Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Design, synthesis, and antiproliferative activity of novel thiazole-based derivatives as tubulin polymerization inhibitors targeting the colchicine binding site
Lamya H. Al-Wahaibia,
Ali M. Elshamsyb,
Taha F. S. Alic,
Bahaa G. M. Youssif *d,
Stefan Bräse
*d,
Stefan Bräse *e,
Mohamed Abdel-Aziz*cf and
Nawal A. El-Koussi
*e,
Mohamed Abdel-Aziz*cf and
Nawal A. El-Koussi g
g
aDepartment of Chemistry, College of Sciences, Princess Nourah Bint Abdulrahman University, Riyadh 11671, Saudi Arabia
bPharmceutical Chemistry Department, Faculty of Pharmacy, Deraya University, Minia, Egypt
cMedicinal Chemistry Department, Faculty of Pharmacy, Minia University, Minia 61519, Egypt. E-mail: abulnil@hotmail.com; Tel: +2101003311327
dDepartment of Pharmaceutical Organic Chemistry, Faculty of Pharmacy, Assiut University, Assiut-71526, Egypt. E-mail: bgyoussif2@gmail.com; Tel: +201098294419
eInstitute of Biological and Chemical Systems, IBCS-FMS, Karlsruhe Institute of Technology, 76131 Karlsruhe, Germany. E-mail: braese@kit.edu
fMedicinal Chemistry Department, Faculty of Pharmacy, International Minia University, Minia, Egypt
gDepartment of Pharmaceutical Medicinal Chemistry, Faculty of Pharmacy, Assiut University, Assiut, Egypt
First published on 12th May 2026
Abstract
The development of novel microtubule-targeting medicines (MTAs) remains a crucial strategy in cancer treatment, as they combat drug resistance and systemic toxicity. A novel series of thiazole-based derivatives was synthesized, characterized, and evaluated as antitubulin agents endowed with antiproliferative action. An IC50 experiment was performed to assess the efficacy of novel compounds 9a–o in suppressing tubulin activity. The antiproliferative effects of the most potent compounds were evaluated. The levels of initiator caspases (Caspase-8 and Caspase-9) and executioner caspase (Caspase-3) were examined to ascertain the degree of apoptosis. Additionally, the expression levels of the mitochondrial regulatory proteins Bax and Bcl-2 were examined to ascertain the importance of the intrinsic apoptotic pathway. Compound 9k exhibited significant inhibition of tubulin with an IC50 of 1.56 µM and showed potent activity against HeLa (cervical), HCT-116 (colorectal), and A-549 (lung) cancer cell lines. The apoptotic assays revealed that 9k effectively triggered the apoptotic cascade, leading to a ninefold increase in Caspase-3 and a significant twenty-onefold rise in Caspase-9, surpassing the effects of the reference Staurosporine. Caspase-8 was activated by an elevenfold increase, primarily via the intrinsic pathway. This was confirmed by a substantial change in the mitochondrial “rheostat”: 9k induced a 38-fold increase in pro-apoptotic Bax and a 5-fold decrease in anti-apoptotic Bcl-2. Molecular docking studies showed that 9k exhibited a favorable binding mode consistent with its tubulin-inhibition profile. In silico ADMET predictions further supported 9k as a drug-like lead with acceptable oral exposure and a beneficial P-gp–related transporter profile. The enhanced apoptotic effects of 9k compared with Staurosporine make 9k an attractive lead candidate for further development as an anti-cancer drug targeting the colchicine-binding site.
1. Introduction
Cancer is a huge concern in global healthcare. It occurs when cells proliferate uncontrollably and fail to undergo programmed cell death (apoptosis).1 A key approach to combating cancer is to modify microtubule function. This remains an essential component of chemotherapy.2,3 Microtubules, composed of α- and β-tubulin heterodimers, are essential for fundamental cellular processes such as intracellular transport, structural integrity, and the formation of the mitotic spindle during cell division.4,5Taxanes and vinca alkaloids are tubulin inhibitors that cause cell death by controlling the assembly of microtubules. This interference triggers the spindle assembly checkpoint, which halts the cell cycle at the G2/M phase.6,7 However, the therapeutic efficacy of existing tubulin-targeting medicines is often limited by the development of multidrug resistance and significant systemic toxicity.8,9 Consequently, there is a critical need to develop new small-molecule tubulin inhibitors with strong antiproliferative effects and a distinct apoptotic mechanism.
Taxanes are therapeutically effective microtubule-targeting agents that act on polymerized microtubules.10 The colchicine binding site (CBS) has distinct pharmacological advantages. The CBS serves as the primary target for many small-molecule destabilizers. It is situated at the junction of the α- and β-tubulin subunits.11 Binding at this juncture inhibits tubulin from adopting the “linear” conformation needed for microtubule assembly. This inhibits polymerization by complicating molecular aggregation.12,13
Targeting the CBS is particularly advantageous, as these inhibitors exhibit poor substrate characteristics for the P-glycoprotein (P-gp) efflux pump, thereby allowing them to remain effective against MDR cancer cells.14,15 Furthermore, some colchicine-site drugs exhibit substantial vascular-disrupting properties, directly targeting established tumor vasculature.16 Colchicine, the primary alkaloid, is highly potent; however, its narrow therapeutic index and notable systemic toxicity have prompted the development of novel synthetic analogs that retain efficacy while minimizing off-target effects.17
The intrinsic apoptotic pathway regulates the transition from mitotic arrest to cell death. The Bcl-2 protein family is integral to this process.18,19 This “mitochondrial rheostat” maintains a precise balance between pro-apoptotic proteins, such as Bax, and anti-apoptotic proteins, such as Bcl-2. When this equilibrium shifts in favor of Bax, mitochondrial outer membrane permeabilization (MOMP) occurs, allowing cytochrome c to escape and activating Caspase-9.20 This initiator caspase subsequently activates the executioner caspase-3, initiating the final phases of cell degradation. Moreover, growing evidence suggests that potent tubulin inhibitors may activate the extrinsic pathway via Caspase-8, either by directly activating death receptors or through intricate pathway interactions.21,22
Thiazole-based derivatives have emerged as effective antitubulin drugs because the thiazole ring acts as a stiff bioisosteres for the double bond found in natural inhibitors such as Combretastatin A-4 (CA-4). These compounds frequently bind to tubulin's colchicine-binding site, blocking microtubule polymerization, slowing cell division during the G2/M phase, and killing cancer cells.23,24
Hashem et al.23 described a series of thiazole-privileged chalcones that impede tubulin polymerization, designating compound I, Fig. 1, as the most significant candidate. In the NCI single-dose screen, compound I showed a low mean growth percentage (22.13), indicating potent antiproliferative activity. Five-dose testing across the panel showed low micromolar growth inhibition, with GI50 values ranging from 1.55 to 2.95 µM against OVCAR-3 and MDA-MB-468. Compound I reduced tubulin polymerization with an IC50 of 7.78 µM (CA-4: 4.93 µM).
Khasawneh et al.25 enhanced this scaffold by integrating a para-sulfamoyl group to develop a dual tubulin/CA IX inhibitor, compound II (Fig. 1). Compound II exhibited significant cytotoxicity against HT-29 cells (IC50 = 0.98 µM) while demonstrating favorable selectivity for normal WI-38 fibroblasts (IC50 = 44.06 µM), and it inhibited tubulin polymerization (IC50 = 2.72 µM, comparable to CA-4 at 2.97 µM). Significantly, it effectively suppressed CA IX (IC50 = 0.021 µM) and triggered apoptosis, as evidenced by elevated p53 and Bax levels, decreased Bcl-2 levels, and activation of caspase-3 and caspase-9.
In our latest study,26 we identified thiazole-2-acetamide derivatives as inhibitors of tubulin polymerization, with compound III (Fig. 1) demonstrating the highest activity. Compound III reduced tubulin polymerization with an IC50 of 2.69 µM and exhibited antiproliferative action in various cancer cell lines, with an average GI50 of approximately 6.0 µM. The series had a favorable profile against normal cells, maintaining cell viability above about 85% at 50 µM. In contrast, mechanistic experiments indicated that apoptosis was induced by elevated Bax and caspase-3/9 levels, alongside concurrent downregulation of Bcl-2.
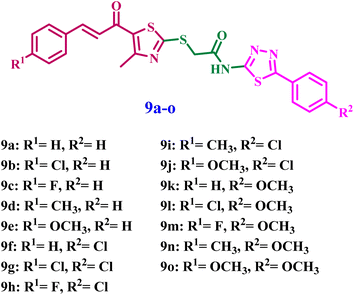
Inspired by prior data and in furtherance of our objective of developing effective tubulin polymerization inhibitors with improved antiproliferative efficacy,26–29 we present the synthesis of novel thiazole-thiadiazole derivatives (9a–o) (Fig. 2) for evaluation as antiproliferative agents targeting the colchicine-binding site in tubulin. The novel 9a–o compounds were assessed for their antitubulin activity, and the most effective derivatives were further examined for antiproliferative activity against a selection of cancer cell lines that express tubulin. The concentrations of initiator caspases (Caspase-8 and Caspase-9) and the executioner caspase (Caspase-3) were analyzed to determine the extent of apoptosis. The expression levels of the mitochondrial regulatory proteins Bax and Bcl-2 were analyzed to determine the significance of the intrinsic apoptotic pathway. Subsequently, docking analysis and ADMET investigation followed.
The rational design of the target compounds focused on developing a structurally stable mimic of combretastatin A-4 (CA-4), a strong tubulin inhibitor whose clinical application is frequently hindered by the metabolic isomerization of its cis-olefinic bridge. To resolve this, we substituted the unstable ethene linker with a rigid thiazole core. This heterocyclic framework serves as a structural “anchor,” keeping the aromatic substituents in a bioactive orientation that emulates the cis-conformation of CA-4 while improving chemical stability. The incorporation of a chalcone-like pharmacophore, recognized for its efficient occupation of the colchicine binding site by aligning its aromatic rings within the deep hydrophobic cavities of the β-tubulin subunit.
Further optimization entailed the integration of a phenyl-1,3,4-thiadiazole terminal moiety as a bioisosteres for the 3,4,5-trimethoxyphenyl A-ring of CA-4. This modification preserves essential hydrophobic interactions while incorporating nitrogen and sulfur heteroatoms that can establish supplementary hydrogen bonds with critical residues in the binding pocket. Finally, the thioacetamide linker connecting these heterocyclic systems imparts the requisite conformational flexibility, enabling the molecule to “thread” into the α/β-tubulin interface, which may augment binding affinity through a synthesis of conventional hydrophobic anchoring and improved polar interactions.
2. Experimental
2.1. Chemistry
General details: refer to Appendix A (SI).3-Chloroacetylacetone (2) (ref. 30), 1-(2-mercapto-4-methylthiazol-5-yl)ethan-1-one and (4a–e),23 (6a–c), and (7a–c)31 were synthesized according to reported procedures.
2.1.1.1. 2-((5-Cinnamoyl-4-methylthiazol-2-yl)thio)-N-(5-phenyl-1,3,4-thiadiazol-2-yl) acetamide (9a). Yellow powder; 0.421 g, 88% yield; mp 244–246 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.07 (s, 1H, NH), 7.91 (dd, J = 6.5, 2.8 Hz, 2H, Ar–H), 7.76 (dd, J = 6.3, 2.6 Hz, 2H, Ar–H), 7.64 (d, J = 15.6 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.52–7.46 (m, 3H, Ar–H), 7.45–7.40 (m, 3H, Ar–H), 7.33 (d, J = 15.5 Hz, 1H, CH
CH), 7.52–7.46 (m, 3H, Ar–H), 7.45–7.40 (m, 3H, Ar–H), 7.33 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 4.44 (s, 2H, CH2), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.07, 168.26, 166.72, 162.72, 158.84, 158.27, 144.26, 134.65, 132.80, 131.43, 131.24, 130.56, 129.91, 129.54, 129.34, 127.48, 124.82, 37.23, 18.81; anal. calcd. For C23H18N4O2S3: C, 57.72%; H, 3.79%; N, 11.71%. Found: C, 57.52%; H, 3.98%; N, 11.53%.
CH), 4.44 (s, 2H, CH2), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.07, 168.26, 166.72, 162.72, 158.84, 158.27, 144.26, 134.65, 132.80, 131.43, 131.24, 130.56, 129.91, 129.54, 129.34, 127.48, 124.82, 37.23, 18.81; anal. calcd. For C23H18N4O2S3: C, 57.72%; H, 3.79%; N, 11.71%. Found: C, 57.52%; H, 3.98%; N, 11.53%.
2.1.1.2. (E)-2-((5-(3-(4-Chlorophenyl)acryloyl)-4-methylthiazol-2-yl)thio)-N-(5-phenyl-1,3,4-thiadiazol-2-yl)acetamide (9b). White powder; 0.416 g, 81% yield; mp 257–260 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.08 (s, 1H, NH), 7.90 (s, 2H, Ar–H), 7.82–7.74 (m, 2H, Ar–H), 7.62 (d, J = 14.8, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.53–7.42 (m, 5H, Ar–H), 7.34 (d, J = 15.4, 1H, CH
CH), 7.53–7.42 (m, 5H, Ar–H), 7.34 (d, J = 15.4, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 4.45 (s, 2H, CH2), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.91, 168.40, 166.69, 162.71, 158.82, 158.43, 142.77, 135.90, 133.62, 132.71, 131.23, 131.04, 130.55, 129.90, 129.56, 127.48, 125.49, 37.23, 18.83; anal. calcd. For C23H17ClN4O2S3: C, 53.85%; H, 3.34%; N, 10.92%. Found: C, 53.94%; H, 3.26%; N, 10.74%.
CH), 4.45 (s, 2H, CH2), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.91, 168.40, 166.69, 162.71, 158.82, 158.43, 142.77, 135.90, 133.62, 132.71, 131.23, 131.04, 130.55, 129.90, 129.56, 127.48, 125.49, 37.23, 18.83; anal. calcd. For C23H17ClN4O2S3: C, 53.85%; H, 3.34%; N, 10.92%. Found: C, 53.94%; H, 3.26%; N, 10.74%.
2.1.1.3. (E)-2-((5-(3-(4-Fluorophenyl)acryloyl)-4-methylthiazol-2-yl)thio)-N-(5-phenyl-1,3,4-thiadiazol-2-yl)acetamide (9c). Yellow powder; 0.447 g, 90% yield; mp 239–241 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.07 (s, 1H, NH), 7.91 (dd, J = 6.4, 2.3 Hz, 2H, Ar–H), 7.87–7.83 (m, 2H, Ar–H), 7.65 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.52–7.47 (m, 3H, Ar–H), 7.30 (d, J = 15.6 Hz, 1H, CH
CH), 7.52–7.47 (m, 3H, Ar–H), 7.30 (d, J = 15.6 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.26 (t, J = 8.8 Hz, 2H, Ar–H), 4.44 (s, 2H, CH2), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.00, 168.32, 166.68, 162.71, 161.95, 158.83, 158.25, 143.02, 134.62, 132.78, 131.43, 131.22, 130.56, 129.91, 129.54, 127.48, 124.81, 116.65, 116.48, 37.23, 18.80; anal. calcd. For C23H17FN4O2S3: C, 55.63%; H, 3.45%; N, 11.28%. Found: C, 55.88%; H, 3.25%; N, 11.07%.
CH), 7.26 (t, J = 8.8 Hz, 2H, Ar–H), 4.44 (s, 2H, CH2), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.00, 168.32, 166.68, 162.71, 161.95, 158.83, 158.25, 143.02, 134.62, 132.78, 131.43, 131.22, 130.56, 129.91, 129.54, 127.48, 124.81, 116.65, 116.48, 37.23, 18.80; anal. calcd. For C23H17FN4O2S3: C, 55.63%; H, 3.45%; N, 11.28%. Found: C, 55.88%; H, 3.25%; N, 11.07%.
2.1.1.4. (E)-2-((4-Methyl-5-(3-(p-tolyl)acryloyl)thiazol-2-yl)thio)-N-(5-phenyl-1,3,4-thiadiazol-2-yl)acetamide (9d). Yellow powder; 0.419 g, 85% yield; mp 268–271 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.08 (s, 1H, NH), 7.94–7.86 (m, 2H, Ar–H), 7.67–7.57 (m, 3H, Ar–H & CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.49 (s, 3H, Ar–H), 7.26 (d, J = 15.5 Hz, 1H, CH
CH), 7.49 (s, 3H, Ar–H), 7.26 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.22 (d, J = 7.3 Hz, 2H, Ar–H), 4.44 (s, 2H, CH2), 2.60 (s, 3H, CH3), 2.30 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.98, 168.03, 166.70, 162.71, 158.83, 158.11, 144.37, 141.61, 132.84, 131.93, 131.23, 130.56, 130.16, 129.89, 129.38, 127.48, 123.73, 37.23, 21.64, 18.78; anal. calcd. For C24H20N4O2S3: C, 58.52%; H, 4.09%; N, 11.37%. Found: C, 58.33%; H, 3.99%; N, 11.55%.
CH), 7.22 (d, J = 7.3 Hz, 2H, Ar–H), 4.44 (s, 2H, CH2), 2.60 (s, 3H, CH3), 2.30 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.98, 168.03, 166.70, 162.71, 158.83, 158.11, 144.37, 141.61, 132.84, 131.93, 131.23, 130.56, 130.16, 129.89, 129.38, 127.48, 123.73, 37.23, 21.64, 18.78; anal. calcd. For C24H20N4O2S3: C, 58.52%; H, 4.09%; N, 11.37%. Found: C, 58.33%; H, 3.99%; N, 11.55%.
2.1.1.5. (E)-2-((5-(3-(4-Methoxyphenyl)acryloyl)-4-methylthiazol-2-yl)thio)-N-(5-phenyl-1,3,4-thiadiazol-2-yl)acetamide (9e). Yellowish white powder; 0.402 g, 79% yield; mp 250–252 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.07 (s, 1H, NH), 7.95–7.87 (m, 2H, Ar–H), 7.73 (d, J = 8.7 Hz, 2H, Ar–H), 7.62 (d, J = 15.4 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.53–7.46 (m, 3H, Ar–H), 7.19 (d, J = 15.4 Hz, 1H, CH
CH), 7.53–7.46 (m, 3H, Ar–H), 7.19 (d, J = 15.4 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 6.97 (d, J = 8.7 Hz, 2H, Ar–H), 4.44 (s, 2H, CH2), 3.78 (s, 3H, OCH3), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.89, 167.74, 166.71, 162.72, 162.14, 158.82, 157.84, 144.36, 132.97, 131.33, 131.23, 130.56, 129.90, 127.48, 127.25, 122.22, 115.04, 55.93, 37.22, 18.75; anal. calcd. For C24H20N4O3S3: C, 56.67%; H, 3.96%; N, 11.02%. Found: C, 56.53%; H, 4.04%; N, 10.80%.
CH), 6.97 (d, J = 8.7 Hz, 2H, Ar–H), 4.44 (s, 2H, CH2), 3.78 (s, 3H, OCH3), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.89, 167.74, 166.71, 162.72, 162.14, 158.82, 157.84, 144.36, 132.97, 131.33, 131.23, 130.56, 129.90, 127.48, 127.25, 122.22, 115.04, 55.93, 37.22, 18.75; anal. calcd. For C24H20N4O3S3: C, 56.67%; H, 3.96%; N, 11.02%. Found: C, 56.53%; H, 4.04%; N, 10.80%.
2.1.1.6. N-(5-(4-Chlorophenyl)-1,3,4-thiadiazol-2-yl)-2-((5-cinnamoyl-4-methylthiazol-2-yl)thio)acetamide (9f). Yellow powder; 0.446 g, 87% yield; mp 275–278 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.11 (s, 1H, NH), 7.96–7.89 (m, 2H, Ar–H), 7.76 (s, 2H, Ar–H), 7.65 (dd, J = 15.6, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.59–7.52 (m, 2H, Ar–H), 7.43 (s, 3H, Ar–H), 7.34 (dd, J = 15.4, 1H, CH
CH), 7.59–7.52 (m, 2H, Ar–H), 7.43 (s, 3H, Ar–H), 7.34 (dd, J = 15.4, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 4.45 (s, 2H, CH2), 2.62 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.05, 168.27, 166.85, 161.49, 159.29, 158.26, 144.25, 135.77, 134.66, 133.85, 132.79, 131.43, 129.94, 129.54, 129.34, 129.14, 124.82, 37.30, 18.80; anal. calcd. For C23H17ClN4O2S3: C, 53.85%; H, 3.34%; N, 10.92%. Found: C, 53.69%; H, 3.49%; N, 11.14%.
CH), 4.45 (s, 2H, CH2), 2.62 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.05, 168.27, 166.85, 161.49, 159.29, 158.26, 144.25, 135.77, 134.66, 133.85, 132.79, 131.43, 129.94, 129.54, 129.34, 129.14, 124.82, 37.30, 18.80; anal. calcd. For C23H17ClN4O2S3: C, 53.85%; H, 3.34%; N, 10.92%. Found: C, 53.69%; H, 3.49%; N, 11.14%.
2.1.1.7. (E)-N-(5-(4-Chlorophenyl)-1,3,4-thiadiazol-2-yl)-2-((5-(3-(4-chlorophenyl)acryloyl)-4-methylthiazol-2-yl)thio)acetamide (9g). White powder; 0.416 g, 76% yield; mp 262–264 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.10 (s, 1H, NH), 7.92 (d, J = 8.5 Hz, 2H, Ar–H), 7.79 (d, J = 8.3 Hz, 2H, Ar–H), 7.62 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.55 (d, J = 8.4 Hz, 2H, Ar–H), 7.47 (d, J = 8.4 Hz, 2H, Ar–H), 7.33 (d, J = 15.5 Hz, 1H, CH
CH), 7.55 (d, J = 8.4 Hz, 2H, Ar–H), 7.47 (d, J = 8.4 Hz, 2H, Ar–H), 7.33 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 4.44 (s, 2H, CH2), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.91, 168.35, 166.76, 161.56, 159.14, 158.41, 142.77, 135.90, 135.80, 133.63, 132.71, 131.03, 129.94, 129.55, 129.43, 129.14, 125.50, 37.24, 18.81; anal. calcd. For C23H16Cl2N4O2S3: C, 50.46%; H, 2.95%; N, 10.23%. Found: C, 50.31%; H, 3.03%; N, 10.34%.
CH), 4.44 (s, 2H, CH2), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.91, 168.35, 166.76, 161.56, 159.14, 158.41, 142.77, 135.90, 135.80, 133.63, 132.71, 131.03, 129.94, 129.55, 129.43, 129.14, 125.50, 37.24, 18.81; anal. calcd. For C23H16Cl2N4O2S3: C, 50.46%; H, 2.95%; N, 10.23%. Found: C, 50.31%; H, 3.03%; N, 10.34%.
2.1.1.8. (E)-N-(5-(4-Chlorophenyl)-1,3,4-thiadiazol-2-yl)-2-((5-(3-(4-fluorophenyl)acryloyl)-4-methylthiazol-2-yl)thio)acetamide (9h). Yellow powder; 0.425 g, 80% yield; mp 246–249 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.13 (s, 1H, NH), 7.92 (d, J = 8.1 Hz, 2H, Ar–H), 7.88–7.81 (m, 2H, Ar–H), 7.64 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.55 (d, J = 8.3 Hz, 2H, Ar–H), 7.33–7.22 (m, 3H, Ar–H & CH
CH), 7.55 (d, J = 8.3 Hz, 2H, Ar–H), 7.33–7.22 (m, 3H, Ar–H & CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 4.43 (s, 2H, CH2), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.94, 168.20, 166.75, 161.57, 159.08, 158.27, 143.05, 135.80, 132.77, 131.79, 131.35, 129.93, 129.42, 129.14, 124.66, 116.65, 116.48, 37.22, 18.80; anal. calcd. For C23H16ClFN4O2S3: C, 52.02%; H, 3.04%; N, 10.55%. Found: C, 51.79%; H, 2.96%; N, 10.46%.
CH), 4.43 (s, 2H, CH2), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.94, 168.20, 166.75, 161.57, 159.08, 158.27, 143.05, 135.80, 132.77, 131.79, 131.35, 129.93, 129.42, 129.14, 124.66, 116.65, 116.48, 37.22, 18.80; anal. calcd. For C23H16ClFN4O2S3: C, 52.02%; H, 3.04%; N, 10.55%. Found: C, 51.79%; H, 2.96%; N, 10.46%.
2.1.1.9. (E)-N-(5-(4-Chlorophenyl)-1,3,4-thiadiazol-2-yl)-2-((4-methyl-5-(3-(p-tolyl)acryloyl) thiazol-2-yl)thio)acetamide (9i). Yellow powder; 0.437 g, 83% yield; mp 271–273 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.09 (s, 1H, NH), 7.93 (d, J = 6.5 Hz, 2H, Ar–H), 7.66–7.57 (m, 3H, Ar–H & CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.55 (d, J = 6.6 Hz, 2H, Ar–H), 7.29–7.20 (m, 3H, Ar–H & CH
CH), 7.55 (d, J = 6.6 Hz, 2H, Ar–H), 7.29–7.20 (m, 3H, Ar–H & CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 4.44 (s, 2H, CH2), 2.60 (s, 3H, CH3), 2.31 (s, 3H, Ar-CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.98, 167.99, 166.40, 161.58, 158.10, 156.85, 144.37, 141.62, 135.80, 132.84, 131.92, 130.16, 129.95, 129.40, 129.16, 123.72, 111.11, 37.21, 21.64, 18.83; anal. calcd. For C24H19ClN4O2S3: C, 54.69%; H, 3.63%; N, 10.63%. Found: C, 54.61%; H, 3.49%; N, 10.85%.
CH), 4.44 (s, 2H, CH2), 2.60 (s, 3H, CH3), 2.31 (s, 3H, Ar-CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.98, 167.99, 166.40, 161.58, 158.10, 156.85, 144.37, 141.62, 135.80, 132.84, 131.92, 130.16, 129.95, 129.40, 129.16, 123.72, 111.11, 37.21, 21.64, 18.83; anal. calcd. For C24H19ClN4O2S3: C, 54.69%; H, 3.63%; N, 10.63%. Found: C, 54.61%; H, 3.49%; N, 10.85%.
2.1.1.10. (E)-N-(5-(4-Chlorophenyl)-1,3,4-thiadiazol-2-yl)-2-((5-(3-(4-methoxyphenyl) acryloyl)-4-methylthiazol-2-yl)thio)acetamide (9j). White powder; 0.402 g, 74% yield; mp 237–239 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.11 (s, 1H, NH), 7.93 (d, J = 7.6 Hz, 2H, Ar–H), 7.72 (d, J = 7.5 Hz, 2H, Ar–H), 7.61 (d, J = 15.4 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.55 (d, J = 7.3 Hz, 2H, Ar–H), 7.18 (d, J = 15.5 Hz, 1H, CH
CH), 7.55 (d, J = 7.3 Hz, 2H, Ar–H), 7.18 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 6.97 (d, J = 7.5 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.78 (s, 3H, OCH3), 2.59 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.88, 167.69, 166.80, 162.14, 161.58, 159.10, 157.83, 144.36, 135.80, 132.98, 131.33, 129.94, 129.43, 129.15, 127.25, 122.21, 115.04, 55.93, 37.21, 18.73; anal. calcd. For C24H19ClN4O3S3: C, 53.08%; H, 3.53%; N, 10.32%. Found: C, 52.99%; H, 3.77%; N, 10.15%.
CH), 6.97 (d, J = 7.5 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.78 (s, 3H, OCH3), 2.59 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.88, 167.69, 166.80, 162.14, 161.58, 159.10, 157.83, 144.36, 135.80, 132.98, 131.33, 129.94, 129.43, 129.15, 127.25, 122.21, 115.04, 55.93, 37.21, 18.73; anal. calcd. For C24H19ClN4O3S3: C, 53.08%; H, 3.53%; N, 10.32%. Found: C, 52.99%; H, 3.77%; N, 10.15%.
2.1.1.11. 2-((5-Cinnamoyl-4-methylthiazol-2-yl)thio)-N-(5-(4-methoxyphenyl)-1,3,4-thiadiazol-2-yl)acetamide (9k). Yellow powder; 0.463 g, 91% yield; mp 277–280 °C; 1H NMR (500 MHz, DMSO-d6) δ 12.99 (s, 1H, NH), 7.84 (d, J = 8.6 Hz, 2H, Ar–H), 7.77–7.72 (m, 2H, Ar–H), 7.64 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.45–7.39 (m, 3H, Ar–H), 7.32 (d, J = 15.5 Hz, 1H, CH
CH), 7.45–7.39 (m, 3H, Ar–H), 7.32 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.03 (d, J = 8.6 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.78 (s, 3H, OCH3), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.03, 168.31, 166.56, 162.52, 161.66, 158.29, 158.18, 144.24, 134.64, 132.78, 131.43, 129.54, 129.33, 129.04, 124.78, 123.08, 115.28, 55.95, 37.24, 18.81; anal. calcd. For C24H20N4O3S3: C, 56.67%; H, 3.96%; N, 11.02%. Found: C, 56.54%; H, 3.75%; N, 11.21%.
CH), 7.03 (d, J = 8.6 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.78 (s, 3H, OCH3), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 182.03, 168.31, 166.56, 162.52, 161.66, 158.29, 158.18, 144.24, 134.64, 132.78, 131.43, 129.54, 129.33, 129.04, 124.78, 123.08, 115.28, 55.95, 37.24, 18.81; anal. calcd. For C24H20N4O3S3: C, 56.67%; H, 3.96%; N, 11.02%. Found: C, 56.54%; H, 3.75%; N, 11.21%.
2.1.1.12. (E)-2-((5-(3-(4-Chlorophenyl)acryloyl)-4-methylthiazol-2-yl)thio)-N-(5-(4-methoxy phenyl)-1,3,4-thiadiazol-2-yl)acetamide (9l). Yellow powder; 0.424 g, 78% yield; mp 255–257 °C; 1H NMR (500 MHz, DMSO-d6) δ 12.99 (s, 1H, NH), 7.84 (d, J = 8.8 Hz, 2H, Ar–H), 7.80 (d, J = 8.4 Hz, 2H, Ar–H), 7.63 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.47 (d, J = 8.4 Hz, 2H, Ar–H), 7.34 (d, J = 15.5 Hz, 1H, CH
CH), 7.47 (d, J = 8.4 Hz, 2H, Ar–H), 7.34 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.04 (d, J = 8.8 Hz, 2H, Ar–H), 4.42 (s, 2H, CH2), 3.79 (s, 3H, OCH3), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.91, 168.45, 166.53, 162.52, 161.66, 158.44, 158.14, 142.77, 135.90, 133.62, 132.70, 131.05, 129.56, 129.05, 125.48, 123.07, 115.28, 55.95, 37.22, 18.82; anal. calcd. For C24H19ClN4O3S3: C, 53.08%; H, 3.53%; N, 10.32%. Found: C, 53.21%; H, 3.66%; N, 10.14%.
CH), 7.04 (d, J = 8.8 Hz, 2H, Ar–H), 4.42 (s, 2H, CH2), 3.79 (s, 3H, OCH3), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.91, 168.45, 166.53, 162.52, 161.66, 158.44, 158.14, 142.77, 135.90, 133.62, 132.70, 131.05, 129.56, 129.05, 125.48, 123.07, 115.28, 55.95, 37.22, 18.82; anal. calcd. For C24H19ClN4O3S3: C, 53.08%; H, 3.53%; N, 10.32%. Found: C, 53.21%; H, 3.66%; N, 10.14%.
2.1.1.13. (E)-2-((5-(3-(4-fluorophenyl)acryloyl)-4-methylthiazol-2-yl)thio)-N-(5-(4-methoxy phenyl)-1,3,4-thiadiazol-2-yl)acetamide (9m). White powder; 0.432 g, 82% yield; mp 263–266 °C; 1H NMR (500 MHz, DMSO-d6) δ 12.99 (s, 1H, NH), 7.88–7.81 (m, 4H, Ar–H), 7.65 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.30 (d, J = 15.6 Hz, 1H, CH
CH), 7.30 (d, J = 15.6 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.28–7.24 (m, 2H, Ar–H), 7.04 (d, J = 8.7 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.79 (s, 3H, OCH3), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.97, 168.26, 163.08, 162.54, 161.67, 158.27, 143.04, 132.76, 131.80, 131.73, 131.36, 129.05, 124.69, 123.09, 116.65, 116.48, 115.29, 55.96, 37.23, 18.81; anal. calcd. For C24H19FN4O3S3: C, 54.74%; H, 3.64%; N, 10.64%. Found: C, 54.50%; H, 3.47%; N, 10.89%.
CH), 7.28–7.24 (m, 2H, Ar–H), 7.04 (d, J = 8.7 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.79 (s, 3H, OCH3), 2.61 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.97, 168.26, 163.08, 162.54, 161.67, 158.27, 143.04, 132.76, 131.80, 131.73, 131.36, 129.05, 124.69, 123.09, 116.65, 116.48, 115.29, 55.96, 37.23, 18.81; anal. calcd. For C24H19FN4O3S3: C, 54.74%; H, 3.64%; N, 10.64%. Found: C, 54.50%; H, 3.47%; N, 10.89%.
2.1.1.14. (E)-N-(5-(4-Methoxyphenyl)-1,3,4-thiadiazol-2-yl)-2-((4-methyl-5-(3-(p-tolyl) acryloyl)thiazol-2-yl)thio)acetamide (9n). Yellowish white powder; 0.449 g, 86% yield; mp 248–250 °C; 1H NMR (500 MHz, DMSO-d6) δ 12.99 (s, 1H, NH), 7.87–7.81 (m, 2H, Ar–H), 7.68–7.59 (m, 3H, Ar–H), 7.27 (d, J = 15.7 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.23 (d, J = 5.9 Hz, 2H, Ar–H), 7.07–7.02 (m, 2H, Ar–H), 4.42 (s, 2H, CH2), 3.79 (s, 3H, OCH3), 2.61 (s, 3H, CH3), 2.31 (s, 3H, Ar–CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.96, 169.40, 168.11, 166.56, 162.52, 161.65, 158.12, 144.37, 141.62, 132.83, 131.91, 130.16, 129.40, 129.05, 123.70, 123.07, 115.28, 55.94, 37.22, 21.65, 18.79; anal. calcd. For C25H22N4O3S3: C, 57.45%; H, 4.24%; N, 10.72%. Found: C, 57.37%; H, 4.02%; N, 10.92%.
CH), 7.23 (d, J = 5.9 Hz, 2H, Ar–H), 7.07–7.02 (m, 2H, Ar–H), 4.42 (s, 2H, CH2), 3.79 (s, 3H, OCH3), 2.61 (s, 3H, CH3), 2.31 (s, 3H, Ar–CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.96, 169.40, 168.11, 166.56, 162.52, 161.65, 158.12, 144.37, 141.62, 132.83, 131.91, 130.16, 129.40, 129.05, 123.70, 123.07, 115.28, 55.94, 37.22, 21.65, 18.79; anal. calcd. For C25H22N4O3S3: C, 57.45%; H, 4.24%; N, 10.72%. Found: C, 57.37%; H, 4.02%; N, 10.92%.
2.1.1.15. (E)-N-(5-(4-Methoxyphenyl)-1,3,4-thiadiazol-2-yl)-2-((5-(3-(4-methoxyphenyl) acryloyl)-4-methylthiazol-2-yl)thio)acetamide (9o). White powder; 0.404 g, 75% yield; mp 240–242 °C; 1H NMR (500 MHz, DMSO-d6) δ 13.05 (s, 1H, NH), 7.87 (d, J = 8.8 Hz, 2H, Ar–H), 7.73 (d, J = 8.7 Hz, 2H, Ar–H), 7.61 (d, J = 15.5 Hz, 1H, CH
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.21 (d, J = 15.5 Hz, 1H, CH
CH), 7.21 (d, J = 15.5 Hz, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.04 (d, J = 8.8 Hz, 2H, Ar–H), 6.97 (d, J = 8.7 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.79 (s, 6H, 2×OCH3), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.90, 167.72, 166.80, 162.52, 162.14, 161.66, 159.10, 157.83, 144.35, 132.98, 131.33, 130.56, 129.90, 129.54, 127.25, 122.22, 115.28, 115.04, 55.94, 37.22, 18.74; anal. calcd. For C25H22N4O4S3: C, 55.75%; H, 4.12%; N, 10.40%. Found: C, 55.83%; H, 4.22%; N, 10.30%.
CH), 7.04 (d, J = 8.8 Hz, 2H, Ar–H), 6.97 (d, J = 8.7 Hz, 2H, Ar–H), 4.43 (s, 2H, CH2), 3.79 (s, 6H, 2×OCH3), 2.60 (s, 3H, CH3); 13C NMR (120 MHz, DMSO-d6) δ 181.90, 167.72, 166.80, 162.52, 162.14, 161.66, 159.10, 157.83, 144.35, 132.98, 131.33, 130.56, 129.90, 129.54, 127.25, 122.22, 115.28, 115.04, 55.94, 37.22, 18.74; anal. calcd. For C25H22N4O4S3: C, 55.75%; H, 4.12%; N, 10.40%. Found: C, 55.83%; H, 4.22%; N, 10.30%.
2.2. Biology
3. Results and discussion
3.1. Chemistry
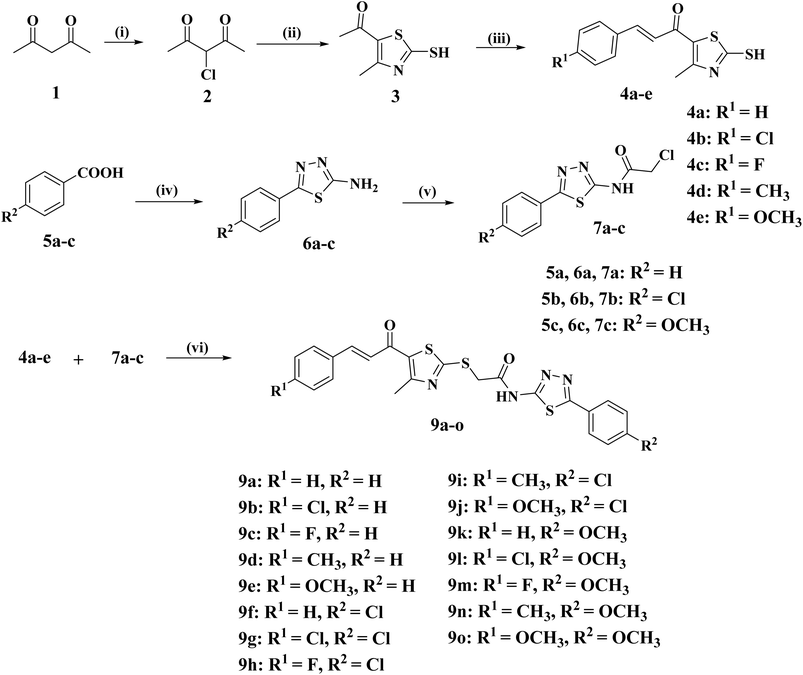
As outlined in Scheme 1, the target compounds 9a–o were synthesized via a convergent synthetic route involving two key intermediates: mercapto-thiazole chalcones (4a–e) and chloroacetamido-thiadiazoles (7a–c). The thiazole core was synthesized through heterocyclization of 3-chloro acetylacetone30 with carbon disulfide and ammonia to yield the thiazole intermediate 3, which subsequently underwent base-catalyzed Claisen–Schmidt condensation with substituted aldehydes to afford the corresponding chalcones (4a–e).23Concurrently, substituted benzoic acids 5a–c were converted into 1,3,4-thiadiazole amines (6a–c) via condensation with thiosemicarbazide followed by POCl3-mediated cyclodehydration. These were then acylated with chloroacetyl chloride to give the electrophilic chloroacetamides (7a–c).31 Final coupling with chalcones (4a–e) was achieved under mild basic conditions through an S-alkylation reaction, affording the final compounds 9a–o.
Compounds 9a–o were characterized by 1H NMR, 13C NMR, and elemental analysis (C, H, N), confirming formation of the targeted thioether-linked thiazole–chalcone/1,3,4-thiadiazole hybrids. In the 1H NMR spectra, all derivatives displayed a diagnostic amide NH as a downfield singlet at δ 12.99–13.13 (1H, s). The α,β-unsaturated enone fragment consistently appeared as two trans-olefinic doublets (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH) at δ 7.61–7.65 (1H, d, J ≈ 14.8–15.6 Hz) and δ 7.18–7.34 (1H, d, J ≈ 15.4–15.7 Hz), and the large coupling constant supports the E configuration across the series. The aromatic protons of the two phenyl rings resonated mainly in the δ 6.97–7.96 region. The characteristic methylene signal was observed as a singlet at δ 4.42–4.45 (2H, s), while the thiazole methyl group appeared as a sharp singlet at δ 2.59–2.62 (3H, s). Compounds containing methoxy groups showed an extra singlet at 3.78–3.79, while p-tolyl derivatives showed an additional singlet at δ 2.30–2.31.
CH) at δ 7.61–7.65 (1H, d, J ≈ 14.8–15.6 Hz) and δ 7.18–7.34 (1H, d, J ≈ 15.4–15.7 Hz), and the large coupling constant supports the E configuration across the series. The aromatic protons of the two phenyl rings resonated mainly in the δ 6.97–7.96 region. The characteristic methylene signal was observed as a singlet at δ 4.42–4.45 (2H, s), while the thiazole methyl group appeared as a sharp singlet at δ 2.59–2.62 (3H, s). Compounds containing methoxy groups showed an extra singlet at 3.78–3.79, while p-tolyl derivatives showed an additional singlet at δ 2.30–2.31.
In the 13C NMR spectra, the chalcone carbonyl was observed in the expected downfield region at δ 181.88–182.07, while the acetamide carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) appeared at δ 166.40–169.40. Aliphatic carbons were also diagnostic, such as the methylene moiety, which appeared at δ 37.21–37.30, the thiazole methyl at δ 18.73–18.83, the methoxy carbon at δ 55.93–55.96 (when present), and the p-tolyl methyl carbon at δ 21.64–21.65 (when present).
O) appeared at δ 166.40–169.40. Aliphatic carbons were also diagnostic, such as the methylene moiety, which appeared at δ 37.21–37.30, the thiazole methyl at δ 18.73–18.83, the methoxy carbon at δ 55.93–55.96 (when present), and the p-tolyl methyl carbon at δ 21.64–21.65 (when present).
As a representative example, compound 9l showed 1H and 13C signals fully consistent with its substitution pattern. In the 1H NMR spectrum, the amide proton appeared at δ 12.99, while the enone moiety gave two trans-coupled vinylic doublets at δ 7.63 and δ 7.34, confirming the E geometry. The aromatic region exhibited two para-disubstituted patterns and appeared as 4 doublets: the p-chlorophenyl chalcone ring resonated at δ 7.80 and δ 7.47, whereas the p-methoxyphenyl thiadiazole ring appeared at δ 7.84 and δ 7.04. The methylene was observed at δ 4.42, the methoxy group at δ 3.79, and the thiazole methyl at δ 2.61. In the 13C NMR spectrum, the expected enone carbonyl appeared at δ 181.91 and the amide carbonyl at δ 168.45. The methoxy carbon appeared at δ 55.95, while the methylene and thiazole methyl were observed at δ 37.22 and δ 18.82, respectively. Taken together, the 1H/13C features of 9l mirror the general spectral fingerprint of the series and further confirm the structures of 9a–o.
3.2. Biology
| Compound | R1 | R2 | Cell viability% | Tubulin inhibition IC50 ± SEM (µM) |
|---|---|---|---|---|
| a –: not applicable. | ||||
| 9a | H | H | 90 | 29.66 ± 1.20 |
| 9b | Cl | H | 93 | 6.07 ± 0.25 |
| 9c | F | H | 91 | 19.45 ± 1.10 |
| 9d | Me | H | 90 | 15.60 ± 0.70 |
| 9e | OMe | H | 92 | 3.04 ± 0.11 |
| 9f | H | Cl | 88 | 22.67 ± 0.81 |
| 9g | Cl | Cl | 91 | 5.33 ± 0.17 |
| 9h | F | Cl | 88 | 8.11 ± 0.30 |
| 9i | Me | Cl | 91 | 2.71 ± 0.09 |
| 9j | OMe | Cl | 93 | 11.60 ± 0.55 |
| 9k | H | OMe | 90 | 1.56 ± 0.05 |
| 9l | Cl | OMe | 89 | 37.21 ± 1.50 |
| 9m | F | OMe | 90 | 5.74 ± 0.20 |
| 9n | Me | OMe | 89 | 26.40 ± 1.05 |
| 9o | OMe | OMe | 90 | 32.15 ± 1.20 |
| CA-4 | — | — | — | 2.80 ± 0.10 |
Compound 9k (R1 = H, R2 = OMe) had the highest potency as an antitubulin agent, with an IC50 value of 1.56 µM, demonstrating double the efficacy of the reference compound CA-4. The anti-tubulin efficacy of compounds 9a–o is markedly influenced by the substitution pattern on the phenyl groups of the chalcone moiety (R1) and at the fourth position of the thiadiazole moiety (R2). For instance, compounds 9a (R1 = R2 = H) and 9f (R1 = H, R2 = Cl) both include an unsubstituted phenyl group on the chalcone moiety, akin to compound 9k, yet possess distinct substituents at the para position of the phenyl group inside the thiadiazole moiety. Compounds 9a and 9f exhibited diminished potency relative to 9k, with IC50 values of 29.66 µM for 9a and 22.67 µM for 9f, representing 19-fold and 15-fold reductions in potency compared to 9k, respectively. The data indicated that, when the phenyl group of the chalcone moiety is unsubstituted, the methoxy group is the optimal substituent on the phenyl group of the thiadiazole moiety, followed by the chlorine atom. Additionally, when both phenyl groups are unsubstituted, it is detrimental to antitubulin activity.
Conversely, compounds 9l (R1 = Cl, R2 = OMe), 9m (R1 = F, R2 = OMe), 9n (R1 = Me, R2 = OMe), and 9o (R1 = OMe, R2 = OMe) possess a methoxy group in the para position of the phenyl group within the thiadiazole moiety, similar to 9k, but feature varying substituents on the phenyl group of the chalcone moiety (R1). Except for 9m, compounds 9l, 9n, and 9o showed a significant reduction in antitubulin activity, with IC50 values of 37.21, 26.4, and 32.15 µM, respectively, indicating at least a 17-fold drop in potency relative to 9k. Compound 9m, containing a fluorine atom, showed significant antitubulin activity with an IC50 of 5.74 µM, 3.7-fold less effective than 9k. These data indicated that the presence of a methoxy group at the para position of the phenyl group in the thiadiazole moiety renders the unsubstituted phenyl group in the chalcone moiety optimum for activity. Substitution with electron-donating groups such as methyl and methoxy, or with electron-withdrawing groups such as chlorine or fluorine, reduces activity, with fluorine ranking second in effect after hydrogen.
Compound 9i (R1 = Me, R2 = Cl) scored second in antitubulin activity, with an IC50 value of 2.71 µM. It was 1.7-fold less effective than compound 9k, but marginally more potent than the standard CA-4. Compounds 9f (R1 = H, R2 = Cl), 9g (R1 = Cl, R2 = Cl), 9h (R1 = F, R2 = Cl), and 9j (R1 = OMe, R2 = Cl) all possess a chlorine atom as R2, similar to 9i, but feature distinct R1 substituents. Excluding 9f, compounds 9g, 9h, and 9j exhibited moderate antitubulin activity with IC50 values of 5.33, 8.11, and 11.60 µM, respectively. Compound 9f exhibited a significant reduction in its antitubulin activity, with an IC50 value of 22.67 µM. The study found that when a chlorine atom is present as an R2 substituent within the thiadiazole moiety, the unsubstituted phenyl group in the chalcone moiety has reduced activity. Still, chlorine and fluorine atoms (as R1, electron-withdrawing groups) are the best for activity. Ultimately, compound 9o (R1 = R2 = OMe) had the lowest potency among all synthesised derivatives as an antitubulin agent. It exhibited an IC50 of 32.15 µM, indicating a 20-fold reduction in activity compared to 9k, suggesting that the presence of a methoxy group on both phenyl rings is detrimental to activity.
| Compd | Antiproliferative activity IC50 ± SEM (µM) | |||
|---|---|---|---|---|
| HCT-116 | A-549 | Hela | GI50 | |
| 9e | 8.91 ± 0.06 | 10.23 ± 0.90 | 7.72 ± 0.05 | 8.98 |
| 9g | 9.64 ± 0.07 | 11.40 ± 0.95 | 7.68 ± 0.05 | 9.57 |
| 9i | 7.51 ± 0.05 | 9.72 ± 0.07 | 8.65 ± 0.06 | 8.62 |
| 9k | 7.29 ± 0.05 | 8.01 ± 0.05 | 5.62 ± 0.03 | 6.97 |
| 9m | 15.22 ± 1.05 | 11.98 ± 1.01 | 8.59 ± 0.06 | 11.93 |
| CA-4 | 2.35 ± 0.002 | 1.12 ± 0.001 | 2.01 ± 0.002 | 1.82 |
The outcomes of the in vitro antiproliferative assay corresponded with those of the antitubulin assay. Compounds 9e, 9g, 9i, 9k, and 9m demonstrated significant antiproliferative activity, with GI50 values ranging from 6.97 to 11.93 µM, compared with the reference medication CA-4, which had a GI50 of 1.82 µM across the three cancer cell lines examined. In every instance, the studied compounds were less potent than the reference CA-4 across all cancer cell lines evaluated. Compound 9k (R1 = H, R2 = OMe), the most effective tubulin inhibitor, exhibited the highest antiproliferative activity with a GI50 value of 6.97 µM. It showed a fourfold decrease in potency compared with CA-4 against the cancer cell lines evaluated.
Compound 9k exhibited the highest potency among derivatives against the cervical (HeLa), colorectal (HCT-116), and lung (A-549) cancer cell lines, with IC50 values of 5.62, 7.29, and 8.01 µM, respectively, suggesting that cervical cancer cells demonstrate greater sensitivity to the mechanism of action of 9k compared to colon or lung cancer models. The increased sensitivity in HeLa cells could be attributable to their rapid proliferation rate, as tubulin inhibitors frequently exhibit higher toxicity in more rapidly dividing cells.37
Compounds 9e (R1 = OMe, R2 = H) and 9i (R1 = Me, R2 = Cl) exhibited the third and second-highest antiproliferative activity, with GI50 values of 8.98 and 8.62 µM, respectively. They exhibited approximately a 1.3-fold reduction in potency relative to 9k. The HeLa (cervical) cancer cell line was the most responsive to compound 9e, while the colorectal (HCT-116) cancer cell line showed the highest sensitivity to compound 9i. Compounds 9g (R1 = Cl, R2 = Cl) and 9m (R1 = Me, R2 = OMe) exhibited the lowest antiproliferative efficacy, with GI50 values of 9.57 and 11.93 µM, respectively, signifying modest antiproliferative activity, which correlates with antitubulin activity reflected by IC50 values of 5.33 and 5.74 µM, respectively, Table 1.
Consequently, the capability of compound 9k to function as an activator of Bax, Caspases-3, -8, and -9, as well as a down-regulator of the anti-apoptotic Bcl-2, was examined.
3.2.4.1. Caspase- 3, -8, and -9 activation assay. Compound 9k, the most effective tubulin inhibitor and antiproliferative agent, was evaluated for its activation of caspases-3, -8, and -9 in the HeLa cervical cancer cell line,44 with results presented in Table 3. The findings showed that compound 9k had significantly higher caspase-3 protein levels (587 ± 4 pg mL−1) than the standard compound, staurosporine (510 ± 4 pg mL−1). Compound 9k significantly increased total caspase-3 protein levels in the HeLa cervical cancer cell line, with values nine times higher than those of untreated control cells and above those of the reference drug, staurosporine.
| Compound number | Caspase-3 | Caspase-8 | Caspase-9 | |||
|---|---|---|---|---|---|---|
| Conc (pg mL−1) | Fold change | Conc (ng mL−1) | Fold change | Conc (ng mL−1) | Fold change | |
| 9k | 589 ± 4 | 9 | 1.12 ± 0.10 | 11 | 21.4 ± 1 | 21 |
| Staurosporine | 510 ± 4 | 8 | 1.90 ± 0.10 | 19 | 20 ± 1 | 20 |
| Control | 65 | 1 | 0.10 | 1 | 1 | 1 |
Furthermore, activation testing of caspase-8 and -9 indicated that compound 9k markedly increased their levels compared with staurosporine. Compound 9k demonstrated substantial overexpression of caspase-9 (21.4 ng mL−1, 21-fold increase), followed by caspase-8 (1.12 ng mL−1, 11-fold increase). The data indicate that apoptosis may contribute to the antiproliferative effects of the tested compound, through activation of both intrinsic and extrinsic pathways, with a greater impact on the intrinsic pathway, as evidenced by elevated caspase-9 levels. This suggests that 21-fold activation of Caspase-9 is sufficient to initiate the proteolytic cascade. Although the Caspase-8 route is reduced, the “apoptotic engine” of 9k remains effective, resulting in a higher concentration of active Caspase-3 (589 ± 4 pg mL−1). In summary, even with reduced caspase-8 activation, 9k remains more proficient in achieving the final stage of apoptosis.
3.2.4.2. Assay for levels of Bax and Bcl-2. We also investigated the impact of compound 9k on the apoptotic marker Bax and the anti-apoptotic Bcl-2 levels in the HeLa cancer cell line, using staurosporine as a ref. 44. The results are presented in Table 4.
| Compound number | Bax | Bcl-2 | ||
|---|---|---|---|---|
| Conc (pg mL−1) | Fold change | Conc (ng mL−1) | Fold reduction | |
| 9k | 307 ± 5 | 38 | 1.05 | 5 |
| Staurosporine | 280 ± 7 | 35 | 1.10 | 5 |
| Control | 8 | 1 | 5 | 1 |
The Bax/Bcl-2 ratio is pivotal in dictating cellular outcomes. Compound 9k significantly shifts the equilibrium towards mortality. Bax is a protein that facilitates apoptosis by creating pores in the outer mitochondrial membrane. A 38-fold increase (307 ± 5 pg mL−1) is substantial, exceeding the effect of staurosporine (35-fold). Bcl-2 is an anti-apoptotic protein that generally inhibits Bax activity. Reducing its concentration to 1.05 ng mL−1 (a fivefold reduction) reduces apoptosis inhibition.
These findings elucidate the mechanism underlying the previously observed 21-fold elevation in Caspase-9 levels. The inhibition of tubulin (IC50 = 1.56 µM) induces mitotic stress. The cell responds by elevating Bax levels and reducing Bcl-2 levels. An elevated Bax/Bcl-2 ratio leads to mitochondrial outer membrane permeabilization (MOMP). Upon the leakage of Cytochrome c, caspase-9 is activated, subsequently activating caspase-3.
3.3. In silico studies
 | ||
| Fig. 3 Superimposition of cocrystallized and redocked colchicine in the colchicine binding site of tubulin (RMSD = 0.9278 Å). | ||
Within this validated setup, docking of the most potent derivative 9k into the colchicine binding site (CBS) revealed a binding affinity of −9.1 kcal mol−1, surpassing that of colchicine (−7.8 kcal mol−1) and supporting its superior antiproliferative activity. Compound 9k fits deeply into the hydrophobic pocket at the α/β-tubulin interface and establishes a dense network of favorable contacts with key residues (Fig. 4).
The p-methoxyphenyl–thiadiazole portion forms a carbon–hydrogen bond with Gln136 and hydrophobic interactions with Ile4, Leu252, Leu242, Thr239, and Val238, residues that define the colchicine pocket and contribute critically to ligand affinity. The thiadiazole and thiazole rings further engage Leu238, Leu248, Leu255, and Ala250 through pi–sigma and pi–alkyl contacts, while the thiazole-methyl group interacts with Ala354, Ile318, Ala316, and Cys241 in the deeper hydrophobic groove. Finally, the terminal chalcone benzene ring interacts with Lys352 at the pocket entrance, helping to stabilize the ligand orientation. Altogether, this interaction pattern, involving hallmark residues such as Leu242, Leu248, Leu252, Leu255, Val238, Ala250, Ala316, Ile318, Ala354, Lys352, and Cys241, is highly consistent with the known pharmacophoric features of colchicine binding site inhibitors and provides a clear structural basis for the marked tubulin-targeted cytotoxicity of compound 9k.
To further address the observed SAR trends and clarify the influence of substitution at R1 and R2, additional comparative docking analyses were performed for compounds 9e, 9i, 9m, and 9l within the same colchicine binding site. The obtained binding modes were generally consistent with the experimental antitubulin data and showed that activity was governed not simply by the number of contacts, but rather by the ability of each analogue to adopt a productive orientation and establish well-positioned polar and hydrophobic interactions within the CBS.
Among the newly examined analogues, 9i displayed the most favorable interaction pattern, in agreement with its high tubulin inhibitory activity. It formed a classical hydrogen bond between the thiadiazole nitrogen and Tyr202, in addition to an extended hydrophobic network involving Leu248, Ala250, Leu255, Ala316, Ala354, Ile318, Leu242, Val238, and Thr239. Moreover, the p-chloro substituent on the thiadiazole-linked phenyl ring occupied a lipophilic region and contributed additional favorable contacts with Phe169, Phe20, and Met235, supporting the beneficial effect of the Me/Cl substitution pattern in compound 9i.
Compound 9e also adopted a productive binding mode consistent with its potent activity. A key feature was the formation of a classical hydrogen bond between the chalcone carbonyl and Asn258, while the aromatic scaffold was further stabilized by several hydrophobic interactions with Lys352, Leu248, Leu255, Ala316, Leu242, Leu252, and Thr239. The p-methoxy substituent on the chalcone phenyl ring also contributed favorably through hydrophobic interaction with Lys352, indicating that R1 = OMe is advantageous when the thiadiazole-linked phenyl ring is unsubstituted.
In the case of 9m, which showed moderate activity, the docked pose retained several favorable hydrophobic interactions with Leu248, Ala316, Lys352, Ala354, Val238, Leu242, Leu255, and Leu252, together with additional contacts involving Ser178 and Asn167. However, compared with 9e and 9i, the interaction pattern of 9m appeared less effectively organized around the key anchoring regions of the colchicine pocket, which may account for its lower potency despite maintaining a reasonable binding profile.
By contrast, compound 9l, which showed the weakest antitubulin activity among the docked comparators, adopted a completely different binding pose from the more active analogues 9e, 9i, and 9m within the colchicine binding site. Although it still exhibited some hydrophobic contacts, its markedly altered orientation prevented the ligand from reproducing the key anchoring pattern observed for the more potent compounds. This distinct binding mode likely led to suboptimal occupation of the favorable regions within the colchicine pocket and therefore provides a structural explanation for the sharp decline in activity of 9l. These findings further indicate that the combination R1 = Cl and R2 = OMe is unfavorable for productive colchicine-site recognition. The interaction diagrams of compounds 9e, 9i, 9m, and 9l are provided in the Supplementary Materials.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S = −5.84 vs. −3.83). As shown in the bioavailability radar plots in Fig. 5, compound 9k maintains a generally acceptable bioavailability-related physicochemical profile, with differences from CA-4 mainly reflecting its distinct structural framework. These features suggest that while CA-4 represents a more compact scaffold, compound 9k still resides within a reasonable drug-like physicochemical space.
S = −5.84 vs. −3.83). As shown in the bioavailability radar plots in Fig. 5, compound 9k maintains a generally acceptable bioavailability-related physicochemical profile, with differences from CA-4 mainly reflecting its distinct structural framework. These features suggest that while CA-4 represents a more compact scaffold, compound 9k still resides within a reasonable drug-like physicochemical space.
Despite these structural differences, the predicted membrane permeation descriptors of 9k remained broadly comparable to those of CA-4. Compound 9k showed predicted Caco-2 and MDCK permeabilities of −5.269 and −4.674, respectively, compared with −5.174 and −4.817 for CA-4, while PAMPA prediction was slightly higher for 9k (0.06) than for CA-4 (0.016). These results indicate that although 9k differs from CA-4 in size and polarity, its predicted cellular permeability remains within a comparable borderline-to-moderate range.
A particularly relevant difference emerged in the predicted transporter interaction profile. Compound 9k was predicted to be a non-substrate of P-glycoprotein (P-gp) and to exhibit a relatively high probability of P-gp inhibition (0.844), whereas CA-4 also behaved as a likely non-substrate but showed a lower probability of P-gp inhibition (0.593). Moreover, 9k was predicted to inhibit MRP1 but not BCRP, while CA-4 showed inhibition of both MRP1 and BCRP. This suggests that the transporter profile of 9k may be advantageous in the context of multidrug resistance, since the combination of predicted P-gp non-substrate behavior and stronger P-gp inhibitory potential may help maintain intracellular drug exposure in resistant tumor cells.
The predicted distribution profile also distinguished the two compounds. Compound 9k showed very high plasma protein binding (PPB = 98.768%) and a very low fraction unbound (Fu = 1.082%), whereas CA-4 displayed lower PPB (74.467%) and a substantially higher Fu (19.266%). In addition, 9k had a very low predicted probability of blood–brain barrier penetration (BBB = 0.007) compared with CA-4 (0.457), indicating that 9k may have more restricted central nervous system distribution. Therefore, relative to CA-4, 9k may circulate predominantly in a protein-bound form and display lower free systemic exposure.
The metabolic liability profile of 9k also differed from that of CA-4. Compound 9k was predicted to inhibit multiple CYP isoforms, including CYP1A2, CYP2C19, CYP2C9, CYP2B6, CYP2C8, and CYP3A4, and was also predicted to behave as a CYP2C9 substrate. In contrast, CA-4 showed a narrower CYP inhibition profile, mainly involving CYP1A2, CYP2B6, and CYP2C8, although it was predicted to act as a substrate for CYP2C9, CYP2D6, and CYP3A4. Furthermore, 9k showed a high probability of human liver microsomal instability (HLM stability = 0.841), suggesting possible metabolic vulnerability. These findings indicate that 9k may carry broader metabolism-related liabilities than CA-4 and would likely benefit from future structural optimization.
With respect to excretion, 9k showed lower predicted plasma clearance than CA-4 (2.29 vs. 8.147 mL min−1 kg−1), while both compounds exhibited a short predicted half-life (1.156 vs. 1.452 h). Lower clearance may partially compensate for the high protein binding of 9k, although its overall exposure profile would still require experimental confirmation.
The toxicity predictions further suggested that 9k may have more pronounced liabilities than CA-4 in several categories. In particular, 9k showed markedly higher predicted risk for drug-induced liver injury (0.999 vs. 0.109), genotoxicity (0.999 vs. 0.083), carcinogenicity (0.815 vs. 0.468), and drug-induced nephrotoxicity (0.852 vs. 0.702). On the other hand, some endpoints were less unfavorable for 9k, such as eye irritation and respiratory toxicity. Overall, these results indicate that 9k does not exhibit a superior virtual ADMET profile relative to CA-4. Nevertheless, its distinct transporter-related behavior, especially its predicted P-gp non-substrate character combined with stronger P-gp inhibitory potential, still supports its value as a mechanistically interesting lead scaffold for further optimization rather than as a direct ADMET-improved analogue of CA-4.
3.4. Structure activity relationship (SAR) analysis
The structure–activity relationship (SAR) analysis revealed the following trends:• Substitution at both R1 and R2 strongly affected activity. The most active analogue was 9k (R1 = H, R2 = OMe), indicating that this substitution pattern provides the best overall fit for the series.
• In the R2 = H series, the activity order was 9e (OMe) > 9b (Cl) > 9d (Me) > 9c (F) > 9a (H), showing that R1 = OMe is the most favorable substituent in this subgroup. This agrees with docking study, where 9e showed a productive pose supported by a good balance of polar anchoring and hydrophobic stabilization.
• In the R2 = Cl series, the order became 9i (Me) > 9g (Cl) > 9h (F) > 9j (OMe) > 9f (H), indicating that R1 = Me is optimal when R2 = Cl. Docking of 9i also suggested a highly favorable binding arrangement, which is consistent with its strong tubulin inhibitory activity.
• In the R2 = OMe series, the trend was 9k (H) >> 9m (F) > 9n (Me) > 9o (OMe) > 9l (Cl), showing that an unsubstituted R1 is most favorable in this subgroup. Docking supported this trend, as 9m retained a reasonably favorable orientation, whereas 9l adopted a less productive pose, explaining its marked loss of activity.
Overall, the results show that the effects of R1 and R2 are interdependent rather than additive. The best activity was observed with substitution patterns that allowed a more favorable docking orientation, as seen for 9k, 9i, and 9e, whereas mismatched combinations such as 9l were detrimental despite the presence of multiple substituents.
4. Conclusion
This study presents thiazole-based derivatives 9a–o as a promising category of colchicine-site tubulin inhibitors that demonstrate low-micromolar antiproliferative action while exhibiting minimal toxicity to normal MCF-10A cells at 50 µM. In this series, compound 9k was identified as the most effective analogue, exhibiting significant tubulin inhibition (IC50 = 1.56 µM), enhanced antiproliferative activity against HeLa, HCT-116, and A-549 cancer cell lines, and the ability to activate the intrinsic apoptotic pathway via Bax upregulation, Bcl-2 downregulation, and substantial caspase-9/3 activation. Molecular docking into the colchicine binding site corroborated our findings by demonstrating a favorable binding conformation for 9k, featuring critical interactions typical of high-affinity CBS ligands. The complementary in silico ADMET analysis further validated 9k as a drug-like candidate, demonstrating good oral bioavailability and a favorable P-gp transporter profile aligned with the expected characteristics of tubulin-targeted anticancer medicines. All of these findings highlight 9k as a prospective framework for targeted cancer therapy. Future research should emphasize in vivo efficacy investigations and structural improvements to augment its potency and selectivity against resistant tumor models.Conflicts of interest
The authors declare no competing interests.Data availability
The authors assert that any data supporting this study can be found in the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra01355d.Acknowledgements
The authors acknowledge the support by Princess Nourah bint Abdulrahman University Researchers Supporting Project Number (PNURSP2026R3), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The authors also acknowledge support from the KIT-Publication Fund of the Karlsruhe Institute of Technology.References
- P. Ghankar, A. Bhowate, T. Tidke and A. Jiddewar, Int. J. Sci. R. Tech, 2025, 2, 671–683 Search PubMed
.
- H. C. Assunção, P. M. Silva, H. Bousbaa and H. Cidade, Molecules, 2025, 30, 3314 CrossRef PubMed
.
- T. Fazil, S. C. Suresh, R. Lavoori and K. Natarajan, Comput. Biol. Med., 2026, 204, 111512 CrossRef CAS PubMed
.
- A. R. Nair, N. Saroj and A. Kunwar, Biomolecules, 2026, 16, 81 CrossRef CAS PubMed
.
- M. Ezzo and S. Etienne-Manneville, Int. J. Mol. Sci., 2025, 26, 7652 CrossRef CAS PubMed
.
- Y. Salinas, S. C. Chauhan and D. Bandyopadhyay, Int. J. Mol. Sci., 2025, 26, 3279 CrossRef CAS PubMed
.
- E. Alday, P. Ruiz-Bustos, A. Garibay-Escobar, H. Astiazaran-Garcia, J. M. Sforcin, D. Valencia, Y. Lipovka, Z. Domínguez-Esquivel, C. Virués-Colorado and J. Monribot-Villanueva, J. Pharm. Pharmacol., 2026, 78, rgaf130 CrossRef CAS PubMed
.
- S. Verma, P. Gupta, R. Narang, S. Lal and V. Singh, Future Med. Chem., 2025, 17, 2043–2066 CrossRef CAS PubMed
.
- A. Kamal, P. A. Yakkala, L. Soukya and S. A. Begum, Expert Opin. Drug Discovery, 2025, 20(8), 991–1029 CrossRef CAS PubMed
.
- Y.-S. Hah and S.-Y. Han, Biomol. Ther., 2025, 33, 804 CrossRef CAS PubMed
.
- Y. Lu, J. Chen, M. Xiao, W. Li and D. D. Miller, Pharm. Res., 2012, 29, 2943–2971 CrossRef CAS PubMed
.
- A. Cario and C. L. Berger, Bioessays, 2023, 45, 2200138 CrossRef CAS PubMed
.
- S. Mondal, E. Bonventre, W. O. Hancock and L. M. Rice, Nat. Commun., 2025, 17, 559 CrossRef PubMed
.
- G. Perazzoli, C. Mesas, F. Quiñonero, K. Doello, M. Peña, A. Cepero, J. Rodríguez-Criado, J. Prados and C. Melguizo, Appl. Sci., 2025, 15, 1125 CrossRef CAS
.
- A. Harish, N. Deepika, V. Joshi and P. S. Goudanavar, Invest. New Drugs, 2025, 43, 1086–1108 CrossRef PubMed
.
- J. Dong, M. Sun, K. Cao and T. Fang, Front. Oncol., 2026, 15, 1723016 CrossRef PubMed
.
- N. V. Dubashynskaya, A. N. Bokatyi, M. M. Galagudza and Y. A. Skorik, Int. J. Mol. Sci., 2025, 26, 7591 CrossRef CAS PubMed
.
- A. Tkachenko and O. Havranek, Int. J. Mol. Sci., 2026, 27, 1495 CrossRef PubMed
.
- B. G. Youssif, M. M. Morcoss, S. Bräse, M. Abdel-Aziz, H. M. Abdel-Rahman, D. A. Abou El-Ella and E. S. M. Abdelhafez, Molecules, 2024, 29, 446 CrossRef CAS PubMed
.
- V. Eisner, M. Picard and G. Hajnóczky, Nat. Cell Biol., 2018, 20, 755–765 CrossRef CAS PubMed
.
- S. H. Dho, M. Cho, W. Woo, S. Jeong and L. K. Kim, Exp. Mol. Med., 2025, 1–12 Search PubMed
.
- G. T. Almutairi, A. G. Almutairi, F. A. N. Alotibi, F. A. Aljohani, B. Alkhalifah, A. A. Alshehri, M. M. Aljohani, A. H. Al Shamary, A. M. O. Albaqami and A. H. Almutairi, Egypt. J. Chem., 2026, 69, 179–189 Search PubMed
.
- H. Hashem, A. Hassan, W. M. Abdelmagid, A. G. Habib, M. A. Abdel-Aal, A. M. Elshamsy, A. El Zawily, I. T. Radwan, S. Bräse and A. S. Abdel-Samea, Pharmaceuticals, 2024, 17, 1154 CrossRef CAS PubMed
.
- H. Rajak, P. Kumar Dewangan, V. Patel, D. Kumar Jain, A. Singh, R. Veerasamy, P. Chander Sharma and A. Dixit, Curr. Pharm. Des., 2013, 19, 1923–1955 CrossRef CAS PubMed
.
- H. E. N. Khasawneh, E. I. Ali, R. M. Elmagzoub, R. F. A. Al-Aouadi, W. T. Almagharbeh, G. Alotaibi, S. Bräse and A. Alkhammash, Front. Chem., 2025, 13, 1606848 CrossRef CAS PubMed
.
- L. H. Al-Wahaibi, A. M. Elshamsy, T. F. Ali, B. G. Youssif, S. Bräse, M. Abdel-Aziz and N. A. El-Koussi, Front. Chem., 2025, 13, 1565699 CrossRef CAS PubMed
.
- M. S. Abdelbaset, M. H. Abdelrahman, S. N. A. Bukhari, A. M. Gouda, B. G. Youssif, M. Abdel-Aziz and G. E.-D. A. Abuo-Rahma, Bioorg. Chem., 2021, 107, 104522 CrossRef CAS PubMed
.
- M. S. Abdelbaset, G. E.-D. A. Abuo-Rahma, M. H. Abdelrahman, M. Ramadan, B. G. Youssif, S. N. A. Bukhari, M. F. Mohamed and M. Abdel-Aziz, Bioorg. Chem., 2018, 80, 151–163 CrossRef CAS PubMed
.
- E. Mahmoud, D. Abdelhamid, A. F. Mohammed, Z. M. Almarhoon, S. Bräse, B. G. Youssif, A. M. Hayallah and M. Abdel-Aziz, Pharmaceuticals, 2025, 18, 275 CrossRef CAS PubMed
.
- M. F. Abo-Ashour, W. M. Eldehna, R. F. George, M. M. Abdel-Aziz, M. M. Elaasser, N. M. A. Gawad, A. Gupta, S. Bhakta and S. M. Abou-Seri, Eur. J. Med. Chem., 2018, 160, 49–60 CrossRef CAS PubMed
.
- T. S. Kokovina, S. Y. Gadomsky, A. A. Terentiev and N. A. Sanina, Molecules, 2021, 26, 5159 CrossRef CAS PubMed
.
- L. H. Al-Wahaibi, Y. A. Mostafa, M. H. Abdelrahman, A. H. El-Bahrawy, L. Trembleau and B. G. Youssif, Pharmaceuticals, 2022, 15, 1006 CrossRef CAS PubMed
.
- M. Ramadan, M. Abd El-Aziz, Y. A. Elshaier, B. G. Youssif, A. B. Brown, H. M. Fathy and A. A. Aly, Bioorg. Chem., 2020, 105, 104392 CrossRef CAS PubMed
.
- F. F. Hagar, S. H. Abbas, H. A. Gomaa, B. G. Youssif, A. M. Sayed, D. Abdelhamid and M. Abdel-Aziz, BMC Chem., 2023, 17, 116 CrossRef CAS PubMed
.
- L. H. Al-Wahaibi, H. A. Abou-Zied, M. Hisham, E. A. Beshr, B. G. Youssif, S. Bräse, A. M. Hayallah and M. Abdel-Aziz, Molecules, 2023, 28, 6586 CrossRef CAS PubMed
.
- L. H. Al-Wahaibi, A. M. Elshamsy, T. F. Ali, B. G. Youssif, S. Bräse, M. Abdel-Aziz and N. A. El-Koussi, ACS Omega, 2024, 9, 34358–34369 CrossRef CAS PubMed
.
- D. Raghav, S. M. Ashraf, L. Mohan and K. Rathinasamy, Biochemistry, 2017, 56, 2594–2611 CrossRef CAS PubMed
.
- D. Fernández-Lázaro, B. Sanz and J. Seco-Calvo, Proteomes, 2024, 12, 3 CrossRef PubMed
.
- C. Cenciarelli, Curr. Med. Chem., 2009, 16(11), 1315–1324 CrossRef PubMed
.
- H. Gali-Muhtasib and N. Bakkar, Curr. Cancer Drug Targets, 2002, 2, 309–336 CrossRef CAS PubMed
.
- S. W. Tait and D. R. Green, Nat. Rev. Mol. Cell Biol., 2010, 11, 621–632 CrossRef CAS PubMed
.
- Z. Z. Chong, J.-Q. Kang and K. Maiese, J. Cereb. Blood Flow Metab., 2003, 23, 320–330 CrossRef CAS PubMed
.
- K. Cain, S. B. Bratton and G. M. Cohen, Biochimie, 2002, 84, 203–214 CrossRef CAS PubMed
.
- B. G. Youssif, A. M. Mohamed, E. E. A. Osman, O. F. Abou-Ghadir, D. H. Elnaggar, M. H. Abdelrahman, L. Treamblu and H. A. Gomaa, Eur. J. Med. Chem., 2019, 177, 1–11 CrossRef CAS PubMed
.
- N. B. Ayrim, A. A. Balakit, S. J. Laftaa, F. F. Alkazazz, Y. Bekkar, L. Bourougaa and B. A. Saleh, ChemistrySelect, 2025, 10, e202405188 CrossRef CAS
.
- A. O. H. Zayed, Discover Chemistry, 2025, 2, 164 CrossRef
.
- M. Suchecki, M. D. Amico, J. Herkt, G. Vergilio, F. Rappa, M. Gorska-Ponikowska and A. Kuban-Jankowska, Biomed. Pharmacother., 2026, 196, 119006 CrossRef CAS PubMed
.
- M. Bağlan, K. Gören, V. Tahiroğlu, G. Kotan, S. Manap and H. Yüksek, ChemistrySelect, 2026, 11, e06617 CrossRef
.
| This journal is © The Royal Society of Chemistry 2026 |