 Open Access Article
Open Access ArticleFabrication and characterization of novel PLA/CH/GelMA/hydroxyapatite electrospun nanofibers for bone tissue regeneration
Mehmet Bozdagab,
Tubanur Avciab,
Sefa Izgorduab,
Dilruba Baykaraab,
Esra Pilavciab,
Burak Elemanc,
Zehra Kanlia,
Canan Dogana,
Denisa Ficaidefgh,
Anton Ficai *defgh and
Oğuzhan Gunduz*a
*defgh and
Oğuzhan Gunduz*a
aMarmara University, Faculty of Technology, Department of Metallurgical and Materials Engineering, Nanotechnology and Biomaterials Research Group, Istanbul, Turkiye
bDepartment of Metallurgical and Materials Engineering, Faculty of Technology, Marmara University, Turkiye
cMarmara University Innovation and Technology Transfer Application and Research Centre Directorate, Marmara University, Turkiye. Tel: +90 216 777 3956
dFaculty of Chemical Engineering and Biotechnologies, National University of Science and Technology Politehnica Bucharest, 1-7 Gh. Polizu St., 011061 Bucharest, Romania
eNational Centre for Micro- and Nanomaterials, National University of Science and Technology Politehnica Bucharest, 313 Independence Boulevard, 060042 Bucharest, Romania
fNational Centre for Food Safety, National University of Science and Technology Politehnica Bucharest, 313 Independence Boulevard, 060042 Bucharest, Romania
gAcademy of Romanian Scientists, 3 Ilfov St., 050045 Bucharest, Romania
hResearch Center for Advanced Materials, Products and Processes, National University of Science and Technology Politehnica Bucharest, 060042 Bucharest, Romania
First published on 20th February 2026
Abstract
This study focuses on the fabrication and comprehensive characterization of electrospun nanofibrous scaffolds composed of polylactic acid (PLA), chitosan (CH), gelatin methacryloyl (GelMA), and varying concentrations of hydroxyapatite (HAP) nanoparticles (0.1, 0.3, and 0.5% w/v) for potential applications in bone tissue engineering. Unlike previous studies utilizing binary or ternary systems, this work introduces a novel quadri-composite scaffold designed to overcome individual material limitations through synergistic effects: PLA provides the mechanical backbone, chitosan ensures hydrophilicity and antimicrobial potential, GelMA enhances cell adhesion via RGD motifs, and HAP serves as the osteoconductive mineral phase. The structural, thermal, mechanical, swelling, degradation, and biological properties of the nanofibers were investigated using SEM, FT-IR, DSC, and tensile testing. Among the tested formulations, the nanofibers containing 0.3% HAP exhibited the most balanced properties, with a tensile strength of 0.50 ± 0.18 MPa and elongation at break of 14.82 ± 6.93%, indicating optimal mechanical flexibility. These fibers also demonstrated improved thermal stability, controlled swelling behavior, and a degradation profile suitable for bone healing. In vitro biocompatibility studies using human osteoblast (hFOB) cells showed that while lower HAP content supported initial attachment, significantly higher cell viability was observed on the 0.3% HAP scaffolds at day 7 (p < 0.0365), supporting their superior long-term osteoblast proliferation. Overall, PLA/CH/GelMA nanofibers containing 0.3% HAP offer a promising strategy for bone regeneration due to their ECM-mimetic structure, tunable biodegradability, mechanical resilience, and bioactivity. Future studies will focus on in vivo evaluation of these scaffolds for bone defect repair.
Introduction
Bones, which play crucial roles in protection, support, movement, and hematopoiesis, can experience a loss of tissue integrity due to factors such as trauma, infection, and tumor removal. Generally, the regenerative capacity of bone tissue is limited, necessitating medical intervention for the healing of bone defects.1Research indicates that bone grafts are among the most common tissue transplantations. Worldwide, the number of orthopedic operations conducted exceeds millions. However, bone grafts involve multiple surgeries and carry the risk of complications and patient trauma. Consequently, there has been a growing focus on the development of artificial bone substitute materials and alternative methods to minimize complications in recent years.2 One such method under investigation is electrospinning, which is a simple, cost-effective, and reproducible process.3 Compared to other fabrication strategies such as solvent casting, phase separation, or 3D printing, electrospinning offers unique advantages including the ability to produce fibers with a high surface area-to-volume ratio and remarkable porosity that closely mimics the native extracellular matrix (ECM) architecture. These structural features promote efficient nutrient transport and cell signaling. However, limitations regarding pore size for deep cellular infiltration and the need for precise control over fiber alignment remain challenges that necessitate careful material selection and parameter optimization (Ref needed: add the “Smart Materials in Medicine” and “Biofabrication” papers suggested by Referee 2 here). Moreover, different components, including natural or synthetic polymers, inorganic ceramics, and composites, can be combined in this method.2 These properties make fibers with mechanical properties and morphology similar to the extracellular matrix (ECM) of native tissue a suitable option for bone substitutes.3,4 Poly(lactic acid) (PLA) is one of the various biodegradable polymers for its application in bone tissue engineering. It is biodegradability, good mechanical strength, and cost-effectiveness, relative to other biodegradable polymers, make PLA a widely utilized material.5,6 However, PLA's inherent hydrophobicity, lack of reactive side chains, and absence of cell recognition sites often limit its initial cell adhesion and biological interaction, necessitating its blending with bioactive polymers to improve the biological response. Chitosan is composed of chitin, a natural polysaccharide found in lobsters, crabs, shrimps, jellyfish, butterflies, ladybugs, and fungi. However, shellfish shells are popularly used as the main source of chitosan production.7 Chitosan has recently been used in many biomedical studies. This can be explained by the fact that chitosan is a non-toxic, bio-adhesive, biocompatible, and biodegradable component that introduces hydrophilic groups and antimicrobial properties to the scaffold system, countering the inert nature of synthetic polymers.8,9 To further enhance the biological performance and mimic the soft hydrogel-like component of the ECM, incorporating chemically modified proteins is essential. GelMA or gelatin methacryloyl is a photosensitive hydrogel that is involved in various biomedical studies due to its biocompatibility, biodegradability, and cost-effective solution.10 GelMA is obtained by subjecting gelatin to the methacrylation process, preserving the RGD (arginine–glycine–aspartic acid) sequences essential for cell attachment while allowing for photocrosslinking to maintain structural stability in aqueous environments.11 Some components can be used to reinforce GelMA during the studies. These include biopolymers, synthetic polymers, and inorganic particles. For example, hydroxyapatite particles can be used with GelMA to increase osteoinductivity.12 Hydroxyapatite (HAP) is the main mineral structure in teeth and bones and is one of the most preferred bioceramic components in bone regeneration processes (Pedrosa et al., 2021).13 The incorporation of HAP nanoparticles into polymer matrices is a widely adopted strategy to mimic the mineral phase of bone, thereby enhancing the mechanical stiffness and promoting osteogenic differentiation of cells.14,15 However, the use of HAP alone is limited by its brittleness and difficulty in shaping into fibrous structures, which underscores the necessity of combining it with flexible polymeric matrices to create composite scaffolds that possess both toughness and bioactivity. Despite the advantages of these individual polymers, binary or ternary combinations often fail to fully satisfy the complex requirements of bone tissue engineering, which demands a delicate balance of mechanical stiffness, osteoinductivity, and controlled degradation. While PLA provides the necessary mechanical strength, it typically lacks sufficient biological recognition sites; GelMA supports cell adhesion but lacks load-bearing capacity; and chitosan contributes to hydrophilicity and structural stability but can exhibit rapid degradation rates alone. Therefore, the design of a novel quadri-composite scaffold by integrating HAP into this tri-polymer system is proposed to impart osteoconductivity and mechanical reinforcement, aiming to create a balanced material that overcomes the limitations of single or dual-component systems.
In this work, a novel quadri-composite system comprising PLA/CH/GelMA electrospun nanofibers containing different concentrations of HAP (0.1, 0.3, and 0.5% w/v) was developed and characterised to evaluate their potential as bone tissue scaffolds, as Fig. 1 presents. Unlike previous studies that utilized these materials separately or in limited combinations, this study aims to exploit the synergistic effects of this specific quadruple formulation: PLA for the backbone, chitosan for hydrophilicity, GelMA for enhanced cell adhesion, and HAP for osteogenic mimicry. It was hypothesized that an optimal HAP concentration would yield a scaffold with enhanced bioactivity, mechanical integrity, and degradation behaviour, suitable for supporting osteoblast proliferation and bone regeneration. The fabricated scaffolds were systematically evaluated through morphological (SEM), chemical (FT-IR, 1H-NMR), thermal (DSC), mechanical, swelling, degradation, and in vitro biocompatibility analyses using human osteoblast (hFOB) cells.
Experimental
Chemicals
Polylactic acid (PLA; molecular weight [MW] = 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Da), chitosan (CS; MW = 50
000 Da), chitosan (CS; MW = 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–190
000–190![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Da), gelatin type A obtained from porcine skin, and hydroxyapatite (powder, >97%) were purchased from Sigma-Aldrich (USA). Acetone (CH3COCH3, >99.8%), formic acid (HCOOH, 98–100%), and chloroform (CHCl3) were obtained from Merck Chemicals (Germany).
000 Da), gelatin type A obtained from porcine skin, and hydroxyapatite (powder, >97%) were purchased from Sigma-Aldrich (USA). Acetone (CH3COCH3, >99.8%), formic acid (HCOOH, 98–100%), and chloroform (CHCl3) were obtained from Merck Chemicals (Germany).
GelMA synthesis
A 0.1 M carbonate bicarbonate buffer (0.1 M CB buffer, 3.18 g sodium carbonate, and 5.86 g sodium bicarbonate in 1 L of distilled water, pH 9) was used to prepare a 10% solution of type A gelatin. After the solution became homogeneous at 60 °C, methacrylic anhydride (MAA) (0.1 mL per gram of gelatin) was added dropwise to the solution and allowed to react at 50 °C with vigorous stirring for 3 h. After that, the pH was adjusted to 7.4 to end the methacryloyl modification reaction. To remove unreacted MAA, the final solution was dialyzed against distilled water for 4 days at 40 °C, using a dialysis membrane (14 kDa molecular-weight-cutoff (MWCO)). The GelMA solution was lyophilized for 3 days. Until further use, dried foam-like GelMA was stored at −20 °C.Preparation of the solutions for electrospinning of nanofibers
First, a base solution was prepared by dissolving 12% PLA in a solvent mixture formic acid, chloroform, and acetone (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) at 70 °C for 1 hour using a magnetic stirrer. Then 2% chitosan was added to the solution and stirred for an additional 30 minutes to obtain a homogeneous solution. Once the polymers were completely dissolved, 5% GelMA and 5 mg of LAP as photoinitiator were incorporated to the structure, the beaker was covered with a light-proof cover and stirred for another hour using a magnetic stirrer. Finally, designated amounts of HAP at concentration 0.1, 0.3, and 0.5% (w/v) were added to the solution and stirring was continued for another 15 minutes to complete the preparation. Table 1 below summarizes the solution samples prepared in total.
2) at 70 °C for 1 hour using a magnetic stirrer. Then 2% chitosan was added to the solution and stirred for an additional 30 minutes to obtain a homogeneous solution. Once the polymers were completely dissolved, 5% GelMA and 5 mg of LAP as photoinitiator were incorporated to the structure, the beaker was covered with a light-proof cover and stirred for another hour using a magnetic stirrer. Finally, designated amounts of HAP at concentration 0.1, 0.3, and 0.5% (w/v) were added to the solution and stirring was continued for another 15 minutes to complete the preparation. Table 1 below summarizes the solution samples prepared in total.
| Solution | Formic acid–chloroform–acetone | Polylactic acid (PLA) | Chitosan (CH) | Gelatin methacryloyl (GelMA) | Hydroxyapatite (HAP) |
|---|---|---|---|---|---|
| Solution-1 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
12% | — | — | — |
| Solution-2 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
12% | 2% | — | — |
| Solution-3 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
12% | 2% | 5% | — |
| Solution-4 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
12% | 2% | 5% | 0.1% |
| Solution-5 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
12% | 2% | 5% | 0.3% |
| Solution-6 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
12% | 2% | 5% | 0.5% |
Fabrication of PLA/CH/GelMA/HAP nanofibers
A laboratory-scale electrospinning device (NS24, Invens Co., Turkey) was used to fabricate the nanofibers at designated concentrations. Each prepared solution was individually loaded into syringes, which were connected to the lab-scale pumping system and go through an optimization process to determine the optimum voltage and flow rate. With the flow rate set at 0.45 mL h−1, nanofibers were deposited onto a rotating collector positioned 12 cm away from the needle, spraying the solution samples on collector. Before the fabrication, the rotating collector was covered with wax paper, and weights were recorded for each trial. Table 2 presents the voltage and flow rate parameters for each successfully fabricated sample.| Sample | Voltage (kV) | Flow rate (mL h−1) |
|---|---|---|
| PLA | 26 | 0.4 |
| PLA/CH | 27.5 | 0.3 |
| PLA/CH/GelMA | 27.5 | 0.45 |
| PLA/CH/GelMA/HAP (0.1, 0.3, and 0.5% (w/v)) | 27 | 0.60 |
Scanning electron microscopy (SEM)
SEM (EVO LS 10, Zeiss) was utilized to analyze the nanofibrous patches' surface morphologies. Using spray coating equipment (Quorum SC7620, USA), a layer of palladium and gold was first applied to the surface of the nanofibers for 120 seconds. Additionally, histogram graphs were created by measuring 100 fibers using imaging software (Olympus AnalySIS, USA) in accordance with the SEM data that were obtained.Fourier transform infrared spectroscopy (FT-IR)
The FTIR (Jasco FT/IR-4700 model machine) was performed to examine the functional groups and physicochemical structures of nanofibers. At room temperature (23 °C), the spectra were acquired in transmission mode, with a scanning range of 4000–400 cm−1, a resolution of 4 cm−1, and a scanning speed of 32.1H-NMR analysis
The functionalization of gelatin with methacryloyl groups was investigated by 1H NMR spectroscopy (Bruker Avance III 600 MHz, Bremen, Germany). For this analysis, gelatin and GelMA samples were separately dissolved in D2O at a concentration of 10 mg L−1. The spectra were recorded at 400 MHz and at room temperature. To determine the degree of substitution (DS), the proton signals of the methacrylate groups were compared with the lysine methylene signals of gelatin. The DS was calculated using the following equation:Differential scanning calorimetry (DSC)
The thermal properties such as glass transition temperature and melting temperature of all nanofibers were analysed by means of differential scanning calorimetry (DSC, Shimadzu). The temperature ranges were set from 20 to 400 °C and the heating rate was 10 °C min−1 during the analysis.Mechanical properties of PLA/CH/GelMA/HAP nanofibers
A tensile testing apparatus (Shimadzu Corporation, EZ-LX, Kyoto, Japan) was used to ascertain the mechanical properties. Nanofibers measuring 10 × 50 mm were cut for the test. The thickness of the nanofibers was measured prior to testing using a digital micrometer (Mitutoyo MTI Corp., USA). A 5 kN load cell was applied during the test, and the test speed was set to 5 mm min−1. Each group was conducted three times to ensure accuracy.Swelling and degradation behavior of PLA/CH/GelMA/HAP nanofibers
Measurements of swelling and degradation were performed to ascertain the nanofibers' ability to absorb water as well as how their mass changed over time. For the swelling tests, phosphate-buffered saline (PBS) with a pH of 7.4 was utilized. The initial weights of the nanofibers with equal weight were noted during this test. Following that, the nanofibrous patches were placed in Eppendorf tubes with one milliliter of PBS and maintained at 37 °C in a thermal shaker (BIOSAN TS-100). The samples were taken out of the extra water every 24 hours, and their wet weights were recorded. In order to calculate the swelling value, utilize eqn (1),16
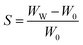 | (1) |
For the degrading procedure, nanofibers were weighed equally and put in 1 mL Eppendorf tubes with PBS in a thermal shaker device. The samples and PBS were separated after a day, and the nanofibers were dried for twenty-four hours in a thermal heater. Eqn (2),16
 | (2) |
Cell viability of human osteoblast (hFOB) cell lines
The biocompatibility of the fabricated nanofiber scaffolds was evaluated using the human fetal osteoblast cell line hFOB 1.19 (CRL-11372™, ATCC, Maryland, USA). Prior to testing, all biomaterial samples were sterilized under ultraviolet (UV) light for 2–3 hours. Cell viability was assessed using the MTT assay (Sigma-Aldrich, Darmstadt, Germany). hFOB cells were seeded into 96-well plates at a density of 5 × 103 cells per well in DMEM/F-12 medium supplemented with 10% fetal bovine serum (FBS) and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 U mL−1 penicillin/streptomycin. Each experimental group was tested in triplicate, with the different biomaterial samples added separately into the wells. Cells were incubated at 37 °C in a humidified atmosphere containing 5% CO2. MTT assays were performed on days 1, 3, and 7 after cell seeding. On the respective days, the culture medium was aspirated and replaced with 100 µL of fresh medium. Subsequently, 10 µL of MTT stock solution (5 mg mL−1) was added to each well and incubated for 4 hours at 37 °C to allow for formazan crystal formation. After incubation, 100 µL of dimethyl sulfoxide (DMSO) was added to dissolve the formazan crystals, followed by an additional 10 minute incubation at 37 °C with gentle shaking. The optical density (OD) was measured at 570 nm using a Multiskan SkyHigh Microplate Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
000 U mL−1 penicillin/streptomycin. Each experimental group was tested in triplicate, with the different biomaterial samples added separately into the wells. Cells were incubated at 37 °C in a humidified atmosphere containing 5% CO2. MTT assays were performed on days 1, 3, and 7 after cell seeding. On the respective days, the culture medium was aspirated and replaced with 100 µL of fresh medium. Subsequently, 10 µL of MTT stock solution (5 mg mL−1) was added to each well and incubated for 4 hours at 37 °C to allow for formazan crystal formation. After incubation, 100 µL of dimethyl sulfoxide (DMSO) was added to dissolve the formazan crystals, followed by an additional 10 minute incubation at 37 °C with gentle shaking. The optical density (OD) was measured at 570 nm using a Multiskan SkyHigh Microplate Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
Statistical analysis
Data were analyzed using two-way ANOVA with Tukey's multiple comparison test, based on n = 3 independent experiments and p < 0.05 was considered statistically significant.Results and discussion
Morphology and fiber diameter (SEM)
SEM images of the nanofibers at different magnifications, along with histogram graphs, are shown in Fig. 2(A1 and A2), PLA nanofiber exhibits a smooth surface and beadles' morphology. It has fiber diameter approximately 270 nm, which is consistent with findings by Duygulu et al., who observed bead-free PLA fibers with average diameters of 300 nm depending on polymer concentration during electrospinning (5–8% w/v PLA).17 In Fig. 2(B1 and B2), PLA/CH nanofiber also exhibit a smooth morphology with a fiber diameter approximately 240 nm. For the PLA/CH/GelMA nanofibers shown in Fig. 2(C1 and C2), the fiber diameter increases significantly to about 700 nm. Pilavci et al. (2023) reported that the addition of GelMA resulted in a deterioration of nanofiber morphology.18,19 Fig. 2(D1 and D2), represents PLA/CH/GelMA/0.1% HAP nanofiber where the addition of HAP decreases to nearly 650 nm. HAP particles were visible in the SEM images. In Fig. 2(E1 and E2), the PLA/CH/GelMA/0.3% HAP nanofiber group has a smaller fiber diameter than PLA/CH/GelMA and PLA/CH/GelMA/0.1% HAP nanofiber groups. It has a fiber diameter of approximately 450 nm. Fig. 2(F1 and F2) depicts the PLA/CH/GelMA/0.5% HAP nanofiber group with a fiber diameter of roughly 400 nm. All nanofiber groups display a beadless morphology. In the groups containing HAP, polymeric particles could be clearly detected from the images. The addition of HAP led to a proportional decrease in fiber diameters. In the study by Stella et al., they produced nanofibers containing different concentrations of HAP–HPC/PLA, reported that the electrospun mats with higher HAP content showed a smaller fiber diameter, while the mats without HAP showed a larger fiber diameter.20 This trend aligns with Kotrotsos et al., who showed that adding HAP to PLA scaffolds during electrospinning decreased fiber diameters and enhanced scaffold uniformity.21 In particular, while the PLA/CH/GelMA nanofibers exhibited a broader diameter range due to the increased fiber thickness, the addition of HAP at different concentrations resulted in a more consistent diameter distribution. This indicates that HAP acts as a structural stabilizer during the electrospinning process, preventing excessive fiber thickening and potentially improving the uniformity of the scaffold, which is crucial for consistent cell–material interactions.22,23 Similar findings were reported by Stella et al. (2022a), where higher HAP concentrations contributed to reduced fiber diameter variability, enhancing the homogeneity of the electrospun mats.20,24Chemical structure (FT-IR and 1H-NMR)
The FTIR measurements were performed to analyze the molecular structure of nanofibers. In Fig. 3A(a), for pure PLA characteristic peaks from the backbone ester group, appeared at 1751, 1182, 1128, and 1085 cm−1. For pure chitosan, in Fig. 3A(b), characteristic peaks from amide I and amide II adsorption bands occurred at 1646 cm−1 and 1529 cm−1, and 1373 cm−1 peaks corresponds to the amide III adsorption band.25 For pure gelatin, in Fig. 3A(c), N–H bending is observed 1519 cm−1 and C–O stretching occurs at 1627 cm−1.26 In case of pure gelatin methacryloyl, GelMA, in Fig. 3A(d), the adsorption bands from amide I, II and III are observed at 1627, 1546, and 1452 cm−1, respectively.27 For pure hydroxyapatite, in Fig. 3A(e), the functional groups PO43− and HPO42− exhibit bending at 599 and 629 cm−1 and stretching at 962 and 1022 cm−1.28 For the nanofibers, all samples shown in Fig. 3B(f–j), shows the resemblance of pure PLA and chitosan. After the addition of GelMA to PLA/CH, characteristic peaks from GelMA are seen at 1540 and 1646 cm−1 in all samples, Fig. 3B(f and h–j), corresponding to amide I and amide II, respectively. In samples containing HAP, Fig. 3B(h–j), the stretching at 1041 cm−1 is also observed which belongs to the PO43−, P–O bond in triply degenerate asymmetric stretching.29 The strong shift of the peak from 1020–1030 cm−1 specific to the stoichiometric/nonstoichiometric HAP to 1041 cm−1 is a proof that HAP strongly interact with the organic phase, especially with the gelatin phase. 1H-NMR analysis was employed to quantify the degree of substitution (DS) of GelMA, which is a critical parameter for photocrosslinking efficiency. As detailed in the Experimental section, the integration of the methacrylate proton peaks (at δ = 5.4 and 5.7 ppm) relative to the lysine methylene peaks indicated a DS of 75.4%. Previous studies have shown that adjusting the DS allows for the tailoring of hydrogel stiffness to favor osteogenic differentiation.30 Furthermore, degrees of functionalization in the range of 70–95% have been reported to support high mesenchymal stem cell viability and proliferation.31 Therefore, the obtained DS of 75.4% represents an optimal modification level, ensuring stable network formation without compromising bioactivity.Thermal properties (DSC)
Thermal measurements are essential for understanding and determining the chemical and physical alterations in the materials under investigation, such as phase transition, glass transition temperature (Tg), and melting temperature (Tm).9 Therefore, thermal characteristics and heating processes of all nanofibers were analyzed using Differential Scanning Calorimetry (DSC) and results were shown in Fig. 3(C).According to the results, all nanofibers exhibited three main peaks at approximately 155, 308 and 350 °C with findings by in agreement with DSC profiles reported for PLA-based composites.32,33 It can be said that these peaks belong to the PLA. In other words, it is the primary component with the highest concentration in all manufactured nanofibers, is mainly responsible for the peaks observed in the DSC analysis. The peak at 155 °C corresponds to the melting temperature (Tm) of PLA,34 while the other peaks demonstrated the degradation temperature between 340 and 360 °C, accompanied by significant polymer volatilization due to the thermal decomposition of the polymer backbone.35 It can be said that all DSC curves were roundly similar. However, the added materials affect the melting and decomposition temperatures with a little bit change. In other words, added materials increase the thermal stability of the PLA by raising its decomposition of temperature. This effect is particularly evident in the sharp peak observed for PLA/CH/GelMA/0.5 HAP. These alterations could be attributed to interactions between the components36 as also supported by the shifts recorded in FTIR.
Mechanical properties
The mechanical properties PLA/CH/GelMA, and PLA/CH/GelMA/HAP samples were investigated to see the durability and resistance of nanofibers. Each tested sample was measured and analyzed in terms of both tensile strength and elongation at break, with the results shown in Fig. 4. The PLA/CH/GelMA nanofiber exhibit tensile strength of 0.84 ± 0.37 MPa and the strain at break value 3.24 ± 0.33%. With addition of HAP to the nanofiber matrix, the PLA/CH/GelMA/HAP-0.1% sample exhibited a tensile strength of 0.78 ± 0.17 MPa and a strain at break value of 6.50 ± 1.64%. The PLA/CH/GelMA/HAP-0.3% and 0.5% samples exhibited similar tensile strength results to the 0.1 sample. For PLA/CH/GelMA/HAP-0.3, a tensile strength of 0.50 ± 0.18 MPa and a strain at break value of 14.82 ± 6.93% was observed, highlighting the classic trade-off between stiffness and flexibility seen in composite electrospun scaffolds.37 Similarly, for the PLA/CH/GelMA/HAP-0.5 sample, a tensile strength of 0.80 ± 0.10 MPa and a strain at break value 3.71 ± 0.44% were recorded. These results indicate that increasing the HAP content enhances the flexibility of the nanofibers up to a certain threshold, as observed in the PLA/CH/GelMA/HAP-0.3 sample, which exhibited the highest elongation at break.38 The observed variations in standard deviation are consistent with the inherent challenges of testing electrospun mats. Previous studies have explicitly stated that the relatively large standard deviations in randomly oriented fiber mats are due to the inherent randomness in the direction of the fiber axes relative to the tension vector.39 Additionally, the wider spread of nanofiber angles in non-aligned sheets creates variable load-bearing paths, which is a key factor contributing to the statistical dispersion of mechanical properties compared to aligned scaffolds.40 Therefore, the reported deviations reflect the structural heterogeneity typical of multi-component electrospun nanocomposites rather than experimental inconsistency.41 However, further increasing the HAP concentration to 0.5% resulted in a decrease in strain at break, suggesting that excessive HAP content may lead to agglomeration and structural inconsistencies. This, in turn, affects mechanical integrity by acting as a stress focal point on fiber cracks.42,43 While HAP is primarily added for its bioactivity rather than mechanical reinforcement, these results—consistent with baseline PLA–ceramic composites reaching up to ≈8 MPa tensile strength when well dispersed—underscore the need to optimize HAP loading. Too little HAP underutilizes its toughening potential, whereas too much promotes agglomeration and diminishes both strength and ductility.44,45Swelling and degradation behavior
Swelling and degradation experiments of the produced nanofiber composites were conducted in a pH 7.4 phosphate buffer solution at 37 °C over a period of 192 hours (8 days). Fig. 5(A) shows that PLA–CS nanofibers exhibit a swelling profile up to 120 hours, after which degradation begins. When GelMA is incorporated into PLA–CS nanofibers, the porous structure leads to a higher swelling profile compared to PLA–CS, resulting in an accelerated degradation process.46,47The addition of HAP to the nanofiber has been observed to enhance its hydrophilic properties due to the presence of hydroxyl (OH−) functional groups and a larger pore size, thereby increasing its water absorption capacity. At a 0.1% HAP concentration, water absorption capacity increased between 24 and 72 hours, followed by slow degradation at the end of this period. On the other hand, increasing the HAP concentration to 5% caused agglomeration on the nanofiber surface leading to a decrease in porosity and a reduction in the scaffold's water absorption capacity.48–50 At a 0.3% HAP concentration, hydrophilic properties increased without significant agglomeration, resulting in the fastest degradation among the tested concentrations. At both 0.5% and 0.3% HAP concentrations, the maximum swelling rate was reached at 24 hours. However, at the 0.3% concentration, rapid degradation was observed by the end of the 24 hour period.
This rapid degradation can be explained by the hydrolytic mechanism typical of PLA-based composites. Previous studies have established that hydrophilic components, such as chitosan and hydroxyapatite, facilitate water penetration into the polymer matrix, thereby accelerating the hydrolytic cleavage of ester bonds.51,52 Specifically, PLA/HAP composites have been reported to exhibit greater in vitro weight loss compared to pure polymers due to this enhanced water uptake and increased surface area for hydrolysis.53 Furthermore, the slight mass fluctuations observed in Fig. 5(B) for HAP-containing groups during later incubation periods could be attributed to the bioactivity of the scaffolds. It is well-documented that in physiological fluids, calcium and phosphate ions can precipitate onto the surface of HAP-containing fibers, initiating a mineralization process that may partially offset the mass loss derived from polymer degradation.53,54 Crucially, studies have confirmed that this apatite formation and specific mineralization kinetics occur even in PBS environments, as the ceramic components dissolve and reprecipitate.55,56
Degradation is crucial in bone tissue engineering, as it allows implanted bone grafts or implants to gradually resorb in the body, facilitating the formation of new bone tissue. The molecular weight, structure, composition, and crystallinity of the polymer can affect its degradation. High concentrations of low molecular weight polymers can lead to rapid degradation.20,48 Before the addition of HAP, an increasing degradation rate was observed in the lower molecular weight nanofiber composite, with the highest degradation rate of 27% occurring at the end of the 192 hour, at Fig. 5(B). Generally, as the concentration of HAP, which consists of minerals with higher molecular weight, is increases, Ca2+ and PO43− ion exchange occurs between the polymer and HAP.20 As a result of the mineralization process that occurs when ions enter the pores, ion exchange decreases, and the degradation rate may also decrease. Degradability is closely related to the swelling process. At 0.1% HAP concentration, the degradation rate remained constant until the 72nd hour and then increased, confirming the increasing swelling rate over 72 hours, followed by the onset of degradation. As the HAP concentration increases to 0.5%, ionization stabilizes, and degradation decreases, as evinced by the 24% degradation rate at the 96th hour.
Cell viability and biocompatibility
To assess cell viability, an MTT assay was performed to evaluate the interaction of hFOB cells with PLA/CH/GelMA nanofiber scaffolds and PLA/CH/GelMA/HAP nanofiber scaffolds containing different concentrations of hydroxyapatite (HAP) over time, as in Fig. 6.On day 1, no statistically significant increase in cell viability was observed in the HAP-containing groups compared to the PLA/CH/GelMA group (compared to the PLA/CH/GelMA nanofiber group, the HAP-containing groups (0.1%, 0.3%, and 0.5%) showed no statistically significant differences, p < 0.3303; p < 0.7990; p < 0.9992; respectively). This finding suggests that the presence of HAP did not markedly affect early cell proliferation rates. However, all scaffold formulations were found to be non-cytotoxic and biocompatible at the initial stage of cell culture.
On day 3, the PLA/CH/GelMA/HAP nanofibers containing 0.1% HAP exhibited significantly higher cell viability compared to the PLA/CH/GelMA scaffold (*p < 0.0396). This biphasic trend can be attributed to the surface topography and fiber diameter of the scaffolds. Previous studies indicate that surface roughness and fiber diameter are critical regulators of osteoblast adhesion and spreading.57,58 While nanoscale roughness generally promotes adhesion, excessive agglomeration seen at higher concentrations can create irregular heterogeneous surfaces that may initially hinder optimal cell spreading compared to the more uniform distribution observed at lower HAP concentrations (0.1%).59,60 Thus, the superior viability of the 0.1% group at day 3 suggests that its surface topography provided the most favorable environment for initial cellular attachment.
However, by day 7, overall cell viability had increased in all nanofiber scaffold groups. Notably, the PLA/CH/GelMA/HAP scaffold containing 0.3% HAP showed a significantly higher level of cell proliferation than the PLA/CH/GelMA group (**p < 0.0076). This shift indicates that once initial adhesion is established, the biological effects of the released ions become the dominant factor driving proliferation. The 0.3% HAP concentration likely provides a more optimal release profile of calcium and phosphate ions, which are known to act as bioactive signaling molecules. Specifically, elevated extracellular calcium levels have been shown to sustain the activation of the ERK signaling pathway, which directly stimulates osteoblast proliferation.61,62
The sample with 0.5% HAP also showed viability very close to the sample PLA/CH/GelMA/HAP (0.3%), confirming that a higher mineral content supports long-term viability. Although a slight decrease or plateau in viability was observed after day 7 in some groups, this is a common phenomenon in static cell culture conditions. Limitations in mass transport can lead to nutrient depletion or hypoxia in the center of the scaffold as cell density increases, which may result in reduced metabolic activity or contact inhibition rather than material cytotoxicity.63,64
These findings are consistent with previous studies showing that polymeric scaffolds containing hydroxyapatite (HAP) enhance osteoblast proliferation and support bone tissue regeneration due to their excellent bioactivity and biocompatibility.
For example, Chuenjitkuntaworn et al., reported that scaffolds composed of polycaprolactone (PCL) and HAP supported cell growth and osteogenic differentiation in cultures with bone marrow-derived mesenchymal stem cells (BMSCs), dental pulp stem cells (DPSCs), and adipose-derived mesenchymal stem cells (ADSCs). DPSCs showed the highest calcium deposition on PCL/HAP scaffolds, indicating that these scaffolds are promising candidates for bone tissue engineering.20 Similarly, Subuki et al., stated that biodegradable scaffolds made from natural polymers and HAP, due to their high biocompatibility and bioactivity, are considered promising materials for bone tissue engineering applications. These composite scaffolds mimic the natural structure and composition of bone tissue, promoting cell adhesion, proliferation, and osteogenic differentiation.20
Conclusions
In this study, a novel quadri-composite nanofibrous scaffold comprising PLA, chitosan, GelMA, and HAP was successfully fabricated via electrospinning for bone tissue engineering applications. Unlike binary or ternary systems, this specific combination was designed to leverage synergistic effects: PLA provided mechanical integrity, chitosan introduced hydrophilicity, GelMA offered RGD-mediated cell attachment, and HAP imparted osteoconductivity.Characterization results confirmed the effective incorporation of GelMA and hydroxyapatite, with SEM revealing a reduction in fiber diameter to ∼400 nm at 0.5% HAP content, indicating that HAP acted as a structural stabilizer improving fiber uniformity. FT-IR and 1H-NMR analyses verified the presence of characteristic functional groups, with a GelMA substitution degree of 75.4% confirming the successful modification for stable network formation. Thermal analysis revealed increased decomposition temperatures with higher HAP levels, indicating improved thermal stability due to strong organic–inorganic interactions.
Mechanical testing demonstrated that the inclusion of 0.3% HAP significantly enhanced the flexibility of the scaffolds, exhibiting the highest strain at break (14.82 ± 6.93%) and a tensile strength of 0.50 ± 0.18 MPa. This highlights a critical balance where 0.3% HAP optimizes flexibility without causing the agglomeration-induced brittleness observed at higher concentrations. Degradation studies demonstrated a mass loss of approximately 27% by 192 hours (8 days) in the 0.3% HAP group, where the hydrolytic degradation was partially offset by surface mineralization, confirming the scaffold's bioactivity in physiological environments.
In vitro biocompatibility tests using hFOB cells showed a time-dependent response: while lower HAP concentrations favored initial adhesion due to surface topography, the 0.3% HAP formulation significantly enhanced long-term proliferation (p < 0.0076) after 7 days, likely driven by the sustained release of bioactive ions. Overall, PLA/CH/GelMA nanofibers containing 0.3% HAP demonstrated optimal mechanical flexibility, biocompatibility, and degradation behavior. While this study established the structural and initial biological suitability of the scaffolds, future studies can focus on quantitative osteogenic differentiation assays and in vivo models to further validate their regenerative potential for bone defect repair.
Author contributions
Mehmet Bozdag: methodology, investigation, writing – original draft, writing – review & editing. Tubanur Avci: methodology, investigation, writing – original draft. Dilruba Baykara: methodology, investigation, writing – original draft. Sefa Izgordu: conceptualization, methodology, writing – original draft. Esra Pilavci: methodology, investigation, writing – original draft. Burak Eleman: methodology, investigation, writing – original draft. Zehra Kanli: methodology, investigation, writing – original draft. Canan Ekinci Dogan: investigation, conceptualization, writing – review & editing. Denisa Ficai: conceptualization, writing – review & editing. Anton Ficai: conceptualization, writing – review & editing. Oguzhan Gunduz: project administration, writing – review & editing.Conflicts of interest
The authors declare that there are no conflicts of interest.Data availability
The data that support the findings of this study are included in this published article.Acknowledgements
The authors would like to thank Marmara University for providing laboratory facilities and technical support. The APC was supported within the Transformative Agreements 2025 supported by Anelis-Plus.Notes and references
- X. Yan, H. Yao, J. Luo, Z. Li and J. Wei, Polymers, 2022, 14, 2940 CrossRef CAS PubMed.
- T. Zhao, J. Zhang, X. Gao, D. Yuan, Z. Gu and Y. Xu, J. Mater. Chem. B, 2022, 10, 6078–6106 Search PubMed.
- G. G. Flores-Rojas, B. Gómez-Lazaro, F. López-Saucedo, R. Vera-Graziano, E. Bucio and E. Mendizábal, Macromol, 2023, 3, 524–553 CAS.
- M. P. Prabhakaran, J. Venugopal and S. Ramakrishna, Acta Biomater., 2009, 5, 2884–2893 CrossRef CAS PubMed.
- C. Hwang, S. Park, I. G. Kang, H. E. Kim and C. M. Han, Mater. Sci. Eng., C, 2020, 115, 111112 CrossRef CAS PubMed.
- D. A. Canales, F. Reyes, M. Saavedra, L. Peponi, A. Leonés, H. Palza, A. R. Boccaccini, A. Grünewald and P. A. Zapata, Int. J. Biol. Macromol., 2022, 210, 324–336 CrossRef CAS PubMed.
- J. Venkatesan and S.-K. Kim, Mar. Drugs, 2010, 8, 2252–2266 CrossRef CAS PubMed.
- K. Kalantari, A. M. Afifi, H. Jahangirian and T. J. Webster, Carbohydr. Polym., 2019, 207, 588–600 Search PubMed.
- M. Bozdag, F. Urek, S. Cesur, A. Sahin and O. Gunduz, J. Drug Delivery Sci. Technol., 2024, 106, 106712 CrossRef.
- K. Man, I. A. Barroso, M. Y. Brunet, B. Peacock, A. S. Federici, D. A. Hoey and S. C. Cox, Int. J. Mol. Sci., 2022, 23, 832 CrossRef CAS PubMed.
- R. Augustine, V. K. Nikolopoulos and G. Camci-Unal, Bioengineering, 2023, 10, 854 CrossRef CAS PubMed.
- S. O. L. de Souza, S. M. de Oliveira, L. M. Silva and R. L. Oréfice, J. Appl. Polym. Sci., 2022, 139(28), e52527 Search PubMed.
- M. C. G. Pedrosa, S. A. dos Anjos, E. Mavropoulos, P. L. Bernardo, J. M. Granjeiro, A. M. Rossi and M. L. Dias, J. Bioact. Compat. Polym., 2021, 36, 314–333 Search PubMed.
- S. R. Gavinho, M. Bozdag, C. Kalkandelen, J. S. Regadas, S. K. Jakka, O. Gunduz, F. N. Oktar and M. P. F. Graça, J. Funct. Biomater., 2023, 14, 279 CrossRef CAS PubMed.
- J. Pasuri, J. Holopainen, H. Kokkonen, M. Persson, K. Kauppinen, P. Lehenkari, E. Santala, M. Ritala and J. Tuukkanen, Colloids Surf., B, 2015, 135, 774–783 CrossRef CAS PubMed.
- E. Ilhan, Z. Karahaliloglu, E. Kilicay, B. Hazer and E. B. Denkbas, Mater. Technol., 2020, 35, 179–194 CrossRef CAS.
- N. E. Duygulu, F. Ciftci and C. B. Ustundag, J. Polym. Res., 2020, 27, 232 CrossRef CAS.
- E. Pilavci, M. Ayran, D. Ulubay, E. Kaya, G. Tinaz, O. Bingol Ozakpinar, A. Sancakli and O. Gunduz, J. Bioact. Compat. Polym., 2023, 38, 3–24 CrossRef CAS.
- A. A. Aldana, L. Malatto, M. A. U. Rehman, A. R. Boccaccini and G. A. Abraham, Nanomaterials, 2019, 9, 120 CrossRef PubMed.
- S. M. Stella, T. M. Sridhar, R. Ramprasath, J. Gimbun and U. Vijayalakshmi, Polymers, 2022, 15, 155 CrossRef PubMed.
- A. Kotrotsos, P. Yiallouros and V. Kostopoulos, Biomimetics, 2020, 5, 43 Search PubMed.
- F. M. Sánchez-Arévalo, L. D. Muñoz-Ramírez, M. Álvarez-Camacho, F. Rivera-Torres, A. Maciel-Cerda, R. Montiel-Campos and R. Vera-Graziano, J. Mater. Sci., 2017, 52, 3353–3367 Search PubMed.
- D. Santos, C. O. Correia, D. M. Silva, P. S. Gomes, M. H. Fernandes, J. D. Santos and V. Sencadas, Mater. Sci. Eng., C, 2017, 75, 1184–1190 CrossRef CAS PubMed.
- C. C. Odili, O. P. Gbenebor, R. U. Obisike and S. O. Adeosun, Discover Polymers, 2025, 2, 13 CrossRef.
- J. Xu, J. Zhang, W. Gao, H. Liang, H. Wang and J. Li, Mater. Lett., 2009, 63, 658–660 Search PubMed.
- E. Ilhan, S. Cesur, R. B. Sulutas, E. Pilavci, B. Dalbayrak, E. Kaya, E. D. Arisan, G. B. Tinaz, M. Sengor, E. Kijeńska-Gawrońska, F. N. Oktar and O. Gunduz, Eur. Polym. J., 2022, 176, 111390 CrossRef CAS.
- M. Ayran, S. Ulag, S. Ervan and O. Gunduz, Mater. Lett., 2024, 363, 136316 CrossRef CAS.
- C. I. Codrea, D. Baykara, R.-A. Mitran, A. C. Ç. Koyuncu, O. Gunduz and A. Ficai, Polymers, 2024, 16, 1932 CrossRef CAS PubMed.
- E. Garskaite, L. Alinauskas, M. Drienovsky, J. Krajcovic, R. Cicka, M. Palcut, L. Jonusauskas, M. Malinauskas, Z. Stankeviciute and A. Kareiva, RSC Adv., 2016, 6, 72733–72743 RSC.
- C. Liu, Q. Yu, Z. Yuan, Q. Guo, X. Liao, F. Han, T. Feng, G. Liu, R. Zhao, Z. Zhu, H. Mao, C. Zhu and B. Li, Bioact. Mater., 2023, 25, 445–459 CAS.
- M. Kirsch, L. Birnstein, I. Pepelanova, W. Handke, J. Rach, A. Seltsam, T. Scheper and A. Lavrentieva, Bioengineering, 2019, 6, 76 CrossRef CAS PubMed.
- V. Nadarajan, S. W. Phang and H. L. Choo, AIP Conf. Proc., 2020, 2233(1), 040004 CrossRef CAS.
- E. Odabaş, O. Öztürk and E. Akarsu, Polym. Compos., 2024, 45, 17280–17293 Search PubMed.
- M. A. M. Hussein, S. Su, S. Ulag, A. Woźniak, M. Grinholc, G. Erdemir, S. Erdem Kuruca, O. Gunduz, M. Muhammed, I. M. El-Sherbiny and M. Megahed, Polymers, 2021, 13, 3630 CrossRef CAS PubMed.
- S. N. Kasa, M. F. Omar and I. N. Ismail, AIP Conf. Proc., 2017, 1901(1), 030014 CrossRef.
- M. S. Izgordu, E. I. Uzgur, S. Ulag, A. Sahin, B. Karademir Yilmaz, B. Kilic, N. Ekren, F. N. Oktar and O. Gunduz, Cartilage, 2021, 13, 626S–635S CrossRef CAS PubMed.
- P. J. Piszko, A. Piszko, S. Kiryk, J. Kiryk, T. Horodniczy, N. Struzik, K. Wiśniewska, J. Matys and M. Dobrzyński, Biomimetics, 2024, 9, 503 CrossRef CAS PubMed.
- T. Feng, B. Liang, H. Bi and Y. Han, Mater. Des., 2021, 209, 109957 Search PubMed.
- M. J. Bertocchi, R. A. Simbana, J. H. Wynne and J. G. Lundin, Macromol. Mater. Eng., 2019, 304(8), 1900186 CrossRef.
- H. M. Pauly, D. J. Kelly, K. C. Popat, N. A. Trujillo, N. J. Dunne, H. O. McCarthy and T. L. Haut Donahue, J. Mech. Behav. Biomed. Mater., 2016, 61, 258–270 CrossRef CAS PubMed.
- T. U. Rashid, R. E. Gorga and W. E. Krause, Adv. Eng. Mater., 2021, 23(9), 2100153 CrossRef.
- R. P. Alanis-Gómez, E. M. Rivera-Muñoz, G. Luna-Barcenas, J. R. Alanis-Gómez and R. Velázquez-Castillo, Materials, 2022, 15, 4718 CrossRef PubMed.
- S. Hernández, C. A. Brebbia and W. P. De Wilde, WIT Trans. Built Environ., 2016, 166, 372–378 Search PubMed.
- M. Subramaniyan, S. Karuppan, S. Helaili and I. Ahmad, J. Mol. Struct., 2024, 1306, 137862 CrossRef CAS.
- S. M. Stella, T. M. Sridhar, R. Ramprasath, J. Gimbun and U. Vijayalakshmi, Polymers, 2022, 15, 155 CrossRef PubMed.
- Y. Piao, H. You, T. Xu, H.-P. Bei, I. Z. Piwko, Y. Y. Kwan and X. Zhao, Eng. Regener., 2021, 2, 47–56 Search PubMed.
- E. Pilavci, M. Ayran, D. Ulubay, E. Kaya, G. Tinaz, O. Bingol Ozakpinar, A. Sancakli and O. Gunduz, J. Bioact. Compat. Polym., 2023, 38, 3–24 CrossRef CAS.
- I. K. H. Dinatha, A. H. Diputra, H. Wihadmadyatami, J. Partini and Y. Yusuf, RSC Adv., 2024, 14, 8222–8239 RSC.
- M. M. Kareem and K. E. Tanner, J. Mater. Sci.: Mater. Med., 2020, 31, 38 CrossRef CAS PubMed.
- B. Liang, T. Feng, X. Yuan, K. Zhao, C. Li and Y. Han, Mater. Des., 2022, 219, 110834 Search PubMed.
- M. A. Elsawy, K.-H. Kim, J.-W. Park and A. Deep, Renewable Sustainable Energy Rev., 2017, 79, 1346–1352 CrossRef CAS.
- Y. Fang, Z. Liu, Y. Jin, Y. Huang, S. Zhou, H. Tian and H. Wu, Int. J. Biol. Macromol., 2024, 277, 133905 CrossRef CAS PubMed.
- C. B. Danoux, D. Barbieri, H. Yuan, J. D. de Bruijn, C. A. van Blitterswijk and P. Habibovic, Biomatter, 2014, 4, e27664 Search PubMed.
- E. Pérez, J. Mater. Sci., 2021, 56, 19915–19935 CrossRef.
- T. Dou, N. Jing, B. Zhou and P. Zhang, J. Mater. Sci., 2018, 53, 8009–8019 CrossRef CAS.
- A. Patlolla and T. L. Arinzeh, Biotechnol. Bioeng., 2014, 111, 1000–1017 CrossRef CAS PubMed.
- A. S. Badami, M. R. Kreke, M. S. Thompson, J. S. Riffle and A. S. Goldstein, Biomaterials, 2006, 27, 596–606 Search PubMed.
- S. Metwally, S. Ferraris, S. Spriano, Z. J. Krysiak, Ł. Kaniuk, M. M. Marzec, S. K. Kim, P. K. Szewczyk, A. Gruszczyński, M. Wytrwal-Sarna, J. E. Karbowniczek, A. Bernasik, S. Kar-Narayan and U. Stachewicz, Mater. Des., 2020, 194, 108915 CrossRef CAS.
- X. Jing, H.-Y. Mi, X.-C. Wang, X.-F. Peng and L.-S. Turng, ACS Appl. Mater. Interfaces, 2015, 7, 6955–6965 Search PubMed.
- R. L. Price, K. M. Haberstroh and T. J. Webster, Nanotechnology, 2004, 15, 892–900 Search PubMed.
- M. Zayzafoon, J. Cell. Biochem., 2006, 97, 56–70 CrossRef CAS PubMed.
- Z. Huang, S.-L. Cheng and E. Slatopolsky, J. Biol. Chem., 2001, 276, 21351–21358 CrossRef CAS PubMed.
- J. Markhoff, J. Wieding, V. Weissmann, J. Pasold, A. Jonitz-Heincke and R. Bader, Materials, 2015, 8, 5490–5507 Search PubMed.
- N. L. Nerurkar, S. Sen, B. M. Baker, D. M. Elliott and R. L. Mauck, Acta Biomater., 2011, 7, 485–491 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |







