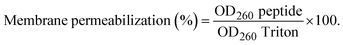Open Access Article
Open Access ArticleIntegrated proteomics, molecular dynamics, and in vitro characterization of antimicrobial peptide from Lactobacillus acidophilus vesicles against Streptococcus mutans
Venkatramanan Mahendrarajan and
Nalini Easwaran
and
Nalini Easwaran *
*
School of Biosciences and Technology, Vellore Institute of Technology, Tiruvalam Road, Katpadi, Vellore-632014, Tamil Nadu, India. E-mail: nalini.e@vit.ac.in
First published on 27th January 2026
Abstract
Membrane vesicles (MVs) released by probiotic bacteria such as Lactobacillus acidophilus have emerged as rich sources of bioactive peptides with antimicrobial properties. These MV-associated peptides represent a natural alternative to conventional synthetic antimicrobials, offering inherent selectivity and enhanced stability within complex microbial ecosystems such as the oral cavity. Given that Streptococcus mutans is a major etiological agent of dental caries due to its biofilm-forming and acidogenic capabilities, targeting it with probiotic-derived antimicrobial peptides (AMPs) presents a promising therapeutic avenue. In this study, the L. acidophilus MVs' peptide was tested against S. mutans, in silico and in vitro. Proteomic profiling of L. acidophilus MVs identified 143 unique peptides, of which several were predicted as AMPs by computational screening. A lead sequence, LTVTKAMNKVNKNAK, was prioritized for its non-hemolytic, non-toxic profile and favorable physicochemical traits. Molecular modelling and dynamics revealed membrane surface associated disruption against S. mutans by increased root mean square deviation (RMSD), root mean square fluctuation (RMSF) of the membrane, and water penetration count. The lead peptide exhibited an EC50 value of 1.1 mM and effectively inhibited biofilm formation while enhancing membrane permeability, thereby corroborating the in silico findings. These findings suggest that MVs from L. acidophilus can harbor AMPs capable of disrupting Gram-positive bacterial membranes through membrane disruption mechanisms.
1. Introduction
Dental caries is a major global public health problem affecting individuals across all age groups.1 According to the Global Burden of Disease Study, untreated dental caries of permanent teeth is the most prevalent condition worldwide, with a high incidence particularly in low and middle income countries.2 The etiopathogenesis of dental caries is primarily attributed to dysbiosis within the oral microbiome, where acidogenic and aciduric bacteria such as Streptococcus mutans play a central role.3 S. mutans contributes to cariogenesis through several virulence mechanisms, including its ability to form robust biofilms, metabolize carbohydrates to organic acids, and synthesize extracellular polysaccharides that enhance adherence and biofilm stability.4–6Traditional caries management relies heavily on mechanical debridement and chemical antimicrobial agents such as chlorhexidine and fluoride.7,8 However, these treatments have limitations, including non-specific action, undesirable side effects, and increasing concerns about antimicrobial resistance and microbiome disruption.9,10 In this context, antimicrobial peptides (AMPs) have emerged as promising therapeutic candidates.11 AMPs are small, typically cationic and amphipathic molecules produced by a wide range of organisms as part of their innate immune defense.12 They exhibit broad-spectrum antimicrobial activity, often by targeting microbial membranes, leading to pore formation, membrane thinning, or lipid disordering.13 From a chemical perspective, AMP activity is governed by its physicochemical properties such as net positive charge, amphipathicity, hydrophobic moment, and secondary structure propensity, which collectively dictate affinity for anionic bacterial membranes.14 Likewise, lipid chemistry, particularly the abundance of anionic phospholipids such as POPG and cardiolipin in the membrane creates electrostatic and hydrophobic interaction landscapes that facilitate selective peptide insertion and disordering.15
Recent studies have focused on identifying natural sources of AMPs from probiotic bacteria, including Lactobacillus acidophilus, which is widely recognized for its probiotic and health-promoting properties.16 While some strains of L. acidophilus have been historically associated with late-stage caries progression, others demonstrate strain-specific antimicrobial effects against oral pathogens, particularly through secreted metabolites and membrane vesicles (MVs).17 MVs are nanoscale, bilayered extracellular structures released by both Gram-positive and Gram-negative bacteria and are increasingly recognized as vehicles for the delivery of bioactive molecules, including peptides, enzymes, and signaling factors.18–20 Proteomic studies have revealed that MVs from probiotic bacteria such as L. acidophilus harbor peptides with antimicrobial, immunomodulatory, and antibiofilm activities.21,22 Beyond their biological origin, MV associated peptides represent a chemically distinct class of AMPs. Unlike canonical AMPs that are often optimized for systemic host defense, MV associated peptides are naturally secreted, shorter, less hydrophobic, and frequently enriched in cationic and polar residues.23 These chemical features make them more compatible with aqueous and biofilm-rich environments such as the oral cavity and may reduce cytotoxicity towards mammalian cells. This chemical novelty positions MV associated AMPs as an underexplored source for selective, biocompatible oral antimicrobials.11 Computational screening of peptides derived from bacterial MVs has become a powerful approach to identify novel AMPs.24 Advanced in silico tools such as the Database of Antimicrobial Activity and Structure of Peptides (DBAASP) and Collection of Antimicrobial Peptides (CAMP) allow high throughput screening based on machine learning algorithms and curated experimental data.25,26 Moreover, toxicity and hemolysis prediction tools such as ToxinPred 3.0 and HemoPI 2.0 are essential for prioritizing peptides with therapeutic potential and minimal risk to host cells.27,28
Molecular dynamics (MD) simulations offer a powerful platform to investigate peptide membrane interactions at atomistic resolution.29 Several studies have demonstrated that AMP activity is highly dependent on peptide orientation, conformation, and physicochemical properties, which determine the mode of membrane disruption ranging from carpet-like surface destabilization to transmembrane pore formation.30 Despite the increasing catalog of bioactive peptides identified from probiotic and commensal bacteria, a critical knowledge gap remains in delineating how such MV associated peptides interact with bacterial membranes at an atomistic level.31 Particularly for oral pathogens like S. mutans, the mechanistic understanding of AMP mediated disruption is scarce, limiting rational design and clinical translation of these molecules. Thus, integrating proteomic discovery with atomistic molecular dynamics and validating in vitro provides a powerful strategy to bridge this gap and accelerate AMP validation as next-generation oral therapeutics. In the present study, we performed proteomic screening of MVs derived from L. acidophilus MTCC 10307 and identified potential AMP candidates. The peptides were evaluated for antimicrobial potential using DBAASP and CAMP, and screened for hemolytic and toxic properties using HemoPI 2.0 and ToxinPred 3.0, respectively. We then constructed a biologically relevant model membrane representing S. mutans, incorporating 1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (POPG), cardiolipin (TOCL2), and phosphatidylethanolamine (POPE), based on established lipidomic data.32,33
Gram positive organisms such as S. mutans possess membranes enriched with cardiolipin and negatively charged phospho glycerol lipids, rendering lipid electrostatics a dominant chemical determinant of AMP binding and its associated disruption. This anionic lipid enrichment contrasts with the predominantly zwitterionic composition of mammalian membranes and provides a basis for selective AMP targeting. All-atom MD simulations were carried out to explore the interaction of the selected peptide with the modeled S. mutans membrane in perpendicular orientation.34 Through comprehensive structural and dynamic analyses, Root Mean Square Deviation (RMSD), z-distance, Center Of Mass (COM) distance, contacts and hydrogen bond, we aimed to elucidate the mechanism by which the peptide perturbs the bacterial membrane. These insights provide mechanistic support for the use of MVs associated AMPs as targeted antimicrobial agents for oral health applications, with potential to combat biofilm-associated infections such as dental caries.
2. Materials and methods
2.1. Proteomics study of L. acidophilus MVs
The MVs isolated from L. acidophilus MVs MTCC 10307 were sonicated for 1 minute to release proteins, with approximately 50 µg of liberated protein treated in 100 µL of 7 M urea. To reduce disulfide bonds, 50 mM tris(2-carboxyethyl)phosphine (TCEP) was added and incubated at 37 °C for 1 hour. Then, 50 mM iodoacetamide (IAA) was introduced, mixed by vortexing, and incubated again at 37 °C for 30 minutes for alkylation. To lower the urea concentration in preparation for trypsin digestion, 25 mM NH4HCO3 was added. Trypsin was then added at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 enzyme-to-substrate ratio, and the samples were incubated at 37 °C for 16 hours. After digestion, samples were dried in a speed vacuum and reconstituted in 20 µL of Milli-Q water with 0.1% formic acid. For desalting, a Thermo Scientific C18 Pierce column was used. The resin was activated and equilibrated in 50% methanol/50% acetonitrile (ACN) and then in 5% ACN/0.5% TFA for 3–5 minutes. The reconstituted protein sample was applied to the resin and centrifuged for 3 minutes, followed by a wash with 0.5% TFA in 5% ACN. Elution was performed with 20 µL of 70% ACN, after which samples were dried in a speed vacuum and stored at −80 °C for LC-MS/MS analysis. For the LC-MS/MS, the mobile phases used were 0.1% formic acid in Milli-Q water (A) and 85
30 enzyme-to-substrate ratio, and the samples were incubated at 37 °C for 16 hours. After digestion, samples were dried in a speed vacuum and reconstituted in 20 µL of Milli-Q water with 0.1% formic acid. For desalting, a Thermo Scientific C18 Pierce column was used. The resin was activated and equilibrated in 50% methanol/50% acetonitrile (ACN) and then in 5% ACN/0.5% TFA for 3–5 minutes. The reconstituted protein sample was applied to the resin and centrifuged for 3 minutes, followed by a wash with 0.5% TFA in 5% ACN. Elution was performed with 20 µL of 70% ACN, after which samples were dried in a speed vacuum and stored at −80 °C for LC-MS/MS analysis. For the LC-MS/MS, the mobile phases used were 0.1% formic acid in Milli-Q water (A) and 85![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 ACN in Milli-Q with 0.1% formic acid (B). Separation was achieved using a PepMap RSLC C18 analytical column (2 µm, 100 Å, 50 cm) and an Acclaim PepMap 100 pre-column (100 µm × 2 cm, nanoViper). The analysis was conducted on a Q Exactive Plus Biopharma mass spectrometer (Thermo Scientific).35 The data were acquired using the software Thermo Scientific Xcalibur (version 4.2.28.14). The thermo raw files were converted into open mzML format using ThermoRawFileParser (1.4.5).36,37 Then peptides were identified using MSGF+(2024.03.26) software by matching them with host protein sequences obtained from UniProt (L. acidophilus, Taxon ID: 1579).38 Peptide identification was performed using MS-GF+ with default search parameters, assuming trypsin specificity and allowing up to two missed cleavages. Carbamidomethylation of cysteine was set as a fixed modification and methionine oxidation as a variable modification. A posterior error probability (PEP) cutoff of 0.01 was applied, corresponding to a 1% peptide level False Discovery Rate (FDR). Peptide confidence was further validated using a target-decoy strategy.39 Then high confidence peptides were filtered using posterior error probability of 0.01 using OpenMS (3.3.0).40 Then the AMPs were predicted in the Database of Antimicrobial Activity and Structure of Peptides (DBAASP) and CAMP server. The AMPs which got predicted in common by both servers are listed in a table and taken for further analysis.25,26
15 ACN in Milli-Q with 0.1% formic acid (B). Separation was achieved using a PepMap RSLC C18 analytical column (2 µm, 100 Å, 50 cm) and an Acclaim PepMap 100 pre-column (100 µm × 2 cm, nanoViper). The analysis was conducted on a Q Exactive Plus Biopharma mass spectrometer (Thermo Scientific).35 The data were acquired using the software Thermo Scientific Xcalibur (version 4.2.28.14). The thermo raw files were converted into open mzML format using ThermoRawFileParser (1.4.5).36,37 Then peptides were identified using MSGF+(2024.03.26) software by matching them with host protein sequences obtained from UniProt (L. acidophilus, Taxon ID: 1579).38 Peptide identification was performed using MS-GF+ with default search parameters, assuming trypsin specificity and allowing up to two missed cleavages. Carbamidomethylation of cysteine was set as a fixed modification and methionine oxidation as a variable modification. A posterior error probability (PEP) cutoff of 0.01 was applied, corresponding to a 1% peptide level False Discovery Rate (FDR). Peptide confidence was further validated using a target-decoy strategy.39 Then high confidence peptides were filtered using posterior error probability of 0.01 using OpenMS (3.3.0).40 Then the AMPs were predicted in the Database of Antimicrobial Activity and Structure of Peptides (DBAASP) and CAMP server. The AMPs which got predicted in common by both servers are listed in a table and taken for further analysis.25,26
2.2. Peptide screening and toxicity evaluation
The hemolytic potential of these peptides was evaluated in silico using the HemoPI 2.0 server, which predicts hemolytic activity based on physicochemical and structural features.28 The regression method was chosen which provides both a binary classification (hemolytic vs. non-hemolytic) and a regression-based HC50 value (µM) using Random Forest models trained on peptide sequence features.41 Toxicity was assessed using the ToxinPred 3.0 webserver, which classifies peptides as toxic or non-toxic based on sequence-derived features.27 Subsequently, the physicochemical properties including net charge, hydrophobicity, penetration depth and tilt angle were analyzed using the DBAASP platform. Further Boman index and GRAVY score was computed using APD6 online server.42 Based on these evaluations, a lead peptide with favorable antimicrobial, non-toxic, and non-hemolytic properties was selected for further structural and functional analysis. Additional properties such as hydrophobic moment, isoelectric point, Boman index, and amphipathicity were also calculated to assess peptide folding propensity and functional selectivity toward bacterial over mammalian membranes.252.3. Protein and Streptococcus mutans membrane modeling
The three-dimensional (3D) structure of the selected lead peptide was predicted using the PEP-FOLD3 webserver, which employs de novo modeling based on structural alphabets. The generated structure was further validated using the Ramachandran plot analysis within BIOVIA Discovery Studio to assess stereochemical quality.43For membrane modeling, a Gram-positive bacterial membrane representing S. mutans was constructed using the CHARMM-GUI membrane builder. A bilayer membrane was generated with 200 lipids in each leaflet to approximate native membrane dimensions. The lipid composition was selected based on literature-reported membrane profiles of S. mutans, comprising 65% POPG, 30% cardiolipin (TOCL2), and 5% POPE, in a 65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30
30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 molar ratio.44–46
5 molar ratio.44–46
2.4. Molecular dynamics simulation of peptide–membrane system
MD simulations were performed using the GROMACS software package (v2024.1) with the CHARMM36m force field parameters, which were generated using the CHARMM-GUI interface.47,48 The system was solvated with the TIP3P water model, and Na+ and Cl− ions were added to neutralize the system through the Monte Carlo ion replacement method. Energy minimization was performed using the steepest descent algorithm, followed by a 1 ns equilibration phase during which the system temperature was gradually increased from 100 K to 310.15 K. The temperature and pressure were maintained using the velocity-rescale thermostat and Berendsen barostat, respectively. Long-range electrostatics were calculated using the Particle Mesh Ewald (PME) method with a real-space cut-off of 1.2 nm, and van der Waals interactions were treated with a 1.2 nm cut-off. All covalent bonds involving hydrogen atoms were constrained using LINCS, allowing a 2 fs integration time step. Semi-isotropic pressure coupling was applied to maintain the membrane system under periodic boundary conditions.An initial 500 ns MD simulation of the membrane alone (without peptide) was conducted in the NPT ensemble at 310.15 K and 1 atm to evaluate membrane stability. Subsequently, the peptide–membrane systems were simulated for 500 ns in perpendicular orientation to the membrane plane, to mimic possible physiological interactions and insertion behavior of the peptide. These simulations allowed for the detailed examination of peptide interactions and membrane disruption mechanisms.49 A simulation length of 500 ns was selected as peptide membrane interactions typically require hundreds of nanoseconds to capture adsorption, reorientation, and insertion events as well as membrane perturbation. Prior atomistic studies of antimicrobial peptides report comparable timescales (200–500 ns) for stabilization and initial pore-forming behavior, while shorter simulations (<100 ns) generally do not reach mechanistic convergence. All MD simulations were run on a local workstation (Intel Core i7-14700K CPU, NVIDIA RTX 4070 GPU, 32 GB DDR5 RAM) using GROMACS 2024.2 with CUDA acceleration.
2.5. Simulation analysis parameters
Membrane specific parameters such as RMSD and root mean square fluctuation (RMSF) has been plotted for the membrane to assess the structural integrity.For the peptide–membrane complexes, additional analyses were conducted including: RMSD, RMSF, peptide SASA, peptide z-distance (vertical insertion depth into the membrane), COM distance between peptide and membrane, number of contacts (<0.6 nm), and hydrogen bond between peptide and membrane throughout the simulation were analyzed. These analyses provided insights into the peptide's structural stability, orientation dynamics, and membrane-disruptive potential under simulated physiological conditions.50
2.6. Bacterium culture conditions, peptide synthesis and chemical characterization
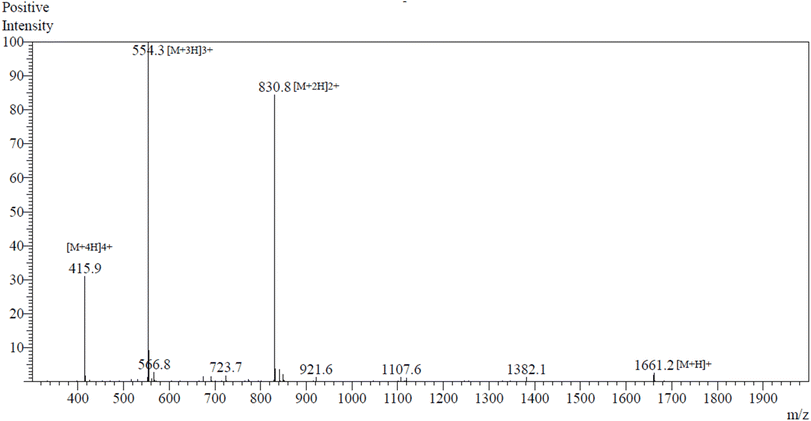
S. mutans (MTCC 497) was sources from Microbial Type Culture Collection (MTCC), Chandigarh, India. They were regularly maintained and cultured on Luria Bertani (LB) media (HiMedia) at 37 °C for 24 hours.The peptide LTVTKAMNKVNKNAK was synthesized via standard Fmoc-based solid-phase peptide synthesis (SPPS) on a rink amide resin using sequential coupling of protected amino acids (GenScript, USA). After chain assembly, the peptide was cleaved from the resin under acidic conditions and obtained as a TFA salt. The crude peptide was purified by reverse-phase high-performance liquid chromatography (RP-HPLC) to >99% purity (retention time: 7.60 min), and purity was verified at 220 nm. Peptide identity was confirmed by electrospray ionization mass spectrometry (ESI-MS) in positive ion mode, with an observed molecular weight of 1659.9 Da closely matching the theoretical mass of 1660.01 Da. The peptide was obtained as a white lyophilized powder and stored at −20 °C until use.
2.7. Hemolysis assay
Hemolytic activity was evaluated by incubating the peptide at a final concentration of 2.2, 1.1 and 0.55 mM in HiMedia, blood agar plates. The plates were incubated at 37 °C for 24 hours and any zone of hemolysis in the spot area was imaged and recorded.512.8. Cytotoxicity assay
The cytotoxic potential of the peptide was assessed in vitro using on 3T3 mouse fibroblast cells at three concentrations: 2.2, 1.1, and 0.55 mM. Cells were cultured in DMEM supplemented with 5% FBS until they reached approximately 80% confluence, then treated with the peptide in fresh DMEM for 24 hours at 37 °C and 5% CO2. Following treatment, 100 µL of MTT solution (0.5 mg mL−1) was added to each well and incubated for an additional 4 hours. The resulting formazan crystals were dissolved in 100 µL DMSO, and cell viability was determined by measuring the absorbance at 570 nm.52 The percentage viability is calculated by the formula: % viability = treated OD/control OD × 100.2.9. MIC and half-maximal effective concentration
The MIC and half-maximal effective concentrations (EC50) for the synthesized peptide were determined as described earlier. 0.1OD culture of S. mutans were added with 6 to 0.047 mM concentration of peptide and incubated at 37 °C for 24 hours.53 The MIC of tetracycline was also performed for positive control of the study. EC50 estimation was performed by fitting percent inhibition versus log-transformed concentration using a four-parameter logistic (4PL) model implemented in Python (SciPy). Model quality was assessed via residual analysis, determination of R2 and RMSE, and bootstrap resampling (n = 2000) to obtain 95% confidence intervals. The top and bottom asymptotes were constrained to 100% and 0% inhibition, respectively. The EC50 obtained from the 4PL model was used to guide the selection of a representative concentration for downstream inhibition assays.542.10. Biofilm inhibition assay
The biofilm inhibition by peptide was screened using CV staining assay. Briefly 2.2, 1.1 and 0.55 mM of peptide was added with 0.1OD of S. mutans and incubated for 24 hours at 37 °C. The planktonic cells were removed carefully and the wells were washed with PBS. Then 0.1% crystal violet was used to stain the biofilm cells and unbound stain were removed by brief washing. The bound CV were eluted in 30% acetic acid and OD were recorded at 595 nm using a spectrophotometer.552.11. Membrane permeability assay
Membrane permeabilization was performed utilizing OD at 260 nm. 0.1OD cells of S. mutans were incubated with 2.2, 1.1 and 0.55 mM of peptide and incubated for 2 hours at 37 °C. The cultures were filtered using 0.22 µm syringe filters and the OD of filtrates were recorded at 260 nm. PBS was used as negative control and Triton X-100 was used as positive control for the experiment.562.12. Bacterial permeability assay
Bacterial membrane permeability was evaluated using propidium iodide (PI) staining and flow cytometry. Mid-log phase S. mutans cells (equivalent to 1OD) were treated with the peptides at concentrations of 2.2 mM, 1.1 mM, and 0.55 mM for 2 hours at 37 °C. Following incubation, the cells were stained with 15 µg mL−1 PI for 15 minutes in the dark. The cells were then centrifuged to remove excess dye and resuspended in PBS. PI-positive events were gated from the FSC/SSC scatter plots and quantified as membrane-permeabilized cells. Flow cytometric analysis was performed using a CytoFLEX (Beckman Coulter, USA) system.572.13. Statistical analysis
In vitro experiments such as cytotoxicity, two-fold serial dilution, biofilm inhibition and membrane permeability assays were performed with three independent biological replicates. Data are presented as mean ± SD. Statistical analyses were performed in GraphPad Prism (version 8). One-way ANOVA was used to compare multiple groups, followed by a post hoc t-test for pairwise comparisons. Statistical significance was indicated using APA-style asterisks as follows: *p < 0.05, **p < 0.01, and ***p < 0.001. Parametric tests were used under the assumption of normally distributed data.3. Results
3.1. Proteomic screening and AMP selection
Proteomic profiling of L. acidophilus MV fractions resulted in 143 unique peptides mapped against the host proteome. To evaluate their antimicrobial potential, peptides were screened through two curated AMP databases: DBAASP, which employs the Moon and Fleming hydrophobicity scale to assess membrane interaction potential, and CAMP, using the Random Forest algorithm both of which have been previously validated as reliable predictors of AMP functionality.Seven peptides predicted to have antimicrobial activity in both databases were shortlisted to improve prediction confidence and reduce false positives. To evaluate the safety profile of the identified AMPs, the peptides were subjected to in silico hemolytic activity screening using HemoPI 2.0, a machine learning-based predictive tool. The results indicated that all peptides were classified as non-hemolytic, with predicted HC50 values ranging from 134 to 221 µM, suggesting minimal risk of erythrocyte lysis at physiologically relevant concentrations. In parallel, potential peptide toxicity was assessed using ToxinPred 3.0, which employs advanced deep learning algorithms trained on experimentally validated toxic and non-toxic peptides. The analysis identified peptide 2 and peptide 4 as potentially toxic, whereas peptides including LTVTKAMNKVNKNAK, were predicted to be non-toxic. Among those peptides, the peptide LTVTKAMNKVNKNAK was prioritized based on its physicochemical characteristics: a net positive charge (+4), and a hydrophobicity profile conducive to membrane interaction, a chemical signature typical of membrane-active cationic peptides. Additionally, its penetration depth and tilt angle values, which are critical parameters in AMP membrane dynamics, suggested its ability to destabilize bacterial membranes.3,4 Interestingly, the identified peptide (15 residues) falls within the ideal size range of cationic AMPs (10–30 residues) that balance penetrative ability with reduced aggregation risk, further supporting its classification as a functional AMP candidate (Table 1).
| ID | Sequences | Length | Normalized hydrophobicity (Moon and Fleming) | Net charge | Penetration depth | Tilt angle | Grand average hydropathy value of the peptide | Boman index (kcal mol−1) | In silico hemolytic activity | In silico toxicity |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | LSSQIQNGAK | 10 | 0.13 | 1 | 22 | 24 | −0.63 | 1.74 | Non hemolytic | Non toxin |
| 2 | RNAIVK | 6 | 0.14 | 2 | 30 | 42 | −0.23 | 2.72 | Non hemolytic | Toxin |
| 3 | KAVREIIRYMKK | 12 | 0.06 | 4 | 30 | 79 | −0.717 | 2.95 | Non hemolytic | Non toxin |
| 4 | SGVNHRFKK | 9 | 1.02 | 3 | 30 | 39 | −1.467 | 3.63 | Non hemolytic | Toxin |
| 5 | YTFVWRKAVEKFHK | 14 | −0.1 | 3 | 17 | 56 | −0.714 | 1.96 | Non hemolytic | Non toxin |
| 6 | LTVTKAMNKVNKNAK | 15 | 0.53 | 4 | 23 | 16 | −0.653 | 1.88 | Non hemolytic | Non toxin |
| 7 | RNIYSR | 6 | 0.11 | 2 | 15 | 89 | −1.68 | 5.85 | Non hemolytic | Non toxin |
3.2. Peptide structure and membrane model construction
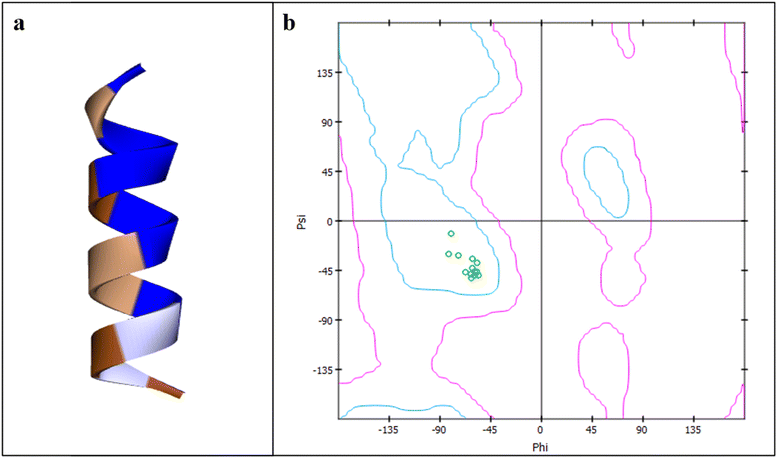
The 3D structure of LTVTKAMNKVNKNAK was modeled using PEP-FOLD3 and subjected to all-atom MD simulations (Fig. 1a). The predicted structure was validated by Ramachandran plot which depicted all the amino acids in the allowed region (Fig. 1b). The target membrane of S. mutans was constructed in CHARMM-GUI, with 200 lipids per leaflet (400 total), ensuring a biologically relevant bilayer size capable of accommodating full peptide insertion and realistic curvature. The membrane lipid composition POPG![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) TOCL2
TOCL2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) POPE in a 65
POPE in a 65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30
30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 molar ratio was chosen based on literature reporting the dominance of anionic phospholipids (particularly POPG and cardiolipin derivatives like TOCL2) in S. mutans membranes.
5 molar ratio was chosen based on literature reporting the dominance of anionic phospholipids (particularly POPG and cardiolipin derivatives like TOCL2) in S. mutans membranes.
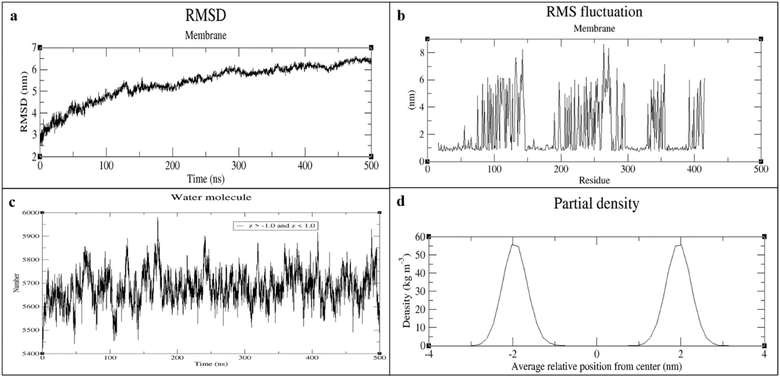
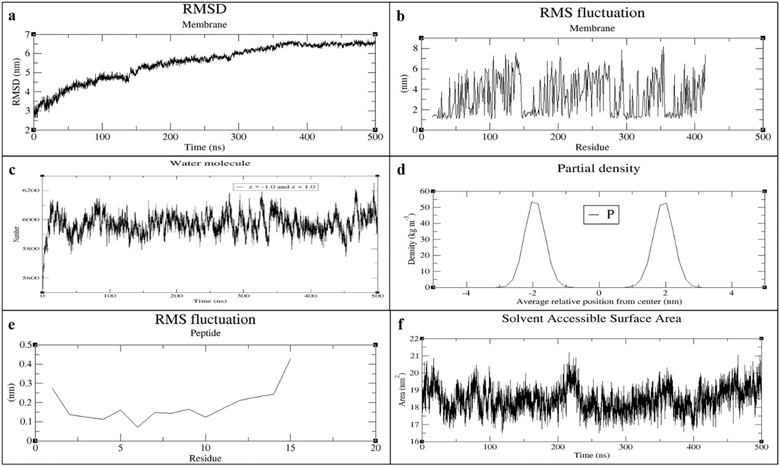
The number of water molecules within the central bilayer region (−1 < z < 1 nm) remained stable throughout the trajectory, fluctuating narrowly between 5400 and 5600 molecules, indicating preservation of membrane integrity (Fig. 2c). Membrane thickness profiles showed two sharp and symmetric peaks, consistent with a well-defined Gram-positive bilayer (Fig. 2d).
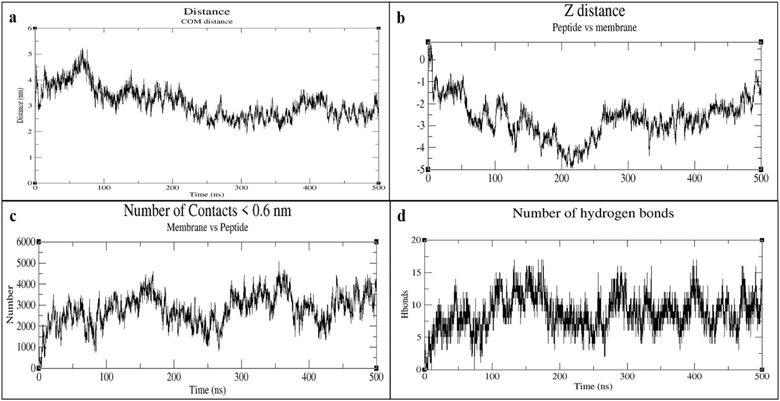
Interactions between the peptide and membrane were quantified by center of mass (COM) distance, z-axis displacement, number of hydrogen bonds, and contact frequency over a 500 ns simulation. The COM distance between the peptide and membrane fluctuated between 2.5 and 5 nm, with relative stability observed throughout the trajectory, indicating persistent peptide association with the bilayer surface (Fig. 4a). z-Axis distance analysis showed progressive peptide insertion, stabilizing at a displacement of approximately −4 nm for much of the simulation, suggesting the peptide penetrated and remained embedded in the membrane (Fig. 4b).
The number of atomic contacts (<0.6 nm) between the peptide and membrane lipids gradually increased and remained elevated (2000–5000 contacts), further supporting close association and substantial interaction surface between the peptide and bilayer lipids (Fig. 4c). Hydrogen bond analysis revealed formation and persistence of multiple peptide–membrane hydrogen bonds, averaging between 5 and 15 throughout the simulation, with transient peaks reaching up to 18. This indicates strong and dynamic interactions likely stabilizing the peptide in the bilayer (Fig. 4d).
Representative snapshots at 0, 250, and 500 ns, shown from both upright and side perspectives, highlight the temporal progression of the peptide's orientation and interaction with the membrane (Fig. 5a–f).
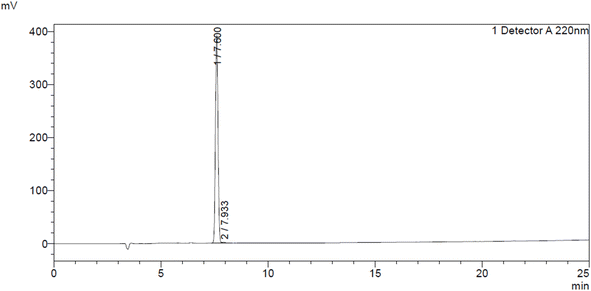 | ||
| Fig. 6 Analytical RP-HPLC trace at 220 nm shows a single peak at ∼7.6 min, confirming high chemical purity (>99%). | ||
3.3. In vitro toxicity screening
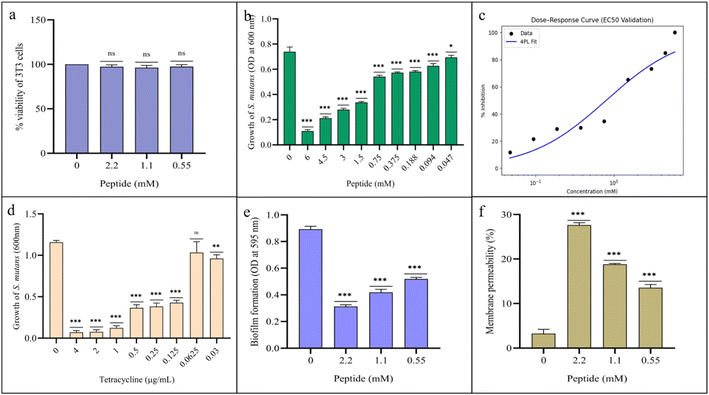
The peptides were evaluated for hemolytic and cytotoxic effects in vitro. At concentrations of 2.2, 1.1, and 0.55 mM, none of the peptides shown hemolysis on blood agar plates (Fig. 8a–c), whereas the positive control 1% Triton X 100 shows prominent hemolytic zone (Fig. 8d). Cytotoxicity assessment on 3T3 cells revealed high cell viability values of 97.29%, 96.33%, and 97.4% at 2.2 mM, 1.1 mM, and 0.55 mM concentrations, respectively (Fig. 9a).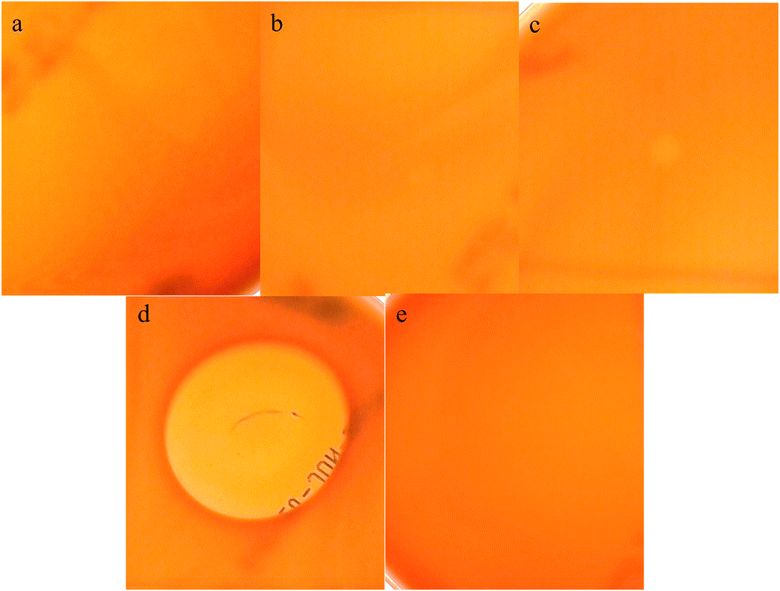 | ||
| Fig. 8 Hemolytic activity of peptides at (a) 0.55 mM (b) 1.1 mM (c) 2.2 mM (d) positive control 1% Triton X 100 (e) negative control (PBS). | ||
3.4. In vitro antibacterial activity
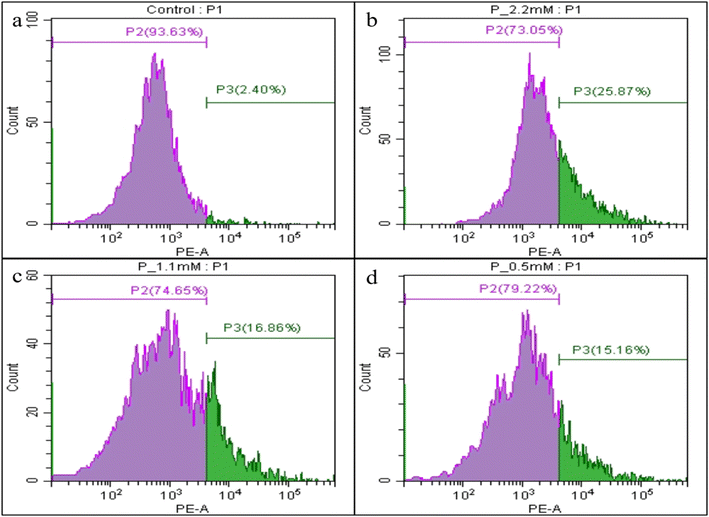
The peptide exhibited an EC50 value of 1.1 mM against S. mutans in vitro (Fig. 9b). A four-parameter logistic (4PL) model was used to estimate the EC50 from the dose response inhibition data. The validated EC50 was 0.80 mM with a 95% bootstrap confidence interval of 0.52–1.13 mM and a Hill slope of 0.89. The model showed a good fit to the data (R2 = 0.93; RMSE = 7.99). The 95% bootstrap confidence interval indicates that EC50 represents a concentration range rather than a single fixed value. Based on this range, an EC50 of 1.10 mM was selected as the representative midpoint and used for subsequent inhibition assays, as it fell within the validated interval and corresponded to the empirically observed midpoint of the inhibition curve (Fig. 9c). Residuals were evenly distributed around zero without systematic deviation, indicating that the 4PL model adequately described the data. The MIC against the positive control tetracycline was found to be 2 µg mL−1 for S. mutans in vitro (Fig. 9d). Biofilm formation was significantly inhibited in a dose-dependent manner, with inhibition rates of 64.91%, 53.03%, and 41.93% observed at 2.2 mM, 1.1 mM, and 0.55 mM peptide concentrations, respectively (Fig. 9e). Correspondingly, membrane permeabilization increased to 27.64%, 18.82%, and 13.56% at the same concentrations after 2 hours of treatment, indicating concentration-dependent disruption of the bacterial membrane integrity (Fig. 9f).Membrane-compromised (dead) cells were quantified using PI staining by flow cytometry. Following 2 hours of peptide exposure at concentrations of 2.2, 1.1, and 0.55 mM, the proportions of dead cells were 25.87%, 16.86%, and 15.16%, respectively (Fig. 10a–d).
4. Discussion
In this study, we discovered a novel AMP, LTVTKAMNKVNKNAK, originating from L. acidophilus membrane vesicles, and validated its activity through computational and in vitro analyses. In silico safety assessment predicted the peptide to be non-hemolytic and non-toxic, supporting its potential therapeutic applicability. All-atom molecular dynamics (MD) simulations further demonstrated its ability to interact with and disrupt a model S. mutans membrane, suggesting a plausible mechanism of action consistent with known AMP membrane disruptive models. These findings are significant because they highlight the potential of probiotic MV derived peptides as biocompatible candidates for oral antimicrobial therapy, offering advantages over synthetic AMPs through natural biogenesis and compatibility with complex biofilm niches.To ensure therapeutic safety, all peptides were assessed in silico for hemolysis and toxicity. HemoPI 2.0 predicted all peptides to be non-hemolytic, with HC50 values between 134 and 221 µM, indicating low potential to lyse human red blood cells. ToxinPred 3.0 identified peptides 2 and 4 as potentially toxic, whereas peptides including the lead candidate LTVTKAMNKVNKNAK, were non-toxic. The cationic and amphipathic nature of the lead peptide likely contributes to its selectivity for bacterial membranes, which are anionic, over zwitterionic mammalian membranes.58 These findings highlight its suitability for further antimicrobial evaluation. Unlike synthetic AMPs, MV-derived peptides benefit from natural biogenesis within probiotic vesicles, which may enhance their stability and functionality in complex biofilm niches.21 Moreover, since L. acidophilus is a Generally Recognized as Safe (GRAS) organism, its MV associated peptides represent a promising source of biocompatible antimicrobials.59 AMPs act primarily by disrupting microbial membranes, either by forming transient pores, inducing membrane thinning, or causing lipid disordering.60–63
In silico analysis revealed that the peptide engages the membrane primarily through surface association and partial insertion, inducing local lipid disorder without forming stable transmembrane pores. Increased lipid mobility and membrane hydration observed in the simulations suggest early-stage destabilization rather than catastrophic membrane rupture. This surface-driven perturbation is consistent with the mechanistic behavior of many cationic antimicrobial peptides that weaken membrane integrity through disordering, thinning, or interfacial disruption instead of pore formation. Importantly, this simulation-derived mode of action agrees with our experimental data, where the peptide caused concentration-dependent membrane permeabilization and PI uptake without overt cytotoxicity. Together, these results support a membrane-disruptive mechanism in which the peptide compromises S. mutans viability by perturbing lipid packing and weakening bilayer stability rather than forming defined pores.
The anionic nature of the membrane likely facilitated electrostatic attraction with the cationic peptide (+4 net charge), promoting initial surface interaction and eventual interactions. This is supported by the peptide's physicochemical properties such as moderate hydrophobicity, amphipathicity, and a suitable hydrophobic moment, all of which are known predictors of membrane activity.64,65 The membrane model used in our study, composed of POPG, TOCL2, and POPE in a 65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30
30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 molar ratio, is consistent with previously reported compositions for Gram-positive organisms, including S. mutans.44–46
5 molar ratio, is consistent with previously reported compositions for Gram-positive organisms, including S. mutans.44–46
The non-toxicity nature of peptide was confirmed in vitro through hemolysis and cytotoxicity assay. In vitro antimicrobial activity of the peptide LTVTKAMNKVNKNAK against S. mutans demonstrates its potential as a naturally occurring antibacterial agent, despite its relatively high effective concentration (EC50 = 1.1 mM). The observed dose-dependent inhibition of biofilm formation and increased membrane permeabilization suggest that the peptide exerts its effect primarily through membrane disruption, consistent with the membrane perturbation seen in molecular dynamics simulations.66 The permeabilization was confirmed through PI staining in flow cytometry which enters the cell with disrupted cytoplasmic membrane.67 Although the inhibitory concentrations are higher compared to some synthetic or chemically modified AMPs, this peptide was isolated directly from L. acidophilus MVs without sequence optimization or chemical modification. Naturally occurring peptides often display moderate activity in their native form, the peptide reported in our study show higher concentration for MIC and EC50 which is a possible limitation of this study but possess desirable features such as biocompatibility and low cytotoxicity within biological environments. Moreover, the moderate potency may reflect a physiological role of MV-associated peptides in maintaining microbial balance rather than complete eradication of commensal species.68,69 Importantly, the central value of this work lies not in immediate therapeutic potency but in demonstrating that probiotic MV-derived peptides can modulate Gram-positive membrane chemistry through a defined, surface-disruptive mechanism. Our peptide demonstrated moderate antimicrobial activity in vitro relative to more potent synthetic AMPs. Such higher potency benchmarks are primarily relevant to systemic antibiotics or optimized pharmaceutical peptides.60 Similarly tetracycline used as a positive control in this study, exhibited a MIC of 2 µg mL−1 against S. mutans in vitro, but clinical use in the oral cavity requires localized formulations at far higher concentrations. So the topical tetracycline-HCl pastes and irrigations are typically applied at 100 mg mL−1 and maintain gingival crevicular fluid concentrations in the µg mL−1 range for ≥1 week in vivo, demonstrating that oral efficacy relies on high local payload, sustained exposure, and direct contact with dental biofilms rather than low MIC values alone.70 Accordingly, the peptide's higher MIC/EC50 values remain within realistic bounds for localized dental delivery systems (e.g., gels, varnishes, pastes, or slow-release coatings) where high concentrations are both feasible and clinically acceptable.71 Importantly, the peptide is non-hemolytic, non-toxic, and membrane-selective, positioning it as a valuable scientific lead for understanding probiotic MV-derived antimicrobial mechanisms and for future peptide optimization rather than immediate clinical translation.72
Overall, the in vitro findings support the computational predictions and suggest that naturally secreted vesicular peptides from L. acidophilus can disrupt S. mutans membranes in a concentration-dependent manner. The observed membrane permeabilization aligns with the chemically predicted mechanism that cationic peptides disrupt Gram positive membranes via charge mediated adsorption followed by bilayer disordering rather than enzymatic inhibition.60 These results lay the foundation for future peptide engineering or formulation approaches to enhance potency while preserving the natural selectivity of probiotic-derived AMPs. The activity of the peptide can be rationalized in terms of its chemical composition. The peptide contains multiple lysine residues contributing to a net +4 charge, enabling electrostatic attraction to the anionic POPG and cardiolipin headgroups in the S. mutans membrane. Moreover, the distribution of hydrophobic and polar residues imparts amphipathicity, a key chemical feature that promotes interfacial alignment and partial insertion into the bilayer. These chemical descriptors are consistent with the MD results, where peptide adopted a surface-oriented topology and induced lipid disordering, increased hydration, and leaflet thinning. The concordance between chemical properties, atomistic simulation, and biological membrane assays indicate a membrane chemistry driven antimicrobial mechanism. While the antimicrobial potency of LTVTKAMNKVNKNAK is moderate compared to synthetic AMPs, the significance of these findings is primarily conceptual. This work identifies probiotic membrane vesicles as natural reservoirs of short cationic peptides capable of modulating Gram-positive membrane electrostatics and bilayer order. Such mechanistic insight advances peptide biology and probiotic chemistry, independent of immediate clinical application. Moreover, future work should investigate whether LTVTKAMNKVNKNAK can synergize with other probiotic-derived antimicrobials or conventional treatments like fluoride, enhancing efficacy while reducing cytotoxicity or resistance pressure. Validation of activity in saliva-mimicking and biofilm conditions will also be essential to approximate the complex oral microenvironment. Future studies should explore sequence optimization, synergistic combinations with oral antimicrobials, and in vivo validation in multispecies oral biofilms to assess therapeutic potential.
5. Conclusion
The peptide LTVTKAMNKVNKNAK, identified from L. acidophilus MVs, demonstrated selective membrane-disruptive and antibacterial activity against S. mutans while remaining non-toxic. These findings support the concept of MV associated AMPs as precision antimicrobials for oral pathogen control. Moving forward, sequence optimization, formulation for topical oral delivery, and in vivo testing in biofilm-based caries models will be essential to enhance potency and translate these peptides toward practical therapeutic applications. Integration of proteomics, MD simulations, and experimental validation pipelines may further accelerate the discovery and development of MV associated AMPs as a new class of biocompatible oral therapeutics.Author contributions
All authors contributed to the study conception and design. V. M. performed all the experiments and wrote the manuscript. N. E. finalized and edited the manuscript. N. E. supervised all the work performed.Conflicts of interest
The authors have no relevant financial or non-financial interests to disclose.Data availability
The data supporting this article have been included within this article.Acknowledgements
The authors thank the Indian Council of Medical Research for the funding provided to this project (Project no. 5/4/2-6/Oral Health/2022-NCD-II). The authors also thank Vellore Institute of Technology for providing infrastructure to carry out the research. The funder played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript.References
- V. Mahendrarajan, H. P. S. Lazarus, G. K. Muthukaliannan, S. Varghese and N. Easwaran, Front. Oral Health, 2025, 6, 1607931, DOI:10.3389/froh.2025.1607931.
- Z. Li, C. Yu and H. Chen, BMC Oral Health, 2025, 25, 715 CrossRef PubMed.
- Y. Zhu, Y. Wang, S. Zhang, J. Li, X. Li, Y. Ying, J. Yuan, K. Chen, S. Deng and Q. Wang, Front. Microbiol., 2023, 14, 1162380 CrossRef PubMed.
- J. A. Lemos, S. R. Palmer, L. Zeng, Z. T. Wen, J. K. Kajfasz, I. A. Freires, J. Abranches and L. J. Brady, Microbiol. Spectrum, 2019, 7 DOI:10.1128/microbiolspec.GPP3-0051-2018.
- Z. Gao, X. Chen, C. Wang, J. Song, J. Xu, X. Liu, Y. Qian and H. Suo, Microbiol. Res., 2024, 278, 127526 Search PubMed.
- W. H. Bowen, R. A. Burne, H. Wu and H. Koo, Trends Microbiol., 2018, 26, 229–242 CrossRef CAS PubMed.
- J. Autio-Gold, Operat. Dent., 2008, 33, 710–716 Search PubMed.
- M. J. Y. Yon, S. S. Gao, K. J. Chen, D. Duangthip, E. C. M. Lo and C. H. Chu, Dent. J., 2019, 7, 37, DOI:10.3390/dj7020037.
- Z. L. S. Brookes, R. Bescos, L. A. Belfield, K. Ali and A. Roberts, J. Dent., 2020, 103, 103497 CrossRef CAS PubMed.
- G. P. Moran, L. Zgaga, B. Daly, M. Harding and T. Montgomery, Toxicol. Lett., 2023, 379, 11–19 CrossRef CAS PubMed.
- Q.-Y. Zhang, Z.-B. Yan, Y.-M. Meng, X.-Y. Hong, G. Shao, J.-J. Ma, X.-R. Cheng, J. Liu, J. Kang and C.-Y. Fu, Mil. Med. Res., 2021, 8, 48 CAS.
- C. Bucataru and C. Ciobanasu, Microbiol. Res., 2024, 286, 127822 Search PubMed.
- F. Asif, S. U. Zaman, M. K. H. Arnab, M. Hasan and M. M. Islam, Microbe, 2024, 2, 100051 Search PubMed.
- Z. Jiang, A. I. Vasil, J. D. Hale, R. E. W. Hancock, M. L. Vasil and R. S. Hodges, Biopolymers, 2008, 90, 369–383 CrossRef CAS PubMed.
- A. L. Martin, P. N. Jemmett, T. Howitt, M. H. Wood, L. R. Cox, T. R. Dafforn, M. Campana, R. J. L. Welbourn, M. W. A. Skoda, L. A. Clifton, H. Hussain, J. L. Rawle, F. Carlà, C. L. Nicklin, T. Arnold and S. L. Horswell, Soft Matter, 2025, 21, 7054–7073 Search PubMed.
- Y. Liu, H. Nawazish, M. S. Farid, K. Abdul Qadoos, U. E. Habiba, M. Muzamil, M. Tanveer, M. Sienkiewicz, A. Lichota and Ł. Łopusiewicz, Fermentation, 2024, 10, 380, DOI:10.3390/fermentation10080380.
- D.-Y. Park, J. Hwang, Y. Kim, D. Lee, Y.-Y. Kim, H.-S. Kim and I. Hwang, Sci. Rep., 2023, 13, 7969 CrossRef CAS PubMed.
- M. Rima, M. Dakramanji, E. El Hayek, T. El Khoury, Z. Fajloun and M. Rima, Heliyon, 2025, 11, e42509 CrossRef PubMed.
- P. Aytar Çelik, K. Erdogan-Gover, D. Barut, B. M. Enuh, G. Amasya, C. T. Sengel-Türk, B. Derkus and A. Çabuk, Pharmaceutics, 2023, 15, 1052, DOI:10.3390/pharmaceutics15041052.
- Q. Li, X. Chen, J. Xie and S. Nie, Engineering, 2025, 54, 291, DOI:10.1016/j.eng.2025.06.042.
- S. N. Dean, M. A. Rimmer, K. B. Turner, D. A. Phillips, J. C. Caruana, W. J. Hervey IV, D. H. Leary and S. A. Walper, Front. Microbiol., 2020, 11, 710 CrossRef PubMed.
- V. Mahendrarajan and E. Nalini, J. Biomol. Struct. Dyn., 2025, 1–11 CrossRef PubMed.
- C. Ciobanasu, Antibiotics, 2025, 14, 414, DOI:10.3390/antibiotics14040414.
- Y. Fang, D. Fan, B. Feng, Y. Zhu, R. Xie, X. Tan, Q. Liu, J. Dong and W. Zeng, Bioact. Mater., 2025, 50, 510–524 CAS.
- M. Pirtskhalava, A. A. Amstrong, M. Grigolava, M. Chubinidze, E. Alimbarashvili, B. Vishnepolsky, A. Gabrielian, A. Rosenthal, D. E. Hurt and M. Tartakovsky, Nucleic Acids Res., 2020, 49, D288–D297 CrossRef PubMed.
- S. Thomas, S. Karnik, R. S. Barai, V. K. Jayaraman and S. Idicula-Thomas, Nucleic Acids Res., 2010, 38, D774–D780 CrossRef CAS PubMed.
- S. Gupta, P. Kapoor, K. Chaudhary, A. Gautam, R. Kumar and G. P. S. Raghava, PLoS One, 2013, 8, e73957 CrossRef CAS PubMed.
- I. Abdelbaky, M. Elhakeem, H. Tayara, E. Badr and M. Abdul Salam, BMC Bioinf., 2024, 25, 368 CrossRef CAS PubMed.
- C. H. Chen, M. C. R. Melo, N. Berglund, A. Khan, C. de la Fuente-Nunez, J. P. Ulmschneider and M. B. Ulmschneider, Curr. Opin. Struct. Biol., 2020, 61, 160–166 CrossRef CAS PubMed.
- M. Pirtskhalava, B. Vishnepolsky, M. Grigolava and G. Managadze, Pharmaceuticals, 2021, 14, 471, DOI:10.3390/ph14050471.
- A. Chaudhary, S. Bhalla, S. Patiyal, G. P. S. Raghava and G. Sahni, Heliyon, 2021, 7, e06668 CrossRef CAS PubMed.
- M. E. MacGilvray, J. D. Lapek, A. E. Friedman and R. G. Quivey, Microbiology, 2012, 158, 2133–2143 CrossRef CAS PubMed.
- K. Murzyn, T. Róg and M. Pasenkiewicz-Gierula, Biophys. J., 2005, 88, 1091–1103 Search PubMed.
- H. Itaya, K. Kasahara, Q. Xie, Y. Yano, K. Matsuzaki and T. Takahashi, ACS Omega, 2021, 6, 11458–11465 CrossRef CAS PubMed.
- R. Prados-Rosales, L. Brown, A. Casadevall, S. Montalvo-Quirós and J. L. Luque-Garcia, MethodsX, 2014, 1, 124–129 CrossRef PubMed.
- N. Hulstaert, J. Shofstahl, T. Sachsenberg, M. Walzer, H. Barsnes, L. Martens and Y. Perez-Riverol, J. Proteome Res., 2020, 19, 537–542 CrossRef CAS PubMed.
- L. Martens, M. Chambers, M. Sturm, D. Kessner, F. Levander, J. Shofstahl, W. H. Tang, A. Römpp, S. Neumann, A. D. Pizarro, L. Montecchi-Palazzi, N. Tasman, M. Coleman, F. Reisinger, P. Souda, H. Hermjakob, P.-A. Binz and E. W. Deutsch, Mol. Cell. Proteomics, 2011, 10, R110.000133, DOI:10.1074/mcp.R110.000133.
- S. Kim and P. A. Pevzner, Nat. Commun., 2014, 5, 5277 CrossRef CAS PubMed.
- J. E. Elias and S. P. Gygi, Nat. Methods, 2007, 4, 207–214 Search PubMed.
- H. L. Röst, T. Sachsenberg, S. Aiche, C. Bielow, H. Weisser, F. Aicheler, S. Andreotti, H.-C. Ehrlich, P. Gutenbrunner, E. Kenar, X. Liang, S. Nahnsen, L. Nilse, J. Pfeuffer, G. Rosenberger, M. Rurik, U. Schmitt, J. Veit, M. Walzer, D. Wojnar, W. E. Wolski, O. Schilling, J. S. Choudhary, L. Malmström, R. Aebersold, K. Reinert and O. Kohlbacher, Nat. Methods, 2016, 13, 741–748 CrossRef PubMed.
- A. S. Rathore, N. Kumar, S. Choudhury, N. K. Mehta and G. P. S. Raghava, Commun. Biol., 2025, 8, 176 Search PubMed.
- G. Wang, C. Schmidt, X. Li and Z. Wang, Nucleic Acids Res., 2025, 54, D363–D374 Search PubMed.
- A. Lamiable, P. Thévenet, J. Rey, M. Vavrusa, P. Derreumaux and P. Tufféry, Nucleic Acids Res., 2016, 44, W449–W454 Search PubMed.
- S. Feng, S. Park, Y. K. Choi and W. Im, J. Chem. Theory Comput., 2023, 19, 2161–2185 CrossRef CAS PubMed.
- J. E. Custer, B. D. Goddard, S. F. Matter and E. S. Kaneshiro, Lipids, 2014, 49, 543–554 Search PubMed.
- M. A. Bojanich and R. O. Calderón, Arch. Oral Biol., 2017, 81, 74–80 Search PubMed.
- J. C. Phillips, D. J. Hardy, J. D. C. Maia, J. E. Stone, J. V Ribeiro, R. C. Bernardi, R. Buch, G. Fiorin, J. Hénin, W. Jiang, R. McGreevy, M. C. R. Melo, B. K. Radak, R. D. Skeel, A. Singharoy, Y. Wang, B. Roux, A. Aksimentiev, Z. Luthey-Schulten, L. V Kalé, K. Schulten, C. Chipot and E. Tajkhorshid, J. Chem. Phys., 2020, 153, 44130 Search PubMed.
- J. Huang, S. Rauscher, G. Nawrocki, T. Ran, M. Feig, B. L. de Groot, H. Grubmüller and A. D. MacKerell, Nat. Methods, 2017, 14, 71–73 CrossRef CAS PubMed.
- D. Sengupta, H. Leontiadou, A. E. Mark and S.-J. Marrink, Biochim. Biophys. Acta, Biomembr., 2008, 1778, 2308–2317 Search PubMed.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graphics, 1996, 14, 33–38 CrossRef CAS PubMed.
- S. H. Lim, S. Mix, V. Anikst, I. Budvytiene, M. Eiden, Y. Churi, N. Queralto, A. Berliner, R. A. Martino, P. A. Rhodes and N. Banaei, Analyst, 2016, 141, 918–925 Search PubMed.
- V. Mahendrarajan, H. Lazarus and N. Easwaran, Heliyon, 2024, 10, e31898 CrossRef CAS PubMed.
- L. Andresen, T. Tenson and V. Hauryliuk, Sci. Rep., 2016, 6, 36549 CrossRef CAS PubMed.
- P. Virtanen, R. Gommers, T. E. Oliphant, M. Haberland, T. Reddy, D. Cournapeau, E. Burovski, P. Peterson, W. Weckesser, J. Bright, S. J. van der Walt, M. Brett, J. Wilson, K. J. Millman, N. Mayorov, A. R. J. Nelson, E. Jones, R. Kern, E. Larson, C. J. Carey, İ. Polat, Y. Feng, E. W. Moore, J. VanderPlas, D. Laxalde, J. Perktold, R. Cimrman, I. Henriksen, E. A. Quintero, C. R. Harris, A. M. Archibald, A. H. Ribeiro, F. Pedregosa and P. van Mulbregt, Nat. Methods, 2020, 17, 261–272 CrossRef CAS PubMed.
- V. Mahendrarajan, G. K. Muthukaliannan and N. Easwaran, Nat. Prod. Res., 2025, 1–8 Search PubMed.
- M. Zhang, L. Wang, W. Tang, Y. Xing, P. Liu and X. Dang, Pest Manage. Sci., 2023, 79, 3681–3692 Search PubMed.
- V. Mahendrarajan and N. Easwaran, Biofouling, 2025, 41, 92–102 Search PubMed.
- L. T. Nguyen, E. F. Haney and H. J. Vogel, Trends Biotechnol., 2011, 29, 464–472 Search PubMed.
- J. María Remes Troche, E. Coss Adame, M. Ángel Valdovinos Díaz, O. Gómez Escudero, M. Eugenia Icaza Chávez, J. Antonio Chávez-Barrera, F. Zárate Mondragón, J. Antonio Ruíz Velarde Velasco, G. Rafael Aceves Tavares, M. Antonio Lira Pedrín, E. Cerda Contreras, R. I. Carmona Sánchez, H. Guerra López and R. Solana Ortiz, Ther. Adv. Gastroenterol., 2020, 13, 1756284820971201 CrossRef PubMed.
- K. A. Brogden, Nat. Rev. Microbiol., 2005, 3, 238–250 CrossRef CAS PubMed.
- W. C. Wimley, ACS Chem. Biol., 2010, 5, 905–917 CrossRef CAS PubMed.
- L. Chen, W. Liu, Y. Li, S. Luo, Q. Liu, Y. Zhong, Z. Jian and M. Bao, Int. Immunopharmacol., 2013, 17, 108–115 CrossRef CAS PubMed.
- J.-L. Velasco-Bolom, G. Corzo and R. Garduño-Juárez, J. Biomol. Struct. Dyn., 2018, 36, 2070–2084 Search PubMed.
- C. D. Fjell, J. A. Hiss, R. E. W. Hancock and G. Schneider, Nat. Rev. Drug Discovery, 2011, 11, 37–51 CrossRef PubMed.
- W. C. Wimley and K. Hristova, J. Membr. Biol., 2011, 239, 27–34 CrossRef CAS PubMed.
- F. Lin, W. Jin, L. Yang, C. Tian, J. Zhang, Z. Jiao, J. Peng and G. Guo, BMC Microbiol., 2025, 25, 707 Search PubMed.
- I. D. Gbala, R. W. Macharia, J. L. Bargul and G. Magoma, Molecules, 2022, 27, 4325, DOI:10.3390/molecules27144325.
- P. Anjana and S. K. Tiwari, Front. Cell. Infect. Microbiol., 2022, 12, 851140 Search PubMed.
- A. Mazumdar, Y. Haddad, V. P. Sur, V. Milosavljevic, S. Bhowmick, H. Michalkova, R. Guran, R. Vesely and A. Moulick, Front. Microbiol., 2020, 11, 1963 CrossRef PubMed.
- L. A. Christersson, O. M. Norderyd and C. S. Puchalsky, J. Clin. Periodontol., 1993, 20, 88–95 Search PubMed.
- J. M. ten Cate, Odontology, 2006, 94, 1–9 Search PubMed.
- R. E. W. Hancock and H.-G. Sahl, Nat. Biotechnol., 2006, 24, 1551–1557 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |