 Open Access Article
Open Access ArticleIntegrated synthesis, antiproliferative potential, and in silico analysis of novel pyrazole-based hydrazones derived from 3-(2-(4-chlorophenyl)hydrazineylidene)-5-phenylfuran-2(3H)-one
Sayed K. Ramadan *a,
Mariam S. H. Kazema,
Mahmoud Kamal
*a,
Mariam S. H. Kazema,
Mahmoud Kamal b,
Ahmed I. Hashema,
Hisham S. M. Abd-Rabboh
b,
Ahmed I. Hashema,
Hisham S. M. Abd-Rabboh c and
Wael S. I. Abou-Elmagda
c and
Wael S. I. Abou-Elmagda
aChemistry Department, Faculty of Science, Ain Shams University, Cairo, 11566, Egypt. E-mail: sayed.karam2008@sci.asu.edu.eg
bEntomology Department, Faculty of Science, Ain Shams University, Cairo, 11566, Egypt
cChemistry Department, College of Science, King Khalid University, P. O. Box 9004, Abha, 62223, Saudi Arabia
First published on 24th March 2026
Abstract
Azofuranone was transformed into a series of pyrazole-based hydrazones with 71–79% yields through hydrazinolysis using hydrazine hydrate at room temperature followed by condensation with selected aromatic aldehydes in refluxing ethanol. The antiproliferative activity of the obtained derivatives was evaluated against MCF-7 (breast) and HCT-116 (colon) cancer cell lines using the MTT assay. Among the tested compounds, the chlorobenzylidene derivative (3a) exhibited the most pronounced cytotoxic activity (IC50 = 6.33 ± 1.3 µM against HCT-116 and IC50 = 8.61 ± 1.6 µM against MCF-7), while the remaining hydrazones also showed moderate to strong effects in both cell models. Computational target predictions suggested possible involvement of kinase-related pathways. Molecular docking studies were performed using CDK2 (PDB ID: 2 A4L) to explore potential binding modes. Compound 3a displayed a favorable docking score compared with the reference ligands and formed hydrogen-bond interactions with key residues within the active site. Molecular dynamics simulations indicated stable ligand–protein interaction patterns over the simulation period. In addition, in silico ADME analysis revealed acceptable physicochemical and pharmacokinetic characteristics, with compound 3a demonstrating the most balanced profile. Overall, the combined biological and computational findings identify compound 3a as a promising scaffold for further optimization in the development of antiproliferative agents.
Introduction
The continuous need for effective anticancer therapies remains a major focus in medicinal chemistry, as cancer continues to be one of the leading causes of mortality worldwide. Cancer was responsible for around ten million deaths in 2020, according to the WHO (world health organization). In 2022, approximately 2.3 million cases of breast cancer and 670![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths ocurred worldwide. Breast cancer was reported for roughly 1 in 6 cancer deaths. Besides, an estimated 866
000 deaths ocurred worldwide. Breast cancer was reported for roughly 1 in 6 cancer deaths. Besides, an estimated 866![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 136 cases of liver cancer and 758
136 cases of liver cancer and 758![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 725 deaths worldwide appeared in 2022. Pyrazoles are the subject of chemical and biological investigations because of their promising pharmacological, agrochemical, and analytical applications. Pyrazole derivatives have demonstrated diverse biological activities, including antitumor, antimicrobial, antioxidant, and antiviral effects.1–9 They are present in many natural products and well known for being helpful in drug design.10 Some pyrazole-based anticancer drugs are presented in Fig. 1. Pyrazofurin is effective against a broad spectrum of cancer cells due to its inhibition of nucleotide synthesis, which is crucial for rapidly dividing tumor cells. Ruxolitinib is a potent, selective inhibitor of Janus kinase 1 and 2 (JAK1/JAK2) with high enzymatic potency (IC50 of 3.3 nM for JAK1 and 2.8 nM for JAK2). Against various cancer cell lines, Ruxolitinib acts as a targeted agent to inhibit proliferation and induce apoptosis, generally displaying values in the micromolar (M) range, typically between 1 µM and 50 µM, reliant on the cell line, duration of exposure (24–72 h), and culture conditions. Crizotinib acts as a potent tyrosine kinase (TK) inhibitor with values normally ranging from 10 nM to 900 nM against ALK-rearranged or MET-amplified cancer cells. Key sensitivities contain lung cancer lines (EBC-1, H1993 ≤ 10 nM; H2228 ≈ 311 nM) and ALCL/gastric lines (5–20 nM).
725 deaths worldwide appeared in 2022. Pyrazoles are the subject of chemical and biological investigations because of their promising pharmacological, agrochemical, and analytical applications. Pyrazole derivatives have demonstrated diverse biological activities, including antitumor, antimicrobial, antioxidant, and antiviral effects.1–9 They are present in many natural products and well known for being helpful in drug design.10 Some pyrazole-based anticancer drugs are presented in Fig. 1. Pyrazofurin is effective against a broad spectrum of cancer cells due to its inhibition of nucleotide synthesis, which is crucial for rapidly dividing tumor cells. Ruxolitinib is a potent, selective inhibitor of Janus kinase 1 and 2 (JAK1/JAK2) with high enzymatic potency (IC50 of 3.3 nM for JAK1 and 2.8 nM for JAK2). Against various cancer cell lines, Ruxolitinib acts as a targeted agent to inhibit proliferation and induce apoptosis, generally displaying values in the micromolar (M) range, typically between 1 µM and 50 µM, reliant on the cell line, duration of exposure (24–72 h), and culture conditions. Crizotinib acts as a potent tyrosine kinase (TK) inhibitor with values normally ranging from 10 nM to 900 nM against ALK-rearranged or MET-amplified cancer cells. Key sensitivities contain lung cancer lines (EBC-1, H1993 ≤ 10 nM; H2228 ≈ 311 nM) and ALCL/gastric lines (5–20 nM).
 | ||
| Fig. 1 Some pyrazole-based anticancer drugs in addition to kinase inhibitor (A) and roscovitine as CDK inhibitor. | ||
Therefore, constructing heterocycles bearing pyrazoles is a strategic synthetic line for drug development.11,12 They are essential moieties in various chemotherapeutic agents with reliable potential.13–16 Cyclin-dependent kinases (CDKs) play a central role in regulating the cell division cycle. Dysregulation of these kinases and their regulatory partners is frequently observed in human tumors. Roscovitine (inhibitor of CDKs) exhibits high efficacy and selectivity (cf. Fig. 1). The purine core of this inhibitor bonds to the adenine binding site of CDK2. Analysis of CDK2/roscovitine complex crystal structure would guide the enterprise of many other potent CDK inhibitors.17
Thus, searching for new antitumor drugs has become an interesting research topic in medicinal chemistry. Structural diversification through the introduction of additional aromatic and heterocyclic fragments is a common medicinal chemistry strategy18,19 aimed at modulating physicochemical properties, enhancing π–π stacking potential, and optimizing hydrophobic and hydrogen-bonding interactions within biological targets. In this context, the present design focused on scaffold expansion of the azofuranone-derived core to explore structure–activity relationships rather than targeting a predefined kinase pharmacophore. Some synthetic approaches reported the preparation of pyrazole-based hydrazone bearing various moieties with good yields (75–90%, depending on the substituent on the aldehyde), through condensation of the acid hydrazide with different aldehydes, which exhibited promising anticancer and antiviral activity.6,13,20
This condensation strategy is the most widely used approach due to its operational simplicity and high efficiency. Some advantages of this strategy included mild reaction conditions, simple work-up (often crystallizes directly from ethanol), good to excellent yields, and no need for expensive catalysts. Some limitations included that reaction rate can decrease with sterically hindered aldehydes, electron-donating substituents may lower yields, and possible E/Z isomer formation in some cases. Engelhardt et al. stated benzimidazole derivative (A) (cf. Fig. 1) as a next generation of EGFR tyrosine kinase inhibitor acquiring reasonable affinity to mutated EGFR (IC50 EGFRL858R T790M C797S = 2100 nM, IC50 EGFRdel19 T790M C797S = 250 nM).21
Bearing in mind these reports and continuing to our work for synthesizing bioactive pyrazoles,22–29 this work focused on the heterocyclic transformation of azofuranone 1, as a common precursor, into pyrazole candidates 3a–3c to investigate their ability to impede the proliferation of MCF-7 (breast) and HCT-116 (colon) cancer cell panels (compared with CDK2 inhibitor, roscovitine). In silico target prediction, molecular docking, and pharmacokinetic modeling were conducted as post-biological mechanistic exploration tools to investigate potential molecular targets associated with the observed antiproliferative activity and to assess drug-likeness properties. These computational analyses were intended to provide mechanistic hypotheses and structural insights rather than to serve as primary design drivers. To further narrative for protein flexibility and overcome the inherent limitations of static docking approaches, molecular dynamics simulations were employed to assess the time-dependent stability and interaction behavior of the ligand–protein complexes.
Results and discussion
Synthesis
The key material, 3-(2-(4-chlorophenyl)hydrazineylidene)-5-phenylfuran-2(3H)-one (1) was easily transformed into pyrazole candidates 3a–3c (cf. Scheme 1). First, hydrazinolysis of azofuranone 1 with hydrazine hydrate (80%) in ethyl alcohol at ambient temperature (RT) afforded the acid hydrazide 2.30 Second, condensation of acid hydrazide with aromatic aldehydes, including 4-chlorobenzaldehyde, anthracene-9-carbaldehyde, and 1,3-diphenylpyrazole-4-carbaldehyde in refluxing ethanol achieved the construction of pyrazoles 3a–3c, respectively. IR of pyrazoles 3a–3c presented absorption bands for an amidic carbonyl and NH groups. Its 1H NMR spectra lacked a signal for NH2 protons and presented a singlet signal for NH proton. The reaction way can be visualized via initial condensation of primary amino set of the hydrazide 2 with carbonyl group of aldehydes to produce a non-isolable intermediate [I] pursued by intramolecular 5-exo-trig ring closure to remove water molecule (Scheme 1).While our current work focuses on small scale laboratory synthesis, several features of the route described indicate that it could be adapted to a Process Chemistry framework with relative ease. The sequence consists of only two fundamental operations, hydrazinolysis and subsequent hydrazone formation, both of which proceed under mild temperatures, use readily available reagents, and rely on operationally simple transformations. These qualities are advantageous for scale up because they minimize the need for tightly controlled conditions or specialized equipment. Additionally, the products often crystallize directly from the reaction mixture, reducing dependence on chromatographic purification and making the route inherently more suitable for process development.
From a Process Chemistry perspective, opportunities for optimization include solvent selection to improve atom economy and waste minimization, evaluation of alternative hydrazine sources with improved safety profiles, and refining crystallization conditions to achieve consistent solid-state properties on larger scales. The clean impurity profile observed at the laboratory scale further suggests that reaction robustness is high, which typically translates well to larger volume reactors. Overall, while dedicated process scale investigations were not within the scope of this study, the simplicity, robustness, and predictable behavior of both key transformations provide a strong foundation for future translation of the method into a Process Chemistry route, especially for the preparation of promising compound 3a.
While the developed two-step sequence proved robust for the aldehydes studied, several opportunities for further improvement were identified, particularly with respect to functional group tolerance and the efficiency of the hydrazinolysis step. The major scope for improvement in our current synthetic approach lies in enhancing structural diversity and optimizing step efficiency, particularly as we move toward generating more advanced analogues. The two step sequence, hydrazinolysis followed by hydrazone formation, proved reliable for the set of simple aromatic aldehydes explored; however, increasing the complexity of substitution or introducing more sensitive functional groups may require further refinement of reaction conditions to ensure consistent yields. Additionally, the hydrazinolysis step, while operationally straightforward, could benefit from optimization to improve atom economy and reduce the excess of hydrazine traditionally used in such transformations.
To address these limitations, we envisage several strategies. First, expanding the substrate scope under diversified solvent systems, catalysts, or mild activating conditions may improve tolerance for more functionalized aldehydes and enable broader exploration of structure–activity relationships (SAR). Second, implementing continuous flow techniques for hydrazinolysis may enhance safety, scalability, and reaction control, thereby reducing excess and narrow impurity profiles. Third, further studies on controlled crystallization could streamline purification workflows, especially for analogues prone to hydrazone formation. Collectively, these enhancements will help transform the current route into a more versatile and scalable platform suitable for generating expanded libraries of pyrazole-based hydrazones and for supporting future medicinal chemistry optimization campaigns.
Antiproliferative activity
The in vitro cytotoxic activity of the synthesized compounds was evaluated using the MTT assay31,32 against colon (HCT-116) and breast (MCF-7) cancer cell lines, with doxorubicin and roscovitine used as reference controls Table 1. Compared with the reference compounds, hydrazone 3a showed the most pronounced activity against both cell lines. The presence of the chlorine substituent and aromatic framework may contribute to its enhanced biological response. The compounds 3b and 3c showed strong effectiveness against two cells. The observed cytotoxic activity may be influenced by the presence of NH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O functional groups, which are capable of participating in hydrogen bond interactions. The contribution of hydrophobic interactions arising from the aryl groups (chlorophenyl, anthracene, and pyrazole rings) may also be consistent with the observed activity trends (cf. Fig. 2).33
O functional groups, which are capable of participating in hydrogen bond interactions. The contribution of hydrophobic interactions arising from the aryl groups (chlorophenyl, anthracene, and pyrazole rings) may also be consistent with the observed activity trends (cf. Fig. 2).33
In silico target prediction
Swiss target prediction analysis34 was used to estimate the most probable biological targets for compounds 3a–3c (cf. Fig. 3). For compound 3a, enzyme-related proteins represented 26.7% of the predicted targets, followed by kinases and G-protein-coupled receptors (13.3%). While substantial enrichments in protein-coupled receptor and kinase-related proteins were perceived for compound 3b with 40.0 and 26.7% of anticipated targets. Also, compound 3c showed considerable prospects in protein-coupled receptors with 46.7%, in addition to enzyme and protease-related proteins with 20.0% of the anticipated targets. These results suggest a possible association with kinase-related pathways. Judging the anticipated targets revealed mutual targets between these compounds as kinase and protein-coupled receptors. Given the role of CDK2 in cell-cycle regulation, it was selected as a representative kinase target for subsequent docking analysis. It is important to note that the computational target prediction indicated multiple potential protein classes, suggesting that the investigated compounds may exert their antiproliferative activity through multi-target mechanisms. The diversity of predicted target classes suggests that the antiproliferative activity may involve multiple biological pathways, providing a broader perspective for subsequent target-oriented studies.Molecular docking simulation
To explore the affinity of the compounds toward the CDK2 binding pocket (PDB ID: 2 A4L),35 molecular docking simulations were performed to estimate their binding energies. Binding affinity was assessed based on calculated S-scores (kcal mol−1) and predicted hydrogen-bonding interactions. More negative docking scores were interpreted as indicating stronger predicted binding affinity within the active site. The binding pocket of native co-crystallized ligand (roscovitine, RRC) was docked for all complexes in the same subset (see Tables 2 and 3).| Compd | S-score (kcal mol−1) | RMSD (Å) | Amino acids included in bonding | |
|---|---|---|---|---|
| H-bonding (bond length, Å) | Hydrophobic interaction (bond length, Å) | |||
| a Shared amino acids interacting with both compounds and RRC were italized. | ||||
| 3a | −9.0453 | 1.1171 | THR 14 (3.06), LYS 89 (3.15) | — |
| 3b | −7.0523 | 1.2722 | — | PHE 80 (4.33) |
| 3c | −8.9010 | 1.5364 | ASP 86 (3.20) | VAL 18 (4.45), GLN 85 (4.30) |
| Doxorubicin | −8.6836 | 1.3266 | GLU 12 (3.19) | VAL 18 (4.20), GLN 131 (4.26) |
| RRC | −8.9902 | 1.4381 | GLN 131 (2.77), LEU 83 (3.31) | ILE 10 (4.21), VAL 18 (4.48), LYS 89 (3.87) |
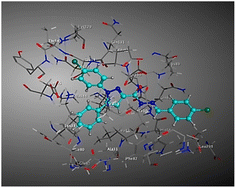
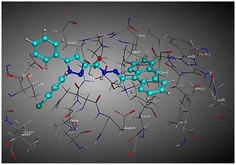
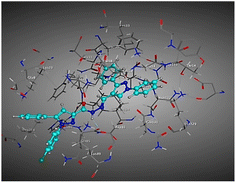
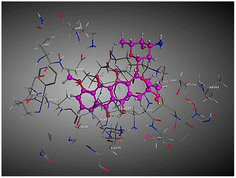
| Compd | 2D | 3D |
|---|---|---|
| a The docked compounds were stained in cyan while doxorubicin and RRC were stained purple. | ||
| 3a | 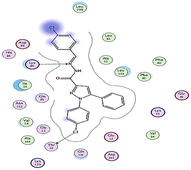 |
 |
| 3b | 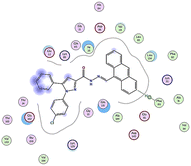 |
 |
| 3c | 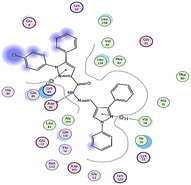 |
 |
| Doxorubicin | 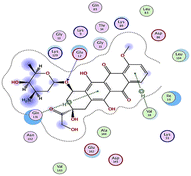 |
 |
| RRC | 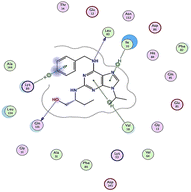 |
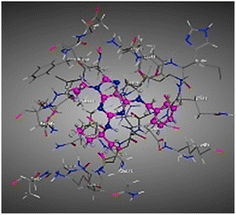 |
The amino acids implied in binding relations e.g., H-bonds and hydrophobic interactions, between the ligands and target CDK2 enzyme were inclined in Table 2. Also, Table 3 appeared graphical 2D and 3D representations of docking simulation. The defined information on the molecular interactions between ligands and CDK2 protein was displayed in a 2D illustration. Besides, the binding interactions between ligands and active pockets of protein are indicated in red by the 3D depiction, that imply hydrogen bond interactions. As detected in Table 2, the ligand binding energies were closer to those of doxorubicin as a reference drug and co-crystallized ligand (roscovitine, RRC) as a CDK2 inhibitor. Some amino acids that interact with compounds also interacted with RRC ligand.
Judging to doxorubicin (S: −8.6836 kcal mol−1) and RRC (S: −8.9902 kcal mol−1), the pyrazole-bearing chlorobenzylidene candidate 3a had the greatest S-score with a binding energy of −9.0453 kcal mol−1 with the lowest RMSD value (1.1171 Å). This docking pose indicated favorable accommodation within the CDK2 active site, with hydrogen-bond interactions involving THR 14 and LYS 89, residues also observed in interactions with the co-crystallized ligand (RRC). Compound 3c provided S-score of −8.9010 kcal mol−1 via one hydrogen bond by NH group with ASP 86 through side chain donor, in addition to two hydrophobic (pi-H) interactions by the two pyrazole rings with VAL 18 (shared with RRC) and GLN 85. Compound 3b developed one hydrophobic (H-pi) interaction by anthracene ring with PHE 80, resulting in S-score of −7.0523 kcal mol−1. These findings highlight the potential of this series as promising antiproliferative scaffolds. The docking outcomes provide structural insight into possible binding modes within the CDK2 active site and may serve as a basis for subsequent experimental evaluation.
The verification of the docking protocol was established using co-crystallized ligands (RRC) with their respective CDK2 protein target, in which the superimposition of native and redocked co-crystallized ligands was envisaged among 2D diagram.36 The RMSD value for superimposition, which indicates the deviation between the two configurations, was calculated to be 1.4381 Å (cf. Fig. 4).
 | ||
| Fig. 4 Superimposition (2D diagram) of native and redocked RRC configurations at CDK2 protein target active pockets (PDB ID: 2 A4L) per RMSD of 1.4381 Å. | ||
As an imperious constraint in drug discovery, the ligand efficiency (LE) parameter of pyrazole-based hydrazones 3a–3c was established and judged to that of RRC and doxorubicin (cf. Table 4). The outcomes clarified that pyrazole-bearing chlorobenzylidene 3a revealed LE value of −0.3015 kcal mol−1 per heavy atom, which was comparable to that of RRC (LE = −0.3457 kcal mol−1 per heavy atom) and higher than that of doxorubicin (LE = −0.2226 kcal mol−1 per heavy atom). While compounds 3b and 3c had LE values comparable to doxorubicin. This indicates that, on a per-heavy-atom basis, the tested compounds display a balanced efficiency profile, supporting their suitability for further optimization in drug design.
| Compd | Ligand efficiency (LE, kcal mol−1 per heavy atom) |
|---|---|
| 3a | −0.3015 |
| 3b | −0.1906 |
| 3c | −0.2225 |
| RRC | −0.3457 |
| Doxorubicin | −0.2226 |
Molecular dynamics (MD) simulation
Through MD simulation, the stability of the ligand–protein complexes over time was evaluated using RMSD analysis.37–39 As illustrated in Fig. 5a–e, all simulated systems reached equilibration early in the trajectory and remained stable over the 100 ns simulation period, with no evidence of major conformational rearrangements. For the tested compounds, RMSD values were mostly held around 2.0 Å for most of the simulation, implying stable positioning within the binding pocket. Doxorubicin showed relatively higher deviations during the initial stage of the simulation, particularly within the first ∼25 ns, before converging to a stable RMSD profile. In contrast, the co-crystallized ligand (RRC) exhibited an overall stable trajectory, with only a short-lived increase in RMSD between approximately 30 and 40 ns, after which stability was preserved. Overall, these observations suggest that both the investigated compounds and the reference ligands maintain stable interaction profiles, in agreement with the docking analysis.Protein flexibility was further evaluated through RMSF analysis (Fig. 5f–j). All systems displayed comparable RMSF profiles, reflecting a similar global fluctuation pattern regardless of the bound ligand. For most residues, RMSF values remained below ∼2.0 Å, proposing that ligand binding did not convince significant structural instability. Higher fluctuations were confined to specific regions, mainly terminal segments and flexible loops, which are inherently mobile and not directly involved in ligand recognition. Residues located within or near the binding pocket showed moderate and largely comparable fluctuations across all systems, indicating that the tested compounds, doxorubicin, and RRC preserve the native flexibility of the protein while maintaining stable interactions.
Analysis of protein-ligand interaction histograms (Fig. 6) revealed that all ligands maintained a consistent network of interactions throughout the simulation. Hydrogen bonds, hydrophobic contacts, and water-mediated interactions were observed for a substantial portion of the trajectory.40,41 In the case of the tested compounds, hydrophobic contacts were particularly prominent, likely facilitated by the presence of multiple aromatic moieties that enhance ligand anchoring within the binding site. Several residues were repeatedly involved in ligand interactions across both the investigated compounds and the reference systems. Among them, LEU83 was identified as a key interacting residue, forming stable hydrogen bonding interactions with the tested compounds. This residue was also engaged by the co-crystallized ligand in the docking analysis, providing further consistency between the docking and MD results.
Overall, the MD simulations indicate that the investigated compounds form stable complexes with the target protein, characterized by balanced RMSD behavior, comparable residue-level flexibility, and persistent interaction patterns. The consistency between molecular dynamics and docking analyses reinforces the proposed binding orientations and suggests that the investigated compounds can maintain interaction patterns comparable to those observed for the reference ligands.
ADME study
ADME profiles indicating physicochemical, lipophilicity, pharmacokinetics, and drug-likeness attributes of substances were depicted by SwissADME free web tool (cf. Fig. S1–S6 and Table S1).42–44 On radar chart, the substrate's pink area was managed to assess its bioavailability. The compounds 3a–3c were implicated in the pink area (suitable physicochemical space for oral bioavailability) and obeyed Veber rule with a number of rotatable bonds less than ten. In contrast to compounds 3b and 3c, compound 3a had a favored molar refractivity (<140). All compounds exhibited Csp3 ratio lower than 0.25. Compound 3a complied with Lipinski's rule of five (with one abuse of M![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Log
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P > 4.15). The bioavailability score was 0.55 for compound 3a and 0.17 for compounds 3b and 3c. Compounds 3b and 3c did not fulfill Lipinski's rule of five as they had two abuses of MW > 500 and MLogP > 4.15. Similar to potential drugs, these compounds exhibited number of hydrogen bond donors (HBD) and acceptors (HBA) within the range (less than 5 and 10, respectively).
P > 4.15). The bioavailability score was 0.55 for compound 3a and 0.17 for compounds 3b and 3c. Compounds 3b and 3c did not fulfill Lipinski's rule of five as they had two abuses of MW > 500 and MLogP > 4.15. Similar to potential drugs, these compounds exhibited number of hydrogen bond donors (HBD) and acceptors (HBA) within the range (less than 5 and 10, respectively).
One foremost factor linked to drug bioavailability is TPSA (topological polar surface area). Fragments that are passively absorbed and show TPSA less than 140 Å2 are reasoned to allow high bioavailability. Thus, compounds 3a–c are probable to expose good passive oral absorption, with TPSA values of 59.28, 59.28, and 77.10 Å2, respectively. Also, their consensus Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Po/w specified that they exhibit good lipophilicity. Whilst compound 3a seemed to have high GI absorption, compounds 3b and 3c established low GI absorption. Their skin permeation (Log
Po/w specified that they exhibit good lipophilicity. Whilst compound 3a seemed to have high GI absorption, compounds 3b and 3c established low GI absorption. Their skin permeation (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp) factors follow this sequence: 3a > 3c > 3b, which expands the skin's ability to pass bioactive chemicals.
Kp) factors follow this sequence: 3a > 3c > 3b, which expands the skin's ability to pass bioactive chemicals.
The cytochrome P450 isoenzymes, which are essential for drug metabolism, were also assessed. None of the studied compounds were predicted to inhibit CYP2D6, indicating a lower likelihood of metabolism-related drug–drug interactions. Evaluation of absorption characteristics using the BOILED-Egg model (Fig. 7) revealed that compound 3a, unlike compounds 3b and 3c, was positioned within the white region, indicating favorable gastrointestinal tract absorption, as illustrated in Fig. 5. In addition, compound 3a was predicted not to be actively effluxed from the central nervous system by P-glycoprotein (PGP), as indicated by its red classification. Screening with the PAINS filter, which detects molecules prone to false-positive biological assay results, showed no alerts for any of the tested compounds. Overall, the computational ADME evaluation supports the suitability of these compounds for early-stage lead exploration. Such predictive analyses provide an initial view of pharmacokinetic behavior and help prioritize candidates for further study. More comprehensive safety and metabolism profiling, including experimental evaluation of hepatic liability and cardiac ion channel interactions, will represent important next steps during advanced preclinical development.
The findings of antiproliferative activity of investigated substrates were judged by their docking scores and pharmacokinetic properties pertaining to doxorubicin and roscovitine (cf. Table 5), which discovered a good relationship.
| Compds | IC50 (µM), (HCT-116/MCF-7) | S-score (kcal mol−1) | TPSA | HBA | HBD | C. Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
GI absorption | Log Kp | Bioavailability score |
|---|---|---|---|---|---|---|---|---|---|
a HBA: hydrogen bond acceptor HBD: hydrogen bond donor, C. Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P: consensus Log P: consensus Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (lipoilicity). P (lipoilicity). |
|||||||||
| 3a | 6.33/8.61 | −9.0453 | 59.28 | 3 | 1 | 5.07 | High | −4.66 | 0.55 |
| 3b | 15.25/17.16 | −7.0523 | 59.28 | 3 | 1 | 6.26 | Low | −3.92 | 0.17 |
| 3c | 10.02/11.52 | −8.9010 | 77.1 | 4 | 1 | 5.83 | Low | −4.6 | 0.17 |
| Dox. | 5.23/4.17 | −8.6836 | 206.07 | 12 | 6 | 2.51 | Low | −8.71 | 0.17 |
| RRC | 12.24/9.32 | −8.9902 | 87.89 | 4 | 3 | 0.52 | High | −6.22 | 0.55 |
Based on biological evaluation, docking analyses, and physicochemical assessment of the synthesized hydrazone series (as discussed above), several preliminary design principles could be proposed to guide further development of this scaffold as an antiproliferative chemotype. First, the results suggest that aryl substitution patterns exert a strong influence on activity, with electron withdrawing substituents, particularly the chloro group in compound 3a, enhancing cytotoxic potency. This indicates that incorporation of moderately lipophilic, σ-accepting groups on the aromatic aldehyde component may strengthen interactions within the CDK2 active site and improve cell-based efficacy. Second, the docking and MD simulations highlight the importance of maintaining the hydrazone linker orientation that enables hydrogen bonding to key CDK2 residues; therefore, structural modifications should preserve the hydrazone moiety and its conjugated alignment with the pyrazole core.
Third, the ADME evaluation shows that balanced lipophilicity and limited polar surface area correlate with favorable pharmacokinetic profiles, suggesting that future analogues should avoid excessive polarity or bulky substituents that may compromise permeability. Lastly, the general scaffold tolerates aromatic modifications well, implying that diversifying the aryl ring with halogens, heterocycles, or additional small electron modulating groups could be an effective strategy to fine tune both activity and physicochemical properties. Together, these design rules provide a conceptual foundation for next generation analogues and support broader structure–activity exploration of pyrazole-based hydrazones as potential anticancer agents.
A concise comparative analysis highlighted the key distinctions between our study and previously reported works on pyrazole-based hydrazones. Unlike earlier reports that primarily focused on conventional condensation-based synthesis and preliminary cytotoxic screening, our work introduces azofuranone as a novel starting precursor, enabling the generation of structurally distinct pyrazole-based hydrazones through a sequential hydrazinolysis-condensation strategy. In addition to evaluating antiproliferative activity against both MCF-7 and HCT-116 cell lines, our study integrates comprehensive computational investigations, including target prediction, molecular docking against CDK2 (PDB ID: 2 A4L), molecular dynamics simulations, and in silico ADME profiling.
Furthermore, compound 3a demonstrated enhanced cytotoxic potency alongside favorable binding interactions and pharmacokinetic characteristics, providing a more mechanistically supported structure–activity rationale compared with previously reported analogues. This combined experimental and computational approach offers a broader and more systematic evaluation than many earlier studies.
Conclusion
In this work, pyrazole-based hydrazone candidates were obtained from azofuranone through hydrazinolysis to give acid hydrazide followed by the condensation with 4-chlorobenzaldehyde, anthracene-9-carbaldehyde, and 1,3-diphenylpyrazole-4-carbaldehyde. The in vitro antiproliferative action versus MCF-7 and HCT-116 cancer cell panels showed the most potency of chlorobenzylidene candidate. The other hydrazones also exhibited strong activity. Based on in silico target prediction suggesting kinase-related pathways, molecular docking analysis was performed toward CDK2 protein, where the chlorobenzylidene derivative showed a docking score comparable to the reference compounds. Also, bis-pyrazole derivative showed a closer docking score through one hydrogen bonding by NH group with ASP 86 through side chain donor, in addition to two hydrophobic interactions by the two pyrazole rings with VAL 18 (shared with RRC) and GLN 85. In turn, the anthracene candidate formed one hydrophobic interaction by anthracene ring with PHE 80. The observed hydrogen-bonding and hydrophobic interactions suggest favorable accommodation of these derivatives within the CDK2 binding pocket. Compound 3a showed a competitive ligand efficiency compared with the reference compounds, supporting its potential as a lead scaffold. Molecular dynamics analysis further indicated stable interaction patterns over the simulation period. Together, the biological and computational findings point toward a possible involvement of kinase-related pathways, offering a structural framework for future mechanistic studies. The in silico ADME profiling of compounds 3a–3c revealed promising physicochemical, pharmacokinetic, and drug-likeness characteristics. Compound 3a showed a balanced drug-like profile, residing within the optimal bioavailability radar region and complying with Veber's and Lipinski's criteria, with a higher bioavailability score compared to compounds 3b and 3c. It also exhibited favorable molar refractivity and strong gastrointestinal (GI) absorption, supported by its position in the BOILED-Egg model's white region. All compounds displayed acceptable TPSA values (<140 Å2), indicating potential for good passive oral absorption, with compound 3a again showing the most desirable profile. Skin permeability followed the trend 3a > 3c > 3b, with 3a being the most permeable. Notably, all compounds passed the PAINS filter, indicating no structural alerts commonly associated with assay interference. Overall, compound 3a presented the most favorable overall ADME balance among the derivatives investigated. The present findings offer a structural and biological framework for continued investigation of pyrazole-based derivatives as antiproliferative agents.Materials and methods
Synthesis and procedures
See SI.Antiproliferative activity
Conflicts of interest
The authors declare that they have no competing interests.Data availability
All data generated in this study can be found in the supplementary information (SI) files. Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra01643j.Acknowledgements
The authors extend their appreciation to the Deanship of Research and Graduate Studies at King Khalid University for funding this work through the Large Groups Project under grant number (RGP.2/137/46).References
- A. O. Abdelhamid, I. E. El Sayed, Y. H. Zaki, A. M. Hussein, M. M. Mangoud and M. A. Hosny, Utility of 5-(furan-2-yl)-3-(p-tolyl)-4,5-dihydro-1H-pyrazole-1-carbothioamide in the synthesis of heterocyclic compounds with antimicrobial activity, BMC Chem, 2019, 13, 48 CrossRef PubMed.
- A. A. Fadda and K. S. Mohamed, Synthesis, Characterization and Cytotoxicity Evaluation of Some Novel Pyrazole and Pyrrole Derivatives Containing Benzothiazole Moiety, Heterocycles, 2015, 91(10), 1937–1954 CrossRef.
- S. Bondock, W. Khalifa and A. A. Fadda, Synthesis and antimicrobial activity of some new 4-hetarylpyrazole and furo[2,3-c]pyrazole derivatives, Eur. J. Med. Chem., 2011, 46(6), 2555–2561 Search PubMed.
- E. A. El-Bordany, A. Abdel Aziz, W. S. I. Abou-Elmagd and A. I. Hashem, Synthesis and Spectroscopic Characterization of some Novel Pyrazoloquinoline, Pyrazolyltetrazine, and Thiazolidinone Derivatives, J. Heterocycl. Chem., 2018, 55, 291–296 CrossRef CAS.
- S. K. Ramadan, A. K. El-Ziaty and R. S. Ali, Synthesis, antiproliferative activity, and molecular docking of some N-heterocycles bearing a pyrazole scaffold against liver and breast tumors, J. Heterocycl. Chem., 2021, 58, 290 CrossRef CAS.
- Y. M. Youssef, M. E. Azab, G. A. Elsayed, A. A. El-Sayed, A. Hassaballah, M. M. El-Safty, R. A. Soliman and E. A. E. El-Helw, Synthesis and antioxidant, antimicrobial, and antiviral activity of some pyrazole-based heterocycles using a 2(3H)-furanone derivative, J. Iran. Chem. Soc., 2023, 20(9), 2203–2216 Search PubMed.
- S. R. Atta-Allah, W. S. I. Abou-Elmagd, K. A. Kandeel and M. M. Hemdan, Synthesis and antimicrobial activity evaluation of some novel hydrazone, pyrazolone, chromenone, 2-pyridone and 2-pyrone derivatives, J. Chem. Res., 2017, 41(11), 617–623 CrossRef CAS.
- S. K. Ramadan and W. S. I. Abou-Elmagd, Synthesis and Anti H5N1 Activities of Some Novel Fused Heterocycles Bearing Pyrazolyl Moiety, Synth. Commun., 2018, 48(18), 2409–2419 CrossRef CAS.
- M. R. Elmorsy, E. Abdel-Latif, H. E. Gaffer, S. E. Mahmoud and A. A. Fadda, Sci. Rep., 2023, 13, 2782 CrossRef CAS PubMed.
- N. Verma and S. Arora, Navigating the Global Regulatory Landscape for Exosome-Based Therapeutics: Challenges, Strategies, and Future Directions, Pharmaceutics, 2025, 17(8), 990 CrossRef PubMed.
- N. Verma, S. Arora, A. K. Singh and A. Kumar, Extracellular Vesicle-Associated miRNAs in Cornea Health and Disease: Diagnostic Potential and Therapeutic Implications, Targets, 2025, 3(4), 32 CrossRef.
- N. Verma, S. Arora, A. K. Singh and J. Ahmed, Unlocking the potential of exosomes ‘extracellular vesicles’: drug delivery advancements and therapeutics in ocular diseases, RSC Pharm., 2025, 2, 1201–1226 RSC.
- A. R. Morsy, S. H. Mahmoud and N. M. Abou Shama, et al., Antiviral activity of pyrazole derivatives bearing a hydroxyquinoline scaffold against SARS-CoV-2, HCoV-229E, MERS-CoV, and IBV propagation, RSC Adv., 2024, 14(38), 27935–27947 Search PubMed.
- A. El-Shafei, A. A. Fadda, A. M. Khalil, T. A. E. Ameen and F. A. Badria, Bioorg. Med. Chem., 2009, 17(14), 5096 CrossRef CAS PubMed.
- S. K. Ramadan, A. T. Ali, S. M. Gomha and E. A. E. El-Helw, Design, synthesis, and in silico studies of pyrazolyl-thiazole and thiazolidinone hybrids as potential antiproliferative agents, Synth. Commun., 2025, 55(12), 931–951 CrossRef CAS.
- S. K. Ramadan, H. S. Abd-Rabboh and W. S. I. Abou-Elmagd, Antiproliferative Activity and Molecular Docking of Some Pyrazole-Based Quinazolinone, Benzimidazole, and Tetrazinethione Derivatives, J. Biochem. Mol. Toxicol., 2025, 39, e70131 CrossRef CAS PubMed.
- L. Meijer, A. Borgne and O. Mulner, et al., Biochemical and Cellular Effects of Roscovitine, a Potent and Selective Inhibitor of the Cyclin-Dependent Kinases cdc2, cdk2 and cdk5, Eur. J. Biochem., 1997, 243, 527–536 CrossRef CAS PubMed.
- S. Arora, J. Liang, S. K. Fullerton-Shirey and J. E. Laaser, Triggerable Ion Release in Polymerized Ionic Liquids Containing Thermally Labile Diels–Alder Linkages, ACS Mater. Lett., 2020, 2(4), 331–335 CrossRef CAS.
- Z. Huo, S. Arora, V. A. Kong, B. J. Myrga, A. Statt and J. E. Laase, Effect of Polymer Composition and Morphology on Mechanochemical Activation in Nanostructured Triblock Copolymers, Macromolecules, 2023, 56(5), 1845–1854 CrossRef CAS PubMed.
- S. K. Ramadan, H. S. Abd-Rabboh, N. M. Gad, W. S. I. Abou-Elmagd and D. S. Haneen, Synthesis and characterization of some chitosan–quinoline nanocomposites as potential insecticidal agents, Polycyclic Aromat. Compd., 2023, 43(8), 7013–7026 CrossRef CAS.
- H. Engelhardt, D. Böse and M. Petronczki, et al., Start Selective and Rigidify: The discovery path toward a next generation of EGFR tyrosine kinase inhibitors, J. Med. Chem., 2019, 62, 10272–10293 CrossRef CAS PubMed.
- E. A. E. El-Helw, M. Asran, M. E. Azab, M. H. Helal and S. K. Ramadan, Synthesis, Cytotoxic, and Antioxidant Activity of Some Benzoquinoline-Based Heterocycles, Polycyclic Aromat. Compd., 2024, 44(9), 5938–5950 CAS.
- M. M. Kaddah, A. A. Fahmi, M. M. Kamel, S. A. Rizk and S. K. Ramadan, Rodenticidal Activity of Some Quinoline-Based Heterocycles Derived from Hydrazide–Hydrazone Derivative. Polycyclic Aromat, Compds, 2023, 43(5), 4231–4241 CAS.
- E. A. E. El-Helw, A. Alzahrani and S. K. Ramadan, Synthesis and antimicrobial activity of thiophene-based heterocycles derived from thiophene-2-carbohydrazide, Future Med. Chem., 2024, 16(5), 439–451 CrossRef CAS PubMed.
- A. Hassaballah, S. K. Ramadan, S. A. Rizk, E. A. E. El-Helw and S. S. Abdelwahab, Ultrasonic promoted regioselective reactions of the novel spiro 3,1-benzoxazon-isobenzofuranone dye toward some organic base reagents, Polycyclic Aromat. Compd., 2023, 43(4), 2973–2989 CrossRef CAS.
- E. A. E. El-Helw, E. M. Hosni, M. Kamal, A. I. Hashem and S. K. Ramadan, Synthesis, insecticidal Activity, and molecular docking analysis of some benzo[h]quinoline derivatives against Culex pipiens L. Larvae, Bioorg. Chem., 2024, 150, 107591 CrossRef CAS PubMed.
- A. K. El-Ziaty, W. S. I. Abou-Elmagd, S. K. Ramadan and A. I. Hashem, Synthesis and biological screening of some chromonyl-substituted heterocycles derived from 2(3H)-furanone derivative, Synth. Commun., 2017, 47(5), 471–480 CrossRef CAS.
- E. A. E. El-Helw, W. S. I. Abou-Elmagd, E. M. Hosni, M. Kamal, A. I. Hashem and S. K. Ramadan, Synthesis of Benzo[h]quinoline Derivatives and Evaluation of Their Insecticidal Activity Against Culex pipiens L. Larvae, Eur. J. Med. Chem., 2025, 290, 117565 CrossRef CAS PubMed.
- M. R. Mahmoud, M. M. El-Shahawi, W. S. I. Abou-Elmagd and M. H. Hekal, Novel Synthesis of Isoquinoline Derivatives Derived from (Z)-4-(1,3-Diphenylpyrazol-4-yl)isochromene-1,3-dione, Synth. Commun., 2015, 45(14), 1632–1641 Search PubMed.
- S. M. El-Kousy, A. M. El-Torgoman, G. M. Salama and A. I. Hashem, A study on rearrangement of arylazofuranones and pyrrolinones, Int. J. Chem., 1994, 5(2), 65–75 Search PubMed.
- T. Mosmann, Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxic assay, J. Immunol. Methods, 1983, 65, 55 Search PubMed.
- S. Arora and C. Mao, Light-regulated RNA interference induced by p-hydroxyphenacyl-modified siRNA in mammalian cells, Nucleosides, Nucleotides Nucleic Acids, 2024, 43(4), 316–339 CrossRef CAS PubMed.
- A. M. El-Naggar, M. A. El-Hashash and E. B. Elkaeed, Eco-friendly sequential one-pot synthesis, molecular docking, and anticancer evaluation of arylidene-hydrazinyl-thiazole derivatives as CDK2 inhibitors, Bioorg. Chem., 2021, 108, 104615 CrossRef CAS PubMed.
- E. A. E. El-Helw, M. Asran, M. E. Azab, M. H. Helal, A. Y. Alzahrani and S. K. Ramadan, Synthesis and in silico studies of certain benzo[f]quinoline-based heterocycles as antitumor agents, Sci. Rep., 2024, 14, 15522 CrossRef CAS PubMed.
- D. Gfeller, A. Grosdidier, M. Wirth, A. Daina, O. Michielin and V. Zoete, SwissTargetPrediction: a web server for target prediction of bioactive small molecules, Nucleic Acids Res., 2014, 42, W32–W38, DOI:10.1093/nar/gku293.
- https://www.rcsb.org/structure/2A4L.
- S. K. Ramadan, W. S. I. Abou-Elmagd, E. M. Hosni, M. Kamal, A. I. Hashem and E. A. E. El-Helw, Synthesis, in vivo evaluation, and in silico molecular docking of benzo[h]quinoline derivatives as potential Culex pipiens L. larvicides, Bioorg. Chem., 2025, 154, 108090 CrossRef CAS PubMed.
- D. Haneen, A. A. Abdalha, M. M. Alkhatib, M. Kamal, A. S. Youssef, W. S. I. Abou-Elmagd and S. S. Samir, Synthesis, comprehensive in silico studies, and cytotoxicity evaluation of novel quinazolinone derivatives as potential anticancer agents, Sci. Rep., 2025, 15, 1–25 Search PubMed.
- M. Kamal, M. Hekal, F. El-Azm, E. M. Hosni, Y. M. Ali, A. Alzahrani and E. H. Rafat, Synthesis and in silico studies of new thiophene-isoquinolinone hybrids as potential larvicides against Culex pipiens, Sci. Rep., 2025, 15, 1–24 Search PubMed.
- S. Arora, J. Rozon and J. E. Laaser, Dynamics of Ion Locking in Doubly-Polymerized Ionic Liquids, Macromolecules, 2021, 54(13), 6466–6476 CrossRef CAS.
- S. Arora and N. Verma, A review: advancing organic electronics through the lens of ionic liquids and polymerized ionic liquids, RSC Appl. Polym., 2024, 2, 317–355 RSC.
- A. H. Abdelrahman, M. E. Azab, M. A. Hegazy, A. Labena, A. Y. Alzahrani and S. K. Ramadan, Synthesis, Computational Analysis, and Exploring Antiproliferative Activity of Triazolo-and Thiazolo-Pyrimidine Derivatives as Potential EGFR Inhibitors, J. Mol. Struct., 2025, 1333, 141789 CrossRef CAS.
- A. Daina, O. Michielin and V. Zoete, SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules, Sci. Rep., 2017, 7, 42717 CrossRef PubMed.
- S. K. Ramadan, S. M. Gomha and E. A. E. El-Helw, Straightforward synthesis and in silico evaluation of pyrazolylthiazolidinone derivatives as prospective antiproliferative agents, Bioorg. Chem., 2025, 165, 109036 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |






