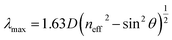Open Access Article
Open Access ArticleRapid, point-of-care, visual detection of RBD of SARS-CoV-2 spike protein using an acrylamide-free photonic crystal aptasensor
Meghana Mary Thomasac,
Parvathy R. Chandrana,
Lekshmi Lekshmananbc,
A. Peer Mohameda and
Saju Pillai *ac
*ac
aMaterials Science and Technology Division, CSIR-National Institute for Interdisciplinary Science and Technology (NIIST), Thiruvananthapuram 695 019, India. E-mail: pillai_saju@niist.res.in
bChemical Sciences and Technology Division, CSIR-National Institute for Interdisciplinary Science and Technology (NIIST), Thiruvananthapuram 695 019, India
cAcademy of Scientific and Innovative Research (AcSIR), Ghaziabad 201 002, India
First published on 5th May 2026
Abstract
As COVID-19 continues to reemerge with new variants, it has become a persistent challenge to public health, causing disruptions in society and the economy. We report a facile photonic crystal-based aptasensor for the rapid visual detection of the receptor-binding domain (RBD) of the severe acute respiratory syndrome (SARS-CoV-2) spike (S) protein. Monodispersed polystyrene@poly(2-hydroxyethyl methacrylate–acrylic acid) [PS@poly(HEMA–AA)] core–shell microspheres were synthesized to fabricate a stimuli-responsive photonic crystal (RPC). The integration of the RBD target-specific aptamer into the poly(HEMA–AA) hydrogel network facilitated the development of a biosensor that enabled RPCs to selectively bind to the SARS-CoV-2 spike protein using coupling chemistry. The interaction between the RBD and a single-stranded DNA aptamer causes the hydrogel to expand, leading to a color change and a shift in the photonic bandgap (PBG). This phenomenon can be used to detect the RBD of the SARS-CoV-2 spike protein. The addition of the RBD of SARS-CoV-2-S-protein to the Apt-RPC aptasensor resulted in a rapid visual color shift from violet to green. The detection range of the developed aptasensor was estimated to be 100–1000 ng with excellent selectivity. Thus, the developed aptasensor offers distinct advantages over conventional detection methods. Our study also facilitates the development of a straightforward point-of-care self-testing kit that does not rely on toxic acrylamide-based chemicals. Furthermore, the sensor eliminates the need for complex sample preparation or signal amplification steps, making it a promising platform for point-of-care diagnosis.
Introduction
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic that broke out in 2019 (COVID-19) has had a devastating impact on people's livelihoods, economy, and social life at large.1–6 According to the World Health Organization (WHO), COVID-19 has affected 778 million people worldwide and has led to 7 million deaths worldwide.7 SARS-CoV-2 has mutated into multiple variants and sub-variants, including, but not limited to, alpha, beta, gamma, delta, epsilon, omicron, and the latest XEC variant, and continues to spread among individuals with enhanced transmissibility and virulence.8–10 Currently, reverse transcription polymerase chain reaction (RT-PCR) using gold as the standard is the predominant method for the detection of SARS-CoV-2, although it is time-consuming. However, rapid on-site diagnosis of SARS-CoV-2 is impeded by intricate sample handling protocols, the need for skilled personnel, and prolonged detection time.11,12 Alternatively, colloidal gold immunochromatography and antigen-based enzyme-linked immunosorbent assays (ELISA) are qualitative detection methods for SARS-CoV-2.13,14 According to several studies, the receptor-binding domain (RBD) of the SARS-CoV-2 spike glycoprotein (S-protein) mediates viral cell entry and disease progression. The RBD of the SARS-CoV-2 S-protein infiltrates human respiratory and digestive epithelial cells via angiotensin-converting enzyme II (ACE2), which plays a key role in the diagnosis and treatment of SARS-CoV-2.15,16 However, the bio-recognition components of most S-protein detection methods use antibodies or ACE-2 enzymes, and their synthesis protocols are time-consuming and involve the exploitation of animal models. Moreover, these methods are expensive, and proteins may be denatured when exposed to elevated temperatures or pH fluctuations.17,18 Therefore, the development of a facile RBD-based approach could serve as a viable option for the rapid, cost-effective PoC diagnosis of SARS-CoV-2 infection.Aptamers are an alternative to antibodies for the detection of the SARS-CoV-2 S protein. They are small, single-stranded DNA or RNA molecules that selectively bind to specific targets, including peptides, proteins, carbohydrates, small toxins, and live cells.19–23 Compared to antibodies, the smaller size of aptamers causes less hindrance to the surface of the virus. In addition, they are stable, cost-effective, possess negligible batch-to-batch variation, and are convenient for transportation.24 Aptamers with affinity for a specific target can be generated via the SELEX approach, which implies the successive evolution of ligands by exponential enrichment using a vast oligonucleotide library.25 Thus, aptamer-based sensors represent an effective therapeutic and prophylactic approach for controlling SARS-CoV-2 infection and potential future pandemics.
Integrating aptamers into responsive photonic crystals (RPCs) is an efficient method for analyte detection.26–31 They can shift the photonic bandgap (PBG) through a volume phase transition (swelling/deswelling) under external stimuli. Based on the variation in lattice spacing, RPCs display a blue or red shift in the PBG position, and the resultant structural color changes enable label-free visual detection of analyte molecules.32–36 Ye et al. reported aptamer-functionalized RPCs for the visual detection of heavy metal ions.37 Xuan et al. demonstrated aptasensor functionalized with P(NIPAM–AA) hydrogel, which is a one-dimensional CPCs, for the colorimetric detection of heavy metal ions.38 The hydrogel network embedded in CPCs shrinks owing to the specific binding of heavy metal ions to the functionalized single-stranded aptamer, which alters the lattice spacing and causes a blue shift in the PBG. Similarly, Murtaza et al. reported a photonic crystal-based aptasensor that requires a complex fabrication procedure, limiting its scalability for the detection of SARS-CoV-2.39 Nevertheless, a simple approach using smart core–shell polymer microspheres of 3D CPCs with the desired functionalities to modulate the PBG within the visible-light region has not been explored for SARS-CoV-2 S-protein detection.
In the present work, we demonstrate RPCs that are more specific to an analyte by attaching the aptamer sequence to the hydrogels through 1-ethyl-3-(3-dimethyl aminopropyl)carbodiimide (EDC)/N-hydroxysuccinimide (NHS) coupling. The designed biosensor integrates the benefits of RPCs and aptamers, providing an efficient and scalable approach for SARS-CoV-2 RBDs detection. We synthesized PS@poly(HEMA–AA) core–shell microspheres via emulsifier-free emulsion polymerization and employed them to fabricate 3D CPCs using a vertical deposition method.40 The rigid PS core imparts structural stability, whereas the soft hydrogel shell contributes to stimuli-responsive characteristics. An aptamer targeting the SARS-CoV-2 RBD was functionalized onto the carboxylic functional group of the hydrogel shell via EDC/NHS chemistry. The specific binding of RBD to a single-stranded DNA aptamer causes swelling of the hydrogel, and the corresponding red shift in the PBG was used to detect the SARS-CoV-2 S-protein (Scheme 1).
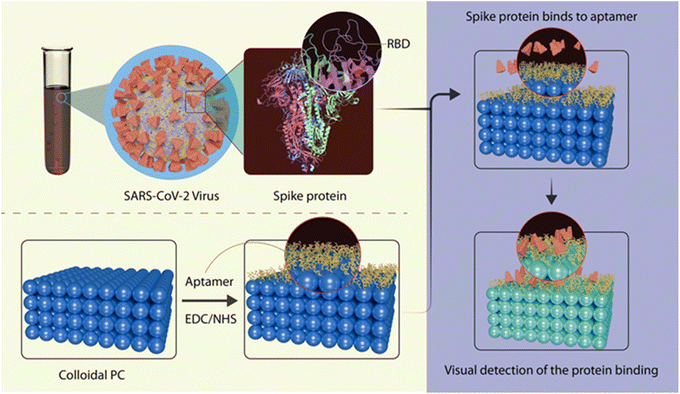 | ||
| Scheme 1 Schematic illustration of the aptasensor RPC for the rapid detection of RBD of the SARS-CoV-2 spike protein. | ||
Results and discussion
Synthesis and characterization of PS@poly(HEMA–AA) microspheres
Monodispersed PS@poly(HEMA–AA) core–shell microspheres were synthesized via emulsifier-free emulsion polymerization. Based on the differences in polarity and hydrophilicity of the co-monomers, the polymerization reaction resulted in the formation of core–shell microspheres with less polar, hydrophobic PS as the core and hydrophilic poly(HEMA–AA) as the shell. Acrylic acid imparts carboxylic functionality to the surface of CS microspheres. Fig. 1A shows the dynamic light scattering (DLS) results of the synthesized particles with an average hydrodynamic diameter of 195 nm and a polydispersity index (PDI) of 0.014, indicating the formation of monodispersed microspheres. The zeta potential value of the solution was −42.7, suggesting the formation of stable colloid. Fig. 1B presents the TEM image of the PS@poly(HEMA–AA) microspheres, where the darker core corresponds to the PS and the lighter area represents the poly(HEMA–AA) shell. The average diameter of the microspheres was 168 ± 10 nm, with a shell thickness of 40 ± 10 nm.The chemical functionality of the synthesized microspheres was analyzed using FT-IR. The IR spectrum of the PS@poly(HEMA–AA) microspheres (Fig. 2A) shows characteristic peaks of polystyrene (PS) at 3065 cm−1, 3023 cm−1 indicating the aromatic C–H stretching vibrations, peaks at 1600 cm−1, 1489 cm−1, and 1449 cm−1 representing aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibrations, aromatic out-of-plane C–H bending at 758 cm−1, and the band at 699 cm−1 corresponding to aromatic ring bending. Furthermore, the peak at 1719 cm−1 indicates the carbonyl stretching frequency of the poly(HEMA–AA) shell, whereas a larger peak near 3435 cm−1 represents the O–H stretching vibration of pHEMA. The peaks at 1165 cm−1 and 1071 cm−1 indicate C–O–C antisymmetric and stretching vibrations, respectively, from pHEMA and pAA. These peaks indicate the formation of a poly(HEMA–AA) shell. XPS was used to investigate the surface chemistry of the PS@poly(HEMA–AA) microspheres. The elemental compositions of the PS and PS@poly(HEMA–AA) microspheres obtained from the XPS survey spectra are summarized in Fig. 2B. Compared to the PS microspheres, the CS microspheres exhibited increased oxygen elemental content, which could indicate the formation of a poly(HEMA–AA) hydrogel in the shell. The deconvoluted XPS C 1s narrow scan of the PS@poly(HEMA–AA) microsphere (Fig. 2C) revealed three components with binding energies of 284.8, 286.2, and 288.8 eV, corresponding to C–C/C–H, C–O, and O–C
C stretching vibrations, aromatic out-of-plane C–H bending at 758 cm−1, and the band at 699 cm−1 corresponding to aromatic ring bending. Furthermore, the peak at 1719 cm−1 indicates the carbonyl stretching frequency of the poly(HEMA–AA) shell, whereas a larger peak near 3435 cm−1 represents the O–H stretching vibration of pHEMA. The peaks at 1165 cm−1 and 1071 cm−1 indicate C–O–C antisymmetric and stretching vibrations, respectively, from pHEMA and pAA. These peaks indicate the formation of a poly(HEMA–AA) shell. XPS was used to investigate the surface chemistry of the PS@poly(HEMA–AA) microspheres. The elemental compositions of the PS and PS@poly(HEMA–AA) microspheres obtained from the XPS survey spectra are summarized in Fig. 2B. Compared to the PS microspheres, the CS microspheres exhibited increased oxygen elemental content, which could indicate the formation of a poly(HEMA–AA) hydrogel in the shell. The deconvoluted XPS C 1s narrow scan of the PS@poly(HEMA–AA) microsphere (Fig. 2C) revealed three components with binding energies of 284.8, 286.2, and 288.8 eV, corresponding to C–C/C–H, C–O, and O–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, respectively. Peaks at binding energy values of 531.3, 532.7, and 534 eV, corresponding to C
O, respectively. Peaks at binding energy values of 531.3, 532.7, and 534 eV, corresponding to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, O–H, and CO–O–C, respectively, were apparent in the deconvoluted spectrum of the XPS O 1s narrow scan (Fig. 2D). Therefore, the existence of the poly(HEMA–AA) shell was confirmed by the XPS results.
O, O–H, and CO–O–C, respectively, were apparent in the deconvoluted spectrum of the XPS O 1s narrow scan (Fig. 2D). Therefore, the existence of the poly(HEMA–AA) shell was confirmed by the XPS results.
Fabrication of PS@poly(HEMA–AA)-based stimuli-RPCs
In contrast to conventional PS microsphere-based CPC synthesis, the integration of a hydrophilic shell into PS microspheres facilitates volume phase transitions via hydrogel swelling or deswelling, making it an efficient approach for the development of stimuli-responsive photonic crystals (RPCs).41,42 Here, the rigid PS core imparts structural stability, whereas the soft poly(HEMA–AA) hydrogel shell contributes to the stimuli-responsive characteristics. PS@poly(HEMA–AA) microspheres were used to fabricate RPC on a pre-cleaned hydrophilic glass substrate via evaporation-induced vertical deposition. The UV-vis reflectance spectrum (Fig. 3A) shows the PBG position centered at 449 nm, and the inset of Fig. 3A depicts its characteristic structural color, bright violet. In Fig. 3B, the corresponding CIE plot is presented. The SEM micrograph of RPC revealed a well-ordered, closely packed arrangement of PS@poly(HEMA–AA) microspheres, as illustrated in Fig. 3C. | ||
| Fig. 3 (A) Reflectance spectra and digital photograph (inset), (B) CIE plot, and (C) SEM image of the fabricated PS@poly(HEMA–AA) RPC. | ||
Fabrication of aptasensor-based on RPC (Apt-RPC) for sensing RBD of SARS CoV-S-protein
In RPCs, stimuli-responsive hydrogels can either swell or de-swell in response to the analyte molecule, thus altering the center-to-center spacing between adjacent colloidal particles, which causes a shift in the structural color.43 This shift in the color of RPCs in response to external stimuli can be employed for the visual detection of analytes in biosensing applications. A photonic crystal-based aptasensor (Apt-RPC) was fabricated by functionalizing PS@poly(HEMA–AA) microspheres with RBD-selective single-stranded DNA aptamer via EDC-NHS coupling chemistry. The designed aptamer was directly functionalized by the carboxylic acid group present in the poly(HEMA–AA) shell. Scheme 1 depicts the covalent coupling of aptamers with the carboxyl group of poly(HEMA–AA) microspheres. The specific binding of Apt-RPC to RBD alters the periodicity of RPC. This study utilized the aptamer sequence reported by Song et al., identified through an ACE2 competition-based selection strategy and a machine learning screening algorithm that selectively binds the RBD of SARS-CoV-2 S-protein with a Kd value of 5.8 nM.44 The selected aptamer sequence was 5′-CAG CAC CGA CCT TGT GCT TTG GGA GTG CTG GTC CAA GGG CGT TAA TGG ACA-3, with 51 bases and a hairpin structure. To prepare Apt-RPC, the PS@poly(HEMA–AA) core–shell was activated with an EDC-NHS solution for 1 h at room temperature. The aptamer solution was added, incubated at room temperature for 30 min, and then refrigerated overnight at 4 °C. The functionalization of aptamers to poly(HEMA–AA) hydrogel shells via amide bonds (–CO–NH–) was confirmed using XPS analysis. The XPS survey spectrum of the Apt-RPC sample revealed that C, O, and N were the major constituents. Compared to RPC, the presence of an additional element, nitrogen, arises from the aptamer bonded to the poly(HEMA–AA) hydrogel shell. The deconvoluted XPS narrow scan of C 1s (Fig. 4A) revealed the presence of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–C, C–N, C–O, and O–C
C, C–C, C–N, C–O, and O–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds with binding energies of 283.5, 284.8, 285.1, 286.5, and 288.5 eV, respectively. The presence of a C–N bond can be attributed to the formation of an amide bond. The deconvoluted XPS narrow scan N 1s (Fig. 4B) indicates the existence of C–N, N–C
O bonds with binding energies of 283.5, 284.8, 285.1, 286.5, and 288.5 eV, respectively. The presence of a C–N bond can be attributed to the formation of an amide bond. The deconvoluted XPS narrow scan N 1s (Fig. 4B) indicates the existence of C–N, N–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and N–H with binding energies of 397.5, 398.6, and 401.8 eV, respectively. Fig. 4C presents an SEM micrograph of an ordered and close-packed arrangement of Apt-RPCs, suggesting the structural stability of the RPC sample even after aptamer functionalization. The PBG of CPC follows modified Bragg's law, where,
O, and N–H with binding energies of 397.5, 398.6, and 401.8 eV, respectively. Fig. 4C presents an SEM micrograph of an ordered and close-packed arrangement of Apt-RPCs, suggesting the structural stability of the RPC sample even after aptamer functionalization. The PBG of CPC follows modified Bragg's law, where,| neff2 = fnp2 + (1 − f)nair2 |
Furthermore, we investigated the stimuli-responsive characteristics of Apt-RPC in relation to the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein. Before conducting the sensing experiments, the Apt-RPC was rinsed with PBS to eliminate any unreacted aptamers. Different concentrations of RBD solution were prepared using 10 mM PBS buffer (pH 7.4). Approximately 30 µL of RBD solution at concentrations of 100, 500, 1000, and 1500 ng was applied to Apt-RPC, and reflectance spectra were recorded after 10 min. The corresponding reflectance spectra shown in Fig. 4E indicate a concentration-dependent spectral shift in response to the RBD. The initial Bragg diffraction wavelength of the RPC at 449 nm was red-shifted to 496, 501, and 507 nm when the concentration of RBD increased to 100, 500, 1000, and 1500 ng, respectively, resulting in an apparent color shift from violet to green. The detection range of the aptasensor was estimated to be, 100–1000 ng and no spectral difference was observed after 1000 ng. Fig. 4F illustrates the CIE plot subsequent to the addition of 1000 ng of RBD in Apt-RPC along with the corresponding visual color shift shown in the inset of Fig. 4F.
The fabricated aptasensor exhibited a rapid response to RBD, and the structure of the Apt-RPC film remained stable in the presence of RBD solution. A control experiment was conducted using non-functionalized RPC to validate the role of the aptamer in RPC binding to RBD. Fig. 5A shows the reflectance spectra of the aptamer-functionalized RPC (Apt-RPC) and non-functionalized RPC in a 1000 ng solution of RBD. The reflection spectra exhibited distinct peaks, indicating the specific interaction between RBD and the aptamer, which resulted in a noticeable red-shift of Apt-RPC in the RBD solution. Based on the above experimental results, the red-shift in the PBG of the Apt-RPC towards the RBD of the SARS-CoV-2 S-protein resulted from a network of hydrogen bonds between T42, T43, and T43 of the aptamer with Thr500, Gln506, and Asn437 from the RBD. Additionally, the conformational changes of the aptamer upon binding with the RBD lead to swelling of the Apt-RPC, resulting in increased interparticle spacing within RPCs and a red-shift.44
 | ||
| Fig. 5 (A) Reflectance spectra showing the response of Apt-RPC and RPC to 1000 ng of RBD, and (B) reflectance spectra of Apt-RPC in response to interferences such as amylase, lysozyme, and HSA. | ||
To detect SARS-CoV-2, swabs were collected from the nasal mucus and saliva. Therefore, selectivity studies were performed using a few model proteins present in nasal mucus and saliva for the practical application of the Apt-RPC sensor. Fig. 5B shows the reflection spectra of HSA, lysozyme and amylase. The HSA, lysozyme, and amylase concentrations were comparable to the range of human physiological conditions (1 mg mL−1). The PBG of Apt-RPC towards HSA at 480 nm, lysozyme at 478 nm, and amylase at 475 nm was observed in the reflection spectra. This indicates that only RBD can selectively bind to Apt-RPC and induce significant red-shift diffraction, indicating the selectivity of the sensor. The reflection spectra revealed that the PBG of the Apt-RPC for HSA at 480 nm, lysozyme at 478 nm, and amylase at 475 nm clearly demonstrate that only the RBD can selectively bind to Apt-RPC, resulting in a noticeable red-shift in diffraction, demonstrating the selectivity of the sensor.
Conclusions
In summary, we adopted a straightforward method for the synthesis and fabrication of PS@poly(HEMA–AA) core–shell microsphere-based stimuli-responsive photonic crystals as an aptasensor platform for the specific detection of the RBD of the SARS-CoV-2 S protein. The specific binding of RBD to the aptamer resulted in hydrogel swelling, accompanied by a rapid color change (1 min) and a corresponding shift in the PBG, which enabled the detection of the RBD of the SARS-CoV-2 S-protein. The CIE plot showed a rapid visual color shift from violet to green upon the addition of RBD of SARS-CoV-2-S-protein to the Apt-RPC aptasensor, with a detection range of, 100–1000 ng. Compared to other COVID-19 detection methods, the developed biosensor uses a simple and facile approach for the visual detection of SARS-CoV-2 S-protein. We demonstrate this method as a qualitative approach for pathogen detection through the integration of a polymer-based aptasensor. Furthermore, the ability to modulate the PBG across the visible spectrum enhances the potential applications of this stimuli-responsive photonic crystal-based aptasensor in future biosensing technologies.Experimental section
Materials and reagents
Styrene, 2-hydroxyethyl methacrylate (HEMA), acrylic acid (AA), ethylene glycol dimethacrylate (EGDMA), potassium persulfate (KPS), human serum albumin, lysozyme, amylase, 1-ethyl-3-(3-dimethyl aminopropyl)carbodiimide (EDC), and N-hydroxysuccinimide (NHS) were purchased from Sigma-Aldrich. All reagents were of analytical grade and used as received, without further purification. Single-stranded-DNA aptamer for the selective binding of RBD of SARS-CoV-2 (5′-CAG CAC CGA CCT TGT GCT TTG GGA GTG CTG GTC CAA GGG CGT TAA TGG ACA-3′) was purchased from Integrated DNA Technologies (IDT, India). All experiments were performed using ultrapure water (18.2 MΩ cm, Merck) from a Milli-Q ultrafiltration system.Synthesis of core–shell microspheres and fabrication of CPCs
Monodispersed core–shell (CS) microspheres of polystyrene@poly(hydroxyethyl methacrylate–acrylic acid) [PS@poly(HEMA–AA)] were synthesized via emulsifier-free emulsion polymerization. Briefly, 5.07 mmol of HEMA, 4.5 mmol of AA, 57.5 mmol of styrene, and 0.53 mmol of EGDMA, which acted as a cross-linker, were mixed with 100 mL deionized water, placed in a round-bottomed flask, and placed in an oil bath at 70 °C. After 30 min, 0.07 mmol of potassium persulfate (KPS) (initiator) was introduced into the reaction mixture. Polymerization was carried out under a N2 gas atmosphere with gentle magnetic stirring at 75 °C for 10 h. The resultant colloidal suspension was purified via centrifugation at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 30 min, and the purified CS microspheres were used to fabricate CPCs using the vertical deposition approach on glass substrates. Prior to deposition, the glass substrates were cleaned with ethanol, followed by N2 plasma treatment to generate hydrophilic surfaces.
000 rpm for 30 min, and the purified CS microspheres were used to fabricate CPCs using the vertical deposition approach on glass substrates. Prior to deposition, the glass substrates were cleaned with ethanol, followed by N2 plasma treatment to generate hydrophilic surfaces.
Fabrication of aptasensor based on RPC (Apt-RPC)
The functionalization of the aptamer on RPC was carried out using EDC/NHS coupling chemistry. The carboxyl functional groups on the CS microspheres of RPC were activated with EDC/NHS using 1 mL of phosphate-buffered saline (PBS, pH 7.4) containing 0.5 mM EDC and 0.5 mM NHS for an hour at ambient temperature. To the activated RPC, 30 µL of 500 nM aptamer in 2 mM PBS buffer solution was added, incubated at room temperature for 30 min, and stored overnight at 4 °C. The synthesized Apt-RPC sensor was subsequently rinsed with PBS to remove unreacted aptamers and used for subsequent investigations. Various quantities of RBD solutions in 10 mM PBS (pH 7.4) were introduced into the Apt-RPC sensor, and the reflectance spectra were recorded.Characterizations
The morphology and dimensions of the CS microspheres were analyzed using transmission electron microscopy (TEM; JEOL JEM F 200). The hydrodynamic diameter was obtained using dynamic light scattering (DLS, Zetasizer Nano ZS, Malvern Instruments). Chemical functionalization studies were conducted using an FT-IR spectrometer (PerkinElmer) and X-ray photoelectron spectroscopy (XPS, PHI 5000 Versa Probe II ULVAC-PHI Inc., USA). Scanning electron microscopy (SEM) was performed using a Zeiss EVO 18 SEM (Germany). The reflectance spectra of the CPCs were obtained using a modular fiber-optic spectrophotometer (Ocean Optics DH-2000-BAL). We adopted the CIE (International Commission on Illumination) color space chromaticity diagram to validate the visual colors of the CPCs.Conflicts of interest
The authors declare no conflicts of interest.Data availability
All data supporting this study are included in this article. Additional data are available from the corresponding author upon reasonable request.Acknowledgements
The authors thank Mr Kiran Mohan and Mr Harish Raj V. for the TEM and SEM acquisitions, respectively. MMT, PRC, and LL thank CSIR-SRF, SERB-National Post-doctoral Fellowship, and DST-INSPIRE fellowship, respectively, for financial support. This work was supported by the Council of Scientific and Industrial Research (CSIR, India).References
- F. Wu, S. Zhao, B. Yu, Y. M. Chen, W. Wang, Z. G. Song, Y. Hu, Z. W. Tao, J. H. Tian, Y. Y. Pei, M. L. Yuan, Y. L. Zhang, F. H. Dai, Y. Liu, Q. M. Wang, J. J. Zheng, L. Xu, E. C. Holmes and Y. Z. Zhang, Nature, 2020, 579, 265–269 CrossRef CAS PubMed.
- L. A. Jackson, E. J. Anderson, N. G. Rouphael, P. C. Roberts, M. Makhene, R. N. Coler, M. P. McCullough, J. D. Chappell, M. R. Denison, L. J. Stevens, A. J. Pruijssers, A. McDermott, B. Flach, N. A. Doria-Rose, K. S. Corbett, K. M. Morabito, S. O'Dell, S. D. Schmidt, P. A. Swanson, M. Padilla, J. R. Mascola, K. M. Neuzil, H. Bennett, W. Sun, E. Peters, M. Makowski, J. Albert, K. Cross, W. Buchanan, R. Pikaart-Tautges, J. E. Ledgerwood, B. S. Graham and J. H. Beigel, N. Engl. J. Med., 2020, 383, 1920–1931 CrossRef CAS PubMed.
- P. Zhou, X. L. Yang, X. G. Wang, B. Hu, L. Zhang, W. Zhang, H. R. Si, Y. Zhu, B. Li, C. L. Huang, H. D. Chen, J. Chen, Y. Luo, H. Guo, R. Di Jiang, M. Q. Liu, Y. Chen, X. R. Shen, X. Wang, X. S. Zheng, K. Zhao, Q. J. Chen, F. Deng, L. L. Liu, B. Yan, F. X. Zhan, Y. Y. Wang, G. F. Xiao and Z. L. Shi, Nature, 2020, 579, 270–273 CrossRef CAS PubMed.
- G. Bonaccorsi, F. Pierri, M. Cinelli, F. Porcelli, A. Galeazzi, A. Flori, A. L. Schmidt, C. M. Valensise, A. Scala, W. Quattrociocchi and F. Pammolli, Sustainability, 2022, 14, 9699 CrossRef.
- E. C. Holmes, Annu. Rev. Virol., 2024, 11, 21–42 CrossRef CAS PubMed.
- S. Steiner, A. Kratzel, G. T. Barut, R. M. Lang, E. A. Moreira, L. Thomann, J. N. Kelly and V. Thiel, Nat. Rev. Microbiol., 2024, 22, 206–225 CrossRef CAS PubMed.
- U. Lele and S. Goswami, Med. Res. Arch., 2024, 12(2) DOI:10.18103/MRA.V12I2.5039.
- J. Y. Choi and D. M. Smith, Yonsei Med. J., 2021, 62, 961 CrossRef CAS PubMed.
- M. Alexandridi, J. Mazej, E. Palermo and J. Hiscott, Cytokine Growth Factor Rev., 2022, 63, 1–9 CrossRef CAS PubMed.
- K. Sun, J. N. Bhiman, S. Tempia, J. Kleynhans, V. S. Madzorera, Q. Mkhize, H. Kaldine, M. L. McMorrow, N. Wolter, J. Moyes, M. Carrim, N. A. Martinson, K. Kahn, L. Lebina, J. D. du Toit, T. Mkhencele, A. von Gottberg, C. Viboud, P. L. Moore, C. Cohen, A. Buys, M. Carrim, L. de Gouveia, M. du Plessis, J. du Toit, F. X. Gómez-Olivé, K. P. Kgasago, R. Kotane, M. L. McMorrow, T. Moloantoa, S. Tollman, A. von Gottberg, F. Wafawanaka and N. Wolter, Nat. Med., 2024, 30, 2805–2812 CrossRef CAS PubMed.
- R. Liu, L. He, Y. Hu, Z. Luo and J. Zhang, Chem. Sci., 2020, 11, 12157–12164 RSC.
- M. L. Wong and J. F. Medrano, Biotechniques, 2005, 39, 75–85 CrossRef CAS PubMed.
- Q. Lin, D. Wen, J. Wu, L. Liu, W. Wu, X. Fang and J. Kong, Anal. Chem., 2020, 92, 9454–9458 CrossRef CAS PubMed.
- J. Huang, J. Wen, M. Zhou, S. Ni, W. Le, G. Chen, L. Wei, Y. Zeng, D. Qi, M. Pan, J. Xu, Y. Wu, Z. Li, Y. Feng, Z. Zhao, Z. He, B. Li, S. Zhao, B. Zhang, P. Xue, S. He, K. Fang, Y. Zhao and K. Du, Anal. Chem., 2021, 93, 9174–9182 CrossRef CAS PubMed.
- Y. Huang, C. Yang, X. f. Xu, W. Xu and S. w. Liu, Acta Pharmacol. Sin., 2020, 41, 1141–1149 CrossRef PubMed.
- R. Yan, Y. Zhang, Y. Li, L. Xia, Y. Guo and Q. Zhou, Science, 2020, 367, 1444–1448 CrossRef CAS PubMed.
- P. Dhar, R. M. Samarasinghe and S. Shigdar, Int. J. Mol. Sci., 2020, 21, 2485 CrossRef CAS PubMed.
- B. Ju, Q. Zhang, J. Ge, R. Wang, J. Sun, X. Ge, J. Yu, S. Shan, B. Zhou, S. Song, X. Tang, J. Yu, J. Lan, J. Yuan, H. Wang, J. Zhao, S. Zhang, Y. Wang, X. Shi, L. Liu, J. Zhao, X. Wang, Z. Zhang and L. Zhang, Nature, 2020, 584, 115–119 CrossRef CAS PubMed.
- G. Yang, Z. Li, I. Mohammed, L. Zhao, W. Wei, H. Xiao, W. Guo, Y. Zhao, F. Qu and Y. Huang, Signal Transduction Targeted Ther., 2021, 6, 1–4 Search PubMed.
- M. Sun, S. Liu, X. Wei, S. Wan, M. Huang, T. Song, Y. Lu, X. Weng, Z. Lin, H. Chen, Y. Song and C. Yang, Angew. Chem., Int. Ed., 2021, 60, 10266–10272 CrossRef CAS PubMed.
- L. Zhang, X. Fang, X. Liu, H. Ou, H. Zhang, J. Wang, Q. Li, H. Cheng, W. Zhang and Z. Luo, Chem. Commun., 2020, 56, 10235–10238 RSC.
- Z. Wang, S. Liu, Z. Shi, D. Lu, Z. Li and Z. Zhu, Talanta, 2024, 277, 126318 CrossRef CAS.
- L. Wei, D. Zhu, Q. Cheng, Z. Gao, H. Wang and J. Qiu, Food Res. Int., 2024, 179, 114005 CrossRef CAS PubMed.
- F. Li, J. Lu, J. Liu, C. Liang, M. Wang, L. Wang, D. Li, H. Yao, Q. Zhang, J. Wen, Z. K. Zhang, J. Li, Q. Lv, X. He, B. Guo, D. Guan, Y. Yu, L. Dang, X. Wu, Y. Li, G. Chen, F. Jiang, S. Sun, B. T. Zhang, A. Lu and G. Zhang, Nat. Commun., 2017, 8, 1–14 CrossRef PubMed.
- Q. Liu, W. Zhang, S. Chen, Z. Zhuang, Y. Zhang, L. Jiang and J. S. Lin, J. Biol. Eng., 2020, 14, 1–13 CrossRef CAS.
- Z. Li, X. Zhou, J. Yang, B. Fu and Z. Zhang, ACS Appl. Mater. Interfaces, 2019, 11, 21417–21423 CrossRef CAS PubMed.
- P. Shen, W. Li, Y. Liu, Z. Ding, Y. Deng, X. Zhu, Y. Jin, Y. Li, J. Li and T. Zheng, Anal. Chem., 2017, 89, 11862–11868 CrossRef CAS PubMed.
- G. Isapour and M. Lattuada, Adv. Mater., 2018, 30, 1707069 CrossRef PubMed.
- Y. Fang, B. M. Phillips, K. Askar, B. Choi, P. Jiang and B. Jiang, J. Mater. Chem. C, 2013, 1, 6031–6047 RSC.
- M. Jia, L. Yu, X. Li, Y. Li, X. He, L. Chen and Y. Zhang, Talanta, 2023, 260, 124638 CrossRef CAS PubMed.
- I. Kalsoom, K. Shehzadi, M. Irfan, L. Qiu, Y. Wang, Z. Xu and Z. Meng, Biosens. Bioelectron., 2025, 273, 117091 CrossRef CAS PubMed.
- C. Fenzl, T. Hirsch and O. S. Wolfbeis, Angew. Chem., Int. Ed., 2014, 53, 3318–3335 CrossRef CAS.
- N. Griffete, H. Frederich, A. Maître, S. Ravaine, M. M. Chehimi and C. Mangeney, Langmuir, 2012, 28, 1005–1012 CrossRef CAS PubMed.
- M. M. Thomas, P. R. Chandran, V. V. Vipin, A. P. Mohamed, P. Kingshott and S. Pillai, React. Funct. Polym., 2021, 158, 104779 CrossRef CAS.
- G. Hwang, G. Bak, Y. Kim, S. H. Jung, H. Na and Y. J. Jung, ACS Omega, 2025, 10, 36582–36591 CrossRef CAS.
- G. Feng, K. Li, G. Li, Z. Zhang, J. Xiang, T. Chen, F. Jiao and H. Zhao, ACS Appl. Polym. Mater., 2025, 7(7), 4270–4278 CrossRef CAS.
- B. F. Ye, Y. J. Zhao, Y. Cheng, T. T. Li, Z. Y. Xie, X. W. Zhao and Z. Z. Gu, Nanoscale, 2012, 4, 5998–6003 RSC.
- H. Xuan, J. Ren, Y. Zhu, B. Zhao and L. Ge, RSC Adv., 2016, 6, 36827–36833 RSC.
- G. Murtaza, A. S. Rizvi, M. Xue, L. Qiu and Z. Meng, Anal. Chem., 2023, 95(2), 917–923 Search PubMed.
- P. A. Lovell and F. J. Schork, Biomacromolecules, 2020, 21, 4396–4441 CrossRef CAS PubMed.
- W. Wang, Y. Zhou, L. Yang, X. Yang, Y. Yao, Y. Meng and B. Tang, Adv. Funct. Mater., 2022, 32(40), 2204744 CrossRef CAS.
- L. Hu, Y. Wan, Q. Zhang and M. J. Serpe, Adv. Funct. Mater., 2020, 30, 1903471 CrossRef CAS.
- X. Jia, T. Zhang, J. Wang, K. Wang, H. Tan, Y. Hu, L. Zhang and J. Zhu, Langmuir, 2018, 34, 3987–3992 CrossRef CAS PubMed.
- Y. Song, J. Song, X. Wei, M. Huang, M. Sun, L. Zhu, B. Lin, H. Shen, Z. Zhu and C. Yang, Anal. Chem., 2020, 92, 9895–9900 CrossRef CAS PubMed.
- F. Wang, Z. Meng, F. Xue, M. Xue, W. Lu, W. Chen, Q. Wang and Y. Wang, Trends Environ. Anal. Chem., 2014, 3–4, 1–6 CAS.
| This journal is © The Royal Society of Chemistry 2026 |