Open Access Article
Open Access ArticleNovel pyridine-heterocycle hybrids: synthesis via Hantzsch and Biginelli reactions, docking simulations, and anticancer activity
Hadeer M. Diaba,
Mostafa E. Salem b,
Ahmed H. M. Elwahy
b,
Ahmed H. M. Elwahy *a,
Mohamed A. Raghebc,
Mahmoud A. Noamaand,
Faisal K. Algethami
*a,
Mohamed A. Raghebc,
Mahmoud A. Noamaand,
Faisal K. Algethami b,
Ismail A. Abdelhamid
b,
Ismail A. Abdelhamid *a and
Huda Kamel Mahmouda
*a and
Huda Kamel Mahmouda
aDepartment of Chemistry, Faculty of Science, Cairo University, Giza 12613, Egypt. E-mail: aelwahy@hotmail.com; aelwhy@cu.edu.eg; ismail_shafy@yahoo.com; ismail_shafy@cu.edu.eg
bChemistry Department, College of Science, Imam Mohammad Ibn Saud Islamic University (IMSIU), P.O. Box 90905, Riyadh 11623, Saudi Arabia
cDepartment of Chemistry (Biochemistry Division), Faculty of Science, Cairo University, Giza, 12613, Egypt
dDepartment of Mathematics, Faculty of Science, Cairo University, Giza 12613, Egypt
First published on 4th February 2026
Abstract
Cancer is a serious global health issue and remains one of the top causes of death worldwide. To overcome the problems of the existing anticancer drugs in terms of specificity and resistance, a new class of hybrid bis-heterocyclic compounds with a pyridine bridge has been designed and synthesized via the Hantzsch reaction. The results of elemental analysis and spectral data were used for the confirmation of the synthesized compounds. Among the tested analogs, the highest cytotoxic activity was shown by compound 7 against the HepG2, A549, and MCF7 cancer cell lines, with IC50 values of 18.07, 14.45, and 30.89 µg mL−1, respectively, while the cytotoxicity against normal fibroblasts was negligible with an IC50 greater than 100 µg mL−1. The structure–activity relationship results emphasized the key role of the molecular planarity and sulfur atom substitution. The results from molecular docking, molecular dynamics simulations, and MM/PBSA and MM/GBSA binding free energy calculations indicated a strong binding of compound 7 with EGFR. The results of the EGFR inhibition assay further strengthened the binding of the EGFR with the mentioned compound. The results from the ADME and toxicological analyses indicated a good pharmacokinetic and safety profile for the compound.
1 Introduction
Cancer, a disease characterized by the unregulated growth of cells that can invade other parts of the body, is identified as one of the most perilous threats to the world's health. According to the GLOBOCAN 2022 database, there were 19.9 million incident cases of cancer as well as 9.6 million deaths due to cancer worldwide. Cancer has proven itself to be one of the principal causes of morbidity as well as mortality rates in the global scenario. Future years will experience a tremendous rise in the incidence of cancer in the global scenario.1 Consequently, it is essential to seek out innovative medications that possess high efficacy, minimal toxicity, and low mutagenicity, along with targeted anticancer effects that can effectively combat the resistance commonly developed against existing treatments. Surgery, radiation therapy, and chemotherapy can be utilized separately or in combination with cancer treatments.2,3Heterocyclic anticancer drugs represent a significant category of medications used in cancer treatments. They are extensively researched in medicinal chemistry because of their various biological effects, particularly their crucial involvement in the development of anticancer drugs. These substances are essential in cancer therapy because of their capacity to target different biological pathways, such as DNA replication, enzyme inhibition, and the induction of apoptosis. The kind or dimension of the ring of heterocycles, together with the substituent groups and structural variations, allows for interactions with distinctly different structural configurations of enzyme binding sites, which significantly affect the physicochemical traits and, in turn, the pharmacological properties of the molecules.4,5 Among various heterocycles, pyridine derivatives have attracted considerable interest as possible anticancer agents because of their capacity to engage with diverse biological targets, such as enzymes, receptors, and DNA.6–10
Fused heterocyclic compounds containing a pyridine element also exhibit a variety of biological activities across different areas. Generally, substances with pyridine and dihydropyridine are predominantly used for diverse applications, including combating microbes, viruses, cancers, hypertension, diabetes, malaria, and inflammation, while also serving as antioxidants and psychopharmacological antagonists, and addressing amoebic infections.11–15 These structures are essential components in the chemical frameworks of many approved drugs available today (Fig. 1).
Molecular hybridization is a method in drug design and development that entails merging pharmacophoric components from different bioactive substances to produce a new hybrid compound with improved affinity and efficacy relative to the original drugs. In recent years, significant advancements have occurred in this domain, with several hybrid compounds featuring strong heterocycles frequently found in the drug discovery field within the pharmaceutical and agrochemical industries.16,17 This study details the creation of new hybrid compounds, including pyridine-derived polyhydroquinoline, bis(hexahydroacridine), bis(hexahydropyrimido[4,5-b]quinoline), and bis(tetrahydrodipyrazolo[3,4-b:4′,3′-e]pyridine), using Hantzsch reactions. The cytotoxic effects and in vitro anticancer activities of the newly created compounds were evaluated against chosen human cancer cell lines. This study builds upon our earlier investigations into generating bioactive heterocycles and bis(heterocycles).18–46
2 Experimental
2.1 Chemistry
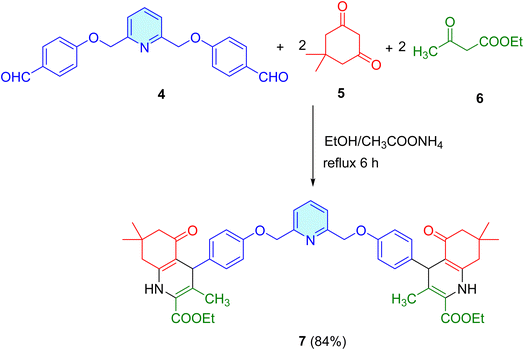
2.1.1.1 Diethyl 4,4′-(((pyridine-2,6-diylbis(methylene))bis(oxy))bis(4,1-phenylene))bis(3,7,7-trimethyl-5-oxo-1,4,5,6,7,8-hexahydroquinoline-2-carboxylate) (7). A mixture of bis-aldehyde 4 (0.35 g, 1 mmol), dimedone (5) (0.28 g, 2 mmol), ethyl acetoacetate (6) (0.26 g, 2 mmol) was heated in absolute ethanol (30 mL) containing ammonium acetate (0.15 g, 2 mmol) for 6 h. The excess solvent was evaporated under reduced pressure, and the collected residue was recrystallized from EtOH.
Yellow powder (0.67 g, 83%); Mp 272–274 °C; IR (KBr): ν 3279 (NH), 1697 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ester), 1643 (C
O ester), 1643 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.82 (s, 6H, 2CH3), 0.97 (s, 6H, 2CH3), 1.09 (t, J = 7.1 Hz, 6H, 2CH3CH2COO), 1.93 (d, 2H, H8, J = 15 Hz) 2.11 (d, 2H, H8, J = 15 Hz), 2.25 (s, 6H, 2CH3), 2.28–2.39 (m, 4H, 2CH2), 3.94 (q, J = 6.9 Hz, 4H, 2CH3CH2COO), 4.77 (s, 2H, 2CH), 5.05 (s, 4H, 2OCH2), 6.82 (d, J = 8.4 Hz, 4H, Ar–H), 7.04 (d, J = 8.2 Hz, 4H, Ar–H), 7.39 (d, J = 7.8 Hz, 2H, pyrdine-H), 7.80 (m, J = 7.9 Hz, 1H, pyrdine-H), 9.03 (s, 2H, 2NH) ppm; 13C NMR (126 MHz, DMSO-d6) δ 14.7, 18.8, 27.1, 29.6, 32.7, 35.6, 40.9, 50.9, 59.6, 70.7, 104.4, 110.7, 114.5, 121.1, 129.1, 138.3, 141.1, 145.3, 149.9, 156.8, 157.1, 167.5, 194.9 ppm; MS (EI, 70 eV): m/z (%) 813 [M+]; anal. calcd for C49H55N3O8: C, 72.30; H, 6.81; N, 5.16. Found: C, 72.26; H, 6.78; N, 5.08%.
O ketone) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.82 (s, 6H, 2CH3), 0.97 (s, 6H, 2CH3), 1.09 (t, J = 7.1 Hz, 6H, 2CH3CH2COO), 1.93 (d, 2H, H8, J = 15 Hz) 2.11 (d, 2H, H8, J = 15 Hz), 2.25 (s, 6H, 2CH3), 2.28–2.39 (m, 4H, 2CH2), 3.94 (q, J = 6.9 Hz, 4H, 2CH3CH2COO), 4.77 (s, 2H, 2CH), 5.05 (s, 4H, 2OCH2), 6.82 (d, J = 8.4 Hz, 4H, Ar–H), 7.04 (d, J = 8.2 Hz, 4H, Ar–H), 7.39 (d, J = 7.8 Hz, 2H, pyrdine-H), 7.80 (m, J = 7.9 Hz, 1H, pyrdine-H), 9.03 (s, 2H, 2NH) ppm; 13C NMR (126 MHz, DMSO-d6) δ 14.7, 18.8, 27.1, 29.6, 32.7, 35.6, 40.9, 50.9, 59.6, 70.7, 104.4, 110.7, 114.5, 121.1, 129.1, 138.3, 141.1, 145.3, 149.9, 156.8, 157.1, 167.5, 194.9 ppm; MS (EI, 70 eV): m/z (%) 813 [M+]; anal. calcd for C49H55N3O8: C, 72.30; H, 6.81; N, 5.16. Found: C, 72.26; H, 6.78; N, 5.08%.
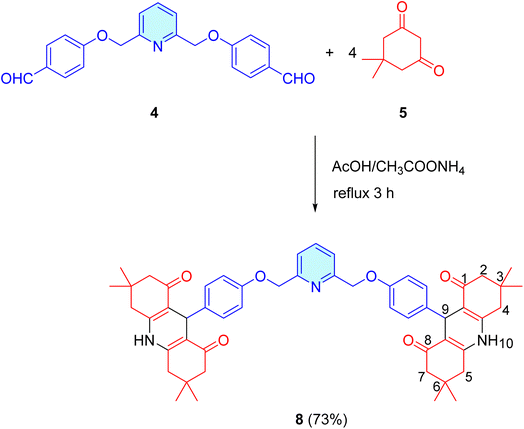
2.1.1.2 9,9′-(((Pyridine-2,6-diylbis(methylene))bis(oxy))bis(4,1-phenylene))bis(3,3,6,6-tetramethyl-3,4,6,7,9,10-hexahydroacridine-1,8(2H,5H)-dione) (8). A mixture of bis-aldehyde 4 (0.35 g, 1 mmol) and dimedone (5) (0.56 g, 4 mmol) was heated in glacial acetic acid (20 mL) containing ammonium acetate (0.15 g, 2 mmol) for 6 h. The excess solvent evaporated under reduced pressure, and the crude product was purified by washing with hot EtOH.
Creamy powder (0.73 g, 88%); Mp 285–287 °C; IR (KBr): ν 3289 (NH), 1636 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.84 (s, 12H, 4CH3), 0.97 (s, 12H, 4CH3), 1.94–2.15 (m, 8H, 4CH2), 2.26–2.44 (m, 8H, 4CH2), 4.73 (s, 2H, 2CH), 5.04 (s, 4H, 2OCH2), 6.79 (d, J = 8.2 Hz, 4H, Ar–H), 7.04 (d, J = 8.2 Hz, 4H, Ar–H), 7.39 (d, J = 7.8 Hz, 2H, pyrdine-H), 7.80 (t, J = 7.8 Hz, 1H, pyrdine-H), 9.23 (s, 2H, 2NH).ppm; 13C NMR (126 MHz, DMSO-d6) δ 27.1, 29.6, 32.5, 32.7, 40.7, 50.9, 70.7, 112.2, 114.4, 121.1, 129.2, 138.3, 140.6, 149.7, 156.6, 157.1, 195.0 ppm; MS (EI, 70 eV): m/z (%) 833 [M+]; anal. calcd for C53H59N3O6: C, 76.32; H, 7.13; N, 5.04. Found: C, 76.24; H, 7.09; N, 5.01%.
O) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.84 (s, 12H, 4CH3), 0.97 (s, 12H, 4CH3), 1.94–2.15 (m, 8H, 4CH2), 2.26–2.44 (m, 8H, 4CH2), 4.73 (s, 2H, 2CH), 5.04 (s, 4H, 2OCH2), 6.79 (d, J = 8.2 Hz, 4H, Ar–H), 7.04 (d, J = 8.2 Hz, 4H, Ar–H), 7.39 (d, J = 7.8 Hz, 2H, pyrdine-H), 7.80 (t, J = 7.8 Hz, 1H, pyrdine-H), 9.23 (s, 2H, 2NH).ppm; 13C NMR (126 MHz, DMSO-d6) δ 27.1, 29.6, 32.5, 32.7, 40.7, 50.9, 70.7, 112.2, 114.4, 121.1, 129.2, 138.3, 140.6, 149.7, 156.6, 157.1, 195.0 ppm; MS (EI, 70 eV): m/z (%) 833 [M+]; anal. calcd for C53H59N3O6: C, 76.32; H, 7.13; N, 5.04. Found: C, 76.24; H, 7.09; N, 5.01%.
2.1.1.3 9,9′-(((Pyridine-2,6-diylbis(methylene))bis(oxy))bis(4,1-phenylene))bis(3,3,6,6-tetramethyl-10-(p-tolyl)-3,4,6,7,9,10-hexahydroacridine-1,8(2H,5H)-dione) (11). Method A. A mixture of bis-aldehyde 4 (0.35 g, 1 mmol), dimedone (5) (0.28 g, 2 mmol), and enaminone 10 (0.46 g, 2 mmol) was heated in pyridine (10 mL) for 6 h. The precipitated solid was collected and washed thoroughly with hot EtOH.
Method B. A mixture of bis-aldehyde 4 (0.35 g, 1 mmol), dimedone (5) (0.56 g, 4 mmol), p-toluidine (12) (0.21 g, 2 mmol) was heated in pyridine (10 mL) for 6 h. The precipitated solid was collected and washed thoroughly with hot EtOH.
Yellow crystals (0.75 g, 74%, method A; 0.72 g, 71%, method B); Mp 300 < °C; IR (KBr): ν 1636 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.68 (s, 12H, 4CH3), 0.84 (s, 12H, 4CH3), 1.72–1.98 (m, 8H, 4CH2), 2.12–2.17 (m, 8H, 4CH2), 2.38 (s, 6H, 2CH3), 4.96 (s, 2H, 2CH), 5.08 (s, 4H, 2OCH2), 6.88 (d, J = 8.4 Hz, 4H, Ar–H), 7.19 (d, J = 8.2 Hz, 8H, Ar–H), 7.37 (d, J = 8.0 Hz, 4H, Ar–H), 7.41 (d, J = 7.7 Hz, 2H, pyrdine-H), 7.82 (t, J = 7.8 Hz, 1H, pyrdine-H).ppm; MS (EI, 70 eV): m/z (%) 1013 [M+]; anal. calcd for C67H71N3O6: C, 79.34; H, 7.06; N, 4.14. Found: C, 79.28; H, 7.03; N, 4.18%.
O) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.68 (s, 12H, 4CH3), 0.84 (s, 12H, 4CH3), 1.72–1.98 (m, 8H, 4CH2), 2.12–2.17 (m, 8H, 4CH2), 2.38 (s, 6H, 2CH3), 4.96 (s, 2H, 2CH), 5.08 (s, 4H, 2OCH2), 6.88 (d, J = 8.4 Hz, 4H, Ar–H), 7.19 (d, J = 8.2 Hz, 8H, Ar–H), 7.37 (d, J = 8.0 Hz, 4H, Ar–H), 7.41 (d, J = 7.7 Hz, 2H, pyrdine-H), 7.82 (t, J = 7.8 Hz, 1H, pyrdine-H).ppm; MS (EI, 70 eV): m/z (%) 1013 [M+]; anal. calcd for C67H71N3O6: C, 79.34; H, 7.06; N, 4.14. Found: C, 79.28; H, 7.03; N, 4.18%.
2.1.2.1 5,5′-(((Pyridine-2,6-diylbis(methylene))bis(oxy))bis(4,1-phenylene))bis(8,8-dimethyl-5,8,9,10-tetrahydropyrimido[4,5-b]quinoline-2,4,6(1H,3H,7H)-trione) (15). Creamy powder (0.62 g, 77%); Mp 300 < °C; IR (KBr): ν 3408, 3269, 3192 (3NH), 1717 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), 1665, 1655 (2C
O ketone), 1665, 1655 (2C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.86 (s, 6H, 2CH3), 0.98 (s, 6H, 2CH3), 1.97–2.17 (m, 4H, 2CH2), 2.40–2.45 (m, 4H, 2CH2), 4.67 (s, 2H, 2CH), 5.06 (s, 4H, 2OCH2), 6.82 (d, J = 8.2 Hz, 4H, Ar–H), 7.07 (d, J = 8.2 Hz, 4H, Ar–H), 7.39 (d, J = 7.8 Hz, 2H, pyrdine-H), 7.81 (t, J = 7.8 Hz, 1H, pyrdine-H), 8.81 (s, 2H, 2NH), 10.26 (s, 2H, 2NH), 10.65 (s, 2H, 2NH).ppm; 13C NMR (126 MHz, DMSO-d6) δ 27.1, 29.5, 32.7, 32.7, 39.5, 39.7, 39.9, 40.0, 40.2, 40.4, 40.5, 40.8, 50.7, 70.7, 90.4, 100.0, 112.0, 114.5, 121.0, 129.1, 138.4, 139.9, 149.5, 156.9, 157.1, 163.3, 163.3, 194.9 ppm; MS (EI, 70 eV): m/z (%) 809 [M+]; anal. calcd for C45H43N7O8: C, 66.74; H, 5.35; N, 12.11. Found: C, 66.71; H, 5.27; N, 12.05%.
O amide) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 0.86 (s, 6H, 2CH3), 0.98 (s, 6H, 2CH3), 1.97–2.17 (m, 4H, 2CH2), 2.40–2.45 (m, 4H, 2CH2), 4.67 (s, 2H, 2CH), 5.06 (s, 4H, 2OCH2), 6.82 (d, J = 8.2 Hz, 4H, Ar–H), 7.07 (d, J = 8.2 Hz, 4H, Ar–H), 7.39 (d, J = 7.8 Hz, 2H, pyrdine-H), 7.81 (t, J = 7.8 Hz, 1H, pyrdine-H), 8.81 (s, 2H, 2NH), 10.26 (s, 2H, 2NH), 10.65 (s, 2H, 2NH).ppm; 13C NMR (126 MHz, DMSO-d6) δ 27.1, 29.5, 32.7, 32.7, 39.5, 39.7, 39.9, 40.0, 40.2, 40.4, 40.5, 40.8, 50.7, 70.7, 90.4, 100.0, 112.0, 114.5, 121.0, 129.1, 138.4, 139.9, 149.5, 156.9, 157.1, 163.3, 163.3, 194.9 ppm; MS (EI, 70 eV): m/z (%) 809 [M+]; anal. calcd for C45H43N7O8: C, 66.74; H, 5.35; N, 12.11. Found: C, 66.71; H, 5.27; N, 12.05%.
2.1.2.2 5,5′-(((Pyridine-2,6-diylbis(methylene))bis(oxy))bis(4,1-phenylene))bis(8,8-dimethyl-2-thioxo-2,3,5,8,9,10-hexahydropyrimido[4,5-b]quinoline-4,6(1H,7H)-dione) (16). Pale yellow powder (0.72 g, 86%); Mp 300 < °C; IR (KBr): ν 3412, 3263, 3200 (3NH), 1670 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), 1653 (C
O ketone), 1653 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide) cm−1;1H NMR (500 MHz, DMSO-d6) δ 0.87 (s, 6H, 2CH3), 0.99 (s, 6H, 2CH3), 1.98–2.20 (m, 4H, 2CH2), 2.34–2.47 (m, 4H, 2CH2), 4.68 (s, 2H, 2CH), 5.07 (s, 4H, 2OCH2), 6.84 (d, J = 8.4 Hz, 4H, Ar–H), 7.09 (d, J = 8.1 Hz, 4H, Ar–H), 7.39 (d, J = 7.6 Hz, 2H, pyrdine-H), 7.81 (t, J = 7.8 Hz, 1H, pyrdine-H), 8.43 (s, 2H, 2NH), 11.64 (s, 2H, 2NH), 12.14 (s, 2H, 2NH).ppm; 13C NMR (126 MHz, DMSO-d6) δ 27.1, 29.4, 32.4, 32.7, 40.7, 50.7, 70.7, 94.9, 100.1, 111.8, 114.6, 121.0, 129.2, 138.4, 139.1, 143.9, 149.0, 157.1, 160.7, 173.9, 194.9 ppm; MS (EI, 70 eV): m/z (%) 841 [M+]; anal. calcd for C45H43N7O6S2: C, 64.19; H, 5.15; N, 11.64. Found: C, 64.12 H, 5.10; N, 11.60%.
O amide) cm−1;1H NMR (500 MHz, DMSO-d6) δ 0.87 (s, 6H, 2CH3), 0.99 (s, 6H, 2CH3), 1.98–2.20 (m, 4H, 2CH2), 2.34–2.47 (m, 4H, 2CH2), 4.68 (s, 2H, 2CH), 5.07 (s, 4H, 2OCH2), 6.84 (d, J = 8.4 Hz, 4H, Ar–H), 7.09 (d, J = 8.1 Hz, 4H, Ar–H), 7.39 (d, J = 7.6 Hz, 2H, pyrdine-H), 7.81 (t, J = 7.8 Hz, 1H, pyrdine-H), 8.43 (s, 2H, 2NH), 11.64 (s, 2H, 2NH), 12.14 (s, 2H, 2NH).ppm; 13C NMR (126 MHz, DMSO-d6) δ 27.1, 29.4, 32.4, 32.7, 40.7, 50.7, 70.7, 94.9, 100.1, 111.8, 114.6, 121.0, 129.2, 138.4, 139.1, 143.9, 149.0, 157.1, 160.7, 173.9, 194.9 ppm; MS (EI, 70 eV): m/z (%) 841 [M+]; anal. calcd for C45H43N7O6S2: C, 64.19; H, 5.15; N, 11.64. Found: C, 64.12 H, 5.10; N, 11.60%.
2.1.2.3 2,6-Bis((4-(3,5-dimethyl-1,4,7,8-tetrahydrodipyrazolo[3,4-b:4′,3′-e]pyridin-4-yl)phenoxy)methyl) pyridine (18). A mixture of bis-aldehyde 4 (0.35 g, 1 mmol), dimedone (5) (0.28 g, 2 mmol), 3-methyl-1H-pyrazol-5(4H)-one (17) (0.20 g, 2 mmol) was heated in absolute ethanol (30 mL) containing ammonium acetate (0.15 g, 2 mmol) for 6 h. The collected precipitate was isolated and washed with hot EtOH.
Orange crystals (0.61 g, 92%); Mp 300 < °C; IR (KBr): ν 3173, 3129 (NH) cm−1; 1H NMR (500 MHz, DMSO-d6) δ 2.05 (s, 12H, 4CH3), 4.74 (s, 2H, 2CH), 5.18 (s, 4H, 2OCH2), 6.85 (d, J = 8.8 Hz, 4H, Ar–H), 7.01 (d, J = 8.5 Hz, 4H, Ar–H), 7.40 (d, J = 7.7 Hz, 2H, pyrdine-H), 7.81 (t, J = 7.8 Hz, 1H, pyrdine-H), 10.84 (br, 6H, 6NH).ppm; MS (EI, 70 eV): m/z (%) 665 [M+]; anal. calcd for C37H35N11O2: C, 66.75; H, 5.30; N, 23.14. Found: C, 66.70; H, 5.21; N, 23.12%.
2.2 In vitro cytotoxicity assay
In the MTT assay47,48 synthetic compounds (16, 15, 8, 7, 11, and 18) were tested for their toxicity on HepG2 (liver cancer), A549 (lung cancer), MCF7 (breast cancer), and HFB4 (normal skin) cell lines. The cell lines used in this study were purchased from the Production Company of Vaccines, Sera, and Drugs (VACSERA), Egypt. Cell adhesion was accomplished by cultivating cells at 5 × 103 per well in 96-well plates for 24 hours. All test substances were administered to cells at varying concentrations (0–100 µg mL−1) for 48 hours. Add 100 µL of MTT solution (0.5 mg mL−1) to each well and incubate for 4 hours at 37 °C. After dissolving the formazan crystals in 100 µL of DMSO, measure the absorbance at 570 nm using a microplate reader. Three experiments on dose–response curves were used to establish the IC50 values, which indicate the concentration required to reduce cell viability by 50%. The selectivity index (SI) was established by calculating the ratio of IC50 values between normal cells (HFB4) and cancer cells (HepG2, A549, or MCF7).The study on the structure–activity relationship (SAR) was conducted to understand how changes in structure within the synthesized chemical series affect their anticancer properties. The examination considered molecular structures, heteroatom modifications, scaffold flexibility and rigidity, as well as the electronic characteristics of the substances. Crucial elements comprised the existence of fused polycyclic structures, hydrogen-bond donor/acceptor ability, characteristics of heteroatoms (such as nitrogen and sulfur), and the equilibrium between hydrophilic and lipophilic areas.49,50 These characteristics were methodically linked to cellular absorption, target-binding capacity, and selectivity. Significant focus was directed toward how scaffold rigidity, the placement of functional groups, and steric effects influence bioactivity and pharmacokinetic properties. This method offered insights into how changes in structure influenced variations in cytotoxic effectiveness and selectivity, providing useful direction for the logical design of enhanced anticancer compounds in upcoming research.
2.3 Molecular docking
In this research, bis-polyhydroquinoline 7 was docked23,51–53 to analyze its binding modes and affinities for five cancer-related protein targets: FGFR4, Aurora A kinase, CDK1, EGFR, and VEGFR2. They were selected due to their essential roles in cancer progression, particularly in hepatocellular carcinoma (HepG2), lung carcinoma (A549), and breast cancer (MCF7) cells, where compound 7 exhibited significant cytotoxicity. Hepatocellular carcinoma is associated with FGFR4 and Aurora A, while the proliferation of lung cancer cells and angiogenesis rely on EGFR and VEGFR2. CDK1 was incorporated due to its regulation of the cell cycle in various tumor types.The Protein Data Bank (PDB) provided the three-dimensional X-ray structures of the selected target. We used the Molecular Operating Environment (MOE 2015.01) program to make the protein.54,55 For protein preparation, each protein structure was examined meticulously to identify any missing residues. Next, any portions that weren't needed were removed, like water molecules, ions, and heteroatoms. Hydrogen atoms were introduced, and the protonation states were adjusted according to the body's pH. For ligand preparation, MOE's Builder module was utilized to construct the 2D structure of compound 7, and then it was changed into a 3D conformer. The MMFF94x force field was employed to minimize the structure's energy and optimize its conformation to the lowest-energy state, ensuring the proper shape and protonation before docking.
The MOE Dock module was utilized to execute docking simulations. The coordinates of the co-crystallized ligand were used to determine the location of the binding site. MOE's site finder (dummy atoms) was employed to find the active pocket. The search region (grid sphere) was altered so that it now includes all the relevant residues that are within roughly 10 Å of the co-crystallized ligand or dummy atom centroid. This made sure that the pocket was fully closed.
The triangle matcher placement method was used for the docking process, with an initial generation of 100 poses ranked by the London dG scoring function. The London dG scoring technique was used to rank these positions, and then the top 50 poses from each simulation were selected in rigid receptor mode for further refinement. After that, the GBVI/WSA dG scoring tool was used to assign a new score to the new poses. This was to find out the free energy of binding.
The best binding pose for each target was picked based on three things: (1) the lowest GBVI/WSA dG score, which shows the best binding affinity; (2) the presence of important non-covalent interactions (hydrogen bonds, hydrophobic contacts, π–π stacking, and salt bridges) with critical active-site residues; and (3) pharmacophoric alignment with the co-crystallized inhibitor or a known active ligand of the target protein. Finally, a visual check was performed to ensure that there were no steric conflicts and that compound 7 was correctly positioned in the active site and was chemically stable.
2.4 Molecular dynamics simulations
The highest consensus docking scores from compound 7 with the protein receptor Epidermal Growth Factor Receptor tyrosine kinase domain (EGFR TKD)56 were subjected to all-atom molecular dynamics (MD) simulations to investigate the relative stabilities of the protein–ligand interactions and to screen the compound for subsequent binding energy calculations. GROMACS V2023 package57–59 and the CHARMM36m60,61 force field were utilized for all simulations. The parameters and topological files for the compound we chose were generated using the latest CHARMM/CGenFF force field using CHARMM-GUI.62–65 The protein–ligand complex was placed in the middle of a box of solvated water molecules with a TIP3P explicit solvation model. To mimic the physiological salt concentrations, 0.154 M ions (98 Na+ and 99 Cl−) were added to provide charge neutralization and electrostatic screening, which extended the edge distance 20 Å from the protein. The rectangular cubic system for CHARMM and the periodic boundary conditions were set to 104.0 Å in the x, y, and z axis.The MD procedures included minimization, equilibration (in two steps), and production. The 200 ns MD production simulations did not limit any atoms. The preliminary energy of the solvated protein–ligand complex was minimized through 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps at a constant temperature of 300 K. This was followed by equilibration in two steps: the first was a canonical (NVT) ensemble with 4
000 steps at a constant temperature of 300 K. This was followed by equilibration in two steps: the first was a canonical (NVT) ensemble with 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps, and the second was an isothermal–isobaric (NPT) ensemble with 4
000 steps, and the second was an isothermal–isobaric (NPT) ensemble with 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps and a 1 fs time integration step. Finally, the production MD simulation step was performed for the isothermal–isobaric (NPT) ensemble with 100
000 steps and a 1 fs time integration step. Finally, the production MD simulation step was performed for the isothermal–isobaric (NPT) ensemble with 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 runs and a 2-fs time integration step for 200 ns. Using NVT and NPT ensembles, the pressure was set to 1 atm. Using the Langevin thermostat, the temperature was regulated to 300.0 K.66 Force-field settings that didn't include scaling of 1.2 nm were used to minimize and balance the complexes in the water box. All atoms, even those of hydrogen, were shown clearly. Trajectory data every 0.05 ns has been obtained. Tools from the GROMACS and VMD packages67 were used to look at the full system residues, utilizing the produced trajectories we got during the manufacturing stage. A distance cut-off of 1.0 nm was used for short-range nonbonded interactions with a pair list distance of 1.2 nm, while Lennard–Jones interactions were smoothly cut off at 1.2 nm (rvdm, rcoulomb = 1.2 nm). The particle-mesh Ewald (PME) approach68,69 was used to handle long-range electrostatic interactions. For the sake of consistency, the same method for all the MD simulations was utilized.
000 runs and a 2-fs time integration step for 200 ns. Using NVT and NPT ensembles, the pressure was set to 1 atm. Using the Langevin thermostat, the temperature was regulated to 300.0 K.66 Force-field settings that didn't include scaling of 1.2 nm were used to minimize and balance the complexes in the water box. All atoms, even those of hydrogen, were shown clearly. Trajectory data every 0.05 ns has been obtained. Tools from the GROMACS and VMD packages67 were used to look at the full system residues, utilizing the produced trajectories we got during the manufacturing stage. A distance cut-off of 1.0 nm was used for short-range nonbonded interactions with a pair list distance of 1.2 nm, while Lennard–Jones interactions were smoothly cut off at 1.2 nm (rvdm, rcoulomb = 1.2 nm). The particle-mesh Ewald (PME) approach68,69 was used to handle long-range electrostatic interactions. For the sake of consistency, the same method for all the MD simulations was utilized.
The binding free energy of the simulated complex was calculated using MD simulations by GROMACS, utilizing snapshots from the system's trajectories over 200 ns, encompassing 4000 trajectories. This was followed by the gmx_MMPBSA tool, based on the AMBER tools MMPBSA.py module with GROMACS files,70 where the ligand (L) binds to the protein receptor (R) to form the complex (RL). We are solely interested in computations of relative binding energy, which is provided by Gibbs relative binding energy:
| ΔGbind = ΔGbind,vaccum + ΔGRL,solvation − (ΔGR − ΔGL) |
2.5 EGFR inhibition assay
Following the manufacturer's instructions, the EGFR kinase inhibitor screening assay kit (Cat. no. 40321, BPS Bioscience, USA) was used to test how well compound 7 worked against EGFR. This colorimetric test measures the phosphorylation of a poly(Glu–Tyr) substrate by EGFR. An HRP-conjugated anti-phosphotyrosine antibody and TMB substrate were used to find it. The concentrations of compound 7 ranged from 7.81 µg mL−1 to 1000 µg mL−1, and sorafenib was used as a positive control. After 30 minutes of incubation at 30 °C with EGFR and ATP, the detection reagents were added sequentially. A microplate reader was utilized to assess absorbance at 450 nm. Percent inhibition was figured out, and IC50 values were evaluated for compound 7 and sorafenib.2.6 ADMET prediction
The pkCSM pharmacokinetics web interface71 evaluates absorption, distribution, metabolism, excretion, and toxicity factors by utilizing graph-based signatures to forecast compound 7's ADMET characteristics. The outcomes were automatically calculated following the input of compound 7's representation into the pkCSM web interface. Furthermore, we utilized in silico permeability data from the ADMETlab 3.0 platform,72 including PAMPA projections, to get a better idea of how well compound 7 may be absorbed.73 To aid research on compound 7's anticancer capabilities, we evaluated its drug-likeness and forecasted its pharmacokinetic and safety profiles in silico to enhance experimental findings and bolster its prospects as an anticancer candidate.2.7 Statistical analysis
Each experiment was conducted in triplicate (n = 3), and results were displayed as mean ± standard deviation (SD). We used GraphPad Prism 9 software for statistical analysis and fitted dose–response curves to get IC50.3 Results and discussion
3.1 Chemistry
To reach our objective, 2,6-bis(bromomethyl)pyridine 3 was chosen as an essential intermediate and can serve as a primary element for the creation of 4,4′-((pyridine-2,6-diylbis(methylene))bis(oxy))dibenzaldehyde 4. The interaction of compound 3 with the potassium salt 2 (obtained from the reaction of 4-hydroxybenzaldehyde 1 with ethanolic KOH) in boiling DMF led to the formation of compound 4 in 88% yield (Scheme 1).74,75Our primary emphasis was on synthesizing bis-polyhydroquinoline 7 through Hantzsch's four-component condensation reactions that include bis-aldehyde 4, dimedone 5, ethyl acetoacetate 6, and ammonium acetate (Scheme 2). The product was obtained with a yield of 84% after being heated in ethanol at reflux for 6 hours.
Spectral information was employed to establish the structure of compound 7. Consequently, the NH group was recognized at ν 3279 cm−1 according to its IR spectra. Additionally, it showed the existence of the carbonyl group at ν 1697 and 1643 cm−1. The 1H NMR spectrum of compound 7 exhibited characteristic singlets at δ 4.77 for H-4, δ 5.05 for OCH2 protons, and δ 9.03 for 2NH. It also showed singlets at 0.82, 0.97, and 2.25 ppm, each integrated to 6H assigned to the different methyl groups, as well as two multiplets for two CH2 groups at δ 1.92–2.16 and 2.27–2.41 ppm. The ethyl protons appeared as a triplet and a quartet at δ 1.09 and 3.94 ppm. Signals for aromatic protons appeared at the expected positions. Moreover, the 13C NMR examination of compound 7 validated its proposed structure, showing characteristic peaks at δ 167.5 and 194.9 ppm for ester and ketonic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, respectively.
O, respectively.
The reaction of bis(aldehyde) 4 with dimedone (5) and ammonium acetate in AcOH under reflux produced a 73% yield of bis(3,3,6,6-tetramethyl-3,4,6,7,9,10-hexahydroacridine-1,8(2H,5H)-dione) 8 (Scheme 3). The structural examination of the target compound was conducted utilizing spectral data.
The composition of product 8 was established through multiple spectral methods. The IR spectra of compound 15 showed NH groups at ν 3289 cm−1, and the carbonyl groups were observed at ν 1636 cm−1. The 1H NMR spectrum exhibited two singlets, totaling 24 protons at δ 0.84 and 0.97 ppm, which correspond to eight CH3 groups. The 16 protons constituting the four methylene groups (H2, H4, H5, H7) were detected as a pair of multiplets at δ 1.94–2.15 and 2.26–2.44 ppm. Furthermore, the singlet signals at δ 4.73 and 5.04 ppm correspond to the acridine-H9 and –OCH2, respectively. The NH group appeared as a singlet at δ 9.23 ppm. All other signals were as expected. The proton at the H-5 position resonates at δ = 4.65 ppm, which is a characteristic signal of the chromene moiety present in this class of compounds.76
Since acridines hold biological significance, we expanded our study to include the synthesis of bis(9-aryl-substituted acridine). Consequently, when bis-aldehyde 4 is reacted with two equivalents each of dimedone 5 and enaminone 10 under reflux conditions in pyridine, it leads to the generation of bis(3,3,6,6-tetramethyl-10-(p-tolyl)-3,4,6,7,9,10-hexahydroacridine-1,8(2H,5H)-dione) 11 in 74% yield (method A) (Scheme 4). The product was obtained with a limited quantity of the target substance when aldehyde 4 reacted with four moles of dimedone 5 and two moles of p-toluidine 12 (method B), as illustrated in Scheme 4. It should be noted that the formation of enamine 10 involved the interaction of dimedone 5 with arylamines 12 in the presence of trichloroacetic acid (Scheme 4).77
The spectral data helped to identify the structure of compound 11. Consequently, the IR spectrum revealed a carbonyl group at ν 1636 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1. The existence of 24 protons was validated in the 1H NMR spectrum of compound 11, displaying two singlets at δ 0.68 and 0.84 ppm that were attributed to eight CH3 groups. Additionally, the singlet peak at δ 4.96 ppm suggests the presence of acridine-H9. All other signals were found exactly where they should have been.
O) cm−1. The existence of 24 protons was validated in the 1H NMR spectrum of compound 11, displaying two singlets at δ 0.68 and 0.84 ppm that were attributed to eight CH3 groups. Additionally, the singlet peak at δ 4.96 ppm suggests the presence of acridine-H9. All other signals were found exactly where they should have been.
A reaction involving three components, namely dibenzaldehyde 4, two equivalents of dimedone 5, and either 6,6-aminopyrimidine-2,4-dione 13 or 6-amino-2-thioxo-2,3-dihydropyrimidin-4(1H)-one 14, produced bis(2,3,5,8,9,10-hexahydropyrimido[4,5-b]quinoline-4,6(1H,7H)diones) 15 and 16 with yields of 91% and 77%, respectively (Scheme 5).
The IR spectra of Compound 15 showed NH groups at ν 3408, 3269, and 3192 cm−1. Furthermore, the carbonyl groups were observed at ν 1717, 1665, and 1655 cm−1. The 1H NMR spectrum of compound 15 showed a singlet peak at 4.67 ppm, which was attributed to H5. The NH group displayed wide signals at δ 8.81, 10.26, and 10.65 ppm. Every other signal was in the proper location. The structure of compound 15 was further supported based on 2D-HMBC NMR spectroscopy that indicated 3J – cross peak correlation between H5 at δ = 4.65 and CO at δ = 195.82 (Fig. S18).
The study was expanded to include the synthesis of bis(3,5-dimethyl-1,4,7,8-tetrahydrodipyrazolo[3,4-b:4′,3′-e]pyridin-4-yl)alkanes 18 via a multicomponent reaction that utilized 3-methyl-1H-pyrazol-5(4H)-one (17), bis(aldehyde) 4, and ammonium acetate in heated ethanol (Scheme 6).
The IR spectrum for compound 18 displayed the NH groups at ν 3173 and 3129 cm−1. The singlet peak in the 1H NMR spectrum of compound 18 exhibited an integration of 12 protons at δ 2.05, which corresponds to four CH3 groups. Moreover, the signals at δ 4.74 and 5.18 ppm correspond to OCH2 and H4, respectively. The NH groups appeared as a broad singlet peak at δ 10.68. All the other signals were seen in their expected positions.
3.2 In vitro cytotoxic activity evaluation
HepG2 (liver cancer), A549 (pulmonary cancer), MCF7 (breast cancer) and normal human skin fibroblast (HFB4) cells were assessed for in vitro cytotoxic effects of synthetic compounds 16, 15, 8, 7, 11, and 18 through the MTT assay after 48 hours of exposure, as shown in Fig. 2. Tables 1 and S1 present pertinent IC50 values and selectivity indices (SI). Certain cell lines (HepG2, A549, and MCF7) were selected for both strategic and scientific purposes. The commonly utilized model of hepatocellular carcinoma, HepG2, represents a substantial worldwide cancer load characterized by treatment resistance and rapid proliferation.78 A549 cells, derived from human alveolar epithelial carcinoma, serve as a recognized model for non-small cell lung cancer (NSCLC), which is one of the leading causes of cancer-related deaths globally.79 MCF7 cells serve as models for hormone-sensitive breast cancer and are widely used to test anticancer compounds that affect the growth of tumor tissue in response to the hormone estrogen.80 These three cell lines offer a solid foundation for assessing the anticancer effects of new compounds sourced from various tissue origins and types of cancer. The evaluated compounds showed a dose-dependent decrease in cancer cell viability according to MTT assay results. Drawing from the bis-polyhydroquinoline framework, 7 exhibited the most significant cytotoxic impact among them. Against HepG2 cells, it exhibited IC50 values of 18.07 ± 1.17 µg mL−1, while against A549 cells and MCF7 cells, it was 14.45 ± 3.73 µg mL−1 and 30.89 ± 6.63, respectively. Notably, compound 7 demonstrated pronounced activity against MCF7 cells comparable to its effects on HepG2 and A549 cells, with exceptional selectivity indices (>5.53 for HepG2, >6.92 for A549, and >3.23 for MCF7), and IC50 values exceeding 100 µg mL−1 ensuring minimal toxicity to normal fibroblast cells (HFB4). This notable selectivity implies that 7 specifically focuses on cancerous cells rather than healthy ones, a highly sought-after trait in anticancer drugs.| Compound | IC50a (µg mL−1) | SIb | |||||
|---|---|---|---|---|---|---|---|
| HepG2 | A549 | MCF7 | HFB4 | HepG2 | A549 | MCF7 | |
| a The data were expressed as the mean ± SD of three independent experiments.b SI = IC50 on HFB4/IC50 on cancer cells. | |||||||
| 16 (PA2) | 39.43 ± 3.25 | >100 | 80.70 ± 10.50 | >100 | >2.53 | >1 | >1.24 |
| 15 (PA3) | 65.78 ± 1.87 | 82.54 ± 6.80 | >100 | >100 | >1.52 | >1.21 | >1 |
| 8 (PA5) | 59.14 ± 8.53 | 44.38 ± 17.84 | >100 | 68.24 ± 4.73 | 1.15 | >1.53 | 0.68 |
| 7 (PA6) | 18.07 ± 1.17 | 14.45 ± 3.73 | 30.89 ± 6.63 | >100 | >5.53 | >6.92 | >3.23 |
| 11 (PA8) | 73.81 ± 14.90 | >100 | >100 | >100 | >1.35 | >1 | >1 |
| 18 (PA11) | 55.82 ± 7.43 | 95.87 ± 3.61 | >100 | >100 | >1.79 | >1.04 | >1 |
| 5-Florouracil | 45.45 ± 5.97 | 48.00 ± 4.32 | 51.04 ± 11.48 | 88.33 ± 2.53 | 1.94 | 1.84 | 1.73 |
In comparison, 16, a sulfur-including bis(hexahydropyrimido[4,5-b]quinolinedione), showed minimal effectiveness on A549 cells (IC50 > 100 µg mL−1), but exhibited moderate cytotoxicity against HepG2 and MCF7 cells (IC50 = 39.43 ± 3.25 µg mL−1 and 80.70 ± 10.50 µg mL−1, respectively). The selectivity index of 16 against HepG2 and MCF7 (>2.53 and 1.24, respectively) highlighted its safer profile in contrast to nonselective cytotoxic drugs. Compound 15, the nitrogen counterpart of 16, demonstrated a similarly diminished effectiveness relative to 16, presenting IC50 values of 65.78 ± 1.87 µg mL−1 for HepG2 and 82.54 ± 6.80 µg mL−1 for A549, and lacked cytotoxic activity against MCF7 cells (IC50 > 100 µg mL−1). Although 15's selectivity indices (>1.52 and >1.21) are satisfactory, replacing sulfur with nitrogen reduced cytotoxic potency, suggesting that sulfur substitution enhances biological activity.81 The compound 8—a bis(hexahydroacridine-1,8(2H,5H)-dione)—demonstrated a slight cytotoxic effect, with IC50 values of 59.14 ± 8.53 µg mL−1 for HepG2 cells and 44.38 ± 17.84 µg mL−1 for A549 cells, and displayed no effective cytotoxicity against MCF7 cells (IC50 > 100 µg mL−1).82 However, the slightly lower selectivity indices (1.15, >1.53, and 0.68, respectively) suggested less cancer specificity when compared to 7. With an IC50 of 73.81 ± 14.90 µg mL−1 for HepG2 and an IC50 > 100 µg mL−1 for A549 and MCF7, 11, featuring bis(9-aryl-substituted acridine) components, exhibited reduced overall activity, suggesting that increased arylation and greater planarity may adversely influence cytotoxic efficacy, possibly due to reduced cellular absorption or impaired binding affinity to the targeted biomolecules.83
The results show that compound 7 has considerable potential to serve as a treatment, considering that it was very detrimental to A549, HepG2, and MCF7 cells, with IC50 values much lower than those of other similar compounds. It also had a high selectivity index against normal fibroblasts, meaning it was only harmful to cancer cells.
3.3 Structure–activity relationship (SAR) analysis
Key structural components influencing anticancer efficacy were identified through a detailed analysis of the structure–activity relationship (SAR) of the studied compounds. Upon examining the structures of the synthesized compounds, it is possible to conclude that there is a pyridine core linked to two fused-heterocyclic units via phenoxymethyl linkers. The heterocyclic systems include hexahydroquinoline, hexahydroacridine, tetrahydropyrimido[4,5-b]quinoline, and tetrahydrodipyrazolo[3,4-b:4′,3′-e]pyridine. The design approach is illustrated in Scheme 7. The bis-polyhydroquinoline scaffold emerged as the most appropriate framework, as suggested by 7. Target binding seems to be enhanced through a fused, semi-rigid tetracyclic design that boosts molecular planarity while maintaining a perfect equilibrium between flexibility and rigidity.84 Moreover, the numerous hydrogen bond donors and acceptors present in the polyhydroquinoline framework are likely to facilitate positive interactions with cellular proteins involved in managing oxidative stress, such as kinases or enzymes linked to cancer development.85,86 Incorporating sulfur atoms, as seen in 16, appears to enhance cytotoxic efficacy compared to 15, a nitrogen analogue. Sulfur atom regulation of electrical properties, enhancement of lipophilicity, and facilitation of membrane permeability, all of which may aid in boosting intracellular accumulation and biological effectiveness.81,86 Conversely, the limited activity of 16 in comparison to 7 suggests a synergistic advantage from the polyhydroquinoline core that goes beyond mere heteroatom influences.87 According to 8 and 11, the derivatives of bis(hexahydroacridine) and bis(aryl-acridine) showed reduced efficacy and selectivity. Either excessive planarity or large aryl groups could lead to suboptimal interactions with biological targets, which may account for the steric hindrance caused by both.88 Additionally, these structures may possess changed pharmacokinetic characteristics and diminished solubility, features considered mainly significant for cytotoxic efficacy. Moreover, 18 demonstrated moderate activity with a pyrazolo-pyridine structure, indicating that nitrogen-dense fused heterocycles hold potential but might need further refinement to achieve the same potency and selectivity as 7.87,89 The inadequate performance of 18 further emphasizes the importance of balancing hydrophilic and hydrophobic elements within the molecule for maximal anticancer effect. The connection with 5-FU was especially beneficial. While 5-FU is frequently employed in therapy, it showed lower selectivity compared to 7, highlighting the need for the creation of next-generation cancer treatments that merge strong cytotoxic effects with minimized off-target effects. In addition, the superior selectivity of 7 compared to 5-FU suggests a distinct or more targeted mechanism of action, necessitating further in-depth mechanistic research. Overall, the SAR analysis reveals that the type and placement of substituents on the bis-heterocyclic structure significantly affects the cytotoxic capability of the compounds. The existence of a central pyridine linker, along with electron-donating groups, seems to boost cellular absorption and enhance interactions with intracellular targets, probably due to better molecular planarity and π–π stacking ability. Moreover, the bis-polyhydroquinoline structure might enhance lipophilicity, promoting membrane penetration and accumulation within cells. These structural characteristics work together to explain the measured cytotoxic effects. Among the synthesized derivatives, compound 7 exhibited strong activity and significant selectivity, underscoring its potential as a key framework for the strategic development of more effective anticancer therapies.3.4 Molecular docking of 7
The results of the MTT assay indicated that 7 was safe for normal HFB4 cells but exhibited significant cytotoxic effects on HepG2 and A549 cancer cells. These findings indicate that 7 could exhibit selective anticancer effects through specific molecular interactions with key oncogenic targets. Through molecular docking90 of bis-polyhydroquinoline 7 with cancer-related targets, a detailed understanding of its binding affinities, interaction characteristics, and potential mechanisms explaining its cytotoxic effects on HepG2, A549, and MCF7 cancer cells was achieved, thereby investigating the molecular processes behind these experimental findings. The choice of the type of protein receptors was determined by how important they were in key signaling pathways linked to hepatocellular carcinoma (HepG2), non-small cell lung cancer (A549), and breast cancer (MCF7). FGFR4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 91 and VEGFR2
91 and VEGFR2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 92 were chosen because they are essential for the growth of hepatocellular carcinoma and the formation of new blood vessels, respectively. EGFR93 and Aurora A kinase94 are often present in high amounts in A549 and MCF7 cells, and they are thought to be important for tumor development and cell cycle progression. CDK1
92 were chosen because they are essential for the growth of hepatocellular carcinoma and the formation of new blood vessels, respectively. EGFR93 and Aurora A kinase94 are often present in high amounts in A549 and MCF7 cells, and they are thought to be important for tumor development and cell cycle progression. CDK1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95 was added because it controls mitosis in all cells, and cancer cells that divide quickly have higher levels, including breast, lung, and liver malignancies. This reasoning guarantees that the docking targets are pathologically and therapeutically relevant to the cancer models used in vitro.
95 was added because it controls mitosis in all cells, and cancer cells that divide quickly have higher levels, including breast, lung, and liver malignancies. This reasoning guarantees that the docking targets are pathologically and therapeutically relevant to the cancer models used in vitro.
The initial validation of docking data was achieved by self-docking of the reference co-crystallized ligand (4-anilinoquinazoline inhibitor, erlotinib) with its respective receptor (EGFR; PDB ID: 1M17). The RMSD value was determined to be under 2 Å, and an impressive superimposition of the re-docked (cyan) and co-crystallized ligand (green) inside the active site was shown, as shown in Fig. 3.
FGFR4 is known to promote cell proliferation, survival, and metastases in hepatocellular carcinoma (HepG2) by activating the MAPK and PI3K/AKT pathways.96 A significant link between 7's strong interaction characteristics within the active area of FGFR4 and the noted reduction in HepG2 cell survival during cytotoxicity assessments indicates that the drug could effectively inhibit FGFR4-mediated signaling. This aligns with reports showing that FGFR4-selective inhibitors reduced HepG2 proliferation in both in vitro and in vivo circumstances.97 The comparable docking score and interaction profile of 7 imply that it may work in the same way as a selective FGFR4 antagonist.
The formation of the mitotic spindle, alignment of chromosomes, and maturation of centrosomes are all largely reliant on Aurora A kinase. The overexpression of Aurora A results in genomic instability and cancer.98,99 The documented interaction of 7 with key residues in the Aurora active site suggests that 7 could lessen the activity of the kinase, consequently promoting mitotic arrest and cell death, outcomes frequently observed in cancer cells during treatment. This clarifies a mechanism through which 7 induces cytotoxicity and modifies cell cycle progression, particularly during the G2/M phase. Similar binding characteristics and inhibitory capabilities have been shown for established Aurora kinase inhibitors, including MLN8237 (alisertib), that exhibited analogous hydrogen bonding with THR residues in preclinical studies of lung and liver malignancies.100,101 This confirms that 7's binding mode is properly configured.
CDK1, often elevated in different tumors, plays a crucial role in regulating the G2/M transition during the cell cycle.102 The identified binding mechanism indicates that 7 attaches directly within the ATP-binding pocket of CDK1, potentially disrupting its kinase function. Even though the docking score was slightly less than that of FGFR4 and EGFR, the nature and abundance of the interactions suggest that 7 could effectively lower CDK1 activity, thereby aiding in the observed inhibition of proliferation and triggering of cell cycle arrest in both HepG2 and A549 cells. Interestingly, the interaction pattern is quite like that described for roscovitine (Seliciclib), which is a well-known CDK1 inhibitor. This suggests that 7 may have a similar inhibitory profile.103
 | ||
| Fig. 4 Docked compound 7 interacts with EGFR (PDB ID: 1M17) in 2D (left panel) and 3D (right panel). Carbon atoms are designated turquoise, nitrogen atoms blue, and oxygen atoms red. | ||
Particularly in non-small cell lung cancer (NSCLC), EGFR plays a vital role in cancer progression by enhancing cell survival, proliferation, and angiogenesis through the activation of downstream pathways such as RAS/RAF/MEK/ERK and PI3K/AKT.104,105 Considering 7's strong binding affinity to EGFR, it appears that it could serve as an effective EGFR inhibitor, thus inhibiting these proliferative signals. For A549 cells, which primarily depend on EGFR signaling, this holds significant importance. Consequently, the strong molecular rationale for the significant antiproliferative effect of 7, shown experimentally against A549 lung cancer cells, is derived from the docking data. The docking pattern resembles that of erlotinib, which is an EGFR inhibitor that has been authorized for use in humans and likewise engages with LYS721 and fills the ATP-binding cleft in a comparable manner.106 This enhances the molecular understanding of 7's antiproliferative activity via EGFR suppression.
| Protein | PDB ID | Ligand | Receptor | Interaction | Distance | E (kcal mol−1) | S (energy score) |
|---|---|---|---|---|---|---|---|
| FGFR4 | 4XCU | N 44 | OD2 ASP 630 | H-donor | 3.15 | −4.7 | −8.18 |
| O 71 | NH2 ARG 483 | H-acceptor | 2.88 | −2.1 | |||
| O 107 | NZ LYS 503 | H-acceptor | 3.21 | −2.5 | |||
| 6-Ring | NH2 ARG 483 | Pi-cation | 3.86 | −0.5 | |||
| 6-Ring | ND2 ASN 557 | Pi-H | 4.71 | −0.8 | |||
| Aurora A | 3H0Z | O 70 | N THR 292 | H-acceptor | 2.93 | −13.6 | −8.04 |
| O 107 | NZ LYS 143 | H-acceptor | 2.90 | −7.2 | |||
| 6-Ring | CG2 THR 292 | Pi-H | 4.16 | −0.5 | |||
| CDK1 | 6GU6 | C 34 | OD2 ASP 86 | H-donor | 3.10 | −0.6 | −7.51 |
| O 70 | N GLU 12 | H-acceptor | 3.23 | −2.2 | |||
| O 97 | N LEU 83 | H-acceptor | 3.11 | −1.6 | |||
| C 99 | 6-Ring PHE 80 | H-pi | 4.27 | −0.6 | |||
| EGFR | 1M17 | O 97 | NZ LYS 721 | H-acceptor | 2.84 | −12.1 | −8.98 |
| VEGFR2 | 3VHE | N 4 | SG CYS 1045 | H-donor | 4.28 | −0.7 | −7.60 |
| C 13 | O ASP 1046 | H-donor | 3.26 | −0.5 | |||
| O 70 | N ASN 923 | H-acceptor | 3.20 | −1.0 | |||
| 6-Ring | CG GLU 885 | Pi-H | 3.73 | −0.6 | |||
| 6-Ring | CG2 VAL 899 | Pi-H | 4.19 | −0.6 |
Tumor angiogenesis, which involves the formation of new blood vessels that provide nutrients and oxygen to growing tumors, relies on VEGFR2. Blocking VEGFR2 leads to restricted growth and dissemination capability by starving tumors of essential nutrients, along with decreased angiogenesis107,108 The identified interaction between 7 and VEGFR2 suggests that it could possess antiangiogenic characteristics, thereby aiding not just in the decrease of primary tumor growth but also in the avoidance of metastasis, which is vital for aggressive cancers like hepatocellular carcinoma and lung adenocarcinoma. This aligns with interaction investigations of established VEGFR2 medications like sorafenib, which additionally associates with ASN923 and CYS1045, supporting 7's expected antiangiogenic effectiveness.109
3.5 Correlation with experimental cytotoxicity
Through hydrogen bonding, π-cation interactions, and hydrophobic contacts, the comprehensive docking analysis indicates that 7 could engage with various significant oncogenic targets. Importantly, the key binding interactions, particularly with EGFR and FGFR4, directly correspond to the established reliance of A549 and HepG2 cells on EGFR and FGFR4 signaling pathways, respectively. This clarifies 7's noticeable cytotoxicity in experimental trials that involve the concurrent inhibition of several pathways essential for the survival, growth, and angiogenesis of cancer cells.Compared to single-target medications, which frequently develop into challenges, compound 7 can block several important targets at once, like multitargeted tyrosine kinase inhibitors (TKIs) such as sunitinib and vandetanib. This makes it more likely to lower resistance to therapeutics and make treatments last longer.110
Moreover, 7's ability to bind to multiple targets may assist in reducing the common problem associated with single-target therapies, as well as the development of drug resistance.111,112 Simultaneously targeting cell cycle regulation (CDK1, Aurora A), receptor tyrosine kinase pathways (EGFR, FGFR4), and angiogenesis (VEGFR2) will enhance 7's overall and enduring anticancer effectiveness.
Consequently, the outcomes of molecular docking not only reinforce the experimental findings but also provide a compelling mechanistic basis for further 7 advancement as a potential multi-targeted cancer therapy.
3.6 Molecular dynamics simulation analysis
The molecular interactions and the aqueous solvent conditions surrounding the protein affect the conformational permanence of the protein–ligand association. The most well-suited (7) posture with the greatest binding capability with EGFR TKD (PDB ID: 1M17)56 was used as the commencing structure, and a molecular dynamics simulation113–115 was used to forecast its association and stability. Consequently, a long-range molecular dynamics simulation lasting 200 ns was conducted on the optimally aligned structure to examine the dynamics, conformational stability, and structural integrity of the protein–ligand combination. We recognized that the whole-protein RMSD was higher because of large-amplitude movements in a distal C-terminal section (residues 964 to end) that contains an uncertain stretch in our coordinates (residues 965 to 976).56 We cut off the model at residue 965 before running MDS and calculated RMSD/RMSF after adapting to the kinase core. This got rid of tail-driven artifacts while keeping the active-site geometry and ligand pose. This is because missing strings of residues in PDB structures usually show intrinsically flexible/disordered termini that are far from and don't affect active-site binding. These shortened segments of the distal C-terminus are being used for ligand pose interaction, active site stability, and conformational analysis.116This study used the Root-Mean-Square Deviation (RMSD) to assess how stable the systems were during the 200 ns simulations compared to the initial frame. The most acceptable range for RMSD values was <5.0 Å, as a lower RMSD value means the system is more stable.115 The average RMSD values for all frames of the protein EGFR TKD, compound 7, and ligand–protein complex systems shown in Fig. 5 were 3.634, 3.714, and 4.062 Å, respectively. The standard deviations of the average RMSD values were 0.603, 0.587, and 0.560 Å, respectively. Fig. S5 shows the Root-Mean-Square Deviation (RMSD) during the 200 ns simulations compared to the frame from 0.05 ns ago. The values were 1.229, 0.926, and 1.245 Å, with standard deviations of 0.126, 0.321, and 0.128 Å, respectively. Considering the composition of the examined ligand and complex, the results indicated that the complex system achieved conformational stability.
 | ||
| Fig. 5 The root means square deviation (RMSD) of solvated ligand (7), receptor, and protein ligand complex (7-1M17) during 200 ns MD simulation time. | ||
To look at how residues behave and how they interact with the ligand during MD simulation creation, you need to look at how flexible the protein structure is when the ligand binds to it.117 The Root-Mean-Square Fluctuation (RMSF) approach was used to look at changes in protein residues across 200 ns of simulations to see how binding inhibitors to the right sites affected them. The average RMSF and the standard deviation of the average RMSF values were 1.787 and 1.574 Å, respectively. Fig. S6 shows how the residues in both systems, protein–ligand complexes, change over time. The terminal residues exhibited greater variations compared to the core residues, which is anticipated due to the flexible characteristics of the bimolecular terminals. The system's average RMSF is less than 5 Å, which means that the complex is quite stable and the ligand molecule has a strong affinity for the receptors.
We used an angle cut = 20° and rcut = 3.0 Å to figure out how many hydrogen bonds formed between the solvated protein and ligands. We then charted the results in contrast to time (200 ns) in Fig. S7a and b. For the solvated protein–ligand complex, the average number of hydrogen bonds per timeframe was 3.1, and for the protein–ligand complex systems, it was 0.26. The total study revealed that ligand–protein interaction dramatically altered the amount of hydrogen bonds, increasing from 1 to 9 HBs per trajectory analysis in the instance of the solvated protein–ligand complex, and from 1 to 2 in the case of protein–ligand complex systems. These findings show that the system becomes relatively stable for interactions between hydrogen bonds.
The radius of gyration (Rg) shows how tightly packed the protein structure is and how stable the simulation is. For all frames of protein EGFR TKD, compound 7, and ligand–protein complex systems as displayed in Fig. S8, the average Rg values were 2.054, 0.776, and 2.049 nm, respectively. The standard deviation of the average Rg values was 0.208, 0.675, and 0.197 Å, respectively. This is clear because of how ligands and bimolecular receptor complexes work. The overall change in the average Rg of the protein–ligand complex shows that it reaches equilibrium quickly, which means that the binding mode and interactions in the system are stable.
The solvent-accessible surface area (SASA) of the protein was computed during molecular dynamics simulation under ligand-bound circumstances, attributable to ligand binding to the protein. The average SASA values for all frames of protein EGFR TKD, compound 7, and ligand–protein complex systems shown in Fig. S9 were 162.541, 12.889, and 163.004 nm2, respectively. The standard deviations of the average SASA values were 3.446, 0.553, and 3.729 nm2, respectively. The study shows the folding configuration of the protein and how stable it is when it binds to a ligand, based on the nature of the complex and the outcomes that show a little change in the protein and its complexes. TKD complex equilibrium rapidly stabilizes, exhibiting oscillations within a restricted range, accompanied by minimal conformational alterations and a stable SASA, which correlates with enhanced hydrophobic interactions.
To quantify the binding between the EGFR TKD receptor and the ligand, a contact frequency (CF) study was conducted using the contact Freq.tcl module on VMD, with a cutoff of 4 Å, as presented in Fig. S10. In the simulated receptor–ligand complex, the following amino acid residues displayed elevated CF values ranging from 93.65% to 43.0%: GLY695, PHE699, SER696, VAL702, ARG817, THY766, CYS773, ASN818, LEU694, ALA719, ASP831, LEU820, LYS721, THR830, LYS830, THR830, and VAL693. These values indicate that the complex system demonstrates a strong binding affinity to TKD inhibition complexes.
The molecular mechanics energy methodology (MM/GBSA and PBSA) is a well-known way to find the binding free energies of receptor–ligand complexes. GBSA merges the generalized Born and surface area continuum solvation models, whereas the one-average molecular mechanics Poisson–Boltzmann surface area (MM-PBSA) method70,118–120 is path-independent and calculates the free energy of binding by examining ensembles of the initial and final states. Consequently, MM-PBSA demonstrates superior efficiency compared to MMGBSA. Both methods could be more dependable, less costly in terms of calculation, and more efficient than docking scores.117,121 Based on molecular dynamics (MD) simulated data, we calculated the binding free energy of the simulated complex to double-check the protein affinity that docking simulation studies had predicted for receptor–ligand complexes. Table 3 shows all the calculated contributions of each energy component in MMGBSA and MMPBSA for both complexes. Strong negative values show advantageous interactions and strong receptor–ligand affinity, whereas positive net binding energy shows low-docked stability. The net binding free energy of the study complex was determined to be −31.84 kcal mol−1 for GBSA and −28.17 kcal mol−1 for PBSA. A detailed analysis of the individual energy contributions indicates that the van der Waals, electrostatic, and free energies for the gas phase are −50.11, −21.83, and −71.94 kcal mol−1, respectively, corresponding to the complex. Notably, the van der Waals and electrostatic energies significantly influenced the binding affinity that facilitated complex formation. Finally, the energy of the surface and non-polar components is very reasonable for complexes, ranging from −6.99 to −6.23 kcal mol−1. The net binding free energy stabilizes the conformation of the receptor–ligand affinity complex, indicating a promising and high affinity between the ligand and receptor. This must be experimentally evaluated to confirm the biological efficacy of the proposed pharmacological inhibition.
| Protein–ligand complexa | ΔEVDW | ΔEEEL | ΔGGAS | GBSA | PBSA | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ΔEGB | ΔESURF | ΔGSOLV | ΔTOTAL | ΔEPB | ΔENPOLAR | ΔGSOLV | ΔTOTAL | ||||
| a ΔEVDW = van der Waals energy; ΔEELE = electrostatic energy; ΔGGAS = gas phase free energy; ΔGSOLV = solvation free energy; ΔESURF = Surface energy; ΔEGB = generalized Born energy; ΔEPB = Poisson–Boltzmann energy; ΔENPOLAR = non-polar energy; ΔGbin = calculated total binding free energy (kcal mol−1). MM-GBSA molecular mechanics generalized Born surface area; MM-PBSA Molecular mechanics Poisson–Boltzmann surface area. | |||||||||||
| 7-1M17 | −50.11 | −21.83 | −71.94 | 47.09 | −6.99 | 40.1 | −31.84 | 50 | −6.23 | 43.77 | −28.17 |
These results provide valuable mechanistic insight into the molecular basis of compound 7's activity. Docking studies indicated favorable initial binding, which was corroborated by MD simulations showing sustained stability and reduced conformational flexibility of the EGFR catalytic domain.122,123 The favorable binding free energy, together with its decomposition by MM/PBSA and MM/GBSA, further supports the robustness of this interaction and offers both structural and energetic justification for the experimentally observed anticancer potential of compound 7.124 The simulations highlight a significant inhibitory capacity, reinforcing compound 7 as a promising framework for the development of novel EGFR-targeted anticancer agents.125,126
3.7 EGFR inhibition assay of compound 7
Following molecular docking and molecular dynamics analyses, compound 7 was selected for biochemical evaluation due to its strong predicted affinity toward EGFR, a well-established therapeutic target in several malignancies.127 In silico studies revealed that compound 7 displayed superior docking scores with EGFR compared to the other proteins investigated, suggesting its potential as a novel EGFR inhibitor.To experimentally validate this prediction, compound 7 was tested using an EGFR kinase inhibition assay at concentrations ranging from 7.81 to 1000 µg mL−1 and compared to sorafenib, a clinically approved multikinase inhibitor.128 As shown in Fig. 6, compound 7 inhibited EGFR in a dose-dependent manner, with an IC50 value of 21.45 µg mL−1, whereas sorafenib exhibited a stronger effect with an IC50 of 7.52 µg mL−1 under the same conditions. Although less potent than sorafenib, compound 7 demonstrated clear inhibitory activity, supporting its ability to interact with the EGFR active site in a biologically relevant manner. The observed inhibition is consistent with docking predictions, which indicated that compound 7 engages key residues within the EGFR binding cleft through hydrogen bonding and hydrophobic interactions critical for kinase inhibition. The alignment between computational and experimental findings further supports EGFR as a plausible molecular target of compound 7. Moreover, the antiproliferative effects observed in cellular assays align with the enzymatic inhibition results, suggesting that EGFR suppression contributes to the cytotoxic activity of compound 7. While less potent than sorafenib, compound 7 possesses a distinct scaffold that may confer unique selectivity or pharmacological properties. Structural optimization and kinase panel screening will be important next steps to refine its potency and broaden its therapeutic relevance. Collectively, these results identify compound 7 as a promising EGFR-targeted scaffold warranting further development as a potential anticancer agent.
3.8 ADMET analysis of 7
Initial chemical assessment in current anticancer drug development heavily depends on monitoring ADMET (absorption, distribution, metabolism, excretion, and toxicity). It assists in pinpointing candidates with outstanding pharmacokinetic attributes and acceptable safety thresholds before significant in vivo or clinical evaluations.129,130 Many of the most promising cytotoxic drugs fail in subsequent phases due to low bioavailability, high toxicity, or metabolic instability.129,130 Employing ADMET analysis129,131 ensures that potential drug candidates like 7 meet necessary efficacy and safety criteria, while also enhancing lead optimization, reducing late-stage attrition, and accelerating development timelines.Initial chemical assessment in current anticancer drug development heavily depends on monitoring ADMET (absorption, distribution, metabolism, excretion, and toxicity). It assists in pinpointing candidates with outstanding pharmacokinetic attributes and acceptable safety thresholds before significant in vivo or clinical evaluations.129,130 Many of the most promising cytotoxic drugs fail in subsequent phases due to low bioavailability, high toxicity, or metabolic instability.130 Employing ADMET analysis129,131 ensures that potential drug candidates like 7 meet necessary efficacy and safety criteria, while also enhancing lead optimization, reducing late-stage attrition, and accelerating development timelines.
Compound 7 demonstrated favorable absorption characteristics, supporting its potential oral bioavailability. As shown in Table 4, it exhibited a high predicted intestinal absorption rate (97.87%), consistent with a moderate Caco-2 permeability value (0.489 log Papp in 10−6 cm s−1), suggesting efficient uptake across intestinal epithelial cells. The compound was also predicted to be both a substrate and dual inhibitor of P-glycoprotein (P-gp), indicating the possibility of efflux-mediated transport and transporter-related drug interactions. This dual behavior may influence intracellular drug accumulation and could play a role in overcoming resistance mechanisms commonly associated with P-gp overexpression in cancer cells. In addition, the predicted skin permeability (−2.735 log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp) indicated negligible dermal absorption, while the low passive PAMPA permeability suggested that active transporters may facilitate uptake in specific tissues. Collectively, these findings highlight a favorable absorption profile for compound 7, supporting its advancement into further preclinical evaluation.
Kp) indicated negligible dermal absorption, while the low passive PAMPA permeability suggested that active transporters may facilitate uptake in specific tissues. Collectively, these findings highlight a favorable absorption profile for compound 7, supporting its advancement into further preclinical evaluation.
| Property | Model name | Predicted value | Unit |
|---|---|---|---|
| Absorption | Water solubility | −4.662 | Numeric (log mol L−1) |
| Caco2 permeability | 0.489 | Numeric (log Papp in 10−6 cm s−1) | |
| Intestinal absorption (human) | 97.873 | Numeric (% absorbed) | |
| Skin permeability | −2.735 | Numeric (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp) Kp) |
|
| P-Glycoprotein substrate | Yes | Categorical (yes/no) | |
| P-Glycoprotein I inhibitor | Yes | Categorical (yes/no) | |
| P-Glycoprotein II inhibitor | Yes | Categorical (yes/no) | |
| Distribution | VDss (human) | 0.098 | Numeric (log L kg−1) |
| Fraction unbound (human) | 0.015 | Numeric (Fu) | |
| BBB permeability | −0.611 | Numeric (log BB) | |
| CNS permeability | −3.077 | Numeric (log PS) | |
| Metabolism | CYP2D6 substrate | No | Categorical (yes/no) |
| CYP3A4 substrate | Yes | Categorical (yes/no) | |
| CYP1A2 inhibitor | No | Categorical (yes/no) | |
| CYP2C19 inhibitor | Yes | Categorical (yes/no) | |
| CYP2C9 inhibitor | Yes | Categorical (yes/no) | |
| CYP2D6 inhibitor | No | Categorical (yes/no) | |
| CYP3A4 inhibitor | Yes | Categorical (yes/no) | |
| Excretion | Total clearance | −0.499 | Numeric (log mL min−1 kg−1) |
| Renal OCT2 substrate | No | Categorical (yes/no) | |
| Toxicity | AMES toxicity | No | Categorical (yes/no) |
| Max. tolerated dose (human) | 0.075 | Numeric (log mg kg−1 per day) | |
| hERG I inhibitor | No | Categorical (yes/no) | |
| hERG II inhibitor | Yes | Categorical (yes/no) | |
| Oral rat acute toxicity (LD50) | 3.128 | Numeric (mol kg−1) | |
| Oral rat chronic toxicity (LOAEL) | 1.422 | Numeric (log mg kg−1_bw per day) | |
| Hepatotoxicity | Yes | Categorical (yes/no) | |
| Skin sensitisation | No | Categorical (yes/no) | |
| T. Pyriformis toxicity | 0.285 | Numeric (log ug L−1) | |
| Minnow toxicity | −2.124 | Numeric (log mM) |
Distribution assessments indicated a relatively low volume of distribution (0.098 log L kg−1) and significant plasma protein binding (percent unbound of 0.015), suggesting restricted tissue infiltration and a low free drug level in plasma. Minimal values for blood–brain barrier permeability (−0.611 log BB) and CNS penetration (−3.077 log PS) suggested insufficient exposure to the central nervous system. This is good for non-CNS anticancer agents because it lowers the risk of neurotoxicity while increasing the drug's exposure to peripheral tumors.132 These traits might lead to a more targeted and safer anticancer profile, especially for solid tumors outside the central nervous system.
Concerning metabolism, compound 7 was identified as a CYP3A4 substrate, indicating liver metabolism through this primary enzymatic pathway.133 Despite the recognized variability among individuals in this enzyme, CYP2D6 was not a substrate. Concerning potential drug–drug interactions that require attention in combination treatment situations, compound 7 showed inhibitory effects on several CYP isoforms, such as CYP3A4, CYP2C19, and CYP2C9.134 This inhibitory behavior requires careful consideration when used with other CYP-metabolized drugs, since it might change their plasma levels and how well they work. The molecule, however, did not inhibit CYP1A2 or CYP2D6, which somewhat lowered its overall metabolic liability. This type of targeted inhibition implies the possibility of preventing extensive metabolic changes and may provide safer multiple therapy regimens.
While compound 7 is not a substrate for renal OCT2, it displayed a low total clearance rate (−0.499 log mL min−1), indicating prolonged systemic retention, which could enhance therapeutic exposure through this transporter.135 This persistence may lead to an extended dosage interval or prolonged pharmacological action, provided that hazardous thresholds are not surpassed.
From a toxicological perspective, compound 7 exhibited a very strong safety profile. The AMES test showed no mutagenicity, minimal genotoxicity, and no hERG I inhibition,136 suggesting a low risk for cardiotoxicity or QT interval prolongation. But its projected suppression of hERG II means that cardiomyocytes or animal models need to be carefully tested for safety using electrophysiology. The highest dosage tolerated in humans was modest (0.075 log mg kg−1 per day); in rats, both acute and chronic toxicity measures (LD50: 3.128 mol kg−1; LOAEL: 1.422 log mg kg−1 per day) remained within acceptable limits. Hepatotoxicity forecasts notably recommend rigorous liver surveillance during the preliminary in vivo studies. Even though hepatotoxicity appeared to require liver attention during development, the substance was not anticipated to cause skin sensitivity.
Ultimately, information regarding environmental toxicity indicated slight to significant effects on aquatic organisms, implying that ecological consequences must be evaluated in subsequent stages. This consideration becomes particularly important when addressing large-scale production and minimizing the environmental impact of manufacturing waste. Overall, compound 7 exhibits a drug-like ADMET profile, characterized by high intestinal absorption, selective metabolic interactions, regulated tissue distribution, and manageable toxicity risks. These integrated pharmacokinetic and toxicological properties support its continued investigation as a promising multi-target anticancer candidate, warranting further validation through experimental pharmacology and in vivo studies.
4 Conclusion
The increasing occurrence of cancer, especially in developing regions, highlights the pressing demand for new therapeutic agents that can tackle resistance to existing treatments. In this study, we created a range of bis-heterocyclic structures connected by a pyridine unit, illustrating a new category of hybrid substances with significant pharmaceutical promise. These derivatives can be produced effectively under gentle conditions using easily accessible starting materials, allowing for the generation of adequate amounts for biological assessment. The synthesized compounds' cytotoxic activities were evaluated on A549 (lung cancer), HepG2 (hepatocellular carcinoma), and MCF7 (breast cancer) cell lines, using human fibroblasts (HSF) as a normal control. Out of the compounds evaluated, compound 7 demonstrated the most potent anticancer effects, significant selectivity for cancerous cells, and low toxicity to normal fibroblasts. Molecular docking analyses indicated that compound 7 could function via various oncogenic pathways, such as EGFR and FGFR4, essential for the survival of cancer cells. Toxicological profiling additionally revealed its beneficial safety features, such as non-mutagenicity and a low risk of cardiotoxicity. Moreover, ADMET evaluations validated its pharmaceutical characteristics, demonstrating promising absorption and tolerable safety measures. Combined, these findings emphasize compound 7 as a potential multi-target anticancer candidate. Subsequent research must encompass thorough in vivo assessments to analyze its pharmacodynamic effectiveness, bioavailability, metabolic stability, and overall safety. These studies will offer crucial translational insights and enhance its prospects as a feasible candidate for preclinical development.Conflicts of interest
There are no conflicts to declare.Data availability
All data generated or analyzed during this study are included in the article and its supplementary information (SI). Supplementary information: molecular docking interactions of compound 7 with FGFR4, Aurora A, CDK1, and VEGFR2; molecular dynamics simulation analyses (RMSD, RMSF, hydrogen bonding, radius of gyration, SASA, and contact frequency); 1H and 13C NMR and IR spectra of compounds 7, 8, 11, 15, 16, and 18; and in vitro cytotoxicity (IC50 values (µM) and selectivity index) data against HepG2, A549, MCF7, and HFB4 cell lines.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6ra00243a.
Acknowledgements
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).References
- F. Bray, M. Laversanne, H. Sung, J. Ferlay, R. L. Siegel, I. Soerjomataram and A. Jemal, Ca-Cancer J. Clin., 2024, 74, 229–263 CrossRef PubMed.
- J. M. Kocarnik, K. Compton, F. E. Dean, W. Fu, B. L. Gaw, J. D. Harvey and H. J. Henrikson, JAMA Oncol., 2022, 8, 420–444 CrossRef PubMed.
- Y. Zheng, Y. Chen, K. Yu, Y. Yang, X. Wang, X. Yang, J. Qian, Z. X. Liu and B. Wu, Infect. Dis. Ther., 2021, 10, 871–895 CrossRef PubMed.
- A. Mushtaq, P. Wu and M. M. Naseer, Pharmacol. Ther., 2024, 254, 108579 CrossRef CAS PubMed.
- M. Hossain, I. Habib, K. Singha and A. Kumar, Heliyon, 2024, 10, e23172 CrossRef CAS PubMed.
- D. Sahu, P. S. R. Sreekanth, P. K. Behera, M. K. Pradhan, A. Patnaik, S. Salunkhe and R. Cep, Eur. J. Med. Chem. Rep., 2024, 12, 100210 CAS.
- S. De, A. Kumar S K, S. K. Shah, S. Kazi, N. Sarkar, S. Banerjee and S. Dey, RSC Adv., 2022, 12, 15385–15406 Search PubMed.
- G. Mohammad Abu-Taweel, M. M. Ibrahim, S. Khan, H. M. Al-Saidi, M. Alshamrani, F. A. Alhumaydhi and S. S. Alharthi, Crit. Rev. Anal. Chem., 2024, 54, 599–616 CrossRef CAS PubMed.
- Y. Ling, Z. Y. Hao, D. Liang, C. L. Zhang, Y. F. Liu and Y. Wang, Drug Des., Dev. Ther., 2021, 15, 4289–4338 CrossRef CAS PubMed.
- M. M. Mehany, O. A. Hammam, A. A. Selim, G. H. Sayed and K. E. Anwer, Sci. Rep., 2024, 14, 2738 CrossRef CAS PubMed.
- A. Manaithiya, O. Alam, V. Sharma, M. J. Naim, S. Mittal, F. Azam, A. Husain, A. A. Sheikh, M. Imran and I. A. Khan, Curr. Top. Med. Chem., 2021, 21, 2292–2349 CrossRef CAS PubMed.
- G. Santhosh Kumar, Y. Poornachandra, S. Kumar Gunda, K. Ratnakar Reddy, J. Mohmed, K. Shaik, C. Ganesh Kumar and B. Narsaiah, Bioorg. Med. Chem. Lett., 2018, 28, 2328–2337 CrossRef CAS PubMed.
- S. R. Alizadeh and M. A. Ebrahimzadeh, Mini-Rev. Med. Chem., 2021, 21, 2584–2611 CrossRef CAS PubMed.
- K. H. Narasimhamurthy, Y. R. Girish, T. R. Swaroop and K. S. Rangappa, Lett. Drug Des. Discovery, 2023, 21, 1904–1912 CrossRef.
- Z. M. Alamshany and E. S. Nossier, J. Mol. Struct., 2024, 1316, 138973 Search PubMed.
- H. M. Sampath Kumar, L. Herrmann and S. B. Tsogoeva, Bioorg. Med. Chem. Lett., 2020, 30, 127514 Search PubMed.
- A. H. Alkhzem, T. J. Woodman and I. S. Blagbrough, RSC Adv., 2022, 12, 19470–19484 Search PubMed.
- H. M. Diab, D. H. Ali, T. A. Abdallah, M. M. Soliman, A. H. M. Elwahy, I. A. Abdelhamid, M. E. Salem and I. M. Z. Fares, J. Heterocycl. Chem., 2025, 62, 1679–1697 Search PubMed.
- H. M. Diab, M. E. Salem, M. Abdellatif, M. M. Soliman, I. A. Abdelhamid, A. H. M. Elwahy, N. Raza and Y. A. El-Gabry, ACS Omega, 2025, 10, 38014–38033 CrossRef CAS PubMed.
- H. H. Mohamed, M. E. Salem, E. B. Abdelazim, A. M. Abdelmoniem, M. Elsabahy, N. Raza, A. H. Abdullah and I. A. Abdelhamid, ACS Omega, 2025, 10, 46868–46883 CrossRef CAS PubMed.
- M. A. Ragheb, M. E. Salem, A. A. Hamed, I. A. Abdelhamid, H. A. Ali, M. Abdel-Megid and A. M. Elgamal, Int. J. Biol. Macromol., 2025, 331, 148298 Search PubMed.
- A. M. Elgamal, M. E. Salem, A. A. Hamed, I. A. Abdelhamid, T. A. Abdallah, H. A. Ali, N. Raza and M. A. Ragheb, Carbohydr. Polym., 2025, 369, 124258 CrossRef CAS PubMed.
- A. M. Abdelmoniem, M. E. Salem, M. H. M. Allam, M. M. Abou-Krisha, I. O. Althobaiti, S. A. S. Ghozlan, K. E. Azmy and I. A. Abdelhamid, Results Chem., 2025, 14, 102065 CrossRef CAS.
- F. G. Mohamed, M. A. Ragheb, A. H. M. Elwahy, H. M. Diab, M. Emara, M. E. Salem, I. O. Althobaiti and I. A. Abdelhamid, Results Chem., 2025, 14, 102119 CrossRef CAS.
- E. M. Fathi, M. E. Salem, F. M. Sroor, F. G. Mohamed, A. H. M. Elwahy, M. Abdel-Megid, I. A. Abdelhamid and M. Emara, ACS Omega, 2025, 10, 25921–25937 CrossRef PubMed.
- N. S. Ibrahim, E. H. Shoukry, M. Sharaky, H. M. Diab, A. H. M. Elwahy and I. A. Abdelhamid, Chem. Biol. Interact., 2025, 416, 111549 CrossRef CAS PubMed.
- M. A. Ragheb, H. E. Abdelrashid, E. M. Elzayat, I. A. Abdelhamid and M. H. Soliman, J. Biomol. Struct. Dyn., 2025, 43, 9139–9157 CrossRef CAS PubMed.
- H. M. Salem, A. H. Orabi, I. A. Abdelhamid, G. R. Saad, D. A. Ismaiel and A. A. Hamed, Int. J. Biol. Macromol., 2025, 311, 144100 CrossRef CAS PubMed.
- M. A. Ragheb, M. S. Ragab, F. Y. Mahdy, M. S. Elsebaie, A. M. Saber, Y. O. AbdElmalak, R. H. Elsafoury, A. A. Elatreby, A. M. Rochdi, A. W. El-Basyouni, M. M. Shoukry, M. A. Eldeeb, R. M. El-Sherif, I. A. Abdelhamid and D. S. Salah-Eldin, Int. J. Biol. Macromol., 2025, 311, 144034 CrossRef CAS PubMed.
- R. E. Abdelwahab, A. H. M. Elwahy, N. S. Ibrahim, A. M. Abdelmoniem and I. A. Abdelhamid, BMC Chem., 2025, 19, 1–14 Search PubMed.
- M. F. Mohamed, A. A. Hamed, A. A. Kashmiry, A. A. Saddiq, I. A. Abdelhamid and A. M. Elgamal, Int. J. Biol. Macromol., 2025, 308, 142048 CrossRef CAS PubMed.
- I. M. Z. Fares, M. E. Salem, M. S. Shafik, I. A. Abdelhamid, A. H. M. Elwahy, N. S. Ibrahim, M. Abdel-Megid and H. M. Diab, J. Mol. Struct., 2025, 1339, 142345 CrossRef CAS.
- I. M. Z. Fares, M. A. Ragheb, I. A. Abdelhamid and A. H. M. Elwahy, J. Mol. Struct., 2025, 1323, 140790 CrossRef CAS.
- A. Shaban, Z. Zakaria, I. A. Abdelhamid, H. K. A. Elhakim and E. R. El-sayed, Process Biochem., 2025, 149, 192–203 CrossRef CAS.
- N. S. Ibrahim, R. A. A. Elghany, M. Sharaky, H. M. Diab, I. A. Abdelhamid and A. H. M. Elwahy, J. Mol. Struct., 2025, 1328, 141424 CrossRef CAS.
- M. M. Soliman, A. H. M. Elwahy, A. M. Sayed, M. Ibrahim, M. A. Dawoud, S. H. M. Ali, M. T. S. Nady, N. A. Hassan, W. Saad, I. A. Abdelhamid and N. Schmiedebergs, Arch. Pharmacol., 2025, 398, 8587–8617 CrossRef CAS PubMed.
- M. A. Ragheb, M. E. Salem, M. M. Ali, I. A. Abdelhamid, N. Raza and M. H. Soliman, ACS Omega, 2025, 43, 51497–51511 Search PubMed.
- Y. A. Ibrahim, A. A. Abbas and A. H. M. Elwahy, J. Heterocycl. Chem., 2004, 41, 135–149 CrossRef CAS.
- Y. A. Ibrahim, A. H. M. Elwahy and A. A. Abbas, Tetrahedron, 1994, 50, 11489–11498 Search PubMed.
- A. H. M. Elwahy and A. A. Abbas, Synth. Commun., 2000, 30, 2903–2921 Search PubMed.
- A. Elwahy and M. Shaaban, Curr. Org. Synth., 2014, 11, 835–873 CrossRef CAS.
- B. N. Barsoum, S. K. Khella, A. H. M. Elwaby, A. A. Abbas and Y. A. Ibrahim, Talanta, 1998, 47, 1215–1222 CrossRef CAS PubMed.
- M. E. Salem, A. F. Darweesh, A. M. Farag and A. H. M. Elwahy, Tetrahedron, 2016, 72, 712–719 Search PubMed.
- Y. A. Ibrahim, A. A. Abbas and A. H. M. Elwahy, Carbohydr. Lett., 1999, 3, 331–338 CAS.
- A. H. M. Elwahy and A. A. Abbas, J. Heterocycl. Chem., 2008, 45, 1–65 CrossRef CAS.
- A. H. M. Elwahy, A. A. Abbas and R. M. Kassab, Synthesis, 2002, 260–264 CAS.
- M. A. Ragheb, M. H. Soliman, I. A. Abdelhamid, M. M. Shoukry, M. Haukka and M. S. Ragab, J. Inorg. Biochem., 2024, 253, 112488 CrossRef CAS PubMed.
- M. S. Ragab, M. R. Shehata, M. M. Shoukry, M. Haukka and M. A. Ragheb, RSC Adv., 2022, 12, 1871–1884 Search PubMed.
- J. Akhtar, A. A. Khan, Z. Ali, R. Haider and M. Shahar Yar, Eur. J. Med. Chem., 2017, 125, 143–189 CrossRef CAS PubMed.
- H. Nada, A. Elkamhawy and K. Lee, Molecules, 2021, 26, 553 CrossRef CAS PubMed.
- N. Abdelaal, M. A. Ragheb, H. M. Hassaneen, E. M. Elzayat and I. A. Abdelhamid, Sci. Rep., 2024, 14, 26647 CrossRef CAS PubMed.
- M. A. Ragheb, R. E. Abdelwahab, A. F. Darweesh, M. H. Soliman, A. H. M. Elwahy and I. A. Abdelhamid, Chem. Biodivers., 2022, 19, e202100958 CrossRef CAS PubMed.
- R. E. Abdelwahab, M. A. Ragheb, A. H. M. Elwahy, I. A. Abdelhamid and A. M. Abdelmoniem, J. Mol. Struct., 2024, 1307, 137946 CrossRef CAS.
- S. Vilar, G. Cozza and S. Moro, Curr. Top. Med. Chem., 2008, 8, 1555–1572 Search PubMed.
- C. R. Corbeil, C. I. Williams and P. Labute, J. Comput. Aided Mol. Des., 2012, 26, 775–786 CrossRef CAS PubMed.
- J. Stamos, M. X. Sliwkowski and C. Eigenbrot, J. Biol. Chem., 2002, 277, 46265–46272 Search PubMed.
- H. J. C. Berendsen, D. van der Spoel and R. van Drunen, Comput. Phys. Commun., 1995, 91, 43–56 Search PubMed.
- S. Páll, M. J. Abraham, C. Kutzner, B. Hess and E. Lindahl, in Solving Software Challenges for Exascale: International Conference on Exascale Applications and Software, EASC 2014, Stockholm, Sweden, April 2-3, 2014, Revised Selected Papers 2, 2015, pp. 3–27 Search PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1, 19–25 CrossRef.
- R. B. Best, X. Zhu, J. Shim, P. E. M. Lopes, J. Mittal, M. Feig and A. D. MacKerell Jr, J. Chem. Theory Comput., 2012, 8, 3257–3273 Search PubMed.
- J. Huang, S. Rauscher, G. Nawrocki, T. Ran, M. Feig, B. L. de Groot, H. Grubmüller and A. D. MacKerell, Nat. Methods, 2017, 14, 71–73 Search PubMed.
- J. Lee, X. Cheng, J. M. Swails, M. S. Yeom, P. K. Eastman, J. A. Lemkul, S. Wei, J. Buckner, J. C. Jeong and Y. Qi, et al., J. Chem. Theory Comput., 2016, 12, 405–413 Search PubMed.
- S. Kim, J. Lee, S. Jo, C. L. Brooks III, H. S. Lee and W. Im, J. Comput. Chem., 2017, 38, 1879–1886 Search PubMed.
- S. Jo, T. Kim, V. G. Iyer and W. Im, J. Comput. Chem., 2008, 29, 1859–1865 Search PubMed.
- B. R. Brooks, C. L. Brooks III, A. D. Mackerell Jr, L. Nilsson, R. J. Petrella, B. Roux, Y. Won, G. Archontis, C. Bartels and S. Boresch, et al., J. Comput. Chem., 2009, 30, 1545–1614 CrossRef CAS PubMed.
- G. S. Grest and K. Kremer, Phys. Rev. A, 1986, 33, 3628 Search PubMed.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graph., 1996, 14, 33–38 CrossRef CAS PubMed.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, J. Chem. Phys., 1995, 103, 8577–8593 CrossRef CAS.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, J. Chem. Theory Comput., 2021, 17, 6281–6291 CrossRef PubMed.
- D. E. V. Pires, T. L. Blundell and D. B. Ascher, J. Med. Chem., 2015, 58, 4066–4072 CrossRef CAS PubMed.
- L. Fu, S. Shi, J. Yi, N. Wang, Y. He, Z. Wu, J. Peng, Y. Deng, W. Wang, C. Wu, A. Lyu, X. Zeng, W. Zhao, T. Hou and D. Cao, Nucleic Acids Res., 2024, 52, W422–W431 Search PubMed.
- K. Sugano, M. Kansy, P. Artursson, A. Avdeef, S. Bendels, L. Di, G. F. Ecker, B. Faller, H. Fischer, G. Gerebtzoff, H. Lennernaes and F. Senner, Nat. Rev. Drug Discovery, 2010, 98(9), 597–614 Search PubMed.
- P. Rajakumar and M. Srisailas, Tetrahedron, 2001, 57, 9749–9754 CrossRef CAS.
- S. M. H. Sanad, R. M. Kassab, I. A. Abdelhamid and A. H. M. Elwahy, Heterocycles, 2016, 92, 910–924 Search PubMed.
- R. Sangsuwan, S. Sangher, T. Aree, C. Mahidol, S. Ruchirawat and P. Kittakoop, RSC Adv., 2014, 4, 13708–13718 Search PubMed.
- S. A. S. Ghozlan, A. G. Ahmed and I. A. Abdelhamid, J. Heterocycl. Chem., 2016, 53, 817–823 Search PubMed.
- Y. R. Lei, X. L. He, J. Li and C. F. Mo, Front. Biosci., 2024, 29, 52 Search PubMed.
- Y. Gao, P. Dorn, S. Liu, H. Deng, S. R. R. Hall, R. W. Peng, R. A. Schmid and T. M. Marti, Cancer Cell Int., 2019, 19, 1–14 Search PubMed.
- G. Chen, W. Liu, B. Yan, G. Chen, W. Liu and B. Yan, J. Cancer Ther., 2022, 13, 117–130 Search PubMed.
- A. Drakontaeidi, I. Papanotas and E. Pontiki, Antioxidants, 2024, 13, 898 Search PubMed.
- P. Nunhart, E. Konkoľová, L. Janovec, R. Jendželovský, J. Vargová, J. Ševc, M. Matejová, B. Miltáková, P. Fedoročko and M. Kozurkova, Bioorg. Chem., 2020, 94, 103393 Search PubMed.
- T. J. Ritchie and S. J. F. Macdonald, Drug Discovery Today, 2009, 14, 1011–1020 CrossRef CAS PubMed.
- A. D. G. Lawson, M. Maccoss and J. P. Heer, J. Med. Chem., 2018, 61, 4383–4389 Search PubMed.
- M. Ismail, R. Ahmad, S. A. Halim, A. A. Khan, S. Ullah, A. Latif, M. Ahmad, A. Khan, F. A. Ozdemir, A. Khalid, A. Al-Harrasi and M. Ali, RSC Adv., 2024, 14, 10978–10994 Search PubMed.
- S. Yoshioka, T. Tosha, S. Takahashi, K. Ishimori, H. Hori and I. Morishima, J. Am. Chem. Soc., 2002, 124, 14571–14579 Search PubMed.
- S. C. Jadhvar, H. M. Kasraliker, S. V. Goswami, A. V. Chakrawar and S. R. Bhusare, Res. Chem. Intermed., 2017, 43, 7211–7221 CrossRef CAS.
- M. Thakur, Nov. Approaches Cancer Stud., 450–454, DOI:10.31031/NACS.2020.05.000604.
- Z. M. Alamshany, E. M. Algamdi, I. M. M. Othman, M. M. Anwar and E. S. Nossier, RSC Adv., 2023, 13, 12889–12905 RSC.
- L. G. Ferreira, R. N. Dos Santos, G. Oliva and A. D. Andricopulo, Molecules, 2015, 20, 13384–13421 Search PubMed.
- D. M. French, B. C. Lin, M. Wang, C. Adams, T. Shek, K. Hötzel, B. Bolon, R. Ferrando, C. Blackmore, K. Schroeder, L. A. Rodriguez, M. Hristopoulos, R. Venook, A. Ashkenazi and L. R. Desnoyers, PLoS One, 2012, 7, e36713 Search PubMed.
- E. Pinto, F. Pelizzaro, F. Farinati and F. P. Russo, Medicina, 2023, 59, 1115 Search PubMed.
- P. Zhou, J. Hu, X. Wang, J. Wang, Y. Zhang and C. Wang, Oncol. Lett., 2018, 15, 5201–5207 Search PubMed.
- D. Galetta and L. Cortes-Dericks, Cancers, 2020, 12, 3371 CrossRef CAS PubMed.
- Q. Wang, A. M. Bode and T. Zhang, npj Precis. Oncol., 2023, 71(7), 1–14 Search PubMed.
- L. Gao, X. Wang, Y. Tang, S. Huang, C. A. A. Hu and Y. Teng, J. Exp. Clin. Cancer Res., 2017, 36, 1–10 Search PubMed.
- J. J. Joshi, H. Coffey, E. Corcoran, J. Tsai, C. L. Huang, K. Ichikawa, S. Prajapati, M. H. Hao, S. Bailey, J. Wu, V. Rimkunas, C. Karr, V. Subramanian, P. Kumar, C. MacKenzie, R. Hurley, T. Satoh, K. Yu, E. Park, N. Rioux, A. Kim, W. G. Lai, L. Yu, P. Zhu, S. Buonamici, N. Larsen, P. Fekkes, J. Wang, M. Warmuth, D. J. Reynolds, P. G. Smith and A. Selvaraj, Cancer Res., 2017, 77, 6999–7013 Search PubMed.
- L. Grisetti, C. J. C. Garcia, A. A. Saponaro, C. Tiribelli and D. Pascut, Cell Prolif., 2024, 57, e13641 CrossRef CAS PubMed.
- M. Kollareddy, D. Zheleva, P. Dzubak, P. S. Brahmkshatriya, M. Lepsik and M. Hajduch, Invest. New Drugs, 2012, 30, 2411–2432 Search PubMed.
- E. C. Dees, R. B. Cohen, M. Von Mehren, T. E. Stinchcombe, H. Liu, K. Venkatakrishnan, M. Manfredi, H. Fingert, H. A. Burris and J. R. Infante, Clin. Cancer Res., 2012, 18, 4775–4784 Search PubMed.
- D. Mahadevan, A. Stejskal, L. S. Cooke, A. Manziello, C. Morales, D. O. Persky, R. I. Fisher, T. P. Miller and W. Qi, Clin. Cancer Res., 2012, 18, 2210–2219 CrossRef CAS PubMed.
- G. Massacci, L. Perfetto and F. Sacco, Br. J. Cancer, 2023, 129, 1707–1716 CrossRef CAS PubMed.
- J. Cicenas, K. Kalyan, A. Sorokinas, E. Stankunas, J. Levy, I. Meskinyte, V. Stankevicius, A. Kaupinis and M. Valius, Ann. Transl. Med., 2015, 3, 135 Search PubMed.
- A. Díaz-Serrano, P. Gella, E. Jiménez, J. Zugazagoitia and L. Paz-Ares Rodríguez, Drugs, 2018, 78, 893–911 CrossRef PubMed.
- Z. Zhao, Z. Yu, J. Li and X. Ouyang, Oncol. Lett., 63–68 Search PubMed.
- A. M. Barbuti, G.-N. Zhang, P. Gupta, S. Narayanan and Z.-S. Chen, Protein Kinase Inhibitors as Sensitizing Agents for Chemotherapy, 2019, pp. 1–11 Search PubMed.
- L. Wang, W. Q. Liu, S. Broussy, B. Han and H. Fang, Front. Pharmacol, 2023, 14, 1307860 CrossRef CAS PubMed.
- Z. L. Liu, H. H. Chen, L. L. Zheng, L. P. Sun and L. Shi, Signal Transduct. Targeted Ther., 2023, 8, 1–39 Search PubMed.
- Y. Wang, C. Peng, G. Wang, Z. Xu, Y. Luo, J. Wang and W. Zhu, Chem. Biol. Drug Des., 2019, 93, 934–948 CrossRef CAS PubMed.
- X. Ou, G. Gao, I. A. Habaz and Y. Wang, MedComm, 2024, 5, e694 CrossRef CAS PubMed.
- G. C. Fletcher, R. D. Brokx, T. A. Denny, T. A. Hembrough, S. M. Plum, W. E. Fogler, C. F. Sidor and M. R. Bray, Mol. Cancer Ther., 2011, 10, 126–137 CrossRef CAS PubMed.
- Y. Li, C. Tan, C. Gao, C. Zhang, X. Luan, X. Chen, H. Liu, Y. Chen and Y. Jiang, Bioorg. Med. Chem., 2011, 19, 4529–4535 Search PubMed.
- S. Mirzaei, F. Eisvand, F. Hadizadeh, F. Mosaffa, A. Ghasemi and R. Ghodsi, Bioorg. Chem., 2020, 98, 103711 Search PubMed.
- T. A. Majrashi, A. Sabt, H. Almahli, M. A. El Hassab, M. A. Noamaan, E. B. Elkaeed, M. F. Hamissa, A. N. Maslamani, M. A. Shaldam and W. M. Eldehna, PLoS One, 2024, 19, e0300035 Search PubMed.
- H. M. Al-Maqtari, A. H. Hasan, M. Suleiman, F. M. A Abid, A. R. Bhat, M. A. A. Zahidi, M. A. Noamaan, S. Murugesan, S. Ahmed and J. Jamalis, ChemistrySelect, 2025, 10, e01227 Search PubMed.
- K. Djinovic-Carugo and O. Carugo, Intrinsically Disord. Proteins, 2015, 3, e1095697 CrossRef PubMed.
- A. H. Hasan, F. A. Abdulrahman, A. J. Obaidullah, H. F. Alotaibi, M. M. Alanazi, M. A. Noamaan, S. Murugesan, S. I. Amran, A. R. Bhat, J. Jamalis, H. Arai, Y. Higashigaki, M. Gotoh and S. Y. S. Yano, Pharmaceuticals, 2023, 16, 5755 CrossRef PubMed.
- R. Kumari, R. Kumar, O. S. D. D. Consortium and A. Lynn, J. Chem. Inf. Model., 2014, 54, 1951–1962 CrossRef CAS PubMed.
- N. A. Baker, D. Sept, S. Joseph, M. J. Holst and J. A. McCammon, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 10037–10041 Search PubMed.
- S. Pronk, S. Páll, R. Schulz, P. Larsson, P. Bjelkmar, R. Apostolov, M. R. Shirts, J. C. Smith, P. M. Kasson and D. Van Der Spoel, et al., Bioinformatics, 2013, 29, 845–854 CrossRef CAS PubMed.
- K. E. Machaba, N. N. Mhlongo and M. E. S. Soliman, Cell Biochem. Biophys., 2018, 76, 345–356 CrossRef CAS PubMed.
- D. D. Li, T. T. Wu, P. Yu, Z. Z. Wang, W. Xiao, Y. Jiang and L. G. Zhao, ACS Omega, 2020, 5, 16307–16314 Search PubMed.
- H. H. Loeffler and M. D. Winn, Proteins: Struct., Funct., Bioinf., 2013, 81, 1931–1943 CrossRef CAS PubMed.
- N. Ornnork, D. Kiriwan, K. Lirdprapamongkol, K. Choowongkomon, J. Svasti and C. Eurtivong, J. Mol. Graphics Modell., 2020, 99, 107639 CrossRef CAS PubMed.
- S. Anwar, J. Alanazi, N. Ahemad, S. Raza, T. A. Chohan and H. Saleem, Front. Pharmacol, 2024, 15, 1399372 CrossRef CAS PubMed.
- Q.-H. Liao, Q.-Z. Gao, J. Wei and K.-C. Chou, Med. Chem., 2010, 7, 24–31 CrossRef PubMed.
- M. A. Ashiru, S. O. Ogunyemi, O. R. Temionu, A. C. Ajibare, N. C. Cicero-Mfon, O. A. Ihekuna, M. O. Jagun, L. Abdulmumin, Q. K. Adisa, Y. E. Asibor, C. J. Okorie, M. O. Lawal, M. O. Babalola, I. T. Abdulrasaq, L. B. Salau, I. O. Olatunji, M. A. Bankole, A. B. Daud and A. O. Adeyemi, J. Mol. Model., 2023, 29, 1–12 CrossRef PubMed.
- F. Morgillo, E. Martinelli, T. Troiani, M. Orditura, F. de Vita and F. Ciardiello, PLoS One, 2011, 6, e28841 CrossRef CAS PubMed.
- F. Wu, Y. Zhou, L. Li, X. Shen, G. Chen, X. Wang, X. Liang, M. Tan and Z. Huang, Front. Chem., 2020, 8, 546712 Search PubMed.
- A. M. B. Amorim, L. F. Piochi, A. T. Gaspar, A. J. Preto, N. Rosário-Ferreira and I. S. Moreira, Chem. Res. Toxicol., 2024, 37, 827–849 Search PubMed.
- L. L. G. Ferreira and A. D. Andricopulo, Drug Discovery Today, 2019, 24, 1157–1165 Search PubMed.
- F. M. G. Cornelissen, G. Markert, G. Deutsch, M. Antonara, N. Faaij, I. Bartelink, D. Noske, W. P. Vandertop, A. Bender and B. A. Westerman, J. Med. Chem., 2023, 66, 7253–7267 Search PubMed.
- Y. Zhang, Z. Wang, Y. Wang, W. Jin, Z. Zhang, L. Jin, J. Qian and L. Zheng, PeerJ, 2024, 12, e18636 Search PubMed.
- N. Muhamad and K. Na-Bangchang, Pharmacol. Res. Perspect., 2023, 11, e01149 CrossRef CAS PubMed.
- L. Zhang, C. M. Brett and K. M. Giacomini, Annu. Rev. Pharmacol. Toxicol., 1998, 38, 431–460 Search PubMed.
- A. Garrido, A. Lepailleur, S. M. Mignani, P. Dallemagne and C. Rochais, Eur. J. Med. Chem., 2020, 195, 112290 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |