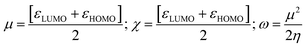Open Access Article
Open Access ArticleToxicological investigation of mycotoxin contaminants and antibiotic residues associated with the poultry industry and their impact on human health
Jahida Akterab,
Kabir Hossainac,
Santanu Deb Nathad,
Hozzatul Islamac,
Khaleda Afrinae,
Hamida Begumaf and
Monir Uzzaman *ae
*ae
aDepartment of Toxicology Research, Computer in Chemistry and Medicine Laboratory, Dhaka, Bangladesh. E-mail: monircu92@gmail.com
bDepartment of Textile Engineering, Port City International University, Chittagong, Bangladesh
cFaculty of Science, Department of Applied Chemistry and Chemical Engineering, University of Chittagong, Chittagong, 4331, Bangladesh
dFaculty of Biological Sciences, Department of Biochemistry and Molecular Biology, Noakhali Science and Technology University, Noakhali, Chittagong, 3814, Bangladesh
eFaculty of Science, Department of Chemistry, University of Chittagong, Chittagong, 4331, Bangladesh
fNational Drug Control Laboratory, Dhaka, Bangladesh
First published on 5th May 2026
Abstract
Contamination of feed with mycotoxins and the intensive use of therapeutic antibiotics in modern poultry farming have resulted in the accumulation of chemical residues in meat and the environment. These residues may contribute to hepatic, renal, and cardiac dysfunctions, leading to significant public-health concerns. Despite growing awareness and extensive studies on individual poultry contaminants, comprehensive molecular-level assessments of all our selected compounds are still limited. Integrated computational analyses combining density functional theory (DFT), molecular docking, molecular dynamics (MD) simulation, absorption, distribution, metabolism, excretion, and toxicity (ADMET), and prediction of activity spectra for substances (PASS) evaluations to disclose their multi-organ toxic mechanisms also remain scarce. This study was designed to evaluate the hepatic, renal, and cardiac toxicity of selected mycotoxins and therapeutic antibiotics through a computational framework. Molecular geometries and electronic properties were assessed using DFT calculations. As per DFT results, ZER, CIP, ENR, and OTA showed better thermodynamic stability due to higher HOMO–LUMO energy gap; and exhibited higher dipole moments in AB1 (8.26 Debye), CIP (9.70 Debye), ENR (9.80 Debye), and OXY (6.89 Debye). The lower HOMO–LUMO energy gap indicates that these compounds have higher chemical reactivity. Molecular docking result revealed strong binding affinities (ranging from −7.5 kcal mol−1 to −9.4 kcal mol−1) within selected ligands and proteins, while MD simulation confirmed stable protein–ligand interactions. Additionally, ADMET and PASS prediction elucidated the potential nephrotoxicity (ENR +0.37, OXY +0.87, and OTA +0.72); hepatotoxicity (ENR +0.94, OXY +0.61, and OTA +0.49); and myocarditis (ENR +0.28, OXY +0.92, and OTA +0.35). Overall, our findings highlighted the potential health risks posed by mycotoxin residues and therapeutic antibiotics used in poultry farms.
1. Introduction
The poultry industry is one of the rapidly expanding sectors in livestock production, with its contribution to the production of proteins, vitamins, and minerals. Over 70% of Bangladesh's animal protein supply comes from this industry, which has grown quickly over the past few decades.1 However, feed additives, antibiotics, and growth promoters are frequently used to maintain poultry health and productivity. Mycotoxin contamination in poultry feed and antibiotic residues in poultry products are notable problems that have arisen in recent years.2 Mycotoxins are produced naturally by Aspergillus flavus and Aspergillus parasiticus found in common food and poultry feedstuffs such as rice, peanuts, corn, nuts, spices, fruits, fishmeal, and soybean meal products.3 Mycotoxins often enter animal feed through fungal contamination, which eventually leads to human exposure. Due to climate change, the occurrence of mycotoxin contamination in poultry feed is expected to increase. As per previous findings, the increase in temperature may favor the growth of thermotolerant fungi such as Aspergillus parasiticus, which could have negatively interfered with crop production or storage.4 Currently, the allowable level of mycotoxins in poultry products is not regulated by any particular guidelines.5Depending on the administered dose and period of exposure, mycotoxins, including Aflatoxin B1 (AB1), Ochratoxin A (OTA), and Zearalenone (ZER), can cause either long-term or short-term illnesses such as carcinogenic, nephrotoxic, hemorrhagic, teratogenic, hemorrhage, nausea, and inflammation in human health.6 Additionally, in poultry farms, various antibiotics, including Oxytetracycline (OXY), Ciprofloxacin (CIP), and Enrofloxacin (ENR), are intentionally administered via feed or intramuscular injection at sub-therapeutic doses to promote growth, prevent diseases, and treat the sick animals.7 Chemical structure of all compounds illustrated in Fig. 1. Although the poultry sector in Bangladesh represents an important regional example where veterinary antibiotics are frequently used to prevent disease and promote growth, similar practices have been widely reported in intensive livestock production systems worldwide. About half of all antibiotics produced worldwide are used in livestock.8 High concentrations of antibiotic residues and resistant bacterial isolates have been found in commercial broiler meat in Bangladesh.9 Overexposure of antibiotics has contributed to the development of multidrug-resistant (MDR) bacteria in poultry farms.10 The presence of antibiotic residues and MDR pathogens in poultry meat poses a severe health threat, such as gastrointestinal disorders, allergic reactions, obesity, carcinogenicity, reproductive effects, and teratogenicity, by entering the human food chain through slaughtering, processing, and consumption.11 Furthermore, recent research has found that exposure to low quantities of antibiotics is associated with human health complications.12 Beyond direct consumption, poultry-origin resistant bacteria can also spread through environmental pathways such as polluted litter, wastewater, and agricultural runoff. It should be noted that antibiotic residues may be present in both poultry and plant-based foods, as crops can become contaminated through polluted soil and water.13 Without targeted interventions, antibiotic-resistant bacteria originating from poultry production will continue to pose a major threat to national and global public health security. Several international regulatory agencies, such as the European Chemicals Agency (ECHA), U.S. Environmental Protection Agency (EPA), and U.S. Food and Drug Administration (FDA), have designed monitoring frameworks and risk-assessment programs to reduce mycotoxin contamination in agricultural and veterinary systems. For instance, the FDA sets action levels and advisory limits for these mycotoxins in food and feed to protect both human and animal health in accordance with the Preventive Controls for Animal Food (PCAF) rule. The ECHA's itself focuses on chemical hazards under REACH (Registration, Evaluation, Authorization and Restriction of Chemicals). The EPA explores the toxicological and environmental concerns of fungal metabolites and other chemical pollutants that may exist in agricultural environments. Although the EPA does not set direct limits for mycotoxins in food or feed, its risk-assessment frameworks, such as the Integrated Risk Information System (IRIS) support the evaluation of environmental toxicants and chemical mixtures. Furthermore, mycotoxin contamination has also been detected in commercial pet foods, suggesting that companion animals may also be exposed to these contaminants through cereal-based diets.
Many experimental methods have been conducted that are unable to explain the molecular-level mechanism behind toxicity. Additionally, it is costly, challenging to use, and takes time.14 The in silico method predicts several kinds of molecular properties of chemical frameworks and also provides a comprehensive study that helps to explain or interpret trends and structure–activity relationships.15 This computational approach helps to predict the characteristics of targeted chemical compounds within a short time. Recent computational studies have successfully predicted toxicity16 and the binding mechanisms of various xenobiotics.17 Most of them were done for single molecules, while some studies were for combined molecules.18 But integrated multiparameter computational evaluation time-dependent density functional theory (TD-DFT), molecular docking, MD simulation, ADMET, and PASS associated with poultry residues and antibiotics for our selected six molecules has not been done yet. This study will help to disclose the toxicological effects of our selected compounds on human health.
To achieve physicochemical and chemical characteristics, an in silico strategy combining DFT, molecular docking, MD, ADMET, and PASS prediction was employed. DFT calculations were employed to assess molecular stability, reactivity, and key electronic descriptors, including frontier molecular orbitals and thermodynamic parameters. Following each part, new toxicological risk evaluations and human health protection are made possible by this computational framework.19 In order to avoid and mitigate the detrimental effects of these compounds, global and international strategies should be taken against mycotoxins, therapeutic antibiotics, and hormonal residues. Diagnostic methods and food quality control measures should be accelerated. Additionally, new maximum permissible limits for these compounds should be established in food and poultry feed for various animal species. Consequently, these options should be congruent and synchronized with the current standards.20 Probiotics, prebiotics, and efficient feed additives should be used more frequently as antibiotic alternatives, which can stop the occurrence of mycotoxins, therapeutic antibiotics, and hormonal residues.21 Control and testing of toxic compound systems are also important to prevent exposure to human and animal health. Most previous studies worked on a single contaminant; our study simultaneously focuses on multiple toxic compounds to better reflect real-world co-occurrence and enables a more realistic assessment of their potential interactions and combined risks in food and feed systems.
2. Materials and methods
2.1. Computational details
The structure of all chemical compounds was collected from the PubChem online chemical structure database. These structures were prepared by GaussView software (version 6.0.16) and quantum chemical calculations, including DFT, Becke, 3-parameter, Lee-Yang-Parr (B3LYP), were done by Gaussian 16 using 6-31G (d,p) basis set.22 The UV-vis absorption spectrum was also predicted using TD-DFT. Furthermore, the Parr and Pearson interpretations of DFT23 and Koopman's theorem was utilized to calculate frontier molecular orbital features, including the highest occupied molecular orbital (HOMO) and the lowest unoccupied molecular orbital (LUMO) energy gap (ΔE), chemical hardness (η), chemical softness (S), chemical potential (µ), electronegativity (χ), and electrophilicity (ω) by the following equations.24Additionally, different convergence parameters such as maximum force, RMS force, maximum displacement, and RMS displacement are also considered in this study. These parameters satisfied the default convergence thresholds limit of 23.14 meV Å−1 for maximum force, 15.43 meV Å−1 for RMS force, 0.95 mÅ for maximum displacement, and 0.63 mÅ for RMS displacement. Therefore, the geometry optimization of our selected compounds was considered converged.
2.2. Ligand selection and preparation
Based on previous reports, the ligands were selected as toxic contaminants in poultry farms and feed formulations. These selected compounds represent different classes, such as mycotoxins and therapeutic antibiotics, which are commonly detected in poultry production systems. AB1,25 OTA,26 and ZER27 were selected as mycotoxins. In addition, OXY,28 CIP,29 and ENR30 were selected as therapeutic antibiotics. All of these compounds were collected in SDF format from the PubChem database and transformed into MOL format using GaussView software. To minimize steric clashes and obtain the most stable molecular conformations, geometry optimization of all compounds was performed using the Gaussian 16 software. Furthermore, molecular docking was done by using PyRx software (version 0.8).312.3. Protein selection and preparation
The target protein selection was performed to identify biologically relevant receptors associated with the toxic effects of the studied compounds. As per ADMET and PASS prediction results, our selected compounds exhibited strong associations with hepatotoxicity,32 nephrotoxicity,33 and cardiac toxicity.34 Based on these findings, an extensive literature review was conducted to identify the key hub genes associated with these diseases. Specific protein structures corresponding to the identified hub genes were collected from the Protein Data Bank (PDB). The selected proteins were 2RJQ for hepatocellular carcinoma, 3P0M for chronic kidney disease, and 4G1C for heart failure. These selected proteins ensured that the absence of mutations or structural abnormalities, almost zero Ramachandran outliers, and were experimentally determined using the X-ray diffraction method. Collected proteins were further prepared by the Discovery Studio Visualizer 2021 software (version 21.1.0.20298) by removing excess water molecules, co-crystallized ligands, heteroatoms, and unexpected chains. Therefore, chain A, active site, and protein group were selected to ensure a stable protein structure, and energy minimization was carried out using Swiss-PDB Viewer (version 4.1.0).2.4. Active site selection
The active site of the protein was identified using a structure-based pocket prediction method. P2Rank (version 2.5) server was utilized to analyze the protein surface topology to detect possible ligand-binding pockets.35 The three-dimensional structures of three selected proteins were uploaded into the binding site prediction tool. The identified active site region was visualized in red on the protein surface, representing the most probable ligand-binding cavity. Among the predicted sites, the top-ranked pocket (rank 1) was selected based on its highest score and probability value, indicating a strong and reliable active site of the protein.36 The pocket center was −43.77, −21.13, 6.23 for 2RJQ, 18.51, 7.60, 22.71 for 4 G1C, and 23.69, 10.02, −0.65 for 3P0M protein.2.5. Molecular docking and non-bonding interaction
Flexible docking was employed by using the PyRx virtual screening tool (version 0.8) and the AutoDock wizard37 of the selected compounds against the receptor proteins, where selected proteins were regarded as macromolecules and compounds as ligands. The grid box dimensions were set to 59.27, 48.51, and 52.12 Å for 2RJQ, 46.07, 48.64, and 56.15 Å for 4G1C, and 50.41, 57.76, and 48.70 Å for 3P0M, along the X, Y, and Z axes, respectively, to cover the entire protein. Ligand and protein complex preparation were performed by PyMOL software (version 2.5.7). At the end, nonbonding interaction results were analyzed by Discovery Studio Visualizer 2021.2.6. Docking validation
Molecular docking was performed on three different computer systems to ensure reproducibility and consistency of the docking scores. The obtained scores were averaged, and the relative standard deviation (RSD) values were calculated and presented as error bars in the graphical analysis. A Ramachandran plot was generated by using a structure validation server, which is PROCHECK UCLA SAVES (version 6.1), to validate the stereochemical structure of proteins. It provided a graphical representation including allowed and disallowed regions with dihedral (φ and ψ) angles of amino acid residues. This analysis helped to evaluate the quality of the protein-ligand complexes.382.7. Molecular dynamics simulation
Our two promising compounds were evaluated on MD simulation to evaluate the stability of protein-ligand complexes. This is a widely accepted approach, which also helps to validate the molecular docking result. Our current study utilized the AMBER 14 force field available in YASARA dynamics software (v 23.9.29). Therefore, a 100 ns MD simulation was performed with a time step of 2.50 fs, and trajectories were saved every 250 ps at constant pressure.39 The primary physiological requirements were set at pH 7.4, 0.9% NaCl, the simulation box size (10.0 Å), and 298 K for the simulation cell. The Berendsen thermostat was employed to regulate these parameters.40 Water molecules at 0.997 g cm−3 were added to neutralize the system. Therefore, the key MD simulation parameters analyzed included hydrogen-bonding interactions within the binding region, root mean square deviation (RMSD), root mean square fluctuation (RMSF), solvent-accessible surface area (SASA), and radius of gyration (Rg).2.8. ADMET and PASS prediction
ADMET prediction plays a significant role in the design and development of novel drugs. Therefore, computational and medicinal chemistry methods have become an initial choice for ADMET prediction over recent decades. In this study, ADMET properties of selected compounds were studied using the ADMET SAR 3.0 online server. PASS prediction was obtained from the PASS online server, which provided probabilities of biological activity (Pa) and inactivity (Pi) for each compound, calculated based on their structural features. Therefore, Pa > 0.7, suggesting a significant potential for activity that is considered a default threshold.412.9. pIC50 value prediction
By using a web-based platform called ChemDes, different molecular fingerprints and descriptors have been designed for quantitative representations of physical, chemical, or functional characteristics of studied compounds in quantitative structure–activity relationship (QSAR) analysis.423. Results and discussion
3.1. Thermodynamic and convergence analysis
Thermodynamic metrics such as Gibbs free energy and internal energy can give information about the spontaneity of a reaction and the stability of the products.43 These variables determined the amount of energy absorbed or released during a chemical reaction. Herein, OTA showed the highest free energy, −1739.06 Hartree, in the presence of electronegative atoms (Cl, O). The free energy values became progressively more negative, with values of ZER (−1075.02 Hartree), AB1 (−1106.15 Hartree), CIP (−1148.12 Hartree), ENR (−1226.70 Hartree), OXY (−1638.93 Hartree), suggesting energetically and configurationally more stable. The free energy, enthalpy, and dipole moments are presented in Fig. 2(a) and (b) and Table S1. The dipole moments are crucial for determining the electrical properties of molecules. Improved dipole moments can enhance polarity, binding affinity, and non-bonding interactions of the drug–protein complex.44 In our analysis, ENR (9.80 Debye) and CIP (9.70 Debye) exhibited higher dipole moments than other compounds, due to the presence of more electronegative (F, Cl, and O) atoms, which enhanced electron-withdrawing effects within the molecules. Conversely, OXY (6.89 Debye) and OTA (5.41 Debye) exhibited moderate dipole moments, indicating less charge separation within these compounds.Regarding convergence parameters, the calculated maximum force (in meV Å−1) of optimized structures were 0.72 (ZER), 0.62 (AB1), 1.13 (CIP), 0.36 (ENR), 2.32 (OXY), and 0.87 (OTA). The RMS force (in meV Å−1) values were 0.21, 0.10, 0.21, 0.10, 0.46, and 0.15, respectively. The maximum displacement (in mÅ) values were 0.30 (ZER), 0.49 (AB1), 0.52 (CIP), 0.64 (ENR), 0.88 (OXY), 0.89 (OTA) and they are respectively followed by this ordering up to conditions of peak displacement as shown in Table S3. The corresponding RMS displacement (in mÅ) values were 0.08, 0.09, 0.12, 0.11, 0.17, and 0.21, respectively. The geometry optimizations for all six compounds were successfully converged, and stable stationary points were obtained. All of the computed parameter values were much smaller than their respective threshold limit. Therefore, our optimized geometries are trustworthy for quantum chemical analyses.
3.2. Frontier molecular orbital analysis
Frontier molecular orbitals are key determinants of molecular properties, stability, reactivity, and energy states of a compound involved in chemical processes. The energy gap between HOMO and LUMO is essential to examine the reactivity and electronic transitions of the compounds.45 The energy of the HOMO determines the nucleophilicity, indicating how easily a molecule can donate electrons, while the energy of the LUMO determines the electrophilicity, indicating how easily a molecule can accept electrons.46 A smaller HOMO–LUMO gap indicates greater electron donation and acceptance ability, suggesting higher reactivity and lower stability of the molecule. Conversely, a large gap exhibits more stability and less reactivity. The HOMO–LUMO gap serves as an indicator of molecular hardness and softness.47 In Fig. 3(a) and Table S2, ZER showed the highest energy gap (4.63 eV), while OXY and AB1 showed the lowest energy gap (4.10 eV). In ZER, the relatively high HOMO–LUMO energy gap indicated that electronic excitation required more energy, implying greater molecular stability and lower chemical reactivity. Our study exhibited that the HOMO–LUMO gap was influenced by the presence of various functional groups. Consequently, electron-donating groups increased the HOMO energy by adding electron density, while electron-withdrawing groups attracted electrons and lowered the LUMO energy. Additionally, differences in bond strength and electron delocalization also influenced the energy gap.48 When a π bond was adjacent to another π bond, the electrons could spread over multiple bonds, allowing easy movement through the molecules. As a result, the HOMO–LUMO gap decreased. Conversely, disruption of conjugation reduced delocalization and increased the gap. AB1 included a furan ring, a lactone group (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O–O), and a conjugated double bond. The π conjugation in this group allowed extensive electron delocalization. The electron-withdrawing carbonyl group reduced the LUMO, causing a small gap (4.10 eV) in AB1. ZER involved a hydroxyl (OH) group, conjugated carbonyl (C
O–O), and a conjugated double bond. The π conjugation in this group allowed extensive electron delocalization. The electron-withdrawing carbonyl group reduced the LUMO, causing a small gap (4.10 eV) in AB1. ZER involved a hydroxyl (OH) group, conjugated carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), and sp3 carbon linkage. This limited conjugation hindered effective electron delocalization. As a result, the HOMO and the LUMO were not significantly increased and decreased, respectively, suggesting a larger gap (4.63 eV). ZER also exhibited the highest chemical hardness (2.32 eV), indicating higher chemical stability and resistance to charge transfer. However, the lowest softness value (0.22 eV−1) remained constant for ZER, CIP, ENR, and OTA. Although the hardness values varied among the compounds, they shared a similar tendency for electronic polarization and charge redistribution.49 In contrast, AB1 and OXY exhibited the highest chemical softness (0.24 eV−1) and the lowest hardness (2.05 eV), presented in Fig. 3(b) and Table S2. Therefore, their chemical structure allowed them to accept or donate electrons easily, and both of them exhibited high reactivity and were chemically less stable.
O), and sp3 carbon linkage. This limited conjugation hindered effective electron delocalization. As a result, the HOMO and the LUMO were not significantly increased and decreased, respectively, suggesting a larger gap (4.63 eV). ZER also exhibited the highest chemical hardness (2.32 eV), indicating higher chemical stability and resistance to charge transfer. However, the lowest softness value (0.22 eV−1) remained constant for ZER, CIP, ENR, and OTA. Although the hardness values varied among the compounds, they shared a similar tendency for electronic polarization and charge redistribution.49 In contrast, AB1 and OXY exhibited the highest chemical softness (0.24 eV−1) and the lowest hardness (2.05 eV), presented in Fig. 3(b) and Table S2. Therefore, their chemical structure allowed them to accept or donate electrons easily, and both of them exhibited high reactivity and were chemically less stable.
Our study also revealed that the electron cloud of the HOMO in all studied compounds was primarily localized over the π electron-rich aromatic ring and heteroatom-containing donor groups, such as hydroxyl (OH), primary amine (NH2), and secondary amine (NH). The electron cloud of the LUMO was distributed over the carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) groups and other oxygen or halogen-rich electron-withdrawing groups.50 During HOMO–LUMO excitation, this intramolecular charge was transferred from the donor region to the acceptor region due to the higher electronegativity of the acceptor groups and the effective π conjugation within the delocalized π system,51 as illustrated in Fig. S1. For AB1, the HOMO electron cloud was mainly localized over the aromatic ring and hydroxyl (OH) substituted π system. The LUMO electron cloud was concentrated over the carbonyl and oxygen-rich groups, indicating charge transfer from aromatic donor to carbonyl acceptor. In OXY, the electron cloud of the HOMO was mainly concentrated over the aromatic ring and amine-containing donor region, which was localized over the carbonyl and adjacent heteroatom-rich sites in LUMO. The electron cloud was shifted from the aromatic ring and nitrogen-containing heterocyclic ring to the carbonyl group and electronegative substituent in CIP, which showed donor–acceptor charge migration. Additionally, in ZER, ENR, and OTA, the electron cloud was transferred from the phenolic and heterocyclic donor region to the carbonyl-centered and oxygen-rich electron-withdrawing groups. This electron redistribution reflected that intramolecular charge transfer occurred due to the difference in electronegativity between various functional groups.
O) groups and other oxygen or halogen-rich electron-withdrawing groups.50 During HOMO–LUMO excitation, this intramolecular charge was transferred from the donor region to the acceptor region due to the higher electronegativity of the acceptor groups and the effective π conjugation within the delocalized π system,51 as illustrated in Fig. S1. For AB1, the HOMO electron cloud was mainly localized over the aromatic ring and hydroxyl (OH) substituted π system. The LUMO electron cloud was concentrated over the carbonyl and oxygen-rich groups, indicating charge transfer from aromatic donor to carbonyl acceptor. In OXY, the electron cloud of the HOMO was mainly concentrated over the aromatic ring and amine-containing donor region, which was localized over the carbonyl and adjacent heteroatom-rich sites in LUMO. The electron cloud was shifted from the aromatic ring and nitrogen-containing heterocyclic ring to the carbonyl group and electronegative substituent in CIP, which showed donor–acceptor charge migration. Additionally, in ZER, ENR, and OTA, the electron cloud was transferred from the phenolic and heterocyclic donor region to the carbonyl-centered and oxygen-rich electron-withdrawing groups. This electron redistribution reflected that intramolecular charge transfer occurred due to the difference in electronegativity between various functional groups.
Fig. 3(c) illustrates the DOS profile of AB1, ZER, and CIP, showing the distribution of occupied and unoccupied electronic states alongside the energy gaps. The HOMO states were distributed in the energy range of −6 eV to −5 eV, while the LUMO states appeared between −3 eV and −2 eV. A clear energy gap between the occupied and unoccupied states was observed. In the lower energy region (around −6 eV to −7 eV), DOS peaks arose from bonding molecular orbitals, reflecting enhanced molecular stability. The higher energy region (approximately above +2 eV) indicated available unoccupied states that affected the electronic activity and intermolecular interactions. The variation in DOS peaks among these three compounds originated from differences in heteroatom contents, functional groups, and π conjugation patterns that altered electronic state distribution and orbital energies. Furthermore, ENR, OXY, and OTA are shown in Fig. S2. The HOMO states of these compounds appeared in the range of −6 eV to −5 eV. The LUMO states were distributed between −3 eV and −1 eV approximately. The DOS peak observed in the lower energy region (−6 eV to −7 eV) indicates bonding molecular orbitals, while the peak in the higher energy region (approximately above +1 eV) corresponds to unoccupied states. Minor differences in the DOS intensity and peaks in the figure reflected the differences within the electronic structure of ENR, OXY, and OTA.
3.3. Electrostatic potential analysis
The electrostatic potential surface mapping is a three-dimensional tool that visualizes the distribution of molecular electron density and helps to identify regions susceptible to electrophilic or nucleophilic interactions.52 In the electrostatic potential surface, the red zone indicates a high electron region favourable for electrophilic attack, the blue zone indicates an electron-deficient region favourable for nucleophilic attack, and the green region indicates neutral regions.53 As observed in the electrostatic potential surface maps shown in Fig. 4, oxygen-containing regions showed strong negative potentials visualized as a red zone. Hydrogen, nitrogen, and fluorine-containing sites were observed as a blue region, indicating an electron-deficient and positively charged area. Among all the compounds, CIP showed the most negative potential value (−9.94 × 10−2 Hartree). This is due to the existence of specific functional groups such as the carboxylic group (COOH), carbonyl group (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), fluorine atom, and a piperazine ring. Among these, the carbonyl group (C
O), fluorine atom, and a piperazine ring. Among these, the carbonyl group (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) was highly electron-withdrawing, which showed a strong negative potential region. The carboxyl group (COOH) contained two oxygen molecules, one in a double-bonded (C
O) was highly electron-withdrawing, which showed a strong negative potential region. The carboxyl group (COOH) contained two oxygen molecules, one in a double-bonded (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) and another in a hydroxyl (OH) functional group as hydrogen bond acceptors and retained electron density, contributing to the overall negative potential region. Therefore, CIP exhibited a more significant electrophilic attack than other compounds. CIP also showed a higher positive value (+9.94 × 10−2 Hartree) than other compounds due to the presence of basic piperazine nitrogen (N–(CH2)2–N). As a result, its charge distribution is highly polarized, resulting in high charge separation across the molecule.
O) and another in a hydroxyl (OH) functional group as hydrogen bond acceptors and retained electron density, contributing to the overall negative potential region. Therefore, CIP exhibited a more significant electrophilic attack than other compounds. CIP also showed a higher positive value (+9.94 × 10−2 Hartree) than other compounds due to the presence of basic piperazine nitrogen (N–(CH2)2–N). As a result, its charge distribution is highly polarized, resulting in high charge separation across the molecule.
 | ||
| Fig. 4 Molecular electrostatic potential map of selected compounds (remaining are presented in Fig. S5). | ||
3.4. FT-IR analysis
FT-IR spectral analysis is a widely employed technique for identifying different functional groups present in chemical compounds.39 Here, the vibrational frequencies of ZER, AB1, CIP, ENR, OXY, and OTA are calculated from 400 cm−1 to 4000 cm−1 and scaled by 0.9627 for improved accuracy (reference for scaling https://cccbdb.nist.gov/vsfx.asp). The calculated frequencies are presented in Fig. 5(a) and Table S4. Except for AB1, in the other studied compounds, O–H bond stretching was found at frequencies ranging from 3133 cm−1 to 3673 cm−1, which corresponds to experimental values ranged 3450–3500 cm−1 (CIP), 3300–3500 cm−1 (ENR), and 3380 cm−1 (OTA). All compounds showed aliphatic and aromatic C–H stretching vibrations in the region 2810–3142 cm−1; experimental peaks were recorded in 2853–2925 cm−1 (AB1), 2950–3000 cm−1 (CIP), and 2988 cm−1 (OTA). C–N bond stretching associated with aryl groups was displayed at 1242–1329 cm−1 in ENR, CIP, OXY, and OTA compounds, with OXY and OTA showing experimental peaks at 1218 cm−1 and 1140 cm−1, respectively. Each compound showed absorption peaks at 1669–1810 cm−1 due to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond stretching from different carbonyl groups such as ketone, amide, acid, or lactone, experimentally these were observed at 1715–1720 cm−1 for ZER, 1706 cm−1 for AB1, 1700–1750 cm−1 for CIP, 1737 cm−1 for ENR, 1621 cm−1 for OXY, and 1723 cm−1 for OTA. The N–H bond stretching vibration of amide functional group was seen at 3377 cm−1 and 3509 cm−1 in OXY and OTA, respectively. In contrast, CIP revealed N–H stretching bands at 3397 cm−1, indicating the presence of a secondary amine group. The C–F bond was observed at 1237 and 1240 cm−1 in ENR and CIP, respectively, while the C–Cl bond was detected at 713 cm−1 in OTA, confirming the presence of halogen functional group.
O bond stretching from different carbonyl groups such as ketone, amide, acid, or lactone, experimentally these were observed at 1715–1720 cm−1 for ZER, 1706 cm−1 for AB1, 1700–1750 cm−1 for CIP, 1737 cm−1 for ENR, 1621 cm−1 for OXY, and 1723 cm−1 for OTA. The N–H bond stretching vibration of amide functional group was seen at 3377 cm−1 and 3509 cm−1 in OXY and OTA, respectively. In contrast, CIP revealed N–H stretching bands at 3397 cm−1, indicating the presence of a secondary amine group. The C–F bond was observed at 1237 and 1240 cm−1 in ENR and CIP, respectively, while the C–Cl bond was detected at 713 cm−1 in OTA, confirming the presence of halogen functional group.
 | ||
| Fig. 5 (a) FT-IR and (b) UV-visible spectra of selected compounds (remaining are presented in Fig. S6). | ||
3.5. UV-visible analysis
In UV-vis spectroscopy analysis, the electronic transition spectra in different orbitals were analyzed. Key molecular transition parameters, including absorption peak wavelength (λmax), excitation energy, oscillator strength, and transition configuration composition (presented in %) for all molecules, are summarized in Table S5. Graphical representations of UV-vis spectra for all molecules are presented in Fig. 5(b) and S6(b). It was noted that all the molecules showed their maximum absorption from 250 nm to 450 nm. Among our selected 6 molecules, OXY and OTA exhibited maximum absorption at the longer wavelength of 431 nm and 396 nm, respectively. This transition occurred when electrons jumped from lower orbitals to higher orbitals, with a configuration composition of 63.49% for OXY and 96.27% for OTA. The measured oscillator strength of this transition state was 0.0047 for OXY and 0.081 for OTA. In contrast, AB1 and ZER showed maximum absorption (λmax) at lower wavelengths of 315 nm and 326 nm, respectively. This transition occurred when electrons jumped from lower orbitals to higher orbitals, with a configuration composition of 75.46% for AB1 and 90.05% for ZER. The measured oscillator strength of this transition state was 0.0721 for AB1 and 0.0952 for ZER. Among all of these molecules, OXY and OTA showed the highest degree of stability as their absorption maxima were at the longer wavelengths, indicating lower energy levels, where the excitation energy was 2.87 eV for OXY and 3.13 eV for OTA.54 Conversely, AB1 and ZER exhibited the lowest stability, with their absorption maxima at the shortest wavelengths, suggesting higher-energy states, where the excitation energy was 3.93 eV for AB1 and 3.80 eV for ZER.55 Among these molecules, AB1 (312 g mol−1) and ZER (318 g mol−1) have a lower molecular weight, while OXY (497 g mol−1) and OTA (404 g mol−1) have a comparatively higher molecular weight. Lower molecular weight exhibited the lower conjugation and shorter wavelength for UV absorption, hence lower stability and vice versa.56 These compounds contained numerous functional groups, where the presence of more functional groups is responsible for higher electron conjugation in the molecular structure. Consequently, the UV peak will appear in a longer wavelength region.57 Conversely, AB1 and ZER had fewer functional groups, indicating less conjugation. Therefore, UV absorption appeared at shorter wavelengths and experienced less molecular stability.3.6. Molecular docking and nonbonding interaction analysis
Molecular docking is a widely used computational approach for evaluating atomic-level interactions between chemical compounds and target proteins. Additionally, in predictive toxicology and risk assessment studies, this method is also used to achieve the optimal docked conformations, including the orientations and placements of ligands within proteins.58 In molecular docking, a more negative value indicates greater binding interactions between proteins and ligands, which is similar to the lock and key model.59 Numerous bonds are responsible for stabilizing docked conformations, such as hydrogen bonds, hydrophobic bonds, and van der Waals forces.60In this study, all of our selected compounds exhibited strong binding affinities within 2RJQ, 4G1C, and 3P0M proteins, illustrated in Fig. 6. ZER exhibited strong binding affinities within 2RJQ, 4G1C, and 3P0M proteins, which were −8.0 kcal mol−1, −8.0 kcal mol−1, and −7.9 kcal mol−1, respectively. Therefore, the ZER compound showed an increased potential for cardiac toxicity. AB1 compound showed stronger binding affinities of −8.9 kcal mol−1 and −8.1 kcal mol−1 within the 4G1C and 3P0M proteins, respectively. Consequently, this compound strongly exhibited cardiotoxicity and nephrotoxicity. Furthermore, CIP and ENR compounds showed a greater likelihood of nephrotoxicity, with higher similar docking scores −8.1 kcal mol−1 within the 3P0M protein. OXY showed extremely negative docking scores, such as −8.8 kcal mol−1 within the 4 G1C protein and −8.7 kcal mol−1 within the 3P0M protein, indicating residues of this drug demonstrated cardiotoxicity and nephrotoxicity, respectively. Therefore, the OTA compound exhibited higher docking scores within three selected proteins. It showed −8.8 kcal mol−1, −9.0 kcal mol−1, and −9.4 kcal mol−1 within the 2RJQ, 4 G1C, and 3P0M proteins, respectively. Therefore, residues of OTA compounds strongly bound with these proteins, leading to hepatotoxicity, cardiac toxicity, and nephrotoxicity (Fig. 7).
 | ||
| Fig. 6 Comparative binding affinity of all compounds with some selected proteins (PDB ID: 2RJQ, 4G1C, and 3P0M). | ||
Hydrogen bonding is crucial in non-bonding interactions for both donor and acceptor in protein ligand interactions. Receptor proteins 2RJQ, 4 G1C, and 3P0M exhibited a heterogeneous donor region presented in pink, which employed lower electron densities, and an acceptor region presented in green, which employed higher electron densities.61 Non-bonding interactions of all ligands and receptor 2RJQ, 4 G1C, and 3P0M proteins are visualized in Fig. 8(a)–(c) and summarized in Table S6. Across these selected proteins, binding was stabilized by the mixture of conventional hydrogen bonds, carbon-hydrogen bonds, π-type interactions (π–alkyl, π–π stacking, π–cation, and π–anion), and alkyl contacts. To exemplify, ZER formed conventional hydrogen bonds with LEU 443 (1.99 Å) within 2RJQ; SER 251 (2.48 Å) within 4G1C; and SER 53 (2.69 Å), and LYS 72 (2.45 Å) within 3P0M. Carbon–hydrogen bonds, π–cation, π–π T-shaped, alkyl, π–alkyl, and π–anion bonds were also associated with different amino acid residues across all selected proteins. AB1 compound also formed carbon–hydrogen bonds, conventional hydrogen bonds, alkyl, π–alkyl, and π–anion bonds, with bond distances ranging from 2.18 Å to 5.41 Å with different amino acid residues. CIP and ENR showed similar interaction patterns within 2RJQ and 4 G1C proteins; both formed conventional hydrogen bonds and multiple carbon–hydrogen bonds to residues lining the pocket. These compounds also engaged in alkyl, π–alkyl, and π–cation contacts. All of these bonds were also found in CIP and ENR across the 3P0M protein with halogen (fluorine). OXY formed conventional hydrogen bonds with numerous amino acid residues throughout all the selected proteins. This compound also formed carbon–hydrogen bonds, π–alkyl, π–cation, and π–sigma bonds. OTA exhibited the hydrogen-bonding pattern across LEU 379 (1.94 Å), LEU 443 (2.26 Å), SER 441 (2.58 Å), and GLY 380 (2.06 Å), combined with additional carbon–hydrogen bonds, π–anion, π–π stacked, and π-interactions for the 2RJQ protein. Almost similar types of bonds were also found within the 4G1C and 3P0M proteins, varying in bond length.
 | ||
| Fig. 8 RMSD and RMSF of the selected complexes during 100 ns MD simulation, for (a and b) 2RJQ, (c and d) 4G1C, and (e and f) 3P0M proteins. | ||
3.7. Molecular dynamics simulation analysis
MD simulation is a computational technique that investigates the physical movements of atoms and molecules over time. It offers an atomistic-level of conformational assessment, structural stability, and interaction mechanism of biomolecular systems such as proteins, nucleic acids, and protein–ligand complexes. To assess the dynamics of protein–ligand complexes, this study employed RMSD, RMSF, Rg, H-bond, and SASA analysis for 100 ns. All statistical analysis presented in Table 1. RMSD is a significant parameter in MD simulation because it assesses structural stability and changes over time. It reflects the conformational changes and overall stability, where a lower RMSD value (1–3 Å) indicates structural stability.62 Within the 2RJQ protein, a small rise in the RMSD fluctuation of OXY was noted at 32.25 ns and 99.95 ns, with values of 3.16 Å and 3.12 Å, respectively, while OTA showed relatively lower fluctuation. Within this protein, the RMSD fluctuation of OTA occurred from 1.47 Å (59 ns) to 2.97 Å (98 ns), while OXY occurred from 1.40 Å (0.25 ns) to 3.16 Å (32.25 ns). Furthermore, within the 4G1C protein, the RMSD fluctuation values ranged from 1.49 Å (0.25 ns) to 3.97 Å (75 ns) for OXY, while the 3P0M protein's fluctuation ranged from 1.02 Å (0.75 ns) to 1.98 Å (76.5 ns), as presented in Fig. 8(a), (c) and (e). After 60 ns, OTA exhibited a sharp rise, indicating the compound was undergoing reorientation or movement in the binding pocket of the 3P0M protein. However, OXY showed the lowest ±0.22 standard deviation within 3P0M and ±0.34 standard deviation within 2RJQ, compared to 4G1C (±0.48) protein. This lower deviation indicated minimal structural fluctuations and reflected the stability of the complexes during the simulation. Therefore, these compounds showed enhanced binding stability within both proteins, suggesting increased conformational stability and stiffness.39| Proteins | Compounds | MD simulation parameters (mean ± SD) | ||||
|---|---|---|---|---|---|---|
| RMSD | RMSF | Rg | H-bonds | SASA | ||
| 2RJQ | OXY | 2.18 ± 0.34 | 1.52 ± 0.64 | 19.64 ± 0.40 | 215 ± 16.33 | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 399 ± 241 399 ± 241 |
| OTA | 2.02 ± 0.31 | 1.40 ± 0.60 | 19.43 ± 0.40 | 211 ± 6.99 | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 085 ± 233 085 ± 233 |
|
| 4G1C | OXY | 2.74 ± 0.48 | 1.21 ± 0.48 | 20.01 ± 0.37 | 203 ± 16.96 | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 ± 276 300 ± 276 |
| OTA | 2.17 ± 0.36 | 1.30 ± 0.50 | 19.95 ± 0.38 | 202 ± 7.10 | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 287 ± 206 287 ± 206 |
|
| 3P0M | OXY | 1.56 ± 0.22 | 1.28 ± 0.54 | 20.57 ± 0.34 | 253 ± 14.81 | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 051 ± 240 051 ± 240 |
| OTA | 2.06 ± 0.55 | 1.24 ± 0.55 | 20.68 ± 0.35 | 252 ± 8.51 | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 157 ± 277 157 ± 277 |
|
Furthermore, an important measure of the compactness and general folding state of a macromolecule, like a protein, is Rg. Lower Rg value indicates a more compact or folded structure, while a higher Rg value provides a more expansive or unfolded form. This value can be tracked throughout an MD simulation to assess conformational dynamics, folding stability, and structural alterations.63 Rg value indicated the compactness of the protein–ligand complexes shown in Fig. S8(a), S9(a), and S10(a). The lower Rg value of OTA observed within 2RJQ (mean 19.43) and 4G1C (mean 19.95) proteins compared to 3P0M protein (mean 20.68), lower standard deviation within 4 G1C (±0.40) and 3P0M (±0.38), with minor fluctuations leading to the compactness and stability of complexes. Likewise, OXY and OTA showed lower Rg values within 2RJQ and 4G1C proteins. Thus, OXY and OTA exhibited reduced Rg values and less variation, resulting in increased stability and compactness of the complexes.
RMSF is a measure of the average deviation of atomic position from its mean position over time in an MD simulation. It provides information about the flexibility and dynamics of a protein structure. Regions with high RMSF values are typically more flexible, while low values indicate more rigid.64 The RMSF values within the 2RJQ protein varied from 0.64 Å to 4.14 Å for OXY and 0.45 Å to 3.5 Å for OTA among 270 and 550 amino acid residues. Slight fluctuations were observed for oxy at 3.07 Å (326 residue), 3.46 Å (356 residue), and 4.14 Å (524 residue). However, among 40 and 300 amino acid residues within the 4G1C protein, the RMSF values varied between 0.62 Å and 3.10 Å. The highest peak was observed at 3.10 Å (174 residue). Furthermore, among 20 and 350 amino acid residues, OXY and OTA displayed steady RMSF values ranging from 0.71 Å to 3.21 Å within the 3P0M protein, presented in Fig. 8(b), (d) and (f). The standard deviation was lower compared to the previous report,65 ranging from ±0.48 to ±0.64 within 2RJQ, 4G1C, and 3P0M proteins for both compounds. Comparatively higher RMSD and lower standard deviation values indicated more flexibility and less fluctuation of our complexes. The continuity of protein structure depends on hydrogen bonding. A greater and constant number of H-bonds indicates a stable protein–ligand combination throughout the system.66 OXY and OTA exhibited a higher number of H-bonds, ranging from 187 to 278 within the 2RJQ and 3P0M proteins, suggesting stronger intermolecular interactions leading to more stability and rigidity. While OXY and OTA exhibited a comparatively lower number of H-bonds, ranging from 179 to 223 within the 4G1C protein, as presented in Fig. S8(b), S9(b), and S10(b).
SASA is utilized to examine the solvent accessibility of docked complexes. SASA is potentially attributable to the interaction between complexes and solvents. The area of a protein that interacts with solvent molecules, usually water, is referred to as SASA.65 A protein with smaller SASA values is less exposed to solvents. In contrast, a protein with greater SASA values is more solvent-accessible and exhibits a more extended structure. OXY and OTA have comparatively higher SASA values within the 2RJQ and 3P0M proteins. The values ranged from 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 226 Å2 to 15
226 Å2 to 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 149 Å2 within 2RJQ protein and 15
149 Å2 within 2RJQ protein and 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 447 Å2 to 16
447 Å2 to 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 930 Å2 within 3P0M protein for OXY and OTA, as presented in Fig. S8(c), S9(c), and S10(c). OXY and OTA have relatively lower SASA values, ranging from 11
930 Å2 within 3P0M protein for OXY and OTA, as presented in Fig. S8(c), S9(c), and S10(c). OXY and OTA have relatively lower SASA values, ranging from 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 757 Å2 to 13
757 Å2 to 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 221 Å2 throughout the protein 4G1C. Consequently, OXY and OTA exhibited more solvent-accessible structures within 2RJQ and 3P0M proteins, while a less accessible structure within 4G1C protein. Additionally, these compounds exhibited more stable and consistent interaction patterns within selected proteins due to lower standard deviation. As per MD simulation results, our most promising compounds, OXY and OTA, showed more favourable MD profiles, making them stable complexes. Consequently, these compounds possess a significant risk of cardiotoxic, hepatotoxic, and nephrotoxic potential.
221 Å2 throughout the protein 4G1C. Consequently, OXY and OTA exhibited more solvent-accessible structures within 2RJQ and 3P0M proteins, while a less accessible structure within 4G1C protein. Additionally, these compounds exhibited more stable and consistent interaction patterns within selected proteins due to lower standard deviation. As per MD simulation results, our most promising compounds, OXY and OTA, showed more favourable MD profiles, making them stable complexes. Consequently, these compounds possess a significant risk of cardiotoxic, hepatotoxic, and nephrotoxic potential.
3.8. ADMET prediction
In a toxicological study, ADMET parameters are necessary for predicting pharmacokinetic properties, involving absorption, distribution, metabolism, excretion, and toxicity. The predicted ADMET parameters are presented in Table 2. All of the compounds exhibited positive intestinal absorption (HIA) and human oral bioavailability (HOB). Consequently, AB1 and CIP had higher HIA values (+0.98), indicating easy absorption from the gastrointestinal tract during oral administration.44 No compounds can be excreted more readily by the colon and the urinary system. In our study, ZER showed the highest HOB (+0.84). Promising positive values of HOB exhibited health issues. Moreover, a positive BBB value points out that the compound may be able to cross the blood–brain barrier, potentially exposing the central nervous system and posing a neurotoxic risk. Herein, ZER (+0.61) and AB1 (+0.93) showed positive blood–brain barrier (BBB) permeability, indicating they easily penetrate through the blood–brain barrier membrane. However, CIP, ENR, OXY, and OTA showed negative BBB permeability, suggesting minimal neurotoxicity. Pharmaceuticals with high plasma protein binding (PPB) have a limited therapeutic index. Regarding metabolism, most compounds did not exhibit any inhibition of the vital cytochrome P450 enzyme CYP2C9, except AB1. Cytochrome P450 enzyme inhibitor (AB1) found in the liver and intestines, which is responsible for decreasing the metabolism of any compounds. All of our compounds exhibited weak inhibition of the human ether-α-go-go-related gene (hERG), raising the risk of long QT syndrome and sudden cardiac death.67| Compounds | Absorption | Distribution | Metabolism | Toxicity | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| HIA | HOB | BBB | PPB | CYP450 2C9 | hERG | HPT | AOT | RAT | GT | TPT | |
| a HIA = human intestinal absorption, HOB = human oral bioavailability, BBB = blood–brain barrier, PPB = plasma protein binding, hERG = human ether-α-go-go-related gene inhibition, HPT = hepatotoxicity, AOT = acute oral toxicity, RAT = rat acute toxicity LD50 (mol kg−1), GT = genotoxicity, TPT = tetrahymena pyriformis toxicity. | |||||||||||
| ZER | +0.95 | +0.84 | +0.61 | 89.98 | NI (0.85) | WI (0.83) | +0.76 | IV (0.61) | 1.33 | 0.52 | +0.95 |
| AB1 | +0.98 | +0.26 | +0.93 | 84.54 | I (0.67) | WI (0.78) | +0.82 | I (0.81) | 3.59 | 0.98 | +1.00 |
| CIP | +0.98 | +0.13 | −0.94 | 25.52 | NI (0.85) | WI (0.75) | +0.98 | III (0.67) | 2.43 | 1.00 | +0.87 |
| ENR | +0.90 | +0.03 | −0.91 | 36.09 | NI (0.79) | WI (0.69) | +0.94 | III (0.68) | 2.27 | 1.00 | +0.98 |
| OXY | +0.85 | +0.68 | −0.99 | 42.65 | NI (0.91) | WI (0.99) | +0.97 | III (0.80) | 2.32 | 0.99 | +0.95 |
| OTA | +0.80 | +0.65 | −0.79 | 98.63 | NI (0.81) | WI (0.95) | +0.93 | I (0.83) | 4.27 | 0.96 | +0.99 |
All compounds exhibited potential hepatotoxic effects, ranging from +0.76 to +0.98. To exemplify, CIP (+0.98) demonstrated the most pronounced hepatotoxic effect, indicating a high likelihood of liver toxicity. AB1 and OTA showed category I acute oral toxicity, suggesting highly toxic and irritating. Additionally, category III (CIP, ENR, and OXY) and category IV (ZER) acute toxicity indicate less toxicity for oral usage. The rodent acute toxicity (LD50) refers to the amount of dose required to kill 50% of a test population of animals, such as mice, after immediate exposure. Furthermore, a lower LD50 dose of ZER (1.33 mol kg−1) compound means a small amount needed to cause death, indicating higher toxicity. All other compounds have considered LD50 values within 2.27 mol kg−1 to 4.27 mol kg−1, suggesting a comparatively lower risk range. In our study, all compounds showed genotoxic effects leading to the mutation or alteration of genetic materials, such as DNA, RNA, and chromosomes in a cell. Tetrahymena pyriformis is used as a tool to assess the toxicity of various chemicals, measuring how different compounds impact the organism's growth and survival. A higher Tetrahymena pyriformis toxicity value indicated a lower effective concentration of compounds upon test populations, suggesting less toxicity.68 Here, all compounds showed these toxicities, ranging from +0.87 to +1.00. Lower tetrahymena pyriformis toxicity value was observed in compound CIP (+0.87). Based on the toxicological evaluation, OXY and OTA exhibit a higher propensity for adverse toxicological effects, whereas ZER, AB1, and CIP display comparatively lower toxicity profiles.
3.9. PASS prediction
PASS is a computerized method that determines possible pharmacological activity as well as toxic effects of compounds based on structure–activity relationships.69 In this study, the probability of all compounds being active (Pa) values are presented in Table 3, including various biological activities. These compounds are used in chicken poultry feed for different purposes, such as treatment or control of infectious diseases, immunity, or production.25,28 Exposure to these compounds has harmful effects on the human body's respiratory, renal, digestive, and nervous systems. We noticed that all compounds showed reproductive dysfunction (Pa = 0.28–0.99), inflammation (Pa = 0.37–0.98), and respiratory failure effects (Pa = 0.25–0.91). Except AB1, all compounds exhibited allergic dermatitis, leukopenia, nausea, nephrotoxicity, and myocarditis syndrome with promising Pa values ranging from 0.32 to 0.90, 0.36 to 0.95, 0.49 to 0.96, 0.37 to 0.87, and 0.26 to 0.92, respectively. Herein, OXY and OTA demonstrated a greater risk of both nephrotoxicity and myocarditis, where OXY showed corresponding values of 0.87 and 0.92, and OTA exhibited values of 0.82 and 0.85, respectively. Moreover, compared to OXY (0.34) and OTA (0.53), ZER (0.80) and AB1 (0.94) were found to be remarkably carcinogenic. These compounds are in contact with the fetus during pregnancy. They can undergo congenital defects, which are known as teratogenic effects. The probable activity values on the teratogenic effect range from 0.55 to 0.94. Additionally, most compounds exhibited hematotoxic effects affecting bone marrow,70 with toxicity values ranging from 0.30 to 0.92. Except for carcinogenicity, the OXY compound showed potential probable activity of all adverse effects. The majority of compounds provide a positive PASS result on different adverse effects, indicating a broad toxicity profile.| Compounds | RD | IN | AD | LKP | HMT | NS | RF | NT | MY | CAR | TT |
|---|---|---|---|---|---|---|---|---|---|---|---|
| a RD = reproductive dysfunction, IN = inflammation, AD = allergic dermatitis, LKP = leukopenia, HMT = hematotoxin, NS = nausea, RF = respiratory failure, NT = nephrotoxic, MY = myocarditis, CAR = carcinogenic, TT = teratogen. | |||||||||||
| ZER | 0.64 | 0.56 | 0.34 | 0.45 | 0.61 | 0.49 | 0.51 | 0.43 | 0.26 | 0.80 | 0.81 |
| AB1 | 0.35 | 0.48 | — | — | — | — | 0.25 | — | 0.26 | 0.94 | 0.82 |
| CIP | 0.28 | 0.37 | 0.44 | 0.69 | 0.30 | 0.57 | 0.37 | 0.40 | — | — | — |
| ENR | 0.69 | 0.38 | 0.88 | 0.79 | 0.39 | 0.61 | 0.64 | 0.37 | 0.28 | — | — |
| OXY | 0.99 | 0.98 | 0.90 | 0.95 | 0.97 | 0.96 | 0.91 | 0.87 | 0.92 | 0.34 | 0.94 |
| OTA | 0.54 | 0.55 | 0.32 | 0.36 | 0.58 | 0.59 | 0.45 | 0.82 | 0.85 | 0.53 | 0.55 |
3.10. pIC50 analysis
The analysis of the complex relationship between independent and dependent variables was done by using a validated multivariate mathematical model. Therefore, descriptors: Chiv5, bcutm1, MRVSA9, MRVSA6, PEOEVSA5, GATSv4, J, and Diameter were regarded as independent variables and pIC50 as the dependent variable. The QSAR method allowed the prediction of key features related to the biological activity of novel compounds.71 The topological and physicochemical characteristics of the molecules are represented by molecular descriptors. A multiple linear regression (MLR) equation was used to examine the pIC50 values, suggesting the expression for the previously indicated complex correlation. The typical pIC50 values for standard drugs fall between 4 and 10.72 In this study, the pIC50 values of all compounds ranged from 4.35 to 4.92, while ZER showed a pIC50 of 4.35 and OXY showed 4.92, as illustrated in Table S7 and Fig. 9.4. Conclusion
This study demonstrated an integrated computational toxicological analysis of mycotoxins and antibiotic residues frequently found in poultry feed and tissues. It also revealed the potential adverse effects of natural and intentional contaminants on human health. Thermodynamic analysis showed that all the compounds had higher negative free energy and enthalpy values, indicating they are energetically stable. OTA (−1739.06 Hartree) and OXY (−1638.93 Hartree) showed the highest negative free energy, indicating higher spontaneity in interaction. A lower HOMO–LUMO energy gap indicates their easy electron transfer, higher reactivity, and kinetically less stable. Furthermore, ZER had the highest chemical hardness (2.32 eV) and the lowest chemical softness (0.22 eV−1), representing resistance to electronic structure deformation. MD simulation revealed that all of the compounds had greater binding affinities. OTA compound exhibited higher docking scores of −8.8 kcal mol−1, −8.9 kcal mol−1, and −9.4 kcal mol−1 within 2RJQ, 4 G1C, and 3P0M proteins, respectively. The most promising compounds, OXY and OTA, were subjected to MD simulation, and their greater stability was demonstrated by favourable RMSD, RMSF, Rg, H-bond, and SASA values. All compounds exhibited a multi-organ toxicity profile. According to the pharmacokinetic analysis, OXY and OTA showed significant hepatotoxicity. These compounds are also responsible for cardiotoxicity (Pa = 0.92 and 0.85, respectively) and nephrotoxicity (Pa = 0.87 and 0.82, respectively), indicating a significant risk of adverse effects. Therefore, the goal of our research is to highlight the endangered condition of poultry farms, to understand biochemical behavior, and the biological effects of our selected compounds. The limitation of our study is dependent on in silico analyses without experimental evaluations. However, our findings provide a foundation for in vitro and in vivo validation. Integrating these findings will improve understanding of toxicity mechanisms and help to disclose other alternative medications.Author contributions
Jahida Akter and Kabir Hossain: writing original draft, methodology, software, formal analysis, and data curation. Santanu Deb Nath: writing original draft, and investigation. Hozzatul Islam: writing original draft, and investigation. Khaleda Afrin: writing original draft, and investigation. Hamida Begum: writing original draft, and investigation. Monir Uzzaman: conceptualization, writing review and editing, validation, supervision, and visualization.Conflicts of interest
The authors declare that there are no conflicts of interest related to the publication of this article. This study was conducted independently, all funding sources were fully disclosed, and had no influence on the study outcomes.Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information (SI) files. Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00856a.Acknowledgements
The authors extend their heartfelt appreciation for supports and laboratory resources provided by the Computer in Chemistry and Medicine Laboratory, Dhaka, Bangladesh (https://sites.google.com/view/computer-chemistry-medicine/).References
- M. A. Hamid, M. A. Rahman, S. Ahmed and K. M. Hossain, Status of poultry industry in Bangladesh and the role of private sector for its development, Asian J. Poult. Sci., 2016, 11, 1–13 CrossRef.
- P. Kumar, D. K. Mahato, M. Kamle, T. K. Mohanta and S. G. Kang, Aflatoxins: A Global Concern for Food Safety, Human Health and Their Management, Front. Microbiol., 2017, 7, 2170 CrossRef PubMed.
- L. Pinotti, M. Ottoboni, C. Giromini, V. Dell'Orto, F. Cheli, L. Pinotti, M. Ottoboni, C. Giromini, V. Dell'Orto and F. Cheli, Mycotoxin Contamination in the EU Feed Supply Chain: A Focus on Cereal Byproducts, Toxins, 2016, 8, 45 CrossRef PubMed.
- T. Lešić, J. Pleadin, N. Kudumija, D. Tomašković and A. Vulić, Mycotoxin Residues in Chicken Breast Muscle and Liver, Foods, 2025, 14, 2017 CrossRef PubMed.
- European Commission, Commission Regulation (EU) 2023/915 of 25 April 2023 on maximum levels for certain contaminants in food and repealing Regulation (EC) No 1881/2006, Off. J. Eur. Union, 2023, 119, 103–157 Search PubMed.
- B. A. Al-Jaal, M. Jaganjac, A. Barcaru, P. Horvatovich and A. Latiff, Aflatoxin, fumonisin, ochratoxin, zearalenone and deoxynivalenol biomarkers in human biological fluids: A systematic literature review, 2001-2018, Food Chem. Toxicol., 2019, 129, 211–228 CrossRef CAS PubMed.
- A. Leiva, G. Méndez, C. Rodríguez, A. Molina and F. Granados-Chinchilla, Chemical assessment of mycotoxin contaminants and veterinary residues in Costa Rican animal feed, Int. J. Food Contam., 2019, 6, 1–26 Search PubMed.
- O. M. Ghimpeeanu, E. N. Pogurschi, D. C. Popa, N. Dragomir, T. Drăgotoiu, O. D. Mihai, C. D. Petcu, O. M. Ghimpeeanu, E. N. Pogurschi, D. C. Popa, N. Dragomir, T. Drăgotoiu, O. D. Mihai and C. D. Petcu, Antibiotic Use in Livestock and Residues in Food – A Public Health Threat: A Review, Foods, 2022, 11, 1430 CrossRef PubMed.
- B. Matubber, S. M. E. Rahman, S. Akter, F. Farzana, M. A. Rahman, J. Wang and D. H. Oh, Prevalence of antibiotic resistance and residue in meats in southwest region of Bangladesh, Food Humanity, 2025, 4, 100636 CrossRef.
- N. T. Nhung, N. Chansiripornchai and J. J. Carrique-Mas, Antimicrobial Resistance in Bacterial Poultry Pathogens: A Review, Front. Vet. Sci., 2017, 4, 284486 Search PubMed.
- D. C. S. De Assis, G. R. Da Silva, I. P. Lanza, A. C. D. S. R. Ribeiro, É. M. Q. Lana, L. J. C. Lara, T. C. De Figueiredo and S. V. De Cançado, Evaluation of the Presence and Levels of Enrofloxacin, Ciprofloxacin, Sulfaquinoxaline and Oxytetracycline in Broiler Chickens after Drug Administration, PLoS One, 2016, 11, e0166402 CrossRef PubMed.
- M. G. Bacanlı, The two faces of antibiotics: an overview of the effects of antibiotic residues in foodstuffs, Arch. Toxicol., 2024, 98, 1717–1725 CrossRef PubMed.
- E. S. Lopes, K. C. Ferreira Santaren, L. C. Araujo de Souza, C. E. T. Parente, R. C. Picão, D. A. Jurelevicius and L. Seldin, Cross-environmental cycling of antimicrobial resistance in agricultural areas fertilized with poultry litter: a One Health approach, Environ. Pollut., 2024, 363, 125177 CrossRef CAS PubMed.
- M. Foroutankhah, M. Toghyani, N. Landy and L. Evaluation of Calendula officinalis, (marigold) flower as a natural growth promoter in comparison with an antibiotic growth promoter on growth performance, carcass traits and humoral immune responses of broilers, Anim. Nutr., 2019, 5, 314–318 CrossRef PubMed.
- J. Martínez, M. Hernández-Rodríguez, A. Méndez-Albores, G. Téllez-Isaías, E. M. Jiménez, M. I. Nicolás-Vázquez, R. M. Ruvalcaba, J. Martínez, M. Hernández-Rodríguez, A. Méndez-Albores, G. Téllez-Isaías, E. M. Jiménez, M. I. Nicolás-Vázquez and R. M. Ruvalcaba, Computational Studies of Aflatoxin B1 (AFB1): A Review, Toxin, 2023, 15, 135 CrossRef PubMed.
- K. Ropejko and M. Twarużek, Zearalenone and Its Metabolites – General Overview, Occurrence, and Toxicity, Toxins, 2021, 13, 35 CrossRef CAS PubMed.
- A. Fiaz, D. Zhu and J. Sun, Environmental fate of tetracycline antibiotics: degradation pathway mechanisms, challenges, and perspectives, Environ. Sci. Eur., 2021, 33, 64 CrossRef CAS.
- Y. Fakhri, F. Mehri, V. Ranaei, Z. Pilevar, F. Soleimani, R. Nasiri and A. Mousavi Khaneghah, The Prevalence and Concentration of Mycotoxins (Aflatoxins, Deoxynivalenol, Zearalenone, and Ochratoxin A) in Domestic Bird Eggs: A Global Systematic Review, Meta-analysis, and Probabilistic Risk Assessment, J. Food Prot., 2025, 88, 100600 CrossRef CAS PubMed.
- R. M. Olariu, N. I. Fiţ, C. M. Bouari, G. C. Nadăş, R. M. Olariu, N. I. Fiţ, C. M. Bouari and G. C. Nadăş, Mycotoxins in Broiler Production: Impacts on Growth, Immunity, Vaccine Efficacy, and Food Safety, Toxins, 2025, 17, 261 CrossRef CAS PubMed.
- A. Filazi, B. Yurdakok-Dikmen, O. Kuzukiran, U. T. Sireli, A. Filazi, B. Yurdakok-Dikmen, O. Kuzukiran and U. T. Sireli, Chemical Contaminants in Poultry Meat and Products, in Poult. Sci., ed. M. Manafi, InTech, 1st edn, 2017, ch. 9, pp. 171–190 Search PubMed.
- J. Łukasik and H. Szajewska, Can probiotics replace antibiotics? Not yet, says the evidence, Expert Rev. Anti-Infect. Ther., 2025, 23, 323–324 CrossRef PubMed.
- G. He, L. Feng and H. Chen, A QSAR Study of the Acute Toxicity of Halogenated Phenols, Procedia Eng., 2012, 43, 204–209 CrossRef CAS.
- J. L. Calais, R. G. Parr and W. Yang, Density-functional theory of atoms and molecules, Int. J. Quantum Chem., 1993, 47, 107–108 CrossRef.
- M. Uzzaman, M. K. Hasan, S. Mahmud, K. Fatema and M. M. Matin, Structure-based design of new diclofenac: Physicochemical, spectral, molecular docking, dynamics simulation and ADMET studies, Inform. Med. Unlocked, 2021, 25, 100677 CrossRef.
- A. M. Fouad, D. Ruan, H. K. El-Senousey, W. Chen, S. Jiang, C. Zheng, A. M. Fouad, D. Ruan, H. K. El-Senousey, W. Chen, S. Jiang and C. Zheng, Harmful Effects and Control Strategies of Aflatoxin B1 Produced by Aspergillus flavus and Aspergillus parasiticus Strains on Poultry: Review, Toxins, 2019, 11, 176 CrossRef CAS PubMed.
- E. Bonerba, A. Manfredi, M. M. Dimuccio, P. Lorusso, A. Pandiscia, V. Terio, A. Di Pinto, S. Panseri, E. Ceci, G. Bozzo, E. Bonerba, A. Manfredi, M. M. Dimuccio, P. Lorusso, A. Pandiscia, V. Terio, A. Di Pinto, S. Panseri, E. Ceci and G. Bozzo, Ochratoxin A in Poultry Supply Chain: Overview of Feed Occurrence, Carry-Over, and Pathognomonic Lesions in Target Organs to Promote Food Safety, Toxins, 2024, 16, 487 CrossRef CAS PubMed.
- L. Sun, R. Li, B. Tai, S. Hussain, G. Wang, X. Liu and F. Xing, Current status of major mycotoxins contamination in food and feed in Asia – a review, ACS Food Sci. Technol., 2023, 3, 231–244 CrossRef CAS.
- M. R. A. Ferdous, M. R. Ahmed, S. H. Khan, M. A. Mukta, T. T. Anika, M. T. Hossain, M. Z. Islam and K. Rafiq, Effect of discriminate and indiscriminate use of oxytetracycline on residual status in broiler soft tissues, Vet. World, 2020, 13, 61–67 CAS.
- M. Gbylik-Sikorska, B. Łebkowska-Wieruszewska, A. Gajda, E. Nowacka-Kozak, A. Lisowski and A. Posyniak, Transfer of enrofloxacin, ciprofloxacin, and lincomycin into eggshells and residue depletion in egg components after multiple oral administration to laying hens, Poult. Sci., 2021, 100, 101341 CrossRef CAS PubMed.
- M. M. I. Bhuiyan, M. S. Islam, M. R. Hasan and K. R. Islam, Thin layer chromatographic detection of enrofloxacin antibiotic residues in poultry tissues, Asian-Australas. J. Food Saf. Secur., 2021, 5, 11–18 CrossRef.
- M. Z. Saif, N. J. I. Esha, S. T. Quayum, S. Rahman, M. A. Al-Gawati, G. Alsowygh, H. Albrithen, A. N. Alodhayb, R. A. Poirier and K. M. Uddin, Sci. Rep., 2024, 14, 1–20 Search PubMed.
- B. Liu, X. Wang, N. Wu, F. Liu and H. Rao, Curr. Med. Chem., 2025, 32, 4786–4804 CrossRef CAS PubMed.
- M. M. Ahmed, Z. Shafat, S. Tazyeen, R. Ali, M. N. Almashjary, R. Al-Raddadi, S. Harakeh, A. Alam, S. Haque and R. Ishrat, Identification of pathogenic genes associated with CKD: An integrated bioinformatics approach, Front. Genet., 2022, 13, 891055 CrossRef CAS PubMed.
- H. Peng, B. Lv, J. Du, Y. Huang, Q. Cui, C. Cui and H. Jin, Identification of Metabolism-Related Hub Genes in Heart Failure via Comprehensive Transcriptome Analysis, Genes, 2025, 16, 305 CrossRef CAS PubMed.
- V. Škrhák, M. Novotný, C. P. Feidakis, R. Krivák and D. Hoksza, CryptoBench: cryptic protein-ligand binding sites dataset and benchmark, Bioinformatics, 2024, 41, btae745 CrossRef PubMed.
- R. Krivák and D. Hoksza, P2Rank: machine learning based tool for rapid and accurate prediction of ligand binding sites from protein structure, J. Chem. Inform., 2018, 10, 1–12 Search PubMed.
- A. Vikhar Danish Ahmad, S. W. Khan, S. A. Ali and Q. Yasar, Network pharmacology combined with molecular docking and experimental verification to elucidate the effect of flavan-3-ols and aromatic resin on anxiety, Sci. Rep., 2024, 14, 9799 CrossRef CAS PubMed.
- V. A. Obakachi, K. K. Govender and P. P. Govender, Targeting Acute Myeloid Leukemia with 1,2,4-triazolo[4,3-b]pyridazine derivatives: a molecular docking, dynamics, and ADMET approach, In Silico Pharmacol., 2025, 13, 1–21 CrossRef PubMed.
- M. O. Farque, R. M. Islam, M. F. Rahman Joni, M. Akter, S. Akter, M. D. Islam, M. J. Bin Salim, A. A. Aziz, E. Kabir and M. Uzzaman, Structural modification of Naproxen; physicochemical, spectral, medicinal, and pharmacological evaluation, Inform. Med. Unlocked, 2025, 53, 101617 CrossRef.
- İ. Yetek, S. Mert, E. Tunca, A. Bayrakdar and R. Kasımoğulları, Synthesis, molecular docking and molecular dynamics simulations, drug-likeness studies, ADMET prediction and biological evaluation of novel pyrazole-carboxamides bearing sulfonamide moiety as potent carbonic anhydrase inhibitors, Mol. Diversity, 2024, 29, 1207–1227 CrossRef PubMed.
- P. Santoso, S. Ilyas, Y. H. Midoen, R. Maliza and D. F. Belahusna, Predictive bioactivity of compounds from Vitis gracilis leaf extract to counteract doxorubicin-induced cardiotoxicity via sirtuin 1 and adenosine monophosphate-activated protein kinase: an in-silico study, J. Appl. Pharm. Sci., 2024, 14, 99–114 CAS.
- J. Dong, D. S. Cao, H. Y. Miao, S. Liu, B. C. Deng, Y. H. Yun, N. N. Wang, A. P. Lu, W. Bin Zeng and A. F. Chen, ChemDes: an integrated web-based platform for molecular descriptor and fingerprint computation, J. Cheminform., 2015, 7, 60 CrossRef PubMed.
- M. J. H. Moniruzzaman, M. N. Uddin, A. Ahsan and T. Mahmud, Quantum Chemical, Molecular Docking, and ADMET Predictions of Ketorolac and its Modified Analogues, Biomed J Sci Tech Res, 2018, 11, 8723–8729 Search PubMed.
- T. Sultana, J. Tasnim, M. W. H. Talukder, M. L. Mia, S. N. Suchana, F. Akter, M. A. Saleh, M. F. Afrin and M. Uzzaman, Physicochemical and toxicological studies of some commonly used triazine-based herbicides; In-silico approach, Inform. Med. Unlocked, 2023, 42, 101378 CrossRef.
- M. N. Uddin, M. N. H. Knock, M. Uzzaman, M. M. H. Bhuiyan, A. F. M. Sanaullah, W. Shumi and H. M. Sadrul Amin, Microwave-assisted synthesis, characterization, molecular docking and pharmacological activities of some new 2′-hydroxychalcone derivatives, J. Mol. Struct., 2020, 1206, 127678 CrossRef CAS.
- M. N. Uddin, S. Khandaker, Moniruzzaman, M. S. Amin, W. Shumi, M. A. Rahman and S. M. Rahman, Synthesis, characterization, molecular modeling, antioxidant and microbial properties of some Titanium (IV) complexes of Schiff bases, J. Mol. Struct., 2018, 1166, 79–90 CrossRef CAS.
- V. Hadigheh Rezvan, Molecular structure, HOMO–LUMO, and NLO studies of some quinoxaline 1,4-dioxide derivatives: Computational (HF and DFT) analysis, Results Chem., 2024, 7, 101437 CrossRef CAS.
- K. I. Williamson, D. J. C. Herr and Y. Mo, Mapping the correlations between bandgap, HOMO, and LUMO trends for meta substituted Zn-MOFs, J. Comput. Chem., 2024, 45, 2119–2127 CrossRef CAS PubMed.
- M. M. Talmaciu, E. Bodoki and R. Oprean, Global chemical reactivity parameters for several chiral beta-blockers from the density functional theory viewpoint, Clujul Med., 2016, 89, 513–518 Search PubMed.
- J. Yu, N. Q. Su and W. Yang, Describing chemical reactivity with frontier molecular orbitalets, JACS Au, 2022, 2, 1383–1394 CrossRef CAS PubMed.
- A. Ullah, M. Ibrahim, A. Yousuf, M. A. Ali, H. L. Xu and M. Arshad, Crafting optical wonders: the interplay of electron push–pull dynamics and π-conjugation in non-linear optics, Next Mater., 2025, 9, 101239 CrossRef CAS.
- K. R. Raghi, D. R. Sherin, M. J. Saumya, P. S. Arun, V. N. Sobha and T. K. Manojkumar, Computational study of molecular electrostatic potential, docking and dynamics simulations of gallic acid derivatives as ABL inhibitors, Comput. Biol. Chem., 2018, 74, 239–246 CrossRef CAS PubMed.
- H. Guan, H. Sun, X. Zhao, H. Guan, H. Sun and X. Zhao, Application of Density Functional Theory to Molecular Engineering of Pharmaceutical Formulations, Int. J. Mol. Sci., 2025, 26, 3262 CrossRef CAS PubMed.
- S. Kyushin and Y. Suzuki, Cooperation of σ–π and σ*–π* Conjugation in the UV/Vis and Fluorescence Spectra of 9,10-Disilylanthracene, Molecules, 2022, 27, 2241 CrossRef CAS PubMed.
- B. Y. Jeon, A. Kidanemariam, J. Noh, C. Hyun, H. J. Mun, K. Park, S. J. Jung, Y. Jeon, P. J. Yoo, J. H. Park, H. T. Jung, T. J. Shin and J. Park, Strong Bathochromic Shift of Conjugated Polymer Nanowires Assembled with a Liquid Crystalline Alkyl Benzoic Acid via a Film Dispersion Process, ACS Omega, 2021, 6, 34876–34888 CrossRef CAS PubMed.
- P. Mahesha, N. S. Shetty and S. D. Kulkarni, Effect of conjugation length on the photophysical properties of naphthyl chalcones: a DFT and experimental study, Discover Appl. Sci., 2025, 7, 1102 CrossRef CAS.
- Z. Fang, X. Zhang, F. Wu, B. Huang, Z. Cao and B. Yi, Substituent effects in 2-hydroxybenzophenone-based ultraviolet absorbers, J. Mol. Liq., 2024, 412, 125630 CrossRef CAS.
- R. Jakhar, M. Dangi, A. Khichi and A. K. Chhillar, Relevance of Molecular Docking Studies in Drug Designing, Curr. Bioinform., 2020, 15, 270–278 CrossRef CAS.
- M. Er-rajy, M. El fadili, S. Mujwar, H. Imtara, O. Al kamaly, S. Z. Alshawwa, F. A. Nasr, S. Zarougui, M. Elhallaoui, M. Er-rajy, M. El fadili, S. Mujwar, H. Imtara, O. Al kamaly, S. Z. Alshawwa, F. A. Nasr, S. Zarougui and M. Elhallaoui, Design of novel anti-cancer agents targeting COX-2 inhibitors based on computational studies, Arabian J. Chem., 2023, 16, 105193 CrossRef CAS.
- D. Shukla, I. Azad, S. Y. Sheikh, S. N. Ali, N. Ahmad, A. Kamal, M. Faiyyaz, A. R. Khan, V. Ahmad, A. A. Alghamdi, M. Nasibullah and F. Hassan, Quantum chemical modelling, molecular docking, and ADMET evaluation of imidazole phenothiazine hybrids, Sci. Rep., 2025, 15, 23413 CrossRef CAS PubMed.
- Z. Ma, A. Ajibade and X. Zou, Docking strategies for predicting protein–ligand interactions and their application to structure-based drug design, Commun. Inf. Syst., 2024, 24, 199–230 CrossRef PubMed.
- L. Martínez, Evaluation of a Simple and Robust Method for Simultaneous Determination of Multiple Mycotoxins in Corn and Corn-Based Products by Liquid Chromatography Coupled to Tandem Mass Spectrometry, PLoS One, 2015, 10, e0119264 CrossRef PubMed.
- M. M. Jwaid and A. A. Adnan, In-silico design, molecular docking, molecular dynamic simulations, molecular mechanics with generalized Born and surface area solvation study, and pharmacokinetic prediction of novel diclofenac as anti-inflammatory compounds, Turk. Comput. Theor. Chem., 2024, 8, 108–121 CrossRef CAS.
- P. K. Arya, K. Barik, A. K. Singh and A. Kumar, Molecular docking and simulation studies of medicinal plant phytochemicals with Leishmania donovani adenosylmethionine decarboxylase, J. Appl. Biol. Biotechnol., 2023, 12, 219–228 Search PubMed.
- K. Hossain, J. Akter, Md. H. Islam, T. Fabiha, Md. M. Hasan, N. Afreen and M. Uzzaman, Computational design and evaluation employed DFT, molecular docking, molecular dynamics, and ADMET analysis of salen-based drug candidates for potential cancer and tuberculosis treatment, In Silico Pharmacol., 2025, 14, 3 CrossRef PubMed.
- A. A. Elfiky, H. A. Mahran, I. M. Ibrahim, M. N. Ibrahim and W. M. Elshemey, Molecular dynamics simulations and MM-GBSA reveal novel guanosine derivatives against SARS-CoV-2 RNA dependent RNA polymerase, RSC Adv., 2022, 12, 2741–2750 RSC.
- V. K. Tran-Nguyen, U. F. Randriharimanamizara and O. Taboureau, HERGAI: an artificial intelligence tool for structure-based prediction of hERG inhibitors, J. Cheminform., 2025, 17, 110 CrossRef CAS PubMed.
- O. V. Tinkov, V. Y. Grigorev and L. D. Grigoreva, QSAR analysis of the acute toxicity of avermectins towards Tetrahymena pyriformis, SAR QSAR Environ. Res., 2021, 32, 541–571 CrossRef CAS PubMed.
- M. F. Afrin, E. Kabir, M. R. O. K. Noyon, N. Akter, T. Sultana, J. Un Nayeem and M. Uzzaman, Spectrochemical, biological, and toxicological studies of DDT, DDD and DDE: An in-silico approach, Inform. Med. Unlocked, 2023, 39, 101254 CrossRef.
- A. P. Challa, A. L. Beam, M. Shen, T. Peryea, R. R. Lavieri, E. S. Lippmann and D. M. Aronoff, Machine learning on drug-specific data to predict small molecule teratogenicity, Reprod. Toxicol., 2020, 95, 148–158 CrossRef CAS PubMed.
- E. Kabir, M. R. O. K. Noyon and M. Uzzaman, Computational and Pharmacokinetic Investigation of Some Heterocyclic Amide Derivatives as Cyclooxygenase Inhibitors: An In-Silico Approach, Pharmacogn. J., 2023, 15, 194–207 CrossRef CAS.
- S. Tukur, G. A. Shallangwa and A. Ibrahim, Theoretical QSAR modelling and molecular docking studies of some 4-hydroxyphenylpyruvate dioxygenase (HPPD) enzyme inhibitors potentially used as herbicides, Heliyon, 2019, 5, e02859 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |