Open Access Article
Open Access ArticleOptimization of phenolics extraction and isolation from gymnosperms growing in Egypt using response surface methodology: Dioon edule Lindl leaves as a case study
Asmaa M. Akara,
Nesrine M. Hegazi b,
Ahmed Zayed
b,
Ahmed Zayed *a and
Souzan M. Ibrahima
*a and
Souzan M. Ibrahima
aDepartment of Pharmacognosy, College of Pharmacy, Tanta University, El-Guish Street (Medical Campus), Tanta 31527, Egypt. E-mail: asmaa.akar@pharm.tanta.edu.eg; ahmed.zayed1@pharm.tanta.edu.eg; suzan.mostafa@pharm.tanta.edu.eg
bDepartment of Phytochemistry and Plant Systematics, Institute of Pharmaceutical Industries, National Research Centre, Dokki, Cairo 12622, Egypt. E-mail: nm.hegazi@nrc.sci.eg
First published on 24th March 2026
Abstract
Gymnosperms are recognized as rich sources of phenolic constituents, particularly biflavonoids, which exhibit a wide range of biological activities, including antioxidant, anti-inflammatory, and anticancer effects. However, conventional extraction techniques such as maceration and Soxhlet extraction are often limited by prolonged extraction times and high solvent consumption. In this study, traditional extraction methods were systematically compared with modern techniques, namely microwave-assisted extraction (MAE) and ultrasound-assisted extraction (UAE), for the recovery of phenolic compounds from the leaves of Dioon edule Lindl. (D. edule) cultivated in Egypt. Among the evaluated methods, UAE afforded the highest total phenolic content (11.28 mg gallic acid equivalents GAE g−1 dry plant matter (DM)), total flavonoid content (8.56 mg rutin equivalents RE g−1 DM), and antioxidant activity (37.1% DPPH radical scavenging activity, IC50 = 151.25 µg mL−1). Based on these findings, the UAE was further optimized using response surface methodology (RSM), considering temperature, extraction time, and solvent-to-plant material ratio as critical variables. Under the optimized conditions (80 °C, 100 min, 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mL g−1), the extraction yield increased by 27%, while phenolic and flavonoid contents increased by 15% and 33%, respectively. Antioxidant activity was significantly enhanced, with DPPH inhibition increasing to 58.2% and IC50 decreasing to 75.6 µg mL−1. In addition, antibacterial and cytotoxic activity assays consistently supported the superiority of UAE. Chromatographic separation of the optimized UAE extract led to the isolation of five bioactive biflavonoids, identified as 7,4′,7′,4‴-tetra-O-methylamentoflavone (1, isolated from dichloromethane fraction (DCM)) and total methanolic extract), isoginkgetin (2, DCM fraction), sciadopitysin (3, ethyl acetate (EtOAc) fraction), amentoflavone (4, EtOAc fraction), and kayaflavone (5, total methanolic fraction). Structural elucidation was achieved through comprehensive spectroscopic analyses and comparison with reported data. These compounds were obtained in higher yields compared with previous reports based on conventional extraction methods. Therefore, the study provides a systematic evaluation and statistical optimization of UAE for phenolic recovery from D. edule, coupled with phytochemical profiling and preliminary biological assessment. The findings indicate that optimized UAE conditions enhance extraction efficiency and phenolic recovery relative to the tested conventional approaches.
1 mL g−1), the extraction yield increased by 27%, while phenolic and flavonoid contents increased by 15% and 33%, respectively. Antioxidant activity was significantly enhanced, with DPPH inhibition increasing to 58.2% and IC50 decreasing to 75.6 µg mL−1. In addition, antibacterial and cytotoxic activity assays consistently supported the superiority of UAE. Chromatographic separation of the optimized UAE extract led to the isolation of five bioactive biflavonoids, identified as 7,4′,7′,4‴-tetra-O-methylamentoflavone (1, isolated from dichloromethane fraction (DCM)) and total methanolic extract), isoginkgetin (2, DCM fraction), sciadopitysin (3, ethyl acetate (EtOAc) fraction), amentoflavone (4, EtOAc fraction), and kayaflavone (5, total methanolic fraction). Structural elucidation was achieved through comprehensive spectroscopic analyses and comparison with reported data. These compounds were obtained in higher yields compared with previous reports based on conventional extraction methods. Therefore, the study provides a systematic evaluation and statistical optimization of UAE for phenolic recovery from D. edule, coupled with phytochemical profiling and preliminary biological assessment. The findings indicate that optimized UAE conditions enhance extraction efficiency and phenolic recovery relative to the tested conventional approaches.
1. Introduction
Phenolic compounds play an essential role in the prevention and management of chronic diseases, such as cardiovascular diseases, diabetes, neurodegenerative disorders and certain cancers.1 The extraction process represents a critical initial step in the isolation and recovery of these bioactive compounds from plant matrices.2 Conventional extraction techniques, such as solvent extraction, Soxhlet extraction, hydrodistillation, and pressing methods, are widely applied; however, they are often associated with several drawbacks, including long extraction times, high solvent consumption, degradation of thermolabile compounds, low extraction efficiency, and potential solvent residues in the final extract.3In recent years, increasing attention has been directed toward alternative extraction approaches that enhance efficiency while reducing processing time. Among these, ultrasound-assisted extraction (UAE), microwave-assisted extraction (MAE), and supercritical fluid extraction (SFE) have emerged as effective methodologies for improving mass transfer and extraction performance.4
MAE is an efficient technique used to enhance the extraction of bioactive compounds from medicinal plants by utilizing microwave energy to heat solvents and plant materials. This method accelerates the extraction process, often resulting in higher yields and reduced extraction times compared with conventional methods. For instance, a study on Panax quinquefolius L. showed that MAE increased the extraction yields of the nine rare ginsenosides significantly compared with the conventional method (as the extraction yields with MAE were at least 10 times higher than those with heat reflux).5
Similarly, the UAE has gained substantial interest as an efficient and sustainable extraction technique. The enhanced extraction efficiency of UAE is primarily attributed to acoustic cavitation, which leads to cell wall disruption and facilitates the release of intracellular compounds into the extraction solvent. Compared to traditional methods, UAE offers several advantages, including reduced extraction time, lower solvent consumption, and improved recovery of phenolic compounds. For instance, UAE (26 kHz, 200 W) achieved a polyphenol yield of 4016 mg gallic acid equivalents GAE 100 g−1 DM from saffron floral biomass, surpassing MAE (800 W), which yielded 3108 mg GAE 100 g−1 DM.6
Despite these advantages, the efficiency of both MAE and UAE is highly dependent on operational parameters such as extraction time, temperature, power, solvent composition, and solvent-to-solid ratio.7,8 Therefore, systematic optimization is essential to maximize extraction efficiency while preserving compound stability.
Response surface methodology (RSM) represents a robust statistical tool for modeling and optimizing multivariable systems. By evaluating the interactions between selected independent variables, RSM enables the development of predictive models while minimizing experimental runs.9,10
Gymnosperms represent an evolutionarily ancient group of seed plants that differ structurally and chemically from angiosperms. Several gymnosperm families, particularly Cycadaceae and Zamiaceae, are recognized as rich sources of phenolic compounds, especially flavonoids and biflavonoids, which exhibit diverse biological activities including antioxidant, antimicrobial, anti-inflammatory, and anticancer effects.11–13 Interestingly, amentoflavone has shown various biological and pharmacological effects, including antioxidant, anti-cancer, anti-inflammatory, antimicrobial, and antiviral.14 In addition, Bilobetin, sciadopitysin and 7,4′,7″4‴ tetra-O-methyl amentoflavone enhance differentiation osteoblast suggesting their therapeutic potential in osteoporosis.15
Although UAE combined with RSM has been widely applied in angiosperm species, fewer studies have systematically evaluated extraction strategies and statistical optimization in biflavonoid-rich gymnosperms. Moreover, the influence of optimized extraction conditions on both phenolic recovery and the isolation profile of major biflavonoids has not been extensively explored in Dioon edule Lindl.
Therefore, the present study aimed to provide a systematic comparison between conventional extraction techniques (maceration and Soxhlet) and modern approaches (MAE and UAE) for phenolic recovery from D. edule Lindl. leaves cultivated in Egypt. The most efficient method was further optimized using RSM to evaluate the effects of temperature, extraction time, and solvent-to-solid ratio. In addition, the optimized extract was subjected to phytochemical investigation to characterize the major biflavonoids and assess the impact of optimized conditions on their recovery. This approach integrates extraction comparison, statistical modeling, and phytochemical profiling to better understand phenolic extraction behavior in a gymnosperm matrix.
2. Materials, apparatus, and methodology
2.1. Materials and chemicals
All solvents, including absolute methanol (MeOH), dichloromethane (DCM, CH2Cl2), and ethyl acetate (EtOAc), were of analytical grade. Sodium carbonate (Na2CO3), sodium nitrite (NaNO2), and sodium hydroxide (NaOH) were also of analytical grade. Aluminum chloride (AlCl3), Folin–Ciocalteu reagent, 1,1-diphenyl-2-picrylhydrazyl (DPPH), gallic acid, rutin, dimethylsulfoxide (DMSO), MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide), ciprofloxacin, doxorubicin, and sorafenib were purchased from Sigma-Aldrich Chemical Co. (St. Louis, MO, USA). Ascorbic acid was obtained from Rasayan Laboratories (OPC) Pvt. Ltd (India).Fetal bovine serum (FBS) and RPMI-1640 medium were obtained from GIBCO (UK). The mammary gland breast cancer cell line (MCF-7) was procured from the American Type Culture Collection (ATCC) through the Holding Company for Biological Products and Vaccines (VACSERA), Cairo, Egypt.
Microbial strains, including Staphylococcus aureus (ATCC 6538) and Pseudomonas aeruginosa (ATCC 27853) were obtained from ATCC, Silica gel 60 and precoated thin-layer chromatography (TLC) silica gel sheets G F254 were purchased from E. Merck (now Merck KGaA, Darmstadt, Germany), while Sephadex LH-20 (Sigma-Aldrich Chemical Co., USA) and α-naphthol (Sigma Chemical Co., St. Louis, USA) for Molisch’s reagent.
In addition, the authentic compounds used in this study were kindly provided by Dr Walaa Negm (Department of Pharmacognosy, Faculty of Pharmacy, Tanta University, Egypt). These compounds had been previously isolated and identified in her PhD work using chromatographic and spectroscopic analyses (1H-NMR, 13C-NMR, and MS), and their identities were confirmed by comparison with reported literature data.16
2.2. Equipment and apparatus
A Soxhlet extraction apparatus, a sonication water bath (Transsonic T 460H, Elma, Germany), a microwave extraction system (MARS, CEM, California, USA), and a rotary evaporator (IKA RV10, Wilmington, NC, USA) were used for extraction and solvent removal. Absorbance measurements were performed using a UV-visible spectrophotometer (UV-1800, Shimadzu, Japan) and a microplate reader (Anthos Zenyth 200RT, Harvard Bioscience, MA, USA). Thin-layer chromatography (TLC) analysis was carried out using a UV cabinet (ENF-260C/FE, Spectroline, Spectronics Corporation, Westbury, NY, USA).Infrared spectra were recorded using an FT-IR spectrometer (FT/IR-6100, Jasco, Japan). Nuclear magnetic resonance (NMR) spectra were obtained using Bruker spectrometers operating at 400 and 600 MHz. In addition, HRMS measurements were carried out on a Waters Acquity UPLC coupled to a Waters QTof Premier mass spectrometer.
Analytical LC-MS analyses were conducted on a Waters system equipped with a 2767 autosampler, a 2545 pump, and a Phenomenex Kinetex C18 column (2.6 µm, 100 Å, 4.6 × 100 mm) fitted with a Phenomenex Security Guard precolumn (Luna C5, 300 Å). The mobile phase was delivered at a flow rate of 1.0 mL min−1 using a 15 min linear gradient from 10% to 90% acetonitrile (0.045% formic acid) in water (0.05% formic acid). Detection was carried out using a Waters 2998 diode array detector (210–600 nm), a Waters 2424 evaporative light scattering detector (ELSD), and a Waters SQD-2 mass detector operating in both ES(+) and ES(−) modes (m/z 100–1000).
2.3. Plant collection and identification
D. edule Lindl. was collected from El Abd Garden at 68 km from the desert Cairo-Alexandria Road, Egypt, in August 2022. The plant specimen was kindly identified by Dr Esraa E. Ammar, lecturer of plant Ecology, Botany department, Faculty of Science, Tanta University. A plant specimen has been archived at the Department of Pharmacy, Tanta University with a voucher number PGG-016 for D. edule Lindl. The leaves of the plant were dried at room temperature (25 ± 2.0 °C), followed by one week in an oven at 40 °C, then reduced to a fine powder, and finally stored in a tightly closed amber colored containers in a refrigerator (4 °C) until further investigations.2.4. Plant extractions
2.4.1.1. Maceration. The dried powdered leaves of D. edule Lindl. (300 g) were extracted by cold maceration with MeOH (5 × 1.5 L, each for 48 h) at room temperature. The combined methanolic extracts were concentrated under reduced pressure at 40 °C using a rotary evaporator until dryness. The experiment was repeated twice to confirm reproducibility under the same extraction conditions.17
2.4.1.2. Soxhlet. Another portion of the dried powdered leaves of D. edule Lindl. (100 g) was extracted using a Soxhlet apparatus with MeOH (500 mL) until exhaustion at 70 °C. The methanolic extract was then concentrated under reduced pressure at 40 °C using a rotary evaporator until dryness. The experiment was repeated twice to verify reproducibility under the same conditions.18
2.4.2.1. UAE. The dried powdered leaves of D. edule Lindl. (10 g) were extracted for 2 h with 500 mL of 80% v/v MeOH at room temperature using a sonication water bath. The resulting suspension was filtered, and the residue was re-extracted under identical conditions. The experiment was repeated twice to confirm reproducibility under the same experimental parameters.15
2.4.2.2. MAE. This extraction was carried out using an open-system microwave apparatus (MARS, 240 V/50 Hz). The dried powdered leaves of D. edule Lindl. (10 g) were extracted with 200 mL of MeOH for 5 min at a power of 800 W. The extraction temperature was maintained at 65–70 °C, taking into account the boiling point of methanol (64.7 °C).19
2.5. Sample preparation for phytochemical and biological assays
All extracts obtained from different extraction techniques and optimized UAE conditions were dried and stored at 4 °C until analysis. For phytochemical and biological evaluation, each extract was accurately weighed and reconstituted in DMSO to prepare a stock solution (1 mg mL−1). Serial dilutions were freshly prepared when required. All assays, including TPC, TFC, antioxidant, antibacterial, and cytotoxic activities, were performed under identical experimental conditions to ensure reliable comparison, reproducibility, and proper data normalization among the different samples.2.6. Estimation of the total flavonoid content (TFC)
TFC was determined using the colorimetric method described by20 with some modifications. About 25 µL of the extract (the residue was reconstituted in DMSO (1 mg mL−1) was mixed with 100 µL of distilled water followed by the addition of 10 µL of 5% NaNO2 solution. After 6 min, 15 µL of a 10% AlCl3 solution was added and allowed to stand for another 5 min before 50 µL of 4% NaOH solution and 50 µL distilled water were added. The absorbance was recorded immediately at 510 nm using a microplate reader. A calibration curve was constructed using rutin (300, 250, 200, 150, 100, and 50 µg mL−1) (Fig. S1). Results were expressed as mg rutin equivalent per g dried plant Material (mg RE g−1 DM).2.7. Estimation of the total phenolic content (TPC)
TPC was determined using Folin–Ciocalteu reagent as previously described20 with some modifications. About 10 µL of extract, after reconstitution of the residue in DMSO at 1 mg mL−1, were mixed with 40 µL of Folin–Ciocalteu reagent, previously diluted 10-fold with distilled water, and allowed to stand at room temperature for 5 min; 50 µL of 20% sodium carbonate solution were added to the mixture. After 90 min at room temperature, absorbance was measured at 750 nm using a microplate reader. A calibration curve was constructed using Gallic acid (300, 250, 200, 150, 100, and 50 µg mL−1) (Fig. S2). Results were expressed as mg gallic acid equivalent per g dried plant material (mg GAE g−1 DM).2.8. Antioxidant activity
The free radical scavenging activity of plant extracts was measured by DPPH˙ using the method described by ref. 21 and 22 with some modifications. Briefly, 1 mL of 0.1 mM solution of DPPH˙ in MeOH was added to 3 mL of extract solution at different serial concentrations (12.5–100 µg mL−1). The mixture was shaken vigorously and allowed to stand at room temperature for 30 min. The absorbance was measured at 517 nm afterwards in microplate reader. Lower absorbance of the reaction mixture indicated higher free radical scavenging activity. The radical scavenging activity was measured using eqn (1).| DPPH radical scavenging effect (%) = 100 − [((A0 − A1)/A0) × 100] | (1) |
2.9. Antibacterial activity assay
The antibacterial activity of D. edule Lindl. Extracts obtained using four different extraction techniques were tested against Gram-positive bacteria (Staphylococcus aureus) and Gram-negative bacteria (Pseudomonas aeruginosa) using the method described by reference with some modifications.23 Each extract was dissolved in DMSO to prepare a solution with a concentration of 1 mg mL−1. Sterile paper discs (5 mm diameter, Whatman filter paper) were prepared and sterilized by autoclaving. The discs were soaked in the desired concentration of the extract solution and aseptically placed in Petri dishes containing nutrient agar medium (20 g agar, 3 g beef extract, and 5 g peptone) seeded with S. aureus and P. aeruginosa. The Petri dishes were incubated at 36 °C for 24 h, and the inhibition zones were recorded. Each extract was tested in triplicate. The antibacterial activity of a standard antibiotic (Ciprofloxacin) was also evaluated using the same procedure, concentration, and solvent. The percentage activity index of the extracts was calculated using eqn (2).
 | (2) |
2.10. Cytotoxic activity
The cytotoxic activity was carried out according to the reported procedures for the four different extracts of D. edule Lindl. against selected cell line using the MTT colorimetric assay.24 The assay was carried out using seven different concentrations (1.56, 3.125, 6.25, 12.5, 25, 50 and 100 µg mL−1) of the total methanol extracts of D. edule Lindl. against the MCF-7 cell line. This assay is based on the reduction of the yellow tetrazolium salt (MTT) to an insoluble purple formazan product by the mitochondrial succinate dehydrogenase enzyme present in metabolically active cells. The cell line was cultured in RPMI-1640 medium with 10% fetal bovine serum. Antibiotics added were 100 units mL−1 penicillin and 100 µg mL−1 streptomycin at 37 °C in a 5% CO2 incubator. The cell lines were seeded in a 96-well plate at a density of 1.0 × 104 cells well−1. At 37 °C for 48 h under 5% CO2. After incubation, the cells were treated with different concentrations of extracts and incubated for 24 h. Afterwards, 20 µL of MTT solution at 5 mg mL−1 was added and incubated for 4 h. DMSO in a volume of 100 µL was added into each well to dissolve the purple formazan formed. The colorimetric assay is measured and recorded at an absorbance of 570 nm using a microplate reader. Doxorubicin and Sorafenib were used as standard anticancer drugs for comparison. The percentage of viable cells was determined according to eqn (3).
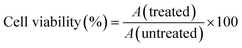 | (3) |
The percentage of cell viability obtained at different extract concentrations was plotted against the corresponding logarithmic concentrations to determine the concentration of the extract required to inhibit cell viability by 50% (IC50) and the cytotoxic effect was assessed according to the classification of Hossan and Abu Melha.25
2.11. Extraction optimization
Experimental design and optimization were performed using Design-Expert software (Version 11, Stat-Ease Inc., Minneapolis, MN, USA). A three-factor, three-level Box–Behnken response surface methodology (BBD-RSM) was employed to optimize the extraction conditions of phenolic compounds from D. edule Lindl. The selected independent variables were extraction temperature (A; °C), extraction time (B; min), and solvent-to-plant material ratio (C; mL g−1). The ranges investigated of these parameters were chosen based on previously published studies concerning the extraction of phenolic compounds from various plant materials.26–28 Three variation levels were considered (Table 1). % Yield, TPC, and TFC were set as the responses.| Factors | Symbols | Coded levels | ||
|---|---|---|---|---|
| −1 | 0 | +1 | ||
| Extraction temperature (°C) | A | 20 | 50 | 80 |
| Extraction time (min) | B | 30 | 75 | 120 |
| Solvent-to-plant material ratio (mL g−1) | C | 30 | 40 | 50 |
A total of 17 experimental runs, including five replicates at the center point (Table 2), were performed to develop a predictive model and optimize the extraction of phenolic compounds. The experimental data obtained were fitted to a second-order polynomial regression equation (eqn (4)):
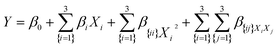 | (4) |
| Actual variables | Observed values | ||||
|---|---|---|---|---|---|
| Extraction temperature (A; °C) | Extraction time (B; min) | Solvent-to-plant material ratio (C; mL g−1) | % Yield | TPCa | TFCb |
| a TPC was expressed as mg GAE g−1 DM.b TFC was expressed as mg RE g−1 DM. | |||||
| 80 | 30 | 40 | 11.84 | 8.42 | 7.09 |
| 50 | 75 | 40 | 11.88 | 8.313 | 6.85 |
| 50 | 75 | 40 | 11.94 | 8.44 | 7.03 |
| 20 | 30 | 40 | 2.84 | 1.836 | 1.22 |
| 20 | 120 | 40 | 6.92 | 4.488 | 3.12 |
| 50 | 75 | 40 | 11.94 | 8.211 | 6.48 |
| 50 | 120 | 50 | 13.88 | 9.614 | 7.7 |
| 80 | 75 | 50 | 17.42 | 12.954 | 11.8 |
| 50 | 30 | 30 | 7.6 | 4.998 | 3.55 |
| 80 | 120 | 40 | 14.28 | 10.251 | 8.8 |
| 50 | 30 | 50 | 10.08 | 6.707 | 4.95 |
| 20 | 75 | 50 | 7 | 4.539 | 3.179 |
| 50 | 120 | 30 | 11.54 | 7.395 | 5.44 |
| 50 | 75 | 40 | 11.28 | 7.905 | 6.48 |
| 50 | 75 | 40 | 10.46 | 7.344 | 6.05 |
| 80 | 75 | 30 | 11.48 | 7.982 | 6.48 |
| 20 | 75 | 30 | 5.94 | 3.851 | 2.69 |
2.12. Ramp function and t-test
The validity of the polynomial model was examined using a ramp function optimization approach, and the predicted values were statistically compared with the experimental results using Student’s t-test. Two sets of extraction conditions were randomly generated from the developed model, different from those included in the RSM design, and were used to verify the agreement between predicted and experimental responses (Table 3). The desirability function was set to 1.00. All validation experiments were performed in duplicate (n = 2), and the obtained data were statistically analyzed using Student’s t-test with IBM SPSS Statistics for Windows (version 26.0, Armonk, NY: IBM Corp.).| Extraction conditions | % Yield | TPCa | TFCb | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Run | Temperature (°C) | Time (min) | Solvent-to-plant material ratio (mL g−1) | Observed | Predicted | Observed | Predicted | Observed | Predicted |
| a TPC was expressed as mg GAE g−1 DM.b TFC was expressed as mg RE g−1 DM. | |||||||||
| 1 | 80 | 116 | 50 | 16.92 | 17.70 | 12.30 | 12.90 | 10.80 | 11.40 |
| 2 | 80 | 100 | 50 | 17.53 | 17.60 | 12.90 | 13.04 | 11.42 | 11.71 |
2.13. Phytochemical study of D. edule Lindl. leaves
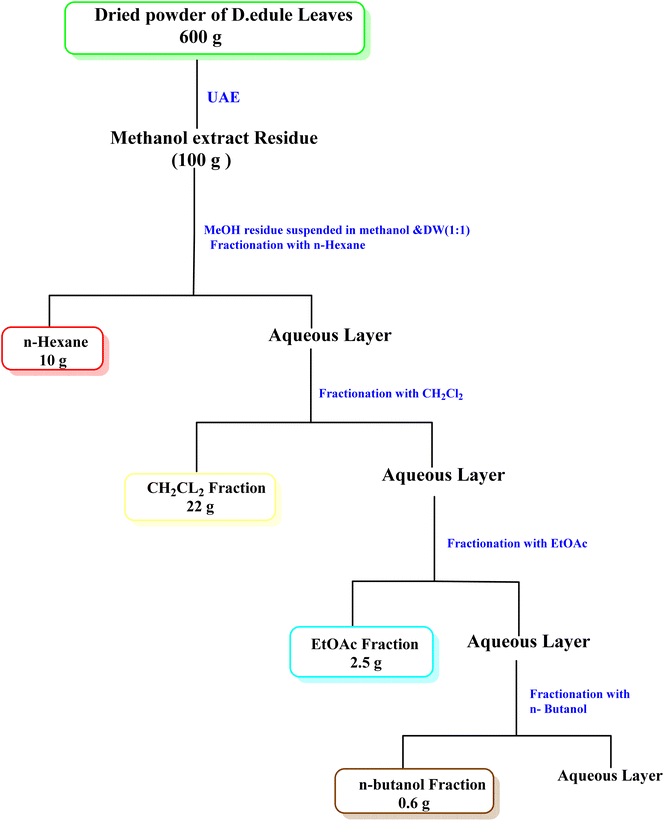
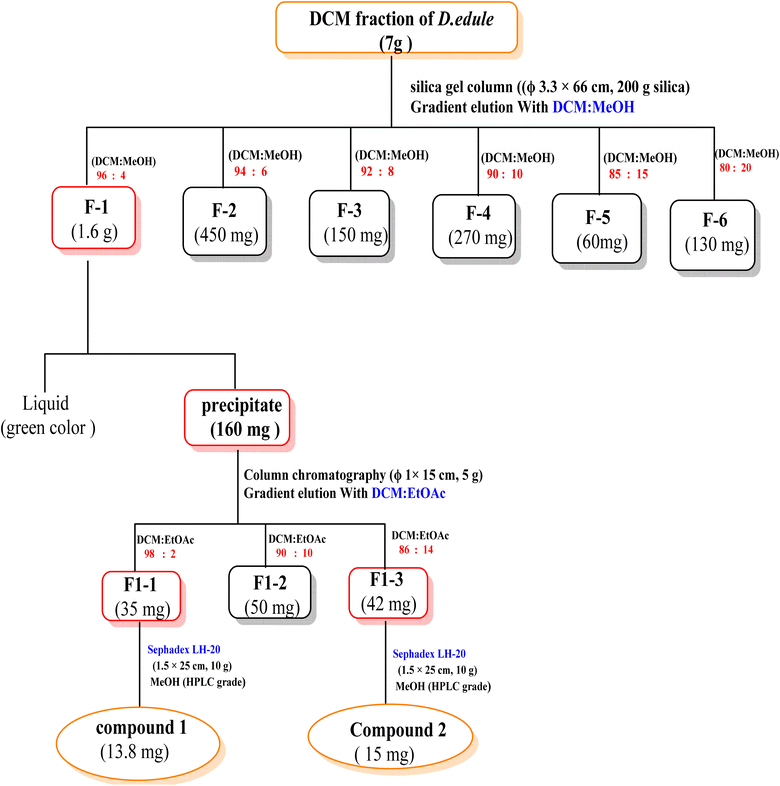
For the phytochemical study of D. edule Lindl. leaves, 600 g of the dried powder were extracted through UAE at the optimized conditions producing the highest TPC, i.e., at a temperature of 80 °C, an extraction time of 100 min, and a solvent-to-plant material ratio of 50 mL g−1 with 80% v/v MeOH. The combined methanolic extract was concentrated under reduced pressure at 40 °C to afford 100 g crude extract. The crude extract was suspended in water and fractionated using n-hexane, DCM, EtOAc, and n-butanol saturated with water to yield 10, 22, 2.5, and 0.6 g, respectively (Scheme 1).Approx. 7 g of the DCM fraction were subjected to a silica gel column chromatography (Φ 3.3 × 66 cm, 200 g silica) using a gradient elution method starting with DCM, then increasing polarity using MeOH. The collecting fractions (30 mL) were screened by TLC to give six fractions (F1 to F6). Fraction F1 (1.6 g) eluted with 4% MeOH in DCM was obtained as a precipitating powder. F1 precipitate (160 mg) was subjected to a silica gel column (Φ 1 × 15 cm, 5 g silica) using a gradient elution method starting with DCM, then increasing polarity using EtOAc in 2% increments to yield three subfractions (F1-1 to F1-3). The subfraction F1-1 (35 mg) was purified on a Sephadex LH-20 (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to give compound 1 (13.8 mg). The subfraction F1-3 (42 mg) was also purified using a Sephadex LH-20 (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to afford compound 2 (15 mg) (Scheme 2).
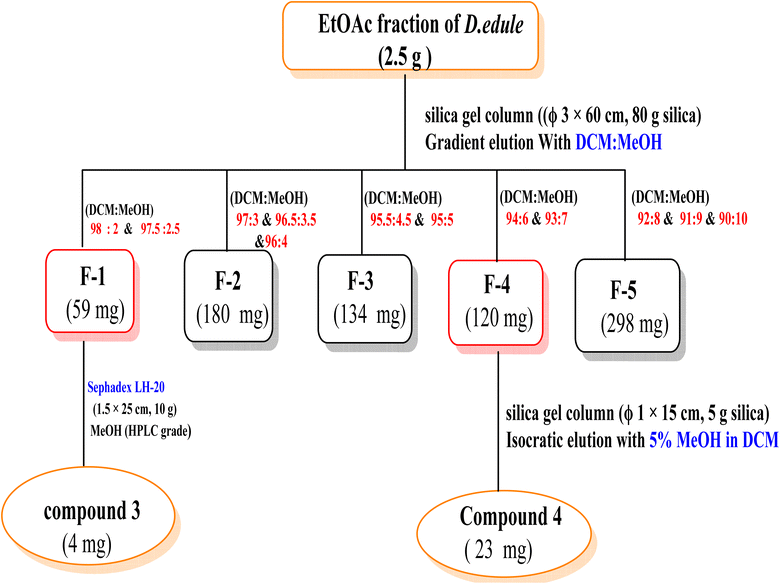
In addition, 2.5 g of the EtOAc fraction were subjected to a silica gel column chromatography (Φ 3 × 60 cm, 80 g silica) using a gradient elution method starting with DCM, then increasing polarity using methanol. The collecting fractions (30 mL) were screened by TLC to give five fractions (F 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5). F1 (59 mg, eluted with 2% MeOH in DCM) was re-chromatographed on a Sephadex-LH-20 column (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to yield compound 3 (4 mg). F4 (120 mg, eluted with 6% MeOH in DCM) was re-chromatographed over silica gel column (Φ 1 × 15 cm, 5 g silica) eluted isocratically with 5% MeOH in DCM to give compound 4 (23 mg) (Scheme 3).
5). F1 (59 mg, eluted with 2% MeOH in DCM) was re-chromatographed on a Sephadex-LH-20 column (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to yield compound 3 (4 mg). F4 (120 mg, eluted with 6% MeOH in DCM) was re-chromatographed over silica gel column (Φ 1 × 15 cm, 5 g silica) eluted isocratically with 5% MeOH in DCM to give compound 4 (23 mg) (Scheme 3).
To confirm the reproducibility of the optimized extraction conditions, an additional batch of D. edule Lindl. leaves (100 g) was extracted using UAE under the same optimized conditions. The combined methanolic extract was concentrated under reduced pressure at 40 °C to afford 16.5 g crude extract. Upon re-dissolving in a small amount of MeOH, yellow amorphous powder (2.5 g) was precipitated. TLC of this precipitate after dissolving in DCM revealed four spots. Column chromatography (Φ 3 × 60 cm, 80 g silica) was carried out using a gradient elution method starting with n-hexane and increasing polarity using EtOAc in 5% increments to afford five subfractions (F1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5), fraction volume was (25 mL). AlCl3 spray reagent was used for detection. F1 (50 mg, eluted with 15% EtOAc in DCM) was re-chromatographed on Sephadex-LH-20 column (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to yield compound 1 (32 mg), which is the same as that previously isolated from DCM fraction. F3 (14 mg, eluted with 25% EtOAc in DCM) was also purified on Sephadex-LH-20 column (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to yield compound 5 (8 mg) (Scheme 4).
5), fraction volume was (25 mL). AlCl3 spray reagent was used for detection. F1 (50 mg, eluted with 15% EtOAc in DCM) was re-chromatographed on Sephadex-LH-20 column (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to yield compound 1 (32 mg), which is the same as that previously isolated from DCM fraction. F3 (14 mg, eluted with 25% EtOAc in DCM) was also purified on Sephadex-LH-20 column (Φ 1.5 × 25 cm, 10 g) using MeOH (HPLC grade) to yield compound 5 (8 mg) (Scheme 4).
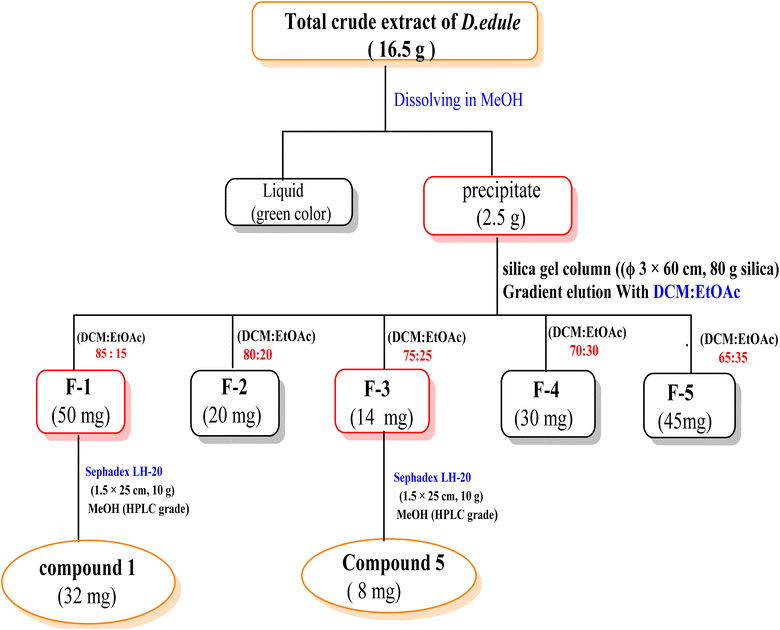 | ||
| Scheme 4 Column chromatography of the precipitate of total crude extract of D. edule Lindl. leaves after redissolving in MeOH. | ||
2.14. Statistical analysis of the data collected
Data were collected, tabulated, and statistically analyzed. Descriptive statistics were expressed as number (n), percentage (%), mean (![[x with combining macron]](https://www.rsc.org/images/entities/i_char_0078_0304.gif) ), and standard deviation (SD). Replicate numbers were as follows: preliminary screening of extraction methods (n = 2), RSM optimization (17 experimental runs including five replicates at the center point), and model verification experiments (n = 2). Variability was reported as standard deviation (SD) and represented as error bars in the relevant figure. Inferential statistical analysis included one-way ANOVA to compare quantitative variables among more than two normally distributed groups, followed by Tukey’s post hoc test for multiple pairwise comparisons. A one-sample t-test was applied to compare the experimental values with the predicted responses from the model. Differences were considered statistically significant at p < 0.05. All statistical analyses were performed using IBM SPSS Statistics for Windows (Version 26.0, IBM Corp., Armonk, NY, USA; released 2019).
), and standard deviation (SD). Replicate numbers were as follows: preliminary screening of extraction methods (n = 2), RSM optimization (17 experimental runs including five replicates at the center point), and model verification experiments (n = 2). Variability was reported as standard deviation (SD) and represented as error bars in the relevant figure. Inferential statistical analysis included one-way ANOVA to compare quantitative variables among more than two normally distributed groups, followed by Tukey’s post hoc test for multiple pairwise comparisons. A one-sample t-test was applied to compare the experimental values with the predicted responses from the model. Differences were considered statistically significant at p < 0.05. All statistical analyses were performed using IBM SPSS Statistics for Windows (Version 26.0, IBM Corp., Armonk, NY, USA; released 2019).
3. Results and discussion
3.1. Extraction yield, TPC, and TFC
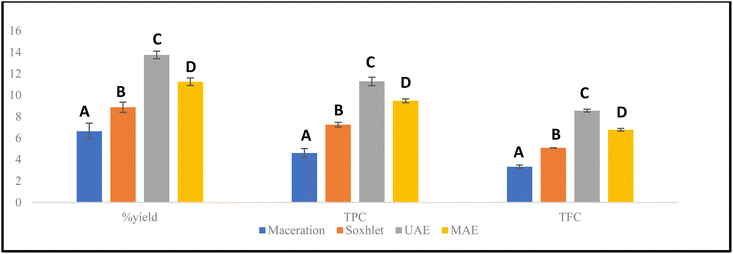
Medicinal plants and their extracts are rich in various secondary metabolites. Some of these compounds exhibit beneficial pharmacological effects, while others can affect the overall activity of the extracts either positively or negatively. Therefore, extraction methods for medicinal plants are designed to maximize the concentration of desired metabolites while minimizing unwanted or harmful substances. The levels of secondary metabolites obtained in the extracts are influenced by their physicochemical characteristics, the solvents used, the extraction techniques applied, and multiple parameters specific to each extraction method.29 In this work, the conventional methods of D. edule Lindl. extraction were compared with two different modern extraction methods to select the most suitable method to be optimized. The extraction efficiency of the different techniques was evaluated in terms of extraction yield, TPC, TFC, and antioxidant activity. Table 4 presents the yield, TPC, TFC, and DPPH of D. edule Lindl. extracts that were obtained using four different extraction techniques, namely maceration, Soxhlet, UAE and MAE. The comparative results are also illustrated in Fig. 1.| Extraction method | Yield (% w/w) | TPC (mg GAE g−1 DM) | TFC (mg RE g−1 DM) |
|---|---|---|---|
| a The results are expressed as the mean of two replicates (n = 2) ± SD; one-way ANOVA with Tukey’s post hoc, p < 0.05. Means followed by different letters are significantly different according to Tukey’s multiple range comparison. | |||
| Maceration | 6.63 ± 0.74a | 4.61 ± 0.41a | 3.33 ± 0.15a |
| Soxhlet | 8.86 ± 0.47b | 7.24 ± 0.21b | 5.08 ± 0.01b |
| UAE | 13.75 ± 0.35c | 11.28 ± 0.39c | 8.56 ± 0.13c |
| MAE | 11.25 ± 0.34d | 9.48 ± 0.17d | 6.78 ± 0.12d |
The extraction yield was calculated as the ratio between the mass of the dried plant material and the mass of the obtained extract. Among the tested techniques, the highest yield was obtained using UAE (13.75%), followed by MAE (11.25%), Soxhlet extraction (8.86%), and maceration (6.63%). Similarly, UAE of D. edule Lindl. produced the highest TPC (11.28 mg GAE g−1 DM), followed by MAE (9.48 mg GAE g−1 DM), Soxhlet (7.24 mg GAE g−1 DM), and maceration (4.61 mg GAE g−1 DM). Moreover, TFC ranged from 3.33 to 8.56 mg RE g−1 DM, with the highest value also observed for UAE. The results of the present study indicate that UAE achieved superior extraction efficiency for phenolic compounds and flavonoids compared with the other tested methods (Table 4). These findings are consistent with previous reports30–32 comparing the extraction efficiency of UAE, MAE, and maceration for medicinal plants. The enhanced extraction efficiency of UAE can be attributed to acoustic cavitation, which disrupts plant cell walls, enhances solvent penetration, and improves mass transfer of phenolic compounds into the extraction medium.33
In addition to improving extraction yield, TPC, and TFC, UAE demonstrated clear practical advantages over conventional techniques. The UAE process required significantly shorter extraction time and lower solvent consumption compared with maceration and Soxhlet extraction, contributing to reduced energy input and operational costs.
3.2. Antioxidant activity of D. edule extracts
Polyphenols are recognized as potent natural antioxidants due to their ability to participate in redox reactions, which are facilitated by resonance stabilization within their phenyl rings.7 Antioxidant properties of natural compounds can be initially assessed by measuring the DPPH scavenging activity, and the mechanism is based on the ability of antioxidants to donate hydrogen atoms or electrons to the DPPH radical, resulting in its reduction.34 The DPPH radical scavenging activities of D. edule Lindl. extracts obtained using different extraction methods are summarized in Table 5. Among the tested methods, UAE exhibited the highest % inhibition (37.1%), followed by MAE (32.5%), Soxhlet extraction (28.63%), and maceration (12.6%). The corresponding IC50 values further confirmed these results, with UAE showing the lowest IC50 (151.25 µg mL−1), indicating the strongest radical scavenging activity among the tested extracts, followed by MAE (186.15 µg mL−1), Soxhlet (193.45 µg mL−1), and Maceration (232.85 µg mL−1). Ascorbic acid was used as a positive control and showed an IC50 of 16.81 ± 0.10 µg mL−1.| Extraction method | DPPH | |
|---|---|---|
| % Inhibition at conc. 100 µg mL−1 | IC50 (µg mL−1) | |
| a The results are expressed as a means of two replicates (n = 2) ± SD; one-way ANOVA with Tukey's post hoc, p < 0.05. Means followed by different letters are significantly different according to Tukey's multiple range comparison. | ||
| Maceration | 12.6 ± 1.10a | 233.85 ± 0.15a |
| Soxhlet | 28.63 ± 0.76b | 193.54 ± 0.09b |
| UAE | 37.10 ± 0.49c | 151.25 ± 0.09c |
| MAE | 32.50 ± 0.73d | 186.15 ± 0.09d |
The observed antioxidant activity, as measured by the DPPH assay, was associated with the higher TPC and TFC obtained for the UAE extract. Several studies have reported similar correlations between increased phenolic and flavonoid contents and enhanced radical scavenging capacity, suggesting that these compounds may play an important role in antioxidant potential. For example, UAE of phenolic compounds from Corchorus olitorius (T8 variety, Tiliaceae) leaves resulted in significantly higher DPPH radical scavenging activity in parallel with increased TPC and TFC compared to conventional extraction methods.34
3.3. Antibacterial activity
Staphylococcus aureus is one of the major causes of morbidity and mortality worldwide, while Pseudomonas aeruginosa is an opportunistic pathogen responsible for severe infections, particularly in immuno-compromised individuals. Both pathogens have high antibiotic resistance, which highlights the need for new antimicrobial agents.Therefore, the antibacterial activities of D. edule Lindl. extracts obtained using different extraction methods were preliminarily assessed using disc diffusion. D. edule Lindl. extracts showed variable antibacterial activities depending on the extraction method (Table 6).
| Extraction technique | P. aeruginosa | S. aureus | ||
|---|---|---|---|---|
| Diameter of IZ* (mm) | % Activity Index | Diameter of IZ* (mm) | % Activity Index | |
| a The results are expressed as a mean of two replicates (n = 2) ± SD; one-way ANOVA with Tukey’s post hoc, p < 0.05. Means followed by different letters are significantly different according to Tukey’s multiple range comparison. | ||||
| Maceration | 4.9 ± 0.40a | 21.4 ± 1.70 | 8.5 ± 0.55a | 35.5 ± 1.30 |
| Soxhlet | 8.3 ± 0.15b | 36.3 ± 0.66 | 10.3 ± 0.1b | 42.9 ± 0.23 |
| UAE | 14.8 ± 0.10c | 64.5 ± 0.25 | 16 ± 0.15c | 67.3 ± 0.50 |
| MAE | 12.05 ± 0.03d | 52.4 ± 0.15 | 13.2 ± 0.14d | 55.1 ± 0.37 |
| Ciprofloxacin | 23 ± 0.02 | 100 ± 0.05 | 24 ± 0.1 | 100 ± 0.25 |
UAE showed the highest inhibition zones against both S. aureus (16 ± 0.15 mm) and P. aeruginosa (14.8 ± 0.1 mm), followed by MAE, Soxhlet and maceration. The enhanced antibacterial activity observed for the UAE extract may be related to the improved extraction efficiency and higher recovery of phenolic and flavonoid compounds. Similar trends have been reported in previous studies, where UAE improved the extraction of antimicrobial phytochemicals from plant materials. For example, flavonoids extracted from Cyclocarya paliurus (Batalin) Iljinskaja. using ultrasound-assisted extraction exhibited stronger antibacterial activity compared to those obtained by conventional methods.35 Likewise, ultrasound-derived extracts of Erodium glaucophyllum L. demonstrated enhanced antimicrobial activity against several pathogenic microorganisms.36
3.4. Cytotoxic activity
Polyphenols have been widely reported to exhibit anticancer-related activities, which are partly attributed to their antioxidant properties and modulation of oxidative stress. The structural diversity of these compounds enables interactions with reactive species and multiple cellular targets involved in cell survival and proliferation.37,38In the present study, the anticancer activity of D. edule Lindl. extracts was preliminarily evaluated in vitro against the human breast cancer cell line MCF-7. The cytotoxic activities of the different extracts are summarized in Table 7.
| Sample | In vitro cytotoxicity IC50 (µg mL−1) |
|---|---|
| MCF-7 | |
| a Results showing IC50 (µg mL−1) ± SD. The results are expressed as a mean of two replicates (n = 2) ± SD; one-way ANOVA with Tukey’s post hoc, p < 0.05. Means followed by different letters are significantly different according to Tukey’s multiple range comparison. IC50 (µg mL−1): 1–10 (very strong), 11–25 (strong), 26–50 (moderate), 51–100 (weak) and above 100 (non-cytotoxic). | |
| DOX | 4.17 ± 0.2 |
| SOR | 7.26 ± 0.3 |
| Maceration | 30.34 ± 2.7a |
| Soxhlet | 17.88 ± 1.5b |
| UAE | 12.61 ± 1.4c |
| MAE | 15.74 ± 1.3d |
The D. edule Lindl. extract obtained by UAE showed the strongest cytotoxic activity against MCF-7 cells (IC50 = 12.61 ± 1.4 µg mL−1), whereas the maceration extract exhibited moderate activity (IC50 = 30.34 ± 2.7 µg mL−1), according to the classification of.25 The higher cytotoxic activity observed for the UAE extract may be associated with its elevated phenolic and flavonoid contents compared with the other extraction methods. The relevance of UAE in cancer-related phytochemical research has been highlighted in previous studies. For example, polyphenols extracted by UAE from Thelephora ganbajun M. Zang demonstrated enhanced antiproliferative activity against several cancer cell lines, including MCF-7, A549, HepG2, and HT-29, compared with maceration and Soxhlet extracts.39
3.5. Fitting the models
Compared with conventional extraction methods, UAE resulted in higher extraction yield and extracts with improved biological activities. However, the objective of the present study was not merely to confirm the superiority of UAE, but to quantitatively define the optimal operational conditions for maximizing phenolic recovery from a gymnosperm matrix characterized by biflavonoid-rich composition. Accordingly, RSM was employed to optimize temperature, extraction time, and solvent-to-sample ratio for maximizing extraction yield, TPC, and TFC.All experimental data were analyzed using ANOVA to evaluate the significance and adequacy of the developed models by examining the corresponding F- and p-values (Tables 8 and 9).10 High coefficients of determination (R2) were obtained for % yield (0.98), TFC (0.98), and TPC (0.99), indicating a strong agreement between the predicted and experimental values (Table 2). In addition, the lack-of-fit was not significant for any of the responses (p > 0.05), confirming that the models adequately described the experimental data. The interactive effects of the independent variables on % yield, TPC, and TFC were visualized using three-dimensional response surface plots generated from the regression equations (Fig. 2). Hence, the developed models showed good reliability and were suitable for optimization. These statistical indicators confirm model validity; however, practical significance must be evaluated based on the magnitude of improvement achieved after optimization. To assess the practical relevance of the optimization process, extraction performance before and after RSM optimization was directly compared. The optimized UAE conditions increased the extraction yield from 13.75% to 16.5% (20% relative improvement). Similarly, TPC increased from 11.28 to 12.3 mg GAE g−1 DM, while TFC increased from 8.56 to 10.8 mg RE g−1 DM. Notably, antioxidant activity showed a substantial enhancement, as evidenced by the reduction in DPPH IC50 values from 151.25 to 75.6 µg mL−1, representing approximately a twofold increase in radical scavenging efficiency. These improvements demonstrate that the statistical significance of the RSM model translated into measurable and practically relevant gains in extract quality, particularly in terms of antioxidant performance.
| Source | Sum of squares | Degree of freedom | Mean square | F-Value | p-Value | |
|---|---|---|---|---|---|---|
| % Yield | ||||||
| Model | 193.46 | 6 | 32.24 | 85.87 | <0.0001 | Significant |
| Residual | 3.76 | 10 | 0.3755 | |||
| Lack of fit | 2.09 | 6 | 0.3489 | 0.8399 | 0.5967 | Not significant |
| Pure error | 1.66 | 4 | 0.4154 | |||
| Cor total | 197.22 | 16 | ||||
| R2 = 0.98; predicted R 2 = 0.94; adjusted R 2 = 0.97 coefficient of variance (%) = 5.84; adequate precision = 34.707 | ||||||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| TPC | ||||||
| Model | 112.66 | 6 | 18.78 | 123.98 | <0.0001 | Significant |
| Residual | 1.51 | 10 | 0.1514 | |||
| Lack of fit | 0.748 | 6 | 0.1247 | 0.6507 | 0.6965 | Not significant |
| Pure error | 0.7664 | 4 | 0.1916 | |||
| Cor total | 114.17 | 16 | ||||
| R2 = 0.99; predicted R 2 = 0.96; adjusted R 2 = 0.98 coefficient of variance (%) = 5.37; adequate precision = 42.78 | ||||||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| TFC | ||||||
| Model | 103.34 | 6 | 17.22 | 106.97 | <0.0001 | Significant |
| Residual | 1.61 | 10 | 0.1610 | |||
| Lack of fit | 1.03 | 6 | 0.1723 | 1.20 | 0.4511 | Not significant |
| Pure error | 0.5763 | 4 | 0.1441 | |||
| Cor total | 104.95 | 16 | ||||
| R2 = 0.98; predicted R 2 = 0.94; adjusted R 2 = 0.98 coefficient of variance (%) = 6.90; adequate precision = 40.75 | ||||||
| Source | Sum of squares | Degree of freedom | Mean square | F-Value | p-Value |
|---|---|---|---|---|---|
| % Yield | |||||
| A-temp | 130.57 | 1 | 130.57 | 347.73 | <0.0001 |
| B-time | 25.42 | 1 | 25.42 | 67.69 | <0.0001 |
| C-ratio | 17.46 | 1 | 17.46 | 46.51 | <0.0001 |
| AC | 5.95 | 1 | 5.95 | 15.86 | 0.0026 |
| A2 | 8.30 | 1 | 8.30 | 22.11 | 0.0008 |
| B2 | 4.99 | 1 | 4.99 | 13.29 | 0.0045 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| TPC | |||||
| A-temp | 77.46 | 1 | 77.46 | 511.47 | <0.0001 |
| B-time | 11.97 | 1 | 11.97 | 79.06 | <0.0001 |
| C-ratio | 11.49 | 1 | 11.49 | 75.88 | <0.0001 |
| AC | 4.59 | 1 | 4.59 | 30.30 | 0.0003 |
| A2 | 2.80 | 1 | 2.80 | 18.50 | 0.0016 |
| B2 | 3.95 | 1 | 3.95 | 26.11 | 0.0005 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| TFC | |||||
| A-temp | 71.77 | 1 | 71.77 | 455.74 | <0.0001 |
| B-time | 8.51 | 1 | 8.51 | 52.84 | <0.0001 |
| C-ratio | 11.21 | 1 | 11.21 | 69.61 | <0.0001 |
| AC | 5.83 | 1 | 5.83 | 36.24 | <0.0001 |
| A2 | 0.861 | 1 | 0.861 | 5.35 | 0.0433 |
| B2 | 4.91 | 1 | 4.91 | 30.52 | 0.0003 |
Compared with conventional maceration, which yielded lower phenolic and flavonoid contents, the optimized UAE protocol demonstrated superior efficiency in recovering bioactive compounds. Therefore, the contribution of this work lies not in introducing new extraction variables, but in providing a statistically validated and plant-specific optimization strategy that enhances both extraction efficiency and functional activity.
3.6. Effect of extraction variables on % yield, TFC, and TPC
The experimental data were analyzed using multiple regression, resulting in a second-order polynomial equation expressed in coded variables as follows (eqn (5)–(7)):| % Yield = 11.66 + 4.04A +1.78B + 1.48C + 1.22AC − 1.40A2 − 1.09B2 | (5) |
| TFC = 6.54 + 3A + 1.03B + 1.18C + 1.21AC − 0.4516A2 − 1.08B | (6) |
| TPC = 8.09 + 3.11A +1.22B + 1.20C + 1.07AC − 0.8147A2 − 0.9677B2 | (7) |
Among the extraction factors studied, temperature exhibited the most significant effect on all three responses, % yield, TPC, and TFC, with p values < 0.0001. This strong influence can be attributed to the role of temperature in reducing the viscosity and surface tension of the solvent, thereby enhancing solvent penetration into plant tissues and improving the diffusion and solubilization of intracellular compounds, which ultimately leads to higher extraction efficiency.40
The high F-values and extremely low p-values indicated that the developed models provided an excellent fit to the experimental data. Moreover, the non-significant lack-of-fit values confirmed that the models were appropriate, accurate, and reliable for predicting the extraction responses (Table 8).
The interaction between temperature and the solvent-to-plant material ratio significantly impacted the extraction process, affecting the extraction yield, TPC, and TFC. Moreover, the notable quadratic effects of temperature and extraction time suggest that their influence was nonlinear. Instead, a curved relationship was observed, pointing to optimal extraction conditions. Exceeding these conditions by increasing temperature or extraction time further may not enhance extraction efficiency and could potentially decrease it (Table 9).
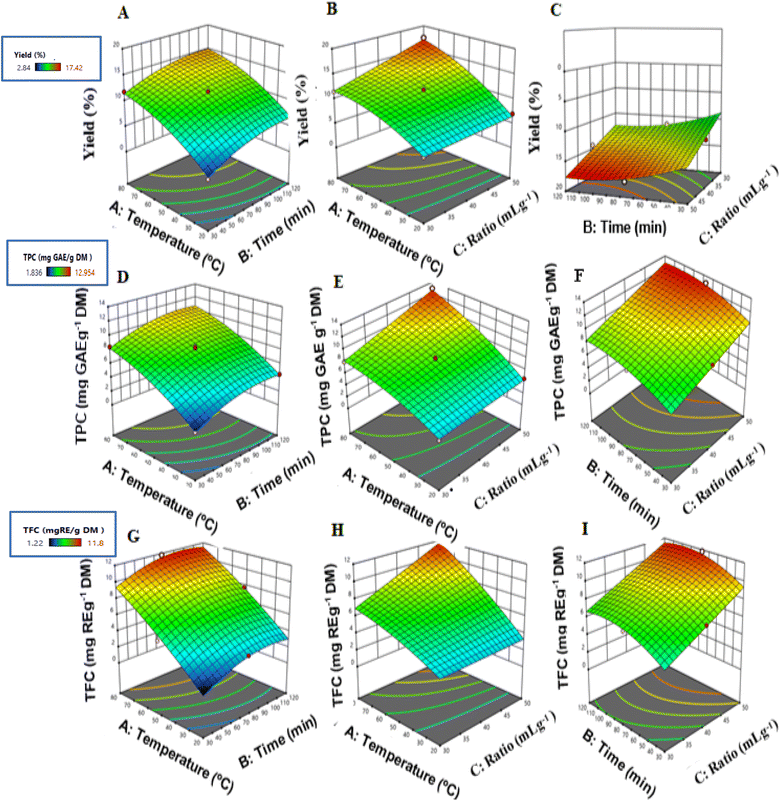
The 3D response surface plots (Fig. 2A–I) supported the results of the multiple regression analysis, confirming that extraction temperature exerted the most significant effect on extraction yield, TPC, and TFC. The responses increased as the temperature rose until reaching an optimal level, after which a slight decrease occurred at higher temperatures. This decline could be due to the thermal breakdown of heat-sensitive phenolic and flavonoid compounds, along with a decrease in the extraction rate constant at elevated temperatures. The latter is likely caused by reduced cavitation intensity resulting from lower surface tension and increased vapor pressure within the cavitation bubbles.33
Regarding the effect of the solvent-to-solid ratio, plots C, F, and I illustrate the interaction between extraction time (B) and solvent ratio (C). The steep gradient observed along the solvent ratio axis indicates that this parameter plays a critical role in improving extraction performance. Increasing the solvent volume enhances the concentration gradient between the plant matrix and the extraction medium, which acts as the main driving force for diffusion according to Fick’s second law.41 As a result, solvent saturation is delayed, maintaining an effective diffusion process and improving the recovery of phenolic and flavonoid compounds.
The elliptical contour patterns observed in these plots confirm the presence of a significant interaction between extraction time and solvent ratio. This finding indicates that extraction efficiency is governed by a synergistic balance between these parameters rather than by maximizing each factor independently.
These results highlight that temperature and solvent ratio are the most critical factors controlling UAE efficiency, while extraction time contributes mainly through its interaction with the other variables. This interpretation further supports the robustness of the developed optimization model.
3.7. Verification of the model
The results of the t-test indicated no statistically significant difference (p > 0.05) between the predicted and experimental values for all tested responses. This lack of significance confirms that the experimental results were in close agreement with the values predicted by the RSM model. Such consistency demonstrates the model's robustness and reliability in describing the relationship between extraction variables and the measured responses. Therefore, the optimized extraction conditions derived from the RSM model can be considered valid and experimentally reproducible. This agreement between predicted and actual data highlights the adequacy of the developed model for accurately predicting extraction performance under similar conditions. It also supports the practical applicability of the model in optimizing extraction parameters to maximize yield and bioactive compound recovery from D. edule leaves.3.8. Phytochemical study of D. edule leaves
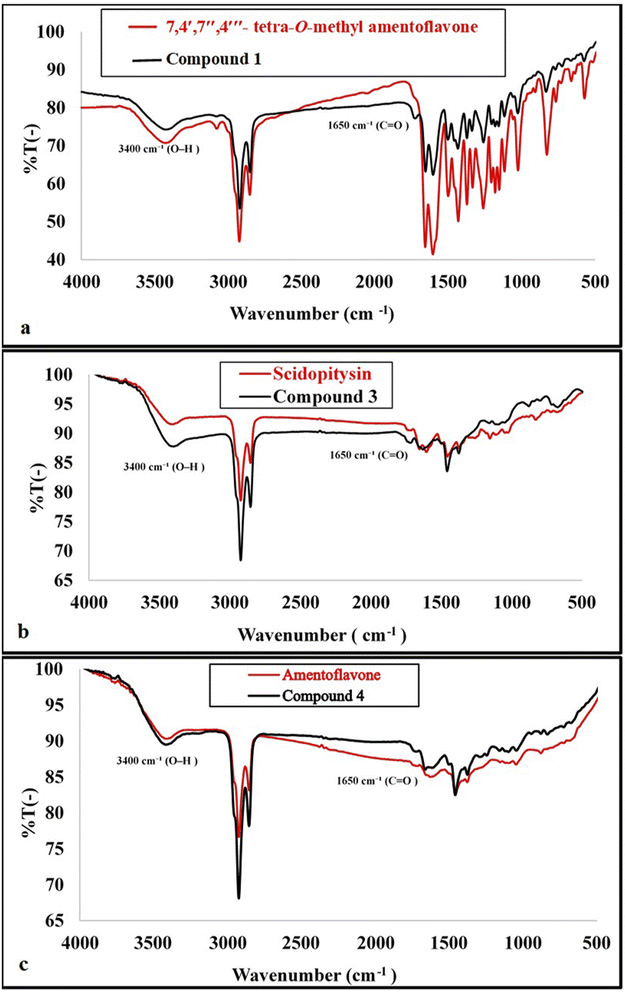
In the present study, five biflavonoids (1–5) were successfully isolated (Fig. 3) and structurally characterized using comprehensive spectroscopic techniques, including UV, IR, ESI-MS, and 1D and 2D NMR analyses (1H, 13C, DEPTQ, and HMBC). Structural elucidation was further supported by comparison with authentic reference compounds (Fig. 4) and previously reported literature data. The isolated biflavonoids were identified as 7,4′,7′′,4‴-tetra-O-methylamentoflavone (1), isoginkgetin (2), sciadopitysin (3), amentoflavone (4), and kayaflavone (5). Interestingly, compound 1 was re-isolated from a different chromatographic fraction.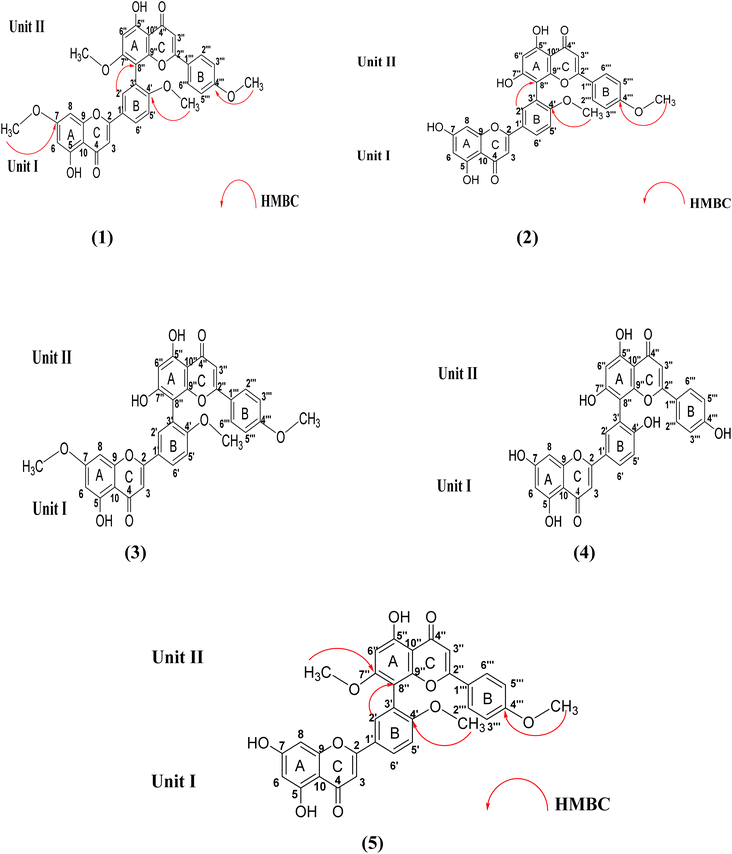 | ||
| Fig. 3 Chemical structures of compounds (1–5) isolated from D. edule using UAE at the optimized conditions. | ||
Compound 1 (isolated from DCM fraction) was obtained as a yellow amorphous powder (13.8 mg), soluble in chloroform and slightly soluble in MeOH. The compound gave yellow color with AlCl3 on TLC with an Rf value of 0.94 using solvent system S1 (CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeOH, 9.5
MeOH, 9.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.5). It gave a negative Molisch's test, indicating a non-glycosidic nature. UV spectrum of compound 1 (MeOH) showed UV λmax (MeOH) at 225, 268, and 327 that referred to the biflavonoids bands42 (Fig. S3). The IR spectrum exhibited a broad band hydroxyl absorption band at 3423 cm−1, a carbonyl band at 1656 cm−1 and an aliphatic C–H stretching band attributable to methoxy groups at 2928 cm−1 (Fig. S4). 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 6.93 (2H, d, J = 9.1 Hz, H-2‴, H-6‴) and δH 7.61 (2H, d, J = 9.1 Hz, H-3‴, H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH7.38 (1H, d, J = 9.0 Hz, H-5′), 8.10 (1H, d, J = 2.5 Hz, H-2′), and 8.24 (1H, dd, J = 9.0, 2.5 Hz, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. The presence of two m-coupled protons in ring A of unit I was evidenced by signals at δH 6.78 (1H, d, J = 2.2 Hz, H-8) and δH 6.37 (1H, d, J = 2.2 Hz, H-6). A singlet resonance at δH 7.01 was assigned to H-6″. Furthermore, the 1H NMR spectrum displayed four aromatic methoxy proton signals at δH 3.82, 3.79, 3.84, and 3.75, indicating a tetra-methoxylated amentoflavone derivative (Fig. S6a, b and Table S1). Analysis of the 13C NMR spectrum confirmed the biflavonoid nature of compound 1, as evidenced by the presence of 30 carbon signals along with four aromatic methoxy carbons resonating at δC 56.04, 56.00, 56.53, and 55.51. The involvement of C-8″ and C-3′ in the interflavonoid linkage was supported by their characteristic downfield shifts relative to apigenin, showing Δδ values of 10.6 ppm for C-8″ (δC 104.63) and 5.46 ppm for C-3′ (δC 121.26). Furthermore, comparison with amentoflavone revealed an up-field shift of C-8, indicative of 7-O-methoxylation, an up-field shift of C-5′ accompanied by a downfield shift of C-1′, supporting 4′-O-methoxylation. Similarly, an up-field shift of C-6″ suggested 7″-O-methoxylation, while up-field shifts of C-3‴ and C-5‴ together with a downfield shift of C-1‴confirmed 4‴-O-methoxylation (Fig. S7a, b and Table S2).
0.5). It gave a negative Molisch's test, indicating a non-glycosidic nature. UV spectrum of compound 1 (MeOH) showed UV λmax (MeOH) at 225, 268, and 327 that referred to the biflavonoids bands42 (Fig. S3). The IR spectrum exhibited a broad band hydroxyl absorption band at 3423 cm−1, a carbonyl band at 1656 cm−1 and an aliphatic C–H stretching band attributable to methoxy groups at 2928 cm−1 (Fig. S4). 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 6.93 (2H, d, J = 9.1 Hz, H-2‴, H-6‴) and δH 7.61 (2H, d, J = 9.1 Hz, H-3‴, H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH7.38 (1H, d, J = 9.0 Hz, H-5′), 8.10 (1H, d, J = 2.5 Hz, H-2′), and 8.24 (1H, dd, J = 9.0, 2.5 Hz, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. The presence of two m-coupled protons in ring A of unit I was evidenced by signals at δH 6.78 (1H, d, J = 2.2 Hz, H-8) and δH 6.37 (1H, d, J = 2.2 Hz, H-6). A singlet resonance at δH 7.01 was assigned to H-6″. Furthermore, the 1H NMR spectrum displayed four aromatic methoxy proton signals at δH 3.82, 3.79, 3.84, and 3.75, indicating a tetra-methoxylated amentoflavone derivative (Fig. S6a, b and Table S1). Analysis of the 13C NMR spectrum confirmed the biflavonoid nature of compound 1, as evidenced by the presence of 30 carbon signals along with four aromatic methoxy carbons resonating at δC 56.04, 56.00, 56.53, and 55.51. The involvement of C-8″ and C-3′ in the interflavonoid linkage was supported by their characteristic downfield shifts relative to apigenin, showing Δδ values of 10.6 ppm for C-8″ (δC 104.63) and 5.46 ppm for C-3′ (δC 121.26). Furthermore, comparison with amentoflavone revealed an up-field shift of C-8, indicative of 7-O-methoxylation, an up-field shift of C-5′ accompanied by a downfield shift of C-1′, supporting 4′-O-methoxylation. Similarly, an up-field shift of C-6″ suggested 7″-O-methoxylation, while up-field shifts of C-3‴ and C-5‴ together with a downfield shift of C-1‴confirmed 4‴-O-methoxylation (Fig. S7a, b and Table S2).
In the HSQC spectrum, the signals at δH 3.82 (3H, s), 3.79 (3H, s), 3.84 (3H, s), and 3.75 (3H,s) were correlated to δC 56.04, 56.00, 56.53, and 55.51 respectively indicating four methoxy groups (Fig. S8a and b).
The HMBC data further supported the proton assignments. In the HMBC spectrum of compound 1, the correlation observed between H-2′ at δH 8.10 and C-8″ at δC 104.63 confirmed the involvement of C-3′ and C-8″ in the interflavonoid linkage. This finding supports the classification of compound 1 as a member of the amentoflavone series. Additionally, singlet signals at δH 3.82, 3.79, 3.84, and 3.75, each integrating for three protons, showed correlations with carbons at δC 165.16, 160.47, 162.64, and 162.33 indicating the presence of four methoxy groups. Accordingly, the methoxy substituents were assigned to C-7, C-4′, C-7″, and C-4‴ (Fig. S9a–S9c). The HPLC-ESI-MS analysis of compound 1 exhibited a pseudo-molecular ion peak at m/z 595.5 [M + H]+ (Fig. S10). This finding was further confirmed by HR-ESI-MS, which showed a pseudo-molecular ion at m/z 595.1597 [M + H]+ (Fig. S11). The observed mass data are consistent with the molecular formula (C34H26O10) and structure of 7,4′,7″,4‴-tetramethylamentoflavone. Comparison of the spectral data of compound 1 with previously reported literature16,43 led to its identification as 7,4′,7″,4‴ tetra-O-methyl amentoflavone.
Interestingly, the same compound was re-isolated from the total extract as a yellow powder (32 mg). It exhibited identical chromatographic behavior, physicochemical properties, and spectral characteristics, including TLC profile, 1H NMR, DEPTQ-135, and mass spectrometry data (Fig. S12–S14). Therefore, this compound was confirmed to be identical to the previously characterized metabolite, indicating that it represents the same compound obtained from a different fraction.
Compound 2 (isolated from DCM fraction) was obtained as a yellow amorphous powder (15 mg), soluble in chloroform and slightly soluble in MeOH. The compound gave a yellow color with AlCl3 on TLC with an Rf value of 0.51 using solvent system S1 (CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeOH, 9.5
MeOH, 9.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.5). It gave a negative Molisch’s test, indicating a non-glycosidic nature. UV spectrum of compound 2 (MeOH) showed UV λmax (MeOH) at 221, 268, and 329 that referred to the biflavonoid bands42 (Fig. S15). 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 7.61 (2H, d, J = 9.0 Hz, H-2‴, H-6‴) and δH 6.92 (2H, d, J = 9.0 Hz, H-3‴, H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH7.35 (1H, d, J = 9.0 Hz, H-5′), 8.05 (1H, d, J = 2.5 Hz, H-2′), and 8.19 (1H, dd, J = 9.0, 2.5 Hz, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. The presence of two m-coupled protons in ring A of unit I was evidenced by signals at δH 6.48 (1H, d, J = 2.2 Hz, H-8) and δH 6.19 (1H, d, J = 2.2 Hz, H-6). A singlet resonance observed at δH 6.42 was assigned to H-6″. 1H-NMR spectrum demonstrated amentoflavone pattern with two aromatic methoxy signals at δH 3.79 and 3.76 ppm suggesting di-methoxy derivatives (Fig. S16a, S16b and Table S3). 13C-NMR spectrum confirmed the biflavonoid structure of compound 2 by showing 30 carbons along with two methoxy signals at δC 55.49 and 55.89. 13C-NMR spectrum showed up-field shift in C-5′, downfield shift in C-1′ indicating 4′-O- methoxylation, also up-field shift in C-3‴, 5‴, downfield shift in C-1‴ indicating 4‴-O methoxylation compared with amentoflavone16 (Fig. S17a, b and Table S4). All spectroscopic data were consistent with the structure of 4′,4‴di-O-methyl amentoflavone.
0.5). It gave a negative Molisch’s test, indicating a non-glycosidic nature. UV spectrum of compound 2 (MeOH) showed UV λmax (MeOH) at 221, 268, and 329 that referred to the biflavonoid bands42 (Fig. S15). 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 7.61 (2H, d, J = 9.0 Hz, H-2‴, H-6‴) and δH 6.92 (2H, d, J = 9.0 Hz, H-3‴, H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH7.35 (1H, d, J = 9.0 Hz, H-5′), 8.05 (1H, d, J = 2.5 Hz, H-2′), and 8.19 (1H, dd, J = 9.0, 2.5 Hz, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. The presence of two m-coupled protons in ring A of unit I was evidenced by signals at δH 6.48 (1H, d, J = 2.2 Hz, H-8) and δH 6.19 (1H, d, J = 2.2 Hz, H-6). A singlet resonance observed at δH 6.42 was assigned to H-6″. 1H-NMR spectrum demonstrated amentoflavone pattern with two aromatic methoxy signals at δH 3.79 and 3.76 ppm suggesting di-methoxy derivatives (Fig. S16a, S16b and Table S3). 13C-NMR spectrum confirmed the biflavonoid structure of compound 2 by showing 30 carbons along with two methoxy signals at δC 55.49 and 55.89. 13C-NMR spectrum showed up-field shift in C-5′, downfield shift in C-1′ indicating 4′-O- methoxylation, also up-field shift in C-3‴, 5‴, downfield shift in C-1‴ indicating 4‴-O methoxylation compared with amentoflavone16 (Fig. S17a, b and Table S4). All spectroscopic data were consistent with the structure of 4′,4‴di-O-methyl amentoflavone.
In the HSQC spectrum, the signals at δH 3.76 (3H, s) and 3, 79 (3H, s) were correlated to δC 55.4 and 55.8, respectively, indicating two methoxy groups (Fig. S18a and S18b). This assignment was further confirmed by HMBC analysis, which showed clear 3JCH correlations from the methoxy protons at H-4′OCH3 (δ 3.79) to C-4′ (δ 160.61) and from H-4‴OCH3 (δ 3.76) to C-4‴ (δ 161.4). In addition, the HMBC spectrum supported the interflavonoid linkage through C-3′ and C-8″, as evidenced by 3JCH correlations from H-2′ (δ 8.05) to C-8″ (δ 103.76) (Fig. S19a–c). These findings confirm that compound 2 is an amentoflavone-member biflavonoid. The HPLC-ESI-MS analysis of compound 2 exhibited a pseudo-molecular ion peak at m/z 565.5 [M − H]− and m/z 567.4 [M + H]+ (Fig. S20). This finding was further confirmed by HR-ESI-MS, which showed a pseudo-molecular ion at m/z 565.1127 [M − H]− (Fig. S21). The observed mass data are consistent with the molecular formula(C32H22O10) and structure of 4′,4‴ di-O-methyl amentoflavone. Comparison of the spectral data of compound 2 with previously reported literature15,16,44 led to its identification as 4′,4‴ di-O-methyl amentoflavone (isoginkgetin).
Compound 3 (isolated from EtOAc fraction) was obtained as a pale-yellow amorphous powder (4 mg), soluble in chloroform and slightly soluble in MeOH. The compound gave a yellow color with AlCl3 reagent on TLC (Rf = 0.92) using solvent system S2 (CH2Cl2: MeOH, 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and showed a negative Molisch’s test, indicating a non-glycosidic nature. The UV spectrum of compound 3 (MeOH) exhibited absorption maxima at λmax 232, 271, and 330 nm, characteristic of the biflavonoid chromophores (Fig. S22). The IR spectrum displayed a broad hydroxyl absorption band at 3402 cm−1, a carbonyl stretching band at 1630 cm−1, and an aliphatic C–H stretching band attributable to methoxy groups at 2923 cm−1 (Fig. S23). Further comparison of compound 3 with an authentic sciadopitysin sample using TLC and CO-TLC analysis showed identical behavior. In addition, superimposition of the IR spectra demonstrated a high degree of similarity, including the fingerprint region (Fig. S24). Based on these combined spectroscopic and chromatographic data, which were consistent with those reported previously,15,45,46compound 3 was confidently identified as sciadopitysin (7,4′,4‴ tri-O- methyl amentoflavone).
1) and showed a negative Molisch’s test, indicating a non-glycosidic nature. The UV spectrum of compound 3 (MeOH) exhibited absorption maxima at λmax 232, 271, and 330 nm, characteristic of the biflavonoid chromophores (Fig. S22). The IR spectrum displayed a broad hydroxyl absorption band at 3402 cm−1, a carbonyl stretching band at 1630 cm−1, and an aliphatic C–H stretching band attributable to methoxy groups at 2923 cm−1 (Fig. S23). Further comparison of compound 3 with an authentic sciadopitysin sample using TLC and CO-TLC analysis showed identical behavior. In addition, superimposition of the IR spectra demonstrated a high degree of similarity, including the fingerprint region (Fig. S24). Based on these combined spectroscopic and chromatographic data, which were consistent with those reported previously,15,45,46compound 3 was confidently identified as sciadopitysin (7,4′,4‴ tri-O- methyl amentoflavone).
Compound 4 (isolated from EtOAc fraction) was obtained as a pale-yellow amorphous powder (4 mg), soluble in chloroform and slightly soluble in MeOH. The compound gave a yellow color with AlCl3 reagent on TLC (Rf = 0.46) using solvent system S2 (CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeOH, 9
MeOH, 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and showed a negative Molisch's test, indicating a non-glycosidic nature. The UV spectrum of compound 4 (MeOH) exhibited absorption maxima at λmax 233, 270, and 337 nm, characteristic of biflavonoid chromophores (Fig. S25). The IR spectrum displayed a broad hydroxyl absorption band at 3415 cm−1, a carbonyl stretching band at 1662 cm−1, and an aliphatic C–H stretching band attributable to methoxy groups at 2923 cm−1 (Fig. S26). In addition, superimposition of the IR spectra with authentic amentoflavone demonstrated a high degree of similarity, including the fingerprint region (Fig. S27). 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 7.58 (2H, d, J = 8 Hz, H-2‴, H-6‴) and at δH 6.73 (2H, d, J = 8 Hz, H-3‴,H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH7.15 (1H, d, J = 8 Hz, H-5′), 8.01(1H, brs, H-2′), and 7.99 (1H, brs, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. Two signals were observed in the up-field aromatic region at δH 6.46 and δH 6.19, each integrating for one proton, and were assigned to the H-8 and H-6 protons of ring A, respectively. A singlet resonance at δH 6.39 was assigned to H-6″ (Fig. S28a, b and Table S5). 13C-NMR spectrum confirmed the biflavonoid nature of compound 4, revealing a total of 30 carbon signals. The involvement of C-8″ and C-3′ in the interflavonoid linkage was evidenced by the downfield shifts of C-8″ (9.81 ppm) and C-3′ (5.29 ppm) compared to the apigenin (Fig. S29a, b and Table S6). The HPLC–ESI-MS analysis of compound 4 exhibited a pseudo-molecular ion peak at m/z 537.4 [M − H]− and m/z 539.4 [M + H]+ (Fig. S30). This finding was further confirmed by HR-ESI-MS, which showed a pseudo molecular ion at m/z 539.0958 [M + H]+ (Fig. S31). The observed mass data is consistent with the molecular formula (C30H18O10) and structure of amentoflavone. Comparison of the spectral data of compound 4 with previously reported literature16,47,48 led to its identification as amentoflavone.
1) and showed a negative Molisch's test, indicating a non-glycosidic nature. The UV spectrum of compound 4 (MeOH) exhibited absorption maxima at λmax 233, 270, and 337 nm, characteristic of biflavonoid chromophores (Fig. S25). The IR spectrum displayed a broad hydroxyl absorption band at 3415 cm−1, a carbonyl stretching band at 1662 cm−1, and an aliphatic C–H stretching band attributable to methoxy groups at 2923 cm−1 (Fig. S26). In addition, superimposition of the IR spectra with authentic amentoflavone demonstrated a high degree of similarity, including the fingerprint region (Fig. S27). 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 7.58 (2H, d, J = 8 Hz, H-2‴, H-6‴) and at δH 6.73 (2H, d, J = 8 Hz, H-3‴,H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH7.15 (1H, d, J = 8 Hz, H-5′), 8.01(1H, brs, H-2′), and 7.99 (1H, brs, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. Two signals were observed in the up-field aromatic region at δH 6.46 and δH 6.19, each integrating for one proton, and were assigned to the H-8 and H-6 protons of ring A, respectively. A singlet resonance at δH 6.39 was assigned to H-6″ (Fig. S28a, b and Table S5). 13C-NMR spectrum confirmed the biflavonoid nature of compound 4, revealing a total of 30 carbon signals. The involvement of C-8″ and C-3′ in the interflavonoid linkage was evidenced by the downfield shifts of C-8″ (9.81 ppm) and C-3′ (5.29 ppm) compared to the apigenin (Fig. S29a, b and Table S6). The HPLC–ESI-MS analysis of compound 4 exhibited a pseudo-molecular ion peak at m/z 537.4 [M − H]− and m/z 539.4 [M + H]+ (Fig. S30). This finding was further confirmed by HR-ESI-MS, which showed a pseudo molecular ion at m/z 539.0958 [M + H]+ (Fig. S31). The observed mass data is consistent with the molecular formula (C30H18O10) and structure of amentoflavone. Comparison of the spectral data of compound 4 with previously reported literature16,47,48 led to its identification as amentoflavone.
Compound 5 (isolated from the total extract) was obtained as a yellow amorphous powder (8 mg), soluble in chloroform and slightly soluble in MeOH. On TLC, the compound gave a yellow color upon spraying with AlCl3 and showed an Rf value of 0.62 using solvent system S1 (CH2Cl2: MeOH, 9.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.5). It gave a negative Molisch's test, indicating its non-glycosidic nature. 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 7.63 (2H, d, J = 9.0 Hz, H-2‴, H-6‴) and δH 6.93 (2H, d, J = 9.0 Hz, H-3‴, H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH 7.37 (1H, d, J = 8.9 Hz, H-5′), 8.08 (1H, d, J = 2.5 Hz, H-2′), and 8.20 (1H, dd, J = 8.9, 2.5 Hz, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. The presence of two m-coupled protons in ring A of unit I was evidenced by signals at δH 6.48 (1H, d, J = 2.1 Hz, H-8) and δH 6.19 (1H, d, J = 2.1 Hz, H-6). A singlet resonance at δH 6.96 was assigned to H-6″. Also, 1H-NMR spectrum revealed amentoflavone pattern with three aromatic methoxy signals at δH 3.79, 3.85, and 3.76 suggesting tri-methoxy derivatives (Fig. S32a, b and Table S7). In addition, the DEPTQ-135 NMR of compound 5 confirmed the biflavonoid nature of compound 5 as it showed 30 carbons in addition to three methoxy signals at δ 56.44, 57.00, and 55.99. Moreover, DEPTQ-135 NMR spectrum revealed different chemical shifts as up-field shift in C-5′, downfield shift in C-1′ indicating 4′-O-methoxylation, up-field shift in C-6″, indicating 7″-O methoxylation, up-field shift in C-3‴,5‴, downfield shift in C-1‴ indicating 4‴-O methoxylation compared with amentoflavone (Fig. S33) and (Table S8). All spectroscopic data were consistent with the structure of 4′,7″,4‴-tri-O-methyl amentoflavone.16
0.5). It gave a negative Molisch's test, indicating its non-glycosidic nature. 1H-NMR spectrum indicated a biflavonoid structure characteristic of an amentoflavone-type compound (biapigenin nucleus). An AA′BB′ spin system corresponding to the para-substituted B ring of unit II was observed at δH 7.63 (2H, d, J = 9.0 Hz, H-2‴, H-6‴) and δH 6.93 (2H, d, J = 9.0 Hz, H-3‴, H-5‴). Additionally, an ABX coupling pattern attributable to ring B of unit I appeared at δH 7.37 (1H, d, J = 8.9 Hz, H-5′), 8.08 (1H, d, J = 2.5 Hz, H-2′), and 8.20 (1H, dd, J = 8.9, 2.5 Hz, H-6′), supporting a C-3′ interflavonoid linkage between the two flavonoid units. The presence of two m-coupled protons in ring A of unit I was evidenced by signals at δH 6.48 (1H, d, J = 2.1 Hz, H-8) and δH 6.19 (1H, d, J = 2.1 Hz, H-6). A singlet resonance at δH 6.96 was assigned to H-6″. Also, 1H-NMR spectrum revealed amentoflavone pattern with three aromatic methoxy signals at δH 3.79, 3.85, and 3.76 suggesting tri-methoxy derivatives (Fig. S32a, b and Table S7). In addition, the DEPTQ-135 NMR of compound 5 confirmed the biflavonoid nature of compound 5 as it showed 30 carbons in addition to three methoxy signals at δ 56.44, 57.00, and 55.99. Moreover, DEPTQ-135 NMR spectrum revealed different chemical shifts as up-field shift in C-5′, downfield shift in C-1′ indicating 4′-O-methoxylation, up-field shift in C-6″, indicating 7″-O methoxylation, up-field shift in C-3‴,5‴, downfield shift in C-1‴ indicating 4‴-O methoxylation compared with amentoflavone (Fig. S33) and (Table S8). All spectroscopic data were consistent with the structure of 4′,7″,4‴-tri-O-methyl amentoflavone.16
In the HSQC spectrum, the signals at δH 3.79 (3H, s), 3.85 (3H, s), and 3,76 (3H,s) were correlated to δC 56.44, 57.00,and 55.99 respectively indicating three methoxy groups (Fig. S34a and b). This assignment was further confirmed by HMBC analysis, which showed clear 3JCH correlations from the methoxy protons at H-4′OCH3 (δ 3.79) to C-4′ (δ 160.80), H-7″OCH3 (δ 3.85) to C-7″ (δ 163.12) and from H-4‴OCH3 (δ 3.76) to C-4‴ (δ 162.81). In addition, the HMBC spectrum supported the interflavonoid linkage through C-3′ and C-8″, as evidenced by 3JCH correlation from H-2′ (δ 8.08) to C-8″ (δ 105.13) (Fig. S35a and S35b). These findings confirmed that compound 5 is an amentoflavone-member biflavonoid. The HR-ESI-MS spectrum of compound 5 exhibited a pseudo molecular ion at m/z 581.46985 [M + H]+ which agrees with the molecular formula (C33H24O10) and structure of 4′,7″,4‴-tri-O-methyl amentoflavone (Fig. S36). Furthermore, Comparison of the spectral data of compound 5 with previously reported literature16,43 led to its identification as kayaflavone (4′,7″,4‴-tri-O-methyl amentoflavone). This is the first report for isolation of kayaflavone from D. edule Lindl.
Although the isolated biflavonoids are known constituents of gymnosperms, their significance in the present study lies in their enhanced recovery at optimized UAE conditions. For example, UAE of D. edule Lindl. at the optimized conditions yielded 32 mg of 7,4′,7″,4‴-tetra-O-methylamentoflavone from 100 g of dried plant material, corresponding to an extraction yield of 0.032%. In contrast, a previous study reported the isolation of only 4 mg of the same compound from 930 g of D. edule Lindl. using conventional percolation, corresponding to a yield of 0.00043%.15 Similarly, 23 mg of amentoflavone was obtained from 600 g of dried plant material under optimized UAE, whereas conventional maceration of Dioon spinulosum Dyer Ex Eichler yielded only 26 mg (0.0013%) from 2 kg.45 These data collectively suggest improved extraction efficiency under optimized UAE parameters.
The improved recovery of these compounds can be attributed to acoustic cavitation, which disrupts plant cell walls, enhances solvent penetration, and accelerates mass transfer between the solvent and plant matrix. As a result, intracellular metabolites such as 7,4′,7″,4‴-tetra-O-methylamentoflavone and amentoflavone are released more efficiently within shorter extraction times.
The higher relative abundance of these bioactive biflavonoids under optimized UAE conditions is consistent with the enhanced antioxidant and cytotoxic activities observed for the extracts. These findings provide quantitative evidence that extraction optimization influences metabolite recovery and may impact biological performance.
Although UAE is often described as a green extraction technique, the use of methanol limits the overall sustainability profile due to its toxicity. However, the significantly reduced extraction time and lower solvent consumption compared with conventional methods represent improvements in process efficiency. Future replacement of methanol with safer solvent systems such as aqueous ethanol may further enhance environmental compatibility.
4. Conclusion
In this study, four extraction techniques, i.e., maceration, Soxhlet, UAE, and MAE, were systematically evaluated for their effectiveness in extracting phenolic compounds from D. edule Lindl. leaves. Evaluation criteria included % extraction yield, TPC, TFC, in addition to antioxidant, antibacterial and cytotoxic activities. Among these methods, the UAE demonstrated superior performance, which justified its selection for further optimization by RSM. Subsequent optimization of UAE parameters allowed for a large-scale extraction that enabled the successful isolation of five key bioactive biflavonoids: 7,4′,7′′,4‴-tetra-O-methylamentoflavone (1, isolated from DCM fraction and total methanolic extract), isoginkgetin (2, DCM fraction), sciadopitysin (3, EtOAc fraction), amentoflavone (4, EtOAc fraction), and kayaflavone (5, total methanolic extract). Interestingly, this study represents the first report of kayaflavone isolation from D. edule. The isolation of these bioflavonoids is consistent with the enhanced antioxidant, cytotoxic, and antibacterial activities observed in the UAE extracts. The cavitation effects generated by ultrasound facilitate more efficient disruption of plant cell walls and improve solvent penetration, resulting in significantly higher yields of these biologically active compounds compared to conventional extraction methods. Based on their well-documented bioactivities, these compounds likely contribute substantially to the superior functional properties exhibited by the UAE extracts.Moreover, although temperature, time, and solvent-to-sample ratio are commonly investigated variables in UAE studies, their combined interactive effects are matrix-dependent and cannot be assumed a priori. The present findings highlight how small adjustments within conventional parameter ranges can substantially influence phenolic recovery in gymnosperm tissues, which are structurally and chemically distinct from many angiosperm species.
Therefore, this study highlights UAE as a technically advanced extraction approach with improved efficiency over conventional methods. The promising antioxidant, cytotoxic, and antibacterial activities associated with the isolated biflavonoids suggest potential applications of UAE-derived extracts in pharmaceutical and nutraceutical fields, warranting further investigation.
Author contributions
Asmaa M. Akar: writing – original draft, writing – review & editing, methodology, visualization, formal analysis, data curation. Nesrine M. Hegazi: methodology, supervision, writing – review & editing. Ahmed Zayed: writing – review & editing, writing – original draft, supervision, resources, project administration, conceptualization. Souzan M. Ibrahim: supervision, writing – review & editing. All authors have read and approved the final submitted version of the manuscript.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00755d.Acknowledgements
This work did not receive any kind of financial support. The authors are greatly appreciated Dr Ahmed Serag (Al-Azhar University, Faculty of Pharmacy, Egypt) for helping model construction and data interpretation, Dr Mohamed I. Elshorbagy and Dr Ahmed S. Arafa (Department of Pharmacognosy, Faculty of Pharmacy, Tanta University, Egypt) for their valuable assistance in analyzing the samples using NMR and mass spectroscopy. In addition, Dr Walaa Negm (Department of Pharmacognosy, Faculty of Pharmacy, Tanta University, Egypt) is acknowledgment for providing us the authentic compounds required for comparsion. During the preparation of this work, ChatGPT (OpenAI) was used in order to improve the readability and language of the manuscript. After using this tool/service, the authors reviewed and edited the content as needed and took full responsibility for the content of the published article.References
- M. M. Rahman, M. S. Rahaman, M. R. Islam, F. Rahman, F. M. Mithi, T. Alqahtani, M. A. Almikhlafi, S. Q. Alghamdi, A. S. Alruwaili and M. S. Hossain, Molecules, 2021, 27, 233 CrossRef PubMed.
- D. Borjan, M. Leitgeb, Ž. Knez and M. K. Hrnčič, Molecules, 2020, 25, 5946 CrossRef CAS PubMed.
- J. Azmir, I. S. M. Zaidul, M. M. Rahman, K. Sharif, A. Mohamed, F. Sahena, M. Jahurul, K. Ghafoor, N. Norulaini and A. Omar, J. Food Eng., 2013, 117, 426–436 Search PubMed.
- Q.-W. Zhang, L.-G. Lin and W.-C. Ye, Chin. Med., 2018, 13, 1–26 Search PubMed.
- H. Yao, X. Li, Y. Liu, Q. Wu and Y. Jin, J. Ginseng Res., 2016, 40, 415–422 Search PubMed.
- C. Da Porto and A. Natolino, Food Chem., 2018, 258, 137–143 Search PubMed.
- C. S. Dzah, Y. Duan, H. Zhang, C. Wen, J. Zhang, G. Chen and H. Ma, Food Biosci., 2020, 35, 100547 Search PubMed.
- P. C. Veggi, J. Martinez and M. A. A. Meireles, in Microwave-assisted Extraction for Bioactive Compounds: Theory and Practice, Springer, 2012, pp. 15–52 Search PubMed.
- W. J. Hill and W. G. Hunter, Technometrics, 1966, 8, 571–590 Search PubMed.
- M. A. Bezerra, R. E. Santelli, E. P. Oliveira, L. S. Villar and L. A. Escaleira, Talanta, 2008, 76, 965–977 CrossRef CAS PubMed.
- A. Moawad, M. Hetta, J. K. Zjawiony, D. Ferreira and M. Hifnawy, Nat. Prod. Res., 2014, 28, 41–47 CrossRef CAS PubMed.
- A. Moawad, M. Hetta, J. K. Zjawiony, M. R. Jacob, M. Hifnawy, J. P. Marais and D. Ferreira, Planta Med., 2010, 76, 796–802 Search PubMed.
- M. K. Mourya, A. Prakash, A. Swami, G. K. Singh and A. Mathur, World J. Sci. Technol., 2011, 1, 11–20 CAS.
- X. Xiong, N. Tang, X. Lai, J. Zhang, W. Wen, X. Li, A. Li, Y. Wu and Z. Liu, Front. Pharmacol, 2021, 12, 768708 CrossRef CAS PubMed.
- A. Moawad and D. Amir, Eur. J. Med. Plants, 2016, 16, 1–7 Search PubMed.
- K. R. Markham, C. Sheppard and H. Geiger, Phytochemistry, 1987, 26, 3335–3337 CrossRef CAS.
- H. M. El-Seadawy, A. E. Ragab, M. El-Aasr, K. A. A. El-Seoud, A. A. Elblihy, E.-S. El-Alfy, H. A. Elgawad, S. Saleh, H. Sheta and R. Elseadawy, Sci. Rep., 2025, 15, 15697 CrossRef CAS PubMed.
- A. Ibrahim, N. F. Mohammad, K. F. Kasim, N. F. M. Nasir, S. S. M. Salleh, M. Sangar, F. D. M. Daud and R. F. Navea, Journal of Advanced Research in Fluid Mechanics and Thermal Sciences, 2025, 127(1), 213–222 CrossRef.
- E. Badawy, M. Vinatoru, I. Calinescu, K. A. Shams, N. S. Abel-Azim, A. A. Fahmi, M. Abdul-Rahman, A. A. Abd-Rabou, A. R. Hamed, K. Mahmoud, T. J. Mason and I. A. Saleh, Biointerface Res. Appl. Chem., 2023, 13(5), 463 CAS.
- A. M. Salih, F. Al-Qurainy, M. Nadeem, M. Tarroum, S. Khan, H. O. Shaikhaldein, A. Al-Hashimi, A. Alfagham and J. Alkahtani, Molecules, 2021, 26, 7454 Search PubMed.
- A. R. Alzahrani, N. Hosny, D. I. Mohamed, H. H. A. Nahas, A. Albogami, T. M. I. Al-Hazani, I. A. A. Ibrahim, A. H. Falemban, G. A. Bamagous and E. M. Saied, RSC Adv., 2024, 14, 19400–19427 RSC.
- N. Kishore, D. Twilley, A. Blom van Staden, P. Verma, B. Singh, G. Cardinali, D. Kovacs, M. Picardo, V. Kumar and N. Lall, J. Nat. Prod., 2018, 81, 49–56 CrossRef CAS PubMed.
- M. M. Labib, A. M. Alqahtani, H. H. Abo Nahas, R. M. Aldossari, B. F. Almiman, S. Ayman Alnumaani, M. El-Nablaway, E. Al-Olayan, M. Alsunbul and E. M. Saied, Biomolecules, 2024, 14, 1018 CrossRef CAS PubMed.
- S. M. Khirallah, H. M. Ramadan, H. A. A. Aladl, N. O. Ayaz, L. A. Kurdi, M. Jaremko, S. Z. Alshawwa and E. M. Saied, Pharmaceuticals, 2022, 15, 1576 CrossRef CAS PubMed.
- A. Hossan and H. Abu-Melha, Synthesis, 2014, 21 Search PubMed.
- A. Christou, F. Nikola and V. Goulas, Chem. Biodivers., 2025, 22, e202401337 CrossRef CAS PubMed.
- T. Ahmed, M. R. Rana, M. R. Maisha, A. Sayem, M. Rahman and R. Ara, Heliyon, 2022, 8, e11109 CrossRef CAS PubMed.
- J. Zhou, L. Zhang, Q. Li, W. Jin, W. Chen, J. Han and Y. Zhang, Molecules, 2018, 24, 112 Search PubMed.
- F. Chemat, M. Abert Vian, H. K. Ravi, B. Khadhraoui, S. Hilali, S. Perino and A.-S. Fabiano Tixier, Molecules, 2019, 24, 3007 CrossRef CAS PubMed.
- B. Feraoun, K. Otmanine, M. Hammoudi, H. Mostfaoui and R. Fellah, South Afr. J. Bot., 2025, 187, 623–631 CrossRef CAS.
- J. H. Shourove, A. R. Kazi, F. U. Emon and G. R. Islam, Discover Food, 2025, 5, 359 Search PubMed.
- L. Kurniasari, M. Djaeni and A. C. Kumoro, Braz. J. Food Technol., 2023, 26, e2022140 CrossRef CAS.
- N. Medina-Torres, T. Ayora-Talavera, H. Espinosa-Andrews, A. Sánchez-Contreras and N. Pacheco, Agronomy, 2017, 7, 47 CrossRef CAS.
- A. Biswas, S. Dey, A. Xiao, Y. Deng, Z. M. Birhanie, R. Roy, D. Akhter, L. Liu and D. Li, Chem. Biol. Technol. Agric., 2023, 10, 64 CrossRef CAS.
- W.-B. Hu, Z.-W. Yang and W.-J. Wang, Ind. Crops Prod., 2019, 130, 615–626 CrossRef.
- R. Abdelkebir, C. Alcántara, I. Falcó, G. Sánchez, J. V. Garcia-Perez, M. Neffati, J. M. Lorenzo, F. J. Barba and M. C. Collado, Innov. Food Sci. Emerg. Technol., 2019, 52, 189–196 CrossRef CAS.
- C. S. Dzah, Afr. J. Food Nutr. Sci., 2014, 14, 9578–9591 Search PubMed.
- E. Brglez Mojzer, M. Knez Hrnčič, M. Škerget, Ž. Knez and U. Bren, Molecules, 2016, 21, 901 CrossRef PubMed.
- D.-P. Xu, J. Zheng, Y. Zhou, Y. Li, S. Li and H.-B. Li, Int. J. Mol. Sci., 2016, 17, 1664 CrossRef PubMed.
- A. P. D. F. Machado, B. R. Sumere, C. Mekaru, J. Martinez, R. M. N. Bezerra and M. A. Rostagno, Int. J. Food Sci. Technol., 2019, 54, 2792–2801 CrossRef CAS.
- A. Sanou, K. Konaté, K. Kabakde, R. Dakuyo, D. Bazié, S. Hemayoro and M. H. Dicko, Sci. Rep., 2023, 13, 358 CrossRef CAS PubMed.
- N. Kumar, B. Singh, P. Bhandari, A. P. Gupta, S. K. Uniyal and V. K. Kaul, Phytochemistry, 2005, 66, 2740–2744 CrossRef CAS PubMed.
- C.-M. Sun, W.-J. Syu, Y.-T. Huang, C.-C. Chen and J.-C. Ou, J. Nat. Prod., 1997, 60, 382–384 CrossRef CAS PubMed.
- P. Sugita, D. D. Agusta, H. Dianhar, I. H. Suparto, K. Kurniawanti, D. U. C. Rahayu and L. Irfana, J. Appl. Pharmaceut. Sci., 2023, 13, 199–209 CAS.
- W. A. Negm, K. A. Abo El-Seoud, A. Kabbash, A. A. Kassab and M. El-Aasr, Nat. Prod. Res., 2021, 35, 5166–5176 Search PubMed.
- M. Kovač Tomas, I. Jurčević and D. Šamec, Plants, 2022, 12, 147 CrossRef PubMed.
- B. R. Nair, Ind. Crop. Prod., 2021, 160, 113099 CrossRef.
- H. M. El-Seadawy, K. A. Abo El-Seoud, M. El-Aasr, H. O. Tawfik and A. E. Ragab, Plants, 2022, 11, 2867 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |