Open Access Article
Open Access ArticleA structural blueprint for antibacterial discovery: microwave- and ultrasound-assisted synthesis of pyrrolidine-fused quinoxalines as novel inhibitors of DNA gyrase and biofilm
Eman A. Fayed *a,
Mazin A. A. Najmb,
Mostafa I. Abdelglilc,
Triveena M. Ramsis*d,
Nirvana A. Gohar
*a,
Mazin A. A. Najmb,
Mostafa I. Abdelglilc,
Triveena M. Ramsis*d,
Nirvana A. Gohar e,
Shimaa A. Metwallyf and
Maha A. Ebrahima
e,
Shimaa A. Metwallyf and
Maha A. Ebrahima
aDepartment of Pharmaceutical Organic Chemistry, Faculty of Pharmacy (Girls), Al-Azhar University, Cairo 11754, Egypt. E-mail: alfayed_e@azhar.edu.eg; alfayed_e@yahoo.com; Tel: +202 01221330523
bDepartment of Pharmacy, Mazaya University College, Nasiriyah 64001, Iraq
cDepartment of Medical Laboratory Techniques, Al-Farahidi University, Baghdad, 10021, Iraq
dDepartment of Pharmaceutical Chemistry, Faculty of Pharmacy, Sinai University, Kantara Branch, Ismailia 41636, Egypt. E-mail: triveena.farid@su.edu.eg; triveenaramsis@gmail.com
eDepartment of Pharmaceutical Organic Chemistry, Faculty of Pharmacy, Modern University for Technology and Information (MTI), Cairo 11571, Egypt
fMicrobiology and Immunology Department Faculty of Pharmacy (Girls), Al-Azhar University, Cairo 11754, Egypt
First published on 17th March 2026
Abstract
In order to lessen the severity of infectious diseases, anti-infective agents—drugs that prevent, combat, or control infections brought on by microorganisms—are essential in contemporary medicine. To tackle antimicrobial resistance, this project intends to design and synthesize hybrid compounds that contain pyrrolidine, quinoxaline and a hydrazinyl bridge, and assess the antimicrobial and antifungal properties of these compounds against a variety of pathogenic strains. The bactericidal properties of hybrids 24, 27, and 29 against E. coli were verified. The MIC of 12.5 µM was shown by hybrids 24, 25, and 31, which suggests bactericidal hybrids are effective against P. aeruginosa at greater concentrations. In comparison to Levofloxacin, treatment with all hybrids produced an 89–92% reduction in biofilm formation at 90% MIC. Eight hybrids' killing kinetics against P. aeruginosa were time-dependent, with an abrupt decrease in CFU number observed at higher concentrations. While 4-fold and 8-fold MICs resulted in nearly total bacterial eradication, primary bacterial elimination happened after three hours. The most effective DNA gyrase inhibitors were hybrids 25, 28, and 31; their IC50 values were significantly less than that of ciprofloxacin (77.3, 87.6, and 65.5 µM, respectively). To determine the best drug-like qualities, the study examined the physicochemical and pharmacokinetic features of active compounds. Molecular docking simulation experiments were also conducted to comprehend the binding interactions and mechanisms of action of these hits.
1. Introduction
Diseases caused by infectious bacteria, viruses, and fungi have appeared and are continuing to grow in tandem with daily life.1–4 The need for novel antimicrobial drugs with lower toxicity and resistance strain activity appears unresolved.5–8Although many antibacterial medications are already accessible, microorganism resistance to antibiotics remains an immense issue.9–12 Compounds that are heterocyclic are frequently included in numerous synthetic and biological fields because of their ubiquitous occurrence in nature.13–16 Amongst the several scaffolds considered for the prospective pharmacological applications of them, nitrogen-containing heterocyclic frameworks have acquired a lot of focus owing to the diverse spectrum of biological processes,17–19 comprising antifungal,20,21 antimicrobial,22,23 anticancer,24,25 antiviral,26,27 and antidiabetic properties.28,29
A useful pharmacophore in medicinal science, quinoxaline, is a bicyclic molecule constructed when a pyrazine ring and a benzene ring combine.30–33 Its distinct structure permits a variety of alterations, producing molecules with ameliorated biological activity.34–37 Due to its capacity to communicate with the essential cellular structures and microbial enzymes, quinoxaline derivatives have recently been shown to potentially be potent antimicrobial drugs.38 The naturally occurring antibiotic phenazinomycin, which has anticancer properties, is another antibacterial medication containing a quinoxaline scaffold.39–43
Additionally, pyrrolidine ring piqued the curiosity of scientists because of its strong antibacterial qualities.44–47 Numerous quinolone antibiotics, including Gemifloxcin and Moxifloxacin, include pyrrolidine.48–50 In addition, the newly approved drug vonoprazan, which destroys Helicobacter-pylori, has a scaffold with a pyrrole ring.51–53 Including an azo part to the compounds which are heterocyclic enhances the drug's biological impact.27,54–57
DNA gyrase is a potential target for antibacterial therapy, a crucial bacterial enzyme associated with transcription, recombination, and DNA replication.58 The effective prevention of bacterial growth is achieved by DNA gyrase inhibitors, which disrupt DNA supercoiling and replication.59 Since DNA gyrase is essential for bacterial life yet is absent from human cells, it is a desirable target for targeted antimicrobial therapy.60
The notion that bacteria must attain a particular cell density in infectious illnesses before they may establish virulence factors and interfere with the host defences is widely recognized.61,62 Pathogenic bacteria have evolved a special mechanism known as “biofilm formation” to withstand the lethal assaults of oxidative stress, nutritional deprivation, and the host immune system or antibiotics. Microorganisms form a biofilm while they latch on to an abiotic or biotic surface in a matrix of extracellular polymeric substance (EPS). The ability of Candida species forge biofilms, in addition to bacteria, increases the possibility of resistance to existing antifungal medications.63,64 The need for novel molecules or hybrids of already-existing chemicals that can suppress or destroy the biofilm matrix is critical because there is currently no treatment option for biofilms in clinical settings.
The purpose of this work is to use pyrrolidine-fused quinoxaline employing azo bridge to produce a variety of novel hybrid compounds, as seen in Fig. 1. The evaluation of the novel hybrids involved examining their DNA gyrase inhibitory, antibacterial, and antifungal properties. The antibiofilm activity of each bactericidal derivative was examined. Each compound was examined for pharmacokinetic and physicochemical characteristics. Lastly, an evaluation of binding mode simulations through docking for the most successful samples were carried out.
 | ||
| Fig. 1 Schematic representation of hybridized substances featuring quinoxaline, pyrrole/pyrrolidine, azo-bridge, and various substitutes. | ||
2. Findings and analysis
2.1. Chemical synthesis
The methods indicated in Schemes 1–3 were used to synthesis the target compounds.Benzene-1,2-diamine 1 was relfuxed with oxalic acid 2 using water as a solvent in presence of conc. HCl affording 1,4-dihydroquinoxaline-2,3-dione 3. Further reaction of 3 with thionyl chloride resulted in 2,3-dichloroquinoxaline 5. Hydrazinylquinoxaline 4 or 6 were obtained by refluxing 3 or 5 with hydrated-hydrazine, respectively, in ethyl alcohol for 24 h, accordingly as represented in Scheme 1.65
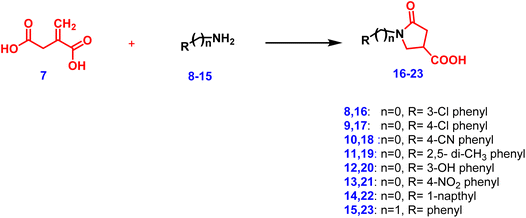
Furthermore, the reaction of itaconic acid 7 with various aniline derivatives 8–15, afforded the 5-oxo-pyrrolidine derivatives 16–23 (Scheme 2). The reaction was conducted using three methods implementing the principles of green chemistry (discussed in the experimental section). Using ultrasound-assisted reaction afforded the highest yield and consumed the least time. The reaction time and yield are given in Table 1. Structure elucidation using 1H NMR confirmed the structures of oxo-pyrrolidine derivatives 16–23. The COOH proton appeared as a singlet-signal at approximately δ 12 part per million (ppm). The hydrogens for the two methylene carbons appeared as four doublet of doublet signals each integrating one proton at approximately δ 2 and 4 ppm. The methine hydrogens from pyrrolidine showed up as a multiplet at approximately delta-(δ) 3 ppm.
| Compound | Reflux in water | Ultrasound | Microwave | |||
|---|---|---|---|---|---|---|
| Time | Yield % | Time | Yield % | Time | Yield % | |
| 16 | 24 h | 56% | 6 min | 91% | 12 min | 73% |
| 17 | 24 h | 44% | 4.5 min | 95% | 13 min | 77% |
| 18 | 24 h | 62% | 5 min | 90% | 12 min | 84% |
| 19 | 24 h | 50% | 5.5 min | 92% | 11 min | 79% |
| 20 | 24 h | 55% | 4 min | 89% | 15 min | 86% |
| 21 | 24 h | 58% | 5.5 min | 88% | 14 min | 73% |
| 22 | 24 h | 65% | 5 min | 92% | 15 min | 88% |
| 23 | 24 h | 57% | 4 min | 90% | 12 min | 84% |
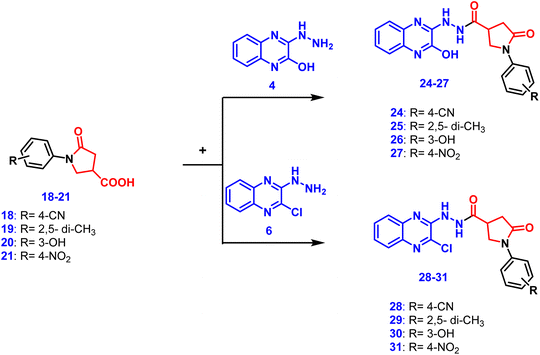
The targeted quinoxalne-pyrrolidine hybrids 24–31 were afforded by refluxing the respective pyrrolidine fragment with hydrazinyl quinoxaline 4 or 6 (Scheme 3). A characteristic singlet appeared in the 1H NMR spectra of hybrids 24–27 at approximately δ 11–12 ppm representing the 2-OH proton on the quinoxaline ring. In all synthesized hybrids, –NH–NH–CO showed as doublet at δ 11 ppm, while –NH–NH–CO showed as a doublet roughly between δ 5 and 8 ppm. The protons of the pyrrolidine ring exhibited the identical signal pattern in all synthesized hybrids. The 13C NMR of all synthesized hybrids revealed two characteristic signals representing the two C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups at δ between 170 and 175 ppm.
O groups at δ between 170 and 175 ppm.
The 1H NMR of hybrids 25 and 29 demonstrated two characteristic signals between δ 2 and 2.3 ppm, each integrating for 3 protons representing the two methyl groups. Additionally, the two methyl groups were shown as two signals in 13C NMR at approximately δ 16 and 19 ppm. The 1H NMR of hybrids 26 and 30 a signal representing the phenolic-OH between delta (δ) 7 and delta (δ) 8 part per million. The 13C NMR of hybrids 24 and 28 confirmed the presence of the cyano groups at approximately δ 118 ppm.
2.2. In vitro assessment
| Cpd. no. | Average inhibition zone diameter (mm) | |||||
|---|---|---|---|---|---|---|
| Bacteria (Gram negative) | Bacteria (Gram positive) | Fungi | ||||
| E. coli | P. aeruginosa | K. pneumonia | A. baumannii | MSSA | C. albicans | |
| a DMSO was used as a solvent with no inhibition zone. | ||||||
| 24 | 20.83 ± 0.60 | 18 ± 0.57 | 20.3 ± 1.2 | — | — | 13.8 ± 0.16 |
| 25 | 16.3 ± 0.33 | 19.6 ± 0.33 | 17.8 ± 0.16 | — | — | — |
| 26 | 19 ± 0.01 | 20.16 ± 0.6 | 20.6 ± 0.33 | — | — | 15 ± 0.01 |
| 27 | 19.3 ± 0.33 | 18.3 ± 0.88 | 15.5 ± 0.28 | 13.8 ± 0.16 | — | 14.6 ± 0.33 |
| 28 | 18.3 ± 0.66 | 22.3 ± 0.3 | 16.6 ± 0.33 | 15.5 ± 0.28 | 15.5 ± 0.28 | 13.8 ± 0.4 |
| 29 | 14.6 ± 0.33 | 14.6 ± 0.66 | 14 ± 0.01 | 13.8 ± 0.16 | 14.6 ± 0.66 | 14 ± 0.57 |
| 30 | 20.6 ± 0.33 | 21.3 ± 0.33 | 20.5 ± 0.28 | 15.3 ± 0.33 | 17 ± 0.01 | 15 ± 0.57 |
| 31 | 18 ± 0.57 | 20.6 ± 0.88 | 15.16 ± 0.16 | 17.16 ± 0.44 | 15.6 ± 0.33 | 14.6 ± 0.6 |
| Levofloxacin | 26.3 ± 0.33 | 19.6 ± 1.3 | 27.5 ± 0.28 | 30.5 ± 0.28 | 41.1 ± 0.01 | NA |
| Amoxicillin | 17.6 ± 0.33 | 15.3 ± 0.33 | 16.3 ± 0.33 | 15.6 ± 0.33 | 25 ± 0.01 | NA |
| Fluconazole | NA | NA | NA | NA | NA | 19 ± 0.57 |
Additionally, the ZOI values of hybrids 26, 28, 30, and 31 ranged from 20 to 23 mm, indicating a greater reduction in P. aeruginosa growth than that of Levofloxacin (19.6 ± 1.3 mm). Out of all the synthetic derivatives, hybrids 24, 25, 26, and 30 showed the highest ZOI against K. pneumoniae, with values ranging from 17 to 21 mm, outperforming Amoxicillin (16.3 ± 0.33 mm). Only hybrid 31 showed the greatest ZOI in the case of A. baumannii, with a value of 17.16 ± 0.44 mm surpassing that of Amoxicillin (15.6 ± 0.33 mm), while only hybrid 28 was on par with Amoxicillin.
Nevertheless, the hybrids under investigation showed modest antibacterial efficacy against MSSA. With ZOI varying from 13 to 15 mm, all hybrids excluding 25 showed considerable antifungal efficacy against C. albicans, which is comparable to the common antifungal medication Fluconazole (19 ± 0.57 mm).
| Cpd. no. | Bacteria (Gram negative) | Bacteria (Gram positive) | Fungi | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E.coli | P. aeruginosa | K. pneumonia | A. baumannii | MSSA | C. albicans | |||||||||||||
| MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | MIC | MFC | R | |
| 24 | 25 | 50 | 2 | 12.5 | 25 | 2 | 12.5 | 50 | 4 | >200 | >200 | — | >200 | >200 | — | 50 | 100 | 2 |
| 25 | 12.5 | 50 | 4 | 12.5 | 100 | 8 | 50 | 200 | 4 | >200 | >200 | — | >200 | >200 | — | 12.5 | 50 | 4 |
| 26 | 12.5 | 50 | 4 | 25 | 50 | 2 | 12.5 | 50 | 4 | >200 | >200 | — | >200 | >200 | — | 50 | 100 | 2 |
| 27 | 25 | 50 | 2 | 50 | 100 | 2 | 50 | 100 | 2 | 50 | 50 | 1 | >200 | >200 | — | 50 | 100 | 2 |
| 28 | 12.5 | 50 | 4 | 6.25 | 25 | 4 | 25 | 50 | 2 | 6.25 | 12.5 | 2 | 12.5 | 25 | 2 | 100 | 200 | 2 |
| 29 | 6.25 | 12.5 | 2 | 100 | 200 | 2 | 25 | 100 | 4 | 50 | 100 | 2 | 6.25 | 12.5 | 2 | 12.5 | 50 | 4 |
| 30 | 12.5 | 50 | 4 | 6.25 | 50 | 8 | 50 | 200 | 4 | 25 | 50 | 2 | 25 | 50 | 2 | 50 | 100 | 2 |
| 31 | 50 | 200 | 4 | 12.5 | 50 | 4 | 25 | 100 | 4 | 100 | 100 | 1 | 6.25 | 25 | 4 | 100 | 200 | 2 |
| Levofloxacin | 8 | 4 | 8 | 2 | <0.5 | NA | ||||||||||||
| Amoxicillin | 1 | 256 | >1024 | >1024 | <0.5 | NA | ||||||||||||
| Fluconazole | — | — | — | — | — | 64 | ||||||||||||
Hybrids 28 and 30 displayed substantial MIC value of 6.25 µM, and MBC value of 25 µM and 50 µM, against P. aeruginosa, resulting in (R) value of 4 and 8, indicating bacteriostatic activity. Moreover, 24, 25 and 31 displayed MIC of 12.5 µM, whereas the calculated (R) indicate bactericidal hybrids at higher concentrations.
In case of K. pneumonia, 24 and 26 displayed excellent MIC value of 12.5 µM, and a much higher MBC value resulting in a ratio above 2 indicating bacteriostatic hybrids. Compounds 28, 29, and 31 demonstrated moderate MIC value of 25 µM. Hybrid 28 emerged as the most potent derivative against A. baumannii, with MIC and MBC values of 6.25 µM and 12.5 µM, respectively, thereby identifying it as a bactericidal contender.
Hybrids 29 and 31 displayed outstanding MIC value of 6.25 µM and MBC value of 12.5 µM and 25 µM, respectively, against MSSA. Hybrid 28, however, demonstrated MIC and MBC values of 12.5 and 25 µM, respectively. MIC/MBC ratio confirmed hybrids 28 and 29 as bactericidal and 31 as bacteriostatic against MSSA.
With the exception of 28 and 31, the assessed hybrids outperformed fluconazole in opposition to C. albicans, according to an evaluation of their fungistatic/fungicidal activities. With MBC values of 50 µM and MIC values of 12.5 µM, hybrids 25 and 29 indicated the strongest anti-fungal activity against C. albicans; nevertheless, the MFC to MIC Rate deemed that they were fungistatic.
| Cpd. no. | DNA gyrase supercoiling IC50 (µM) |
|---|---|
| 24 | 172.7 ± 15.1 |
| 25 | 77.3 ± 15.5 |
| 26 | 144.7 ± 15.2 |
| 27 | 200.8 ± 16.4 |
| 28 | 87.6 ± 16.0 |
| 29 | 182.2 ± 16.5 |
| 30 | 147.8 ± 15.6 |
| 31 | 65.5 ± 16.8 |
| Ciprofloxacin | 143.1 ± 11.2 |
2.3. The study of the structure–activity relationship (SAR)
This work exploited the novel compounds' biological processes to understand the connection between structure and the action (Fig. 5). The impact of the most potent chemicals on DNA Gyrase inhibitory verified that the 3-chloro-quinoxalinyl bearing hybrids are generally more active than 3-hydroxy-quinoxalinyl derivatives. Substitution on the ring of the N-phenyl demonstrated a pattern in the inhibitory effect of DNA gyrase, where electron-withdrawing groups on the para position demonstrated superior activity to Ciprofloxacin as observed with 28 and 31. The remaining 3-chloro-quinoxalinyl hybrids bearing electron-donating groups on the N-phenyl ring (30) and (29) demonstrated DNA gyrase inhibitory activity either equipotent or lower than Ciprofloxacin, respectively. This finding confirms the importance of bearing electron-withdrawing groups at both R and R1. Most importantly, Presence of 3-hydroxy substitution on the N-phenyl ring generally resulted in lower DNA gyrase inhibitory activity, however equipotent to Ciprofloxacin. Such a trend was not noted in hybrid 26, bearing two hydroxyl groups. A plausible explanation for this exception suggests the formation of an extra compensatory hydrogen bond with DNA gyrase. It was noted, however that hybrids bearing both R groups as electron-donating still exhibited equipotent or superior DNA gyrase inhibitory activity to Ciprofloxacin (hybrids 26 and 25, respectively).2.4. In silico ADME
To assess the physicochemical characteristics of the produced hybrids in comparison to levofloxacin, amoxicillin, and fluconazole, a simulated study was conducted.68–70 In terms of their physicochemical characteristics, all compounds and reference drugs—aside from 27—display zero breaches of the Lipinski's requirements regarding oral medications, as stated by Table 5. With the exception of 27 and amoxicillin, all examined derivatives and references adhere to Veber's requirements. Additionally, all hybrids exhibit high flexibility with no more than six rotatable bonds. This implies that oral consumption of the eight hybrids is possible. Each analogue was given a score between 3.05 and 3.76 for ease of manufacturing, suggesting that a large number of them can be produced on a large scale.| Cpd. no. | HBD | HBA | MW | M![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
No. of rotatable bonds | Lipinski's violations | Veber's violations | Synthetic accessibility |
|---|---|---|---|---|---|---|---|---|
| 24 | 3 | 6 | 388.38 | 1.09 | 5 | 0 | 0 | 3.64 |
| 25 | 3 | 5 | 391.42 | 2.16 | 5 | 0 | 0 | 3.76 |
| 26 | 4 | 6 | 379.37 | 1.2 | 5 | 0 | 0 | 3.67 |
| 27 | 3 | 7 | 408.37 | 0.88 | 6 | 1 | 1 | 3.71 |
| 28 | 2 | 5 | 406.83 | 1.68 | 5 | 0 | 0 | 3.05 |
| 29 | 2 | 4 | 409.87 | 2.76 | 5 | 0 | 0 | 3.31 |
| 30 | 3 | 5 | 397.82 | 1.8 | 5 | 0 | 0 | 3.30 |
| 31 | 2 | 6 | 426.81 | 1.47 | 6 | 0 | 0 | 3.49 |
| Levofloxacin | 2 | 7 | 379.38 | 0.2 | 2 | 0 | 0 | 3.74 |
| Amoxicillin | 4 | 6 | 365.40 | 0.23 | 5 | 0 | 1 | 4.17 |
| Fluconazole | 1 | 7 | 306.27 | 1.47 | 5 | 0 | 0 | 2.45 |
| Ciprofloxacin | 2 | 5 | 331.34 | 1.28 | 3 | 0 | 0 | 2.51 |
Additionally, the percentage was computed using TPSA as follows: % ABS = 109 − (0.354 × TPSA).71,72 All of the prospective antibacterial agents possess estimated absorption percentages between 87 and 154%, which indicates strong oral bioavailability analogous to reference medications. The percentage of ABS ranges from 56 to 79% (Table 6).
| Cpd. no. | TPSA | % ABS |
|---|---|---|
| 24 | 131.240 | 63.720 |
| 25 | 107.450 | 71.920 |
| 26 | 127.680 | 64.950 |
| 27 | 153.270 | 56.120 |
| 28 | 111.010 | 70.700 |
| 29 | 87.220 | 78.900 |
| 30 | 107.450 | 71.920 |
| 31 | 133.040 | 63.100 |
| Levofloxacin | 84.240 | 79.180 |
| Amoxicillin | 158.260 | 52.980 |
| Fluconazole | 81.650 | 80.090 |
| Ciprofloxacin | 74.570 | 82.600 |
The findings of monitoring the new antibacterial hybrids for pharmacokinetic and medicinal chemistry characteristics suggest that, with the exception of 27 and amoxicillin, every other compound has demonstrated adequate GI tract absorption. They can't overcome all, however, overcome the brain–blood barrier. Every hybrid has a high probability of effluxing outside the cell and is a P-gp (P-glycoprotein) substrate.
Furthermore, inhibition of metabolic enzymes by investigated hybrids was computationally assessed. Predictions revealed that hybrid 26 did not inhibit any of the assessed metabolic enzymes, whereas 27 only inhibited CYP2C9.
All investigated hybrids received a bioavailability score of 0.55, with the exception of derivative 24, which received a score of 0.56, analogous to the other reference drugs. We assessed every hit, and PAINS showed no warnings. In order to avoid inaccurate outcomes, PAINS are important considerations while developing drugs (Table 7). The evaluated substances' bioavailability radar is illustrated within Fig. 6. The intended hybrids revealed promise as attractive agents for antibacterial treatment by exhibiting typically very good physicochemical properties.
| Cpd. no. | GI absorption | BBB permeation | Pgp substrate | CYP1A2 inhibitor | CYP2C19 inhibitor | CYP2C9 inhibitor | CYP2D6 inhibitor | CYP3A4 inhibitor | Bioavailability score | PAINS alerts |
|---|---|---|---|---|---|---|---|---|---|---|
| 24 | High | No | Yes | Yes | No | Yes | No | No | 0.56 | 0 |
| 25 | High | No | Yes | Yes | No | Yes | Yes | Yes | 0.55 | 0 |
| 26 | High | No | Yes | No | No | No | No | No | 0.55 | 0 |
| 27 | Low | No | Yes | No | No | Yes | No | No | 0.55 | 0 |
| 28 | High | No | Yes | Yes | No | Yes | Yes | Yes | 0.55 | 0 |
| 29 | High | No | Yes | Yes | Yes | Yes | Yes | Yes | 0.55 | 0 |
| 30 | High | No | Yes | Yes | No | Yes | Yes | Yes | 0.55 | 0 |
| 31 | High | No | Yes | No | Yes | Yes | No | Yes | 0.55 | 0 |
| Levofloxacin | High | No | Yes | No | No | No | No | No | 0.55 | 0 |
| Amoxicillin | Low | No | No | No | No | No | No | No | 0.55 | 0 |
| Fluconazole | High | No | Yes | No | Yes | No | No | No | 0.55 | 0 |
| Ciprofloxacin | High | No | Yes | No | No | No | No | No | 0.55 | 0 |
2.5. Toxicity assessment in silico
The pkCSM online tools confirm the security of the most active hits since, as shown in Table 8, with the exception of 31, none have demonstrated the potential for mutagenicity according to AMES toxicity, equivalent to that of Levofloxacin and Fluconazole. Furthermore, compared to Fluconazole (0.114![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log mg per kg per day) and other hits, hybrid 26 demonstrated a maximum acceptable human dose of 0.261
log mg per kg per day) and other hits, hybrid 26 demonstrated a maximum acceptable human dose of 0.261![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log mg per kg per day. The LD50 value of compound 31 has been estimated to be 2.587 mol kg−1, which is considerably higher than that of Fluconazole and equivalent to that of Levofloxacin. Additionally, compound 26 had an LOAEL value of 2.798
log mg per kg per day. The LD50 value of compound 31 has been estimated to be 2.587 mol kg−1, which is considerably higher than that of Fluconazole and equivalent to that of Levofloxacin. Additionally, compound 26 had an LOAEL value of 2.798![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log mg per kg bw per day, surpassing the values of all hits and the reference drugs. The potassium channels encoded by the hERG gene control the repolarization of cardiac action potentials. Pharmaceutical companies are concerned because blocking. These channels may cause cardiovascular problems that could be fatal. While every potent ingredient was expected to have an inhibitory impact on hERG-II, it was shown that they had no effect on hERG-I.
log mg per kg bw per day, surpassing the values of all hits and the reference drugs. The potassium channels encoded by the hERG gene control the repolarization of cardiac action potentials. Pharmaceutical companies are concerned because blocking. These channels may cause cardiovascular problems that could be fatal. While every potent ingredient was expected to have an inhibitory impact on hERG-II, it was shown that they had no effect on hERG-I.
| Test | Most active compounds | |||||
|---|---|---|---|---|---|---|
| 25 | 26 | 30 | 31 | Levofloxacin | Fluconazole | |
| pkCSM73 | ||||||
| AMES toxicity | No | No | No | Yes | No | No |
| Max. tolerated dose (human) | −0.11 | 0.261 | 0.144 | 0.021 | 0.965 | 0.114 |
| hERG I inhibitor | No | No | No | No | No | No |
| hERG II inhibitor | Yes | Yes | Yes | Yes | No | No |
| Oral rat acute toxicity (LD50) | 2.046 | 2.089 | 2.116 | 2.587 | 2.59 | 2.328 |
| Oral rat chronic toxicity (LOAEL) | 1.504 | 2.798 | 2.046 | 2.07 | 1.791 | 1.033 |
| Hepatotoxicity | Yes | Yes | Yes | Yes | Yes | Yes |
| Skin sensitization | No | No | No | No | No | No |
| T. Pyriformis toxicity | 0.405 | 0.346 | 0.357 | 0.326 | 0.285 | 0.312 |
| Minnow toxicity | 0.812 | 1.777 | 0.187 | −0.601 | 1.273 | 3.872 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| ProTox-II prediction74 | ||||||
| LD50 (mg kg−1) | 2000 | 1600 | 1600 | 2000 | 1478 | 1271 |
| Toxicity class | IV | IV | IV | IV | IV | IV |
| Neurotoxicity | +(0.72 | +(0.73) | +(0.9) | +(0.57) | +(0.93) | +(0.91) |
| Nephrotoxicity | −(0.53) | −(0.53) | +(0.58) | −(0.54) | +(0.85) | −(0.61) |
| Immunotoxicity | −(0.99) | −(0.98) | −(0.96) | −(0.98) | −(0.99) | −(0.83) |
| Mutagenicity | −(0.57) | −(0.58) | −(0.54) | +(0.79) | +0.66) | −(0.52) |
| Cytotoxicity | −(0.57) | −(0.6) | −(0.6) | −(0.53) | −(0.69) | −(0.76) |
| Clinical toxicity | +(0.66) | +(0.68) | +(0.73) | +(0.58) | +(0.62) | +(0.55) |
| Phosphoprotein (tumor suppressor) p53 | −(0.88) | −(0.88) | −(0.87) | −(0.86) | −(0.98) | −(0.97) |
| Voltage gated sodium channel (VGSC) | −(0.89) | −(0.76) | −(0.7) | −(0.82) | −(0.95) | −(0.95) |
The pkCSM server was used to examine hepatotoxicity. All active hits, including reference medications, were predicted to be hepatotoxic. Skin discomfort was not evident in any of the hits. It was expected that Levofloxacin, Fluconazole, and active hybrids would be categorized as a class IV (GHS) utilizing the-ProTox-II identification webtool (Table 8). It was discovered that all active hybrids had LD50 values exceeding the reference medications. None of the active hits were supposed to be immunotoxic, cytotoxic, or mutagenic. The active hybrids and reference drugs were expected to exhibit both neurotoxicity and clinical toxicity. Neither the voltage-gated sodium channel (VGSC) nor the phosphoprotein (tumor suppressor) p53 were suppressed by those active hybrids.
2.6. Molecular docking
The investigated hybrids were docked against DNA gyrase (PDB ID: 8BN6) to predict binding mode and interactions. The bound ligand was separately redocked to validate the docking procedure Validation was confirmed by visualizing the overlay of the original and the redocked pose (Fig. 7) and calculated RMSD value of 0.79 Å. The investigated hybrids were docked implementing the same parameters. The 2D interactions and energy scores are recorded (Table 9). | ||
| Fig. 7 (A) Overlay of bound ligand (cyan) and redocked ligand (yellow) and bound ligand interactions, (B) binding mode and ligand interactions of Ciprofloxacin. | ||
| Cpd. no. | S (kcal mol−1) | Type of interaction | Amino acid | Length (Å) | Cpd. no. | S (kcal mol−1) | Type of interaction | Amino acid | Length (Å) |
|---|---|---|---|---|---|---|---|---|---|
| 24 | −8.35 | Amide–Pi stacked | Asn48 | 5.01 | 28 | −8.05 | Pi–alkyl | Ile80 | 4.44 |
| Amide–Pi stacked | Asn48 | 4.66 | Pi–alkyl | Ile80 | 4.51 | ||||
| Pi–alkyl | Ile80 | 4.41 | Amide–Pi stacked | Gly79 | 4.50 | ||||
| Pi–alkyl | Ile80 | 4.40 | H-bond | Val122 | 4.85 | ||||
| H-bond | Thr167 | 2.17 | Pi–alkyl | Ile96 | 5.55 | ||||
| H-bond | Asp75 | 1.97 | Pi–alkyl | Ile96 | 7.25 | ||||
| Amide–Pi stacked | Gly79 | 5.10 | |||||||
| Pi–alkyl | Ile96 | 4.78 | |||||||
| Pi–alkyl | Val122 | 5.35 | |||||||
| H-bond | Arg138 | 2.96 | |||||||
| 25 | −8.12 | Amide–Pi stacked | Asn48 | 4.64 | 29 | −7.66 | Alkyl | Ile96 | 5.35 |
| Amide–Pi stacked | Asn48 | 5.00 | Alkyl | Pro81 | 4.41 | ||||
| Pi–alkyl | Ile80 | 4.41 | H-bond | Arg138 | 2.82 | ||||
| Pi–alkyl | Ile80 | 4.44 | Pi–alkyl | Ile80 | 4.25 | ||||
| H-bond | Asp75 | 1.95 | Pi–alkyl | Ile80 | 4.23 | ||||
| H-bond | Asp75 | 3.12 | Amide–Pi stacked | Gly79 | 4.80 | ||||
| Amide–Pi stacked | Gly79 | 5.12 | |||||||
| Pi–alkyl | Ile96 | 4.46 | |||||||
| Alkyl | Ile96 | 4.52 | |||||||
| H-bond | Thr167 | 2.16 | |||||||
| Pi–alkyl | Val122 | 5.32 | |||||||
| 26 | −7.81 | H-bond | Arg138 | 2.23 | 30 | −10.20 | H-bond | Asp51 | 1.96 |
| H-bond | Asp75 | 2.79 | Amide–Pi stacked | Asn48 | 5.07 | ||||
| H-bond | Gly79 | 2.27 | Amide–Pi stacked | Asn48 | 4.69 | ||||
| Amide–Pi-stacked | Gly79 | 4.36 | Pi–alkyl | Val122 | 5.25 | ||||
| Pi–alkyl | Ile80 | 4.10 | Pi–alkyl | Val169 | 5.43 | ||||
| Pi–alkyl | Ile80 | 4.37 | Pi–alkyl | Ile80 | 5.32 | ||||
| H-bond | Glu52 | 2.85 | Pi–alkyl | Ile80 | 4.29 | ||||
| 27 | −7.41 | Pi–alkyl | ILE96 | 4.48 | 31 | −10.80 | H-bond | GLU52 | 3.07 |
| Pi–alkyl | VAL122 | 5.34 | Pi–alkyl | ILE80 | 4.34 | ||||
| H-bond | ARG138 | 3.06 | Pi–alkyl | ILE80 | 5.42 | ||||
| Amide Pi-stacked | ASN48 | 4.99 | Pi–alkyl | VAL169 | 5.35 | ||||
| Amide Pi-stacked | ASN48 | 4.65 | Amide Pi-stacked | ASN48 | 5.08 | ||||
| Pi–alkyl | ILE80 | 4.43 | Amide Pi-stacked | ASN48 | 4.65 | ||||
| Pi–alkyl | ILE80 | 4.41 | Pi–alkyl | VAL122 | 5.22 | ||||
| H-bond | ASP75 | 3.12 | Pi–alkyl | ILE96 | 4.66 | ||||
| H-bond | ASP75 | 1.95 | |||||||
| Amide Pi-stacked | GLY79 | 5.11 | |||||||
| H-bond | THR167 | 2.16 | |||||||
| Co-crystalized | −9.34 | H-bond | ARG138 | 2.17 | Ciprofloxacin | −9.70 | H-bond | ARG138 | 7.22 |
| Pi–cation | ARG78 | 4.05 | Pi–anion | GLU52 | 6.62 | ||||
| Pi–alkyl | PRO81 | 4.21 | Pi–anion | GLU52 | 5.34 | ||||
| Pi–alkyl | PRO81 | 4.37 | Pi–alkyl | ILE80 | 4.88 | ||||
| Pi–anion | GLU52 | 5.18 | Alkyl | ILE80 | 5.73 | ||||
| Pi–anion | GLU52 | 3.78 | H-bond | THR167 | 3.93 | ||||
| Pi–alkyl | ILE96 | 5.23 | Halogen | ASP75 | 5.58 | ||||
| H-bond | THR167 | 2.94 | Amide–pi stacked | ASN48 | 6.59 | ||||
| Alkyl | VAL73 | 4.87 | Alkyl | ILE96 | 4.64 | ||||
| Amide–Pi stacked | ASN48 | 4.50 | H-bond | ARG78 | 5.40 | ||||
| Alkyl | VAL169 | 4.95 | |||||||
| Alkyl | VAL169 | 4.41 | |||||||
| Alkyl | VAL45 | 5.03 | |||||||
| Alkyl | VAL45 | 4.18 | |||||||
| Alkyl | VAL122 | 3.77 |
The interaction of co-crystalized ligand with DNA Gyrase involved six alkyl-type van der Waals interactions with Val73, Val169, Val45 and Val122. Two H-bonds were noted between the co-crystalized ligand and key amino acids: Arg138 and Thr167. Three pi–alkyl interactions were observed with Pro81 and Ile96. Glu52 interacted with the ligand through two pi–anion interactions, whereas Arg78 interacted through one pi–cation interactions. Amide-pi stacked interaction was involved with Asn48.
Docking of ciprofloxacin as a reference against DNA Gyrase demonstrated that the binding energy was −9.70 kcal per mol. Ciprofloxacin was able to interact with key amino acids; including three H-bonds with The167, Arg138 and Arg78, two alkyl-type van der Waals interactions with Ile96 and Ile80, one pi–alkyl interaction with Ile80, one amide–pi stacked interaction with Asn48, and two pi–anion interactions with Glu52.
Simulation of the hybrid 25 binding showed that the binding energy was −8.12 kcal mol−1. Key amino acids were involved in the interaction of 25 with DNA Gyrase. Such interactions included two amide-Pi stacked interactions with Asn48, two pi–alkyl-type van der Waals interactions with Ile96 and Val122, one H-bond with Thr167 and one alkyl-type van der Waals with Ile96. Additional interactions included two pi–alkyl interactions with Ile80, two H-bonds with Asp75 and one amide–pi stacked interaction with Gly79.
Binding simulation of hybrid 28 demonstrated a binding energy of −8.05 kcal mol−1. Key amino acids were involved in the interaction of 28 with DNA Gyrase. Such interactions included two pi-alkyl-type van der Waals interactions with Ile96, one H-bond with Val122. Additional interactions involved two pi–alkyl interactions with Ile 80, and one amide–pi stacked interaction with Gly79.
Binding simulation of hybrid 31 revealed that the binding energy score was −10.80 kcal mol−1. Although docking scores do not strictly follow experimental IC50 values, the binding energy of 31 still confirms superiority to Ciprofloxacin in silico. Key amino acids were involved in the interaction of 31 with DNA Gyrase. Such interactions included three pi–alkyl-type van der Waals interactions with Val169, Val122 and Ile96, one H-bond with Glu52, and two amide–pi stacked interactions with Asn48. Additional interactions involved two pi–alkyl interactions with Ile80.
The binding mode and energies of the remaining hybrids are noted in (Table 9, Fig. 8 and 9).
3. Experimental
3.1. Chemistry
3.1.1.1 Method A. In a flask with a round bottom, 1 mmol (0.13 g) of itaconic acid (1) was mixed with 1 mmol of each of the respective amines 8–15 with water. The mixture of the reaction underwent reflux heating for 24 hours. The resulting precipitates were filtered recrystallized from ethanol.75–78
3.1.1.2 Method B. In a beaker of 250 mL, 1 mmol (0.13 g) itaconic acid (1) was dissolved in the least amount of ethanol and mixed with 1 mmol of each of the respective amines 8–15. The reaction solution was sonicated until reaction completion, monitored by TLC.
3.1.1.3 Method C. In a 250 mL beaker, 1 mmol (0.13 g) itaconic acid (1) was dissolved in the least amount of ethanol and mixed with 1 mmol of each of the respective amines 8–15. The reaction solution was microwaved in domestic microwave until reaction completion, monitored by TLC.
3.1.1.4 1-(3-Chlorophenyl)-5-oxopyrrolidine-3-carboxylic acid, 8. Yellowish powder; yield: 91%, m.p. 187–189 °C; 1H NMR (400 MHz, CDCl3): δ = 12.38 (s, 1H, COOH), 7.28–7.24 (m, 3H, A–H), 7.20–7.15 (m, 1H, A–H), 4.19 (dd, J = 11.9, 4.4 Hz, 1H, pyrrolidine CH2), 3.93 (dd, J = 11.9, 4.4 Hz, 1H, pyrrolidine CH2), 3.33–3.24 (m, 1H, pyrrolidine CH), 2.78 (dd, J = 15.0, 5.6 Hz, 1H, pyrrolidine CH2), 2.53 (dd, J = 15.0,5.6 Hz, 1H, pyrrolidine CH2).
3.1.1.5 1-(4-Chlorophenyl)-5-oxopyrrolidine-3-carboxylic acid, 9. Yellowish powder; yield: 95%, m.p. 184–185 °C; 1H NMR (400 MHz, CDCl3): δ = 12.14 (s, 1H, COOH), 7.31–7.22 (m, 2H, A–H), 7.08–7.00 (m, 2H, A–H), 4.15 (dd, J = 11.8, 4.5 Hz, 1H, pyrrolidine CH2), 3.90 (dd, J = 11.8, 4.2 Hz, 1H, pyrrolidine CH2), 3.37–3.27 (m, 1H, pyrrolidine CH), 2.76 (dd, J = 14.9, 5.5 Hz, 1H, pyrrolidine CH2), 2.51 (dd, J = 14.9,5.5 Hz, 1H, pyrrolidine CH2).
3.1.1.6 1-(4-Cyanophenyl)-5-oxopyrrolidine-3-carboxylic acid, 10. Orange powder; yield: 90%, m.p. 181–183 °C; 1H NMR (400 MHz,CDCl3): δ = 12.38 (s, 1H, COOH), 7.58 (d, J = 7.5 Hz, 2H, A–H), 7.30 (d, J = 7.1 Hz, 2H, A–H), 4.17 (dd, J = 11.7, 4.2 Hz, 1H, pyrrolidine CH2), 3.92 (dd, J = 12.0,4.4 Hz, 1H, pyrrolidine CH2), 3.29 (p, J = 4.9 Hz, 1H, pyrrolidine CH), 2.78 (dd, J = 14.8,5.6 Hz, 1H, pyrrolidine CH2), 2.53 (dd, J = 14.8,5.6 Hz, 1H, pyrrolidine CH2).
3.1.1.7 1-(2,5-Dimethylphenyl)-5-oxopyrrolidine-3-carboxylic acid, 11. Ligh yellow powder; yield: 92%, m.p. 196–198 °C; 1H NMR (400 MHz, CDCl3): δ = 12.13 (s, 1H, COOH), 6.97 (d, J = 2.8 Hz, 1H, A–H), 6.87 (d, J = 8.5 Hz, 1H, A–H), 6.72 (dd, J = 8.6,2.9 Hz, 1H, A–H), 4.32 (dd, J = 11.6, 4.3 Hz, 1H, pyrrolidine CH2), 4.07 (dd, J = 11.6, 4.3 Hz, 1H, pyrrolidine CH2), 3.04–2.93 (m, 1H, pyrrolidine CH), 2.70 (dd, J = 15.0, 5.6 Hz, 1H, pyrrolidine CH2), 2.44 (dd, J = 14.9, 5.6 Hz, 1H, pyrrolidine CH2), 2.17 (s, 3H, CH3), 2.09 (s, 3H, CH3).
3.1.1.8 1-(3-Hydroxyphenyl)-5-oxopyrrolidine-3-carboxylic acid, 12. Yellowish powder; yield: 89%, m.p. 179–181 °C; 1H NMR (400 MHz,CDCl3): δ = 12.38 (s, 1H, COOH), 7.98 (s, 1H, OH), 7.11 (t, J = 7.9 Hz, 1H, A–H), 7.02 (d, J = 7.1 Hz, 1H, A–H), 6.65 (d, J = 8.3 Hz, 1H, A–H), 6.59 (d, J = 2.6 Hz, 1H, A–H), 4.19 (dd, J = 11.9, 4.3 Hz, 1H, pyrrolidine CH2), 3.93 (dd, J = 11.9, 4.4 Hz, 1H, pyrrolidine CH2), 3.29 (p, J = 4.8 Hz, 1H, pyrrolidine CH), 2.78 (dd, J = 15.0, 5.4 Hz, 1H, pyrrolidine CH2), 2.53 (dd, J = 15.1, 5.6 Hz, 1H, pyrrolidine CH2).
3.1.1.9 1-(4-Nitrophenyl)-5-oxopyrrolidine-3-carboxylic acid, 13. Orange powder; yield: 88%, m.p. 198–200 °C; 1H NMR (400 MHz, CDCl3): δ = 13.14 (s, 1H, COOH), 8.05 (d, J = 7.2 Hz, 2H, A–H), 7.43 (d, J = 7.4 Hz, 2H, A–H), 3.98 (dd, J = 11.9, 4.4 Hz, 1H, pyrrolidine CH2), 3.73 (dd, J = 11.8, 4.3 Hz, 1H, pyrrolidine CH2), 3.38 (p, J = 4.8 Hz, 1H, pyrrolidine CH), 2.79 (dd, J = 14.9, 5.6 Hz, 1H, pyrrolidine CH2), 2.54 (dd, J = 14.8, 5.6 Hz, 1H, pyrrolidine CH2).
3.1.1.10 1-(Naphthalen-1-yl)-5-oxopyrrolidine-3-carboxylic acid, 14. Light yellow powder; yield: 92%, m.p. 195–196 °C; 1H NMR (400 MHz, CDCl3): δ = 12.13 (s, 1H, COOH), 7.78–7.64 (m, 3H, A–H), 7.42–7.22 (m, 3H, A–H), 7.14 (d, J = 7.8 Hz, 1H, A–H), 4.32 (dd, J = 11.5, 4.3 Hz, 1H, pyrrolidine CH2), 4.06 (dd, J = 11.6, 4.4 Hz, 1H, pyrrolidine CH2), 2.99 (p, J = 4.6 Hz, 1H, pyrrolidine CH), 2.70 (dd, J = 15.0, 5.7 Hz, 1H, pyrrolidine CH2), 2.45 (dd, J = 15.0, 5.6 Hz, 1H, pyrrolidine CH2).
3.1.1.11 5-Oxo-1-phenylpyrrolidine-3-carboxylic acid, 15. Yellow powder; yield: 90%, m. p. 186–187 °C; 1H NMR (400 MHz,CDCl3): δ = 12.38 (s, 1H, COOH), 7.36–7.15 (m, 5H, A–H), 4.17 (dd, J = 11.9, 4.2 Hz, 1H, pyrrolidine CH2), 3.92 (dd, J = 11.9, 4.4 Hz, 1H, pyrrolidine CH2), 3.33–3.24 (m, 1H, pyrrolidine CH), 2.78 (dd, J = 15.0, 5.6 Hz, 1H, pyrrolidine CH2), 2.53 (dd, J = 15.0, 5.6 Hz, 1H, pyrrolidine CH2).
3.1.2.1 1-(4-Cyanophenyl)-N′-(3-hydroxyquinoxalin-2-yl)-5-oxopyrrolidine-3-carbohydrazide, 24. Orange powder; yield: (59%), m.p. 264–265 °C; IR (KBr, cm−1) = 3474–3359 (2NH & OH), 3073 (CH aromatic), 2214 (CN), 1687 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, CDCl3): δ = 12.87 (s, 1H, OH), 11.04 (d, J = 6.6 Hz, 1H, –NH–NH–CO), 8.18 (d, J = 6.8 Hz, 1H, NH–NH–CO), 8.05 (dd, J = 7.7, 2.0 Hz, 1H, A–H), 7.73–7.52 (m, 5H, A–H), 7.30 (d, J = 7.8 Hz, 2H, A–H), 4.17 (dd, J = 12.4, 3.6 Hz, 1H, pyrrolidine CH2), 3.92 (dd, J = 12.5, 3.5 Hz, 1H, pyrrolidine CH2), 3.12 (m, 1H, pyrrolidine CH), 2.52 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.33 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.77 (C
O); 1H NMR (400 MHz, CDCl3): δ = 12.87 (s, 1H, OH), 11.04 (d, J = 6.6 Hz, 1H, –NH–NH–CO), 8.18 (d, J = 6.8 Hz, 1H, NH–NH–CO), 8.05 (dd, J = 7.7, 2.0 Hz, 1H, A–H), 7.73–7.52 (m, 5H, A–H), 7.30 (d, J = 7.8 Hz, 2H, A–H), 4.17 (dd, J = 12.4, 3.6 Hz, 1H, pyrrolidine CH2), 3.92 (dd, J = 12.5, 3.5 Hz, 1H, pyrrolidine CH2), 3.12 (m, 1H, pyrrolidine CH), 2.52 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.33 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.77 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 171.24 (C
O), 171.24 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 146.52, 141.54, 140.22, 136.97, 136.51, 132.41, 128.93, 126.45, 125.38, 123.42, 118.23 (CN), 117.28, 102.21, 48.51 (pyrrolidine CH2), 38.02 (pyrrolidine CH), 36.42 (pyrrolidine CH2); MS(EI, 70 eV): m/z (%) = 388 [M˙+] (18%), 356.46 (100%); anal. calc. for: C20H16N6O3 (388.38): C, 61.85; H, 4.15; N, 21.64; found: C, 61.80; H, 4.17; N, 21.68.
O), 146.52, 141.54, 140.22, 136.97, 136.51, 132.41, 128.93, 126.45, 125.38, 123.42, 118.23 (CN), 117.28, 102.21, 48.51 (pyrrolidine CH2), 38.02 (pyrrolidine CH), 36.42 (pyrrolidine CH2); MS(EI, 70 eV): m/z (%) = 388 [M˙+] (18%), 356.46 (100%); anal. calc. for: C20H16N6O3 (388.38): C, 61.85; H, 4.15; N, 21.64; found: C, 61.80; H, 4.17; N, 21.68.
3.1.2.2 1-(2,5-Dimethylphenyl)-N′-(3-hydroxyquinoxalin-2-yl)-5-oxopyrrolidine-3-carbohydrazide, 25. Light brown powder; yield: (77%), m.p. 271–272 °C; IR (KBr, cm−1) = 3369–3280 (br. 2NH & OH), 3060 (CH aromatic), 2971, 2860 (CH aliphatic), 1698 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O; 1H NMR (400 MHz,CDCl3): δ = 12.86 (s, 1H, OH), 10.79 (d, J = 6.8 Hz, 1H, –NH–NH–CO), 8.88 (d, J = 6.6 Hz, 1H, NH–NH–CO), 7.91 (dd, J = 8.0, 1.6 Hz, 1H, A–H), 7.68 (dd, J = 8.0, 1.8 Hz, 1H, A–H), 7.65–7.52 (m, 2H, A–H), 7.12 (d, J = 8.2 Hz, 1H, A–H), 6.98 (d, J = 8.2 Hz, 2H, A–H), 4.21 (dd, J = 12.2, 3.6 Hz, 1H, pyrrolidine CH2), 3.96 (dd, J = 12.3, 3.7 Hz, 1H, pyrrolidine CH2), 3.34–3.25 (m, 1H, pyrrolidine CH), 2.75 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.53 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.28 (s, 3H, CH3), 2.10 (s, 3H, CH3); 13C NMR (100 MHz,CDCl3): δ = 172.15 (C
O; 1H NMR (400 MHz,CDCl3): δ = 12.86 (s, 1H, OH), 10.79 (d, J = 6.8 Hz, 1H, –NH–NH–CO), 8.88 (d, J = 6.6 Hz, 1H, NH–NH–CO), 7.91 (dd, J = 8.0, 1.6 Hz, 1H, A–H), 7.68 (dd, J = 8.0, 1.8 Hz, 1H, A–H), 7.65–7.52 (m, 2H, A–H), 7.12 (d, J = 8.2 Hz, 1H, A–H), 6.98 (d, J = 8.2 Hz, 2H, A–H), 4.21 (dd, J = 12.2, 3.6 Hz, 1H, pyrrolidine CH2), 3.96 (dd, J = 12.3, 3.7 Hz, 1H, pyrrolidine CH2), 3.34–3.25 (m, 1H, pyrrolidine CH), 2.75 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.53 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.28 (s, 3H, CH3), 2.10 (s, 3H, CH3); 13C NMR (100 MHz,CDCl3): δ = 172.15 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 171.54 (C
O), 171.54 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 145.99, 140.73, 137.35, 136.84, 135.02, 132.42, 129.63, 129.36, 129.05, 126.42, 125.73, 123.93, 123.84, 121.82, 49.31 (pyrrolidine CH2), 38.12 (pyrrolidine CH), 36.32 (pyrrolidine CH2), 19.52 (CH3), 16.53 (CH3); MS (EI, 70 eV): m/z (%) = 391.94 [M˙+] (16.89%), 74.07 (100%); anal. calc. for: C21H21N5O3 (391.42): C, 64.44; H, 5.41; N, 17.89; found: C, 64.48; H, 5.38; N, 17.85.
O), 145.99, 140.73, 137.35, 136.84, 135.02, 132.42, 129.63, 129.36, 129.05, 126.42, 125.73, 123.93, 123.84, 121.82, 49.31 (pyrrolidine CH2), 38.12 (pyrrolidine CH), 36.32 (pyrrolidine CH2), 19.52 (CH3), 16.53 (CH3); MS (EI, 70 eV): m/z (%) = 391.94 [M˙+] (16.89%), 74.07 (100%); anal. calc. for: C21H21N5O3 (391.42): C, 64.44; H, 5.41; N, 17.89; found: C, 64.48; H, 5.38; N, 17.85.
3.1.2.3 1-(3-Hydroxyphenyl)-N′-(3-hydroxyquinoxalin-2-yl)-5-oxopyrrolidine-3-carbohydrazide, 26. Dark yellow powder; yield: (62%), m.p. 243–244 °C; IR (KBr, cm−1) = 3422–3269 (br. 2NH & 2OH), 3070 (CH aromatic), 1656–1703 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O; 1H NMR (400 MHz,CDCl3): δ = 11.03 (s, 1H, OH), 8.80 (d, J = 6.6 Hz, 1H, –NH–NH–CO), 8.60 (s, 1H, OH), 8.17 (d, J = 8.1 Hz, 1H, A–H), 7.73 (dd, J = 8.1, 1.6 Hz, 1H, 1H, A–H), 7.70–7.54 (m, 2H, 1H, A–H), 7.09 (t, J = 7.9 Hz, 1H, 1H, A–H), 6.83 (d, J = 6.9 Hz, 1H, A–H), 6.68 (d, J = 8.6 Hz, 1H, A–H), 6.54 (t, J = 2.3 Hz, 1H), 5.90 (d, J = 6.8 Hz, 1H, –NH–NH–CO), 3.90 (dd, J = 12.5, 3.5 Hz, 1H, pyrrolidine CH2), 3.65 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.08–3.00 (m, 1H, pyrrolidine CH), 2.54 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.40 (dd, J = 15.8, 4.8 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 173.48 (C
O; 1H NMR (400 MHz,CDCl3): δ = 11.03 (s, 1H, OH), 8.80 (d, J = 6.6 Hz, 1H, –NH–NH–CO), 8.60 (s, 1H, OH), 8.17 (d, J = 8.1 Hz, 1H, A–H), 7.73 (dd, J = 8.1, 1.6 Hz, 1H, 1H, A–H), 7.70–7.54 (m, 2H, 1H, A–H), 7.09 (t, J = 7.9 Hz, 1H, 1H, A–H), 6.83 (d, J = 6.9 Hz, 1H, A–H), 6.68 (d, J = 8.6 Hz, 1H, A–H), 6.54 (t, J = 2.3 Hz, 1H), 5.90 (d, J = 6.8 Hz, 1H, –NH–NH–CO), 3.90 (dd, J = 12.5, 3.5 Hz, 1H, pyrrolidine CH2), 3.65 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.08–3.00 (m, 1H, pyrrolidine CH), 2.54 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.40 (dd, J = 15.8, 4.8 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 173.48 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 172.48 (C
O), 172.48 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 158.81, 147.23, 147.07, 140.85, 136.93, 135.59, 131.70, 131.16, 129.32, 125.08, 122.44, 117.35, 109.00, 106.54, 49.02 (pyrrolidine CH2), 40.83 (pyrrolidine CH), 38.00 (pyrrolidine CH2); MS (EI, 70 eV): m/z (%) = 379.40 [M˙+] (19.76%), 190.16 (100%); anal. calc. for: C19H17N5O4 (379.37): C, 60.15; H, 4.52; N, 18.46; found: C, 60.19; H, 4.50; N, 18.48.
O), 158.81, 147.23, 147.07, 140.85, 136.93, 135.59, 131.70, 131.16, 129.32, 125.08, 122.44, 117.35, 109.00, 106.54, 49.02 (pyrrolidine CH2), 40.83 (pyrrolidine CH), 38.00 (pyrrolidine CH2); MS (EI, 70 eV): m/z (%) = 379.40 [M˙+] (19.76%), 190.16 (100%); anal. calc. for: C19H17N5O4 (379.37): C, 60.15; H, 4.52; N, 18.46; found: C, 60.19; H, 4.50; N, 18.48.
3.1.2.4 N′-(3-Hydroxyquinoxalin-2-yl)-1-(4-nitrophenyl)-5-oxopyrrolidine-3-carbohydrazide, 27. Light brown powder; yield: (73%), m.p. 267–268 °C; IR (KBr, cm−1) = 3480, 3359, 3219 (br. 2NH & OH), 3078 (CH aromatic), 1698 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O; 1H NMR (400 MHz,CDCl3): δ = 12.86 (s, 1H, OH), 10.79 (d, J = 6.8 Hz, 1H, –NH–NH–CO), 8.88 (d, J = 6.6 Hz, 1H, –NH–NH–CO), 8.09 (d, J = 7.8 Hz, 2H, A–H), 7.91 (dd, J = 7.7, 1.8 Hz, 1H, A–H), 7.73–7.42 (m, 5H, A–H), 4.20 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.94 (dd, J = 12.5, 3.6 Hz, 1H, pyrrolidine CH2), 3.28–3.16 (m, 1H, pyrrolidine CH), 2.71 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.49 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.73 (C
O; 1H NMR (400 MHz,CDCl3): δ = 12.86 (s, 1H, OH), 10.79 (d, J = 6.8 Hz, 1H, –NH–NH–CO), 8.88 (d, J = 6.6 Hz, 1H, –NH–NH–CO), 8.09 (d, J = 7.8 Hz, 2H, A–H), 7.91 (dd, J = 7.7, 1.8 Hz, 1H, A–H), 7.73–7.42 (m, 5H, A–H), 4.20 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.94 (dd, J = 12.5, 3.6 Hz, 1H, pyrrolidine CH2), 3.28–3.16 (m, 1H, pyrrolidine CH), 2.71 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.49 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.73 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 171.24 (C
O), 171.24 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 146.52, 143.90, 141.54, 139.45, 136.97, 136.51, 128.93, 126.45, 125.38, 124.48, 123.42, 119.11, 48.60 (pyrrolidine CH2), 38.03 (pyrrolidine CH), 36.42 (pyrrolidine CH2); MS(EI, 70 eV): m/z (%) = 408.60 [M˙+] (13.86%), 147.50 (100%); anal. calc. for: C19H16N6O5 (408.37): C, 55.88; H, 3.95; N, 20.58; found: C, 55.86; H, 3.96; N, 20.55.
O), 146.52, 143.90, 141.54, 139.45, 136.97, 136.51, 128.93, 126.45, 125.38, 124.48, 123.42, 119.11, 48.60 (pyrrolidine CH2), 38.03 (pyrrolidine CH), 36.42 (pyrrolidine CH2); MS(EI, 70 eV): m/z (%) = 408.60 [M˙+] (13.86%), 147.50 (100%); anal. calc. for: C19H16N6O5 (408.37): C, 55.88; H, 3.95; N, 20.58; found: C, 55.86; H, 3.96; N, 20.55.Conjugates 28–31 were afforded by reacting 2-chloro-3-hydrazinylquinoxaline (6) with fragments 18–21.
3.1.2.5 N′-(3-Chloroquinoxalin-2-yl)-1-(4-cyanophenyl)-5-oxopyrrolidine-3-carbohydrazide, 28. Brown powder; yield: (66%), m.p. 258–259 °C; IR (KBr, cm−1) = 3367–3260 (br. 2NH), 3070 (CH aromatic), 2213 (CN), 1696 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz,CDCl3): δ = 10.33 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 8.97 (d, J = 7.0 Hz, 1H, NH–NH–CO), 8.01–7.86 (m, 2H, A–H), 7.61–7.51 (m, 4H, A–H), 7.30 (d, J = 7.7 Hz, 2H, A–H), 4.19 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.94 (dd, J = 12.5, 3.6 Hz, 1H, pyrrolidine CH2), 3.26–3.17 (m, 1H, pyrrolidine CH), 2.71 (dd, J = 15.7, 5.0 Hz, 1H, pyrrolidine CH2), 2.49 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.24 (C
O); 1H NMR (400 MHz,CDCl3): δ = 10.33 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 8.97 (d, J = 7.0 Hz, 1H, NH–NH–CO), 8.01–7.86 (m, 2H, A–H), 7.61–7.51 (m, 4H, A–H), 7.30 (d, J = 7.7 Hz, 2H, A–H), 4.19 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.94 (dd, J = 12.5, 3.6 Hz, 1H, pyrrolidine CH2), 3.26–3.17 (m, 1H, pyrrolidine CH), 2.71 (dd, J = 15.7, 5.0 Hz, 1H, pyrrolidine CH2), 2.49 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.24 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 171.54 (C
O), 171.54 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 146.11, 141.32, 140.42, 137.80, 132.33, 131.31, 129.74, 127.06, 126.42, 125.81, 117.89, 117.18 (CN), 102.13, 48.78 (pyrrolidine CH2), 38.09 (pyrrolidine CH), 36.37 (pyrrolidine CH2); anal. calc. for: C20H15ClN6O2 (406.83): C, 59.05; H, 3.72; N, 20.66; found: C, 59.08; H, 3.70; N, 20.65.
O), 146.11, 141.32, 140.42, 137.80, 132.33, 131.31, 129.74, 127.06, 126.42, 125.81, 117.89, 117.18 (CN), 102.13, 48.78 (pyrrolidine CH2), 38.09 (pyrrolidine CH), 36.37 (pyrrolidine CH2); anal. calc. for: C20H15ClN6O2 (406.83): C, 59.05; H, 3.72; N, 20.66; found: C, 59.08; H, 3.70; N, 20.65.
3.1.2.6 N′-(3-Chloroquinoxalin-2-yl)-1-(2,5-dimethylphenyl)-5-oxopyrrolidine-3-carbohydrazide, 29. Light brown powder; yield: (75%), m.p. 274–276 °C; IR (KBr, cm−1) = 3349 (br. 2NH), 3044 (CH aromatic), 2904, 2840 (CH aliphatic), 1656 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O; 1H NMR (400 MHz, CDCl3): δ = 10.50 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 8.16–8.01 (m, 2H, A–H), 7.94 (d, J = 7.0 Hz, 1H, NH–NH–CO), 7.61 (dd, J = 6.0, 3.6 Hz, 2H, A–H), 7.14 (d, J = 8.7 Hz, 1H, A–H), 7.00 (d, J = 7.3 Hz, 2H, A–H), 4.18 (dd, J = 12.2, 3.6 Hz, 1H, pyrrolidine CH2), 3.93 (dd, J = 12.3, 3.6 Hz, 1H, pyrrolidine CH2), 3.18–3.09 (m, 1H, pyrrolidine CH), 2.53 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.34 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.26 (s, 3H, CH3), 2.09 (s, 3H, CH3); 13C NMR (100 MHz,CDCl3): δ = 172.15 (C
O; 1H NMR (400 MHz, CDCl3): δ = 10.50 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 8.16–8.01 (m, 2H, A–H), 7.94 (d, J = 7.0 Hz, 1H, NH–NH–CO), 7.61 (dd, J = 6.0, 3.6 Hz, 2H, A–H), 7.14 (d, J = 8.7 Hz, 1H, A–H), 7.00 (d, J = 7.3 Hz, 2H, A–H), 4.18 (dd, J = 12.2, 3.6 Hz, 1H, pyrrolidine CH2), 3.93 (dd, J = 12.3, 3.6 Hz, 1H, pyrrolidine CH2), 3.18–3.09 (m, 1H, pyrrolidine CH), 2.53 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.34 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.26 (s, 3H, CH3), 2.09 (s, 3H, CH3); 13C NMR (100 MHz,CDCl3): δ = 172.15 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 171.54 (C
O), 171.54 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 146.11, 140.42, 137.80, 137.35, 132.42, 131.31, 129.74, 129.63, 129.36, 127.06, 126.42, 125.81, 123.84, 121.82, 49.31 (pyrrolidine CH2), 38.12 (pyrrolidine CH), 36.32 (pyrrolidine CH2), 19.52 (CH3), 16.53 (CH3); MS(EI, 70 eV): m/z (%) = 411.38 [M + 2] (24.70%), 409.44 [M˙+] (33.19%), 259.30 (100%); anal. calc. for: C21H20ClN5O2 (409.87): C, 61.54; H, 4.92; N, 17.09; found: C, 61.58; H, 4.90; N, 17.14.
O), 146.11, 140.42, 137.80, 137.35, 132.42, 131.31, 129.74, 129.63, 129.36, 127.06, 126.42, 125.81, 123.84, 121.82, 49.31 (pyrrolidine CH2), 38.12 (pyrrolidine CH), 36.32 (pyrrolidine CH2), 19.52 (CH3), 16.53 (CH3); MS(EI, 70 eV): m/z (%) = 411.38 [M + 2] (24.70%), 409.44 [M˙+] (33.19%), 259.30 (100%); anal. calc. for: C21H20ClN5O2 (409.87): C, 61.54; H, 4.92; N, 17.09; found: C, 61.58; H, 4.90; N, 17.14.
3.1.2.7 N′-(3-Chloroquinoxalin-2-yl)-1-(3-hydroxyphenyl)-5-oxopyrrolidine-3-carbohydrazide, 30. Brown powder; yield: (63%), m.p. 278–279 °C; IR (KBr, cm−1) = 3418–3397 (br. 2NH & 2OH), 3076 (CH aromatic), 1651 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz,CDCl3): δ = 10.27 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 9.38 (d, J = 7.0 Hz, 1H, NH–NH–CO), 7.81–7.72 (m, 1H, A–H), 7.71–7.61 (m, 1H, A–H), 7.49–7.39 (m, 3H, A–H & OH), 7.06–6.97 (m, 2H, A–H), 6.55–6.49 (m, 1H, A–H), 6.43–6.34 (m, 1H, A–H), 4.24 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.98 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.17–3.09 (m, 1H, pyrrolidine CH), 2.70 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.45 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3):δ = 173.48 (C
O); 1H NMR (400 MHz,CDCl3): δ = 10.27 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 9.38 (d, J = 7.0 Hz, 1H, NH–NH–CO), 7.81–7.72 (m, 1H, A–H), 7.71–7.61 (m, 1H, A–H), 7.49–7.39 (m, 3H, A–H & OH), 7.06–6.97 (m, 2H, A–H), 6.55–6.49 (m, 1H, A–H), 6.43–6.34 (m, 1H, A–H), 4.24 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.98 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.17–3.09 (m, 1H, pyrrolidine CH), 2.70 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2), 2.45 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3):δ = 173.48 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 172.44 (C
O), 172.44 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 158.81, 149.69, 141.26, 140.85, 139.36, 131.53, 131.16, 130.66, 129.32, 126.83, 124.68, 117.35, 109.00, 106.54, 49.02 (pyrrolidine CH2), 40.79 (pyrrolidine CH), 38.00 (pyrrolidine CH2); MS(EI, 70 eV):m/z (%) = 397.20 [M + 2] (33.98%), 399.22 [M˙+] (25.48%), 379.93 (100%); anal. calc. for: C19H16ClN5O3 (397.82): C, 57.37; H, 4.05; N, 17.60; found: C, 57.33; H, 4.09; N, 17.57.
O), 158.81, 149.69, 141.26, 140.85, 139.36, 131.53, 131.16, 130.66, 129.32, 126.83, 124.68, 117.35, 109.00, 106.54, 49.02 (pyrrolidine CH2), 40.79 (pyrrolidine CH), 38.00 (pyrrolidine CH2); MS(EI, 70 eV):m/z (%) = 397.20 [M + 2] (33.98%), 399.22 [M˙+] (25.48%), 379.93 (100%); anal. calc. for: C19H16ClN5O3 (397.82): C, 57.37; H, 4.05; N, 17.60; found: C, 57.33; H, 4.09; N, 17.57.
3.1.2.8 N′-(3-Chloroquinoxalin-2-yl)-1-(4-nitrophenyl)-5-oxopyrrolidine-3-carbohydrazide, 31. Dark yellow powder; yield: (51%), m.p. 266–267 °C; IR (KBr, cm−1) = 3420–3342 (br. 2NH & OH), 3067 (CH aromatic), 1686 (br. 2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz,CDCl3): δ = 10.50 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 8.15–8.01 (m, 4H, A–H), 7.94 (d, J = 7.0 Hz, 1H, NH–NH–CO), 7.61 (dd, J = 6.0, 3.5 Hz, 2H, A–H), 7.40 (d, J = 7.8 Hz, 2H, A–H), 4.17 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.92 (dd, J = 12.5, 3.6 Hz, 1H, pyrrolidine CH2), 3.12 (p, J = 4.2 Hz, 1H, pyrrolidine CH), 2.52 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.33 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.65 (C
O); 1H NMR (400 MHz,CDCl3): δ = 10.50 (d, J = 7.0 Hz, 1H, –NH–NH–CO), 8.15–8.01 (m, 4H, A–H), 7.94 (d, J = 7.0 Hz, 1H, NH–NH–CO), 7.61 (dd, J = 6.0, 3.5 Hz, 2H, A–H), 7.40 (d, J = 7.8 Hz, 2H, A–H), 4.17 (dd, J = 12.5, 3.7 Hz, 1H, pyrrolidine CH2), 3.92 (dd, J = 12.5, 3.6 Hz, 1H, pyrrolidine CH2), 3.12 (p, J = 4.2 Hz, 1H, pyrrolidine CH), 2.52 (dd, J = 15.7, 4.9 Hz, 1H, pyrrolidine CH2), 2.33 (dd, J = 15.6, 4.9 Hz, 1H, pyrrolidine CH2); 13C NMR (100 MHz,CDCl3): δ = 172.65 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 171.12 (C
O), 171.12 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 146.45, 143.82, 139.83, 139.37, 138.67, 130.84, 129.01, 126.36, 126.02, 125.18, 124.41, 119.04, 48.53 (pyrrolidine CH2), 37.94 (pyrrolidine CH), 36.35 (pyrrolidine CH2); MS (EI, 70 eV): m/z (%) = 428.41 [M + 2] (10.83%), 426.69 [M˙+] (19.33%), 263.76 (100%); anal. calc. for: C19H15ClN6O4 (426.82): C, 53.47; H, 3.54; N, 19.69; Found: C, 53.44; H, 3.50; N, 19.67.
O), 146.45, 143.82, 139.83, 139.37, 138.67, 130.84, 129.01, 126.36, 126.02, 125.18, 124.41, 119.04, 48.53 (pyrrolidine CH2), 37.94 (pyrrolidine CH), 36.35 (pyrrolidine CH2); MS (EI, 70 eV): m/z (%) = 428.41 [M + 2] (10.83%), 426.69 [M˙+] (19.33%), 263.76 (100%); anal. calc. for: C19H15ClN6O4 (426.82): C, 53.47; H, 3.54; N, 19.69; Found: C, 53.44; H, 3.50; N, 19.67.
3.2. Biology
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 v/v. The concentrations varied between 1.5 and 200 µg mL−1, with fluconazole, levofloxacin, and amoxicillin serving as drug controls. Briefly, in a microtiter plate, 100 µL of the chemicals to be tested were combined with 100 µL of TSB and 5 µL of 1 × 108 CFU mL−1 of the strain under investigation. For bacteria or fungi, incubation of the plates lasted at 35 ± 2 °C for 24 or 48 hours, respectively. MIC was the least amount of concentration of substance which showed no signs of growth.87–90
10 v/v. The concentrations varied between 1.5 and 200 µg mL−1, with fluconazole, levofloxacin, and amoxicillin serving as drug controls. Briefly, in a microtiter plate, 100 µL of the chemicals to be tested were combined with 100 µL of TSB and 5 µL of 1 × 108 CFU mL−1 of the strain under investigation. For bacteria or fungi, incubation of the plates lasted at 35 ± 2 °C for 24 or 48 hours, respectively. MIC was the least amount of concentration of substance which showed no signs of growth.87–90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 2
1 and 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, the investigated conjugate was deemed bactericidal or fungicidal; if the ratio is more than 2
1, the investigated conjugate was deemed bactericidal or fungicidal; if the ratio is more than 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, it was deemed bacteriostatic or fungistatic.91–95
1, it was deemed bacteriostatic or fungistatic.91–95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 CFU mL−1 vs. time (h) was used to depict the bactericidal kinetics of conjugates.104–109
10 CFU mL−1 vs. time (h) was used to depict the bactericidal kinetics of conjugates.104–1093.3. In silico ADMET
Pharmacokinetics prediction was performed as reported.116–1233.4. Molecular docking
The 3D layout for DNA gyrase (PDB.: 8BN6) had been extracted through data bank of the protein (https://www.rcsb.org/) and prepared using UCSF Chimera.124–127 Briefly, the mol2 file of protein was set by deleting water molecules, adding and charges and energy minimization. The generated compounds and respective co-crystallized ligands were sketched in Avogadro software, and their geometries were optimized.128,129 The prepared protein was imported in AutoDock. Hydrogens were added and Kollman charges and AD4 atom parameters were assigned. The grid calculations were set up; −5.5 × 23.5 × 1.1 Å grid with dimensions of 35, 35, 35 respectively, at a resolution of 0.375 Å, was created around the compound. Docking commenced using genetic algorithm search parameters. During docking, partial Gasteiger charges were applied. Docked conformations were generated using the Lamarckian evolutionary method and arranged according to their docking energy in descending order.130–134 Results visualizations were performed using Biovia Discovery Studio. The same steps were repeated for the co-crystallized ligand in order to perform validation. The RMSD value of the redocked pose was calculated using DockRMSD online tool.135,1364. Conclusion
This study successfully synthesized and studied several novel hybrid chemicals, such as quinoxaline, pyrrolidine, and a hydrazinyl bridge, indicating their potential as strong antibacterial and antifungal medicines. Several compounds had strong efficacy against different pathogenic strains, according to the in vitro tests, with eight derivatives demonstrating notable bactericidal effects. Eight novel conjugates were tested for antibacterial activity at a dosage of 5 mM. All synthesized derivatives caused E. coli growth ZOI ≥15 mm, with hybrids 26, 28, 30, and 31 showing greater reduction than Amoxicillin and Levofloxacin. Hybrids 24, 27, and 29 were confirmed as bactericidal agents against E. coli. Hybrids 24, 25, and 31 displayed MIC of 12.5 µM, whereas they indicate bactericidal hybrids at higher concentrations against P. aeruginosa. Treatment with involving all hybrids resulted in 89–92% decrease in biofilm formation at 90% MIC, outperforming Levofloxacin. The killing kinetics of eight hybrids against P. aeruginosa were time-dependent, with higher concentrations leading to a rapid decrease in CFU number. Primary bacterial elimination occurred after 3 hours, while 4-fold and 8-fold MICs led to almost complete elimination. Hybrids 25, 28, and 31 were the most efficient DNA gyrase inhibitors; their IC50 values were significantly lower than those of ciprofloxacin (77.3, 87.6, and 65.5 µM). The study reveals that molecular docking studies reveal effective compounds' binding interactions, confirming their biological activities. Compounds having drug-like properties are identified by examining their physicochemical and pharmacokinetic properties, indicating the possibility of further development. With future research concentrating on optimizing lead compounds for clinical applications, the findings support the continuing attempts to develop novel treatment medicines for infectious disorders and resistant microbial strains.Conflicts of interest
The author(s) report no conflicts of interest in this work.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00675b.References
- T. M. Rawson, R. C. Wilson and A. Holmes, Understanding the role of bacterial and fungal infection in COVID-19, Clin. Microbiol. Infection, 2021, 27, 9–11 CrossRef CAS PubMed.
- J. R. Köhler, B. Hube, R. Puccia, A. Casadevall and J. R. Perfect, Fungi that Infect Humans, Microbiol. Spectr., 2017, 5(3) DOI:10.1128/microbiolspec.FUNK-0014-2016.
- J. Verhoef, K. van Kessel and H. Snippe, Immune Response in Human Pathology: Infections Caused by Bacteria, Viruses, Fungi, and Parasites, in Nijkamp and Parnham's Principles of Immunopharmacology, ed. M. J. Parnham, F. P. Nijkamp and A. G. Rossi, Springer International Publishing, Cham, 2019, pp. 165–178 Search PubMed.
- B. M. Peters, M. A. Jabra-Rizk, G. A. O'May, J. W. Costerton and M. E. Shirtliff, Polymicrobial Interactions: Impact on Pathogenesis and Human Disease, Clin. Microbiol. Rev., 2012, 25, 193–213 CrossRef PubMed.
- G. Muteeb, M. T. Rehman, M. Shahwan and M. Aatif, Origin of Antibiotics and Antibiotic Resistance, and Their Impacts on Drug Development: A Narrative Review, Pharmaceuticals, 2023, 16, 1615 CrossRef CAS PubMed.
- M. Terreni, M. Taccani and M. Pregnolato, New Antibiotics for Multidrug-Resistant Bacterial Strains: Latest Research Developments and Future Perspectives, Molecules, 2021, 26, 2671 CrossRef CAS PubMed.
- R. L. Monaghan and J. F. Barrett, Antibacterial drug discovery—Then, now and the genomics future, Biochem. Pharmacol., 2006, 71, 901–909 CrossRef CAS PubMed.
- J. Murugaiyan, P. A. Kumar, G. S. Rao, K. Iskandar, S. Hawser and J. P. Hays, et al., Progress in Alternative Strategies to Combat Antimicrobial Resistance: Focus on Antibiotics, Antibiotics, 2022, 11, 200 CrossRef CAS PubMed.
- M. Frieri, K. Kumar and A. Boutin, Antibiotic resistance, J. Infect. Public Health, 2017, 10, 369–378 CrossRef PubMed.
- A. Tarín-Pelló, B. Suay-García and M.-T. Pérez-Gracia, Antibiotic resistant bacteria: current situation and treatment options to accelerate the development of a new antimicrobial arsenal, Expert Rev. Anti-infect. Ther., 2022, 20, 1095–1108 CrossRef PubMed.
- D. Chinemerem Nwobodo, M. C. Ugwu, C. Oliseloke Anie, M. T. S. Al-Ouqaili, J. Chinedu Ikem and U. Victor Chigozie, et al., Antibiotic resistance: The challenges and some emerging strategies for tackling a global menace, J. Clin. Lab. Anal., 2022, 36, e24655 CrossRef PubMed.
- M. A. Salam, M. Y. Al-Amin, M. T. Salam, J. S. Pawar, N. Akhter and A. A. Rabaan, et al., Antimicrobial Resistance: A Growing Serious Threat for Global Public Health, Healthcare, 2023, 11, 1946 CrossRef PubMed.
- T. Ramsis, H. M. Refat, M. Selim, H. Elseedy and E. A. Fayed, The role of current synthetic and possible plant and marine phytochemical compounds in the treatment of acne, RSC Adv., 2024, 14, 24287–24321 RSC.
- E. A. Fayed, S. A. El-Sebaey, M. A. Ebrahim, K. Abu-Elfotuh, R. El-Sayed Mansour and E. K. Mohamed, et al., Discovery of novel bicyclic and tricyclic cyclohepta[b]thiophene derivatives as multipotent AChE and BChE inhibitors, in-Vivo and in-Vitro assays, ADMET and molecular docking simulation, Eur. J. Med. Chem., 2025, 284, 117201 CrossRef CAS PubMed.
- M. S. Abusaif, A. Ragab, E. A. Fayed, Y. A. Ammar, A. M. H. Gowifel and S. O. Hassanin, et al., Exploring a novel thiazole derivatives hybrid with fluorinated-indenoquinoxaline as dual inhibitors targeting VEGFR2/AKT and apoptosis inducers against hepatocellular carcinoma with docking simulation, Bioorg. Chem., 2025, 154, 108023 CrossRef CAS PubMed.
- R. Hassan, R. Mohi-ud-din, M. O. Dar, A. J. Shah, P. A. Mir and M. Shaikh, et al., Bioactive Heterocyclic Compounds as Potential Therapeutics in the Treatment of Gliomas: A Review, Anti-Cancer Agents Med. Chem., 2022, 22, 551–565 CrossRef CAS PubMed.
- E. A. Fayed, A. Thabet, S. M. A. El-Gilil, H. M. A. Elsanhory and Y. A. Ammar, Fluorinated thiazole–thiosemicarbazones hybrids as potential PPAR-γ agonist and α-amylase, α-glucosidase antagonists: Design, synthesis, in silico ADMET and docking studies and hypoglycemic evaluation, J. Mol. Struct., 2024, 1301, 137374 CrossRef CAS.
- S. E. Desouky, M. Abu-Elghait, E. A. Fayed, S. Selim, B. Yousuf and Y. Igarashi, et al., Secondary Metabolites of Actinomycetales as Potent Quorum Sensing Inhibitors Targeting Gram-Positive Pathogens: In Vitro and In Silico Study, Metabolites, 2022, 12, 246 CrossRef CAS PubMed.
- B. A. AbdelFattah, M. M. A. Khalifa, H. El-Sehrawi, E. Fayed, A. Bayoumi and M. Said, Synthesis and Anxiolytic Activity of Some Novel 5-oxo-1, 4-oxazepine Derivatives, Lett. Drug Des. Discovery, 2011, 8, 330–338 CrossRef CAS.
- H. Li, Y.-H. He, Y.-M. Hu, Q.-R. Chu, Y.-J. Chen and Z.-R. Wu, et al., Design, Synthesis, and Structure–Activity Relationship Studies of Magnolol Derivatives as Antifungal Agents, J. Agric. Food Chem., 2021, 69, 11781–11793 CrossRef CAS PubMed.
- N. J. C. Oliveira, I. N. S. Teixeira, P. O. Fernandes, G. C. Veríssimo, A. D. Valério and C. P. D. S. Moreira, et al., Computer-aided molecular design, synthesis and evaluation of antifungal activity of heterocyclic compounds, J. Mol. Struct., 2022, 1267, 133573 CrossRef CAS.
- B. A. Abdel Fattah, M. M. A. Khalifa, H. El-Sehrawi and E. Fayed, Synthesis, Anxiolytic and Antimicrobial Activity of Some Novel 5-oxo-1, 4-Oxazepine Derivatives, Egypt. J. Biomed. Sci., 2010, 33, 269 Search PubMed.
- Y. U. Cebeci, Ö. Ö. Batur and H. Boulebd, Design, synthesis, theoretical studies, and biological activity evaluation of new nitrogen-containing poly heterocyclic compounds as promising antimicrobial agents, J. Mol. Struct., 2024, 1299, 137115 CrossRef CAS.
- U. Fathy, H. A. Abd El Salam, E. A. Fayed, A. M. Elgamal and A. Gouda, Erratum to “Facile synthesis and in vitro anticancer evaluation of a new series of tetrahydroquinoline” [Heliyon 7(10) (October 2021) e08117] (Heliyon (2021) 7(10), (S2405844021022209), Heliyon, 2024, 10(1), e23189, DOI:10.1016/j.heliyon.2021.e08117.
- T. Z. Shawer and E. A. Fayed, Synthesis, docking and anticancer activity of some new thienopyrimidine and thienooxazine derivatives, Nat. Sci., 2014, 12(12), 171–181 Search PubMed.
- M. Montana, V. Montero, O. Khoumeri and P. Vanelle, Quinoxaline derivatives as antiviral agents: a systematic review, Molecules, 2020, 25, 2784 CrossRef CAS PubMed.
- T. M. Ramsis, S. E. Abdel Karim, N. Vassilaki, E. Frakolaki, A. A. M. Kamal and G. Zoidis, et al., Expanding the chemical space of anti-HCV NS5A inhibitors by stereochemical exchange and peptidomimetic approaches, Arch. Pharmazie, 2018, 351, 1800017 CrossRef PubMed.
- N. A. Gohar, E. A. Fayed, Y. A. Ammar, O. A. Abu Ali, A. Ragab and A. M. Mahfoz, et al., Fluorinated indeno-quinoxaline bearing thiazole moieties as hypoglycaemic agents targeting α-amylase, and α-glucosidase: synthesis, molecular docking, and ADMET studies, J. Enzyme Inhib. Med. Chem., 2024, 39, 2367128 CrossRef PubMed.
- A. Thabet, M. Samir Abusaif, Y. A. Ammar, S. M. Abd-Elgilil and H. Elsanhory, Bis-thiazole hybrids as promising candidates in controlling diabetes mellitus in vivo, in vitro evaluation, in silico ADMET, and docking studies, Al-Azhar Int. J. Pharm. Med. Sci., 2025, 5(2), 87–105 Search PubMed.
- S. Tariq, K. Somakala and M. Amir, Quinoxaline: An insight into the recent pharmacological advances, Eur. J. Med. Chem., 2018, 143, 542–557 CrossRef CAS PubMed.
- E. A. Fayed, N. A. Gohar, A. M. Farrag and Y. A. Ammar, Upregulation of BAX and caspase-3, as well as downregulation of Bcl-2 during treatment with indeno[1,2-b]quinoxalin derivatives, mediated apoptosis in human cancer cells, Arch. Pharmazie, 2022, 355, 2100454 CrossRef CAS PubMed.
- A. Mishra, V. Jha and H. Rajak, Molecular structural investigations of quinoxaline derivatives through 3D-QSAR, molecular docking, ADME prediction and pharmacophore modeling studies for the search of novel antimalarial agent, J. Indian Chem. Soc., 2022, 99, 100343 CrossRef CAS.
- Y. Singh, N. Kumar, S. Kulkarni, S. Singh and S. Thareja, Pharmacophore derived 3D-QSAR, molecular docking, and simulation studies of quinoxaline derivatives as ALR2 inhibitors, J. Biomol. Struct. Dyn., 2024, 42, 10452–10488 CrossRef CAS PubMed.
- S. K. Suthar, N. S. Chundawat, G. P. Singh, J. M. Padrón and Y. K. Jhala, Quinoxaline: A comprehension of current pharmacological advancement in medicinal chemistry, Eur. J. Med. Chem. Rep., 2022, 5, 100040 CAS.
- S. ElKalyoubi and E. Fayed, Synthesis and Evaluation of Antitumour Activities of Novel Fused Tri- and Tetracyclic Uracil Derivatives, J. Chem. Res., 2016, 40, 771–777 CrossRef.
- U. Fathy, H. A. Abd El Salam, E. A. Fayed, A. M. Elgamal and A. Gouda, Facile synthesis and anticancer evaluation of a new series of tetrahydroquinoline, Heliyon, 2021, 7, e08117 CrossRef CAS PubMed.
- E. Fayed and H. Ahmed, Synthesis, characterization and pharmacological evaluation of some new 1, 4-diazepine derivatives as anticancer agents, Pharma Chem., 2016, 8, 77–90 CAS.
- A. Sharma, A. Deep, M. G. Marwaha and R. K. Marwaha, Quinoxaline: A chemical moiety with spectrum of interesting biological activities, Mini Rev. Med. Chem., 2022, 22, 927–948 CrossRef CAS PubMed.
- K. Otoguro, A. Ishiyama, M. Iwatsuki, M. Namatame, A. Nishihara-Tukashima and T. Nakashima, et al., In vitro and in vivo anti-Trypanosoma brucei activities of phenazinomycin and related compounds, J. Antibiotics, 2010, 63, 579–581 CrossRef CAS PubMed.
- F. Xu and J. A. McCauley, Discovery and Chemical Development of Grazoprevir: An HCV NS3/4a Protease Inhibitor for the Treatment of the Hepatitis C Virus Infection, in Complete Accounts of Integrated Drug Discovery and Development: Recent Examples from the Pharmaceutical Industry Volume 3, American Chemical Society, 2020, vol. 1369, pp. 285–312 Search PubMed.
- S. Wang, Y. Wang, J. Wang, T. Sato, K. Izawa and V. A. Soloshonok, et al., The Second-generation of Highly Potent Hepatitis C Virus (HCV) NS3/4A Protease Inhibitors: Evolutionary Design Based on Tailor-made Amino Acids, Synthesis and Major Features of Bio-activity, Curr. Pharm. Des., 2017, 23, 4493–4554 CAS.
- J. G. Taylor, S. Zipfel, K. Ramey, R. Vivian, A. Schrier and K. K. Karki, et al., Discovery of the pan-genotypic hepatitis C virus NS3/4A protease inhibitor voxilaprevir (GS-9857): A component of Vosevi®, Bioorg. Med. Chem. Lett., 2019, 29, 2428–2436 CrossRef CAS PubMed.
- D. Nageswara Rao, J. Zephyr, M. Henes, E. T. Chan, A. N. Matthew and A. K. Hedger, et al., Discovery of Quinoxaline-Based P1–P3 Macrocyclic NS3/4A Protease Inhibitors with Potent Activity against Drug-Resistant Hepatitis C Virus Variants, J. Med. Chem., 2021, 64, 11972–11989 CrossRef CAS PubMed.
- L. Yurttaş, Y. Özkay, Z. A. Kaplancıklı, Y. Tunalı and H. Karaca, Synthesis and antimicrobial activity of some new hydrazone-bridged thiazole-pyrrole derivatives, J. Enzyme Inhib. Med. Chem., 2013, 28, 830–835 CrossRef PubMed.
- R. H. Abd El-Hameed, A. I. Sayed, S. Mahmoud Ali, M. A. Mosa, Z. M. Khoder and S. S. Fatahala, Synthesis of novel pyrroles and fused pyrroles as antifungal and antibacterial agents, J. Enzyme Inhib. Med. Chem., 2021, 36, 2183–2198 CrossRef CAS PubMed.
- P. Rawat, R. Singh, A. Ranjan, A. Gautam, S. Trivedi and M. Kumar, Study of antimicrobial and antioxidant activities of pyrrole-chalcones, J. Mol. Struct., 2021, 1228, 129483 CrossRef CAS.
- E. A. Fayed, N. A. Gohar, A. H. Bayoumi and Y. A. Ammar, Novel fluorinated pyrazole-based heterocycles scaffold: cytotoxicity, in silico studies and molecular modelling targeting double mutant EGFR L858R/T790M as antiproliferative and apoptotic agents, Med. Chem. Res., 2023, 32, 369–388 CrossRef CAS.
- C. Y. Hong, Discovery of gemifloxacin (Factive, LB20304a): a quinolone of new a generation, Il Farmaco, 2001, 56, 41–44 CrossRef CAS PubMed.
- J.-P. Caeiro and P. B. Iannini, Moxifloxacin (Avelox®): a novel fluoroquinolone with a broad spectrum of activity, Expert Rev. Anti-infect. Ther., 2003, 1, 363–370 CrossRef CAS PubMed.
- U. Fathy, M. N. M. Yousif, E. M. Mohi El-Deen and E. Fayed, Design, Synthesis, and biological evaluation of a novel series of thiazole derivatives based on pyrazoline as anticancer agents, Egypt. J. Chem., 2022, 65, 1241–1252 Search PubMed.
- S. Kiyotoki, J. Nishikawa and I. Sakaida, Efficacy of vonoprazan for Helicobacter pylori eradication, Intern. Med., 2020, 59, 153–161 CrossRef CAS PubMed.
- A. Rusu, I.-M. Moga, L. Uncu and G. Hancu, The Role of Five-Membered Heterocycles in the Molecular Structure of Antibacterial Drugs Used in Therapy, Pharmaceutics, 2023, 15, 2554 CrossRef CAS PubMed.
- T. M. Ramsis, S. Hussein, M. S. Abusaif, A. Ragab, Y. A. Ammar and O. Ali, et al., Bridging the gap with amide linkers: rational design, synthesis, and multi-target evaluation of sulfonamide/acetamide-NSAID hybrids as dual COX-2/5-LOX inhibitors, Future Med. Chem., 2026, 18, 131–148 CrossRef CAS PubMed.
- M. N. Matada, K. Jathi, M. M. Rangappa, K. Geoffry, S. R. Kumar and R. B. Nagarajappa, et al., A new sulphur containing heterocycles having azo linkage: Synthesis, structural characterization and biological evaluation, J. King Saud Univ. Sci., 2020, 32, 3313–3320 CrossRef.
- T. Tahir, M. Ashfaq, M. Saleem, M. Rafiq, M. I. Shahzad and K. Kotwica-Mojzych, et al., Pyridine scaffolds, phenols and derivatives of azo moiety: current therapeutic perspectives, Molecules, 2021, 26, 4872 CrossRef CAS PubMed.
- K. Mezgebe and E. Mulugeta, Synthesis and pharmacological activities of azo dye derivatives incorporating heterocyclic scaffolds: a review, RSC Adv., 2022, 12, 25932–25946 RSC.
- S. A. El-Kalyoubi, E. A. Fayed and A. S. Abdel-Razek, One pot synthesis, antimicrobial and antioxidant activities of fused uracils: pyrimidodiazepines, lumazines, triazolouracil and xanthines, Chem. Cent. J., 2017, 11, 66 CrossRef PubMed.
- T. Khan, K. Sankhe, V. Suvarna, A. Sherje, K. Patel and B. Dravyakar, DNA gyrase inhibitors: Progress and synthesis of potent compounds as antibacterial agents, Biomed. Pharmacother., 2018, 103, 923–938 CrossRef CAS PubMed.
- A. Duprey and E. A. Groisman, DNA supercoiling differences in bacteria result from disparate DNA gyrase activation by polyamines, PLoS Genet., 2020, 16, e1009085 CrossRef CAS PubMed.
- G. Sanyal and P. Doig, Bacterial DNA replication enzymes as targets for antibacterial drug discovery, Expet Opin. Drug Discov., 2012, 7, 327–339 CrossRef CAS PubMed.
- G. Seneviratne, J. S. Zavahir, W. Bandara and M. Weerasekara, Fungal-bacterial biofilms: their development for novel biotechnological applications, World J. Microbiol. Biotechnol., 2008, 24, 739–743 CrossRef CAS.
- D. C. Sheppard and P. L. Howell, Biofilm Exopolysaccharides of Pathogenic Fungi: Lessons from Bacteria, J. Biol. Chem., 2016, 291, 12529–12537 CrossRef CAS PubMed.
- F. M. Carvalho, A. Azevedo, M. M. Ferreira, F. J. M. Mergulhão and L. C. Gomes, Advances on Bacterial and Fungal Biofilms for the Production of Added-Value Compounds, Biology, 2022, 11, 1126 CrossRef CAS PubMed.
- H. Van Acker, P. Van Dijck and T. Coenye, Molecular mechanisms of antimicrobial tolerance and resistance in bacterial and fungal biofilms, Trends Microbiol., 2014, 22, 326–333 CrossRef CAS PubMed.
- E. A. Fayed, M. A. Ebrahim, U. Fathy, H. S. El Saeed and W. S. Khalaf, Evaluation of quinoxaline derivatives as potential ergosterol biosynthesis inhibitors: design, synthesis, ADMET, molecular docking studies, and antifungal activities, J. Mol. Struct., 2022, 1267, 133578 CrossRef CAS.
- R. M. Donlan and J. W. Costerton, Biofilms: survival mechanisms of clinically relevant microorganisms, Clin. Microbiol. Rev., 2002, 15, 167–193 CrossRef CAS PubMed.
- S. T. Asma, K. Imre, A. Morar, V. Herman, U. Acaroz and H. Mukhtar, et al., An overview of biofilm formation–combating strategies and mechanisms of action of antibiofilm agents, Life, 2022, 12, 1110 CrossRef CAS PubMed.
- A. R. Bhat, R. S. Dongre, A. H. Shalla, G. A. Naikoo and I. U. Hassan, Computational analysis for antimicrobial active pyrano [2, 3-d] pyrimidine derivatives on the basis of theoretical and experimental ground, J. Assoc. Arab Univ. Basic Appl. Sci., 2016, 20, 19–25 Search PubMed.
- M. A. Ebrahim, T. M. Ramsis, N. A. Gohar, S. A. metwally, A. Rushdi and E. A. Fayed, Novel Pyrrolidine-bearing quinoxaline inhibitors of DNA Gyrase, RNA polymerase and spike glycoprotein, Bioorg. Chem., 2025, 156, 108218 CrossRef CAS PubMed.
- K. H. Oudah, M. A. A. Najm, T. M. Ramsis, M. A. Ebrahim, N. A. Gohar and K. Abu-Elfotuh, et al., Unlocking Therapeutic Potential of Novel Thieno-Oxazepine Hybrids as Multi-Target Inhibitors of AChE/BChE and Evaluation Against Alzheimer's Disease: In Vivo, In Vitro, Histopathological, and Docking Studies, Pharmaceuticals, 2025, 18, 1214 CrossRef CAS PubMed.
- A. S. Oyedele, D. N. Bogan and C. O. Okoro, Synthesis, biological evaluation and virtual screening of some acridone derivatives as potential anticancer agents, Bioorg. Med. Chem., 2020, 28, 115426 CrossRef CAS PubMed.
- M. Y. Refai, A. M. Elazzazy, S. E. Desouky, M. Abu-Elghait, E. A. Fayed and S. M. Alajel, et al., Interception of Epoxide ring to quorum sensing system in Enterococcus faecalis and Staphylococcus aureus, AMB Express, 2023, 13, 126 CrossRef CAS PubMed.
- D. E. V. Pires, T. L. Blundell and D. B. Ascher, pkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Properties Using Graph-Based Signatures, J. Med. Chem., 2015, 58, 4066–4072 CrossRef CAS PubMed.
- P. Banerjee, A. O. Eckert, A. K. Schrey and R. Preissner, ProTox-II: a webserver for the prediction of toxicity of chemicals, Nucleic Acids Res., 2018, 46, W257–W263 CrossRef CAS PubMed.
- S. A. Serkov, N. N. Kostikova, N. V. Sigay, A. P. Tyurin, N. G. Kolotyrkina and G. A. Gazieva, The synthesis and study of antimicrobial activity of 5-oxo-1-(thiazol-2-yl)pyrrolidine-3-carboxylic acids, Chem. Heterocycl. Compd., 2024, 60, 365–370 CrossRef CAS.
- P. L. Paytash, M. J. Thompson and M. E. Fykes, Itaconic Acid Derivatives of Sulfanilamide, J. Am. Chem. Soc., 1952, 74, 4549–4552 CrossRef CAS.
- K. Inoguchi, T. Morimoto and K. Achiwa, Asymmetric reactions catalyzed by chiral metal complexes: XXXII. Efficient asymmetric hydrogenation of itaconic acid derivatives catalyzed by rhodium complexes of improved (2S,4S)-N-(t-butoxycarbonyl)-4-(diphenylphosphino)-2-[(diphenylphosphino)methyl]pyrrolidine (BPPM) analogues, J. Organomet. Chem., 1989, 370, C9–C12 CrossRef CAS.
- G. J. Noordzij and C. H. Wilsens, Cascade aza-Michael addition-cyclizations; toward renewable and multifunctional carboxylic acids for melt-polycondensation, Front. Chem., 2019, 7, 729 CrossRef CAS PubMed.
- I. Shabeeb, L. Al-Essa, M. Shtaiwi, E. Al-Shalabi, E. Younes and R. Okasha, et al., New Hydrazide-hydrazone Derivatives of Quinoline 3-Carboxylic Acid Hydrazide: Synthesis, Theoretical Modeling and Antibacterial Evaluation, Lett. Org. Chem., 2019, 16, 430–436 CrossRef CAS.
- Ł. Popiołek, B. Rysz, A. Biernasiuk and M. Wujec, Synthesis of promising antimicrobial agents: hydrazide-hydrazones of 5-nitrofuran-2-carboxylic acid, Chem. Biol. Drug Des., 2020, 95, 260–269 CrossRef PubMed.
- K. Paruch, Ł. Popiołek, A. Biernasiuk, A. Berecka-Rycerz, A. Malm and A. Gumieniczek, et al., Novel Derivatives of 4-Methyl-1,2,3-Thiadiazole-5-Carboxylic Acid Hydrazide: Synthesis, Lipophilicity, and In Vitro Antimicrobial Activity Screening, Appl. Sci., 2021, 11, 1180 CrossRef CAS.
- M. Y. Kornev, V. S. Moshkin, O. S. Eltsov and V. Y. Sosnovskikh, Reactions of chromone-3-carboxylic acid and chromone-3-carboxamides with cyanoacetic acid hydrazide, Mendeleev Commun., 2016, 26, 72–74 CrossRef CAS.
- E. M. Sarshira, N. M. Hamada, Y. M. Moghazi and M. M. Abdelrahman, Synthesis and Biological Evaluation of Some Heterocyclic Compounds from Salicylic Acid Hydrazide, J. Heterocycl. Chem., 2016, 53, 1970–1982 CrossRef CAS.
- Clinical and L. S. Institute, Performance Standards for Antimicrobial Susceptibility Testing, Clinical and Laboratory Standards Institute, Wayne, PA, 2020 Search PubMed.
- S. A. El-Kalyoubi, E. A. Fayed and A. S. Abdel-Razek, Erratum: One pot synthesis, antimicrobial and antioxidant activities of fused uracils: Pyrimidodiazepines, lumazines, triazolouracil and xanthines," [Chem Cent J 11, (2017) (66)], Chem. Cent. J., 2017, 11(1), 69, DOI:10.1186/s13065-017-0294-0.
- Y. Ammar, E. Fayed, A. Bayoumi and M. Saleh, Synthesis and biological evaluation of new amides pro-drugs containing naproxen moiety as anti-inflammatory and antimicrobial agents, Pharma Chem., 2016, 8, 495–508 Search PubMed.
- A. J. Cartagena, K. L. Taylor, J. T. Smith, A. L. Manson, V. M. Pierce and A. M. Earl, The carbapenem inoculum effect provides insight into the molecular mechanisms underlying carbapenem resistance in Enterobacterales, bioRxiv, 2023, preprint, DOI:10.1101/2023.05.23.541813.
- N. Sasoni, B. Caracciolo, M. S. Cabeza, S. Gamarra, S. Carnovale and G. Garcia-Effron, Antifungal susceptibility testing following the CLSI M27 document, along with the measurement of MFC/MIC ratio, could be the optimal approach to detect amphotericin B resistance in Clavispora (Candida) lusitaniae. Susceptibility patterns of contemporary isolates of this species, Antimicrob. Agents Chemother., 2024, 68, e00968–23 CrossRef PubMed.
- C. S. Makade, P. R. Shenoi, B. A. Bhongade, S. A. Shingane, P. C. Ambulkar and A. M. Shewale, Estimation of MBC: MIC Ratio of Herbal Extracts against Common Endodontic Pathogens, J. Pharm. BioAllied Sci., 2024, 16, S1414–S1416 CrossRef CAS PubMed.
- R. D. de Castro, T. M. P. A. de Souza, L. M. D. Bezerra, G. L. S. Ferreira, E. M. M. de Brito Costa and A. L. Cavalcanti, Antifungal activity and mode of action of thymol and its synergism with nystatin against Candida species involved with infections in the oral cavity: an in vitro study, BMC Compl. Alternative Med., 2015, 15, 417 CrossRef PubMed.
- G. Morace, M. Drago, M. M. Scaltrito, S. Conti, F. Fanti and L. Polonelli, In Vitro Activity (MIC and MFC) of Voriconazole, Amphotericin B, and Itraconazole Against 192 Filamentous Fungi: The GISIA-2 Study, J. Chemother., 2007, 19, 508–513 CrossRef CAS PubMed.
- G. Anand, M. Ravinanthan, R. Basaviah and A. V. Shetty, In vitro antimicrobial and cytotoxic effects of Anacardium occidentale and Mangifera indica in oral care, J. Pharm. BioAllied Sci., 2015, 7, 69–74 CrossRef PubMed.
- P. Arulmozhi, S. Vijayakumar and T. Kumar, Phytochemical analysis and antimicrobial activity of some medicinal plants against selected pathogenic microorganisms, Microb. Pathog., 2018, 123, 219–226 CrossRef CAS PubMed.
- S. Rehman, S. M. Asiri, F. A. Khan, B. R. Jermy, H. Khan and S. Akhtar, et al., Biocompatible Tin Oxide Nanoparticles: Synthesis, Antibacterial, Anticandidal and Cytotoxic Activities, ChemistrySelect, 2019, 4, 4013–4017 CrossRef CAS.
- M. Jalal, M. A. Ansari, M. A. Alzohairy, S. G. Ali, H. M. Khan and A. Almatroudi, et al., Biosynthesis of Silver Nanoparticles from Oropharyngeal Candida glabrata Isolates and Their Antimicrobial Activity against Clinical Strains of Bacteria and Fungi, Nanomaterials, 2018, 8, 586 CrossRef PubMed.
- H. Liu, Y. Zhao, D. Zhao, T. Gong, Y. Wu and H. Han, et al., Antibacterial and anti-biofilm activities of thiazolidione derivatives against clinical staphylococcus strains, Emerg. Microb. Infect., 2015, 4, 1–6 Search PubMed.
- G. Spengler, A. Kincses, T. Mosolygó, M. A. Marć, M. Nové and M. Gajdács, et al., Antiviral, Antimicrobial and Antibiofilm Activity of Selenoesters and Selenoanhydrides, Molecules, 2019, 24, 4264 CrossRef CAS PubMed.
- Y. Liu, L. Wu, J. Han, P. Dong, X. Luo and Y. Zhang, et al., Inhibition of Biofilm Formation and Related Gene Expression of Listeria monocytogenes in Response to Four Natural Antimicrobial Compounds and Sodium Hypochlorite, Front. Microbiol., 2020, 11, 617473 CrossRef PubMed.
- N. Tripathi and M. K. Goshisht, Recent Advances and Mechanistic Insights into Antibacterial Activity, Antibiofilm Activity, and Cytotoxicity of Silver Nanoparticles, ACS Appl. Bio Mater., 2022, 5, 1391–1463 CrossRef CAS.
- E. Sánchez, C. Rivas Morales, S. Castillo, C. Leos-Rivas, L. García-Becerra and D. M. Ortiz Martínez, Antibacterial and Antibiofilm Activity of Methanolic Plant Extracts against Nosocomial Microorganisms, Evid. Based Complement. Alternat. Med., 2016, 2016, 1572697 CrossRef PubMed.
- M. M. Bazargani and J. Rohloff, Antibiofilm activity of essential oils and plant extracts against Staphylococcus aureus and Escherichia coli biofilms, Food Control, 2016, 61, 156–164 CrossRef CAS.
- A. Majumder, C. Sarkar, I. Das, S. Sk, S. Bandyopadhyay and S. Mandal, et al., Design, Synthesis and Evaluation of a Series of Zinc(II) Complexes of Anthracene-Affixed Multifunctional Organic Assembly as Potential Antibacterial and Antibiofilm Agents against Methicillin-Resistant Staphylococcus aureus, ACS Appl. Mater. Interfaces, 2023, 15, 22781–22804 CrossRef CAS PubMed.
- R. Gondru, K. Sirisha, S. Raj, S. K. Gunda, C. G. Kumar and M. Pasupuleti, et al., Design, Synthesis, In Vitro Evaluation and Docking Studies of Pyrazole-Thiazole Hybrids as Antimicrobial and Antibiofilm Agents, ChemistrySelect, 2018, 3, 8270–8276 CrossRef CAS.
- Z. Zhang, K. Li, G.-Y. Zhang, Y.-Z. Tang and Z. Jin, Design, synthesis and biological activities of novel pleuromutilin derivatives with a substituted triazole moiety as potent antibacterial agents, Eur. J. Med. Chem., 2020, 204, 112604 CrossRef CAS PubMed.
- A. M. Shehabeldine, B. H. Amin, F. A. Hagras, A. A. Ramadan, M. R. Kamel and M. A. Ahmed, et al., Potential Antimicrobial and Antibiofilm Properties of Copper Oxide Nanoparticles: Time-Kill Kinetic Essay and Ultrastructure of Pathogenic Bacterial Cells, Appl. Biochem. Biotechnol., 2023, 195, 467–485 CrossRef CAS.
- T. Appiah, Y. D. Boakye and C. Agyare, Antimicrobial Activities and Time-Kill Kinetics of Extracts of Selected Ghanaian Mushrooms, Evid. Based Complement. Alternat. Med., 2017, 2017, 4534350 CrossRef PubMed.
- S. Foerster, M. Unemo, L. J. Hathaway, N. Low and C. L. Althaus, Time-kill curve analysis and pharmacodynamic modelling for in vitro evaluation of antimicrobials against Neisseria gonorrhoeae, BMC Microbiol., 2016, 16, 216 CrossRef PubMed.
- L. de Sousa Eduardo, T. C. Farias, S. B. Ferreira, P. B. Ferreira, Z. N. Lima and S. B. Ferreira, Antibacterial Activity and Time-kill Kinetics of Positive Enantiomer of α-pinene Against Strains of Staphylococcus aureus and Escherichia coli, Curr. Top. Med. Chem., 2018, 18, 917–924 CrossRef CAS PubMed.
- S. Liu, R. J. Ono, H. Wu, J. Y. Teo, Z. C. Liang and K. Xu, et al., Highly potent antimicrobial polyionenes with rapid killing kinetics, skin biocompatibility and in vivo bactericidal activity, Biomaterials, 2017, 127, 36–48 CrossRef CAS PubMed.
- J. Li, Y. Wei, J. Wang, Y. Li, G. Shao and Z. Feng, et al., Characterization of Mutations in DNA Gyrase and Topoisomerase IV in Field Strains and In Vitro Selected Quinolone-Resistant Mycoplasma hyorhinis Mutants, Antibiotics, 2022, 11, 494 CrossRef CAS PubMed.
- M. D. Huband, P. A. Bradford, L. G. Otterson, G. S. Basarab, A. C. Kutschke and R. A. Giacobbe, et al., In Vitro Antibacterial Activity of AZD0914, a New Spiropyrimidinetrione DNA Gyrase/Topoisomerase Inhibitor with Potent Activity against Gram-Positive, Fastidious Gram-Negative, and Atypical Bacteria, Antimicrob. Agents Chemother., 2015, 59, 467–474 CrossRef PubMed.
- L. Coronado, X.-Q. Zhang, D. Dorta, N. Escala, L. M. Pineda and M. G. Ng, et al., Semisynthesis, Antiplasmodial Activity, and Mechanism of Action Studies of Isocoumarin Derivatives, J. Nat. Prod., 2021, 84, 1434–1441 CrossRef CAS.
- S. Roychoudhury, K. M. Makin, T. L. Twinem, D. T. Stanton, S. L. Nelson and C. E. Catrenich, Development and Use of a High-Throughput Bacterial DNA Gyrase Assay to Identify Mammalian Topoisomerase II Inhibitors with Whole-Cell Anticancer Activity, SLAS Discov., 2003, 8, 157–163 CrossRef CAS PubMed.
- M. Mora-Pale, N. Bhan, S. Masuko, P. James, J. Wood and S. McCallum, et al., Antimicrobial mechanism of resveratrol-trans-dihydrodimer produced from peroxidase-catalyzed oxidation of resveratrol, Biotechnol. Bioeng., 2015, 112, 2417–2428 CrossRef CAS PubMed.
- E. A. Fayed, M. A. Ebrahim, U. Fathy, A. M. Elawady, W. S. Khalaf and T. M. Ramsis, Pyrano-coumarin hybrids as potential antimicrobial agents against MRSA strains: Design, synthesis, ADMET, molecular docking studies, as DNA gyrase inhibitors, J. Mol. Struct., 2024, 1295, 136663 CrossRef CAS.
- E. A. Fayed, E. M. E. Al-Arab, A. S. Saleh, A. H. Bayoumi and Y. A. Ammar, Design, synthesis, in silico studies, in vivo and in vitro assessment of pyridones and thiazolidinones as anti-inflammatory, antipyretic and ulcerogenic hits, J. Mol. Struct., 2022, 1260, 132839 CrossRef CAS.
- M. Ganesan, J. Sekar, S. Palani Kandasamy and P. Srinivasan, Design, synthesis, spectral characterization, in silico ADMET studies, molecular docking, antimicrobial activity, and anti breast cancer activity of 5,6-dihydrobenzo[H]quinazolines, J. Mol. Struct., 2024, 1296, 136771 CrossRef CAS.
- S. Ramkumar and R. Ramarajan, Design, Synthesis, Spectral Characterization, Antioxidant Activity, Molecular Docking and in silico ADMET Studies of 1, 3 Oxazepines, ChemistrySelect, 2023, 8, e202204818 CrossRef CAS.
- S. Ramkumar and R. Ramarajan, Design, synthesis, spectral characterization, multiple-biological activities, docking and in silico ADMET studies of thiazolidinones, Results Chem., 2023, 5, 100861 CrossRef CAS.
- A. K. Das, P. Paul, M. P. Pranto, M. J. Hassan, K. Saha and M. E. Hossain, Design, synthesis, characterization, antimicrobial activity, cytotoxicity, molecular docking, and in-silico ADMET analysis of the novel cefuroxime derivatives, Eur. J. Med. Chem. Rep., 2024, 10, 100129 CAS.
- P. F. Lamie and J. N. Philoppes, Design, synthesis, stereochemical determination, molecular docking study, in silico pre-ADMET prediction and anti-proliferative activities of indole-pyrimidine derivatives as Mcl-1 inhibitors, Bioorg. Chem., 2021, 116, 105335 CrossRef CAS PubMed.
- S. Hussein, E. A. Fayed, A. Ragab, M. S. Abusaif, Y. A. Ammar and R. El-Sayed Mansou, et al., Flufenamic acid-based sulfonohydrazide and acetamide derivatives NSAI as inhibitors of multi-targets COX-1/COX-2/5-LOX: design, synthesis, in silico ADMET and binding mode studies, BMC Chem., 2025, 19, 271 CrossRef CAS PubMed.
- E. A. Fayed, M. A. A. Najm, K. H. Oudah, M. A. Ebrahim, N. A. Gohar and K. Abu-Elfotuh, et al., From bench to brain: novel thieno-oxazine hybrids as potent pleiotropic anti-Alzheimer’s agents with in vivo/in vitro validation and in silico insights, J. Enzyme Inhib. Med. Chem., 2026, 41, 2598741 CrossRef PubMed.
- S. Hertig, T. D. Goddard, G. T. Johnson and T. E. Ferrin, Multidomain Assembler (MDA) Generates Models of Large Multidomain Proteins, Biophys. J., 2015, 108, 2097–2102 CrossRef CAS PubMed.
- J. E. Chen, C. C. Huang and T. E. Ferrin, RRDistMaps: a UCSF Chimera tool for viewing and comparing protein distance maps, Bioinformatics, 2015, 31, 1484–1486 CrossRef CAS PubMed.
- C. C. Huang, E. C. Meng, J. H. Morris, E. F. Pettersen and T. E. Ferrin, Enhancing UCSF Chimera through web services, Nucleic Acids Res., 2014, 42, W478–W484 CrossRef CAS PubMed.
- Z. Yang, K. Lasker, D. Schneidman-Duhovny, B. Webb, C. C. Huang and E. F. Pettersen, et al., UCSF Chimera, MODELLER, and IMP: an integrated modeling system, J. Struct. Biol., 2012, 179, 269–278 CrossRef CAS PubMed.
- M. D. Hanwell, D. E. Curtis, D. C. Lonie, T. Vandermeersch, E. Zurek and G. R. Hutchison, Avogadro: an advanced semantic chemical editor, visualization, and analysis platform, J. Cheminf., 2012, 4, 17 CAS.
- A. M. Korkor, H. S. Abbass, A. H. Ahmed, A. M. Mansour, T. M. Ramsis and R. H. Elkousy, et al., Insulinotropic and anti-obesity properties of ethno-medicinal plants: pharmacology-based and in-silico predictions, Nat. Prod. Res., 2025, 1–10 CrossRef PubMed.
- R. Huey and G. M. Morris, Using AutoDock 4 with AutoDocktools: a Tutorial, The Scripps Research Institute, USA, 2008, vol. 8, pp. 54–56 Search PubMed.
- Y. A. Ammar, E. A. Fayed, A. H. Bayoumi, M. A. Saleh and M. E. El-Araby, Design and synthesis of pyridine-amide based compounds appended naproxen moiety as anti-microbial and anti-inflammatory agents, Am. J. PharmTech Res., 2015, 5, 245–273 CAS.
- H. I. Ali, K. Tomita, E. Akaho, H. Kambara, S. Miura and H. Hayakawa, et al., Antitumor studies. Part 1: Design, synthesis, antitumor activity, and AutoDock study of 2-deoxo-2-phenyl-5-deazaflavins and 2-deoxo-2-phenylflavin-5-oxides as a new class of antitumor agents, Bioorg. Med. Chem., 2007, 15, 242–256 CrossRef CAS PubMed.
- A. Mai, S. Massa, R. Ragno, I. Cerbara, F. Jesacher and P. Loidl, et al., 3-(4-Aroyl-1-methyl-1H-2-pyrrolyl)-N-hydroxy-2-alkylamides as a New Class of Synthetic Histone Deacetylase Inhibitors. 1. Design, Synthesis, Biological Evaluation, and Binding Mode Studies Performed through Three Different Docking Procedures, J. Med. Chem., 2003, 46, 512–524 CrossRef CAS PubMed.
- H. A. Elseedy, C. Kiriacos and T. M. Ramsis, Molecular Modeling and Drug Development, in Applied Biotechnology and Bioinformatics, 2024, pp. 109–137 Search PubMed.
- E. W. Bell and Y. Zhang, DockRMSD: an open-source tool for atom mapping and RMSD calculation of symmetric molecules through graph isomorphism, J. Cheminf., 2019, 11, 40 Search PubMed.
- S. Bondock, T. Albarqi, M. M. Abdou and N. M. Mohamed, Computational insights into novel benzenesulfonamide-1,3,4-thiadiazole hybrids as a possible VEGFR-2 inhibitor: design, synthesis and anticancer evaluation with molecular dynamics studies, New J. Chem., 2023, 47, 20602–20618 RSC.
| This journal is © The Royal Society of Chemistry 2026 |