = 7.1 Hz, 12H, 8H, H8, H8′, H10 và H10′), 1.12 (d, J = 6.5 Hz, 6H, H14 và H14′). 13C NMR (150 MHz, CDCl3) δ 187.5, 148.1, 136.2, 132.9, 129.6, 122.9, 111.1, 53.4, 50.9, 44.4, 18.9, 12.7. FTIR (KBr, νmax cm−1): 2868, 1654, 1582, 1518, 1042, 704. ESI-MS (m/z): 460.3330 [M + H]+ calcd for C30H42N3O ([M + H]+ = 460.3328).
2.2.1.8 (3E,5E)-3,5-bis(benzo[d][1,3]dioxol-5-ylmethylene)-1-isopropylpiperidin-4-one (KD2h). Yellow powder. Yield 95.1%. 1H NMR (600 MHz, CDCl3) δ 7.68 (s, 2H, Hβ và Hβ′), 6.94 (m, 2H, H–Ar), 6.90–6.84 (m, 4H, H–Ar), 6.01 (s, 4H, H7 và H7′), 3.83 (s, 4H, H9 và H9′), 2.94 (hept, J = 6.5 Hz, 1H, H10), 1.08 (d, J = 6.5 Hz, 6H, H11 và H11′). 13C NMR (150 MHz, CDCl3) δ 187.7, 148.3, 147.9, 135.6, 132.6, 129.7, 125.8, 110.0, 108.6, 101.5, 53.6, 50.5, 18.6. FTIR (KBr, νmax cm−1): 3071, 2920, 1667, 1590, 1574, 1039, 769. ESI-MS (m/z): 406.1648 [M + H]+ calcd for C24H24NO5 ([M + H]+ = 406.1654).
2.2.1.9 1-Isopropyl-3,5-bis((E)-4-methoxybenzylidene)piperidin-4-one (KD2i). Yellow powder. Yield 93.4%. 1H NMR (600 MHz, CDCl3) δ 7.75 (s, 2H, Hβ và Hβ′), 7.41–7.35 (m, 4H, H–Ar), 6.98–6.92 (m, 4H, H–Ar), 3.87 (s, 4H, H8 và H8′), 3.85 (s, 6H, OCH3), 2.94 (hept, J = 6.5, 1H), 1.08 (d, J = 6.5 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 135.6, 132.3, 128.3, 114.1, 55.4, 53.5, 50.6, 18.7. FTIR (KBr, νmax cm−1): 2958, 1665, 1601, 1501, 1031, 730. ESI-MS (m/z): 378.2033 [M + H]+ calcd for C24H28NO3 ([M + H]+ = 378.2069).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, v/v) as the eluent until the disappearance of the aldehyde spot was observed. Subsequently, the second aldehyde (R′CHO, 1.0 mmol), aqueous NaOH (20%, 2 mL), and ethanol (5 mL) were added to the reaction mixture, and ultrasonic irradiation was continued under the same conditions until TLC indicated completion of the reaction. The resulting precipitate was collected by filtration and recrystallized from ethanol. Final purification was achieved by column chromatography, affording the desired asymmetric monoketone curcuminoid derivatives. The purity of all synthesized compounds was estimated to be >95% based on 1H NMR integration; further HPLC-based purity assessment is planned for lead optimization stages.
10, v/v) as the eluent until the disappearance of the aldehyde spot was observed. Subsequently, the second aldehyde (R′CHO, 1.0 mmol), aqueous NaOH (20%, 2 mL), and ethanol (5 mL) were added to the reaction mixture, and ultrasonic irradiation was continued under the same conditions until TLC indicated completion of the reaction. The resulting precipitate was collected by filtration and recrystallized from ethanol. Final purification was achieved by column chromatography, affording the desired asymmetric monoketone curcuminoid derivatives. The purity of all synthesized compounds was estimated to be >95% based on 1H NMR integration; further HPLC-based purity assessment is planned for lead optimization stages.
2.2.2.1 3-((E)-4-chlorobenzylidene)-5-((E)-3,4-dimethoxybenzylidene)-1-isopropylpiperidin-4-one (KB2a). Yellow powder. Yield 54.3%. 1H NMR (600 MHz, CDCl3) δ 7.74 (s, 1H, Hβ), 7.72 (s, 1H, Hβ′), 7.40 (d, J = 8.5 Hz, 2H, H–Ar), 7.33 (d, J = 8.5 Hz, 2H, H–Ar), 7.03 (dd, J = 8.3, 2.0 Hz, 1H, H–Ar), 6.96 (d, J = 2.0 Hz, 1H, H–Ar), 6.93 (d, J = 8.3 Hz, 1H, H–Ar), 3.93 (s, 3H, OCH3), 3.92 (s, 3H, OCH3), 3.90 (s, 2H, H8), 3.83 (s, 2H, H8′), 2.94 (hept, J = 6.5 Hz, 1H, H9), 1.07 (d, J = 6.5 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 187.6, 150.0, 148.9, 136.4, 134.8, 134.6, 134.2, 134.0, 132.2, 131.5, 128.9, 128.3, 123.9, 113.8, 111.1, 56.0, 55.9, 53.5, 50.6, 50.5, 18.6. FTIR (KBr, νmax cm−1): 2930, 1666, 1602, 1511, 1027, 736. ESI-MS (m/z): 412.1660 [M + H]+ calcd for C24H27ClNO3 ([M + H]+ = 412.1679).
2.2.2.2 3-((E)-4-chlorobenzylidene)-1-isopropyl-5-((E)-2-methoxybenzylidene)piperidin-4-one (KB2b). Yellow powder. Yield 54.6%. 1H NMR (600 MHz, CDCl3) δ 8.04 (s, 1H, Hβ), 7.73 (s, 1H, Hβ′), 7.42–7.37 (m, 2H, H–Ar), 7.37–7.30 (m, 3H, H–Ar), 7.22 (dd, J = 7.7, 1.7 Hz, 1H, H–Ar), 6.98 (td, J = 7.6, 0.9 Hz, 1H, H–Ar), 6.93 (dd, J = 8.3, 1.0 Hz, 1H, H–Ar), 3.86 (s, 3H, OCH3), 3.83 (s, 2H, H8), 3.78 (s, 2H, H8′), 2.90 (hept, J = 6.7 Hz, 1H, H9), 1.02 (d, J = 6.5 Hz, 6H, H10 và H10′). 13C NMR (150 MHz, CDCl3) δ 158.5, 134.4, 132.2, 131.5, 130.5, 130.2, 128.8, 120.1, 110.9, 55.5, 53.4, 50.6, 50.5, 18.6. FTIR (KBr, νmax cm−1): 1666, 1605, 1570, 1031, 707. ESI-MS (m/z): 382.1549 [M + H]+ calcd for C23H25ClNO2 ([M + H]+ = 382.1574).
2.2.2.3 (E)-3-(benzo[d][1,3]dioxol-5-ylmethylene)-5-((E)-4-chlorobenzylidene)-1-isopropylpiperidin-4-one (KB2c). Yellow powder. Yield 56.3%. 1H NMR (600 MHz, CDCl3) δ 7.71 (s, 1H, Hβ), 7.70 (s, 1H, Hβ′), 7.41–7.37 (m, 2H, H–Ar), 7.34–7.30 (m, 2H, H–Ar), 6.97–6.93 (m, 1H, H–Ar), 6.90 (d, J = 1.7 Hz, 1H, H–Ar), 6.88 (d, J = 8.1 Hz, 1H, H–Ar), 6.03 (s, 2H, H7), 3.85 (s, 2H, H9), 3.81 (s, 2H, H9′), 2.94 (hept, J = 6.5 Hz, 1H, H10), 1.26 (s, 1H), 1.07 (d, J = 6.5 Hz, 6H, H11 và H11′). 13C NMR (150 MHz, CDCl3) δ 136.2, 134.2, 131.5, 128.9, 126.0, 110.0, 108.7, 101.5, 53.6, 50.7, 50.3, 18.6. FTIR (KBr, νmax cm−1): 1666, 1602, 1501, 1033, 769. ESI-MS (m/z): 396.1343 [M + H]+ calcd for C23H23ClNO3 ([M + H]+ = 396.1366).
2.2.2.4 (E)-3-(benzo[d][1,3]dioxol-5-ylmethylene)-1-isopropyl-5-((E)-3-nitrobenzylidene)piperidin-4-one (KB2d). Yellow powder. Yield 56.7%. 1H NMR (600 MHz, CDCl3) δ 8.24 (m, 1H, H–Ar), 8.22–8.20 (m, 1H, H–Ar), 7.76 (s, 1H, Hβ′), 7.72 (s, 1H, Hβ), 7.72–7.67 (m, 1H, H–Ar), 7.61 (m, 1H, H–Ar), 6.97 (m, 1H, H–Ar), 6.91 (m, 1H, H–Ar), 6.89 (m, 1H, H–Ar), 6.03 (s, 2H, H7), 3.87 (s, 2H, H9′), 3.84 (s, 2H, H9), 2.95 (hept, J = 6.5 Hz, 1H, H10), 1.08 (d, J = 6.5 Hz, 6H, H11 và H11′). 13C NMR (150 MHz, CDCl3) δ 187.3, 148.6, 148.4, 148.0, 137.2, 136.8, 136.6, 135.9, 132.2, 132.0, 129.6, 129.4, 126.2, 124.3, 123.2, 110.0, 108.7, 101.6, 53.7, 50.6, 50.1, 18.5. FTIR (KBr, νmax cm−1): 1675, 1581, 1502, 1036, 702. ESI-MS (m/z): 407.1592 [M + H]+ calcd for C23H22N2O5 ([M + H]+ = 407.1607).
2.3. Antimicrobial activity
Chloramphenicol (2 mg mL−1) was used as the positive control, while the solvent was used as the negative control. The plates were incubated at 37 °C for 24 h. Antimicrobial activity was assessed by measuring the diameter of the inhibition zones surrounding each well. Experiments were performed in triplicate and the results were expressed as the mean inhibition zone diameter (mm). A well diameter of 6 mm was considered as no antibacterial activity.
2.4. Anti-inflammatory activity evaluation
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3.
3.2.5. In vitro cytotoxicity assay against MCF-7 cells
The cytotoxic activity of the samples was evaluated against the human breast cancer cell line MCF-7 using the sulforhodamine B (SRB) assay following the protocol established by the U.S. National Cancer Institute (NCI) with minor modifications.29 This colorimetric method determines total cellular protein content as an indirect measurement of cell proliferation and viability.MCF-7 cells were harvested by trypsinization and counted using a hemocytometer. Cells were seeded into 96-well plates at a density of 190 µL per well in complete culture medium supplemented with 5% fetal bovine serum (FBS) and incubated for 18–20 h at 37 °C in a humidified 5% CO2 atmosphere to allow cell attachment and stabilization.
Test samples were dissolved in 100% DMSO to obtain stock solutions (20 mM for pure compounds or 20 mg mL−1 for extracts) and subsequently diluted with serum-free culture medium to generate four serial concentrations. 10 µL of each diluted sample was added to the wells containing cells (final volume 200 µL). Wells containing cells treated with 1% DMSO served as the negative control, while wells fixed immediately after treatment were used as the day-0 reference control.
After 48 h incubation, cells were fixed with cold 20% (w/v) trichloroacetic acid for 1 h. Plates were gently washed with distilled water and air-dried at room temperature. The fixed cells were stained with 0.4% (w/v) SRB in 1% acetic acid for 30 min at 37 °C, followed by washing three times with 1% acetic acid to remove unbound dye. After drying, the bound dye was solubilized with 200 µL of 10 mM unbuffered Tris base, and absorbance was measured at 540 nm using a microplate reader (BioTek).
Cell growth inhibition was calculated according to the eqn (1):
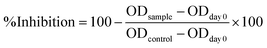 | (1) |
All experiments were performed in triplicate. Ellipticine (10, 2, 0.4, and 0.08 µg mL−1) was used as a positive control. The IC50 values (concentration causing 50% inhibition of cell growth) were determined using TableCurve 2D v4 software. According to the NCI criteria, crude extracts are considered active when IC50 ≤ 20 µg mL−1, while pure compounds are regarded as active when IC50 ≤ 5 µM.
2.6. Molecular docking study
Molecular docking simulations were performed to investigate the binding interactions between the studied compounds and selected target proteins. All calculations were conducted using AutoDockTools 1.5.6 and MGLTools 1.5.6, while Avogadro and Discovery Studio Visualizer 2021 were employed for structure preparation and visualization. The crystal structures of the target proteins (PDB IDs: 1NMT, 1JIK, 1ZI0, 3EQM, and 4WCU) were retrieved from the Protein Data Bank (PDB).30–34 Prior to docking, protein structures were prepared by removing crystallographic water molecules, co-crystallized ligands, and unnecessary ions using Discovery Studio. Polar hydrogen atoms were added and Kollman charges were assigned to the protein structures, which were then saved in PDBQT format.The chemical structures of the investigated ligands were constructed using ChemDraw and converted into optimized 3D geometries in Avogadro. Energy minimization was performed using the MMFF94 force field. Rotatable bonds were defined and Gasteiger charges were assigned using AutoDockTools, and the optimized ligands were exported in PDBQT format. The docking grid box was defined based on the binding pocket of the co-crystallized ligand or the known active site residues. The grid box center coordinates (x, y, z) were set to encompass the catalytic region, with grid dimensions of 0.6 × 0.6 × 0.6 Å and a grid spacing of 0.5–0.7 Å.35,36 Semi-flexible docking was performed with 500 independent runs, where the protein was kept rigid while the ligand was allowed full rotational flexibility. The docking conformations with the lowest binding energies were selected for further analysis. Ligand–protein interactions, including hydrogen bonding, hydrophobic interactions, and π–π stacking, were analyzed and visualized using Discovery Studio Visualizer.
2.7. Molecular dynamics (MD) simulation
Molecular dynamics simulations were performed using GROMACS 2023 to evaluate the stability of the protein–ligand complexes obtained from docking.37–39 Simulations were executed on a GPU-accelerated high-performance computing system equipped with an RTX 3050 GPU. The protein–ligand complex with the lowest docking binding energy was selected as the initial structure for MD simulations. The CHARMM27 force field was applied to generate the protein topology using the pdb2gmx module in GROMACS. Ligand topology parameters were generated using a compatible external server consistent with the selected force field.The complex was placed in the center of a simulation box with sufficient distance between the protein surface and box boundaries and solvated using the TIP3P water model. Counter ions (Na+ and Cl−) were added using the genion module to neutralize the system and mimic physiological ionic conditions. Energy minimization was performed using the steepest descent algorithm for up to 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps to eliminate steric clashes and unfavorable contacts. Subsequently, system equilibration was conducted in two stages with positional restraints applied to the heavy atoms of the protein and ligand: an NVT ensemble simulation for 5 ns at 300 K, followed by an NPT ensemble simulation for 5 ns at 1 bar pressure.
000 steps to eliminate steric clashes and unfavorable contacts. Subsequently, system equilibration was conducted in two stages with positional restraints applied to the heavy atoms of the protein and ligand: an NVT ensemble simulation for 5 ns at 300 K, followed by an NPT ensemble simulation for 5 ns at 1 bar pressure.
After equilibration, positional restraints were removed and a production MD simulation of 200 ns was carried out with a time step of 2 fs. Trajectory analyses were conducted using built-in GROMACS analysis tools after correcting for periodic boundary conditions. The structural stability and dynamic behavior of the complexes were evaluated using several parameters, including root-mean-square deviation (RMSD), root-mean-square fluctuation (RMSF), radius of gyration (Rg), hydrogen bond analysis, and solvent-accessible surface area (SASA). The binding free energy of the complexes was further estimated using the MM-PBSA method implemented in the gmx_MMPBSA package.
The in silico pharmacokinetic properties and drug-likeness of the synthesized MACs were evaluated using the SwissADME web tool (Swiss Institute of Bioinformatics).40 For ADME prediction (absorption, distribution, metabolism, and excretion), the 2D structures of the ligands were converted into SMILES format and submitted to the SwissADME server. Key physicochemical parameters, including topological polar surface area (TPSA), water solubility (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S), and lipophilicity (log
S), and lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), were calculated to estimate human intestinal absorption (HIA) and blood–brain barrier (BBB) permeability. Furthermore, the drug-likeness of the MACs was rigorously assessed based on Lipinski's Rule of Five, in combination with Veber and Egan molecular descriptors. Structural alerts, including pan-assay interference compounds (PAINS) and Brenk filters, were also analyzed to eliminate potential false positives and reactive functional groups, thereby ensuring the reliability and quality of the selected candidate compounds.
P), were calculated to estimate human intestinal absorption (HIA) and blood–brain barrier (BBB) permeability. Furthermore, the drug-likeness of the MACs was rigorously assessed based on Lipinski's Rule of Five, in combination with Veber and Egan molecular descriptors. Structural alerts, including pan-assay interference compounds (PAINS) and Brenk filters, were also analyzed to eliminate potential false positives and reactive functional groups, thereby ensuring the reliability and quality of the selected candidate compounds.
3. Results and discussion
3.1. Synthesis
In this study, MAC derivatives bearing N-isopropyl-4-piperidone were synthesized via a base-catalyzed Claisen–Schmidt condensation reaction (Fig. 2). The reaction efficiency and product distribution were found to depend strongly on the structural symmetry of the target compounds as well as the electronic characteristics of substituents on the benzaldehyde rings. Since structural symmetry and substituent effects are known to significantly influence the biological activity of MAC derivatives, the synthesized compounds were systematically evaluated for antibacterial, anti-inflammatory, and anticancer activities. To further elucidate the molecular basis of their bioactivity, the most potent derivatives were subjected to molecular docking analysis with relevant biological targets. In addition, molecular dynamics (MD) simulations combined with MM-GBSA binding free-energy calculations were employed to provide a more reliable estimation of ligand–protein binding affinity under physiologically relevant conditions.For synthesis of MACs, the symmetric derivatives (KD2a–KD2i) were obtained in excellent yields (>92%). This high efficiency can be attributed to the thermodynamic stabilization associated with the formation of an extended cross-conjugated π-system, which favors the generation of the symmetric bis-benzylidene structure. In contrast, the preparation of asymmetric derivatives (KB2a–KB2e) required stepwise condensation, resulting in significantly lower yields (54.3–63.2%). The presence of strong electron-donating substituents such as methoxy groups at the ortho or para positions (e.g., KB2a and KB2b) further reduced the electrophilicity of the aldehyde carbonyl carbon and introduced steric hindrance, thereby slowing the nucleophilic attack of the enolate intermediate and lowering the overall reaction efficiency.
To overcome the moderate yields of the asymmetric KB series (54.3–56.7%) caused by the reduced nucleophilicity of the mono-arylidene intermediate and steric hindrance, several optimization strategies can be implemented in future scale-up efforts. First, transitioning from ultrasonic irradiation to microwave-assisted organic synthesis (MAOS) could provide the localized and rapid thermal energy required to surpass the elevated activation energy barrier of the second condensation step. Second, the application of specific Lewis acid catalysts (e.g., BF3 OEt2 or TiCl4) could be explored to strongly activate the carbonyl carbon of the second, electron-rich aldehyde. However, careful optimization will be required to minimize competitive coordination with electron-rich oxygen donors present in substrates such as piperonaldehyde. Lastly, optimizing the reaction medium with specific polar aprotic solvent systems, coupled with continuous water removal techniques, could favorably shift the reaction equilibrium, thereby enhancing the overall thermodynamic efficiency.
From a pharmaceutical safety and CMC (Chemistry, Manufacturing, and Controls) perspective, the risk assessment of potential mutagenic impurities, particularly nitrosamines, is crucial when secondary amines (e.g., pyrrolidine used as a catalyst) and tertiary amines (e.g., the starting material 1-isopropyl-4-piperidinone) are involved in the synthetic route. However, the risk of N-nitrosamine formation in our study is considered negligible. The sequential Claisen–Schmidt condensation was strictly conducted under basic conditions without the introduction of any nitrosating agents or nitrite/nitrate sources. Furthermore, the rigorous purification protocol, involving silica gel column chromatography and subsequent recrystallization from ethanol, ensures the efficient removal of any unreacted residual amines. This strict quality control aligns with the safety requirements for early-stage drug discovery.
The structures of all synthesized MAC derivatives were confirmed by FTIR, NMR, and HRMS analyses. In the 1H NMR spectra, the characteristic vinylic protons (Hβ/Hβ′) were clearly observed for all compounds. For the symmetric KD series, these protons appeared as a singlet integrating for two protons at δ 7.68–7.88 ppm, whereas in the asymmetric KB series the signal was split into two singlets corresponding to one proton each. The pronounced downfield chemical shift indicates the formation of the thermodynamically favored (E, E) configuration, in which the vinylic protons are positioned cis to the carbonyl groups and experience strong deshielding effects from the anisotropic magnetic field of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds. The N-isopropyl substituent exhibited characteristic signals consisting of a septet at δ 2.86–2.97 ppm (1H, J 6.5 Hz) corresponding to the methine proton and a doublet at δ 0.97–1.12 ppm (6H, J 6.5 Hz) attributed to the two methyl groups. In addition, the piperidone ring protons (H8/H8′) appeared as a singlet integrating for four protons at δ 3.69–3.90 ppm in the symmetric series. The 13C NMR spectra further confirmed the conjugated ketone structure, with the carbonyl carbon resonating at δ 186.8–188.0 ppm, characteristic of an α, β-unsaturated ketone system. The vinylic carbons (Cβ attached to the phenyl ring and Cα within the piperidone framework) appeared in the range δ 131–137 ppm. Collectively, these spectroscopic data are fully consistent with the proposed 3,5-bis(benzylidene)-4-piperidone framework of the synthesized MACs.
O bonds. The N-isopropyl substituent exhibited characteristic signals consisting of a septet at δ 2.86–2.97 ppm (1H, J 6.5 Hz) corresponding to the methine proton and a doublet at δ 0.97–1.12 ppm (6H, J 6.5 Hz) attributed to the two methyl groups. In addition, the piperidone ring protons (H8/H8′) appeared as a singlet integrating for four protons at δ 3.69–3.90 ppm in the symmetric series. The 13C NMR spectra further confirmed the conjugated ketone structure, with the carbonyl carbon resonating at δ 186.8–188.0 ppm, characteristic of an α, β-unsaturated ketone system. The vinylic carbons (Cβ attached to the phenyl ring and Cα within the piperidone framework) appeared in the range δ 131–137 ppm. Collectively, these spectroscopic data are fully consistent with the proposed 3,5-bis(benzylidene)-4-piperidone framework of the synthesized MACs.
3.2. Biological activity evaluation
| Entry | Compounds | Escherichia coli | Staphylococcus aureus | Candida albicans | |||
|---|---|---|---|---|---|---|---|
| Inhibition zone (mm) | MIC (mg mL−1) | Inhibition zone (mm) | MIC (mg mL−1) | Inhibition zone (mm) | MIC (mg mL−1) | ||
| 1 | KD2a | 6.0 | — | 6.0 | — | 6.0 | — |
| 2 | KD2b | 6.1 ± 0.2 | 2.0 | 6.2 ± 0.2 | 1.5 | 9.0 ± 0.1 | 0.75 |
| 3 | KD2c | 6.0 | — | 6.3 ± 0.1 | 1.5 | 10.0 ± 0.1 | 0.50 |
| 4 | KD2d | 6.0 | — | 6.2 ± 0.1 | 1.5 | 8.0 ± 0.1 | 0.50 |
| 5 | KD2e | 6.2 ± 0.1 | 2.0 | 6.1 ± 0.1 | 1.5 | 10.0 ± 0.2 | 0.50 |
| 6 | KD2f | 9.0 ± 0.2 | 1.0 | 6.2 ± 0.1 | 1.5 | 6.0 | — |
| 7 | KD2g | 6.0 | — | 6.3 ± 0.2 | 2.0 | 8.0 ± 0.2 | 0.75 |
| 8 | KD2h | 6.0 | — | 6.0 | — | 6.0 | — |
| 9 | KD2i | 6.0 | — | 6.0 | — | 6.0 | — |
| 10 | KB2a | 15.0 | 0.3 | 6.0 | — | 6.0 | — |
| 11 | KB2b | 6.0 | — | 6.0 | — | 6.0 | — |
| 12 | KB2c | 6.0 | — | 6.0 | — | 6.0 | — |
| 13 | KB2d | 6.0 | — | 6.0 | — | 6.0 | — |
| 14 | Chloramphenicol (2.0 µg mL−1) | 24.0 ± 1.5 | 0.002 | 22.0 ± 1.5 | 0.002 | 18.0 ± 0.5 | |
Among the tested compounds, KD2f and KB2a demonstrated the most pronounced antibacterial activity against E. coli, producing inhibition zones of 9.0 and 15.0 mm, respectively, with corresponding MIC values of 1.0 and 0.3 mg mL−1. The preferential activity toward E. coli may indicate that these derivatives interact with biological targets more relevant to Gram-negative bacteria or possess physicochemical properties that facilitate penetration through the outer membrane composed of lipopolysaccharides. In contrast, the halogenated derivatives KD2c (4-F) and KD2d (2,4-diCl) showed the strongest antifungal activity against C. albicans, with inhibition zones of 10.0 mm and MIC values of 0.50 mg mL−1. The incorporation of halogen substituents is known to increase molecular lipophilicity, which may enhance the ability of compounds to interact with or penetrate fungal cell membranes enriched with ergosterol.41
In the asymmetric derivative series (KB), the order of activity is determined as KB2d > KB2a > KB2b > KB2c. Of these, KB2d (containing the combination R2,3 = methylenedioxy and R5 = NO2) is the most potent compound in the entire series with an IC50 = 1.54 ± 0.35 µM. This value is close to the reference drug ellipticine (IC50 = 0.34 µM) and is about 10–15 times more potent than natural curcumin (15–30 µM).42 SAR analysis shows that the superiority of the KB series stems from synergistic electronic effects. The combination of the strongly electron-withdrawing nitro group and the hydrophobic methylenedioxy bond creates a pronounced electronic asymmetry, enhancing molecular polarization, thereby facilitating interactions with biological targets in tumor cells.43,44
The superior pharmacological efficacy of asymmetric (KB) compounds compared to symmetric (KD) analogs can be attributed to intramolecular push–pull electronic effects. In these asymmetric structures, an electron-donating group (EDG), such as the methylenedioxy group found in KB2d, is located on a distant aromatic ring, while an electron-withdrawing group (EWG), like the nitro group, is positioned at the opposite end. This arrangement creates a significant electronic gradient along the π-conjugated pentadienone system.
This push–pull structure promotes extensive intramolecular charge transfer (ICT), resulting in considerable molecular polarization. From a pharmaceutical chemistry standpoint, this increased polarity elevates the dipole moment, which could enhance electrostatic interactions and hydrogen bonding with amino acid residues in the binding pockets of target proteins, such as aromatase (3EQM). Additionally, the asymmetry reduces the molecular rigidity typically seen in symmetrical curcuminoids, allowing for greater conformational flexibility and a better fit within the hydrophobic clefts of bioreceptors.
This electronic fine-tuning helps explain why KB2d, with its robust push–pull system, demonstrates ten times greater antiproliferative activity against MCF-7 cells compared to natural curcumin. The arrangement of the LUMO and HOMO energy levels, influenced by this asymmetric electronic distribution, likely facilitates more effective covalent or non-covalent interactions with intracellular nucleophiles, thereby enhancing its overall therapeutic efficacy.
A key factor making KB2d promising is its ability to maintain efficacy at low concentrations (submicromolar). At a concentration of 0.8 µM, while other potent samples such as KD2f or KB2a only showed negligible inhibition (<10%), KB2d maintained an impressive rate of cell death (only 8.17 ± 0.46% of cells survived). Notably, at fixed concentrations (4 µM and 0.8 µM), KB2d even showed higher inhibition percentages than ellipticine. This difference suggests that although ellipticine has a lower overall IC50, KB2d possesses a more favorable dose–response curve (Table 2). This provides compelling experimental grounds for selecting KB2d as a representative candidate for subsequent docking simulations, molecular dynamics (MD), and drug-like characterization studies. Furthermore, although the reference drug ellipticine has a lower absolute IC50 (0.34 µM) than the most potent synthesized compound, KB2d (1.54 µM), the preference for the diarylpentanoid framework aligns with the core principles of modern targeted drug design. Ellipticine is a highly planar alkaloid that acts as a non-selective DNA intercalator and Topoisomerase II inhibitor, a non-discriminatory mechanism that frequently causes severe adverse effects in normal tissues and has historically hindered the clinical advancement of its derivatives. In contrast, KB2d's asymmetric MAC framework features a non-planar, conformationally adaptable structure that is not conducive to DNA intercalation. Molecular docking and MD simulations suggest selective affinity for the aromatase active site (3EQM), supporting a targeted enzyme-inhibition hypothesis for its antiproliferative effect against the ER-positive MCF-7 cell line. This targeted mode of action is expected to confer a broader therapeutic window and a more favorable safety profile than non-selective cytotoxic agents. Consequently, the combination of low-micromolar potency with a putative enzyme-targeted, non-DNA-intercalating mechanism positions KB2d as a more viable and structurally optimizable lead candidate for further development. However, this mechanistic hypothesis is derived from computational evidence and requires experimental validation through direct CYP19A1 enzyme inhibition assays and DNA intercalation displacement studies in future work.
| Entry | Comp | Ia (%) (0.8 µM) | IC50 (µM) |
|---|---|---|---|
| a I (%) (0.8 µM): inhibition percentage at a solution concentration of 0.8 µM. | |||
| 1 | KD2a | 5.39 ± 0.34 | 8.65 ± 0.51 |
| 2 | KD2b | 2.97 ± 0.21 | 28.25 ± 0.97 |
| 3 | KD2c | 2.34 ± 0.28 | 33.27 ± 2.02 |
| 4 | KD2d | 8.28 ± 0.40 | 2.81 ± 0.22 |
| 6 | KD2f | 9.45 ± 0.61 | 2.16 ± 0.28 |
| 7 | KD2g | 2.93 ± 0.26 | >100 |
| 8 | KD2h | 13.67 ± 0.98 | 2.57 ± 0.33 |
| 9 | KD2i | 1.05 ± 0.13 | 61.29 ± 2.73 |
| 10 | KB2a | 2.61 ± 0.29 | 2.54 ± 0.24 |
| 11 | KB2b | 6.98 ± 0.58 | 3.06 ± 0.20 |
| 12 | KB2c | 16.20 ± 0.66 | 3.96 ± 0.28 |
| 13 | KB2d | 33.36 ± 0.91 | 1.54 ± 0.35 |
| 14 | Ellipticine | 22.73 ± 1.09 | 0.34 ± 0.02 |
The choice of 3EQM as a molecular docking target is biologically relevant in the context of MCF-7, as 3EQM represents the crystal structure of human placental aromatase (CYP19A1) in complex with androstenedione. This structure provides a significant model for understanding estrogen biosynthesis. Aromatase is the only enzyme in vertebrates that converts androgens into estrogens, and inhibitors of aromatase are a key therapeutic approach for treating estrogen-dependent breast cancer.
This relevance is particularly important for MCF-7, which serves as a classic model of estrogen-responsive and estrogen receptor-positive (ER-positive) breast cancer, where the cancerous cell proliferation is inhibited by reduced endocrine signaling or by treatment with anti-estrogen drugs. However, it should be noted that the use of 3EQM in this study is proposed as a hypothetical mechanism focusing on the aromatase–estrogen axis, and it cannot yet be regarded as conclusive evidence that CYP19A1 is the only intracellular target of KB2d in MCF-7 cells.
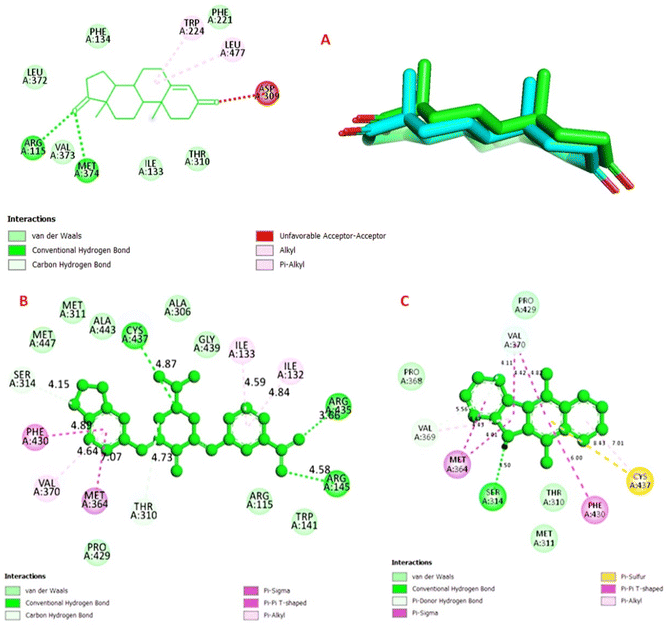
The redocking results of the co-crystallized ligand on 3EQM yielded a heavy atom root mean square deviation (RMSD) of 0.664 Å (Fig. 3A). This confirms that the docking parameters used are capable of reliably reproducing the experimental binding posture. Building on this confirmed foundation, KB2d demonstrates a highly favorable binding posture (pose 192), with a predicted binding energy (ΔGbind) of −10.96 kcal mol−1 and an estimated inhibition constant (Ki) of 9.18 nM (Table 3).
| Entry | Compounds | Predicted binding energy ΔGbind (kcal mol−1) | Estimated inhibition constant Ki (nM) |
|---|---|---|---|
| 1 | KB2d | −10.96 | 9.18 |
| 2 | Ellipticine | −8.48 | 612.04 |
The 2D interaction map (Fig. 3B) illustrates that KB2d establishes a multi-point contact network in the active region. On the right side of the molecule, the ligand forms polar contacts through hydrogen bonds with basic residues such as Arg435 and Arg145, and the central nitrogen atom also forms a hydrogen bond with Cys437. In the central and left regions, the residues Ser314, Thr310, Cys437, Met364, Val370, Ile132, Ile133, and Phe430 contribute to ligand fixation through a combination of weakly polarized contacts, aromatic packing, and van der Waals interactions.
The structural and activity interpretation from this model is quite consistent: the relatively flat aromatic framework, combined with the nitro group and the methylenedioxy ring, enables KB2d to achieve both electron complementation and to take advantage of the hydrophobic-aromatic environment of the attachment cavity. This may explain why KB2d is the most potent sample in MCF-7 in vitro testing.
Conversely, the docking results for ellipticine at 3EQM were less favorable (pose 380), with ΔGbind = −8.48 kcal mol−1 and Ki = 612.04 nM (Table 3). The interaction map (Fig. 3C) displays a less diverse binding pattern, primarily dominated by hydrophobic and aromatic contacts. Notably, this less favorable outcome on 3EQM does not contradict the fact that ellipticine remained more potent in the MCF-7 cell model. The difference between cellular potency and docking score likely reflects variations in the mechanism of action: KB2d seems to be better suited for the aromatase-oriented binding site, while ellipticine exhibits MCF-7 cytotoxicity through other intracellular targets or multiple mechanisms that are not fully captured by the current aromatase-oriented docking model.
The RMSD plot indicates that the protein backbone oscillated mainly around 1.4–2.0 Å throughout the 200 ns simulation, demonstrating that the structural core of 3EQM was well-preserved in the simulated environment (Fig. 4A). The ligand KB2d exhibited a more pronounced oscillation in the early stages, with several peaks around 4–5 Å in the first 20 ns; however, this decreased and stabilized around 1.5–2.5 Å for most of the remainder of the simulation. This pattern is typical for a ligand undergoing initial relaxation and posture optimization, ultimately achieving a relatively stable binding state. Therefore, the RMSD data indicate that KB2d was not ejected from the docking site; rather, it suggests that the initial docking pose was refined during molecular dynamics (MD) simulations, reaching a more favorable equilibrium state.
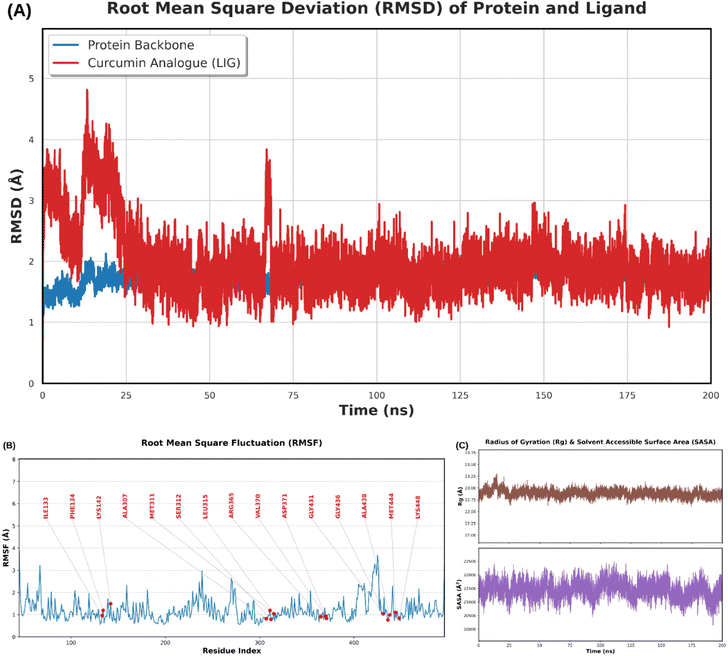 | ||
| Fig. 4 RMSD of backbone protein and KB2d ligand over 200 ns (A), RMSF according to protein residue (B), radius of gyration (Rg) and solvent-accessible surface area (SASA) (C). | ||
The root mean square fluctuation (RMSF) data of the protein (Fig. 4B) show that most residues fluctuate only moderately, with some local peaks occurring at loop regions or chain ends. Importantly, residues associated with the binding cavity generally do not fall within regions of maximum fluctuation, indicating a relatively stable environment around the ligand. Additionally, two structural indices, the radius of gyration (Rg) and solvent-accessible surface area (SASA), do not exhibit unusual fluctuations (Fig. 4C). Rg remains stable within a narrow range, suggesting that the protein does not become stretched or collapse; SASA fluctuates within a moderate amplitude, consistent with a well-equilibrated system in water. The combination of these four indices leads us to conclude that the 3EQM–KB2d complex is stable throughout the 200 ns simulation.
Fig. 5A illustrates that the number of hydrogen bonds between the protein and ligand typically varies from 1 to 2, occasionally increasing to 3 or 4. Kinetically, this model is reasonable for a stable protein–ligand complex, as the system does not need to maintain numerous hydrogen bonds continuously. Instead, one stable anchor bond, complemented by a few transient secondary bonds, is sufficient to retain the ligand within the binding pocket.
 | ||
| Fig. 5 Number of hydrogen bonds over time (A), timeline of donor–acceptor pairs (B), occupancy of principal hydrogen bonds (C). | ||
Fig. 5(B and C) indicate that the hydrogen bond between TRP97-NE1 and LIG453-O1 is the most prominent, with an occupancy of 73.7%, significantly surpassing all other pairs. Other notable interactions include ARG331-NE with LIG453-O (4.2%), TRP97-NE1 with LIG453-O (2.2%), ARG331-NE with LIG453-O1 (1.8%), ARG101-NE with LIG453-O1 (1.3%), SER270-OG with LIG453-O3 (1.0%), along with several weaker interactions.
This data leads to a clear conclusion: Trp97 serves as the primary hydrogen bond anchor residue in KB2d throughout its entire trajectory. This finding presents a significantly stronger result than static docking, as it demonstrates a directional interaction that can be maintained for the majority of the simulation time, rather than occurring only in transient conformations.
This data leads to a clear conclusion: Trp97 serves as the primary hydrogen bond anchor residue in KB2d throughout its entire trajectory. This finding presents a significantly stronger result than static docking, as it demonstrates a directional interaction that can be maintained for the majority of the simulation time, rather than occurring only in transient conformations.
This interpretation is strongly supported by the protein–ligand contact timeline (Fig. 6A and B), which indicates that KB2d is surrounded by a stable and extensive interaction layer. Many residues exhibit very high contact frequencies, including Thr310 (100.0%), Cys437 (100.0%), Arg435 (99.9%), Ile133 (99.4%), Phe430 (99.4%), Met311 (99.3%), Ala443 (95.9%), Ser314 (94.1%), Met364 (92.5%), Ala306 (91.5%), Met447 (91.1%), and Val370 (90.0%). This pattern suggests that KB2d is not stabilized by a single residue or a specific type of interaction; rather, it relies on a coordinated network of hydrophobic, aromatic, and polar interactions. Additionally, the incomplete overlap between the static contacts observed during docking and the dominant interactions identified in molecular dynamics (MD) simulations indicates that the ligand underwent a refinement of its initial docking position. This refinement led to a more favorable orientation, converging to a kinematically optimal binding microstate rather than remaining “frozen” in its initial conformation.
 | ||
| Fig. 6 Timeline of protein–ligand contact with frequency > 5% (A), occupancy of residues in contact with KB2d (B). | ||
The MM/PBSA analysis revealed that the total binding energy consistently remained in the negative range, fluctuating between −30 and −40 kcal mol−1 across the analyzed frames (Fig. 7A). The most significant favorable contribution came from van der Waals interactions, while electrostatic energy also supported the positive binding dynamics. On the other hand, polarized solvation components were unfavorable, which is commonly observed when a ligand of a specific polarity enters an organized binding cavity.
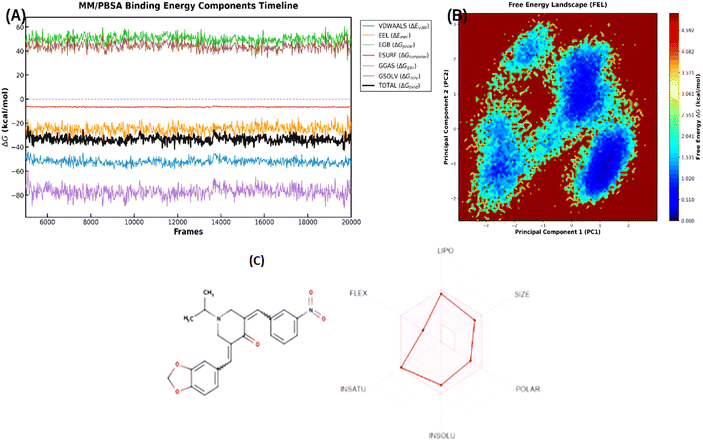 | ||
| Fig. 7 Evolution of MM/PBSA binding energy components over time (A), free energy landscape (FEL) according to PC1 and PC2 (B), SwissADME profile and drug-likeness of KB2d (C). | ||
The free-energy landscape map (Fig. 7B) demonstrates that the system is not concentrated in a single minimum; instead, it is distributed across multiple low-energy basins. This indicates that the 3EQM–KB2d complex exists as a collection of closely spaced semi-stable microconformations. Rather than diminishing the model's strength, this characteristic supports the contemporary perspective that a ligand can bind stably while maintaining controlled conformational adaptability.
In silico ADME prediction results showed that KB2d possesses a relatively balanced drug-like profile (Fig. 7C). The compound falls within the acceptable physicochemical parameter range for a small molecule anticancer lead, with a molecular weight of 406.43 g mol−1, a polar surface area (TPSA) of 84.59 Å2, a consensus log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of 3.33, and only four rotational bonds. SwissADME predicted high gastrointestinal absorption, no crossing of the blood–brain barrier, and no interaction with P-glycoprotein, suggesting suitable membrane permeability with minimal risk of central nervous system exposure.
P of 3.33, and only four rotational bonds. SwissADME predicted high gastrointestinal absorption, no crossing of the blood–brain barrier, and no interaction with P-glycoprotein, suggesting suitable membrane permeability with minimal risk of central nervous system exposure.
However, there are concerns regarding average solubility and potential inhibition of CYP2C19, CYP2C9, and CYP3A4, as well as PAINS/Brenk warnings related to the enone-type Michael acceptor and nitro group. These factors indicate that while KB2d shows promise as a bioactive lead, it should not yet be considered a fully optimized drug candidate. These limitations do not detract from the current value of the molecule; rather, they highlight the need for further pharmaceutical optimization in subsequent steps.
In summary, all available data support KB2d as the most promising antiproliferative lead in the series of synthesized compounds on the MCF-7 line. In vitro data place KB2d in the low micromolar activity region and indicate that the nitro–methylenedioxy motif is the most favorable pharmacophore within this structural framework class. The validated docking model shows that KB2d has strong geometric and energy compatibility with the aromatase target 3EQM, while MD and MM/PBSA analyses demonstrate that this binding is kinetically stable and thermodynamically favorable over a sufficiently long simulation period. Finally, the predictive ADME profile suggests that KB2d lies within a chemical space that can be further optimized, although some structural liabilities remain to be addressed. Overall, KB2d can be considered a potential breast cancer prototype, having both experimental cell-based evidence and being supported by appropriate computational data, making it worthy of further in-depth study on the MCF-7 model. In summary, all available data indicate that KB2d is the most promising antiproliferative compound among the series of synthesized compounds tested on the MCF-7 cell line. In vitro studies place KB2d in the low micromolar activity range and suggest that the nitro–methylenedioxy motif is the most favorable pharmacophore within this class of structures. The validated docking model shows that KB2d has strong geometric and energetic compatibility with the aromatase target 3EQM. Additionally, molecular dynamics (MD) and molecular mechanics/Poisson–Boltzmann surface area (MM/PBSA) analyses demonstrate that this binding is kinetically stable and thermodynamically favorable over a sufficiently long simulation period. Finally, the predictive ADME profile suggests that KB2d exists within a chemical space that can be further optimized, even though some structural liabilities need to be addressed. Overall, KB2d can be considered a potential prototype for breast cancer treatment, supported by both experimental cell-based evidence and relevant computational data, warranting further in-depth study in the MCF-7 model.
The results (Table 4) showed that MAC compounds had a broad range of NO inhibitory activity. Notably, asymmetric derivatives (KB chains) demonstrated significantly stronger inhibitory effects compared to symmetric derivatives (KD chains). Specifically, compound KB2c was the most potent among those tested, with an IC50 of 2.51 ± 0.10 µM. This value is roughly 4.8 times lower than that of dexamethasone, a known positive control, which has an IC50 of 12.11 ± 0.98 µM. Additionally, compounds KD2b, KD2h, KD2i, and KB2d also exhibited inhibitory activity at low micromolar concentrations, indicating that the isopropylpiperidone framework is a promising structure for developing anti-inflammatory drugs.
| Entry | Comp | IC50 | Cell viability at 20 µM (%) |
|---|---|---|---|
| 1 | KD2a | NA | 51.92 ± 1.12 |
| 2 | KD2b | 3.29 ± 0.45 | 81.26 ± 1.16 |
| 3 | KD2c | 11.69 ± 0.47 | 98.52 ± 2.14 |
| 4 | KD2d | 7.09 ± 0.50 | 90.14 ± 2.02 |
| 5 | KD2e | NA | 66.91 ± 0.86 |
| 6 | KD2f | NA | 8.01 ± 0.42 |
| 7 | KD2g | NA | 63.83 ± 0.77 |
| 8 | KD2h | 3.30 ± 0.40 | 76.92 ± 1.72 |
| 9 | KD2i | 4.08 ± 0.12 | 97.87 ± 1.82 |
| 10 | KB2a | NA | 42.31 ± 1.68 |
| 11 | KB2b | NA | 32.04 ± 1.44 |
| 12 | KB2c | 2.51 ± 0.10 | 76.47 ± 1.87 |
| 13 | KB2d | 4.63 ± 0.13 | 91.63 ± 1.24 |
| 14 | Dexa. | 12.11 ± 0.98 | 99.35 ± 1.07 |
To ensure that the reduction in NO production was not due to cytotoxicity, we evaluated the compounds alongside cell survival rates. At 20 µM (about eight times the IC50), cells treated with KB2c maintained a survival rate of 76.47 ± 1.87%. This suggests that the decrease in NO levels was due to regulation of inflammatory signaling pathways rather than cytotoxic effects on macrophages. However, at higher concentrations (100 µM), KB2c began to significantly decrease cell viability. This indicates that although KB2c is a potent “lead compound,” further structural modifications are needed to improve the safety margin for cell viability.
Preliminary structure–activity relationship (SAR) analysis revealed that combining the hydrophobic chloride substituent (R1 = Cl) with the methylenedioxy bridge (R2 = –O–CH2–O–) greatly enhances anti-inflammatory activity. Replacing the hydrophobic chloride group in KB2c with a polar nitro group (KB2d) nearly doubled its IC50 (from 2.51 ± 0.10 µM to 4.63 ± 0.13 µM). This indicates that the enzyme's binding pocket prefers hydrophobic interactions over polar ones. Therefore, strategically placing hydrophobic substituents on the aromatic ring will be key to achieving optimal inhibitory activity.
The structure-activity relationship (SAR) analysis based on nitric oxide (NO) inhibition data provided valuable insights into the factors that influence the efficacy of compounds within the KB framework. Our findings indicate that the combination of a methylenedioxy ring with a chlorinated aryl nucleus, as seen in the structure of KB2c, creates a particularly effective substituent for NO inhibition activity. Specifically, KB2c achieved an inhibition efficiency of 96.88 ± 1.70% at a concentration of 20 µM and maintained an inhibition level of 64.38 ± 1.26% at a concentration of 4 µM. This level of inhibition was significantly higher than that of KB2d at the same concentration.
Although KB2a and KB2b also demonstrated high percentage inhibition at certain concentrations, the lack of sufficient data to determine their IC50 values prevents us from directly comparing their efficacy with that of KB2c on the same quantitative scale.
Overall, the data suggest that compounds with strongly modulating substituents, such as chlorine (Cl), nitro (NO2), methylenedioxy, and methoxy, generally exhibit better activity than other samples. This indicates that the electronic effects, along with the geometric complementarity of the aromatic-enone framework, are key factors influencing the anti-inflammatory effects in this series.
From an experimental standpoint, KB2c emerges as an ideal candidate for further investigation, including molecular docking analysis, molecular dynamics studies, and predictions of drug-likeness properties. The potential of KB2c in developing novel anti-inflammatory agents is substantial, and we will explore this further in our upcoming studies.
The results from redocking the cocrystallized ligand into the 4WCU construct yielded a heavy atom root mean square deviation (RMSD) value of 0.470 Å (Fig. 8). This low RMSD value indicates that the chosen docking parameters can accurately reproduce the experimental binding posture with high reliability. Additionally, the molecular docking results obtained support the activity trend observed in the in vitro experiments.
In this study, the target protein selected was the X-ray crystal structure of human cAMP-specific 3′,5′-cyclic phosphodiesterase 4D (PDE4D) (PDB code: 4WCU), which is in complex with an inhibitor and has a resolution of 2.35 Å.45 The selection of PDE4D/4WCU as a target for the anti-inflammatory model is based on strong biological rationale, as PDE4 is a key enzyme responsible for cAMP hydrolysis and plays a central role in regulating inflammatory signaling. Inhibiting PDE4 results in increased intracellular cAMP levels, leading to reduced inflammatory responses in various immune cell types.46,47 Furthermore, the 4WCU construct has been referenced in studies aimed at developing topical PDE4 inhibitors for the treatment of atopic dermatitis,45 which further supports its suitability as a target in the search for novel anti-inflammatory agents.
The docking results for KB2c at pose 392 indicated a predicted binding energy (ΔGbind) of −10.78 kcal mol−1 and an inhibition constant (Ki) of 12.49 nM (Table 5). These values demonstrate a highly favorable binding affinity of KB2c for the target protein 4WCU/PDE4D. An analysis of the docking energy components revealed that the primary contributions to the binding energy stemmed from van der Waals forces (vdW), hydrogen bonding (H-bond), and desolvation, while electrostatic interactions played a negligible role. This suggests that KB2c's binding is largely influenced by optimal spatial fit within the binding cavity, supported by hydrophobic and aromatic interactions, as well as some directional hydrogen bonds, rather than strong electrostatic forces.
| Entry | Compounds | Predicted binding energy ΔGbind (kcal mol−1) | Estimated inhibition constant Ki (nM) |
|---|---|---|---|
| 1 | KB2c | −10.78 | 12.49 |
| 2 | Dexamethasone | −7.79 | 1960 |
The 2D interaction map illustrates that KB2c is anchored within the binding cavity through a network of multiple contacts with key amino acid residues, including Asp318, Leu319, Asn321, Tyr329, Ile336, Phe340, Phe372, Met273, Met357, and Ile376. This network of interactions generates a binding pattern with both chemical and biological significance. The structure of KB2c is particularly well-suited to the binding cavity due to its relatively extensive aromatic ring system, moderate hydrophobicity, and the presence of oxygen atoms that can participate in hydrogen bonding. This allows KB2c to be stabilized by the surrounding hydrophobic environment while remaining properly oriented through local polar interactions.
In contrast, dexamethasone at pose 21 only achieved a ΔGbind of −7.79 kcal mol−1 and a Ki of 1960 nM (Table 5), indicating a significantly lower binding affinity compared to KB2c within the same docking system. Although dexamethasone formed some compatible interactions with the binding cavity, its overall geometric-energy compatibility was inferior, as evidenced by weaker intermolecular energies and higher conformational costs upon binding.
This difference in binding affinity is significant and aligns with the observed experimental trend: in the NO inhibition assay, KB2c demonstrated stronger activity than dexamethasone in terms of IC50 value. In other words, the docking results not only provided a favorable binding energy value but also accurately reflected the difference in activity between the lead compound (KB2c) and the positive control. The correlation between in vitro and docking data strongly supports the hypothesis that the anti-inflammatory efficacy of KB2c is directly linked to its superior interaction with the target PDE4D (4WCU) chosen for this study.45–47
Molecular dynamics (MD) simulations conducted over 200 ns revealed the dynamic binding behavior of the protein–KB2c complex while maintaining the thermodynamic properties of the complex. Specifically, the Root Mean Square Deviation (RMSD) of the protein backbone (Fig. 9A) showed a slight increase from the beginning, stabilizing within the range of 3.2–3.8 Å. This indicates that the protein undergoes conformational changes upon binding to the ligand but still achieves a relatively stable dynamic equilibrium.
 | ||
| Fig. 9 Structural stability and dynamic behavior of the protein–KB2c complex during 200 ns MD simulation. | ||
In contrast, the ligand's RMSD increased significantly after approximately 50 ns and fluctuated steadily in the range of 6.6–7.2 Å for the remainder of the simulation. Based solely on the ligand's RMSD, one might easily conclude that the ligand was destabilized. However, when considered alongside other indicators, a more plausible interpretation is that KB2c has transitioned from its initial docking pose to an alternative binding state or a more stable micromorphism within the binding cavity, rather than completely detaching from the protein.
This finding is further supported by the root mean square fluctuation (RMSF) data of the protein, which show that large fluctuations are primarily concentrated in certain mobile regions. These fluctuations do not indicate an overall structural collapse of the system. Additionally, the radius of gyration (Rg) and the solvent-accessible surface area (SASA) remained almost constant within a narrow range of fluctuation, confirming that the overall structure of the protein is preserved throughout the simulation (Fig. 9B and C).
Hydrogen bond analysis indicated that KB2c is not stabilized by a dense network of hydrogen bonds, but rather through several localized, directional hydrogen interactions. The most prominent of these is the bond between GLN273-NE2 and LIG323-O1, which has an occupancy rate of 21.2%. Other bonding pairs, such as SER272-OG–LIG323-O, GLN273-NE2–LIG323-O2, and TYR233-OH–LIG323-O1, contribute only at a secondary level (Fig. 10). The relatively low occupancy rates of these hydrogen bonds align with the dynamic nature of the system: the ligand is not “locked” in a single configuration but continually adjusts its position and orientation within the binding cavity. However, the presence of hydrogen bonds with Gln273 remains significant. This suggests that even after detaching from its original docking site, KB2c retains at least one specific anchor point to the target protein. Consequently, despite the increased RMSD of the ligand, the system does not exhibit signs of complete de-binding.
The significance of the dynamic binding model is further highlighted through an analysis of protein–ligand contact timelines. In Fig. 11, KB2c shows very high contact frequencies with a variety of residues, including TYR159, HIS160, MET273, LEU319, ASN321, PRO322, TYR329, ILE336, GLU339, PHE340, GLN343, VAL365, SER368, GLN369, PHE372, ILE373, and VAL377. Many of these residues have an occupancy exceeding 80–95%, indicating that the ligand is surrounded by a stable and extensive interaction “shell”.
This data is extremely important because it suggests that an increase in the root mean square deviation (RMSD) of the ligand should not be interpreted as a failure of the docking model. Instead, it represents a reorganization of the system into a different binding state that still maintains rich interactions. In other words, while KB2c may no longer retain its original binding conformation, it remains within the associated protein region and continues to interact closely with many key residues. For relatively planar aromatic ligands like KB2c, this mechanism of “stable but flexible binding” is entirely chemically plausible.
MM/PBSA and free-energy landscape analyses further support the favorable nature of the post-reordering binding. In Fig. 12A, the total binding energy (ΔGbind) consistently remains in the negative range, fluctuating mainly between −25 and −35 kcal mol−1. This indicates that the ligand remains thermodynamically bound to the protein throughout the analysis. The primary contributor to this binding energy is van der Waals forces, while electrostatic energy contributes little, and may even have a slight negative effect. This suggests that hydrophobic packing and aromatic stabilization are the dominant driving forces in the formation of the protein–KB2c complex.
In Fig. 12B, the free-energy landscape does not present a single minimum; rather, it displays multiple low-energy basins. This reflects that the system exists as a collection of stable or semi-stable bound microstructures. These results are fully consistent with RMSD, hydrogen bond occupancy, and contact timeline analyses. This allows us to conclude that KB2c forms a thermodynamically stable yet positionally flexible complex, indicating a highly adaptable binding behavior within the binding cavity.
SwissADME predictions indicate that KB2c has a relatively favorable drug-like profile as a small molecule lead; however, further optimization is still necessary (Fig. 13). In terms of physicochemical properties, KB2c has a molecular weight (MW) of 395.88 g mol−1, a topological polar surface area (TPSA) of 38.77 Å2, a consensus log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of 4.46, four hydrogen bond acceptors, no hydrogen bond donors, and only three rotational bonds. These parameters suggest that KB2c has a lipophilic tendency, low mobility, and a low polar surface area, which are advantageous for passive diffusion across membranes. Indeed, SwissADME predicts that KB2c has high gastrointestinal (GI) absorption, is not a substrate of P-glycoprotein (P-gp), and can cross the blood–brain barrier. However, these membrane-penetrating characteristics also pose some disadvantages: low water solubility, the risk of non-selective distribution, and the potential for off-target effects on the central nervous system if developed as a peripheral anti-inflammatory drug. Additionally, the molecule is predicted to inhibit multiple cytochrome P450 isoenzymes, including CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4, which raises concerns about drug–drug interactions. The presence of PAINS/Brenk alerts related to the ene-one/Michael acceptor structure suggests that KB2c should be evaluated cautiously for the risk of non-specific reactions. Therefore, while KB2c is a highly promising lead in terms of activity, it cannot yet be regarded as a fully optimized drug candidate.
P of 4.46, four hydrogen bond acceptors, no hydrogen bond donors, and only three rotational bonds. These parameters suggest that KB2c has a lipophilic tendency, low mobility, and a low polar surface area, which are advantageous for passive diffusion across membranes. Indeed, SwissADME predicts that KB2c has high gastrointestinal (GI) absorption, is not a substrate of P-glycoprotein (P-gp), and can cross the blood–brain barrier. However, these membrane-penetrating characteristics also pose some disadvantages: low water solubility, the risk of non-selective distribution, and the potential for off-target effects on the central nervous system if developed as a peripheral anti-inflammatory drug. Additionally, the molecule is predicted to inhibit multiple cytochrome P450 isoenzymes, including CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4, which raises concerns about drug–drug interactions. The presence of PAINS/Brenk alerts related to the ene-one/Michael acceptor structure suggests that KB2c should be evaluated cautiously for the risk of non-specific reactions. Therefore, while KB2c is a highly promising lead in terms of activity, it cannot yet be regarded as a fully optimized drug candidate.
In summary, KB2c stands out as the most significant anti-inflammatory lead among the isopropylpiperidone derivatives investigated, based on both its nitric oxide (NO) inhibitory efficacy and the computational model supporting it. Experimental data place KB2c in the low micromolar activity range, demonstrating stronger activity than the positive control dexamethasone under the same experimental conditions. Docking results show that KB2c exhibits excellent geometric and energetic compatibility with the protein 4WCU, which corresponds to cAMP-specific 3′,5′-cyclic phosphodiesterase 4D (PDE4D). PDE4 is a well-established anti-inflammatory target, as it regulates cAMP metabolism, and its inhibition is strongly linked to anti-inflammatory effects. Moreover, the availability of a crystallographically resolved PDE4–inhibitor complex structure (4WCU) enhances KB2c's value as a suitable docking model for PDE4 inhibitor design studies. Additionally, molecular dynamics (MD), hydrogen bond, contact occupancy, and MM/PBSA analyses indicate that, although the ligand undergoes repositioning within the binding site, the system stabilizes in a thermodynamically favorable binding state characterized by rich specific interactions.
From a pharmacochemical standpoint, KB2c is situated in a favorable structural space for further optimization, although enhancements are needed in solubility, cytochrome P450 liability, and potential safety concerning the Michael acceptor. Overall, KB2c not only demonstrates high bioactivity but also serves as a valuable prototype for further development into novel anti-inflammatory agents, especially if optimized to reduce cytotoxicity and improve pharmacokinetic selectivity.
4. Conclusion
In this study, we successfully designed and synthesized a series of curcuminoid monoketones based on the N-isopropyl-4-piperidone framework. The sequential Claisen–Schmidt condensation strategy proved effective, allowing for the synthesis of both symmetric and asymmetric diarylpentanoids with generally good to excellent yields and high structural purity. This synthesis platform enables systematic tuning of intramolecular electron distribution and spatial environment, facilitating meaningful analyses of structure-activity relationships.Biological screening revealed that incorporating structural asymmetry—especially when combined with electron-donating substituents—significantly improved pharmacological efficacy. Among the synthesized derivatives, compound KB2d, which features a nitro-methylenedioxy “push–pull” electron system, exhibited the most potent cytotoxic activity against MCF-7 breast cancer cells, with an IC50 value of 1.54 ± 0.35 µM. This represents a significant enhancement in efficacy compared to natural curcumin. Moreover, the dose–response profile was favorable when compared to the reference drug ellipticine.
Additionally, compound KB2c demonstrated strong inhibition of nitric oxide production in LPS-stimulated macrophages (IC50 = 2.51 ± 0.10 µM), suggesting promising anti-inflammatory potential with an acceptable safety margin. In antimicrobial evaluations, KB2a exhibited selective activity against Escherichia coli, with a minimum inhibitory concentration (MIC) of 0.3 mg mL−1, indicating its relevance to Gram-negative bacterial targets.
Insights into the mechanism were gained through molecular coupling, molecular dynamics simulations at long timescales (200 ns), and MM-PBSA binding free energy calculations, further supporting the experimental observations. The asymmetric ligand KB2d forms a stable yet conformationally adaptable complex within the aromatase active site (PDB code: 3EQM). This complex is characterized by low ligand RMSD fluctuations and a favorable binding free energy (ΔGbind ≈ −33.5 kcal mol−1), arising from a network of hydrophobic and coordinate polarization interactions.
Overall, these findings demonstrate that incorporating controlled electronic asymmetry into piperidone-based diarylpentanoid frameworks is an effective pharmacochemical strategy. This approach enhances bioactivity while addressing inherent limitations associated with curcumin. Consequently, the compounds led by KB2d and KB2c have been identified as promising drug-like prototypes for further optimization, including improvements in metabolic stability, water solubility, and in vivo pharmacological validation, towards the development of targeted anticancer and anti-inflammatory agents. While the current study successfully established the initial structure-activity relationships of the synthesized MACs with high structural purity suitable for early-stage in vitro screening, we acknowledge that comprehensive impurity profiling is essential for pharmaceutical development. As the promising lead compounds (KB2d and KB2c) advance toward pre-clinical stages, future investigations will mandatorily encompass systematic impurity profiling, the isolation of trace synthetic by-products, and the rigorous toxicological characterization of these impurities in strict accordance with pharmaceutical regulatory guidelines (e.g., ICH Q3A/Q3B) to ensure a comprehensive safety margin.
Author contributions
Huy-Khoa Tran: investigation, formal analysis, validation, software, originate manuscript, Truc-Vy Mai: investigation, formal analysis, Van-Dung Le, Tran-Nguyen Minh-An, Dinh-Tri Mai, Quoc-Tuan Tran, Thi-Quyen Vu: software, visualization, validation; Thanh-Danh Nguyen: writing–review & editing; Chi-Hien Dang: conceptualization, supervisor, writing–review & editing.Conflicts of interest
The authors of this paper state that they have no competing financial interests or personal relationships that could have influenced the reported work.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra02446g.Acknowledgements
This research is funded by Vietnam Academy of Science and Technology under grant number NCXS02.04/24-25.References
- S. Fuloria, J. Mehta, A. Chandel, M. Sekar, N. N. I. M. Rani, M. Y. Begum, V. Subramaniyan, K. Chidambaram, L. Thangavelu and R. Nordin, Front. Pharmacol, 2022, 13, 820806 CrossRef CAS PubMed.
- M. M. Rahaman, A. Rakib, S. Mitra, A. M. Tareq, T. B. Emran, A. Shahid-Ud-Daula, M. N. Amin and J. Simal-Gandara, Plants, 2020, 10, 63 CrossRef PubMed.
- W. Liu, Y. Zhai, X. Heng, F. Y. Che, W. Chen, D. Sun and G. Zhai, J. Drug Targeting, 2016, 24, 694–702 CrossRef CAS PubMed.
- P. Anand, A. B. Kunnumakkara, R. A. Newman and B. B. Aggarwal, Mol. Pharm., 2007, 4, 807–818 CrossRef CAS.
- R. Tabanelli, S. Brogi and V. Calderone, Pharmaceutics, 2021, 13, 1715 CrossRef CAS PubMed.
- P. E. Hansen, Pharmaceuticals, 2021, 14, 1189 CrossRef CAS PubMed.
- A. A. Adeniyi and J. Conradie, J. Mol. Graphics Modell., 2018, 85, 25–39 CrossRef CAS PubMed.
- Z. Hussain, H. E. Thu, M. W. Amjad, F. Hussain, T. A. Ahmed and S. Khan, Mater. Sci. Eng., C, 2017, 77, 1316–1326 CrossRef CAS PubMed.
- C. Zhao, Z. Liu and G. Liang, Curr. Pharm. Des., 2013, 19, 2114–2135 CAS.
- T. M. Vieira, L. S. Tanajura, V. C. Heleno, L. G. Magalhães and A. E. Crotti, Future Pharmacol., 2024, 4, 54–76 CrossRef.
- A. A. Nagargoje, S. P. Panchgalle, M. M. Siddiqui, M. H. Shaikh, D. M. Dipti and B. B. Shingate, Arch. Pharm., 2025, 358, e70164 CrossRef CAS PubMed.
- N. Negi, G. Chand, D. Kholia, R. Anand, S. K. Upadhyay, G. Tewari and P. Joshi, Pharmacogn. Rev., 2023, 17(34), 247–254 CrossRef CAS.
- Z. H. Loh, I. Jantan, S. Laphookhieo, Z. W. Lai, L. Y. Fong and S. H. Mah, Phytochem. Rev., 2025, 1–21 Search PubMed.
- A. Anthwal, K. Singh, M. SM Rawat, A. K Tyagi, A. Haque, I. Ali and D. S Rawat, Anti-Cancer Agents Med. Chem., 2016, 16, 841–851 CrossRef CAS PubMed.
- Y. Mardianis, C. Anwar and W. Haryadi, Mater. Sci. Forum, 2017, 901, 110–117 Search PubMed.
- M. A. Obregón-Mendoza, W. Meza-Morales, Y. Alvarez-Ricardo, M. M. Estévez-Carmona and R. G. Enríquez, Molecules, 2022, 28, 289 CrossRef PubMed.
- Y.-J. Cheng, C.-W. Li, C.-L. Kuo, T.-L. Shih and J.-J. Chen, Molecules, 2022, 27, 2547 CrossRef CAS PubMed.
- Q. Li, J. Chen, S. Luo, J. Xu, Q. Huang and T. Liu, Eur. J. Med. Chem., 2015, 93, 461–469 CrossRef CAS.
- T. Zhang, P. Zheng, X. Shen, R. Shao, B. Wang, H. Shen, J. Zhang, Y. Xia and P. Zou, Free Radicals Biol. Med., 2019, 141, 93–102 CrossRef CAS PubMed.
- D. R. Jonathan, E. DravidaThendral, M. K. Priya, D. A. Shirmila, A. A. Fathima, R. Yuvashri and G. Usha, J. Mol. Struct., 2023, 1292, 136063 CrossRef CAS.
- J. Li, A. Fu and L. Zhang, Interdiscip. Sci.:Comput. Life Sci., 2019, 11, 320–328 CrossRef CAS PubMed.
- Z. Khanjiwala, A. Khale and A. Prabhu, J. Mol. Graphics Modell., 2019, 93, 107451 CrossRef CAS PubMed.
- T. Hou, J. Wang, Y. Li and W. Wang, J. Chem. Inf. Model., 2011, 51, 69–82 CrossRef CAS PubMed.
- U. Dasmahapatra, C. K. Kumar, S. Das, P. T. Subramanian, P. Murali, A. E. Isaac, K. Ramanathan, B. Mm and K. Chanda, Front. Chem., 2022, 10, 991369 CrossRef CAS PubMed.
- B. Cao, Y. Wang, K. Ding, N. Neamati and Y.-Q. Long, Org. Biomol. Chem., 2012, 10, 1239–1245 RSC.
- Y. Zhang, C. Zhao, W. He, Z. Wang, Q. Fang, B. Xiao, Z. Liu, G. Liang and S. Yang, Drug Des., Dev. Ther., 2014, 373–382 Search PubMed.
- K.-P. T. Dang, N.-V. Ngo, H.-P. Nguyen, T. N.-H. Pham, H. V. Quang and T.-D. Nguyen, Russ. J. Appl. Chem., 2025, 98, 201–212 CrossRef CAS.
- N. T. T. Tu, T. L.-A. Vo, T. T.-T. Ho, K.-P. T. Dang, V.-D. Le, P. N. Minh, C.-H. Dang, V.-T. Tran, V.-S. Dang and T. T. K. Chi, Beilstein J. Nanotechnol., 2023, 14, 781–792 CrossRef CAS PubMed.
- P. Houghton, R. Fang, I. Techatanawat, G. Steventon, P. J. Hylands and C. Lee, Methods, 2007, 42, 377–387 CrossRef CAS.
- S. V. Ramalingam, S. Bakthavatchalam, K. Ramachandran, V. Gnanarani Soloman, A. K. Ajmal, M. K. Al-Sadoon and R. Vinayagam, Antibiotics, 2024, 13, 1193 CrossRef CAS PubMed.
- O. A. Ojedele, N. Hamdi, M. H. Mughram, E. M. Alyami, M. Dauelbait and O. A. Almohammed, BMC Cancer, 2025, 25, 1812 CrossRef CAS PubMed.
- M. D. Tri, N. T. Phat, P. N. Minh, M. T. Chi, B. X. Hao, T. N. M. An, M. Alam, N. Van Kieu, V.-S. Dang and T. T. N. Mai, RSC Adv., 2023, 13, 5324–5336 RSC.
- Z. Digafie, Y. Melaku, Z. Belay and R. Eswaramoorthy, Adv. Pharmacol. Pharm. Sci., 2021, 2021, 6635270 CAS.
- S. Behera, A. Behera, S. K. Mekap, C. C. Behera, A. Kadam and P. K. Mohanty, J. Biomol. Struct. Dyn., 2022, 40, 9931–9947 CrossRef CAS PubMed.
- A. Thakur, P. R. S. Gupta, P. Pathak and A. Kumar, Int. J. ChemTech Res., 2016, 9, 575–588 CAS.
- N. D. Khanh, N. T. H. Anh, N. Van Son, N. T. M. Tho, N. T. N. Thang, T. T. Quyen and T. N. M. An, RSC Adv., 2026, 16, 2342–2362 RSC.
- M. Bugnon, M. Goullieux, U. F. Rohrig, M. A. Perez, A. Daina, O. Michielin and V. Zoete, J. Chem. Inf. Model., 2023, 63, 6469–6475 CrossRef CAS PubMed.
- A. F. Wacha and J. A. Lemkul, J. Chem. Inf. Model., 2023, 63, 4246–4252 CrossRef CAS PubMed.
- J. A. Lemkul, J. Phys. Chem. B, 2024, 128, 9418–9435 CrossRef CAS PubMed.
- A. Daina, O. Michielin and V. Zoete, Sci. Rep., 2017, 7, 42717 CrossRef PubMed.
- D. Parolaro, T. Rubino, D. Vigano, P. Massi, C. Guidali and N. Realini, Curr. Drug Targets, 2010, 11, 393–405 CrossRef CAS PubMed.
- S. Kwiecien, M. Magierowski, J. Majka, A. Ptak-Belowska, D. Wojcik, Z. Sliwowski, K. Magierowska and T. Brzozowski, Int. J. Mol. Sci., 2019, 20, 1477 CrossRef CAS.
- H. M. Faidallah, S. S. Panda, J. C. Serrano, A. S. Girgis, K. A. Khan, K. A. Alamry, T. Therathanakorn, M. J. Meyers, F. M. Sverdrup and C. S. Eickhoff, Bioorg. Med. Chem., 2016, 24, 3527–3539 CrossRef CAS PubMed.
- G. Liang, X. Li, L. Chen, S. Yang, X. Wu, E. Studer, E. Gurley, P. B. Hylemon, F. Ye and Y. Li, Bioorg. Med. Chem. Lett., 2008, 18, 1525–1529 CrossRef CAS PubMed.
- J. Felding, M. D. Sørensen, T. D. Poulsen, J. Larsen, C. Andersson, P. Refer, K. Engell, L. G. Ladefoged, T. Thormann, A. M. Vinggaard, P. Hegardt, A. Søhoel and S. F. Nielsen, J. Med. Chem., 2014, 57(13), 5893–5903, DOI:10.1021/jm500378a.
- H. Li, J. Zuo and W. Tang, Front. Pharmacol, 2018, 9, 1048 CrossRef CAS . https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2018.01048/full.
- T. Fan, W. Wang, Y. Wang, M. Zeng, M. Zeng, Y. Liu, S. Zhu and L. Yang, Front. Pharmacol, 2024, 15, 1407871 CrossRef CAS PubMed . https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2024.1407871/full.
| This journal is © The Royal Society of Chemistry 2026 |