 Open Access Article
Open Access ArticleDevelopment, anti-proliferative activity, multi-target kinase inhibition against CHK1, PIM1, and CDK-2, and computational insights of new thiazole-based hybrids
Najla A. Altwaijry a,
Ismail M. M. Othmanb,
Manal M. Anwar
a,
Ismail M. M. Othmanb,
Manal M. Anwar *c,
Soha S. Nosierd,
Asmaa Saleh
*c,
Soha S. Nosierd,
Asmaa Saleh a,
Nada Y. Tashkandie and
Eman S. Nossier*f
a,
Nada Y. Tashkandie and
Eman S. Nossier*f
aDepartment of Pharmaceutical Sciences, College of Pharmacy, Princess Nourah bint Abdulrahman University, P.O. Box 84428, Riyadh 11671, Saudi Arabia
bDepartment of Chemistry, Faculty of Science, Al-Azhar University, Assiut 71524, Egypt
cDepartment of Therapeutic Chemistry, Pharmaceutical and Drug Industries Research Institute, National Research Centre, El-Bohouth Street, Dokki, P.O. Box 12622, Cairo, Egypt. E-mail: manal.hasan52@live.com
dMedical Biochemistry Department, Faculty of Medicine, Damietta University, Damietta, 34511, Egypt
eDepartment of Chemistry, Faculty of Science, King Abdulaziz University, P.O. Box 42805, Jeddah 21551, Saudi Arabia
fPharmaceutical Medicinal Chemistry and Drug Design Department, Faculty of Pharmacy (Girls), Al-Azhar University, Cairo, 11754, Egypt. E-mail: dr.emannossier@gmail.com; dremannossier@azhar.edu.eg
First published on 13th March 2026
Abstract
In the current medical landscape, multi-targeting by a single small molecule is recognized as an effective strategy in the fight against cancer. This study contributes to the global effort to combat cancer by focusing on the synthesis of novel thiazolopyrazoles 2–7, thiazolodiazenylyhiazoles 9a–c, and thiazolotriazolopyrimidines 11a–c. The diazonium salt of 2-aminothiazole was reacted with 3-chloroacetyl acetone, resulting in the formation of 2-oxo-N′-(thiazol-2-yl)propanehydrazonoyl chloride (1). This compound served as a key intermediate precursor for synthesizing the latter compounds, which were proposed as anti-proliferative candidates with multi-kinase inhibitory activities against CHK1, PIM1, and CDK-2. Most of the derivatives exhibited significant cytotoxic activities when assessed for their antiproliferative effects against various tumor cell lines, including lung (EKVX), breast (MCF-7), and colon (HCT116). Derivative 11c, featuring a triazolo[4,3-a]pyrimidine scaffold, exhibited significant antiproliferative activity against the evaluated cell lines (IC50 = 4.8, 5.6, and 6.50 µM, respectively). Furthermore, 11c demonstrated a favorable safety profile against FHC and MCF10A normal cells and showed notable inhibitory activity against the kinases PIM1, CDK-2, CK2α, and CHK1 (IC50 = 0.49 ± 0.02, 0.845 ± 0.05, 4.87 ± 0.18, and 0.032 ± 0.002 µM, respectively). Biological assays investigated the ability of compound 11c to induce apoptosis in MCF-7 cells, arrest the cell cycle at the G1/S phase, and suppress the growth activity of MCF-7 (the wound closure % = 67.407 ± 2.17%, compared to 94.815 ± 3.05% for untreated cells). Docking simulations suggested potential binding modes for 11c, which aligned closely with findings from enzymatic examinations. The in silico physicochemical properties, drug-likeness metrics, and ligand efficiency of 11c appeared to be promising.
1 Introduction
Cancer is a significant problem impacting the health of all human societies and stands as one of the leading causes of mortality.1,2 This disease is linked to genetic instability, which leads to it interfering with the normal regulation of the cell cycle, eventually causing cells to grow and spread in an abnormal way.3,4 Several studies have shown that selective anticancer drugs must effectively eliminate target malignancies while minimizing damage to healthy cells.5–7 The growing understanding of kinase proteins has driven the development of kinase inhibitors, which serve as targeted therapies that exhibit minimal toxicity to normal cells, in contrast to traditional cytotoxic chemotherapies.8–10Activin receptor-like kinase-2 (ALK2) plays an important role in controlling cell growth, proliferation, and survival and, in cancer, oncogenic transformation and immune evasion.11–13 There are various FDA-approved ALK inhibitors used for cancer treatment. Crizotinib is the first ALK inhibitor approved by the US FDA,14–16 ceritinib, alectinib, and brigatinib represent the second generation, and the third-generation ALK TKI, lorlatinib, has shown enhanced activity against commonly known resistance mutations.16
PIM (Provirus Integration in Maloney) kinases are serine/threonine kinases.17,18 Three isoforms—PIM-1, PIM-2, and PIM-3—compose the PIM kinase family.19,20 PIM kinases are pathologically overexpressed in various solid tumors and hematological malignancies. Notably, PIM kinases are absent in benign tumors.21–25 The upregulation of Pim-1 kinase results in increased levels of Bcl-2 and decreases the release of the pro-apoptotic protein Bax, which leads to the inhibition of apoptosis.26 Most research on Pim inhibitors has concentrated on Pim-1 inhibitors, largely due to Pim-2's significantly lower Km for ATP, which is 100 times lower than that of Pim-1 and Pim-3.27 Currently, several PIM kinase inhibitors are undergoing clinical trials in phase I/II28–31 (Fig. S1, SI).
Furthermore, cyclin-dependent kinases (CDKs) are another type of serine/threonine kinases that play a vital role in regulating transcription, the cell cycle, and various other cellular processes.32,33 The dysregulation of cyclin-dependent kinases (CDKs), often due to overexpression or hyperactivation, is a hallmark of various malignancies. This dysregulation results in uncontrolled growth, genomic instability, and tumor progression.34 CDK-2 is rendered as a prospective therapeutic target because it regulates the cell G1/S phase transition and is also involved in many signaling pathways that are essential for the survival and proliferation of cancer cells.35 Roscovitine (V), dinaciclib (VI), and fadraciclib (VII) are FDA-approved drugs that exhibit improved potency and selectivity for various CDKs, specifically CDK-2.34,36,37 It has been documented that multi-kinase inhibitors improve anticancer efficacy by simultaneously inhibiting many oncogenic drivers, in contrast to single kinase inhibitors, which frequently result in compensatory pathway activation and drug resistance.38
Heterocyclic nuclei possess the ability to engage in various intermolecular interactions, including hydrogen bonding, pi stacking, metal coordination bonds, van der Waals forces, and hydrophobic interactions. This versatility enables them to bind to different enzymes in diverse ways.39–41 Various thiazole-based compounds play significant roles as anticancer agents due to their ability to bind to multiple cancer-specific protein targets.41 Several FDA-approved anticancer drugs, currently in therapeutic use, primarily feature the thiazole nucleus. These include dabrafenib,42 dasatinib,43 ixabepilone,44 alpelisib,45 bleomycin,46 and tiazofurin.47 Additionally, pyrazole and [1,2,4]triazolo[4,3-a]pyrimidine ring systems represent other heterocyclic examples with potential anticancer activity, targeting various cancer-specific proteins48,49 (Fig. S2, SI).
The structure–activity relationships (SAR) shared by the most effective multi-kinase inhibitors have been clarified as follows: (a) hydrophobic interactions with several amino acids that line the hydrophobic pocket of the ATP binding site, (b) fused planar heterocycles that occupy the same spatial configuration as the adenine ring of ATP, and (c) hydrogen bond interactions with the hinge region (Fig. 1). When designing the new compounds 2–11, drugs I–IV served as templates. Based on the pharmacophoric cores variation approach, various substituted heterocyclic rings, such as thiazole, pyrazole, and triazolo[4,3-a]pyrimidine, were utilized to interact with the hinge region and replace adenine in the ATP-interacting site. Molecular optimization was performed around these scaffolds by selecting different substituents at various positions that contain nitrogen or oxygen atoms, including acetyl, amino, carbonitrile, carboxylate, and carboxamide aiming to facilitate binding with the active sites of the target kinases through hydrogen bonding. Additionally, several substituted phenyl and methyl groups were incorporated into the novel compounds to engage with the hydrophobic pocket and strengthen the hydrophobic interactions in the hinge area. A nitrile group is known to affect the physicochemical and pharmacokinetic properties of compounds, which enhances both bioavailability and selectivity toward target enzymes by promoting the formation of various types of bonds. Moreover, the carbonitrile group improves compound stability by blocking metabolically labile sites.50 So, a cyano substituent was incorporated into compounds 2, 5, and 11a–c (Fig. 3). Consistent with earlier findings and aiming to obtain new antitumor agents with synergistic efficacy, the strategy of this study was focused on designing and synthesizing new thiazolopyrazole and thiazolotriazolo[4,3-a]pyrimidine-based agents that comply with the pharmacophoric characteristics of protein kinase inhibitors (Fig. 3). All the new analogs were assessed for their cytotoxic efficacy against lung (EKVX), breast (MCF-7), and colon (HCT116) cancer cells, utilizing MTT assay.51,52 The most potent cytotoxic triazolo[4,3-a]pyrimidine congener, 11c was further assessed as a multi-targeting suppressor against ALK2, PIM1, CDK-2, CK2α, and CHK1 kinase enzymes and its effect on the cell cycle, apoptosis induction, and wound healing. Based on the encouraging enzyme inhibition by 11c, a docking simulation and prediction of ADMET were established.
2 Experimental
2.1. Chemistry
The experimental details of the synthesized compounds and their spectral characterization are transferred to the SI section.
2.2. Biological activities
The methods used for antiproliferative investigations kinase inhibitory activities, cell cycle, apoptosis, and wound healing analyses, were described in details in the SI.2.3. In silico studies
The computational analyses included molecular docking with MOE software (Molecular Operating Environment) (MOE-Dock) version 2024.0601 (Chemical Computing Group, Montreal, Canada) and ADMET prediction via the free online SwissADME and AdmetSAR 1.0 websites. The SI contained further information on molecular docking.3 Results and discussion
3.1. Chemistry
The synthetic strategy for preparing the target congeners is depicted in Schemes 1–3. The synthesis began with the treatment of 2-aminothiazole with concentrated hydrochloric acid (HCl) and sodium nitrite in an ice bath at temperatures between 0 and 5 °C, which facilitated the formation of the diazonium salt. This salt was subsequently added to a solution of 3-chloroacetyl acetone in ethanol containing sodium acetate, leading to the formation of the corresponding 2-oxo-N′-(thiazol-2-yl)propanehydrazonoyl chloride (1). The first group of thiazolopyrazoles, 2–7, was formed by mixing the important propanehydrazonoyl chloride derivative 1 with a number of active methylene compounds, such as malononitrile, ethyl 2-cyanoacetate, 2-cyanoacetamide, 3-oxo-3-phenylpropanenitrile, pentane-2,4-dione, and ethyl 3-oxobutanoate, in a sodium ethoxide solution for 8 hours.The proposed mechanism for the formation of the substituted pyrazole ring involves a nucleophilic attack by active methylene derivatives on hydrazonoyl chloride 1, resulting in the formation of adduct A. This adduct subsequently undergoes intramolecular cyclization. Specifically, route A leads to the formation of 5-aminothiazol-1H-pyrazole derivatives 2–4 when malononitrile, ethyl 2-cyanoacetate, or 2-cyanoacetamide is used. In contrast, the 5-phenylthiazol-1H-pyrazole derivative 5 is formed through route B, utilizing 3-oxo-3-phenylpropanenitrile (Scheme S1, SI).
Furthermore, the cyclocondensation reaction of the starting hydrazonoyl chloride derivative 1 with various thiourea derivatives—specifically thiourea, 1-methyl-2-thiourea, and N-phenyl-2-thiourea—was conducted through the Hantzsch reaction, followed by dehydration. This process resulted in the formation of the corresponding thiazolodiazenylthiazol-2-amine derivatives 9a-c (Schemes 2 and S2 (SI)).
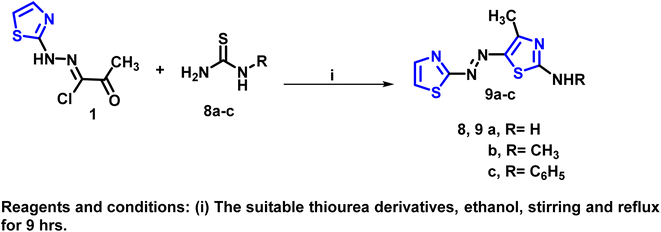 | ||
| Scheme 2 Synthetic approach for preparation of the target thiazolodiazenylthiazole derivatives 9a–c. | ||
Moreover, the thiazolotriazolo[4,3-a]pyrimidine derivatives 11a–c were synthesized through the reaction of the key hydrazonoyl chloride derivative 1 with various 2-thioxo-pyrimidinone derivatives, including 5-isocyano-6-(phenyl/4-methoxyphenyl/4-chlorophenyl)-2-thioxo-2,3-dihydropyrimidin-4(1H)-ones 10a–c, in ethanol that was alkalinized with piperidine. The reaction can proceed via nucleophilic attack by either the nitrogen atom or the sulfur atom of the thiopyrimidine ring (routes A and B, respectively) on the carbon atom of the hydrazonyl chloride that contains the chlorine atom, which results in the elimination of HCl. This nucleophilic substitution is typically followed by intramolecular cyclization, leading to the release of H2S and the formation of the target thiazolotriazolo[4,3-a]pyrimidines 11a–c (Schemes 3 and S3 (SI)).
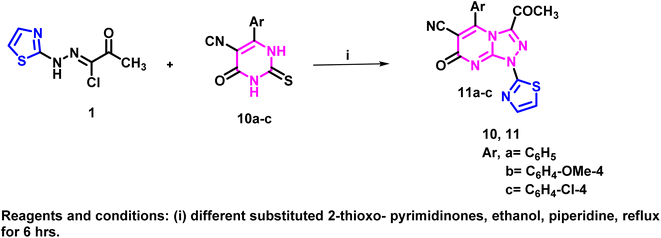 | ||
| Scheme 3 Synthetic approach for preparation of the target thiazolotriazolo[4,3-a]pyrimidine derivatives 11a–c. | ||
All the final compounds, 2–11a–c, exhibited satisfactory elemental analyses, and the obtained spectroscopic data (IR, 1H NMR, and 13C NMR) were consistent with their assigned structures (experimental section).
3.2. Biological activity assessment
In terms of cytotoxic potency against the HCT116 cell line, the tested thiazole-based derivatives 5–11a–c exhibited strong activity, surpassing that of the reference drug, with IC50 values ranging from 2.9 to 6.4 µM and 7.3 µM, respectively. In contrast, compounds 1–4 demonstrated noticeably lower activity, with IC50 values ranging from 13.5 to 18.6 µM.
For the MCF-7 cell line, the thiazolodiazenylthiazole derivative 9c and the triazolo[4,3-a]pyrimidine derivatives 11a–c displayed similar potency to that of doxorubicin, showing IC50 values between 5.6 and 7.9 µM and 4.3 µM, respectively. The other derivatives were less effective, with IC50 values ranging from 8.1 to 17.3 µM.
Regarding cytotoxic activity against EKVX, derivative 11c exhibited 1.6-fold greater activity compared to doxorubicin (IC50 = 4.8 and 7.8 µM, respectively). The remaining derivatives showed lower activity, with IC50 values ranging from 15.1 to 25.7 µM. The results are illustrated in Fig. 2.
 | ||
| Fig. 2 Cytotoxic activity of the new derivatives 1–7, 9a–c, and 11a–c against MCF-7, HCT116, and EKVX cancer cell lines. | ||
The cytotoxicity data indicated that compound 11c represented the most potent antiproliferative activity against the three examined cell lines. Consequently, the in vitro cytotoxicity of 11c was further examined against the normal colon cell line (FHC) and the normal breast cell line (MCF10A). Compound 11c showed a good safety profile, with high IC50 values of 65.90 ± 0.80 µM and 71.55 ± 1.45 µM against the FHC and MCF10A cells, respectively. It also displayed selectivity indexes (SIs) of 10.26 and 13.5, indicating a better safety profile compared to doxorubicin, which has SIs of 4.97 and 6.90, respectively (Table S1 in the SI file).
Based on the obtained results, the following points can be investigated:
• The thiazolotriazolo[4,3-a]pyrimidine scaffold is favorable for potent antiproliferative activity.
• The carbonyl oxygen at position 7 of the triazolo[4,3-a]pyrimidine ring is considered an H-bond acceptor for the target proteins, resulting in potent activity.
• Additionally, the chlorine atom is considered an H-bond donor with different amino acid residues in the target proteins and has a satisfactory fit and potent activity.
| Compound no. | IC50 (µM) (mean ± SEM) | ||||
|---|---|---|---|---|---|
| ALK2 | PIM1 | CDK-2 | CK2α | CHK1 | |
| a IC50: compound concentration resulted in suppression the enzyme activity by 50%, SEM: standard error mean; each value is the mean of three independent tests. | |||||
| 11c | 3.61 ± 0.15 | 0.49 ± 0.02 | 0.845 ± 0.05 | 4.87 ± 0.18 | 0.032 ± 0.002 |
| Lorlatinib | 0.25 ± 0.01 | — | — | — | — |
| Staurosporine | — | 0.32 ± 0.05 | — | — | — |
| Roscovitine | — | — | 0.64 ± 0.012 | — | — |
| Silmitasertib | — | — | — | 0.074 ± 0.002 | — |
| SCH900776 | — | — | — | — | 0.040 ± 0.003 |
As a result, dual inhibitors targeting both enzymes demonstrate a synergistic anticancer effect. CHK1 is a serine/threonine kinase (STK) that plays a crucial role in regulating the cell cycle. In addition to its function in facilitating the repair of DNA damage, CHK1 may also contribute to resistance against chemotherapy and radiation. It is particularly significant during the G2/M phase of the cell cycle.60,61 Reports have indicated that both CHK1 and CDK-2 are upregulated in various cancer types, and there are also dual inhibitors available, such as Dinaciclib III and SCH900776 IV.62,63 Consequently, since compound 11c demonstrated promising inhibitory effects against PIM1 and CDK-2, it warranted further investigation of its inhibitory activity against CK2α and CHK1, with the hope that this derivative could serve as a multitarget inhibitor.11 The results are presented in Table 1. Compound 11c showed notable inhibitory activities against CHK1, with an IC50 value that was very close to that of the reference drug SCH900776 (IC50 = 0.032 ± 0.002 and 0.040 ± 0.003 µM, respectively). The suppression potency against CK2α was significantly lower, with compound 11c displaying an IC50 value of 4.87 ± 0.18 µM. In comparison, the standard drug silmitasertib had an IC50 value of 0.074 ± 0.002 µM (Table 1 and Fig. 3).
As shown in Fig. 4, treatment with 11c resulted in a noticeable decrease in the protein expression levels of PIM1 and CDK-2 in MCF-7 cells compared to the untreated control. A modest decrease in CHK1 expression levels was detected. The obtained protein level downregulation of the abovementioned kinases supported the cellular mechanism of action for 11c that goes beyond direct enzymatic inhibition, potentially leading to its antiproliferative effects.
MCF-7 cells treated with an IC50 of 11c (4.30 µM) experienced cell cycle arrest in the G1/S stages following a 24 hours incubation period [64, 65]. The DNA content increased to 63.44% in the G1 phase and 35.28% in the S phase, compared to the control values of 55.07% and 31.71%, respectively. This increase in DNA content was accompanied by a decrease in the G2/M phase percentage, which dropped to 1.02% compared to the control level of 13.21% (Fig. 5B and S3A).
MCF-7 cells were treated with the IC50 of compound 11c (4.30 µM) for 24 hours. The findings indicated an increase in the proportion of total apoptotic and necrotic cells in MCF-7 (33.21%) compared to the control cells (2.94%). Compound 11c increased early apoptosis from 0.37% (control) to 6.49% and also elevated late apoptosis from 0.025% (control) to 19.57%. Additionally, 11c enhanced the necrotic effect from 2.32% (control) to 7.15% in the treated cells (see Fig. 5A and S3B, SI).
The results suggest that effectively initiating and concluding the apoptotic process, which leads to the breakdown and death of tumor cells, as well as activating apoptosis, are among the promising cytotoxic effects of 11c.
| Compound no. | Wound healing |
|---|---|
| % Closure | |
| 11c/MCF-7 | 67.407 ± 2.17 |
| Cont. MCF-7 | 94.815 ± 3.05 |
3.3. In silico studies
| Compd. no. | Docking score (kcal mol−1) | Amino acid residues (bond length Å) | Atoms of compound | Type of bond |
|---|---|---|---|---|
| 26L/PIM-1 | −10.23 | Lys67(2.88) | Pyrazole(N1) | H-acc |
| Asp128(2.33) | NH2 | H-don | ||
| Glu171(2.39) | H-don | |||
| 11c/PIM-1 | −9.88 | Lys67(3.21) | Triazolo[4,3-a]pyrimidinone (CO) | H-acc |
| Glu171(3.43) | Cl | Halogen-don | ||
| Ile185 | Triazole | Arene-H | ||
| 22K/CHK1 | −9.96 | Leu15 | pyrazolo[1,5-a]pyrimidine | Arene-H |
| Cys87(3.03, 2.87) | pyrazolo[1,5-a]pyrimidine(N1,NH2) | H-acc, H-don | ||
| Glu91, Glu134(2.84) | C6H10(NH2) | Ionic, H-don | ||
| 11c/CHK1 | −9.65 | Glu55(3.06) | Cl | Halogen-don |
| Cys87(3.03, 2.87) | Triazolo[4,3-a]pyrimidinone (CO, N8) | H-don, H-acc | ||
| Roscovitine/CDK-2 | −9.75 | Ile10 | Benzyl | Arene-H |
| Glu81(3.24) | Purine(C8) | H-don | ||
| Leu83(3.18, 2.75) | Purine(C7), benzylamine(NH) | H-acc, H-don | ||
| 11c/CDK-2 | −10.13 | Glu51(2.75, 3.41) | Cl | Halogen-don |
| Phe82(3.26) | Triazolo[4,3-a]pyrimidinone (CO) | H-acc | ||
| Leu83(2.76) | Triazolo[4,3-a]pyrimidinone (CO) | H-acc | ||
| Ile10 | Triazolo[4,3-a]pyrimidinone | Arene-H |
As shown in Fig. 7–9, the whole structure of triazolo[4,3-a]pyrimidine 11c was embedded nicely within the screened receptors' binding sites PIM1, CHK1, and CDK-2, affording significant energy scores of −9.88, −9.65, and −10.13 kcal mol−1, respectively. The carbonyl oxygen at p-7 of triazolo[4,3-a]pyrimidine scaffold gave H-bonds with the key amino acids Lys67 in PIM1, Cys87 in CHK1, and Leu83 in CDK-2, resembling their co-crystallized inhibitors (distances: 3.21, 2.86, and 2.76 Å, respectively). Additionally, the chlorine atom revealed Halogen-bonding with the sidechains of the following amino acids: one with Glu171 in PIM1, another one with Glu55 in CHK1, and two H-bonds with Glu51 in CDK-2 (distances: 3.43, 3.06, 2.75, and 3.41 Å, respectively). Further binding was demonstrated to strengthen the fitting of triazolo[4,3-a]pyrimidine 11c within the screened receptors. The triazole centroid formed an arene–H interaction with Ile185 in PIM1. N-8 at triazolo[4,3-a]pyrimidine moiety revealed an H-bond acceptor with Cys87 backbone (distance: 3.53 Å) in CHK1. Lastly, in CDK-2, the pyrimidine scaffold displayed an arene–H interaction with Ile10 and the carbonyl oxygen at p-7 of triazolo[4,3-a]pyrimidine accepted an H-bond from the Phe82 backbone (distance: 3.26 Å) (Table 3).
 | ||
| Fig. 7 A and B views demonstrate two and three dimensional binding poses of promising triazolo[4,3-a]pyrimidine 11c within the active site of PIM1 (PDB code: 4MBL). | ||
 | ||
| Fig. 8 A and B views demonstrate two and three-dimensional binding poses of promising triazolo[4,3-a]pyrimidine 11c within the active site of CHK1 (PDB code: 3OT3). | ||
It was demonstrated that triazolo[4,3-a]pyrimidine 11c under examination revealed no violation and complied with the Veber and Lipinski's guidelines that could be used to determine which medication behaves best when taken orally (Table 4).
| Compd | Violationsa | MWb | nHBDc | nHBA d | nRBe | TPSAf (Å2) | MLogPg |
|---|---|---|---|---|---|---|---|
| a Violations from Lipinski and Veber Rules.b Molecular weight.c Number of hydrogen bond donor.d Number of hydrogen bond acceptor.e Number of rotatable bond.f Calculated lipophilicity (MLog Po/w).g Topological polar surface area. | |||||||
| Rule | — | ≤500 | ≤5 | ≤10 | ≤10 | ≤140 | ≤4.15 |
| 11c | 0 (Lipinski & Veber) | 396.81 | 0 | 6 | 3 | 134.18 | 1.27 |
The assessed triazolo[4,3-a]pyrimidine 11c was in the optimal range (pink region) of the essential characteristics (lipophilicity, size, solubility, polarity, and flexibility), with the exception of saturation, according to the bioavailability radar chart (Fig. 10A). As a result, this study offered strong evidence regarding the compound under investigation's oral bioavailability.
Fig. 10B and Table S8 demonstrate that potential triazolo[4,3-a]pyrimidine 11c has a higher likelihood of gastrointestinal absorption with no blood–brain barrier penetration. Analog 11c can therefore be used to treat peripheral disorders without resulting in central problems. The tested triazolo[4,3-a]pyrimidine 11c, being a non-substrate for P-glycoprotein, a drug efflux transporter, suggests that the cells may exhibit maximum activity with only a slight efflux.
Additionally, it's predicted to be distributed and localized within the mitochondria. The target 11c is a non-inhibitor to the majority of CYPs, anticipating its little effect upon other active molecules and a low incidence of drug–drug interactions. Since 11c did not block the potassium channel linked to hERG as expected, cardiotoxicity or other cardiac side effects are unlikely. Furthermore, it exhibited no signs of Ames toxicity, which suggests that there is no chance of genotoxicity.
It produced a value of 728.2 mg kg−1, which was categorized as a harmless substance and fell into the third category based on the estimation of acute oral toxicity. At a value of 532.1 mg kg−1 body weight per day, the carcinogenicity descriptor suggests that it may also be classified as non-carcinogenic and non-required. It was predicted that it would be a non-biodegradable compound upon evaluation of its capacity to break down in the environment.
4 Conclusion
In this study, new derivatives of thiazolopyrazoles 2–7, thiazolodiazenylthiazoles 9a–c, and thiazolotriazolopyrimidine 11a–c were designed and synthesized as antiproliferative agents with potential multitarget kinase inhibition against CHK1, PIM1, and CDK-2.The diazonium salt of 2-aminothiazole was allowed to react with 3-chloroacetyl acetone, resulting in the formation of 2-oxo-N′-(thiazol-2-yl)propanehydrazonoyl chloride (1). This compound served as a key intermediate precursor for formation of the target derivatives 2–11a–c. The antiproliferative potency of these new compounds was evaluated against three cancer cell lines: lung (EKVX), breast (MCF-7), and colon (HCT116), using doxorubicin as a reference drug. The thiazolotriazolo[4,3-a]pyrimidine derivative 11c exhibited the most potent antiproliferative activity among the tested compounds, with IC50 values of 4.8, 5.6, and 6.50 µM for the respective cell lines. In contrast, doxorubicin showed IC50 values of 7.85, 4.30, and 7.35 µM for the same cell lines. Derivative 11c surpassed the cytotoxic activity of the reference drug against EKVX by a factor of 1.6. Furthermore, when compared to normal colon cells (FHC) and normal breast cells (MCF10A), it displayed remarkable selectivity—approximately 10-fold for colon cancer cells (HCT-116) and 14-fold for breast cancer cells (MCF-7). This significant selectivity suggests a potential safety profile that may be beneficial for therapeutic applications.
Moreover, compound 11c demonstrated significant inhibitory activity against the kinases PIM1 and CDK-2, with IC50 values of 0.49 ± 0.02 and 0.845 ± 0.05 µM, respectively. In comparison, the standard drugs staurosporine and roscovitine showed IC50 values of 0.32 ± 0.05 and 0.64 ± 0.012 µM. Additionally, 11c exhibited notable inhibitory effects against CK2α and CHK1, with IC50 values that were comparable to those of the reference drugs silmitasertib and SCH900776, which had IC50 values of 4.87 ± 0.18, 0.032 ± 0.002, 0.074 ± 0.002, and 0.040 ± 0.003 µM, respectively.
The ability of compound 11c to induce apoptosis in MCF-7 cells, to arrest cell cycle progression at the G1/S phase, and to suppress the growth activity of MCF-7 cancer cells—demonstrated by cell migration percentage of 67.407 ± 2.17% compared to untreated cells with cell migration percentage of 94.815 ± 3.05%—supports its potential as an anticancer agent. Furthermore, molecular docking and in silico ADMET studies provided valuable insights into the binding modes of 11c with the examined kinases and its pharmacokinetic properties, further validating the experimental findings. The findings indicated that 11c deserves further in vivo pharmacokinetic and pharmacodynamic studies.
Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The data supporting this article have been included as part of the Supplementary information (SI). Supplementary information: Tables S1 and S2, NMR spectra and further experimental details. See DOI: https://doi.org/10.1039/d6ra00253f.Acknowledgements
This research was funded by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R89), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.References
- N. Zhang, J. Wu, Q. Wang, Y. Liang, X. Li, G. Chen, L. Ma, X. Liu and F. Zhou, Global burden of hematologic malignancies and evolution patterns over the past 30 years, Blood Cancer J., 2023, 13(1), 82, DOI:10.1038/s41408-023-00853-3.
- W. M. Eldehna, H. O. Tawfik, M. H. Abdulla, M. S. Nafie, H. Aref, M. A. Shaldam, N. S. Alhassan, O. Al Obeed, Z. M. Elsayed and H. A. Abdel-Aziz, Identification of indole-grafted pyrazolopyrimidine and pyrazolopyridine derivatives as new anti-cancer agents: Synthesis, biological assessments, and molecular modeling insights, Bioorg. Chem., 2024, 153, 107804, DOI:10.1016/j.bioorg.2024.107804.
- D. G. Albertson, C. Collins, F. McCormick and J. W. Gray, Chromosome aberrations in solid tumors, Nat. Genet., 2003, 34(4), 369–376, DOI:10.1038/ng1215.
- Y. Godiyal, D. Maheshwari, H. Taniguchi, S. S. Zinzuwadia, Y. Morera-Díaz, D. Tewari and A. Bishayee, Role of PD-1/PD-L1 signaling axis in oncogenesis and its targeting by bioactive natural compounds for cancer immunotherapy, Mil. Med. Res., 2024, 11(1), 82, DOI:10.1186/s40779-024-00581-0.
- H. A. Gomaa, M. E. Shaker, S. I. Alzarea, O. M. Hendawy, F. A. Mohamed, A. M. Gouda, A. T. Ali, M. M. Morcoss, M. H. Abdelrahman, L. Trembleau and B. G. Youssif, Optimization and SAR investigation of novel 2,3-dihydropyrazino[1,2-*a*]indole-1,4-dione derivatives as EGFR and BRAFV600E dual inhibitors with potent antiproliferative and antioxidant activities, Bioorg. Chem., 2022, 120, 105616, DOI:10.1016/j.bioorg.2022.105616.
- Z. M. Alamshany, N. Y. Tashkandi, I. M. Othman, M. M. Anwar and E. S. Nossier, New thiophene, thienopyridine and thiazoline-based derivatives: Design, synthesis and biological evaluation as antiproliferative agents and multitargeting kinase inhibitors, Bioorg. Chem., 2022, 127, 105964, DOI:10.1016/j.bioorg.2022.105964.
- N. M. Raghavendra, D. Pingili, S. Kadasi, A. Mettu and S. V. Prasad, Dual or multi-targeting inhibitors: The next generation anticancer agents, Eur. J. Med. Chem., 2018, 143, 1277–1300, DOI:10.1016/j.ejmech.2017.10.021.
- S. A. El-Metwally, M. M. Abou-El-Regal, I. H. Eissa, A. B. Mehany, H. A. Mahdy, H. Elkady, A. Elwan and E. B. Elkaeed, Discovery of thieno[2,3-*d*]pyrimidine-based derivatives as potent VEGFR-2 kinase inhibitors and anti-cancer agents, Bioorg. Chem., 2021, 112, 104947, DOI:10.1016/j.bioorg.2021.104947.
- S. Z. Khan and C. G. Lengyel, Challenges in the management of colorectal cancer in low-and middle-income countries, Cancer Treat. Res. Commun., 2023, 35, 100705, DOI:10.1016/j.ctarc.2023.100705.
- X. J. Liu, H. C. Zhao, S. J. Hou, H. J. Zhang, L. Cheng, S. Yuan, L. R. Zhang, J. Song, S. Y. Zhang and S. W. Chen, Recent development of multi-target VEGFR-2 inhibitors for the cancer therapy, Bioorg. Chem., 2023, 133, 106425, DOI:10.1016/j.bioorg.2023.106425.
- M. A. Al-Qadhi, H. A. Allam, S. H. Fahim, T. A. Yahya and F. A. Ragab, Design and synthesis of certain 7-Aryl-2-Methyl-3-Substituted Pyrazolo{1,5-a}Pyrimidines as multikinase inhibitors, Eur. J. Med. Chem., 2023, 262, 115918, DOI:10.1016/j.ejmech.2023.115918.
- T. Ullrich, L. Arista, S. Weiler, S. Teixeira-Fouchard, V. Broennimann, N. Stiefl, V. Head, I. Kramer and S. Guth, Discovery of a novel 2-aminopyrazine-3-carboxamide as a potent and selective inhibitor of Activin Receptor-Like Kinase-2 (ALK2) for the treatment of fibrodysplasia ossificans progressiva, Bioorg. Med. Chem. Lett., 2022, 64, 128667, DOI:10.1016/j.bmcl.2022.128667.
- M. A. Mansour, G. S. Hassan, R. A. Serya, M. Y. Jaballah and K. A. Abouzid, Advances in the discovery of activin receptor-like kinase 5 (ALK5) inhibitors, Bioorg. Chem., 2024, 147, 107332, DOI:10.1016/j.bioorg.2024.107332.
- L. Friboulet, N. Li, R. Katayama, C. C. Lee, J. F. Gainor, A. S. Crystal, P. Y. Michellys, M. M. Awad, N. Yanagitani, S. Kim and A. C. Pferdekamper, The ALK inhibitor ceritinib overcomes crizotinib resistance in non–small cell lung cancer, Cancer Discovery, 2014, 4(6), 662–673, DOI:10.1158/2159-8290.CD-13-0846.
- R. C. Doebele, A. B. Pilling, D. L. Aisner, T. G. Kutateladze, A. T. Le, A. J. Weickhardt, K. L. Kondo, D. J. Linderman, L. E. Heasley, W. A. Franklin and M. Varella-Garcia, Mechanisms of resistance to crizotinib in patients with ALK gene rearranged non–small cell lung cancer, Clin. Cancer Res., 2012, 18(5), 1472–1482, DOI:10.1158/1078-0432.CCR-11-2906.
- R. Katayama, A. T. Shaw, T. M. Khan, M. Mino-Kenudson, B. J. Solomon, B. Halmos, N. A. Jessop, J. C. Wain, A. T. Yeo, C. Benes and L. Drew, Mechanisms of acquired crizotinib resistance in ALK-rearranged lung cancers, Sci. Transl. Med., 2012, 4(120), 120ra17, DOI:10.1126/scitranslmed.3003316.
- G. G. Jinesh, S. Mokkapati, K. Zhu and E. E. Morales, Pim kinase isoforms: devils defending cancer cells from therapeutic and immune attacks, Apoptosis, 2016, 21(11), 1203–1213, DOI:10.1007/s10495-016-1289-3.
- E. Assirelli, J. Ciaffi, V. Scorcu, S. Naldi, V. Brusi, L. Mancarella, L. Lisi, F. Pignatti, F. Ursini and S. Neri, PIM kinases as potential biomarkers and therapeutic targets in inflammatory arthritides, Int. J. Mol. Sci., 2024, 25(6), 3123, DOI:10.3390/ijms25063123.
- F. Jin, Y. Lin, W. Yuan, S. Wu, M. Yang, S. Ding, J. Liu and Y. Chen, Recent advances in c-Met-based dual inhibitors in the treatment of cancers, Eur. J. Med. Chem., 2024, 272, 116477, DOI:10.1016/j.ejmech.2024.116477.
- A. K. Rout, B. Dehury, S. N. Parida, S. S. Rout, R. Jena, N. Kaushik, N. K. Kaushik, S. K. Pradhan, C. R. Sahoo, A. K. Singh and M. Arya, A review on structure-function mechanism and signaling pathway of serine/threonine protein PIM kinases as a therapeutic target, Int. J. Biol. Macromol., 2024, 270, 132030, DOI:10.1016/j.ijbiomac.2024.132030.
- M. Narlik-Grassow, C. Blanco-Aparicio and A. Carnero, The PIM family of serine/threonine kinases in cancer, Med. Res. Rev., 2014, 34(1), 136–159, DOI:10.1002/med.21284.
- J. Xu, G. Xiong, Z. Cao, H. Huang, T. Wang, L. You, L. Zhou, L. Zheng, Y. Hu, T. Zhang and Y. Zhao, PIM-1 contributes to the malignancy of pancreatic cancer and displays diagnostic and prognostic value, J. Exp. Clin. Cancer Res., 2016, 35(1), 133, DOI:10.1186/s13046-016-0410-3.
- R. Jiang, X. Wang, Z. Jin and K. Li, Association of nuclear PIM1 expression with lymph node metastasis and poor prognosis in patients with lung adenocarcinoma and squamous cell carcinoma, J. Cancer, 2016, 7(3), 324, DOI:10.7150/jca.13790.
- L. Chen, W. Mao, C. Ren, J. Li and J. Zhang, Comprehensive insights that targeting PIM for cancer therapy: prospects and obstacles, J. Med. Chem., 2024, 67(1), 38–64, DOI:10.1021/acs.jmedchem.3c01443.
- M. Bellon and C. Nicot, Targeting Pim kinases in hematological cancers: molecular and clinical review, Mol. Cancer, 2023, 22(1), 18, DOI:10.1186/s12943-023-01722-0.
- L. Chen, W. Mao, C. Ren, J. Li and J. Zhang, Comprehensive insights that targeting PIM for cancer therapy: prospects and obstacles, J. Med. Chem., 2024, 67(1), 38–64, DOI:10.1021/acs.jmedchem.3c01443.
- Z. M. Alamshany, N. Y. Tashkandi, I. M. Othman, M. M. Anwar and E. S. Nossier, New thiophene, thienopyridine and thiazoline-based derivatives: Design, synthesis and biological evaluation as antiproliferative agents and multitargeting kinase inhibitors, Bioorg. Chem., 2022, 127, 105964, DOI:10.1016/j.bioorg.2022.105964.
- A. Siddiqui-Jain, D. Drygin, N. Streiner, P. Chua, F. Pierre, S. E. O'Brien, J. Bliesath, M. Omori, N. Huser, C. Ho and C. Proffitt, CX-4945, an orally bioavailable selective inhibitor of protein kinase CK2, inhibits prosurvival and angiogenic signaling and exhibits antitumor efficacy, Cancer Res., 2010, 70(24), 10288–10298, DOI:10.1158/0008-5472.CAN-10-1893.
- F. Pierre, E. Stefan, A. S. Nédellec, M. C. Chevrel, C. F. Regan, A. Siddiqui-Jain, D. Macalino, N. Streiner, D. Drygin, M. Haddach and S. E. O'Brien, 7-(4H-1,2,4-Triazol-3-yl)benzo[*c*][2,6]naphthyridines: a novel class of Pim kinase inhibitors with potent cell antiproliferative activity, Bioorg. Med. Chem. Lett., 2011, 21(22), 6687–6692, DOI:10.1016/j.bmcl.2011.09.058.
- P. D. Garcia, J. L. Langowski, Y. Wang, M. Chen, J. Castillo, C. Fanton, M. Ison, T. Zavorotinskaya, Y. Dai, J. Lu and X. H. Niu, Pan-PIM kinase inhibition provides a novel therapy for treating hematologic cancers, Clin. Cancer Res., 2014, 20(7), 1834–1845, DOI:10.1158/1078-0432.CCR-13-2062.
- M. T. Burger, W. Han, J. Lan, G. Nishiguchi, C. Bellamacina, M. Lindval, G. Atallah, Y. Ding, M. Mathur, C. McBride and E. L. Beans, Structure guided optimization, in vitro activity, and in vivo activity of pan-PIM kinase inhibitors, ACS Med. Chem. Lett., 2013, 4(12), 1193–1197, DOI:10.1021/ml400321k.
- S. Lim and P. Kaldis, Cdks, cyclins and CKIs: roles beyond cell cycle regulation, Development, 2013, 140(15), 3079–3093, DOI:10.1242/dev.091744.
- E. Jamasbi, M. Hamelian, M. A. Hossain and K. Varmira, The cell cycle, cancer development and therapy, Mol. Biol. Rep., 2022, 49(11), 10875–10883, DOI:10.1007/s11033-022-07778-3.
- Q. Hao, W. Zhao, Z. Li, Y. Lai, Y. Wang, Q. Yang and L. Zhang, Combination therapy and dual-target inhibitors based on cyclin-dependent kinases (CDKs): Emerging strategies for cancer therapy, Eur. J. Med. Chem., 2025, 289, 117465, DOI:10.1016/j.ejmech.2024.117465.
- W. Kolch, M. Halasz, M. Granovskaya and B. N. Kholodenko, The dynamic control of signal transduction networks in cancer cells, Nat. Rev. Cancer, 2015, 15(9), 515–527, DOI:10.1038/nrc3984.
- M. M. Mita, A. A. Joy, A. Mita, K. Sankhala, Y. M. Jou, D. Zhang, P. Statkevich, Y. Zhu, S. L. Yao, K. Small and R. Bannerji, Randomized phase II trial of the cyclin-dependent kinase inhibitor dinaciclib (MK-7965) versus capecitabine in patients with advanced breast cancer, Clin. Breast Cancer, 2014, 14(3), 169–176, DOI:10.1016/j.clbc.2013.10.016.
- J. J. Stephenson, J. Nemunaitis, A. A. Joy, J. C. Martin, Y. M. Jou, D. Zhang, P. Statkevich, S. L. Yao, Y. Zhu, H. Zhou and K. Small, Randomized phase 2 study of the cyclin-dependent kinase inhibitor dinaciclib (MK-7965) versus erlotinib in patients with non-small cell lung cancer, Lung Cancer, 2014, 83(2), 219–223, DOI:10.1016/j.lungcan.2013.11.020.
- W. Xu, C. Ye, X. Qing, S. Liu, X. Lv, W. Wang, X. Dong and Y. Zhang, Multi-target tyrosine kinase inhibitor nanoparticle delivery systems for cancer therapy, Mater. Today Bio, 2022, 16, 100358, DOI:10.1016/j.mtbio.2022.100358.
- R. Callaghan, F. Luk and M. Bebawy, Inhibition of the multidrug resistance P-glycoprotein: time for a change of strategy?, Drug Metab. Dispos., 2014, 42(4), 623–631, DOI:10.1124/dmd.113.056176.
- R. Bansal and A. Malhotra, Therapeutic progression of quinazolines as targeted chemotherapeutic agents, Eur. J. Med. Chem., 2021, 211, 113016, DOI:10.1016/j.ejmech.2020.113016.
- F. A. Dos Santos, M. C. Pereira, T. B. de Oliveira, F. J. Junior, M. D. de Lima, M. G. da Rocha Pitta, I. da Rocha Pitta, M. J. de Melo Rêgo and M. G. da Rocha Pitta, Anticancer properties of thiophene derivatives in breast cancer MCF-7 cells, Anti-Cancer Drugs, 2018, 29(2), 157–166, DOI:10.1097/CAD.0000000000000583.
- S. Bowyer, R. Lee, A. Fusi and P. Lorigan, Dabrafenib and its use in the treatment of metastatic melanoma, Melanoma Manag., 2015, 2(3), 199–208, DOI:10.2217/mmt.15.11.
- X. Li, Y. He, C. H. Ruiz, M. Koenig and M. D. Cameron, Characterization of dasatinib and its structural analogs as CYP3A4 mechanism-based inactivators and the proposed bioactivation pathways, Drug Metab. Dispos., 2009, 37(6), 1242–1250, DOI:10.1124/dmd.108.02613.
- J. Li, J. Ren and W. Sun, Systematic review of ixabepilone for treating metastatic breast cancer, Breast Cancer, 2017, 24(2), 171–179, DOI:10.1007/s12282-016-0735-y.
- T. Wilhoit, J. M. Patrick and M. B. May, Alpelisib: a novel therapy for patients with PIK3CA-mutated metastatic breast cancer, J. Adv. Pract. Oncol., 2020, 11(7), 768, DOI:10.6004/jadpro.2020.11.7.7.
- S. M. Hecht, Bleomycin: new perspectives on the mechanism of action, J. Nat. Prod., 2000, 63(1), 158–168, DOI:10.1021/np990549f.
- P. Franchetti, S. Marchetti, L. Cappellacci, H. N. Jayaram, J. A. Yalowitz, B. M. Goldstein, J. L. Barascut, D. Dukhan, J. L. Imbach and M. Grifantini, Synthesis, conformational analysis, and biological activity of C-thioribonucleosides related to tiazofurin, J. Med. Chem., 2000, 43(7), 1264–1270, DOI:10.1021/jm990524p.
- D. Yevale, N. Teraiya, T. Lalwani, M. Dalasaniya, S. K. Patel, N. Dixit, C. B. Sangani, S. Kumar, N. S. Mulukuri, T. Huang and Y. T. Duan, Discovery of new pyrazole-4-carboxamide analogues as potential anticancer agents targeting dual aurora kinase A and B, Eur. J. Med. Chem., 2024, 280, 116917, DOI:10.1016/j.ejmech.2024.116917.
- H. A. Adawy, S. S. Tawfik, A. A. Elgazar, K. B. Selim and F. E. Goda, Design, synthesis, and in vitro and in vivo biological evaluation of triazolopyrimidine hybrids as multitarget directed anticancer agents, RSC Adv., 2024, 14(48), 35239–35254, 10.1039/D4RA06312H.
- J. Wang and H. Liu, Application of nitrile in drug design, Chin. J. Org. Chem., 2012, 32(9), 1643 CrossRef CAS.
- T. Mosmann, Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays, J. Immunol. Methods, 1983, 65(1–2), 55–63, DOI:10.1016/0022-1759(83)90303-4.
- F. A. Ragab, A. A. Eissa, S. H. Fahim, M. A. Salem, M. A. Gamal and Y. M. Nissan, Novel coumarin–pyrazoline hybrids: Synthesis, cytotoxicity evaluation and molecular dynamics study, New J. Chem., 2021, 45(40), 19043–19055, 10.1039/D1NJ03472A.
- L. Brault, C. Gasser, F. Bracher, K. Huber, S. Knapp and J. Schwaller, PIM serine/threonine kinases in the pathogenesis and therapy of hematologic malignancies and solid cancers, Haematologica, 2010, 95(6), 1004, DOI:10.3324/haematol.2009.017079.
- G. K. Gray, B. C. McFarland, A. L. Rowse, S. A. Gibson and E. N. Benveniste, Therapeutic CK2 inhibition attenuates diverse prosurvival signaling cascades and decreases cell viability in human breast cancer cells, Oncotarget, 2014, 5(15), 6484, DOI:10.18632/oncotarget.2248.
- K. Izeradjene, L. Douglas, A. Delaney and J. A. Houghton, Casein kinase II (CK2) enhances death-inducing signaling complex (DISC) activity in TRAIL-induced apoptosis in human colon carcinoma cell lines, Oncogene, 2005, 24(12), 2050–2058, DOI:10.1038/sj.onc.1208391.
- B. Yang, J. Yao, B. Li, G. Shao and Y. Cui, Inhibition of protein kinase CK2 sensitizes non-small cell lung cancer cells to cisplatin via upregulation of PML, Mol. Cell. Biochem., 2017, 436(1), 87–97, DOI:10.1007/s11010-017-3081-2.
- P. Kumar, A. Nagarajan and P. D. Uchil, Analysis of Cell Viability by the MTT Assay, Cold Spring Harb Protoc., 2018, 2018(6) DOI:10.1101/pdb.prot095505.
- J. Sambrook, E. F. Fritsch and T. Maniatis, Molecular Cloning: a Laboratory Manual, Cold Spring Harbor Laboratory, 1989 Search PubMed.
- K. Qian, L. Wang, C. L. Cywin, B. T. Farmer, E. Hickey, C. Homon, S. Jakes, M. A. Kashem, G. Lee, S. Leonard and J. Li, Hit to lead account of the discovery of a new class of inhibitors of Pim kinases and crystallographic studies revealing an unusual kinase binding mode, J. Med. Chem., 2009, 52(7), 1814–1827, DOI:10.1021/jm801401s.
- M. D. Garrett and I. Collins, Anticancer therapy with checkpoint inhibitors: what, where and when?, Trends Pharmacol. Sci., 2011, 32(5), 308–316, DOI:10.1016/j.tips.2011.02.007.
- V. Y. Dudkin, K. Rickert, C. Kreatsoulas, C. Wang, K. L. Arrington, M. E. Fraley, G. D. Hartman, Y. Yan, M. Ikuta, S. M. Stirdivant and R. A. Drakas, Pyridyl aminothiazoles as potent inhibitors of Chk1 with slow dissociation rates, Bioorg. Med. Chem. Lett., 2012, 22(7), 2609–2612, DOI:10.1016/j.bmcl.2012.01.076.
- K. Paruch, M. P. Dwyer, C. Alvarez, C. Brown, T. Y. Chan, R. J. Doll, K. Keertikar, C. Knutson, B. McKittrick, J. Rivera and R. Rossman, Discovery of dinaciclib (SCH 727965): a potent and selective inhibitor of cyclin-dependent kinases, ACS Med. Chem. Lett., 2010, 1(5), 204–208, DOI:10.1021/ml100051d.
- A. Daud, G. M. Springett, D. S. Mendelson, P. N. Munster, J. W. Goldman, J. R. Strosberg, G. Kato, T. Nesheiwat, R. Isaacs and L. S. Rosen, A phase I dose-escalation study of SCH 900776, a selective inhibitor of checkpoint kinase 1 (CHK1), in combination with gemcitabine (Gem) in subjects with advanced solid tumors, J. Clin. Oncol., 2010, 28(15_suppl), 3064 Search PubMed.
- I. Nicoletti, G. Migliorati, M. C. Pagliacci, F. Grignani and C. Riccardi, A rapid and simple method for measuring thymocyte apoptosis by propidium iodide staining and flow cytometry, J. Immunol. Methods, 1991, 139(2), 271–279, DOI:10.1016/0022-1759(91)90198-o.
- I. Vermes, C. Haanen, H. Steffens-Nakken and C. Reutellingsperger, A novel assay for apoptosis flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labelled annexin V, J. Immunol. Methods, 1995, 184(1), 39–51, DOI:10.1016/0022-1759(95)00072-i.
- W. M. Eldehna, H. O. Tawfik, M. H. Abdulla, M. S. Nafie, H. Aref, M. A. Shaldam, N. S. Alhassan, O. Al Obeed, Z. M. Elsayed and H. A. Abdel-Aziz, Identification of indole-grafted pyrazolopyrimidine and pyrazolopyridine derivatives as new anti-cancer agents: Synthesis, biological assessments, and molecular modeling insights, Bioorg. Chem., 2024, 153, 107804, DOI:10.1016/j.bioorg.2024.107804.
- A. E. Mohamed, W. E. Elgammal, A. G. Ibrahim, H. M. Dawaba, E. S. Nossier and A. M. Dawaba, Thiadiazole chitosan conjugates as a novel cosmetic ingredient for rinse-off hair conditioners: design, formulation, characterization and in silico-molecular docking studies, BMC Chem., 2025, 19(1), 104, DOI:10.1186/s13065-025-01419-7.
- G. O. Moustafa, A. Shalaby, A. M. Naglah, M. M. Mounier, H. El-Sayed, M. M. Anwar and E. S. Nossier, Synthesis, characterization, in vitro anticancer potentiality, and antimicrobial activities of novel peptide–glycyrrhetinic-acid-based derivatives, Molecules, 2021, 26(15), 4573, DOI:10.3390/molecules26154573.
- A. E. Amr, R. E. Mageid, M. El-Naggar, A. M. Naglah, E. S. Nossier and E. A. Elsayed, Chiral Pyridine-3,5-bis-(L-phenylalaninyl-L-leucinyl) Schiff Base Peptides as Potential Anticancer Agents: Design, synthesis, and molecular docking studies targeting lactate dehydrogenase-A, Molecules, 2020, 25(5), 1096, DOI:10.3390/molecules25051096.
- R. R. Khattab, A. A. Hassan, D. A. Osman, F. M. Abdel-Megeid, H. M. Awad, E. S. Nossier and W. A. El-Sayed, Synthesis, anticancer activity and molecular docking of new triazolo[4,5-*d*]pyrimidines based thienopyrimidine system and their derived N-glycosides and thioglycosides, Nucleosides Nucleotides Nucleic Acids, 2021, 40(11), 1090–1113, DOI:10.1080/15257770.2021.1966801.
- M. P. Dwyer, K. Keertikar, K. Paruch, C. Alvarez, M. Labroli, C. Poker, T. O. Fischmann, R. Mayer-Ezell, R. Bond, Y. Wang and R. Azevedo, Discovery of pyrazolo[1,5-*a*]pyrimidine-based Pim inhibitors: a template-based approach, Bioorg. Med. Chem. Lett., 2013, 23(22), 6178–6182, DOI:10.1016/j.bmcl.2013.09.001.
- M. Labroli, K. Paruch, M. P. Dwyer, C. Alvarez, K. Keertikar, C. Poker, R. Rossman, J. S. Duca, T. O. Fischmann, V. Madison and D. Parry, Discovery of pyrazolo[1,5-*a*]pyrimidine-based CHK1 inhibitors: A template-based approach—Part 2, Bioorg. Med. Chem. Lett., 2011, 21(1), 471–474, DOI:10.1016/j.bmcl.2010.10.091.
- A. A. El-Sayed, E. S. Nossier, A. A. Almehizia and A. E. Amr, Design, synthesis, anticancer evaluation and molecular docking study of novel 2,4-dichlorophenoxymethyl-based derivatives linked to nitrogenous heterocyclic ring systems as potential CDK-2 inhibitors, J. Mol. Struct., 2022, 1247, 131285, DOI:10.1016/j.molstruc.2021.131285.
- A. F. Kassem, A. Younis, E. S. Nossier, H. M. Awad and W. A. El-Sayed, Pyridine-based glycosides bearing 1,2,4-triazole and their 1,3,4-oxadiazole analogues as potential EGFR and CDK-2 inhibitors: Design, synthesis, antiproliferative activity and in silico studies, J. Mol. Struct., 2024, 1313, 138741, DOI:10.1016/j.molstruc.2024.138741.
- H. E. Hashem, A. E. Amr, A. A. Almehizia, A. M. Naglah, B. M. Kariuki, H. A. Eassa and E. S. Nossier, Nanoparticles of a pyrazolo-pyridazine derivative as potential EGFR and CDK-2 inhibitors: design, structure determination, anticancer evaluation and in silico studies, Molecules, 2023, 28(21), 7252, DOI:10.3390/molecules28217252.
- E. M. Mohi El-Deen, E. S. Nossier and E. A. Karam, New quinazolin-4(3H)-one derivatives incorporating hydrazone and pyrazole scaffolds as antimicrobial agents targeting DNA gyrase enzyme, Sci. Pharm., 2022, 90(3), 52, DOI:10.3390/scipharm90030052.
- D. H. Dawood, M. M. Sayed, S. T. Tohamy and E. S. Nossier, New Thiophenyl-pyrazolyl-thiazole hybrids as DHFR inhibitors: Design, synthesis, antimicrobial evaluation, molecular modeling, and Biodistribution studies, ACS Omega, 2023, 8(42), 39250–39268, DOI:10.1021/acsomega.3c04513.
- A. A. Alsfouk, I. M. Othman, M. M. Anwar, A. Saleh, N. Y. Tashkandi and E. S. Nossier, New quinazolinone-based heterocyclic compounds as promising antimicrobial agents: development, DNA gyrase B/topoisomerase IV inhibition activity, and in silico computational studies, J. Mol. Struct., 2025, 142953, DOI:10.1016/j.molstruc.2024.142953.
- H. S. El-Hema, S. M. Soliman, W. El-Dougdoug, M. H. Ahmed, A. Abdelmajeid, E. S. Nossier, M. F. Hussein, A. A. Alrayes, M. Hassan, N. A. Ahmed and A. Sabry, Design, Characterization, Antimicrobial Activity, and In Silico Studies of Theinothiazoloquinazoline Derivatives Bearing Thiazinone, Tetrazole, and Triazole Moieties, ACS Omega, 2025, 10(9), 9703–9717, DOI:10.1021/acsomega.4c09502.
| This journal is © The Royal Society of Chemistry 2026 |








