Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Design, synthesis and antiproliferative, apoptotic, and immunomodulatory properties of new heteroaryl pyridine-linked 1,2,4-oxadiazoles as prospective dual EGFR/BRAFV600E inhibitors
Hesham A. M. Gomaa *a,
Mohamed E. Shakera,
Sami I. Alzareaa,
Eid Alatwia,
Fatma A. M. Mohamedb,
Abdullah Yahya Abdullah Alzahranic,
Bandar A. Alyamid,
Stefan Bräse
*a,
Mohamed E. Shakera,
Sami I. Alzareaa,
Eid Alatwia,
Fatma A. M. Mohamedb,
Abdullah Yahya Abdullah Alzahranic,
Bandar A. Alyamid,
Stefan Bräse *e,
Safwat M. Rabeaf and
Bahaa G. M. Youssif
*e,
Safwat M. Rabeaf and
Bahaa G. M. Youssif *g
*g
aDepartment of Pharmacology, College of Pharmacy, Jouf University, Sakaka 72388, Saudi Arabia. E-mail: hasoliman@ju.edu.sa
bDepartment of Clinical Laboratory Sciences, College of Applied Medical Sciences at Al-Qurayyat, Jouf University, Al-Qurayyat 72388, Saudi Arabia
cDepartment of Chemistry, Faculty of Science, King Khalid University, Abha 61413, Saudi Arabia
dDepartment of Pharmaceutical Chemistry, College of Pharmacy, Najran University, Najran, Saudi Arabia
eInstitute of Biological and Chemical Systems, IBCS-FMS, Karlsruhe Institute of Technology, 76131 Karlsruhe, Germany. E-mail: braese@kit.edu
fMedicinal Chemistry Department, Faculty of Pharmacy, Minia University, Minia 61519, Egypt
gPharmaceutical Organic Chemistry Department, Faculty of Pharmacy, Assiut University, Assiut 71526, Egypt. E-mail: bgyoussif2@gmail.com; Tel: +201044353895
First published on 16th March 2026
Abstract
A novel series of heteroaryl pyridine-linked 1,2,4-oxadiazole compounds (5, 9, 14, 20a–c, 21a–c, and 22a–c) was developed, synthesized, and investigated as prospective inhibitors of EGFR and BRAFV600E. The new compounds were investigated for antiproliferative activity against four human cancer cell lines and for safety in normal mammary epithelial cells (MCF-10A) and a normal human diploid cell line (WI-38). Compounds 20c, 21a–c, and 22b demonstrated significant antiproliferative action, with compounds 20c and 21c exhibiting the highest efficacy. Compounds 20c and 21c exhibited potent inhibition of EGFR, with IC50 values of 71 and 64 nM, respectively, surpassing the reference erlotinib (IC50 = 80 nM). Moreover, compounds 20c and 21c exhibited BRAFV600E inhibitory action with IC50 values of 49 and 41 nM, respectively, which are somewhat less potent than the reference drug Vemurafenib. Assays for apoptotic markers (Caspases, Bax, Bcl-2, and p53) demonstrated that apoptosis plays a role in the reported antiproliferative effects. Compounds 20c and 21c showed a notable decrease in TNF-α and IL-6 levels compared with dexamethasone, suggesting an immunomodulatory effect. Molecular docking further validated the favorable orientation of 20c and 21c within the ATP-binding pocket of EGFR and BRAFV600E. These findings underscore compounds 20c and 21c as innovative dual-target scaffolds with significant promise for anticancer drug development.
1. Introduction
Cancer is a primary cause of morbidity and mortality globally, and its epidemiological profile is becoming increasingly complex.1 The American Cancer Society projects that by 2026, over 2.1 million new cancer cases will be diagnosed in the United States. That amounts to over 5800 new cases daily.2,3 Breast and colorectal malignancies are two cancer types frequently targeted by novel heterocyclic scaffolds. These malignancies remain a significant public health issue, with an expected 321![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 910 and 158
910 and 158![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 850 new cases, respectively.4 The cumulative five-year relative survival rate has reached an unprecedented 70%. The rising incidence rates in younger populations and the persistence of metastatic disease highlight the urgent need for innovative treatments that might deliver sustained clinical outcomes.5,6
850 new cases, respectively.4 The cumulative five-year relative survival rate has reached an unprecedented 70%. The rising incidence rates in younger populations and the persistence of metastatic disease highlight the urgent need for innovative treatments that might deliver sustained clinical outcomes.5,6
Despite advances in precision medicine and immunotherapy, many modern medications continue to have substantial difficulties. Traditional chemotherapeutic drugs can occasionally lack specificity, resulting in severe systemic toxicities such as myelosuppression and cardiotoxicity, which significantly limit the maximum tolerated dose.7,8 Furthermore, the establishment of multidrug resistance (MDR) is a major factor of treatment failure and illness recurrence. These issues call for the development of new chemical frameworks.9 These frameworks may improve metabolic stability and cellular permeability, and enable more precise interactions with cancer signaling pathways.10 The utilization of multiple drugs in cancer therapy simultaneously targets various pathways; yet, this approach may result in detrimental drug–drug interactions and the development of drug resistance.11 As a result, using a single treatment that can influence more than one target is considered a different approach to overcoming the aforementioned limitations.12,13 The FDA has authorized several multitarget therapies for cancer, including Dasatinib, a highly effective multitargeted kinase inhibitor.14
BRAFV600E is a somatic point mutation that converts thymine to adenine at nucleotide 1799. This modification results in the substitution of valine (V) with glutamic acid (E) at codon 600 inside the kinase domain. This alteration renders the BRAF protein a monomer that is perpetually active and autonomously transmits signals, independent of prior RAS activation. The V600E mutation induces uncontrolled cellular proliferation, evasion of apoptosis, and the emergence of an aggressive oncogenic phenotype by perpetually activating the mitogen-activated protein kinase (MAPK/ERK) pathway. In colorectal cancer (CRC), this mutation serves as a critical molecular marker for a distinct clinical cohort characterized by right-sided primary tumors, poor differentiation, and significantly reduced overall survival rates compared to BRAF wild-type tumors.15,16
Although BRAF inhibitors are effective in other V600E-mutant cancers, they are not effective as monotherapy in colorectal cancer due to the disease's rapid adaptation.17 Colorectal cancer cells, conversely, react to BRAF inhibition by initiating a feedback loop that elevates the levels of epidermal growth factor receptor (EGFR). This reactivation via EGFR circumvents the obstructed BRAF protein and reinstates signaling through the MAPK/ERK pathway, hence sustaining tumor proliferation and viability.18,19 To achieve a durable therapeutic blockade, it is essential to utilize both a BRAF inhibitor and an EGFR inhibitor. This combination therapy addresses intrinsic resistance by simultaneously targeting both the mutant driver and its primary escape mechanism.15 This results in improved clinical outcomes and establishes a new standard of care for this challenging patient population.
Pyridine derivatives constitute a significant category of anticancer treatments, featuring multiple FDA-approved medications (e.g., Sorafenib, Vemurafenib) and a multitude of investigational compounds designed as EGFR and BRAF inhibitors.20,21 The pyridine scaffold is a crucial structural element of numerous kinase inhibitors because of its advantageous electronic properties and ability to form robust interactions within the ATP-binding sites of target enzymes.22 Sorafenib (Compound I, Fig. 1) is an FDA-approved pyridine derivative used for advanced renal cell carcinoma and hepatocellular carcinoma, functioning as a multikinase inhibitor targeting VEGFR and BRAF.23 Vemurafenib (Compound II, Fig. 1), a pyridine derivative, is approved to treat metastatic melanoma by inhibiting the V600E-mutated BRAF kinase specifically.24 Researchers are continually developing novel pyridine derivatives, frequently as hybrid compounds, to target both EGFR and BRAF pathways. This dual-targeting strategy seeks to enhance efficacy and diminish resistance mechanisms.25,26
Meanwhile, literature reviews demonstrate that 1,2,4-oxadiazoles hold considerable significance in bioorganic and medicinal chemistry. They are recognized for their diverse pharmacological characteristics.27,28 The 1,2,4-oxadiazole exhibits bioisosteric equivalence to ester and amide features. Under unstable conditions (e.g., hydrolysis), 1,2,4-oxadiazole offers a highly effective alternative.29 The substantial biological impact of 1,2,4-oxadiazole derivatives on cancer cells is due to multiple mechanisms of action. We recently30 displayed a group of 1,2,4-oxadiazole/quinazoline-4-one hybrids designed as antiproliferative agents targeting EGFR/BRAFV600E. The results indicated that compound III (Fig. 2) exhibited the highest antiproliferative activity. Compound III demonstrated significant efficacy as an EGFR and BRAFV600E inhibitor, with pronounced apoptotic action.
In a separate publication from our laboratory,29 we reported the synthesis of a novel class of dihydropyrimidine-5-carbonitrile/1,2,4-oxadiazole hybrids that serve as dual inhibitors of EGFR and VEGFR-2 and exhibit antioxidant properties. Compound IV (Fig. 2) exhibited the most significant antiproliferative activity, exceeding the efficacy of erlotinib against Panc-1 (pancreatic) and MCF-7 (breast) cancer cell lines. Compound IV had the highest inhibitory action against EGFR and VEGFR-2, with IC50 values of 57 nM for EGFR and 21 nM for VEGFR-2. Additionally, compound IV exhibited significant apoptotic activity by upregulating caspases-3, 8, and 9, along with Bax and p53, and downregulating the anti-apoptotic protein Bcl-2.
In light of the previously discussed anticancer attributes of pyridine and 1,2,4-oxadiazole derivatives as EGFR and/or BRAFV600E inhibitors, and to advance our goal of developing novel inhibitors that target both EGFR and BRAFV600E,15,20,31–35 here, we detail the development, and antiproliferative testing of a handful of novel compounds (20a–c, 21a–c, and 22a–c, Fig. 3) that simultaneously targeting EGFR and BRAFV600E.
The molecular structure was developed using a dual-binding approach that included various pharmacophores tailored to the conserved and divergent characteristics of the target kinases. The aryl-pyridine core was selected as the primary hinge-binding motif to facilitate anchoring interactions within the ATP-binding pockets of both EGFR and BRAFV600E. A 1,2,4-oxadiazole moiety functions as a rigid linker to ensure optimal spatial configuration. It provides the appropriate configuration to link the hinge and supplementary binding regions. Finally, a substituted phenyl tail was included to enhance lipophilicity and accommodate the configuration of distal hydrophobic areas, particularly the “back pocket” of BRAF, hence augmenting potency and selectivity.
The structural confirmation of the novel compounds was achieved using 1H NMR, 13C NMR, elemental analysis, and LC-MS for selected typical instances. The new compounds will be assessed for antiproliferative activity using the MTT assay against four different cancer cell lines. The most effective compounds will be evaluated as dual EGFR/BRAFV600E inhibitors. Additionally, several selected compounds will be evaluated for their effects on normal cell lines, and their apoptotic and immunomodulatory capabilities. Ultimately, docking research will be employed to examine the binding interactions of some selected derivatives with the designated receptor sites.
2. Experimental
2.1. Chemistry
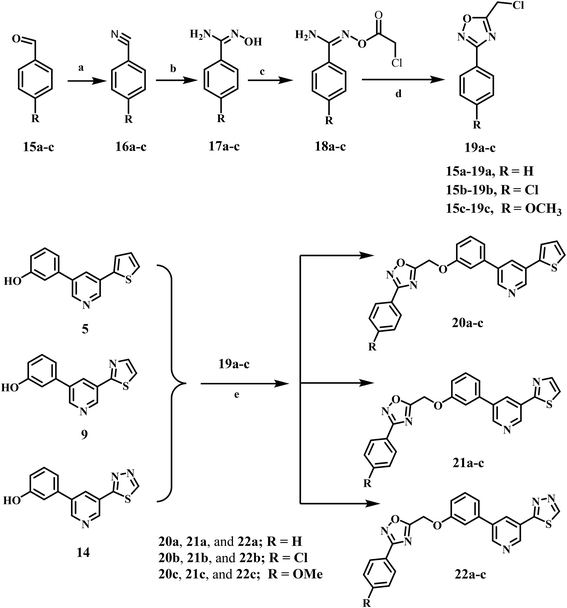
General details: see Appendix A (SI).The starting materials, 3-aryl-5-(chloromethyl)-1,2,4-oxadiazoles 19a–c, were prepared according to literature methods.30
2.1.9.1. 3-Phenyl-5-((3-(5-(thiophen-2-yl)pyridin-3-yl)phenoxy)methyl)-1,2,4-oxadiazole (20a). Yield: 0.17 g (69%), brown solid, m.p: 157–159 °C. 1H NMR (400 MHz, δ ppm CDCl3): 8.89 (d, J = 2.1 Hz, 1H, Ar–H), 8.75 (d, J = 2.1 Hz, 1H, Ar–H), 8.13 (dd, J = 7.8, 1.5 Hz, 1H, Ar–H), 8.02 (t, J = 2.1 Hz, 1H, Ar–H), 7.52–7.47 (m, 4H, Ar–H), 7.44 (dd, J = 3.5, 1.2 Hz, 1H, Ar–H), 7.41 (dd, J = 5.2, 1.2 Hz, 1H, Ar–H), 7.34–7.30 (m, 2H, Ar–H), 7.28 (s, 1H, Ar–H), 7.17 (dd, J = 5.2, 3.5 Hz, 1H, Ar–H), 7.11 (ddd, J = 8.3, 2.4, 1.2 Hz, 1H, Ar–H), 5.46 (s, 2H, O–CH2); 13C NMR (100 MHz, δ ppm CDCl3): 174.54, 168.61, 158.16, 146.92, 146.04, 140.13, 139.39, 138.34, 137.73, 136.14, 131.52, 130.58, 130.42, 128.35, 126.25, 126.09, 124.76, 124.59, 121.32, 114.38, 114.28, 61.29; Anal. Calc. (%) For C24H17N3O2S: C, 70.06; H, 4.16; N, 10.21; S, 7.79. Found: C, 70.15; H, 4.20; N, 10.32; S, 7.71.
2.1.9.2. 3-(4-Chlorophenyl-5-((3-(5-(thiophen-2-yl)pyridin-3-yl)phenoxy)methyl)-1,2,4-oxadiazole (20b). Yield: 0.20 g (75%), brown solid, m.p: 160–162 °C. 1H NMR (400 MHz, δ ppm CDCl3): 8.89 (d, J = 2.2 Hz, 1H, Ar–H), 8.75 (d, J = 2.2 Hz, 1H, Ar–H), 8.07 (d, J = 8.7 Hz, 2H, Ar–H 4-Cl C6H5), 8.02 (t, J = 2.2 Hz, 1H, Ar–H), 7.49 (d, J = 8.7 Hz, 2H, Ar–H 4-Cl C6H5), 7.44 (dd, J = 3.6, 1.2 Hz, 1H, Ar–H), 7.41 (dd, J = 5.1, 1.1 Hz, 1H, Ar–H), 7.35–7.32 (m, 2H, Ar–H), 7.28 (s, 1H, Ar–H), 7.17 (dd, J = 5.1, 3.5 Hz, 1H, Ar–H), 7.11 (ddd, J = 8.3, 2.4, 1.1 Hz, 1H, Ar–H), 5.46 (s, 2H, O–CH2); 13C NMR (100 MHz, δ ppm CDCl3): 174.83, 167.89, 158.12, 146.94, 146.04, 140.14, 139.40, 137.72, 136.15, 131.47, 130.54, 130.44, 129.31, 128.89, 128.39, 126.32, 124.76, 124.56, 121.39, 114.34, 114.25, 61.20; anal. calc. (%) For C24H16ClN3O2S: C, 64.64; H, 3.62; N, 9.42; S, 7.19. Found: C, 64.72; H, 3.67; N, 9.40; S, 7.10.
2.1.9.3. 3-(4-Methoxyphenyl-5-((3-(5-(thiophen-2-yl)pyridin-3-yl)phenoxy)methyl)-1,2,4-oxadiazole (20c). Yield: 0.18 g (68%), brown solid, m.p: 165–167 °C. 1H NMR (400 MHz, δ ppm DMSO-d6): 9.32 (s, 1H, Ar–H), 8.98 (s, 1H, Ar–H), 8.43 (s, 1H, Ar–H), 7.98 (d, J = 8.8 Hz, 2H, Ar–H), 7.68 (dd, J = 6.0, 1.5 Hz, 1H, Ar–H), 7.61 (dd, J = 6.0, 1.5 Hz, 1H, Ar–H), 7.33 (t, J = 7.8 Hz, 1H, Ar–H), 7.26 (d, J = 7.8 Hz, 1H, Ar–H), 7.20 (d, J = 5.5 Hz, 1H, Ar–H), 7.17 (d, J = 8.8 Hz, 2H, Ar–H), 7.10 (s, 1H, Ar–H), 6.82 (d, J = 7.8 Hz, 1H, Ar–H), 5.61 (s, 2H, O–CH2), 3.81 (s, 3H, O–CH3); 13C NMR (100 MHz, δ ppm DMSO-d6): 174.80, 167.86, 158.10, 146.95, 146.03, 140.15, 139.42, 137.75, 136.14, 131.48, 130.53, 130.44, 128.67, 128.38, 126.32, 124.74, 124.53, 121.36, 114.30, 114.21, 113.34, 61.19, 55.39; anal. calc. (%) For C25H19N3O3S: C, 68.01; H, 4.34; N, 9.52; S, 7.26. Found: C, 68.13; H, 4.30; N, 9.60; S, 7.18.
2.1.9.4. 3-Phenyl-5-((3-(5-(thiazol-2-yl)pyridin-3-yl)phenoxy)methyl)-1,2,4-oxadiazole (21a). Yield: 0.18 g (73%), Yellow solid, m.p: 160–162 °C. 1H NMR (400 MHz, δ ppm CDCl3): 9.16 (d, J = 2.0 Hz, 1H, Ar–H), 8.97 (d, J = 2.0 Hz, 1H, Ar–H), 8.55 (t, J = 2.0 Hz, 1H, Ar–H), 8.12 (dd, J = 7.7, 1.5 Hz, 1H, Ar–H), 7.98 (d, J = 3.3 Hz, 1H, Ar–H), 7.52–7.49 (m, 4H, Ar–H), 7.46 (d, J = 3.3 Hz, 1H, Ar–H), 7.35 (dd, J = 7.4, 1.2 Hz, 2H, Ar–H), 7.27 (s, 1H, Ar–H), 7.16–7.12 (m, 1H, Ar–H), 5.46 (s, 2H, O–CH2); 13C NMR (100 MHz, δ ppm CDCl3): 174.41, 168.60, 164.99, 158.22, 156.58, 151.70, 150.54, 147.85, 138.31, 136.58, 133.29, 131.50, 130.70, 128.94, 127.55, 126.23, 126.05, 121.31, 114.76, 114.21, 61.20; anal. calc. (%) For C23H16N4O2S: C, 66.98; H, 3.91; N, 13.58; S, 7.77. Found: C, 66.72; H, 3.98; N, 13.49; S, 7.82.
2.1.9.5. 3-(4-Chlorophenyl-5-((3-(5-(thiazol-2-yl)pyridin-3-yl)phenoxy)methyl)-1,2,4-oxadiazole (21b). Yield: 0.19 g (72%), yellow solid, m.p: 168–170 °C. 1H NMR (400 MHz, δ ppm CDCl3): 9.18 (d, J = 2.2 Hz, 1H, Ar–H), 8.90 (d, J = 2.2 Hz, 1H, Ar–H), 8.46 (t, J = 2.2 Hz, 1H, Ar–H), 8.08 (d, J = 8.6 Hz, 2H, Ar–H 4-Cl C6H5), 7.98 (d, J = 3.2 Hz, 1H, Ar–H), 7.49 (d, J = 8.6 Hz, 2H, Ar–H 4-Cl C6H5), 7.47 (d, J = 3.2 Hz, 1H, Ar–H), 7.37 (dd, J = 4.0, 1.8 Hz, 2H, Ar–H), 7.29 (s, 1H, Ar–H), 7.12 (dd, J = 7.9, 3.1 Hz, 1H, Ar–H), 5.46 (s, 2H, O–CH2); 13C NMR (100 MHz, δ ppm CDCl3): 174.78, 167.89, 164.65, 158.16, 149.21, 146.74, 144.22, 138.92, 137.71, 136.30, 131.95, 130.59, 129.65, 129.30, 128.90, 124.78, 121.40, 119.88, 114.57, 114.23, 61.20; Anal. Calc. (%) For C23H15ClN4O2S: C, 61.81; H, 3.38; N, 12.54; S, 7.17. Found: C, 61.78; H, 3.32; N, 12.46; S, 7.30.
2.1.9.6. 3-(4-Methoxyphenyl-5-((3-(5-(thiazol-2-yl)pyridin-3-yl)phenoxy)methyl)-1,2,4-oxadiazole (21c). Yield: 0.19 g (73%), yellow solid, m.p: 176–178 °C. 1H NMR (400 MHz, δ ppm DMSO-d6): 9.32 (s, 1H, Ar–H), 8.98 (s, 1H, Ar–H), 8.43 (s, 1H, Ar–H), 7.98 (d, J = 8.6 Hz, 2H, Ar–H 4-OCH3 C6H4), 7.59 (d, J = 5.4 Hz, 1H, Ar–H), 7.44 (d, J = 5.4 Hz, 1H, Ar–H), 7.33 (t, J = 7.7 Hz, 1H, Ar–H), 7.26 (d, J = 7.8 Hz, 1H, Ar–H), 7.17 (d, J = 8.6 Hz, 2H, Ar–H 4-OCH3 C6H4), 7.10 (s, 1H, Ar–H), 6.82 (d, J = 7.8 Hz, 1H, Ar–H), 5.61 (s, 2H, O–CH2), 3.81 (s, 3H, O–CH3); 13C NMR (100 MHz, δ ppm DMSO-d6): 174.76, 167.88, 164.63, 158.16, 149.20, 146.75, 144.23, 138.91, 137.73, 136.32, 131.90, 130.57, 129.66, 128.69, 124.80, 121.43, 119.86, 114.84, 114.54, 114.20, 113.34, 61.20, 55.38; anal. calc. (%) For C24H18N4O3S: C, 65.15; H, 4.10; N, 12.66; S, 7.25. Found: C, 65.18; H, 4.03; N, 12.75; S, 7.39.
2.1.9.7. 5-((3-(5-(1,3,4-Thiadiazol-2yl)pyridin-3-yl)phenoxy)methyl)-3-phenyl-1,2,4-oxadiazole (22a). Yield: 0.17 g (70%), orange solid, m.p: 165–167 °C. 1H NMR (400 MHz, δ ppm CDCl3): 9.23 (s, 1H, Ar–H), 9.17 (d, J = 2.2 Hz, 1H, Ar–H), 8.98 (d, J = 2.2 Hz, 1H, Ar–H), 8.56 (t, J = 2.2 Hz, 1H, Ar–H), 8.13 (dd, J = 7.8, 1.6 Hz, 1H, Ar–H), 7.57–7.46 (m, 4H, Ar–H), 7.36 (dd, J = 7.5, 1.3 Hz, 2H, Ar–H), 7.28 (s, 1H, Ar–H), 7.17–7.13 (m, 1H, Ar–H), 5.47 (s, 2H, O–CH2); 13C NMR (100 MHz, δ ppm CDCl3): 174.49, 168.66, 165.03, 158.26, 151.76, 150.56, 147.88, 138.34, 136.62, 133.31, 131.52, 130.72, 128.96, 127.58, 126.25, 126.09, 121.33, 114.81, 114.29, 61.23; anal. calc. (%) For C22H15N5O2S: C, 63.91; H, 3.66; N, 16.94; S, 7.75. Found: C, 64.03; H, 3.71; N, 16.99; S, 7.68.
2.1.9.8. 5-((3-(5-(1,3,4-Thiadiazol-2-yl)pyridin-3-yl)phenoxy)methyl)-3-(4-chlorophenyl)-1,2,4-oxadiazole (22b). Yield: 0.20 g (75%), orange solid, m.p: 173–175 °C. 1H NMR (400 MHz, δ ppm CDCl3): 9.24 (s, 1H, Ar–H), 9.17 (d, J = 2.1 Hz, 1H, Ar–H), 8.98 (d, J = 2.1 Hz, 1H, Ar–H), 8.58 (t, J = 2.1 Hz, 1H, Ar–H), 8.08 (d, J = 8.8 Hz, 2H, Ar–H 4-Cl C6H5), 7.50 (d, J = 8.8 Hz, 2H, Ar–H 4-Cl C6H5), 7.39–7.36 (m, 2H, Ar–H), 7.28 (s, 1H, Ar–H), 7.14 (ddd, J = 8.2, 2.4, 1.0 Hz, 1H, Ar–H), 5.47 (s, 2H, O–CH2); 13C NMR (100 MHz, δ ppm CDCl3): 174.71, 167.91, 164.99, 158.22, 151.74, 150.56, 147.94, 138.38, 137.74, 136.60, 133.26, 130.74, 129.32, 128.90, 126.12, 124.75, 121.38, 114.83, 114.25, 61.18; anal. calc. (%) For C22H14ClN5O2S: C, 59.00; H, 3.15; N, 15.64; S, 7.16. Found: C, 59.08; H, 3.24; N, 15.57; S, 7.10.
2.1.9.9. 5-((3-(5-(1,3,4-Thiadiazol-2-yl)pyridin-3-yl)phenoxy)methyl)-3-(4-methoxyphenyl)-1,2,4-oxadiazole (22c). Yield: 0.19 g (72%), orange solid, m.p: 182–184 °C. 1H NMR (400 MHz, δ ppm DMSO-d6): 9.73 (s, 1H, Ar–H), 9.32 (s, 1H, Ar–H), 8.98 (s, 1H, Ar–H), 8.43 (s, 1H, Ar–H), 7.98 (d, J = 8.5 Hz, 2H, Ar–H 4-OCH3 C6H4), 7.33 (t, J = 7.7 Hz, 1H, Ar–H), 7.26 (d, J = 7.8 Hz, 1H, Ar–H), 7.17 (d, J = 8.5 Hz, 2H, Ar–H 4-OCH3 C6H4), 7.10 (s, 1H, Ar–H), 6.82 (d, J = 7.9 Hz, 1H, Ar–H), 5.61 (s, 2H, O–CH2), 3.81 (s, 3H, O–CH3); 13C NMR (100 MHz, δ ppm DMSO-d6): 174.72, 167.92, 165.00, 158.73, 155.41, 150.30, 147.46, 137.74, 136.74, 133.44, 130.83, 128.68, 126.36, 121.37, 118.31, 116.31, 114.84, 114.46, 113.35, 61.18, 55.38; anal. calc. (%) For C23H17N5O3S: C, 62.29; H, 3.86; N, 15.79; S, 7.23. Found: C, 62.35; H, 3.77; N, 15.85; S, 7.18.
2.2. Biology
3. Results and discussion
3.1. Chemistry
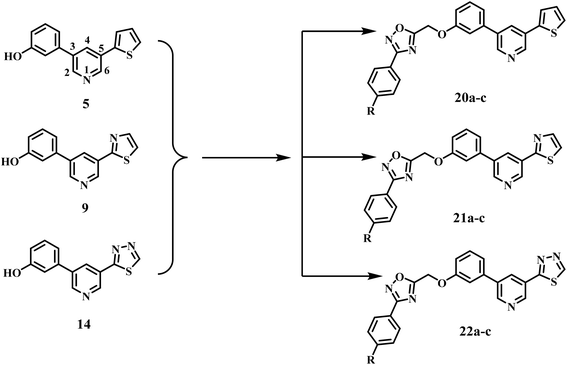
A convergent method for synthesising intermediate 5 (thiophene–pyridine–phenol) was achieved via two successive Suzuki–Miyaura cross-couplings, utilising carefully selected catalyst/ligand complexes to control regioselectivity and functional-group tolerance (Scheme 1). Starting with 3,5-dibromopyridine 1, we decided to insert the thiophene ring via coupling with 2-thienylboronic acid 2. To suppress the formation of the undesired symmetrically disubstituted (bis-coupled) analogue, several experimental conditions were tried using different palladium catalysts and ligand sets. Under these optimized conditions, tetrakis(triphenylphosphine)palladium (Pd(PPh3)4 and cesium carbonate (Cs2CO3) in 1,4-dioxane at 100 °C, we obtain 3-bromo-5-(thiophen-2-yl)pyridine 3 in 40% yield and recover the unreacted dibromopyridine without observing the formation of the bis-adduct byproduct. It is worth noting that an increase in the reaction temperature to more than 120 °C and/or employing two equivalents of the starting 2-thienylboronic 2 led to the formation of the bis-coupling product. The structure of compound 3 was confirmed using the spectral (1H NMR and MS) data. The 1H NMR spectrum exhibited three new aromatic signals at 7.76–7.70 and 7.22–7.19 ppm assigned to the protons of the thiophene ring.In the next step, we employed [1,1′-bis(diphenylphosphino)ferrocene]palladium(II) dichloride (Pd(dppf)Cl2) as a catalyst in presence of Cs2CO3 as a base for the cross-coupling of intermediate 3 with (3-hydroxyphenyl)boronic acid 4. The reaction proceeded in a 1,4-dioxane/H2O system at 90 °C, yielding the target scaffold 5 at 63% efficiency. When the reaction was conducted under the same conditions but without the aqueous component, similar product was obtained in slightly lower yield. The bidentate ligand 1,1′-bis(diphenylphosphino)ferrocene (dppf), together with the dioxane/aqueous medium, proved optimal for trans-metalation from a phenylboronic acid bearing a free phenolic group. The 1H NMR spectrum of compound 5 displayed four new signals in the aromatic region at δ 7.33, 7.23–7.20, 7.15, and 6.88–6.86 ppm, corresponding to the protons of the phenyl ring, along with a phenolic OH signal at δ 9.66 ppm.
Key intermediate 9 (thiazole–pyridine–phenol) was obtained in the same fashion using two Suzuki–Miyaura cross-couplings designed to retain chemo-selectivity and functional-group tolerance (Scheme 1). In the first step, 2-bromothiazole 6 was coupled with 5-bromopyridine-3-boronic acid 7 under Pd(PPh3)4/Cs2CO3 in 1,4-dioxane at 100 °C, furnishing 2-(5-bromopyridin-3-yl)thiazole 8 in 24% yield. It should be noted that the use of an electron-rich phosphine catalyst with a non-nucleophilic carbonate base enabled the trans-metalation from the pyridinyl boronic acid while leaving the C–Br bond on the pyridine ring intact for subsequent derivatization. Next, compound 8 was coupled with (3-hydroxyphenyl)boronic acid 4 using Pd(PPh3)4 and K2CO3 in a mixed 1,4-dioxane/H2O medium at 90 °C to furnish the target scaffold, 3-(5-(thiazol-2-yl)pyridin-3-yl)phenol 9, in 46% yield. The 1H NMR spectrum of compound 9 displayed four new signals in the aromatic region at δ 7.35, 7.24, 7.16, and 6.89–6.87 ppm, corresponding to the protons of the phenyl ring, along with a phenolic OH signal at δ 9.69 ppm.
Finally, Scheme 1 outlines the synthetic route for key intermediate 14 (thiadiazole–pyridine–phenol). The route began with the hydrazinolysis of methyl 5-bromonicotinate 10 with hydrazine monohydrate in ethanol at 60 °C, yielding hydrazide 11 in quantitative yield. Next, formylation of hydrazide 11 with formic acid gave N-formyl hydrazide intermediate 12. The 1H NMR spectrum of compound 12 exhibited a signal at δ 8.19 ppm due to the CH proton of the formyl group. Subsequent cyclization of N-formyl hydrazide 12 using phosphorus pentasulfide (P2S5) in pyridine resulted in the formation of a 1,3,4-thiadiazole derivative 13. In this reaction, phosphorus pentasulfide functions as both a thionating and dehydrating agent, converting the carbonyl oxygen to sulfur and facilitating the subsequent intramolecular cyclization. The 1H NMR spectrum confirmed the formation of thiadiazole derivatives 13. The formyl CH protons were not detected. Instead, the C–H peak of the thiadiazole ring was observed in the aromatic region at δ 9.76 ppm. The key intermediate 14 was synthesized via a Suzuki–Miyaura cross-coupling reaction of compound 13 with the corresponding (3-hydroxyphenyl)boronic acid 4, employing Pd(PPh3)4 and K2CO3 as the catalyst in 1,4-dioxane at 80 °C. The 1H NMR spectrum of compound 14 displayed four new signals in the aromatic region at δ 7.35, 7.27–7.25, 7.19, and 6.91–6.88 ppm, corresponding to the protons of the phenyl ring, along with a phenolic OH signal at δ 9.76 ppm.
Comparatively, amidoxime intermediates 17a–c, were synthesized in two steps with yields ranging from 50 to 60%. The first stage involves reacting the various aldehydes (15a–c) with 28% liquid ammonia and iodine in THF for two to three hours. This produces the aryl nitriles 16a–c in 76–80% yields.45 The second stage in the synthesis of 17a–c involved refluxing 16a–c in methanol with hydroxylamine chloride and sodium carbonate for 12 to 18 h. Benzimidamides 18a–c were synthesized by reacting 17a–c with chloroacetyl chloride in anhydrous acetone. The compounds were then cyclized by refluxing in toluene, resulting in the formation of 3-aryl-5-(chloromethyl)-1,2,4-oxadiazoles 19a–c as a yellow oil,30 Scheme 2. Oxadiazoles 19a–c were purified by column chromatography utilizing a hexane: ethyl acetate (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) eluent.
1) eluent.
The novel compounds 20a–c, 21a–c, and 22a–c were synthesized by reacting intermediates 5, 9, and 14 with oxadiazoles 19a–c in DMF, using K2CO3 and KI as catalysts, as depicted in Scheme 2. The reaction mixture was stirred overnight and then poured onto crushed ice. The formed precipitate was recrystallized from ethanol, yielding pure 20a–c, 21a–c, and 22a–c. The structures of 20a–c, 21a–c, and 22a–c were validated through 1H NMR, 13C NMR, and elemental microanalysis. The 1H NMR spectrum of compound 22c exhibited two distinct singlet signals: one at δ 5.61 ppm, attributed to two protons of the O–CH2 group, and another at δ 3.81 ppm, corresponding to three protons of the p-methoxy group. The spectrum additionally exhibited two doublets at 7.98 (d, J = 8.5 Hz, 2H) and 7.17 (d, J = 8.5 Hz, 2H) corresponding to a p-disubstituted benzene ring. The 13C NMR spectrum of 22c exhibited two signals at δ 61.18 and 55.38 ppm, corresponding to OCH2 and OCH3, respectively. Furthermore, all aromatic signals manifested as anticipated.
3.2. Biology
| Comp. | Cell viability% | Antiproliferative activity IC50 ± SEM (nM) | ||||
|---|---|---|---|---|---|---|
| A-549 | MCF-7 | Panc-1 | HT-29 | Average (mean IC50) | ||
| a ND: not determined. | ||||||
| 5 | 89 | 64 ± 5 | 69 ± 5 | 67 ± 5 | 67 ± 5 | 67 |
| 9 | 90 | 61 ± 5 | 66 ± 5 | 64 ± 5 | 63 ± 5 | 64 |
| 14 | 91 | 69 ± 5 | 74 ± 5 | 72 ± 6 | 71 ± 6 | 72 |
| 20a | 91 | 49 ± 4 | 55 ± 5 | 53 ± 5 | 50 ± 4 | 52 |
| 20b | 90 | 45 ± 3 | 49 ± 3 | 47 ± 3 | 46 ± 3 | 47 |
| 20c | 92 | 23 ± 1 | 26 ± 1 | 24 ± 1 | 23 ± 1 | 24 |
| 21a | 89 | 31 ± 2 | 35 ± 2 | 34 ± 2 | 32 ± 2 | 33 |
| 21b | 90 | 27 ± 1 | 31 ± 2 | 29 ± 1 | 28 ± 1 | 29 |
| 21c | 91 | 20 ± 1 | 23 ± 1 | 21 ± 1 | 21 ± 1 | 21 |
| 22a | 91 | 41 ± 3 | 47 ± 3 | 45 ± 3 | 43 ± 3 | 44 |
| 22b | 90 | 36 ± 2 | 39 ± 2 | 38 ± 2 | 36 ± 2 | 38 |
| 22c | 92 | 53 ± 5 | 58 ± 5 | 56 ± 5 | 56 ± 5 | 56 |
| Erlotinib | ND | 30 ± 3 | 40 ± 3 | 30 ± 3 | 30 ± 3 | 33 |
Generally, compounds 5, 9, 14, 20a–c, 21a–c, and 22a–c exhibited significant antiproliferative activity, with mean IC50 values ranging from 21 to 72 nM across the four cancer cell lines, compared with the reference Erlotinib, which had a mean IC50 of 33 nM. Furthermore, all evaluated compounds demonstrate greater affinity for the lung (A-549) and colon (HT-29) cancer cell lines than for the other cell lines examined. Compounds 20c, 21a–c, and 22b exhibited the most significant antiproliferative activity, with mean IC50 values ranging from 21 to 38 nM. Compounds 20c, 21b, and 21c exhibited superior efficacy than erlotinib against the A-549 lung cancer cell line, with IC50 values of 23, 27, and 20 nM, respectively, compared to 30 nM for Erlotinib. Furthermore, derivatives 20c, 21b, and 21c demonstrated enhanced activity relative to erlotinib against the HT-29 (colon) cancer cell line. Their IC50 values were 23, 28, and 21 nM, respectively, whereas erlotinib demonstrated an IC50 value of 30 nM.
Compound 21c (Thiazole scaffold, R = 4-OMe) outperformed all other evaluated compounds. It had a mean IC50 of 21 nM, therefore being 1.5-fold more effective than erlotinib (mean IC50 = 33 nM) against the four cancer cell lines studied. Compound 21c showed significant antiproliferative activity against the A-549 lung cancer cell line, with an IC50 of 20 nM, which is 1.5 times more effective than erlotinib's IC50 of 30 nM. Furthermore, compound 21c exhibits a potency 1.4-fold superior to erlotinib against pancreatic (Panc-1) and colon (HT-29) cancer cell lines, and 1.7-fold superior against the breast (MCF-7) cancer cell line, as indicated in Table 1.
The structural features of the heterocyclic component at the fifth position of the pyridine ring (thiophene, thiazole, or thiadiazole), as well as the substitution pattern at the fourth position of the phenyl group in the 1,2,4-oxadiazole moiety, have a significant impact on the novel compounds' antiproliferative activity. For instance, compounds 20c (thiophene scaffold, R = 4-OMe) and 22c (thiadiazole scaffold, R = 4-OMe), which possess thiophene and thiadiazole groups at the fifth position of the pyridine moiety, exhibited decreased efficacy as antiproliferative agents relative to the thiazole derivative, 21c (thiazole scaffold, R = 4-OMe). Compound 20c demonstrated a mean IC50 of 24 nM, signifying a potency comparable to that of 21c (mean IC50 = 21 nM), while compound 22c displayed a mean IC50 value of 56 nM, rendering it 2.7-fold less potent than compound 21c. The data demonstrate that both thiophene and thiazole groups at the 5-position of the pyridine moiety are more advantageous for antiproliferative activity than the thiadiazole group, with activity increasing in the sequence thiazole > thiophene > thiadiazole. Compound 20c (Thiophene scaffold, R = 4-OMe) had the second–highest activity and demonstrated more potency than erlotinib across the four cancer cell lines evaluated, as shown in Table 1.
Additionally, the substitution pattern at the para position of the phenyl group in the 1,2,4-oxadiazole moiety may substantially affect the antiproliferative activity of the newly developed compounds. Compounds 21a (thiazole scaffold, R = H) and 21b (Thiazole scaffold, R = 4-Cl), possessing identical structural attributes to 21c but featuring an unsubstituted phenyl group in 21a or a p-chlorophenyl group in 21b, had IC50 values of 33 nM and 29 nM, respectively. Compound 21a exhibited a GI50 value of 33 nM, making it 1.6-fold less effective than 21c. The observations suggest that an unsubstituted phenyl group in the 1,2,4-oxadiazole structure is unfavorable to antiproliferative efficacy. Compound 21b, a p-chloro-phenyl derivative of the 1,2,4-oxadiazole moiety, exhibited a mean IC50 value of 29 nM, which was slightly less potent than the p-methoxy derivative, 21c (mean IC50 = 21 nM). This suggests that substitution with either an electron-withdrawing group (chlorine) or an electron-donating group (methoxy) enhances antiproliferative activity, with the methoxy group demonstrating superior efficacy.
Compounds 5 (thiophene-based derivative), 9 (thiazole-based derivative), and 14 (thiadiazole-based derivative), which possess free phenolic hydroxyl groups unassociated with the phenyl-1,2,4-oxadiazole moiety, were identified as the least potent derivatives, exhibiting mean IC50 values of 67, 64, and 72 nM, respectively. These values are at least threefold less effective than compound 21c and approximately twofold less potent than erlotinib. The data indicated the potential impact of the 1,2,4-oxadiazole moiety on antiproliferative activity.
| Compound | EGFR inhibition IC50 ± SEM (nM) | BRAFV600E inhibition IC50 ± SEM (nM) |
|---|---|---|
| 20c | 71 ± 4 | 49 ± 2 |
| 21a | 82 ± 5 | 61 ± 4 |
| 21b | 76 ± 4 | 54 ± 3 |
| 21c | 64 ± 3 | 41 ± 2 |
| 22b | 87 ± 5 | 69 ± 4 |
| Erlotinib | 80 ± 5 | — |
| Vemurafenib | — | 30 ± 2 |
The results of this assay correspond with those of the antiproliferative assay, demonstrating that compounds 20c (thiophene scaffold, R = 4-OMe) and 21c (thiazole scaffold, R = 4-OMe), recognized as the most potent antiproliferative agents, were the most effective derivatives of EGFR inhibitors, displaying IC50 values of 71 ± 4 and 64 ± 3 nM, respectively. They demonstrated 1.2- and 1.3-fold greater efficacy than erlotinib (IC50 = 80 nM). Moreover, compounds 21a (Thiazole scaffold, R = H) and 21b (thiazole scaffold, R = 4-Cl) have significant EGFR inhibitory action, with IC50 values of 82 and 76 nM, respectively, demonstrating equivalent potency to the reference erlotinib. Ultimately, compound 22b (thiadiazole scaffold, R = 4-Cl) exhibited a slightly reduced EGFR inhibitory efficacy relative to erlotinib, with an IC50 value of 87 nM. The data from these in vitro investigations suggested that compounds 20c, 21b, and 21c were effective antiproliferative agents, potentially acting as EGFR inhibitors.
Compound 21c, the most efficacious antiproliferative and EGFR inhibitor, demonstrated the highest potency as a BRAFV600E inhibitor, with an IC50 of 41 nM, which is 1.4-fold lower than vemurafenib. The results indicated that compounds 20c, 21b, and 21c function as antiproliferative agents targeting both EGFR and mutant BRAF. Nevertheless, they warrant structural modifications to enhance their effectiveness against their molecular targets.
| Compound | Cytotoxicity (WI-38) IC50 (nM) | Selectivity index (SI) | |
|---|---|---|---|
| HCT-116 | A-549 | ||
| 20c | > 200 | > 8 | > 8 |
| 21c | > 200 | > 10 | > 10 |
Additionally, the dual inhibition of EGFR and BRAFV600E efficiently inhibits the MAPK signaling pathway, and the therapeutic benefit of this combination is increasingly being related to its function in restoring the tumor immune milieu. Dual inhibition greatly reduces the release of major pro-inflammatory cytokines such as IL-6 and TNF-α. These cytokines promote bypass signaling via the STAT3 pathway and increase the recruitment of immunosuppressive myeloid cells; thus, studying their regulation is critical for understanding how dual inhibitors overcome initial resistance and restore anti-tumor immunity.
The impact of the most active compounds 20c and 21c on the levels of immunomodulatory proteins (TNF-α and IL-6) was assessed using a qRT-PCR assay test.51 A-549 cells were administered compounds 20c and 21c for 24 hours at concentrations of 23 nM and 20 nM (IC50 against A-549), respectively. The reference molecule was dexamethasone, a pharmaceutical agent that consistently modulates the immune system.
Compounds 20c and 21c had substantial immunomodulatory effects, reducing the production of pro-inflammatory cytokines TNF-α and IL-6 by 86% and 96%, respectively (Table 4). This degree of inhibition is particularly noteworthy since it surpasses the efficacy of the potent corticosteroid Dexamethasone (83% and 93% inhibition). The superior efficacy of 21c demonstrates that concurrently inhibiting both EGFR and BRAFV600E effectively halts the ERK-dependent activation of NF-κB and AP-1, hence dismantling the cytokine-mediated survival niche that resistant tumor cells frequently exploit for sustenance.
| Compound | TNF-α (% inhibition) | IL-6 (% inhibition) |
|---|---|---|
| 20c | 79 | 89 |
| 21c | 86 | 96 |
| Dexamethasone | 83 | 93 |
| Compound no. | Bcl-2 (ng mL−1) | Fold reduction | Bax (pg mL−1) | Fold change | p53 (pg mL−1) | Fold change |
|---|---|---|---|---|---|---|
| 20c | 1.70 ± 0.001 | 3 | 485 ± 2 | 8 | 330 ± 2 | 5 |
| 21c | 1.25 ± 0.001 | 4 | 550 ± 3 | 9 | 450 ± 3 | 7 |
| Control | 5 | 1 | 60 | 1 | 65 | 1 |
Apoptosis is basically controlled by the Bcl-2 protein family, which include anti-apoptotic proteins like Bcl-2 and pro-apoptotic proteins like Bax.55,56 Multiple investigations have demonstrated a considerable correlation between high Bcl-2 levels and lowered Bax levels, which contribute to cancer cell proliferation.57,58 Thus, we evaluated the Bcl-2 and Bax protein expression levels in A-549 lung cancer cells that were exposed to compounds 20c and 21c.42
In comparison to control, untreated cells, compound 20c caused an 8-fold increase in Bax levels and a 3-fold decrease in Bcl-2 levels, as shown in Table 5. Additionally, compound 21c reduced Bcl-2 levels by four times and raised Bax levels nine times. These findings suggest that apoptosis may play a role in the antiproliferative properties of 20c and 21c.
Cancer cells frequently inactivate p53 enzymes during transformation, which could be explained by p53 overexpression's ability to trigger apoptosis.59 Cancer cells treated with 20c and 21c demonstrated a significant increase in p53 levels, which were at least five times greater than those of untreated control cells. This data shows that enhanced p53 protein levels may control the apoptotic process in these new drugs.
Moreover, caspase activity dictates the initiation and termination of the apoptosis process.60 Caspase-3, an essential enzyme, triggers apoptotic cell death by cleaving numerous proteins within the cell.61 Using the A-549 lung cancer cell line, the effects of compounds 20c and 21c on caspase-3 were evaluated and contrasted with staurosporine as a reference drug (Table 6). The findings indicated that 21c was the most potent derivative, exhibiting a significant increase in caspase-3 protein levels (580 ± 4 pg mL−1) compared with the reference, staurosporine (465 ± 4 pg mL−1). Compound 21c exhibited a 9-fold elevation in active caspase-3 levels compared to control A-549 cells, producing caspase-3 levels that surpassed those induced by staurosporine, the reference medication. Compound 20c demonstrated a sevenfold increase in active caspase-3 levels (455 ± 3 pg mL−1) compared to the control untreated A-549 lung cells, as shown in Table 6.
| Compd. no. | Caspase-3 | Caspase-8 | Caspase-9 | |||
|---|---|---|---|---|---|---|
| Conc. (Pg ml−1) | Fold change | Conc. (ng ml−1) | Fold change | Conc. (ng ml−1) | Fold change | |
| 20c | 455 ± 3 | 7 | 2.05 ± 0.10 | 20 | 22 ± 1 | 22 |
| 21c | 580 ± 4 | 9 | 2.80 ± 0.20 | 28 | 25 ± 1 | 25 |
| Staurosporine | 465 ± 4 | 7 | 1.90 ± 0.10 | 19 | 20 ± 1 | 20 |
| Control | 65 | 1 | 0.10 | 1 | 1 | 1 |
The effects of compounds 20c and 21c on caspase-8 and caspase-9 were investigated in order to determine whether these medicines cause apoptosis via the intrinsic or extrinsic pathways. In comparison to the control untreated A-549 cancer cells, the results showed that compound 20c elevates caspase-8 and caspase-9 levels by 20 and 22 times, respectively, while compound 21c increases caspase-8 and caspase-9 levels by 28 and 25 times. This signifies the activation of both intrinsic and extrinsic pathways, as shown in Table 6.
3.3. In silico studies
The traditional hinge-region hydrogen bond between the docked ligands' pyrimidine nitrogen and EGFR Met769 was reliably detected, demonstrating its crucial function in compound stabilization within the ATP-binding pocket. Erlotinib established an additional hydrogen bond with Lys721 via the ethoxy group, as well as a network of hydrophobic bonds comprising Leu694, Phe699, Val702, Ala719, and Leu820 (Fig. 4).
Compound 21c possesses a distinct pharmacophoric structure with two principal components: the phenyl-1,2,4-oxadiazole ring and the pyridine ring. Each characteristic specifically enhances the elevated binding affinity within the ATP-binding pocket of EGFR. The pyridine ring functions as the primary anchoring motif, forming a crucial hydrogen bond with the hinge residue Met769, a conserved interaction necessary for efficient ATP-competitive kinase inhibition. This ring is preserved in a planar conformation, essential for its accurate alignment within the tight hinge region. Planarity is enhanced by the neighboring thiazole group, which stabilizes the entire compound's binding via two pi-sulfur interactions with Cys751 and Met742 (Fig. 5). Adjacent to the pyridine ring, the p-methoxyphenyl-1,2,4-oxadiazole ring facilitates molecular recognition by two hydrogen bonds with Lys721 and Cys773, via the methoxy group and the oxygen atom of the 1,2,4-oxadiazole ring. Furthermore, the phenyl ring of the 1,2,4-oxadiazole moiety engages in a pi–anion interaction with Asp831, hence augmenting binding stability and directional specificity. Ultimately, the entire molecule contributes to a network of hydrophobic contacts with Leu694, Phe699, Ala719, and Leu820, as illustrated in Fig. 5. These combined effects clarify the strong binding affinity (−9.5 kcal mol−1 and RMSD of 1.46) observed in the docking tests and the significant in vitro inhibitory activity demonstrated by 21c, which surpassed that of the reference erlotinib.
Compound 20c's docking investigation within the EGFR binding site revealed that it has a binding interaction pattern identical to compound 21c, with the difference that, unlike compound 21c, the binding with residue Cys773 is accomplished via a pi-sulfur link rather than a hydrogen bond, Fig. 6. This elucidates the decreased binding affinity of 20c (−9.3 kcal mol−1) relative to 21c (−9.5 kcal mol−1) and, consequently, its reduced efficacy. Nonetheless, both 20c and 21c have greater potency than erlotinib.
With a docking score of −9.20 kcal mol−1 and an RMSD of 1.57 Å from the reference pose, 21c's docking into the ATP-binding site of BRAFV600E exhibited a well-aligned and energetically favorable binding configuration. The ligand is firmly positioned within the active site, where the pyridine ring forms a hydrogen bond with the backbone of Cys532 and engages in hydrophobic interactions with Trp531, Ala481, and Phe583, as depicted in Fig. 8. Additionally, the oxadiazole ring forms a hydrogen bond with Ser536 via the p-methoxy group, consistent with structure–activity relationship trends indicating increased potency with p-methoxy substitution. Furthermore, it engages in a π–anion interaction with Asp594 and a π–π-alkyl interaction with Val471, thereby stabilizing the scaffold's position within the hinge region.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 4.44) exhibiting high polarity (TPSA 111.40 Å2), comprising seven rotatable bonds, seven hydrogen-bond acceptors, and no hydrogen-bond donors. All solubility models indicate low to moderate aqueous solubility (ESOL logS −5.62; Ali −6.58; SILICOS-IT −9.61), and gastrointestinal absorption is anticipated to be minimal. The compound serves as a P-glycoprotein substrate and lacks blood–brain barrier permeability; yet it is anticipated to inhibit CYP2C9 and CYP3A4, indicating a potential risk for drug–drug interactions. The criteria for rule-based drug-likeness are fully met Lipinski's criteria are satisfied without violations (MW < 500), and the filters set by Ghose, Veber, Egan, and Muegge are satisfied (TPSA < 140 Å2 and < 10 rotors); the bioavailability score is 0.55. Alerts in medicinal chemistry are negligible (PAINS 0; Brenk alert 0). A synthetic accessibility (SA) score of 3.70 is deemed low to moderate, suggesting that the molecule is anticipated to be comparatively easy to synthesize.
P 4.44) exhibiting high polarity (TPSA 111.40 Å2), comprising seven rotatable bonds, seven hydrogen-bond acceptors, and no hydrogen-bond donors. All solubility models indicate low to moderate aqueous solubility (ESOL logS −5.62; Ali −6.58; SILICOS-IT −9.61), and gastrointestinal absorption is anticipated to be minimal. The compound serves as a P-glycoprotein substrate and lacks blood–brain barrier permeability; yet it is anticipated to inhibit CYP2C9 and CYP3A4, indicating a potential risk for drug–drug interactions. The criteria for rule-based drug-likeness are fully met Lipinski's criteria are satisfied without violations (MW < 500), and the filters set by Ghose, Veber, Egan, and Muegge are satisfied (TPSA < 140 Å2 and < 10 rotors); the bioavailability score is 0.55. Alerts in medicinal chemistry are negligible (PAINS 0; Brenk alert 0). A synthetic accessibility (SA) score of 3.70 is deemed low to moderate, suggesting that the molecule is anticipated to be comparatively easy to synthesize.Erlotinib, used as a reference, has an oral-drug-like profile (MW 393.4 g mol−1, consensus log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 3.20; TPSA 74.7 Å2; 10 rotatable bonds) with model-dependent but acceptable solubility (ESOL log
P 3.20; TPSA 74.7 Å2; 10 rotatable bonds) with model-dependent but acceptable solubility (ESOL log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S −4.11; Ali −4.56; SILICOS-IT −7.26) and predicted high GI absorption. It is not a P-gp substrate and is BBB permeant. Erlotinib meets the Lipinski, Ghose, Veber, Egan, and Muegge criteria (bioavailability score 0.55), has no PAINS alerts and one Brenk alert (alkyne), and is synthetically accessible (3.19). However, broad CYP inhibition (1A2/2C19/2C9/2D6/3A4) is envisaged for this chemotype.
S −4.11; Ali −4.56; SILICOS-IT −7.26) and predicted high GI absorption. It is not a P-gp substrate and is BBB permeant. Erlotinib meets the Lipinski, Ghose, Veber, Egan, and Muegge criteria (bioavailability score 0.55), has no PAINS alerts and one Brenk alert (alkyne), and is synthetically accessible (3.19). However, broad CYP inhibition (1A2/2C19/2C9/2D6/3A4) is envisaged for this chemotype.
Taken together, the ADME comparison reveals that 21c's potency is accompanied by a suboptimal physicochemical envelope, moderate MW, high lipophilicity, and high TPSA, all of which contribute to poor solubility and low predicted intestinal absorption and increase the likelihood of CYP2C9/3A4 interactions.
3.4. Structural activity relationship (SAR) analysis
The subsequent points delineate the structure–activity relationship of the newly synthesized compounds 5, 9, 14, 20a–c, 21a–c, and 22a–c.1- The phenyl-1,2,4-oxadiazole moiety is crucial for activity. Compounds 5 (thiophene-based derivative), 9 (thiazole-based derivative), and 14 (thiadiazole-based derivative), which are characterized by the presence of free phenolic moiety at position-3 of pyridine, were identified to be the least potent derivatives.
2- The pyridine ring is vital for activity, as the nitrogen atom forms a hydrogen bond with the crucial amino acid Met769, underscoring its significant role in stabilizing compounds within the ATP-binding pocket. It similarly establishes a hydrogen bond with Cys352, an essential interaction for the inhibitory activity of BRAFV600E, akin to that observed in Vemurafenib.
3- The pyridine ring engaged in a series of hydrophobic interactions within the binding sites of both EGFR and BRAFV600E, therefore consolidating the compounds in their receptor binding sites and augmenting their efficacy.
4- The coexistence of thiophene and thiazole groups at the 5-position of the pyridine moiety enhances antiproliferative activity more effectively than the thiadiazole group, with activity ranking as thiazole > thiophene > thiadiazole.
5- The unsubstituted phenyl group in the 1,2,4-oxadiazole structure reduces antiproliferative efficacy. Substitution with either an electron-withdrawing group (chlorine atom) or an electron-donating group (methoxy group) enhances antiproliferative activity, with the methoxy group demonstrating superior efficacy.
6- The methoxy group in the phenyl-1,2,4-oxadiazole moiety engages in additional hydrogen bonds with Lys712 in the EGFR binding site and Ser536 in the BRAFV600E binding pocket, thereby further stabilizing the compounds within the receptor binding sites and enhancing their efficacy.
4. Conclusion
This work presents the successful design and synthesis of novel thiophene/thiazole/thiadiazole-1,2,4-oxadiazole derivatives that are potential dual inhibitors of EGFR and BRAFV600E, introducing a new scaffold with considerable therapeutic significance in cancer treatment. Among the synthesized compounds, 20c and 21c emerged as the most promising lead compounds, demonstrating significant antiproliferative activity against specific human cancer cell lines while preserving a favorable safety profile in normal human cell lines. Compared to Erlotinib, compounds 20c and 21c exhibited superior inhibitory potency against EGFR and comparable efficacy to Vemurafenib against BRAFV600E. Compounds 20c and 21c exhibited apoptotic and immunomodulatory characteristics. Molecular docking experiments validated the dual-targeting strategy of the examined compounds via essential interactions within the active sites of EGFR and BRAFV600E.These findings underscore 20c and 21c as promising lead compounds for subsequent preclinical development. Subsequent research will focus on the structural optimization of analogous compounds and in vivo assessments to improve potency, selectivity, and therapeutic efficacy, thereby advancing innovative dual-targeted anticancer treatments.
Conflicts of interest
The authors disclose no conflicting interests.Data availability
The authors assert that any data supporting this study can be found in the supplementary materials (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00207b.Acknowledgements
This work was funded by the Deanship of Graduate Studies and Scientific Research at Jouf University under grant no. (DGSSR-2025-01-01299), and the KIT-Publication Fund of the Karlsruhe Institute of Technology.References
- L. H. Al-Wahaibi, S. A. Othman, H. A. Abou-Zied, S. Bräse, B. G. Youssif and S. M. Rabea, RSC Adv., 2025, 15, 47710–47734 RSC.
- R. L. Siegel, T. B. Kratzer, N. S. Wagle, H. Sung and A. Jemal, Ca, 2026, 76, e70043 Search PubMed.
- P. N. Lara Jr and D. L. Hershman, Ca, 2026, 76, e70061 Search PubMed.
- N. A. Elhawary, S. N. Ekram, H. A. Sembawa, E. Tashkandi, S. Bannani, Z. A. Azher, I. S. Abumansour, R. M. Almuqati, R. Attieh and I. A. Sindi, Int. J. Environ. Health Res., 2025, 35, 3648–3662 Search PubMed.
- C. Xiao, M. Ji and D. Lai, Sage Open Pediatr., 2025, 12, 30502225251370801 CrossRef PubMed.
- A. Berkman, M. Roth and J. A. Livingston, JAMA Oncol., 2026, 12, 26–27 CrossRef PubMed.
- S. D. Bhandare, Toxicol. Res., 2026, 15, tfaf184 CrossRef CAS PubMed.
- A. Hota and B. K. Mandal, Med. Oncol., 2025, 43, 75 CrossRef PubMed.
- M. Ge, X.-Y. Chen, P. Huang, J. S. Fleishman, D.-H. Yang, Z.-X. Wu, Z.-F. Ke and Z.-S. Chen, Nat. Rev. Clin. Oncol., 2025, 22, 760–780 CrossRef PubMed.
- J. Li, J. Hu, Y. Yang, H. Zhang, Y. Liu, Y. Fang, L. Qu, A. Lin, P. Luo and A. Jiang, Mol. Biomed., 2025, 6, 111 CrossRef CAS PubMed.
- Y. A. Abdelrady, H. S. Thabet and A. M. Sayed, Pharmacol. Rep., 2025, 77, 1–20 CrossRef PubMed.
- L. H. Al-Wahaibi, H. A. Abou-Zied, M. A. Mahmoud, B. G. Youssif, S. Bräse and S. M. Rabea, J. Enzyme Inhib. Med. Chem., 2025, 40, 2516075 CrossRef PubMed.
- L. H. Al-Wahaibi, H. M. Hafez, F. H. Edrees, H. A. Abou-Zied, B. G. Youssif and S. Bräse, J. Mol. Struct., 2025, 1339, 142364 CrossRef CAS.
- M. Keerikkadu, P. D. Bangera, V. K. Tippavajhala and M. Rathnanand, AAPS PharmSciTech, 2025, 26, 131 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, H. A. Abou-Zied, E. A. Beshr, B. G. Youssif, A. M. Hayallah and M. Abdel-Aziz, Int. J. Mol. Sci., 2023, 24, 9104 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, A. M. Gouda, O. F. Abou-Ghadir, O. I. Salem, A. T. Ali, H. S. Farghaly, M. H. Abdelrahman, L. Trembleau, H. H. Abdu-Allah and B. G. Youssif, Bioorg. Chem., 2020, 104, 104260 CrossRef CAS PubMed.
- S. Napolitano, D. Ciardiello, E. Cioli, E. Martinelli, T. Troiani, M. G. Zampino, N. Fazio, F. De Vita, F. Ciardiello and G. Martini, Cancer Treat. Rev., 2025, 134, 102905 CrossRef CAS PubMed.
- U. Częścik, M. Gryglas, A. Szterk and S. Flis, Oncol. Res., 2026, 34, 2 CrossRef PubMed.
- M. Y. Abdel-Hady, M. M. Morcoss, B. A. Alyami, B. G. Youssif, E. S. M. Abdelhafez and M. Abdel-Aziz, J. Mol. Struct., 2025, 144971 Search PubMed.
- L. H. Al-Wahaibi, H. A. Abou-Zied, M. Hisham, E. A. Beshr, B. G. Youssif, S. Bräse, A. M. Hayallah and M. Abdel-Aziz, Molecules, 2023, 28, 6586 Search PubMed.
- L. H. Al-Wahaibi, M. Hisham, H. A. Abou-Zied, H. A. Hassan, B. G. Youssif, S. Bräse, A. M. Hayallah and M. Abdel-Aziz, Pharmaceuticals, 2023, 16, 1522 Search PubMed.
- X. Wei, D. Fan, H. Dong, M. Li, Q. Lv, P. Zheng and L. Wang, Curr. Pharmaceut. Des., 2025, 31, 1521–1536 Search PubMed.
- S.-X. Yao, Y.-J. Huang, Y.-X. Zhang, Z.-X. Cui, H.-Y. Lu, R. Wang and L. Shi, J.Drug Target., 2025, 1–24 Search PubMed.
- M. Zhang, C. Zhang, K. Liu, X. Yang, X. Liu and F. Ge, ACS Omega, 2025, 10, 12170–12184 Search PubMed.
- M. Alrooqi, S. Khan, F. A. Alhumaydhi, S. A. Asiri, M. Alshamrani, M. M. Mashraqi, A. Alzamami, A. M. Alshahrani and A. A. Aldahish, Anti-Cancer Agents Med. Chem., 2022, 22, 2775–2787 Search PubMed.
- J. Liu, X. Zhang, Y. Zhuang, Z. Xu and W. Zhou, Future Med. Chem., 2025, 1–18 Search PubMed.
- M. Kučerová-Chlupáčová, ChemMedChem, 2025, 20, e202400971 CrossRef PubMed.
- D. Nagineni, P. Murugesan, N. P. Abburi, R. K. Bollikanda, B. Sridhar, A. D. Tangutur and S. Kantevari, Bioorg. Chem., 2025, 156, 108208 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, A. M. Mohamed, H. A. Abou-Zied, A. Y. A. Alzahrani, S. Bräse and B. G. Youssif, RSC Adv., 2025, 15, 38998–39027 RSC.
- A. M. Mohamed, O. M. Abou-Ghadir, Y. A. Mostafa, K. A. Dahlous, S. Bräse and B. G. Youssif, Front. Chem., 2024, 12, 1447618 CrossRef CAS PubMed.
- H. A. Abou-Zied, E. A. Beshr, H. A. Gomaa, Y. A. Mostafa, B. G. Youssif, A. M. Hayallah and M. Abdel-Aziz, Arch. Pharm., 2023, 356, 2200464 CrossRef CAS PubMed.
- M. B. Alshammari, A. A. Aly, B. G. Youssif, S. Bräse, A. Ahmad, A. B. Brown, M. A. Ibrahim and A. H. Mohamed, Front. Chem., 2022, 10, 1076383 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, E. M. El-Sheref, A. A. Hassan, S. Bräse, M. Nieger, B. G. Youssif, M. A. Ibrahim and H. N. Tawfeek, Pharmaceuticals, 2023, 16, 1014 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, A. M. Mohassab, S. M. Rabea, B. G. Youssif, S. Bräse and E. M. El-Sheref, RSC Adv., 2025, 15, 38229–38242 RSC.
- L. H. Al-Wahaibi, H. A. Abou-Zied, B. G. Youssif, S. Bräse and M. Hisham, Bioorg. Chem., 2025, 109329 Search PubMed.
- M. Ramadan, M. Abd El-Aziz, Y. A. Elshaier, B. G. Youssif, A. B. Brown, H. M. Fathy and A. A. Aly, Bioorg. Chem., 2020, 105, 104392 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, A. M. Elshamsy, T. F. Ali, B. G. Youssif, S. Brase, M. Abdel-Aziz and N. A. El-Koussi, ACS Omega, 2024, 9, 34358–34369 CrossRef CAS PubMed.
- H. A. Gomaa, A. M. Atta, M. E. Shaker, S. I. Alzarea, T. G. Alsahli, E. Alatwi, A. Musa, F. A. Mohamed, H. A. Alzahrani and A. M. Mohassab, Front. Chem., 2025, 13, 1595997 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, H. A. Abou-Zied, M. Nieger, S. Bräse, B. G. Youssif and H. Mohamed, Front. Chem., 2025, 13, 1638489 CrossRef CAS PubMed.
- B. G. Youssif, A. M. Gouda, A. H. Moustafa, A. A. Abdelhamid, H. A. Gomaa, I. Kamal and A. A. Marzouk, J. Mol. Struct., 2022, 1253, 132218 CrossRef CAS.
- D. P. Chandler, C. A. Wagnon and H. Bolton Jr, Appl. Environ. Microbiol., 1998, 64, 669–677 CrossRef CAS PubMed.
- M. Hisham, B. G. Youssif, E. E. A. Osman, A. M. Hayallah and M. Abdel-Aziz, Eur. J. Med. Chem., 2019, 176, 117–128 CrossRef CAS PubMed.
- N. El-Hachem, B. Haibe-Kains, A. Khalil, F. H. Kobeissy and G. Nemer, in Neuroproteomics: Methods and Protocols, Springer, 2017, pp. 391–403 Search PubMed.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, Biol. Dir., 2020, 15, 12 CrossRef PubMed.
- A. M. Mohamed, O. M. Abou-Ghadir, Y. A. Mostafa, Z. M. Almarhoon, S. Bräse and B. G. Youssif, RSC Adv., 2024, 14, 38403–38415 RSC.
- Y. A. Mostafa, J. A. Assoud, A. Y. Desoky, S. Mohamady, N. M. Mohamed, O. I. Salem, Z. M. Almarhoon, S. Bräse and B. G. Youssif, Front. Chem., 2024, 12, 1498104 CrossRef CAS PubMed.
- K. Shohdy, I. S. Chin and F. Thistlethwaite, in Treatment of Cancer, CRC Press, 2025, pp. 636–652 Search PubMed.
- A. Kharbanda, Discovery of Small Molecule Anti-cytokines to Improve Treatment Outcomes in Oncology, University of Arkansas for Medical Sciences, 2021 Search PubMed.
- P. Priya, A. Kumar, A. K. Ghosh, R. Chauhan, R. Kumar and D. Kumar, Chem. Biol. Lett., 2025, 12, 1254 CrossRef.
- Z. Mozooni, R. Ghadyani, S. Soleimani, E. R. Ahangar, M. Sheikhpour, M. Haghighi, M. Motallebi, A. Movafagh and S. M. Aghaei-Zarch, Pathol. Res. Pract., 2024, 263, 155665 CrossRef CAS PubMed.
- Y. Liu, Y. Gao and T. Lin, Ann. Palliat. Med., 2021, 10, 127592766–127512766 Search PubMed.
- A. R. Safa, Cancer Drug Res., 2022, 5, 850 CrossRef CAS PubMed.
- E. Dogan, H. G. Kara, B. Kosova and V. B. Cetintas, Targeting Apoptosis to Overcome Chemotherapy Resistance, Exon Publications, 2022, pp. 163–180 Search PubMed.
- M. Mosadegh, N. Noori Goodarzi and Y. Erfani, Cancer Invest., 2025, 43, 33–58 CrossRef PubMed.
- C. M. Croce, A. Strasser, J. T. Opferman, P. E. Czabotar and S. W. Fesik, Cell Death & Differ., 2025, 1–13 Search PubMed.
- T. Yasmin and Y. Kabir, in Cancerous Cells: Immunobiology of Tumors and Metastasis, Springer, 2025, pp. 193–217 Search PubMed.
- N. Kunac, N. Filipović, S. Kostić and K. Vukojević, Medicina, 2022, 58, 1135 CrossRef PubMed.
- M. Wang and X. Wang, J. Biochem. Mol. Toxicol., 2025, 39, e70125 CrossRef CAS PubMed.
- H. Wang, M. Guo, H. Wei and Y. Chen, Signal Transduction Targeted Ther., 2023, 8, 92 Search PubMed.
- G. Sahoo, D. Samal, P. Khandayataray and M. K. Murthy, Mol. Neurobiol., 2023, 60, 5805–5837 CrossRef CAS PubMed.
- M. Asadi, S. Taghizadeh, E. Kaviani, O. Vakili, M. Taheri-Anganeh, M. Tahamtan and A. Savardashtaki, Biotechnol. Appl. Biochem., 2022, 69, 1633–1645 CrossRef CAS PubMed.
- A. Najib, R. A. S. Rahmawati, A. U. A. Hasnaeni and A. Sakti, J. Pharm. Pharmacogn. Res., 2025, 13, S58–S66 CrossRef CAS.
- R. Y. El-Mahdy, N. Galal, R. Lotfy and R. K. Arafa, Comput. Biol. Med., 2025, 186, 109640 CrossRef CAS PubMed.
- S. Bhambri and P. C. Jha, Mol. Diversity, 2025, 1–17 Search PubMed.
- T. Peanlikhit, U. Aryal, J. S. Welsh, K. R. Shroyer and K. N. Rithidech, Int. J. Mol. Sci., 2025, 26, 2548 CrossRef CAS PubMed.
- X. Tian, C. Huang, Y. Lin and X. Yu, in Drug Discovery Stories, Elsevier, 2025, pp. 121–138 Search PubMed.
- N. R. Tshiluka, D. T. Mbedzi, M. V. Bvumbi and S. S. Mnyakeni-Moleele, ChemistryOpen, 2025, 14, e202400119 CrossRef CAS PubMed.
- A. Seghir, M. Mokhtar, K. M. A. Azzam, B. Nadia, E. C. Aytar, A. Saad and T. Boumediene, Biomed. Chromatogr., 2025, 39, e70110 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |