Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Synthesis and antimitotic activity of 2-phenyl-6-pyridinyl-2H-pyrazolo[4,3-c]pyridines
Vaida Aleksienė†
a,
Eva Řezníčková† b,
Aurimas Bieliauskas
b,
Aurimas Bieliauskas a,
Veronika Vojáčková
a,
Veronika Vojáčková b,
Veronika Molitorováb,
Austėja Šalvytė-Nikliauzienėc,
Sergey Belyakovd,
Asta Žukauskaitė
b,
Veronika Molitorováb,
Austėja Šalvytė-Nikliauzienėc,
Sergey Belyakovd,
Asta Žukauskaitė e,
Eglė Arbačiauskienė
e,
Eglė Arbačiauskienė c,
Vladimír Kryštof
c,
Vladimír Kryštof *b and
Algirdas Šačkus*ac
*b and
Algirdas Šačkus*ac
aInstitute of Synthetic Chemistry, Faculty of Chemical Technology, Kaunas University of Technology, K. Baršausko g. 59, LT-51423 Kaunas, Lithuania. E-mail: algirdas.sackus@ktu.lt
bDepartment of Experimental Biology, Faculty of Science, Palacký University, Šlechtitelů 27, CZ-77900 Olomouc, Czech Republic. E-mail: vladimir.krystof@upol.cz
cDepartment of Organic Chemistry, Faculty of Chemical Technology, Kaunas University of Technology, Radvilėnų pl. 19, LT-50254 Kaunas, Lithuania
dLatvian Institute of Organic Synthesis, Aizkraukles 21, LV-1006 Riga, Latvia
eDepartment of Chemical Biology, Faculty of Science, Palacký University, Šlechtitelů 27, CZ-77900 Olomouc, Czech Republic
First published on 12th May 2026
Abstract
An efficient synthetic route to 2-phenyl-6-pyridinyl-2H-pyrazolo[4,3-c]pyridines, alongside comprehensive structural elucidation and biological evaluation, is reported. Among the newly synthesized compounds, 7f, which contains 4-fluorophenyl and pyridin-2-yl substituents at the 2- and 6-positions, respectively, exhibited the strongest submicromolar cytotoxicity across various cancer cell lines. This compound compromised microtubule integrity, induced mitotic defects and aberrant cytokinesis, triggered endoreduplication, and ultimately resulted in cell death. Our findings highlight the potential of these pyrazolo[4,3-c]pyridine derivatives as antimitotic agents, providing a basis for further development of anticancer therapeutics.
Introduction
Pyrazoles are a class of important nitrogen-containing heterocyclic compounds that cover a range of natural products and synthetic derivatives.1–4 Due to excellent pharmacological properties, the pyrazole moiety is frequently used in drug discovery,5 resulting in numerous approved pharmaceuticals and veterinary drugs.3,6 For instance, sildenafil is used to treat erectile dysfunction7 and pulmonary arterial hypertension,8 while celecoxib, tepoxalin, epirizole, and lonazolac are non-steroidal anti-inflammatory drugs.9–11Pyrazole-based derivatives are also widely investigated for their anticancer and antitumor activities.12,13 Among natural products, pyrazofurin (I) (Fig. 1A), a C-nucleoside analog produced by Streptomyces candidus and other actinobacteria, exhibits antitumor properties, though a phase I clinical trial primarily revealed toxicity without therapeutic effect.14–16 Synthetic pyrazole drugs include several clinically approved kinase inhibitors. For instance, tyrosine kinase inhibitors selpercatinib (II), avapritinib (III), and crizotinib (IV) target the RET, KIT/PDGFRA, and ALK/ROS1 pathways, respectively, and are used to treat thyroid, multidrug-resistant gastrointestinal stromal tumors, and non-small-cell lung cancers.17–20 Indazole moiety-containing pazopanib (V) acts as a multi-targeted tyrosine kinase inhibitor and is approved for the treatment of renal cell carcinoma.21 Beyond kinase inhibition, pyrazole drugs also work via other mechanisms. For instance, lonidamine (VI) selectively inhibits aerobic glycolysis and energy metabolism in tumor cells and is capable of sensitizing tumors to chemo-, radio-, and photodynamic-therapy and hyperthermia,22–24 while darolutamide (VII) is a nonsteroidal androgen receptor antagonist used to treat non-metastatic castration-resistant prostate cancer.25 Finally, the poly(ADP-ribose) polymerase (PARP) inhibitor niraparib (VII) represses DNA damage repair in cancer cells and is used for the maintenance treatment of epithelial ovarian, fallopian tube, and primary peritoneal cancer.26
 | ||
| Fig. 1 Pyrazole moiety-containing anticancer agents (A) and the structural framework of our previous study that inspired the present work (B). | ||
Although anticancer pyrazoles exhibit a wide range of structural diversity, recent research increasingly focuses on condensed systems that combine the pharmacological properties of the pyrazole ring with other heterocyclic frameworks.27,28 Among condensed pyrazole derivatives, pyrazolopyridines, which can exist in either the more prevalent 1H- or synthetically more demanding 2H-tautomeric form, are renowned for their anticancer properties.29 1H-Pyrazolo[3,4-b]pyridines, in particular, are reported as potent inhibitors of various kinases30 and antileukemic agents.31 Although less prevalent in the scientific literature, the bioactivities of other pyrazolopyridines are also notable. 1H-Pyrazolo[3,4-c]pyridines were identified as antiproliferative pro-apoptotic agents32,33 and kinase inhibitors.34 1H-Pyrazolo[4,3-b]pyridines were evaluated as selective c-Met35 or dual FLT3/CDK4 inhibitors.36 In the case of 2H-pyrazolo[4,3-c]pyridines, inhibitory activity against p38α, aurora A, CK1δ37 and p90 ribosomal S6 kinases 2 (RSK2)38 was reported, while in our recent works, we also studied antiproliferative,39 photodynamic anticancer40 and antimitotic41 properties of their derivatives. Our previous study investigated the structure–activity relationships of 6-alkyl- and 6-aryl-2-phenyl-2H-pyrazolo[4,3-c]pyridines IX (Fig. 1B). Several of them showed promising anticancer activity in vitro, including cell cycle arrest in mitosis and induction of apoptosis.41 One of the most potent compounds, 2,6-diphenyl-2H-pyrazolo[4,3-c]pyridine IXa, exhibited in vitro cytotoxicity in the K562 cell line with a micromolar GI50 value of 3.4 µM. Building on these results, in the current work, we explore the synthesis and antimitotic activity of new 2H-pyrazolo[4,3-c]pyridines X bearing pyridinyl substituents at the 6-position and variously substituted phenyl groups at the 2-position, as structural modifications to enhance the properties of the compounds.42,43
Results and discussion
Synthesis
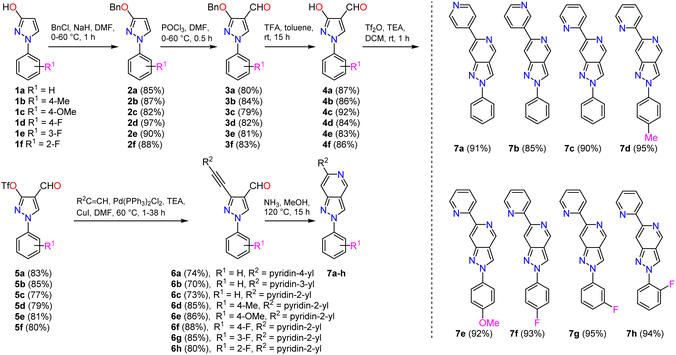
For the synthesis of 2-phenyl-6-pyridinyl-2H-pyrazolo[4,3-c]pyridines 7a–h (Scheme 1), the starting 1-phenyl-1H-pyrazol-3-ols 1a–f were prepared via addition–cyclisation reaction of commercially available variously 2-, 3- and 4-substituted phenyl hydrazine hydrochlorides and ethyl acrylate in the presence of a base and a subsequent oxidation of the obtained pyrazolidin-3-ones by FeCl3 as previously described.44–47 1-Phenyl-1H-pyrazol-3-ols 1a–f were deprotonated by NaH and subjected to O-benzylation by benzyl chloride to provide 3-O-protected pyrazoles 2a–f in very good yields (82–97%). The compounds were further formylated under Vilsmeier–Haack reaction conditions to yield pyrazole-4-carbaldehydes 3a–f in good yields (79–84%). For the O-deprotection, trifluoroacetic acid was employed, and corresponding 3-hydroxy-1H-pyrazole-4-carbaldehydes 4a–f were obtained in very good yields (83–92%). The latter compounds underwent the triflation reaction smoothly with triflic anhydride in the presence of TEA, stirring the reaction mixture in DCM at room temperature. Obtained pseudohalides 5a–f were coupled with pyridin-2-yl, pyridin-3-yl, or pyridin-4-yl acetylenes under the standard sonogashira cross-coupling reaction conditions (Pd(PPh3)2Cl2, CuI, TEA in DMF) to yield appropriate 3-alkynylpyrazoles 6a–h in yields of 70–88%. We have demonstrated previously that 3-alkynyl-1H-pyrazole-4-carbaldehydes can serve as excellent starting materials for the synthesis of various pyrazole-containing fused heterocyclic systems, including but not limited to pyrazolo[4,3-c]pyridines, pyrazolo[4,3-f][1,2,3]triazolo[5,1-c][1,4]oxazepines, pyrazolo[4′,3′:3,4]pyrido[1,2-a]benzimidazoles, and related compounds.48–50 In this work, compounds 6a–h were subjected to dry ammonia-induced cyclisation, and 2-phenyl-6-pyridinyl-2H-pyrazolo[4,3-c]pyridines 7a–h were formed in excellent yields (85–95%).NMR spectroscopic investigations
The structures of all targeted 2-phenyl-6-pyridinyl-2H-pyrazolo[4,3-c]pyridines 7a–h were unambiguously confirmed using various spectroscopic methods, including multinuclear NMR spectroscopy, infrared spectroscopy (IR), and high-resolution mass spectrometry (HRMS). The main information for structure elucidation was obtained through a combination of standard and advanced NMR techniques, including 1H–13C HMBC, 1H–13C HSQC, 1H–13C H2BC, 1H–15N HMBC, 1H–1H COSY, 1H–1H TOCSY, 1H–1H NOESY, 1D selective NOESY, and 1,1-ADEQUATE experiments, which provided an unambiguous assignment of the signals. The NMR data for all studied compounds are provided in the Experimental section and SI; moreover, an in-depth data analysis showed that the chemical shift values were highly consistent across the 2-phenyl-6-pyridinyl-2H-pyrazolo[4,3-c]pyridine series, confirming the shifts for each position. The NMR data for the representative compounds 7a, 7b, and 7f are shown in Fig. 2.Key information about the formation of the pyrazolo[4,3-c]pyridine ring system was easily obtained through various one- and two-dimensional 1H and 15N NMR techniques. This was mainly due to three clearly distinguishable methine protons, which are separated from other aromatic protons and are, most importantly, next to two “pyridine-like” and one “pyrrole-like” nitrogen atoms. In all targeted compounds 7a–h, these methine protons did not show any 1H–1H COSY correlations but showed clear 1H–1H TOCSY cross-peaks in the spectral data, enabling the identification of two distinct spin systems within the pyrazolo[4,3-c]pyridine moiety. Specifically, in the 1H–1H TOCSY spectra, the most downfield methine proton 4-H, resonating from δ 9.25 to 9.34 ppm in compounds 7a–h, shared a TOCSY cross-peak with the methine proton 7-H. Then, 7-H shared a TOCSY cross-peak with the methine proton 3-H. This was supported by the 1H NMR spectra of compounds 7a and 7b, where long-range 5J couplings were clearly seen.
The methine protons 3-H (d, 5JH,H = 0.9 Hz) and 4-H (d, 5JH,H = 1.3 Hz) appeared as doublets, while the proton 7-H (dd, 5JH,H = 1.3, 0.9 Hz) appeared as a doublet of doublets. In compounds 7c–h, a noticeable broadening of the 1H NMR spectral lines was seen throughout the 6-pyridinyl-2H-pyrazolo[4,3-c]pyridine ring system.
Next, the NOESY spectral data provided additional insights into connectivity based on through-space correlations in compounds 7a–h. For instance, a clear NOE was observed between the most downfield 4-H proton and the nearby 3-H proton, followed by an NOE correlation between the 3-H and the nearby phenyl group protons, confirming their close proximity in space. Meanwhile, the methine proton 7-H showed distinct NOEs only in compounds 7a and 7b with the nearby protons from the pyridin-4-yl (δ 7.98–8.00 ppm) and pyridin-3-yl (δ 8.37 and 9.29–9.31 ppm) groups attached at C-6 of the pyrazolo[4,3-c]pyridine ring system. Interestingly, for compounds 7c–h with a pyridin-2-yl group at site 6, NOE correlations were also expected but not observed with the methine proton 7-H.
Furthermore, after successfully identifying the methine protons of the pyrazolo[4,3-c]pyridine ring system, the 15N NMR spectral analysis of compounds 7a–h was straightforward. The 1H–15N HMBC spectra showed that the proton 3-H had long-range correlations with neighboring N-2 “pyrrole-like” (from δ −144.2 to −148.3 ppm, except for compound 7h where it was at δ −157.4 ppm due to the 2-fluorophenyl group) and N-1 “pyridine-like” (from δ −95.3 to −98.9 ppm) nitrogen atoms, while the “pyridine-like” N-5 (from δ −86.6 to −88.5 ppm) nitrogen atom correlated with the nearby H-4 and H-7 protons. The most downfield 15N resonances appeared in the 6-pyridinyl groups, ranging from δ −70.7 to −77.4 ppm.
Finally, assigning 13C resonances for the rest of the pyrazolo[4,3-c]pyridine ring system was straightforward using a combination of 1H–13C HSQC and 1H–13C HMBC techniques. For the representative compounds 7a, 7b, and 7f, a 1,1-ADEQUATE experiment was also performed. Specifically, for compound 7a, the multiplicity-edited 1H–13C HSQC spectrum showed that the distinct methine protons 3-H, 4-H, and 7-H each connect by one bond to carbons C-3 (δ 121.1 ppm), C-4 (δ 146.5 ppm), and C-7 (δ 107.6 ppm), respectively. This finding, along with data from the 1,1-ADEQUATE and 1H–13C HMBC experiments, helped us to unambiguously assign the signals of the quaternary carbons C-3a (δ 119.4 ppm), C-6 (δ 147.3 ppm), and C-7a (δ 150.2 ppm). In all the compounds analyzed, a quaternary carbon C-7a was a bit more downfield than C-6, while C-3a was the most upfield. For compounds 7c–h containing a pyridin-2-yl group at site 6, a significant broadening of the 13C NMR spectral lines was also observed, similar to what was seen in the 1H NMR spectra. However, an in-depth 2D NMR analysis allowed us to identify the derivatives obtained. In the case of a representative compound 7f, the long-range 1H–13C HMBC spectral data clearly confirmed the connectivity between the quaternary carbon C-6 (δ 150.1 ppm) in the pyrazolo[4,3-c]pyridine ring system and the methine proton 3-H″ (δ 8.43 ppm) from the neighboring pyridin-2-yl group. Furthermore, this was supported by the 1,1-ADEQUATE and 1H–13C HSQC experiments, which clearly showed that the methine carbon C-3″ (δ 121.4 ppm) was adjacent to the quaternary carbon C-2″ (δ 156.6 ppm), consistent with data reported for compounds containing the pyridin-2-yl moiety.51
Derivatives 7c–h containing a pyridin-2-yl group at site 6 can be described as “2,2′-bipyridine”-like compounds, existing in an equilibrium between s-trans and s-cis conformers in solution.52,53 Many “2,2′-bipyridine”-like compounds related to metal complexes favor the s-cis conformer, but it is well known that the main form of unchelated “2,2′-bipyridine”-like compounds is the s-trans conformer.51 Some biheterocycles, including 2,2′-bipyridine derivatives, can form weak intramolecular hydrogen bonds between their heterocyclic fragments, which leads to a characteristic downfield shift in the 1H NMR spectrum.54,55 However, intramolecular hydrogen bonds are very weak; thus, they are insufficient to stabilize the corresponding conformations. Comparing the 1H NMR spectral data for all pyridin-2-yl derivatives 7c–h with those of pyridin-4-yl 7a and pyridin-3-yl 7b derivatives, substituted at site 6, clearly shows more key differences. In addition to the significant broadening of NMR lines in the 7c–h series, the distinctive methine proton 7-H (δ 8.64–8.75 ppm) is noticeably downfield by 0.5–0.7 ppm compared to derivatives 7a and 7b, suggesting possible intramolecular hydrogen bonding. Overall, the absence of NOE correlations, along with other key differences in the 1H NMR spectral data between the pyridin-2-yl derivatives 7c–h and derivatives 7a and 7b, is mainly attributed to the equilibrium between s-trans and s-cis conformers, with the s-trans form being predominant, as reported in the literature.
Single-crystal X-ray diffraction analysis
To objectively establish the structure of the prepared derivatives, a single-crystal X-ray diffraction analysis of the most active compound 7f was carried out. Fig. 3A illustrates a perspective view of the molecule with thermal ellipsoids and the atom-numbering scheme followed in the text. In the molecular structure, the 4-fluorophenyl substituent forms a dihedral angle of 36.5(2)° with the plane of the pyrazolo[4,3-c]pyridine system. The 2-pyridyl substituent forms an even smaller dihedral angle of 16.0(2)° with this system. Low dihedral angle values promote conjugation throughout the molecule. | ||
| Fig. 3 ORTEP diagram for compound 7f (A) and a fragment of its molecular packing in the crystal structure showing CH⋯N hydrogen bonds (B). | ||
The C4 carbon atom has increased electronegativity; this makes the C4–H group capable of forming CH⋯ N-type hydrogen bonds. In the crystal structure, a moderate intermolecular C4–H⋯N1 hydrogen bond with a length of 3.322(2) Å (H⋯N = 2.75(2) Å, C–H⋯O = 117(3)°) was detected. These interactions lead to the formation of molecular chains along the crystallographic direction [010] in the crystal structure. Fig. 3B shows a molecular chain formed by CH⋯N hydrogen bonds. The crystal structure is chiral (space group is P212121) despite the absence of asymmetric atoms.
Biology
Prepared derivatives 7a–h were evaluated for their cytotoxicity against three human cancer cell lines: K562 (chronic myeloid leukemia), MV4-11 (acute myeloid leukemia), and CEM (acute lymphoblastic leukemia). The compound IXa from the previous series, along with two standards (taxol and vinflunine), were included for comparison purposes. The majority of the compounds exhibited good cytotoxicity, with GI50 values in the low micromolar or submicromolar range (Table 1).| Cmpd. | GI50 (µM) | ||
|---|---|---|---|
| K562 | MV4-11 | CEM | |
| 7a | 23.07 ± 2.73 | >25 | >25 |
| 7b | 8.93 ± 3.52 | 17.94 ± 2.46 | 18.57 ± 1.03 |
| 7c | 0.61 ± 0.23 | 0.55 ± 0.32 | 0.39 ± 0.08 |
| 7d | 1.76 ± 0.38 | 2.18 ± 0.55 | 6.28 ± 2.47 |
| 7e | 19.41 ± 4.36 | 20.47 ± 5.24 | 22.77 ± 3.16 |
| 7f | 0.03 ± 0.02 | 0.05 ± 0.00 | 0.06 ± 0.01 |
| 7g | 0.04 ± 0.00 | 0.15 ± 0.03 | 0.18 ± 0.07 |
| 7h | 0.65 ± 0.20 | 0.44 ± 0.08 | 0.38 ± 0.06 |
| IXa | 3.83 ± 0.86 | 3.86 ± 0.96 | 7.24 ± 3.28 |
| Taxol | 0.02 ± 0.00 | 0.01 ± 0.00 | 0.02 ± 0.01 |
| Vinflunine | 0.22 ± 0.02 | 0.10 ± 0.01 | 0.23 ± 0.12 |
The cytotoxicity of the compounds was strongly influenced by the position of the nitrogen atom in the pyridine substituent. Specifically, among unsubstituted 2-phenyl derivatives 7a–c, the pyridin-2-yl derivative 7c exhibited significantly higher cytotoxicity, whereas pyridin-3-yl and pyridin-4-yl derivatives 7b and 7a only possessed moderate and low cytotoxicity, respectively. The enhanced activity of pyridin-2-yl derivatives (e.g., 7c, 7f, 7g) compared to their pyridin-3-yl (7b) and pyridin-4-yl (7a) analogues may be attributed to the ability of the 2-pyridyl moiety to adopt conformations enabling more favourable intramolecular interactions and potentially improved binding to the biological target.
Compound 7c was therefore chosen for further optimization of its biological activity. Introduction of substituents on the phenyl ring at the 2-position revealed a clear substituent effect. Addition of electron-donating groups such as methyl and methoxy (7d, 7e) diminished the activity relative to the parent compound 7c, whereas incorporation of fluorine resulted in enhanced cytotoxicity. Specifically, introduction of fluorine at the ortho-position on the 2-phenyl ring did not affect the cytotoxicity of the compound, while substitution with both meta- and especially para-fluorine resulted in the most active compounds of the series, 7g and 7f, respectively. The increased activity of fluorinated derivatives can be explained by a combination of electronic and physicochemical effects. In particular, the electron-withdrawing nature of fluorine, together with increased lipophilicity, may enhance cell membrane permeability and improve interactions with the cellular target, for example, by modulating π–π stacking or hydrogen bonding interactions.
Subsequently, compound 7f, which displayed the strongest antiproliferative activity in the panel of selected cancer cell lines, was selected for further evaluation of its biological effects in vitro. The potential of 7f was further confirmed by evaluating its effects on two non-cancer cell lines, MRC-5 and BJ fibroblasts. Viability of confluent cultures of these cells, mimicking the state of nonproliferating healthy cells, was not significantly affected by 7f up to the concentration of 10 µM. Initial flow cytometry analysis of K562 leukemic cells treated with increasing concentrations of 7f showed a marked enrichment of G2/M cell population after 24 h treatment (Fig. 4A). Moreover, parallel microscopic observations of 7f-treated cells (Fig. 4B) revealed significant morphological alterations characterized by a substantial number of strikingly elongated cells together with a concentration- and time-dependent increase in the population of cells showing signs of cell death.
 | ||
| Fig. 4 Cell cycle analysis (A) and microscopic observations (B) of K562 cells treated with compound 7f for 24 and 48 h. | ||
To distinguish between the G2 and M phase of the cell cycle, several markers of mitotic progression were further evaluated at the protein level by immunoblotting (Fig. 5). A crucial step of mitotic entry56 is the activation of several mitotic kinases, including cyclin-dependent kinase 1 (CDK1) and polo-like kinase 1 (Plk-1). Activation of CDK1 requires phosphorylation of T161 residue by CAK complex and dephosphorylation of T14/Y15 by Cdc25 phosphatases. Activating phosphorylation of Plk-1 at T210 is necessary for the proper function of this kinase.
 | ||
| Fig. 5 Immunoblotting analysis of mitotic and apoptotic markers in K562 cells treated with compound 7f. α-tubulin levels were detected to verify equal protein loading. | ||
Changes in the phosphorylation of these residues in CDK1 as well as Plk-1 were detected after 24 h treatment with 7f in a concentration-dependent manner (Fig. 5). Analysis of Bcl-2 protein via immunoblotting showed a slowly migrating band attributed to phosphorylated Bcl-2. This result was further confirmed using a specific antibody against S70-phosphorylated Bcl-2, clearly showing a dose-dependent increase of this form of Bcl-2, which belongs to another well-known marker of ongoing mitosis,57 as well as histone H3 phosphorylated at S10, whose levels were also rising with increasing concentrations of 7f.
The induction of mitotic arrest, accompanied by morphological changes, may indicate a disruption of cytoskeletal integrity. Although pyrazole derivatives are not commonly associated with tubulin inhibition, a few have been reported to act as tubulin-targeting anticancer agents.58–61 Therefore, immunofluorescence labelling of α-tubulin after 7f treatment was performed in K562 leukemic cells as well as in MCF-7 breast cancer cells (Fig. 6). The initial experiment demonstrated that 7f at micromolar concentrations disrupts the structure of microtubules and acts as a destabilising agent.
To further evaluate the concentration-dependent effect of 7f on the microtubule cytoskeleton, K562 cells were seeded on poly-L-lysine-coated cover slides to improve their attachment to the surface and subsequently treated with 7f. Immunofluorescence labelling of α-tubulin (Fig. 7) showed that nanomolar concentrations of 7f induced multipolar spindle formation, a common consequence of disruption of microtubule dynamics leading to aberrant mitosis. At higher concentrations, microtubules were disrupted, and the usual round cell morphology of K562 cells was altered to a significantly elongated form with nuclei located at one pole of the cell.
 | ||
| Fig. 7 Immunofluorescence staining of K562 cells upon 24 h treatment with indicated concentrations of 7f. α-tubulin was visualized by Alexa Fluor™ 488-conjugated antibody, and cell nuclei by DAPI. | ||
In addition to changes in cell cycle phase distribution, flow cytometry analysis revealed that 7f treatment increased the number of sub-G1 cells undergoing cell death (Fig. 4A). This was further confirmed via detection of massive PARP-1 cleavage, a common marker of ongoing apoptosis, and also increased phosphorylation of histone H2AX at S139 (γH2AX) in the treated samples (Fig. 5). The presence of γH2AX highlights a cellular response to DNA damage that in this case can be a consequence of genomic instability caused by alterations in microtubule dynamics and mitotic defects. Cell death probably also affected expression of several important proteins including common oncogene c-MYC whose levels dramatically decreased after treatment with nanomolar concentrations of 7f (Fig. 5). Moreover, already after 24 h treatment with 7f we observed an increasing population of cells characterized by >4N DNA amount in comparison to the untreated control (Fig. 4A). As the mitotic defects can prevent the proper chromosome segregation and lead to endoreduplication, the duration of treatment was prolonged to 48 h and subsequent analysis revealed that longer exposure of K562 cells to 7f expanded the population having 8N DNA content (Fig. 4A) proving the hypothesis. Analysis of BrdU-pulse labelled K562 cells treated with 0.313 µM 7f for 24 and 48 h also confirmed that DNA replication is maintained (BrdU-FITC-positive cells) in a significant fraction of the cells leading, in the absence of proper cell division, to polyploidy (SI, Fig. S1).
Experimental
Chemistry
Diffraction data for compound 7f were collected at low temperature (150 K) on a Rigaku, XtaLAB Synergy, Dualflex, HyPix diffractometer using monochromated Cu-Kα radiation (λ = 1.54184 Å). The crystal structure was solved with the SIR2011 structure solution program63 using direct methods and refined with the ShelXL refinement package64 using least squares minimisation. All nonhydrogen atoms were refined in anisotropic approximation. The hydrogen atom involved in the formation of H-bond were refined isotropically; all other H-atoms were refined by riding model with Uiso(H) = 1.2Ueq(C). Crystal data: orthorhombic, a = 5.90879(4), b = 10.20119(8), c = 21.8526(2) Å; V = 1317.20(2) Å3, Z = 4, µ = 0.828 mm−1, Dcalc = 1.464 g cm−3; space group is P212121. The final R1 was 0.0284 (I > 2σ(I)) and wR2 was 0.0767 (all data). For further details, see crystallographic data for compound 7f deposited at the Cambridge Crystallographic Data Centre. Deposition Number (https://www.ccdc.cam.ac.uk/services/structures) CCDC 2524805.
General procedure for the synthesis of compounds 2a–f
A solution of appropriate 1-phenyl-1H-pyrazol-3-ol (1a–f)44–47 (7.2 mmol) in dry DMF (20 mL) was cooled to 0 °C under an inert atmosphere, and NaH (60% dispersion in mineral oil, 288 mg, 7.2 mmol) was added portion-wise. After stirring for 15 min, benzyl chloride (0.82 mL, 7.2 mmol) was added dropwise. The reaction mixture was stirred at 60 °C for 1 h, then poured into water and extracted with ethyl acetate. The organic layers were combined, washed with brine, dried over Na2SO4, filtered, and the solvent was evaporated. The residue was purified by column chromatography (SiO2, eluent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate/n-hexane, 1
ethyl acetate/n-hexane, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 7, v/v) to give pure compounds 2a–f.
7, v/v) to give pure compounds 2a–f.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 1266, 1234, 1051, 1022, 828, 806, 733, 698 (CH
C, C–N), 1266, 1234, 1051, 1022, 828, 806, 733, 698 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 2.35 (s, 3H, CH3), 5.30 (s, 2H, OCH2), 5.85–5.88 (m, 1H, 4-H), 7.16–7.22 (m, 2H, NPh 3,5-H), 7.29–7.34 (m, 1H, CPh 4-H), 7.34–7.40 (m, 2H, CPh 3,5-H), 7.44–7.52 (m, 4H, NPh 2,6-H, CPh 2,6-H), 7.65–7.68 (m, 1H, 5-H). 13C NMR (176 MHz, CDCl3): δ 20.9 (OCH3), 70.9 (OCH2), 93.6 (C-4); 117.9 (NPh C-2,6), 127.7 (C-5), 128.10 (CPh C-4), 128.15 (CPh C-2,6), 128.5 (CPh C-3,5), 129.9 (NPh C-3,5), 135.1 (NPh C-4), 137.1 (CPh C-1), 138.1 (NPh C-1), 164.2 (C-3). 15N NMR (41 MHz, CDCl3): δ −185.7 (N-1), N-2 was not found. MS m/z (%): 265 ([M + H]+, 100). HRMS (ESI) for C17H16N2ONa ([M + Na]+): calcd 287.1154, found 287.1155.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 2.35 (s, 3H, CH3), 5.30 (s, 2H, OCH2), 5.85–5.88 (m, 1H, 4-H), 7.16–7.22 (m, 2H, NPh 3,5-H), 7.29–7.34 (m, 1H, CPh 4-H), 7.34–7.40 (m, 2H, CPh 3,5-H), 7.44–7.52 (m, 4H, NPh 2,6-H, CPh 2,6-H), 7.65–7.68 (m, 1H, 5-H). 13C NMR (176 MHz, CDCl3): δ 20.9 (OCH3), 70.9 (OCH2), 93.6 (C-4); 117.9 (NPh C-2,6), 127.7 (C-5), 128.10 (CPh C-4), 128.15 (CPh C-2,6), 128.5 (CPh C-3,5), 129.9 (NPh C-3,5), 135.1 (NPh C-4), 137.1 (CPh C-1), 138.1 (NPh C-1), 164.2 (C-3). 15N NMR (41 MHz, CDCl3): δ −185.7 (N-1), N-2 was not found. MS m/z (%): 265 ([M + H]+, 100). HRMS (ESI) for C17H16N2ONa ([M + Na]+): calcd 287.1154, found 287.1155.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 913, 834, 826, 746, 735, 670, 549 (CH
C, C–N), 913, 834, 826, 746, 735, 670, 549 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 3.83 (s, 3H, OCH3), 5.30 (s, 2H, OCH2), 5.87–5.91 (m, 1H, 4-H), 6.95–6.97 (m, 2H, NPh 3,5-H), 7.32–7.35 (m, 1H, CPh 4-H), 7.36–7.42 (m, 2H, CPh 3,5-H), 7.46–7.50 (m, 2H, CPh 2,6-H), 7.50–7.54 (m, 2H, NPh 2,6-H), 7.61–7.64 (m, 1H, 5-H). 13C NMR (176 MHz, CDCl3): δ 55.7 (OCH3), 71.0 (OCH2), 93.3 (C-4), 114.6 (NPh C-3,5), 119.8 (NPh C-2,6), 127.8 (C-5), 128.1 (CPh C-4), 128.2 (CPh 2,6), 128.6 (CPh C-3,5), 134.2 (NPh C-1), 137.2 (CPh C-1), 157.6 (NPh C-4), 164.2 (C-3). 15N NMR (71 MHz, CDCl3): δ −186.4 (N-1), N-2 was not found. MS m/z (%): 281 ([M + H]+, 100). HRMS (ESI) for C17H16N2O2Na ([M + Na]+): calcd 303.1104, found 303.1104.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 3.83 (s, 3H, OCH3), 5.30 (s, 2H, OCH2), 5.87–5.91 (m, 1H, 4-H), 6.95–6.97 (m, 2H, NPh 3,5-H), 7.32–7.35 (m, 1H, CPh 4-H), 7.36–7.42 (m, 2H, CPh 3,5-H), 7.46–7.50 (m, 2H, CPh 2,6-H), 7.50–7.54 (m, 2H, NPh 2,6-H), 7.61–7.64 (m, 1H, 5-H). 13C NMR (176 MHz, CDCl3): δ 55.7 (OCH3), 71.0 (OCH2), 93.3 (C-4), 114.6 (NPh C-3,5), 119.8 (NPh C-2,6), 127.8 (C-5), 128.1 (CPh C-4), 128.2 (CPh 2,6), 128.6 (CPh C-3,5), 134.2 (NPh C-1), 137.2 (CPh C-1), 157.6 (NPh C-4), 164.2 (C-3). 15N NMR (71 MHz, CDCl3): δ −186.4 (N-1), N-2 was not found. MS m/z (%): 281 ([M + H]+, 100). HRMS (ESI) for C17H16N2O2Na ([M + Na]+): calcd 303.1104, found 303.1104.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 1029, 836, 756, 745, 733, 695, 612 (CH
C, C–N), 1029, 836, 756, 745, 733, 695, 612 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (400 MHz, CDCl3): δ 5.30 (s, 2H, CH2), 5.89–5.93 (m, 1H, 4-H), 7.06–7.14 (m, 2H, NPh 3,5-H), 7.29–7.35 (m, 1H, CPh 4-H), 7.35–7.42 (m, 2H, CPh 3,5-H), 7.46–7.51 (m, 2H, CPh 2,6-H), 7.53–7.59 (m, 2H, NPh 2,6-H), 7.63–7.67 (m, 1H, 5-H). 13C NMR (101 MHz, CDCl3): δ 71.0 (CH2), 94.2 (C-4), 116.2 (d, 2JC,F = 22.9 Hz, NPh C-3,5), 119.6 (d, 3JC,F = 8.2 Hz, NPh C-2,6), 127.9 (C-5), 128.14 (CPh C-2,6), 128.18 (CPh C-4), 128.6 (CPh C-3,5), 136.7 (d, 4JC,F = 2.7 Hz, NPh C-1), 137.0 (CPh C-1), 160.5 (d, 1JC,F = 244.5 Hz, NPh C-4), 164.4 (C-3). 15N NMR (41 MHz, CDCl3): δ −187.7 (N-1), N-2 was not found. MS m/z (%): 269 ([M + H]+, 100). HRMS (ESI) for C16H13FN2ONa ([M + Na]+): calcd 291.0904, found 291.0904.
CH of benzenes). 1H NMR (400 MHz, CDCl3): δ 5.30 (s, 2H, CH2), 5.89–5.93 (m, 1H, 4-H), 7.06–7.14 (m, 2H, NPh 3,5-H), 7.29–7.35 (m, 1H, CPh 4-H), 7.35–7.42 (m, 2H, CPh 3,5-H), 7.46–7.51 (m, 2H, CPh 2,6-H), 7.53–7.59 (m, 2H, NPh 2,6-H), 7.63–7.67 (m, 1H, 5-H). 13C NMR (101 MHz, CDCl3): δ 71.0 (CH2), 94.2 (C-4), 116.2 (d, 2JC,F = 22.9 Hz, NPh C-3,5), 119.6 (d, 3JC,F = 8.2 Hz, NPh C-2,6), 127.9 (C-5), 128.14 (CPh C-2,6), 128.18 (CPh C-4), 128.6 (CPh C-3,5), 136.7 (d, 4JC,F = 2.7 Hz, NPh C-1), 137.0 (CPh C-1), 160.5 (d, 1JC,F = 244.5 Hz, NPh C-4), 164.4 (C-3). 15N NMR (41 MHz, CDCl3): δ −187.7 (N-1), N-2 was not found. MS m/z (%): 269 ([M + H]+, 100). HRMS (ESI) for C16H13FN2ONa ([M + Na]+): calcd 291.0904, found 291.0904.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–O–C, C–F), 981, 951, 844, 761, 751, 735, 696, 522 (CH
C, C–N, C–O–C, C–F), 981, 951, 844, 761, 751, 735, 696, 522 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.33 (s, 2H, CH2), 5.95 (d, 3J = 2.6 Hz, 1H, 4-H), 6.88–6.92 (m, 1H, NPh 4-H), 7.33–7.36 (m, 3H, CPh 3,4,5-H), 7.39–7.41 (m, 3H, NPh 2,5,6-H), 7.49–7.51 (m, 2H, CPh 2,6-H), 7.73 (d, 3J = 2.6 Hz, 1H, 5-H). 13C NMR (176 MHz, CDCl3): δ 71.0 (CH2), 95.0 (C-4), 105.6 (d, 2JC,F = 26.4 Hz, NPh C-2), 112.0 (d, 2JC,F = 21.4 Hz, NPh C-4), 112.8 (d, 4JC,F = 2.8 Hz, NPh C-6), 128.0 (CPh C-4), 128.21 (CPh C-2,6), 128.22 (5JC,F = 2.6 Hz, C-5), 128.6 (CPh C-3,5), 130.7 (d, 3JC,F = 9.2 Hz, NPh C-5), 137.0 (CPh C-1), 141.7 (d, 3JC,F = 7.9 Hz, NPh C-1), 163.5 (d, 1JC,F = 245.8 Hz, NPh C-3), 164.5 (C-3). 19F NMR (376 MHz, CDCl3): δ −111.2. MS m/z (%): 269 ([M + H]+, 100). HRMS (ESI) for C16H13FN2ONa ([M + Na]+): calcd 291.0905, found 291.0904.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.33 (s, 2H, CH2), 5.95 (d, 3J = 2.6 Hz, 1H, 4-H), 6.88–6.92 (m, 1H, NPh 4-H), 7.33–7.36 (m, 3H, CPh 3,4,5-H), 7.39–7.41 (m, 3H, NPh 2,5,6-H), 7.49–7.51 (m, 2H, CPh 2,6-H), 7.73 (d, 3J = 2.6 Hz, 1H, 5-H). 13C NMR (176 MHz, CDCl3): δ 71.0 (CH2), 95.0 (C-4), 105.6 (d, 2JC,F = 26.4 Hz, NPh C-2), 112.0 (d, 2JC,F = 21.4 Hz, NPh C-4), 112.8 (d, 4JC,F = 2.8 Hz, NPh C-6), 128.0 (CPh C-4), 128.21 (CPh C-2,6), 128.22 (5JC,F = 2.6 Hz, C-5), 128.6 (CPh C-3,5), 130.7 (d, 3JC,F = 9.2 Hz, NPh C-5), 137.0 (CPh C-1), 141.7 (d, 3JC,F = 7.9 Hz, NPh C-1), 163.5 (d, 1JC,F = 245.8 Hz, NPh C-3), 164.5 (C-3). 19F NMR (376 MHz, CDCl3): δ −111.2. MS m/z (%): 269 ([M + H]+, 100). HRMS (ESI) for C16H13FN2ONa ([M + Na]+): calcd 291.0905, found 291.0904.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–O–C, C–F), 931, 771, 525 (CH
C, C–N, C–O–C, C–F), 931, 771, 525 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.31 (s, 2H, CH2), 5.94 (d, J = 2.6 Hz, 1H, 4-H), 7.14–7.22 (m, 3H, NPh 3,4,5-H), 7.31–7.34 (m, 1H, CPh 4-H), 7.37–7.40 (m, 2H, CPh 3,5-H), 7.47–7.50 (m, 2H, CPh 2,6-H), 7.84 (d, 3J = 2.6 Hz, 1H, 5-H), 7.87–7.90 (m, 1H, NPh 6-H). 13C NMR (176 MHz, CDCl3): δ 71.0 (CH2), 94.4 (d, 5JC,F = 2.4 Hz, C-4), 116.8 (d, 2JC,F = 20.4 Hz, NPh C-3), 123.3 (CPh C-4), 125.0 (d, 4JC,F = 3.6 Hz, NPh C-5), 126.3 (d, 3JC,F = 7.8 Hz, NPh C-6), 128.15 (CPh C-2,6), 128.16 (d, 4JC,F = 1.7 Hz, C-5), 128.5 (d, 2JC,F = 8.9 Hz, NPh C-1), 128.6 (CPh C-3,5), 132.6 (d, 3J = 12.4 Hz, NPh C-4), 137.0 (CPh C-1), 153.2 (d, 1JC,F = 247.6 Hz, NPh C-2), 164.2 (C-3). MS m/z (%): 269 ([M + H]+, 100). HRMS (ESI) for C16H13N2OFNa ([M + Na]+): calcd 291.0904, found 291.0909.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.31 (s, 2H, CH2), 5.94 (d, J = 2.6 Hz, 1H, 4-H), 7.14–7.22 (m, 3H, NPh 3,4,5-H), 7.31–7.34 (m, 1H, CPh 4-H), 7.37–7.40 (m, 2H, CPh 3,5-H), 7.47–7.50 (m, 2H, CPh 2,6-H), 7.84 (d, 3J = 2.6 Hz, 1H, 5-H), 7.87–7.90 (m, 1H, NPh 6-H). 13C NMR (176 MHz, CDCl3): δ 71.0 (CH2), 94.4 (d, 5JC,F = 2.4 Hz, C-4), 116.8 (d, 2JC,F = 20.4 Hz, NPh C-3), 123.3 (CPh C-4), 125.0 (d, 4JC,F = 3.6 Hz, NPh C-5), 126.3 (d, 3JC,F = 7.8 Hz, NPh C-6), 128.15 (CPh C-2,6), 128.16 (d, 4JC,F = 1.7 Hz, C-5), 128.5 (d, 2JC,F = 8.9 Hz, NPh C-1), 128.6 (CPh C-3,5), 132.6 (d, 3J = 12.4 Hz, NPh C-4), 137.0 (CPh C-1), 153.2 (d, 1JC,F = 247.6 Hz, NPh C-2), 164.2 (C-3). MS m/z (%): 269 ([M + H]+, 100). HRMS (ESI) for C16H13N2OFNa ([M + Na]+): calcd 291.0904, found 291.0909.General procedure for the synthesis of compounds 3a–f
A solution of phosphorus oxychloride (0.37 mL, 4 mmol) in DMF (0.30 mL, 4 mmol) was cooled to 0 °C for 15 min, then appropriate pyrazole 2a–f (1 mmol) was added. The reaction mixture was stirred at 60 °C for 1 hour, then diluted with a 10% Na2CO3 solution and extracted with ethyl acetate (three times, 10 mL each). The organic layers were combined, washed with brine, dried over Na2SO4, filtered, and the solvent was evaporated. The residue was purified by flash column chromatography (SiO2, eluent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate/n-hexane, 1
ethyl acetate/n-hexane, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, v/v) to give pure compounds 3a–f.
2, v/v) to give pure compounds 3a–f.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1558, 1506, 1364, 1225, 1204 (C–O–C, C
O), 1558, 1506, 1364, 1225, 1204 (C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 1010, 817, 735 (CH
C, C–N), 1010, 817, 735 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (400 MHz, CDCl3): δ 2.39 (s, 3H, CH3), 5.44 (s, 2H, CH2), 7.23–7.29 (m, 2H, NPh 3,5-H), 7.32–7.38 (m, 1H, CPh 4-H), 7.38–7.44 (m, 2H, CPh 3,5-H), 7.49–7.55 (m, 4H, NPh 2,6-H, CPh 2,6-H), 8.22 (s, 1H, 5-H), 9.87 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 21.1 (CH3), 71.2 (CH2), 111.3 (C-4), 118.9 (NPh C-2,6), 128.3 (CPh C-2,6), 128.4 (CPh C-4), 128.6 (CPh C-3,5), 129.2 (C-5), 130.2 (NPh C-3,5), 136.3 (CPh C-1), 136.9 (NPh C-1), 137.4 (NPh C-4), 163.7 (C-3), 183.4 (CHO). 15N NMR (41 MHz, CDCl3): δ −178.8 (N-1), N-2 was not found. MS m/z (%): 293 ([M + H]+, 100). HRMS (ESI) for C18H16N2O2Na ([M + Na]+): calcd 315.1104, found 315.1104.
CH of benzenes). 1H NMR (400 MHz, CDCl3): δ 2.39 (s, 3H, CH3), 5.44 (s, 2H, CH2), 7.23–7.29 (m, 2H, NPh 3,5-H), 7.32–7.38 (m, 1H, CPh 4-H), 7.38–7.44 (m, 2H, CPh 3,5-H), 7.49–7.55 (m, 4H, NPh 2,6-H, CPh 2,6-H), 8.22 (s, 1H, 5-H), 9.87 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 21.1 (CH3), 71.2 (CH2), 111.3 (C-4), 118.9 (NPh C-2,6), 128.3 (CPh C-2,6), 128.4 (CPh C-4), 128.6 (CPh C-3,5), 129.2 (C-5), 130.2 (NPh C-3,5), 136.3 (CPh C-1), 136.9 (NPh C-1), 137.4 (NPh C-4), 163.7 (C-3), 183.4 (CHO). 15N NMR (41 MHz, CDCl3): δ −178.8 (N-1), N-2 was not found. MS m/z (%): 293 ([M + H]+, 100). HRMS (ESI) for C18H16N2O2Na ([M + Na]+): calcd 315.1104, found 315.1104.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1557, 1520, 1505, 1497, 1359, 1249, 1212, 1202 (C–O–C, C
O), 1557, 1520, 1505, 1497, 1359, 1249, 1212, 1202 (C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N) 829, 627 (CH
C, C–N) 829, 627 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 3.85 (s, 3H, CH3), 5.43 (s, 2H, CH2), 6.95–7.00 (m, 2H, NPh 3,5-H), 7.33–7.37 (m, 1H, CPh 4-H), 7.39–7.43 (m, 2H, CPh 3,5-H), 7.50–7.53 (m, 2H, CPh 2,6-H), 7.54–7.57 (m, 2H, NPh 2,6-H), 8.16 (s, 1H, 5-H), 9.86 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 55.7 (CH3), 71.2 (CH2), 111.2 (C-4), 114.8 (NPh C-3,5), 120.6 (NPh C-2,6), 128.3 (CPh C-2,6), 128.4 (CPh C-4), 128.6 (CPh C-3,5), 129.2 (C-5), 132.8 (NPh C-1), 136.3 (CPh C-1), 158.9 (NPh C-4), 163.7 (C-3), 183.4 (CHO). 15N NMR (71 MHz, CDCl3): δ −179.1 (N-1), N-2 was not found. MS m/z (%): 309 ([M + H]+, 100). HRMS (ESI) for C18H16N2O3Na ([M + Na]+): calcd 331.1053, found 331.1053.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 3.85 (s, 3H, CH3), 5.43 (s, 2H, CH2), 6.95–7.00 (m, 2H, NPh 3,5-H), 7.33–7.37 (m, 1H, CPh 4-H), 7.39–7.43 (m, 2H, CPh 3,5-H), 7.50–7.53 (m, 2H, CPh 2,6-H), 7.54–7.57 (m, 2H, NPh 2,6-H), 8.16 (s, 1H, 5-H), 9.86 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 55.7 (CH3), 71.2 (CH2), 111.2 (C-4), 114.8 (NPh C-3,5), 120.6 (NPh C-2,6), 128.3 (CPh C-2,6), 128.4 (CPh C-4), 128.6 (CPh C-3,5), 129.2 (C-5), 132.8 (NPh C-1), 136.3 (CPh C-1), 158.9 (NPh C-4), 163.7 (C-3), 183.4 (CHO). 15N NMR (71 MHz, CDCl3): δ −179.1 (N-1), N-2 was not found. MS m/z (%): 309 ([M + H]+, 100). HRMS (ESI) for C18H16N2O3Na ([M + Na]+): calcd 331.1053, found 331.1053.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1559, 1519, 1505, 1495, 1454, 1357, 1297, 1226, 1218, 1205, (C–F, C–O–C, C
O), 1559, 1519, 1505, 1495, 1454, 1357, 1297, 1226, 1218, 1205, (C–F, C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 836, 830, 208, 696 (CH
C, C–N), 836, 830, 208, 696 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.44 (s, 2H, CH2), 7.14–7.20 (m, 2H, NPh 3,5-H), 7.34–7.38 (m, 1H, CPh 4-H), 7.39–7.43 (m, 2H, CPh 3,5-H), 7.49–7.54 (m, 2H, CPh 2,6-H), 7.60–7.65 (m, 2H, NPh 2,6-H), 8.20 (s, 1H, 5-H), 9.88 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 71.3 (CH2), 111.7 (C-4), 116.6 (d, 2JC,F = 23.2 Hz, NPh C-3,5), 120.8 (d, 3JC,F = 8.4 Hz, NPh C-2,6), 128.3 (CPh C-2,6), 128.5 (CPh C-4), 128.7 (CPh C-3,5), 129.5 (C-5), 135.5 (d, 4JC,F = 3.0 Hz, NPh C-1), 136.2 (CPh C-1), 161.6 (d, 1JC,F = 247.7 Hz, NPh C-4), 163.8 (C-3), 183.4 (CHO). 15N NMR (71 MHz, CDCl3): δ −181.0 (N-1), N-2 was not found. MS m/z (%): 297 ([M + H]+, 100). HRMS (ESI) for C17H13FN2O2Na ([M + Na]+): calcd 319.0853, found 319.0853.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.44 (s, 2H, CH2), 7.14–7.20 (m, 2H, NPh 3,5-H), 7.34–7.38 (m, 1H, CPh 4-H), 7.39–7.43 (m, 2H, CPh 3,5-H), 7.49–7.54 (m, 2H, CPh 2,6-H), 7.60–7.65 (m, 2H, NPh 2,6-H), 8.20 (s, 1H, 5-H), 9.88 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 71.3 (CH2), 111.7 (C-4), 116.6 (d, 2JC,F = 23.2 Hz, NPh C-3,5), 120.8 (d, 3JC,F = 8.4 Hz, NPh C-2,6), 128.3 (CPh C-2,6), 128.5 (CPh C-4), 128.7 (CPh C-3,5), 129.5 (C-5), 135.5 (d, 4JC,F = 3.0 Hz, NPh C-1), 136.2 (CPh C-1), 161.6 (d, 1JC,F = 247.7 Hz, NPh C-4), 163.8 (C-3), 183.4 (CHO). 15N NMR (71 MHz, CDCl3): δ −181.0 (N-1), N-2 was not found. MS m/z (%): 297 ([M + H]+, 100). HRMS (ESI) for C17H13FN2O2Na ([M + Na]+): calcd 319.0853, found 319.0853.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1605, 1565, 1509, 1451, 1362,1264, 1234, 1195 (C–F, C–O–C, C
O), 1605, 1565, 1509, 1451, 1362,1264, 1234, 1195 (C–F, C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 877, 743, 733, 674, 458 (CH
C, C–N), 877, 743, 733, 674, 458 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.43 (s, 2H, CH2), 6.99–7.02 (m, 1H, NPh 4-H), 7.33–7.36 (m, 1H, CPh 4-H), 7.38–7.45 (m, 5H, CPh 3,5-H, NPh 2,5,6-H), 7.50–7.52 (m, 2H, CPh 2,6-H), 8.25 (s, 1H, 5-H), 9.87 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 71.3 (CH2), 106.8 (d, 2JC,F = 26.5 Hz, NPh C-2), 112.0 (C-4), 113.8 (d, 4JC,F = 3.2 Hz, NPh C-6), 114.0 (d, 2JC,F = 21.3 Hz, NPh C-4), 128.3 (CPh C-2,6), 128.5 (CPh C-4), 128.6 (CPh C-3,5), 129.6 (C-5), 131.0 (d, 3JC,F = 9.1 Hz, NPh C-5), 136.1 (CPh C-1), 140.4 (d, 3JC,F = 10.2 Hz, NPh C-1), 163.3 (d, 1JC,F = 247.3 Hz, NPh C-3), 163.6 (C-3), 183.3 (CHO). 19F NMR (376 MHz, CDCl3): δ −110.1. MS m/z (%): 297 ([M + H]+, 100). HRMS (ESI) for C17H13FN2O2Na ([M + Na]+): calcd 319.0853, found 319.0853.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.43 (s, 2H, CH2), 6.99–7.02 (m, 1H, NPh 4-H), 7.33–7.36 (m, 1H, CPh 4-H), 7.38–7.45 (m, 5H, CPh 3,5-H, NPh 2,5,6-H), 7.50–7.52 (m, 2H, CPh 2,6-H), 8.25 (s, 1H, 5-H), 9.87 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 71.3 (CH2), 106.8 (d, 2JC,F = 26.5 Hz, NPh C-2), 112.0 (C-4), 113.8 (d, 4JC,F = 3.2 Hz, NPh C-6), 114.0 (d, 2JC,F = 21.3 Hz, NPh C-4), 128.3 (CPh C-2,6), 128.5 (CPh C-4), 128.6 (CPh C-3,5), 129.6 (C-5), 131.0 (d, 3JC,F = 9.1 Hz, NPh C-5), 136.1 (CPh C-1), 140.4 (d, 3JC,F = 10.2 Hz, NPh C-1), 163.3 (d, 1JC,F = 247.3 Hz, NPh C-3), 163.6 (C-3), 183.3 (CHO). 19F NMR (376 MHz, CDCl3): δ −110.1. MS m/z (%): 297 ([M + H]+, 100). HRMS (ESI) for C17H13FN2O2Na ([M + Na]+): calcd 319.0853, found 319.0853.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1561, 1501, 1454, 1361, 1230, 1208, 1199, 1112, (C–F, C–O–C, C
O), 1561, 1501, 1454, 1361, 1230, 1208, 1199, 1112, (C–F, C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 978, 968, 940, 756, 739, 697, 612 (CH
C, C–N), 978, 968, 940, 756, 739, 697, 612 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.44 (s, 2H, CH2), 7.22–7.31 (m, 3H, NPh 3,4,5-H), 7.34–7.37 (m, 1H, CPh 4-H), 7.39–7.42 (m, 2H, CPh 3,5-H), 7.50–7.53 (m, 2H, CPh 2,6-H), 7.89 (m, 1H, NPh 6-H), 8.37 (d, J = 1.8 Hz, 1H, 5-H), 9.89 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 71.3 (CH2), 111.8 (d, 5JC,F = 1.9 Hz, C-4), 117.1 (d, 2JC,F = 20.2 Hz, NPh C-3), 124.0 (C-5), 125.2 (d, 4JC,F = 3.7 Hz, NPh C-5), 127.3 (d, 2JC,F = 8.5 Hz, NPh C-1), 128.3 (CPh C-2,6), 128.4 (d, 3JC,F = 7.9 Hz, NPh C-6), 128.5 (CPh C-4), 128.7 (CPh C-3,5), 134.5 (d, 3JC,F = 12.3 Hz, NPh C-4), 136.2 (CPh C-1), 153.6 (d, 1JC,F = 249.6 Hz, NPh C-2), 163.3 (C-3), 183.4 (CHO). 19F NMR (376 MHz, CDCl3): δ −109.84. MS m/z (%): 297 ([M + H]+, 100). HRMS (ESI) for C17H13FN2O2Na ([M + Na]+): calcd 319.0853, found 319.0853.
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 5.44 (s, 2H, CH2), 7.22–7.31 (m, 3H, NPh 3,4,5-H), 7.34–7.37 (m, 1H, CPh 4-H), 7.39–7.42 (m, 2H, CPh 3,5-H), 7.50–7.53 (m, 2H, CPh 2,6-H), 7.89 (m, 1H, NPh 6-H), 8.37 (d, J = 1.8 Hz, 1H, 5-H), 9.89 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 71.3 (CH2), 111.8 (d, 5JC,F = 1.9 Hz, C-4), 117.1 (d, 2JC,F = 20.2 Hz, NPh C-3), 124.0 (C-5), 125.2 (d, 4JC,F = 3.7 Hz, NPh C-5), 127.3 (d, 2JC,F = 8.5 Hz, NPh C-1), 128.3 (CPh C-2,6), 128.4 (d, 3JC,F = 7.9 Hz, NPh C-6), 128.5 (CPh C-4), 128.7 (CPh C-3,5), 134.5 (d, 3JC,F = 12.3 Hz, NPh C-4), 136.2 (CPh C-1), 153.6 (d, 1JC,F = 249.6 Hz, NPh C-2), 163.3 (C-3), 183.4 (CHO). 19F NMR (376 MHz, CDCl3): δ −109.84. MS m/z (%): 297 ([M + H]+, 100). HRMS (ESI) for C17H13FN2O2Na ([M + Na]+): calcd 319.0853, found 319.0853.General procedure for the synthesis of compounds 4a–f
Into the solution of appropriate 3-benzyloxypyrazole 3a–f (1 mmol) in toluene (10 mL), TFA (10 mL) was added. The mixture was stirred at room temperature for 18 h. Toluene and TFA were evaporated. The residue was purified by flash column chromatography (SiO2, eluent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate/n-hexane, 1
ethyl acetate/n-hexane, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) to give pure compounds 4a–f.
1, v/v) to give pure compounds 4a–f.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1601, 1587, 1537, 1524, 1319, 1220, (C–O–C, C
O), 1601, 1587, 1537, 1524, 1319, 1220, (C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 812 (CH
C, C–N), 812 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 2.40 (s, 3H, CH3), 7.23–7.27 (m, 2H, Ph 3,5-H), 7.49–7.56 (m, 2H, Ph 2,6-H), 8.12 (s, 1H, 5-H), 9.89 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 21.1 (CH3), 109.8 (C-4), 119.3 (Ph C-2,6), 129.6 (C-5), 130.3 (Ph C-3,5), 136.6 (Ph C-1), 137.9 (Ph C-4), 163.7 (C-3), 186.0 (CHO). 15N NMR (41 MHz, CDCl3): δ −178.3 (N-1), N-2 was not found. MS m/z (%): 203 ([M + H]+, 100). HRMS (ESI) for C11H10N2O2Na ([M + Na]+): calcd 225.0635, found 225.0634.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 2.40 (s, 3H, CH3), 7.23–7.27 (m, 2H, Ph 3,5-H), 7.49–7.56 (m, 2H, Ph 2,6-H), 8.12 (s, 1H, 5-H), 9.89 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 21.1 (CH3), 109.8 (C-4), 119.3 (Ph C-2,6), 129.6 (C-5), 130.3 (Ph C-3,5), 136.6 (Ph C-1), 137.9 (Ph C-4), 163.7 (C-3), 186.0 (CHO). 15N NMR (41 MHz, CDCl3): δ −178.3 (N-1), N-2 was not found. MS m/z (%): 203 ([M + H]+, 100). HRMS (ESI) for C11H10N2O2Na ([M + Na]+): calcd 225.0635, found 225.0634.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1574, 1517, 1498, 1459, 1428, 1304, 1252, 1177, 1169, (C–O–C, C
O), 1574, 1517, 1498, 1459, 1428, 1304, 1252, 1177, 1169, (C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N) 1042, 1024, 828, 794 (CH
C, C–N) 1042, 1024, 828, 794 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 3.85 (s, 1H, CH3), 6.96–7.01 (m, 2H, Ph 3,5-H), 7.53–7.59 (m, 2H, Ph 2,6-H), 8.06 (s, 1H, 5-H), 8.59–9.25 (br s, 1H, OH), 9.89 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 55.7 (OCH3), 109.6 (C-4), 114.9 (Ph C-3,5), 121.0 (Ph C-2,6), 129.5 (C-5), 132.5 (Ph C-1), 159.2 (Ph C-4), 163.7 (C-3), 186.1 (CHO). 15N NMR (41 MHz, CDCl3): δ −178.4 (N-1), N-2 was not found. MS m/z (%): 219 ([M + H]+, 100). HRMS (ESI) for C11H10N2O3Na ([M + Na]+): calcd 241.0584, found 241.0584.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 3.85 (s, 1H, CH3), 6.96–7.01 (m, 2H, Ph 3,5-H), 7.53–7.59 (m, 2H, Ph 2,6-H), 8.06 (s, 1H, 5-H), 8.59–9.25 (br s, 1H, OH), 9.89 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 55.7 (OCH3), 109.6 (C-4), 114.9 (Ph C-3,5), 121.0 (Ph C-2,6), 129.5 (C-5), 132.5 (Ph C-1), 159.2 (Ph C-4), 163.7 (C-3), 186.1 (CHO). 15N NMR (41 MHz, CDCl3): δ −178.4 (N-1), N-2 was not found. MS m/z (%): 219 ([M + H]+, 100). HRMS (ESI) for C11H10N2O3Na ([M + Na]+): calcd 241.0584, found 241.0584.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1594, 1539, 1521, 1316, 1223, (C–F, C–O–C, C
O), 1594, 1539, 1521, 1316, 1223, (C–F, C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 1156, 827 (CH
C, C–N), 1156, 827 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.12–7.22 (m, 2H, Ph 3,5-H), 7.61–7.68 (m, 2H, Ph 2,6-H), 8.11 (s, 1H, 5-H), 8.30–9.12 (br s, 1H, OH), 9.91 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 110.0 (C-4), 116.7 (d, 2JC,F = 23.2 Hz, Ph C-3,5), 121.3 (d, 3JC,F = 8.6 Hz, Ph C-2,6), 129.8 (C-5), 135.3 (d, 4JC,F = 2.9 Hz, Ph C-1), 161.9 (d, 1JC,F = 284.4 Hz, Ph C-4), 163.7 (C-3), 186.2 (CHO). 15N NMR (41 MHz, CDCl3): δ −180.2 (N-1), N-2 was not found. MS m/z (%): 207 ([M + H]+, 100). HRMS (ESI) for C10H7FN2O2Na ([M + Na]+): calcd 229.0384, found 229.0384.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.12–7.22 (m, 2H, Ph 3,5-H), 7.61–7.68 (m, 2H, Ph 2,6-H), 8.11 (s, 1H, 5-H), 8.30–9.12 (br s, 1H, OH), 9.91 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 110.0 (C-4), 116.7 (d, 2JC,F = 23.2 Hz, Ph C-3,5), 121.3 (d, 3JC,F = 8.6 Hz, Ph C-2,6), 129.8 (C-5), 135.3 (d, 4JC,F = 2.9 Hz, Ph C-1), 161.9 (d, 1JC,F = 284.4 Hz, Ph C-4), 163.7 (C-3), 186.2 (CHO). 15N NMR (41 MHz, CDCl3): δ −180.2 (N-1), N-2 was not found. MS m/z (%): 207 ([M + H]+, 100). HRMS (ESI) for C10H7FN2O2Na ([M + Na]+): calcd 229.0384, found 229.0384.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 828, 812, 722, 646, 608, 506 (CH
C, C–N), 828, 812, 722, 646, 608, 506 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.04–7.08 (m, 1H, Ph 4-H), 7.26 (s, 1H, Ph 2-H), 7.44–7.46 (m, 2H, Ph 5,6-H), 8.18 (s, 1H, 5-H), 8.69–8.86 (br s, 1H, OH), 9.92 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 107.3 (d, 2JC,F = 26.6 Hz, Ph C-2), 110.3 (C-4), 114.4 (d, 4JC,F = 3.2 Hz, Ph C-6), 114.7 (d, 2JC,F = 21.1 Hz, Ph C-4), 129.9 (C-5), 131.2 (d, 3JC,F = 9.1 Hz, Ph C-5), 140.3 (d, 3JC,F = 10.1 Hz, Ph C-1), 163.4 (d, 1JC,F = 247.9 Hz, Ph C-3), 163.7 (C-3), 186.3 (CHO). 19F NMR (376 MHz, CDCl3): δ −125.3. MS m/z (%): 207 ([M + H]+, 100). HRMS (ESI) for C10H7FN2O2Na ([M + Na]+): calcd 229.0384, found 229.0385.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.04–7.08 (m, 1H, Ph 4-H), 7.26 (s, 1H, Ph 2-H), 7.44–7.46 (m, 2H, Ph 5,6-H), 8.18 (s, 1H, 5-H), 8.69–8.86 (br s, 1H, OH), 9.92 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 107.3 (d, 2JC,F = 26.6 Hz, Ph C-2), 110.3 (C-4), 114.4 (d, 4JC,F = 3.2 Hz, Ph C-6), 114.7 (d, 2JC,F = 21.1 Hz, Ph C-4), 129.9 (C-5), 131.2 (d, 3JC,F = 9.1 Hz, Ph C-5), 140.3 (d, 3JC,F = 10.1 Hz, Ph C-1), 163.4 (d, 1JC,F = 247.9 Hz, Ph C-3), 163.7 (C-3), 186.3 (CHO). 19F NMR (376 MHz, CDCl3): δ −125.3. MS m/z (%): 207 ([M + H]+, 100). HRMS (ESI) for C10H7FN2O2Na ([M + Na]+): calcd 229.0384, found 229.0385.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 813, 795, 763, 655, 615, 465 (CH
C, C–N), 813, 795, 763, 655, 615, 465 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.24–7.37 (m, 3H, Ph 3,4,5-H), 7.88 (t, 3J = 7.9 Hz, 1H, Ph 6-H), 8.31 (s, 1H, 5-H), 9.91 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 110.3 (C-4), 117.1 (d, 2JC,F = 20.4 Hz, Ph C-3), 124.3 (C-5), 125.4 (d, 4JC,F = 3.8 Hz, Ph C-5), 127.0 (d, 2JC,F = 8.5 Hz, Ph C-1), 128.8 (d, 3JC,F = 7.9 Hz, Ph C-6), 134.7 (d, 3JC,F = 12.3 Hz, Ph C-4), 153.7 (d, 1JC,F = 249.3 Hz, Ph C-2), 163.3 (C-3), 186.2 (CHO). MS m/z (%): 207 ([M + H]+, 100). HRMS (ESI) for C10H7FN2O2Na ([M + Na]+): calcd 229.0384, found 229.0385.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.24–7.37 (m, 3H, Ph 3,4,5-H), 7.88 (t, 3J = 7.9 Hz, 1H, Ph 6-H), 8.31 (s, 1H, 5-H), 9.91 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 110.3 (C-4), 117.1 (d, 2JC,F = 20.4 Hz, Ph C-3), 124.3 (C-5), 125.4 (d, 4JC,F = 3.8 Hz, Ph C-5), 127.0 (d, 2JC,F = 8.5 Hz, Ph C-1), 128.8 (d, 3JC,F = 7.9 Hz, Ph C-6), 134.7 (d, 3JC,F = 12.3 Hz, Ph C-4), 153.7 (d, 1JC,F = 249.3 Hz, Ph C-2), 163.3 (C-3), 186.2 (CHO). MS m/z (%): 207 ([M + H]+, 100). HRMS (ESI) for C10H7FN2O2Na ([M + Na]+): calcd 229.0384, found 229.0385.General procedure for the synthesis of compounds 5a–f
Appropriate 3-hydroxypyrazole 4a–f (1 mmol), trifluoromethanesulfonic anhydride (0.17 mL, 1 mmol), and TEA (0.17 mL, 1.2 mmol) were dissolved in DCM (5 mL), and stirred at room temperature for 1 h. The reaction mixture was poured into water and extracted with ethyl acetate. The combined organic layers were washed with brine and dried over Na2SO4; the solvent was evaporated. The residue was purified by flash column chromatography (SiO2, eluent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate/n-hexane, 1
ethyl acetate/n-hexane, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6, v/v) to give pure compounds 5a–f.
6, v/v) to give pure compounds 5a–f.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1557, 1520, 1461, 1429, 1236, 1215, 1136 (C–O–C, C
O), 1557, 1520, 1461, 1429, 1236, 1215, 1136 (C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 885, 603 (CH
C, C–N, C–F), 885, 603 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 2.42 (s, 3H, CH3), 7.28–7.35 (m, 2H, Ph 3,5-H), 7.50–7.58 (m, 2H, Ph 2,6-H), 8.35 (s, 1H, 5-H), 9.91 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 21.2 (CH3), 114.6 (C-4), 118.7 (d, 1JC,F = 321.3 Hz, CF3), 119.6 (Ph C-2,6), 130.5 (Ph C-3,5), 130.8 (C-5), 136.1 (Ph C-1), 139.1 (Ph C-4), 152.2 (C-3), 181.1 (CHO). 15N NMR (41 MHz, CDCl3): δ −170.3 (N-1), N-2 was not found. MS m/z (%): 335 ([M + H]+, 100). HRMS (ESI) for C12H9F3N2O4SNa ([M + Na]+): calcd 357.0127, found 357.0127.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 2.42 (s, 3H, CH3), 7.28–7.35 (m, 2H, Ph 3,5-H), 7.50–7.58 (m, 2H, Ph 2,6-H), 8.35 (s, 1H, 5-H), 9.91 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 21.2 (CH3), 114.6 (C-4), 118.7 (d, 1JC,F = 321.3 Hz, CF3), 119.6 (Ph C-2,6), 130.5 (Ph C-3,5), 130.8 (C-5), 136.1 (Ph C-1), 139.1 (Ph C-4), 152.2 (C-3), 181.1 (CHO). 15N NMR (41 MHz, CDCl3): δ −170.3 (N-1), N-2 was not found. MS m/z (%): 335 ([M + H]+, 100). HRMS (ESI) for C12H9F3N2O4SNa ([M + Na]+): calcd 357.0127, found 357.0127.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1556, 1518, 1465, 1427, 1254, 1233, 1175, 1135 (C–O–C, C
O), 1556, 1518, 1465, 1427, 1254, 1233, 1175, 1135 (C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 1027, 883, 832, 604 (CH
C, C–N, C–F), 1027, 883, 832, 604 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 3.87 (s, 3H, CH3), 6.98–7.04 (m, 2H, Ph 3,5-H), 7.54–7.59 (m, 2H, Ph 2,6-H), 8.29 (s, 1H, 5-H), 9.90 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 55.8 (CH3), 114.5 (C-4), 115.0 (Ph C-3,5), 118.77 (d, 1JC,F = 321.5 Hz, CF3), 121.3 (Ph C-2,6), 130.8 (C-5), 131.8 (Ph C-1), 152.1 (C-3), 160.0 (Ph C-4), 181.2 (CHO). 15N NMR (71 MHz, CDCl3): δ −170.7 (N-1), N-2 was not found. MS m/z (%): 351 ([M + H]+, 100). HRMS (ESI) for C12H19F3N2O5SNa ([M + Na]+): calcd 373.0078, found 373.0076.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 3.87 (s, 3H, CH3), 6.98–7.04 (m, 2H, Ph 3,5-H), 7.54–7.59 (m, 2H, Ph 2,6-H), 8.29 (s, 1H, 5-H), 9.90 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 55.8 (CH3), 114.5 (C-4), 115.0 (Ph C-3,5), 118.77 (d, 1JC,F = 321.5 Hz, CF3), 121.3 (Ph C-2,6), 130.8 (C-5), 131.8 (Ph C-1), 152.1 (C-3), 160.0 (Ph C-4), 181.2 (CHO). 15N NMR (71 MHz, CDCl3): δ −170.7 (N-1), N-2 was not found. MS m/z (%): 351 ([M + H]+, 100). HRMS (ESI) for C12H19F3N2O5SNa ([M + Na]+): calcd 373.0078, found 373.0076.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1557, 1517, 1463, 1434, 1392, 1252, 1229, 1203 (C–F, C–O–C, C
O), 1557, 1517, 1463, 1434, 1392, 1252, 1229, 1203 (C–F, C–O–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 1146, 954, 887, 838, 802, 768, 733, 617, 516 (CH
C, C–N), 1146, 954, 887, 838, 802, 768, 733, 617, 516 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.19–7.25 (m, 2H, Ph 3,5-H), 7.63–7.69 (m, 2H, Ph 2,6-H), 8.35 (s, 1H, 5-H), 9.92 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 114.9 (C-4), 117.0 (d, 2JC,F = 23.4 Hz, Ph C-3,5), 118.7 (1JC,F = 321.3 Hz, CF3), 121.7 (d, 3JC,F = 8.7 Hz, Ph C-2,6), 131.1 (C-5), 134.6 (d, 4JC,F = 3.1 Hz, NPh C-1), 152.4 (C-3), 162.5 (d, 1J = 249.8 Hz, Ph C-4), 181.0 (CHO). 15N NMR (41 MHz, CDCl3): δ −172.6 (N-1), N-2 was not found. MS m/z (%): 339 ([M + H]+, 100). HRMS (ESI) for C11H6F4N2O4SNa ([M + Na]+): calcd 360.9877, found 360.9877.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.19–7.25 (m, 2H, Ph 3,5-H), 7.63–7.69 (m, 2H, Ph 2,6-H), 8.35 (s, 1H, 5-H), 9.92 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 114.9 (C-4), 117.0 (d, 2JC,F = 23.4 Hz, Ph C-3,5), 118.7 (1JC,F = 321.3 Hz, CF3), 121.7 (d, 3JC,F = 8.7 Hz, Ph C-2,6), 131.1 (C-5), 134.6 (d, 4JC,F = 3.1 Hz, NPh C-1), 152.4 (C-3), 162.5 (d, 1J = 249.8 Hz, Ph C-4), 181.0 (CHO). 15N NMR (41 MHz, CDCl3): δ −172.6 (N-1), N-2 was not found. MS m/z (%): 339 ([M + H]+, 100). HRMS (ESI) for C11H6F4N2O4SNa ([M + Na]+): calcd 360.9877, found 360.9877.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 909, 854, 800, 787, 677, 657, 591, 513 (CH
C, C–N), 909, 854, 800, 787, 677, 657, 591, 513 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.13–7.16 (m, 1H, Ph 4-H), 7.44–7.52 (m, 3H, Ph 2,5,6-H), 8.41 (s, 1H, 5-H), 9.92 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 107.7 (d, 2JC,F = 26.7 Hz, Ph C-2), 115.2 (C-4), 114.8 (d, 4JC,F = 3.3 Hz, Ph C-6), 115.9 (d, 2JC,F = 21.1 Hz, Ph C-4), 118.8 (q, 1JC,F = 321.3 Hz, CF3), 131.2 (C-5), 131.5 (d, 3JC,F = 9.0 Hz, Ph C-5), 139.5 (d, 3JC,F = 10.1 Hz, Ph C-1), 152.5 (C-3), 163.3 (d, 1JC,F = 249.5 Hz, Ph C-3), 181.1 (CHO). 19F NMR (376 MHz, CDCl3): δ −109.1. MS m/z (%): 339 ([M + H]+, 100). HRMS (ESI) for C11H6F4N2O2SNa ([M + Na]+): calcd 360.9877, found 360.9876.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.13–7.16 (m, 1H, Ph 4-H), 7.44–7.52 (m, 3H, Ph 2,5,6-H), 8.41 (s, 1H, 5-H), 9.92 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 107.7 (d, 2JC,F = 26.7 Hz, Ph C-2), 115.2 (C-4), 114.8 (d, 4JC,F = 3.3 Hz, Ph C-6), 115.9 (d, 2JC,F = 21.1 Hz, Ph C-4), 118.8 (q, 1JC,F = 321.3 Hz, CF3), 131.2 (C-5), 131.5 (d, 3JC,F = 9.0 Hz, Ph C-5), 139.5 (d, 3JC,F = 10.1 Hz, Ph C-1), 152.5 (C-3), 163.3 (d, 1JC,F = 249.5 Hz, Ph C-3), 181.1 (CHO). 19F NMR (376 MHz, CDCl3): δ −109.1. MS m/z (%): 339 ([M + H]+, 100). HRMS (ESI) for C11H6F4N2O2SNa ([M + Na]+): calcd 360.9877, found 360.9876.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 879, 821, 787, 761, 652, 607, 505 (CH
C, C–N), 879, 821, 787, 761, 652, 607, 505 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.26–7.35 (m, 2H, Ph 3,6-H), 7.39–7.45 (m, 1H, Ph 4-H), 7.83–7.87 (m, 1H, Ph 5-H), 8.49 (s, 1H, 5-H), 9.93 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 114.8 (C-4), 117.4 (d, 2JC,F = 20.1 Hz, Ph C-3), 118.8 (q, 1JC,F = 321.4 Hz, CF3), 124.5 (C-5), 125.6 (d, 4JC,F = 3.8 Hz Ph C-5), 126.5 (d, 2JC,F = 9.1 Hz, Ph C-1), 130.2 (d, 3JC,F = 8.0 Hz, Ph C-4), 135.9 (d, 3JC,F = 11.6 Hz, C-4), 152.3 (C-3), 153.8 (d, 1JC,F = 250.7 Hz, Ph C-2), 181.1 (CHO). 19F NMR (376 MHz, CDCl3): δ −125.0. MS m/z (%): 339 ([M + H]+, 100). HRMS (ESI) for C11H6F4N2O2SNa ([M + Na]+): calcd 360.9877, found 360.9880.
CH of benzene). 1H NMR (400 MHz, CDCl3): δ 7.26–7.35 (m, 2H, Ph 3,6-H), 7.39–7.45 (m, 1H, Ph 4-H), 7.83–7.87 (m, 1H, Ph 5-H), 8.49 (s, 1H, 5-H), 9.93 (s, 1H, CHO). 13C NMR (101 MHz, CDCl3): δ 114.8 (C-4), 117.4 (d, 2JC,F = 20.1 Hz, Ph C-3), 118.8 (q, 1JC,F = 321.4 Hz, CF3), 124.5 (C-5), 125.6 (d, 4JC,F = 3.8 Hz Ph C-5), 126.5 (d, 2JC,F = 9.1 Hz, Ph C-1), 130.2 (d, 3JC,F = 8.0 Hz, Ph C-4), 135.9 (d, 3JC,F = 11.6 Hz, C-4), 152.3 (C-3), 153.8 (d, 1JC,F = 250.7 Hz, Ph C-2), 181.1 (CHO). 19F NMR (376 MHz, CDCl3): δ −125.0. MS m/z (%): 339 ([M + H]+, 100). HRMS (ESI) for C11H6F4N2O2SNa ([M + Na]+): calcd 360.9877, found 360.9880.General procedure for the synthesis of compounds 6a–h
To a solution of appropriate pyrazolyl trifluoromethanesulfonate 5a–f (0.5 mmol) in dry DMF (1 mL) under an inert atmosphere, TEA (4.0 mL, 2.5 mmol), appropriate pyridine acetylene (0.75 mmol), Pd(PPh3)2Cl2 (35 mg, 0.05 mmol), and CuI (18 mg, 0.1 mmol) were added. The reaction mixture was irradiated (100 W) at 130 °C for 1 h. After the completion of the reaction, as indicated by TLC, the mixture was quenched with water (10 mL) and extracted with ethyl acetate (3 × 10 mL). The organic layers were combined, washed with brine, dried over Na2SO4, and concentrated under reduced pressure. The obtained residue was purified by column chromatography (SiO2, eluent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate/n-hexane, 1
ethyl acetate/n-hexane, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8, v/v) to yield compounds 6a–h.
8, v/v) to yield compounds 6a–h.
![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 1680 (CHO), 1595, 1531, 1507, 1406, 1359, 1217 (C
C), 1680 (CHO), 1595, 1531, 1507, 1406, 1359, 1217 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 780, 748, 689, 681, 590 (CH
C, C–N), 780, 748, 689, 681, 590 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.42–7.44 (m, 1H, Ph 4-H), 7.48–7.49 (m, 2H, Pyr 3,5-H), 7.51–7.54 (m, 2H, Ph 3,5-H), 7.74–7.75 (m, 2H, Ph 2,6-H), 8.46 (s, 1H, 5-H), 8.67 (d, J = 5.5 Hz, 2H, Pyr 2,6-H), 10.01 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 83.3 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.42–7.44 (m, 1H, Ph 4-H), 7.48–7.49 (m, 2H, Pyr 3,5-H), 7.51–7.54 (m, 2H, Ph 3,5-H), 7.74–7.75 (m, 2H, Ph 2,6-H), 8.46 (s, 1H, 5-H), 8.67 (d, J = 5.5 Hz, 2H, Pyr 2,6-H), 10.01 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 83.3 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 91.7 (Pyr-C
C), 91.7 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 120.1 (Ph C-2,6), 125.8 (Pyr C-3,5), 130.1 (Pyr C-1), 126.3 (C-4), 128.8 (Ph C-4), 129.3 (C-5), 130.0 (Ph C-3,5), 137.2 (C-3), 138.8 (Ph C-4), 150.1 (Pyr C-2,6), 183.8 (CHO). 15N NMR (71 MHz, CDCl3): δ −156.1 (N-1), −67.5 (Pyr N-1), N-2 was not found. MS m/z (%): 274 ([M + H]+, 100). HRMS (ESI) for C17H12N3O ([M + H]+): calcd 274.0975, found 274.0975.
C), 120.1 (Ph C-2,6), 125.8 (Pyr C-3,5), 130.1 (Pyr C-1), 126.3 (C-4), 128.8 (Ph C-4), 129.3 (C-5), 130.0 (Ph C-3,5), 137.2 (C-3), 138.8 (Ph C-4), 150.1 (Pyr C-2,6), 183.8 (CHO). 15N NMR (71 MHz, CDCl3): δ −156.1 (N-1), −67.5 (Pyr N-1), N-2 was not found. MS m/z (%): 274 ([M + H]+, 100). HRMS (ESI) for C17H12N3O ([M + H]+): calcd 274.0975, found 274.0975.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 1688 (C
C), 1688 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1596, 1566, 1527, 1500, 1412, 1360, 1300, 1236, 1226, 1159, 1022 (C
O), 1596, 1566, 1527, 1500, 1412, 1360, 1300, 1236, 1226, 1159, 1022 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 957, 867, 810, 783, 768, 702, 695, 688, 619, 512 (CH
C, C–N), 957, 867, 810, 783, 768, 702, 695, 688, 619, 512 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.38–7.40 (m, 1H, Pyr 5-H), 7.41–7.43 (m, 1H, Ph 4-H), 7.51–7.53 (m, 2H, Ph 3,5-H), 7.74–7.75 (m, 2H, Ph 2,6-H), 7.95–7.97 (m, 1H, Pyr 4-H), 8.46 (s, 1H, 5-H), 8.64 (br s, 1H, Pyr 6-H), 8.88 (br s, 1H, Pyr 2-H), 10.10 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 82.8 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.38–7.40 (m, 1H, Pyr 5-H), 7.41–7.43 (m, 1H, Ph 4-H), 7.51–7.53 (m, 2H, Ph 3,5-H), 7.74–7.75 (m, 2H, Ph 2,6-H), 7.95–7.97 (m, 1H, Pyr 4-H), 8.46 (s, 1H, 5-H), 8.64 (br s, 1H, Pyr 6-H), 8.88 (br s, 1H, Pyr 2-H), 10.10 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 82.8 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 90.9 (Pyr-C
C), 90.9 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 119.6 (Pyr C-3), 120.1 (Ph C-2,6), 123.6 (Pyr C-5), 126.2 (C-4), 128.7 (Ph C-4), 129.3 (C-5), 129.9 (Ph C-3,5), 137.5 (C-3), 138.8 (Ph C-1), 139.7 (Pyr C-4), 148.9 (Pyr C-6), 151.9 (Pyr C-2), 183.8 (CHO). 15N NMR (71 MHz, CDCl3): δ −157.4 (N-1), −97.5 (N-2), −69.6 (Pyr N-1). MS m/z (%): 274 ([M + H]+, 100). HRMS (ESI) for C17H12N3O ([M + H]+): calcd 274.0975, found 274.0974.
C), 119.6 (Pyr C-3), 120.1 (Ph C-2,6), 123.6 (Pyr C-5), 126.2 (C-4), 128.7 (Ph C-4), 129.3 (C-5), 129.9 (Ph C-3,5), 137.5 (C-3), 138.8 (Ph C-1), 139.7 (Pyr C-4), 148.9 (Pyr C-6), 151.9 (Pyr C-2), 183.8 (CHO). 15N NMR (71 MHz, CDCl3): δ −157.4 (N-1), −97.5 (N-2), −69.6 (Pyr N-1). MS m/z (%): 274 ([M + H]+, 100). HRMS (ESI) for C17H12N3O ([M + H]+): calcd 274.0975, found 274.0974.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 1682 (C
C), 1682 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1598, 1581, 1562, 1530, 1505, 1486, 1465, 1425, 1363, 1318, 1285, 1221, 1150, 1074, 1053 (C
O), 1598, 1581, 1562, 1530, 1505, 1486, 1465, 1425, 1363, 1318, 1285, 1221, 1150, 1074, 1053 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 864, 780, 761, 755, 703, 686, 621, 525, 509, 438 (CH
C, C–N), 864, 780, 761, 755, 703, 686, 621, 525, 509, 438 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.32–7.33 (m, 1H, Pyr 5-H), 7.41–7.43 (m, 1H, Ph 4-H), 7.51–7.53 (m, 2H, Ph 3,5-H), 7.64–7.65 (m, 1H, Pyr 3-H), 7.73–7.76 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.46 (s, 1H, 5-H), 8.63–8.74 (m, 1H, Pyr 6-H), 10.15 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.5 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.32–7.33 (m, 1H, Pyr 5-H), 7.41–7.43 (m, 1H, Ph 4-H), 7.51–7.53 (m, 2H, Ph 3,5-H), 7.64–7.65 (m, 1H, Pyr 3-H), 7.73–7.76 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.46 (s, 1H, 5-H), 8.63–8.74 (m, 1H, Pyr 6-H), 10.15 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.5 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 93.6 (Pyr-C
C), 93.6 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 120.1 (Ph C-2,6), 123.9 (Pyr C-5), 126.5 (C-4), 127.9 (Pyr C-3), 128.6 (C-5), 128.7 (Ph C-4), 129.9 (Ph C-3,5), 136.4 (Pyr C-4), 137.8 (C-3), 138.9 (Ph C-1), 142.4 (Pyr C-2), 150.5 (Pyr C-6), 184.4 (CHO). 15N NMR (71 MHz, CDCl3): δ −157.2 (N-1), −65.8 (Pyr N-1), N-2 was not found. MS m/z (%): 274 ([M + H]+, 100). HRMS (ESI) for C17H12N3O ([M + H]+): calcd 274.0975, found 274.0975.
C), 120.1 (Ph C-2,6), 123.9 (Pyr C-5), 126.5 (C-4), 127.9 (Pyr C-3), 128.6 (C-5), 128.7 (Ph C-4), 129.9 (Ph C-3,5), 136.4 (Pyr C-4), 137.8 (C-3), 138.9 (Ph C-1), 142.4 (Pyr C-2), 150.5 (Pyr C-6), 184.4 (CHO). 15N NMR (71 MHz, CDCl3): δ −157.2 (N-1), −65.8 (Pyr N-1), N-2 was not found. MS m/z (%): 274 ([M + H]+, 100). HRMS (ESI) for C17H12N3O ([M + H]+): calcd 274.0975, found 274.0975.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 1682 (C
C), 1682 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1581, 1535, 1522, 1485, 1426, 1362, 1271, 1234, 1223, 1155, 1052 (C
O), 1581, 1535, 1522, 1485, 1426, 1362, 1271, 1234, 1223, 1155, 1052 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 959, 866, 813, 784, 772, 703, 628, 612, 506 (CH
C, C–N, C–F), 959, 866, 813, 784, 772, 703, 628, 612, 506 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 2.16 (s, 3H, CH3), 7.28–7.32 (m, 3H, Pyr 5-H, Ph 3,5-H), 7.60–7.65 (m, 3H, Pyr 3-H, Ph 2,6-H), 7.71–7.74 (m, 1H, Pyr 4-H), 8.40 (s, 1H, 5-H), 8.65–8.69 (m, 1H, Pyr 6-H), 10.12 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 21.2 (CH3), 78.0 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 2.16 (s, 3H, CH3), 7.28–7.32 (m, 3H, Pyr 5-H, Ph 3,5-H), 7.60–7.65 (m, 3H, Pyr 3-H, Ph 2,6-H), 7.71–7.74 (m, 1H, Pyr 4-H), 8.40 (s, 1H, 5-H), 8.65–8.69 (m, 1H, Pyr 6-H), 10.12 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 21.2 (CH3), 78.0 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 93.5 (Pyr-C
C), 93.5 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 119.9 (Ph C-2,6), 123.8 (Pyr C-5), 126.3 (C-4), 127.9 (Pyr C-3), 128.6 (C-5), 130.4 (Ph C-3,5), 136.4 (Pyr C-4), 136.6 (Ph C-4), 137.5 (C-3), 138.7 (Ph C-1), 142.4 (Pyr C-2), 150.4 (Pyr C-6), 184.4 (CHO). MS m/z (%): 288 ([M + H]+, 100). HRMS (ESI) for C18H14N3O ([M + H]+): calcd 288.1131, found 288.1125.
C), 119.9 (Ph C-2,6), 123.8 (Pyr C-5), 126.3 (C-4), 127.9 (Pyr C-3), 128.6 (C-5), 130.4 (Ph C-3,5), 136.4 (Pyr C-4), 136.6 (Ph C-4), 137.5 (C-3), 138.7 (Ph C-1), 142.4 (Pyr C-2), 150.4 (Pyr C-6), 184.4 (CHO). MS m/z (%): 288 ([M + H]+, 100). HRMS (ESI) for C18H14N3O ([M + H]+): calcd 288.1131, found 288.1125.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 1683 (C
C), 1683 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1533, 1518, 1487, 1455, 1437, 1257, 1219, 1173, 1025 (C
O), 1533, 1518, 1487, 1455, 1437, 1257, 1219, 1173, 1025 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 959, 825, 777, 758, 721, 696, 617, 541, 520, 442 (CH
C, C–N), 959, 825, 777, 758, 721, 696, 617, 541, 520, 442 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 3.86 (s, 3H, CH3), 7.00–7.02 (m, 2H, Ph 3,5-H), 7.31–7.34 (m, 1H, Pyr 5-H), 7.63–7.67 (m, 3H, Pyr 3-H, Ph 2,6-H), 7.72–7.75 (m, 1H, Pyr 4-H), 8.36 (s, 1H, 5-H), 8.67–8.68 (m, 1H, Pyr 6-H), 10.13 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 55.8 (CH3), 78.6 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 3.86 (s, 3H, CH3), 7.00–7.02 (m, 2H, Ph 3,5-H), 7.31–7.34 (m, 1H, Pyr 5-H), 7.63–7.67 (m, 3H, Pyr 3-H, Ph 2,6-H), 7.72–7.75 (m, 1H, Pyr 4-H), 8.36 (s, 1H, 5-H), 8.67–8.68 (m, 1H, Pyr 6-H), 10.13 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 55.8 (CH3), 78.6 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 93.4 (Pyr-C
C), 93.4 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 114.9 (Ph C-3,5), 121.6 (Ph C-2,6), 123.8 (Pyr C-5), 126.3 (C-4), 127.9 (Pyr C-3), 128.5 (C-5), 132.4 (Ph C-1), 136.4 (Pyr C-4), 137.5 (C-3), 142.4 (Pyr C-2), 150.4 (Pyr C-6), 159.8 (Ph C-4), 184.4 (CHO). MS m/z (%): 304 ([M + H]+, 100). HRMS (ESI) for C13H13N3O2Na ([M + Na]): calcd 326.0900, found 326.0904.
C), 114.9 (Ph C-3,5), 121.6 (Ph C-2,6), 123.8 (Pyr C-5), 126.3 (C-4), 127.9 (Pyr C-3), 128.5 (C-5), 132.4 (Ph C-1), 136.4 (Pyr C-4), 137.5 (C-3), 142.4 (Pyr C-2), 150.4 (Pyr C-6), 159.8 (Ph C-4), 184.4 (CHO). MS m/z (%): 304 ([M + H]+, 100). HRMS (ESI) for C13H13N3O2Na ([M + Na]): calcd 326.0900, found 326.0904.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 1683 (C
C), 1683 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1586, 1534, 1488, 1480, 1471, 1362, 1241, 1226, 1157, 1112, 1054 (C
O), 1586, 1534, 1488, 1480, 1471, 1362, 1241, 1226, 1157, 1112, 1054 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 958, 931, 875, 830, 793, 774, 762, 747, 657, 513, 435 (CH
C, C–N, C–F), 958, 931, 875, 830, 793, 774, 762, 747, 657, 513, 435 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.19–7.22 (m, 2H, Ph 3,5-H), 7.31–7.34 (m, 1H, Pyr 5-H), 7.63–7.64 (m, 1H, Pyr 3-H), 7.71–7.75 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.40 (s, 1H, 5-H), 8.66–8.69 (m, 1H, Pyr 6-H), 10.13 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.3 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.19–7.22 (m, 2H, Ph 3,5-H), 7.31–7.34 (m, 1H, Pyr 5-H), 7.63–7.64 (m, 1H, Pyr 3-H), 7.71–7.75 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.40 (s, 1H, 5-H), 8.66–8.69 (m, 1H, Pyr 6-H), 10.13 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.3 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 93.7 (Pyr-C
C), 93.7 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 116.9 (d, 2JC,F = 23.3 Hz, Ph C-3,5), 122.0 (d, 3JC,F = 8.6 Hz, Ph C-2,6), 123.9 (Pyr C-5), 126.5 (C-4), 127.9 (Pyr C-3), 128.8 (C-5), 135.2 (d, 4JC,F = 3.0 Hz, Ph C-1), 136.5 (Pyr C-4), 137.8 (C-3), 142.3 (Pyr C-2), 150.5 (Pyr C-6), 162.4 (d, 1JC,F = 249.2 Hz, Ph C-2), 184.4 (CHO). MS m/z (%): 292 ([M + H]+, 100). HRMS (ESI) for C17H10N3OFNa ([M + Na]+): calcd 314.0700, found 314.0699.
C), 116.9 (d, 2JC,F = 23.3 Hz, Ph C-3,5), 122.0 (d, 3JC,F = 8.6 Hz, Ph C-2,6), 123.9 (Pyr C-5), 126.5 (C-4), 127.9 (Pyr C-3), 128.8 (C-5), 135.2 (d, 4JC,F = 3.0 Hz, Ph C-1), 136.5 (Pyr C-4), 137.8 (C-3), 142.3 (Pyr C-2), 150.5 (Pyr C-6), 162.4 (d, 1JC,F = 249.2 Hz, Ph C-2), 184.4 (CHO). MS m/z (%): 292 ([M + H]+, 100). HRMS (ESI) for C17H10N3OFNa ([M + Na]+): calcd 314.0700, found 314.0699.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 898, 860, 775, 673, 611, 454 (CH
C, C–N, C–F), 898, 860, 775, 673, 611, 454 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 7.07–7.11 (m, 1H, Ph 4-H), 7.30–7.32 (m, 1H, Pyr 5-H), 7.50–7.55 (m, 3H, Ph 2,5,6-H), 7.62–7.64 (m, 1H, Pyr 3-H), 7.71–7.74 (m, 1H, Pyr 4-H), 8.46 (s, 1H, 5-H), 8.64–8.66 (m, 1H, Pyr 6-H) 10.12 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.2 (Pyr-C
CH of benzenes). 1H NMR (700 MHz, CDCl3): δ 7.07–7.11 (m, 1H, Ph 4-H), 7.30–7.32 (m, 1H, Pyr 5-H), 7.50–7.55 (m, 3H, Ph 2,5,6-H), 7.62–7.64 (m, 1H, Pyr 3-H), 7.71–7.74 (m, 1H, Pyr 4-H), 8.46 (s, 1H, 5-H), 8.64–8.66 (m, 1H, Pyr 6-H) 10.12 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.2 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 93.8 (Pyr-C
C), 93.8 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 107.9 (d, 2JC,F = 26.6 Hz, Ph C-2), 115.1 (d, 4JC,F = 3.2 Hz, Ph C-6), 115.4 (d, 2JC,F = 21.1 Hz, Ph C-4), 123.9 (Pyr C-5), 126.6 (C-4), 127.9 (Pyr C-3), 128.5 (C-5), 131.3 (d, 3JC,F = 9.1 Hz, Ph C-5), 136.4 (Pyr C-4), 137.8 (C-3), 139.9 (d, 3JC,F = 10.1 Hz, Ph C-1), 142.2 (Pyr C-2), 150.5 (Pyr C-6), 163.3 (d, 1JC,F = 248.6 Hz, Ph C-3), 184.2 (CHO). MS m/z (%): 292 ([M + H]+, 100). HRMS (ESI) for C17H10N3OFNa ([M + Na]+): calcd 314.0700, found 314.0698.
C), 107.9 (d, 2JC,F = 26.6 Hz, Ph C-2), 115.1 (d, 4JC,F = 3.2 Hz, Ph C-6), 115.4 (d, 2JC,F = 21.1 Hz, Ph C-4), 123.9 (Pyr C-5), 126.6 (C-4), 127.9 (Pyr C-3), 128.5 (C-5), 131.3 (d, 3JC,F = 9.1 Hz, Ph C-5), 136.4 (Pyr C-4), 137.8 (C-3), 139.9 (d, 3JC,F = 10.1 Hz, Ph C-1), 142.2 (Pyr C-2), 150.5 (Pyr C-6), 163.3 (d, 1JC,F = 248.6 Hz, Ph C-3), 184.2 (CHO). MS m/z (%): 292 ([M + H]+, 100). HRMS (ESI) for C17H10N3OFNa ([M + Na]+): calcd 314.0700, found 314.0698.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 990, 960, 850, 817, 780, 773, 754, 720, 675, 634, 607 (CH
C, C–N, C–F), 990, 960, 850, 817, 780, 773, 754, 720, 675, 634, 607 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.28–7.34 (m, 3H, Ph 3,4,5-H), 7.39–7.41 (m, 1H, Pyr 5-H), 7.65 (d, J = 7.8 Hz, 1H, Pyr 3-H), 7.73–7.76 (m, 1H, Pyr 4-H), 7.95–7.98 (m, 1H, Ph 6-H), 8.55 (d, J = 2.1 Hz, 1H, 5-H), 8.68 (d, J = 4.3 Hz, 1H, Pyr 6-H), 10.15 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.3 (Pyr-C
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.28–7.34 (m, 3H, Ph 3,4,5-H), 7.39–7.41 (m, 1H, Pyr 5-H), 7.65 (d, J = 7.8 Hz, 1H, Pyr 3-H), 7.73–7.76 (m, 1H, Pyr 4-H), 7.95–7.98 (m, 1H, Ph 6-H), 8.55 (d, J = 2.1 Hz, 1H, 5-H), 8.68 (d, J = 4.3 Hz, 1H, Pyr 6-H), 10.15 (s, 1H, CHO). 13C NMR (176 MHz, CDCl3): δ 78.3 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 93.7 (Pyr-C
C), 93.7 (Pyr-C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) C), 117.2 (d, 2JC,F = 20.2 Hz, Ph C-3), 123.9 (Pyr C-5), 124.8 (Ph C-6), 125.4 (d, 3JC,F = 3.8 Hz, Ph C-5), 126.3 (C-4), 127.0 (d, 2JC,F = 9.1 Hz, Ph C-1), 127.9 (Pyr C-3), 129.8 (d, 3JC,F = 8.0 Hz, Ph C-4), 133.1 (d, 3JC,F = 11.1 Hz, C-5), 136.4 (Pyr C-4), 137.3 (C-3), 142.3 (Pyr C-2), 150.4 (Pyr C-6), 153.7 (d, 1JC,F = 250.9 Hz, Ph C-2), 184.2 (CHO). MS m/z (%): 292 ([M + H]+, 100). HRMS (ESI) for C17H10N3OFNa ([M + Na]+): calcd 314.0700, found 314.0704.
C), 117.2 (d, 2JC,F = 20.2 Hz, Ph C-3), 123.9 (Pyr C-5), 124.8 (Ph C-6), 125.4 (d, 3JC,F = 3.8 Hz, Ph C-5), 126.3 (C-4), 127.0 (d, 2JC,F = 9.1 Hz, Ph C-1), 127.9 (Pyr C-3), 129.8 (d, 3JC,F = 8.0 Hz, Ph C-4), 133.1 (d, 3JC,F = 11.1 Hz, C-5), 136.4 (Pyr C-4), 137.3 (C-3), 142.3 (Pyr C-2), 150.4 (Pyr C-6), 153.7 (d, 1JC,F = 250.9 Hz, Ph C-2), 184.2 (CHO). MS m/z (%): 292 ([M + H]+, 100). HRMS (ESI) for C17H10N3OFNa ([M + Na]+): calcd 314.0700, found 314.0704.General procedure for the synthesis of compounds 7a–h
A solution of compound 6a–h (0.5 mmol) in dry ammonia and methanol (NH3/MeOH 2 M, 8 mL) was heated at 120 °C for 15 h in a steel reactor. The solvent was evaporated, and the crude was purified by flash chromatography (SiO2, eluent![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate/n-hexane, 1
ethyl acetate/n-hexane, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, v/v) to yield compounds 7a–h.
2, v/v) to yield compounds 7a–h.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 925, 829, 756, 746, 691, 501, 432 (CH
C, C–N), 925, 829, 756, 746, 691, 501, 432 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of monosubstituted benzene). 1H NMR (700 MHz, CDCl3): δ 7.47–7.51 (m, 1H, Ph 4-H), 7.56–7.60 (m, 2H, Ph 3,5-H), 7.92–7.95 (m, 2H, Ph 2,6-H), 7.98–8.00 (m, 2H, Pyr 3,5-H), 8.14 (dd, 5J = 1.3, 0.9 Hz, 1H, 7-H), 8.63 (d, 5J = 0.9 Hz, 1H, 3-H), 8.71–8.76 (m, 2H, Pyr 2,6-H), 9.34 (d, J = 1.3 Hz, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 107.6 (C-7), 119.4 (C-3a), 120.2 (Pyr C-3,5), 120.4 (Ph C-2,6), 121.1 (C-3), 128.1 (Ph C-4), 128.9 (Ph C-3,5), 138.8 (Ph C-1), 146.1 (Pyr C-4), 146.5 (C-4), 147.3 (C-6), 149.2 (Pyr C-2,6), 150.2 (C-7a). 15N NMR (71 MHz, CDCl3): δ −144.2 (N-2), −98.0 (N-1), −86.8 (N-5), −72.3 (Pyr N-1). MS m/z (%): 273 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1135.
CH of monosubstituted benzene). 1H NMR (700 MHz, CDCl3): δ 7.47–7.51 (m, 1H, Ph 4-H), 7.56–7.60 (m, 2H, Ph 3,5-H), 7.92–7.95 (m, 2H, Ph 2,6-H), 7.98–8.00 (m, 2H, Pyr 3,5-H), 8.14 (dd, 5J = 1.3, 0.9 Hz, 1H, 7-H), 8.63 (d, 5J = 0.9 Hz, 1H, 3-H), 8.71–8.76 (m, 2H, Pyr 2,6-H), 9.34 (d, J = 1.3 Hz, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 107.6 (C-7), 119.4 (C-3a), 120.2 (Pyr C-3,5), 120.4 (Ph C-2,6), 121.1 (C-3), 128.1 (Ph C-4), 128.9 (Ph C-3,5), 138.8 (Ph C-1), 146.1 (Pyr C-4), 146.5 (C-4), 147.3 (C-6), 149.2 (Pyr C-2,6), 150.2 (C-7a). 15N NMR (71 MHz, CDCl3): δ −144.2 (N-2), −98.0 (N-1), −86.8 (N-5), −72.3 (Pyr N-1). MS m/z (%): 273 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1135.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 810, 771, 751, 708, 695, 580 (CH
C, C–N), 810, 771, 751, 708, 695, 580 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.40 (dd, J = 7.9, 4.7 Hz, 1H, Pyr 5-H), 7.45–7.48 (m, 1H, Ph 4-H), 7.53–7.57 (m, 2H, Ph 3,5-H), 7.90–7.93 (m, 2H, Ph 2,6-H), 8.03 (dd, 5J = 1.3, 0.9 Hz, 1H, 7-H), 8.37 (dt, J = 7.9, 1.9 Hz, 1H, Pyr 4-H), 8.60 (d, 5J = 0.9 Hz, 1H, 3-H), 8.61–8.66 (m, 1H, Pyr 6-H), 9.29–9.31 (m, 1H, Pyr 2-H), 9.31 (d, J = 1.3 Hz, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 107.7 (C-7), 119.9 (C-3a), 121.2 (Ph C-2,6), 122.0 (C-3), 123.5 (Pyr C-5), 129.0 (Ph C-4), 129.8 (Ph C-3,5), 134.4 (Pyr C-4), 135.5 (Pyr C-3), 139.8 (Ph C-1), 147.6 (C-4), 148.3 (Pyr C-2), 148.4 (C-6), 149.3 (Pyr C-6), 151.3 (C-7a). 15N NMR (71 MHz, CDCl3): δ −145.0 (N-2), −98.9 (N-1), −86.6 (N-5), −70.7 (Pyr N-1). MS m/z (%): 273 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1134.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.40 (dd, J = 7.9, 4.7 Hz, 1H, Pyr 5-H), 7.45–7.48 (m, 1H, Ph 4-H), 7.53–7.57 (m, 2H, Ph 3,5-H), 7.90–7.93 (m, 2H, Ph 2,6-H), 8.03 (dd, 5J = 1.3, 0.9 Hz, 1H, 7-H), 8.37 (dt, J = 7.9, 1.9 Hz, 1H, Pyr 4-H), 8.60 (d, 5J = 0.9 Hz, 1H, 3-H), 8.61–8.66 (m, 1H, Pyr 6-H), 9.29–9.31 (m, 1H, Pyr 2-H), 9.31 (d, J = 1.3 Hz, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 107.7 (C-7), 119.9 (C-3a), 121.2 (Ph C-2,6), 122.0 (C-3), 123.5 (Pyr C-5), 129.0 (Ph C-4), 129.8 (Ph C-3,5), 134.4 (Pyr C-4), 135.5 (Pyr C-3), 139.8 (Ph C-1), 147.6 (C-4), 148.3 (Pyr C-2), 148.4 (C-6), 149.3 (Pyr C-6), 151.3 (C-7a). 15N NMR (71 MHz, CDCl3): δ −145.0 (N-2), −98.9 (N-1), −86.6 (N-5), −70.7 (Pyr N-1). MS m/z (%): 273 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1134.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 918, 871, 804, 788, 765, 754, 740, 690, 673, 652, 622, 553, 492, 432, 406 (CH
C, C–N), 918, 871, 804, 788, 765, 754, 740, 690, 673, 652, 622, 553, 492, 432, 406 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.27–7.31 (m, 1H, Pyr 5-H), 7.46–7.48 (m, 1H, Ph 4-H), 7.56–7.58 (m, 2H, Ph 3,5-H), 7.81–7.83 (m, 1H, Pyr 4-H), 7.94–7.95 (m, 2H, Ph 2,6-H), 8.42 (d, J = 7.7 Hz, 1H, Pyr 3-H), 8.61 (s, 1H, 3-H), 8.71–8.75 (m, 2H, 7-H, Pyr 6-H), 9.32 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.9 (C-7), 120.7 (C-3a), 121.5 (Pyr C-3), 121.6 (Ph C-2,6), 122.0 (C-3), 123.2 (Pyr C-5), 129.0 (Ph C-4), 129.9 (Ph C-3,5), 137.0 (Pyr C-4), 140.2 (Ph C-1), 146.9 (C-4), 149.6 (Pyr C-6), 150.2 (C-6), 151.8 (C-7a), 156.8 (Pyr C-2). 15N NMR (71 MHz, CDCl3): δ −144.7 (N-2), −97.2 (N-1), −88.5 (N-5), −77.4 (Pyr N-1). MS m/z (%): 273 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1135.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.27–7.31 (m, 1H, Pyr 5-H), 7.46–7.48 (m, 1H, Ph 4-H), 7.56–7.58 (m, 2H, Ph 3,5-H), 7.81–7.83 (m, 1H, Pyr 4-H), 7.94–7.95 (m, 2H, Ph 2,6-H), 8.42 (d, J = 7.7 Hz, 1H, Pyr 3-H), 8.61 (s, 1H, 3-H), 8.71–8.75 (m, 2H, 7-H, Pyr 6-H), 9.32 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.9 (C-7), 120.7 (C-3a), 121.5 (Pyr C-3), 121.6 (Ph C-2,6), 122.0 (C-3), 123.2 (Pyr C-5), 129.0 (Ph C-4), 129.9 (Ph C-3,5), 137.0 (Pyr C-4), 140.2 (Ph C-1), 146.9 (C-4), 149.6 (Pyr C-6), 150.2 (C-6), 151.8 (C-7a), 156.8 (Pyr C-2). 15N NMR (71 MHz, CDCl3): δ −144.7 (N-2), −97.2 (N-1), −88.5 (N-5), −77.4 (Pyr N-1). MS m/z (%): 273 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1135.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N), 918, 871, 804, 788, 765, 754, 740, 690, 673, 652, 622, 553, 492, 432, 406 (CH
C, C–N), 918, 871, 804, 788, 765, 754, 740, 690, 673, 652, 622, 553, 492, 432, 406 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of disubstituted benzene). 1H NMR (400 MHz, CDCl3): δ 2.38 (s, 3H, CH3), 7.21–7.24 (m, 1H, Pyr 5-H), 7.28–7.30 (m, 2H, Ph 3,5-H), 7.74–7.76 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.35 (d, 1H, J = 8.0 Hz, Pyr 3-H), 8.57 (s, 1H, 3-H), 8.65–8.67 (m, 2H, 7-H, Pyr 6-H), 9.25 (s, 1H, 4-H). 13C NMR (101 MHz, CDCl3): δ 21.2 (CH3), 108.7 (C-7), 120.5 (C-3a), 121.29 (Ph C-2,6), 121.32 (Pyr C-3), 121.8 (C-3), 123.1 (Pyr C-5), 130.3 (Ph C-3,5), 136.8 (Pyr C-4), 137.8 (Ph C-4), 139.1 (Ph C-1), 146.6 (C-4), 149.5 (Pyr C-6), 149.9 (C-6), 151.6 (C-7a), 156.7 (Pyr C-2). MS m/z (%): 287 ([M + H]+, 100). HRMS (ESI) for C18H15N4 ([M + H]+): calcd 287.1291, found 287.1295.
CH of disubstituted benzene). 1H NMR (400 MHz, CDCl3): δ 2.38 (s, 3H, CH3), 7.21–7.24 (m, 1H, Pyr 5-H), 7.28–7.30 (m, 2H, Ph 3,5-H), 7.74–7.76 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.35 (d, 1H, J = 8.0 Hz, Pyr 3-H), 8.57 (s, 1H, 3-H), 8.65–8.67 (m, 2H, 7-H, Pyr 6-H), 9.25 (s, 1H, 4-H). 13C NMR (101 MHz, CDCl3): δ 21.2 (CH3), 108.7 (C-7), 120.5 (C-3a), 121.29 (Ph C-2,6), 121.32 (Pyr C-3), 121.8 (C-3), 123.1 (Pyr C-5), 130.3 (Ph C-3,5), 136.8 (Pyr C-4), 137.8 (Ph C-4), 139.1 (Ph C-1), 146.6 (C-4), 149.5 (Pyr C-6), 149.9 (C-6), 151.6 (C-7a), 156.7 (Pyr C-2). MS m/z (%): 287 ([M + H]+, 100). HRMS (ESI) for C18H15N4 ([M + H]+): calcd 287.1291, found 287.1295.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–O–C), 920, 832, 792, 748, 698, 634, 521 (CH
C, C–N, C–O–C), 920, 832, 792, 748, 698, 634, 521 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 3.90 (s, 3H, CH3), 7.06–7.08 (m, 2H, Ph 3,5-H), 7.27–7.32 (m, 1H, Pyr 5-H), 7.81–7.87 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.39–8.44 (m, 1H, Pyr 3-H), 8.53 (s 1H, 3-H), 8.71–8.73 (m, 2H, 7-H, Pyr 6-H), 9.31 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 55.8 (CH3), 108.8 (C-7), 115.0 (Ph C-3,5), 120.7 (C-3a), 121.5 (Pyr C-3), 121.8 (C-3), 123.0 (Ph C-2,6), 123.2 (Pyr C-5), 133.7 (Ph C-1), 137.0 (Pyr C-4), 146.6 (C-4), 149.6 (Pyr C-6), 150.1 (C-6), 151.7 (C-7a), 157.0 (Pyr C-2), 160.1 (Ph C-4). 15N NMR (71 MHz, CDCl3): δ −144.8 (N-2), N-1, N-5, Pyr N-1 were not found. MS m/z (%): 303 ([M + H]+, 100). HRMS (ESI) for C18H15N4O ([M + H]+): calcd 303.1240, found 303.1242.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 3.90 (s, 3H, CH3), 7.06–7.08 (m, 2H, Ph 3,5-H), 7.27–7.32 (m, 1H, Pyr 5-H), 7.81–7.87 (m, 3H, Ph 2,6-H, Pyr 4-H), 8.39–8.44 (m, 1H, Pyr 3-H), 8.53 (s 1H, 3-H), 8.71–8.73 (m, 2H, 7-H, Pyr 6-H), 9.31 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 55.8 (CH3), 108.8 (C-7), 115.0 (Ph C-3,5), 120.7 (C-3a), 121.5 (Pyr C-3), 121.8 (C-3), 123.0 (Ph C-2,6), 123.2 (Pyr C-5), 133.7 (Ph C-1), 137.0 (Pyr C-4), 146.6 (C-4), 149.6 (Pyr C-6), 150.1 (C-6), 151.7 (C-7a), 157.0 (Pyr C-2), 160.1 (Ph C-4). 15N NMR (71 MHz, CDCl3): δ −144.8 (N-2), N-1, N-5, Pyr N-1 were not found. MS m/z (%): 303 ([M + H]+, 100). HRMS (ESI) for C18H15N4O ([M + H]+): calcd 303.1240, found 303.1242.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 930, 844, 810, 788, 757, 620 (CH
C, C–N, C–F), 930, 844, 810, 788, 757, 620 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.25–7.28 (m, 2H, Ph 3,5-H), 7.30 (t, J = 6.1 Hz, 1H, Pyr 5-H), 7.84 (t, J = 7.8 Hz, 1H, Pyr 4-H), 7.91–7.95 (m, 2H, Ph 2,6-H), 8.43 (d, J = 7.8 Hz, 1H, Pyr 3-H), 8.56 (s, 1H, 3-H), 8.72 (s, 1H, 7-H), 8.73–8.76 (m, 1H, Pyr 6-H), 9.33 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.6 (C-7), 116.7 (d, 2JC,F = 23.1 Hz, Ph C-3,5), 120.6 (C-3a), 121.4 (Pyr C-3), 121.9 (C-3), 123.2 (Pyr C-5), 123.3 (d, 3JC,F = 8.6 Hz, Ph C-2,6), 136.3 (d, 4JC,F = 2.3 Hz, Ph C-1), 136.9 (Pyr C-4), 146.7 (C-4), 149.5 (Pyr C-6), 150.1 (C-6), 151.7 (C-7a), 156.6 (Pyr C-2), 162.6 (d, 1JC,F = 249.6 Hz, Ph C-4). 15N NMR (71 MHz, CDCl3): δ −146.9 (N-2), −96.7 (N-1), −87.7 (N-5), −77.2 (Pyr N-1). 19F NMR (376 MHz, CDCl3): δ −112.1. MS m/z (%): 291 ([M + H]+, 100). HRMS (ESI) for C17H12N4F ([M + H]+): calcd 291.1041, found 291.1038.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.25–7.28 (m, 2H, Ph 3,5-H), 7.30 (t, J = 6.1 Hz, 1H, Pyr 5-H), 7.84 (t, J = 7.8 Hz, 1H, Pyr 4-H), 7.91–7.95 (m, 2H, Ph 2,6-H), 8.43 (d, J = 7.8 Hz, 1H, Pyr 3-H), 8.56 (s, 1H, 3-H), 8.72 (s, 1H, 7-H), 8.73–8.76 (m, 1H, Pyr 6-H), 9.33 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.6 (C-7), 116.7 (d, 2JC,F = 23.1 Hz, Ph C-3,5), 120.6 (C-3a), 121.4 (Pyr C-3), 121.9 (C-3), 123.2 (Pyr C-5), 123.3 (d, 3JC,F = 8.6 Hz, Ph C-2,6), 136.3 (d, 4JC,F = 2.3 Hz, Ph C-1), 136.9 (Pyr C-4), 146.7 (C-4), 149.5 (Pyr C-6), 150.1 (C-6), 151.7 (C-7a), 156.6 (Pyr C-2), 162.6 (d, 1JC,F = 249.6 Hz, Ph C-4). 15N NMR (71 MHz, CDCl3): δ −146.9 (N-2), −96.7 (N-1), −87.7 (N-5), −77.2 (Pyr N-1). 19F NMR (376 MHz, CDCl3): δ −112.1. MS m/z (%): 291 ([M + H]+, 100). HRMS (ESI) for C17H12N4F ([M + H]+): calcd 291.1041, found 291.1038.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 934, 875, 793, 755, 721, 695, 542 (CH
C, C–N, C–F), 934, 875, 793, 755, 721, 695, 542 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.15–7.19 (m, 1H, Ph 4-H), 7.29–7.31 (m, 1H, Pyr 5-H), 7.51–7.56 (m, 1H, Ph 5-H), 7.72–7.76 (m, 2H, Ph 2,6-H), 7.80–7.84 (m, 1H, Pyr 4-H), 8.42 (d, J = 7.7 Hz, 1H, Pyr 3-H), 8.61 (s 1H, 3-H), 8.69–8.75 (m, 2H, 7-H, Pyr 6-H), 9.32 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.6 (C-7), 109.3 (d, 2JC,F = 26.3 Hz, Ph C-2), 115.8 (d, 2JC,F = 21.2 Hz, Ph C-4), 116.6 (d, 4JC,F = 3.2 Hz, Ph C-6), 120.6 (C-3a), 121.4 (Pyr C-3), 121.9 (C-3), 123.2 (Pyr C-5), 131.1 (d, 3JC,F = 8.9 Hz, Ph C-5), 136.9 (Pyr C-4), 141.3 (d, 3JC,F = 10.0 Hz, Ph C-1), 146.9 (C-4), 149.5 (Pyr C-6), 150.3 (C-6), 151.7 (C-7a), 156.6 (Pyr C-2), 163.2 (d, 1JC,F = 248.5 Hz, Ph C-3). 15N NMR (71 MHz, CDCl3): δ −147.6 (N-2), −96.9 (N-1), −86.8 (N-5), −77.0 (Pyr N-1). 19F NMR (376 MHz, CDCl3): δ −109.7. MS m/z (%): 291 ([M + H]+, 100). HRMS (ESI) for C17H12N4F ([M + H]+): calcd 291.1041, found 291.1043.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.15–7.19 (m, 1H, Ph 4-H), 7.29–7.31 (m, 1H, Pyr 5-H), 7.51–7.56 (m, 1H, Ph 5-H), 7.72–7.76 (m, 2H, Ph 2,6-H), 7.80–7.84 (m, 1H, Pyr 4-H), 8.42 (d, J = 7.7 Hz, 1H, Pyr 3-H), 8.61 (s 1H, 3-H), 8.69–8.75 (m, 2H, 7-H, Pyr 6-H), 9.32 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.6 (C-7), 109.3 (d, 2JC,F = 26.3 Hz, Ph C-2), 115.8 (d, 2JC,F = 21.2 Hz, Ph C-4), 116.6 (d, 4JC,F = 3.2 Hz, Ph C-6), 120.6 (C-3a), 121.4 (Pyr C-3), 121.9 (C-3), 123.2 (Pyr C-5), 131.1 (d, 3JC,F = 8.9 Hz, Ph C-5), 136.9 (Pyr C-4), 141.3 (d, 3JC,F = 10.0 Hz, Ph C-1), 146.9 (C-4), 149.5 (Pyr C-6), 150.3 (C-6), 151.7 (C-7a), 156.6 (Pyr C-2), 163.2 (d, 1JC,F = 248.5 Hz, Ph C-3). 15N NMR (71 MHz, CDCl3): δ −147.6 (N-2), −96.9 (N-1), −86.8 (N-5), −77.0 (Pyr N-1). 19F NMR (376 MHz, CDCl3): δ −109.7. MS m/z (%): 291 ([M + H]+, 100). HRMS (ESI) for C17H12N4F ([M + H]+): calcd 291.1041, found 291.1043.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–N, C–F), 930, 876, 831, 806, 793, 673, 434, 408 (CH
C, C–N, C–F), 930, 876, 831, 806, 793, 673, 434, 408 (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.27–7.37 (m, 3H, Pyr 5-H, Ph 3,5-H), 7.41–7.47 (m, 1H, Ph 4-H), 7.81–7.83 (m, 1H, Pyr 4-H), 8.13–8.16 (m, 1H, Ph 6-H), 8.42–8.44 (m, 1H, Pyr 3-H), 8.70–8.75 (m, 3H, 3,7-H, Pyr 6-H), 9.33 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.6 (C-7), 117.2 (d, 2JC,F = 20.4 Hz, Ph C-3), 120.6 (C-3a), 121.5 (Pyr C-3), 123.3 (Pyr C-5), 125.4 (d, 4JC,F = 3.8 Hz, Ph C-5), 126.3 (C-3), 126.33 (d, 3JC,F = 11.0 Hz, Ph C-6), 128.6 (d, 2JC,F = 12.3 Hz, Ph C-1), 130.1 (d, 3JC,F = 8.0 Hz, Ph C-4), 137.0 (Pyr C-4), 147.2 (C-4), 149.5 (Pyr C-6), 150.3 (C-6), 151.0 (C-7a), 154.2 (d, 1JC,F = 251.3 Hz, Ph C-2), 156.7 (Pyr C-2). 15N NMR (71 MHz, CDCl3): δ −157.4 (N-2), −95.3 (N-1), −87.8 (N-5), −77.4 (Pyr N-1). 19F NMR (376 MHz, CDCl3): δ −124.1. MS m/z (%): 291 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1135.
CH of benzene). 1H NMR (700 MHz, CDCl3): δ 7.27–7.37 (m, 3H, Pyr 5-H, Ph 3,5-H), 7.41–7.47 (m, 1H, Ph 4-H), 7.81–7.83 (m, 1H, Pyr 4-H), 8.13–8.16 (m, 1H, Ph 6-H), 8.42–8.44 (m, 1H, Pyr 3-H), 8.70–8.75 (m, 3H, 3,7-H, Pyr 6-H), 9.33 (s, 1H, 4-H). 13C NMR (176 MHz, CDCl3): δ 108.6 (C-7), 117.2 (d, 2JC,F = 20.4 Hz, Ph C-3), 120.6 (C-3a), 121.5 (Pyr C-3), 123.3 (Pyr C-5), 125.4 (d, 4JC,F = 3.8 Hz, Ph C-5), 126.3 (C-3), 126.33 (d, 3JC,F = 11.0 Hz, Ph C-6), 128.6 (d, 2JC,F = 12.3 Hz, Ph C-1), 130.1 (d, 3JC,F = 8.0 Hz, Ph C-4), 137.0 (Pyr C-4), 147.2 (C-4), 149.5 (Pyr C-6), 150.3 (C-6), 151.0 (C-7a), 154.2 (d, 1JC,F = 251.3 Hz, Ph C-2), 156.7 (Pyr C-2). 15N NMR (71 MHz, CDCl3): δ −157.4 (N-2), −95.3 (N-1), −87.8 (N-5), −77.4 (Pyr N-1). 19F NMR (376 MHz, CDCl3): δ −124.1. MS m/z (%): 291 ([M + H]+, 100). HRMS (ESI) for C17H12N4 ([M + H]+): calcd 273.1135, found 273.1135.Biology
For the viability assays, cells were treated with the tested compounds for 72 h. After treatments, resazurin (Sigma-Aldrich) solution was added for 4 h, and fluorescence of resorufin, corresponding to live cells, was measured at 544 nm/590 nm (excitation/emission) using a Fluoroskan Ascent microplate reader (Labsystems). The GI50 value, the drug concentration lethal to 50% of the cells, was calculated from the dose–response curves that resulted from the assays.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) acetone (1
acetone (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). Then, slides were rehydrated with PBS, blocked in the solution of 0.5% BSA, and incubated with primary antibody against α-tubulin. Subsequently, cells were washed and incubated with secondary antibody conjugated with Alexa Fluor™ 488, and stained with DAPI. Observations were performed using a fluorescence microscope, Olympus IX51.
1). Then, slides were rehydrated with PBS, blocked in the solution of 0.5% BSA, and incubated with primary antibody against α-tubulin. Subsequently, cells were washed and incubated with secondary antibody conjugated with Alexa Fluor™ 488, and stained with DAPI. Observations were performed using a fluorescence microscope, Olympus IX51.Conclusions
In this study, we successfully enhanced the antimitotic properties of 2H-pyrazolo[4,3-c]pyridines by systematically modifying the 2- and 6-positions of the system. Our efforts culminated in the identification of compound 7f, bearing 4-fluorophenyl and pyridin-2-yl substituents at the 2- and 6-positions, respectively, which demonstrated potent submicromolar cytotoxic activity against a panel of cancer cell lines. The biological evaluation revealed that compound 7f affects the integrity of microtubules, leading to the induction of mitotic defects, disruption of proper cytokinesis, and endoreduplication, resulting in cell death. The pronounced activity of 7f underscores the effectiveness of our chemical modifications and validates the approach of using bioisosteric replacements to optimize biological activity. These findings pave the way for further exploration of 2H-pyrazolo[4,3-c]pyridines as potential anti-cancer agents, with compound 7f serving as a possible lead for future drug development.Author contributions
Conceptualization, A. Š., A. Ž., V. K., and E. A.; investigation, V. A., E. Ř., A. Š.-N.; V. V., V. M., S. B., and A. B.; formal analysis, V. A., E. Ř., and A. Š.-N.; data curation, V. K., and E. A.; funding acquisition, A. Š., and V. K.; methodology, V. K., and E. A.; resources, A. Š., V. K., and E. A.; supervision, V. K., and E. A.; writing—original draft, E. Ř., A. B., S. B., A. Ž., and E. A.; writing—review and editing, A. Š., and V. K. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: 1H, 13C, 1H–15N HMBC, 19F NMR, HRMS data, BrdU incorporation-based cellular proliferation analysis and raw western blot data. See DOI: https://doi.org/10.1039/d5ra09208f.CCDC 2524805 contains the supplementary crystallographic data for this paper.66
Acknowledgements
This work was supported by the Research Council of Lithuania (Project No. S-MIP-23-51), Palacký University Olomouc (Project No. IGA_PrF_2025_011), and by the European Union - Next Generation EU (The project National Institute for Cancer Research, Programme EXCELES, Project No. LX22NPO5102).References
- M. Faisal, A. Saeed, S. Hussain, P. Dar and F. A. Larik, J. Chem. Sci., 2019, 131, 1–30 Search PubMed.
- M. F. Khan, M. M. Alam, G. Verma, W. Akhtar, M. Akhter and M. Shaquiquzzaman, Eur. J. Med. Chem., 2016, 120, 170–201 Search PubMed.
- G. Li, Y. Cheng, C. Han, C. Song, N. Huang and Y. Du, RSC Med. Chem., 2022, 13, 1300–1321 Search PubMed.
- V. Kumar, K. Kaur, G. K. Gupta and A. K. Sharma, Eur. J. Med. Chem., 2013, 69, 735–753 CrossRef CAS PubMed.
- A. Ansari, A. Ali, M. Asif and Shamsuzzaman, New J. Chem., 2017, 41, 16–41 RSC.
- M. A. Alam, Future Med. Chem., 2023, 15, 2011–2023 CrossRef CAS PubMed.
- I. Goldstein, T. F. Lue, H. Padma-Nathan, R. C. Rosen, W. D. Steers and P. A. Wicker, N. Engl. J. Med., 1998, 338, 1397–1404 CrossRef CAS PubMed.
- N. Galiè, H. A. Ghofrani, A. Torbicki, R. J. Barst, L. J. Rubin, D. Badesch, T. Fleming, T. Parpia, G. Burgess, A. Branzi, F. Grimminger, M. Kurzyna and G. Simonneau, N. Engl. J. Med., 2005, 353, 2148–2157 CrossRef PubMed.
- N. M. Davies, A. J. McLachlan, R. O. Day and K. M. Williams, Clin. Pharmacokinet., 2000, 38, 225–242 CrossRef CAS PubMed.
- J. Panés, M. Mollà, M. Casadevall, A. Salas, M. Sans, C. Conill, D. C. Anderson, J. Roselló-Catafau, D. N. Granger and J. M. Piqué, Aliment. Pharmacol. Ther., 2000, 14, 841–850 CrossRef PubMed.
- T. K. Shahin Muhammed, D. Amitkumar, C. Sarath Chandran, B. Revanasiddappa, K. Sreeraj and K. Shijith, J. Mol. Struct., 2022, 1255, 132415 CrossRef.
- Y. Zhang, C. Wu, N. Zhang, R. Fan, Y. Ye and J. Xu, Int. J. Mol. Sci., 2023, 24, 12724 CrossRef CAS PubMed.
- H. Kumar, D. Saini, S. Jain and N. Jain, Eur. J. Med. Chem., 2013, 70, 248–258 CrossRef CAS PubMed.
- Z. Zheng, D. Ren, Y. Ko and H. Liu, J. Am. Chem. Soc., 2025, 147, 11425–11431 CrossRef CAS PubMed.
- N. E. Santos, A. R. F. Carreira, V. L. M. Silva and S. S. Braga, Molecules, 2020, 25, 1364 CrossRef CAS PubMed.
- P. A. Salem, G. P. Bodey, M. A. Burgess, W. K. Murphy and E. J. Freireich, Cancer, 1977, 40, 2806–2809 CrossRef CAS PubMed.
- A. Markham, Drugs, 2020, 80, 1119–1124 CrossRef CAS PubMed.
- C. Zhou, B. Solomon, H. H. Loong, K. Park, M. Pérol, E. Arriola, S. Novello, B. Han, J. Zhou, A. Ardizzoni, M. P. Mak, F. C. Santini, Y. Y. Elamin, A. Drilon, J. Wolf, N. Payakachat, M. K. Uh, D. Rajakumar, H. Han, T. Puri, V. Soldatenkova, A. B. Lin, B. K. Lin and K. Goto, N. Engl. J. Med., 2023, 389, 1839–1850 CrossRef CAS PubMed.
- A. T. Shaw, B. Y. Yeap, B. J. Solomon, G. J. Riely, J. Gainor, J. A. Engelman, G. I. Shapiro, D. B. Costa, S.-H. I. Ou, M. Butaney, R. Salgia, R. G. Maki, M. Varella-Garcia, R. C. Doebele, Y.-J. Bang, K. Kulig, P. Selaru, Y. Tang, K. D. Wilner, E. L. Kwak, J. W. Clark, A. J. Iafrate and D. R. Camidge, Lancet Oncol., 2011, 12, 1004–1012 CrossRef CAS PubMed.
- S. Dhillon, Drugs, 2020, 80, 433–439 CrossRef CAS PubMed.
- A. L. Mederle, L. G. Stana, A. C. Ilie, C. Borza, C. G. Streian, D. Nistor, T. Cerbulescu, B. Belovan and A. Lascu, Biomedicines, 2024, 12, 2820 CrossRef CAS PubMed.
- K. Nath, L. Guo, B. Nancolas, D. S. Nelson, A. A. Shestov, S.-C. Lee, J. Roman, R. Zhou, D. B. Leeper, A. P. Halestrap, I. A. Blair and J. D. Glickson, Biochim. Biophys. Acta, Rev. Cancer, 2016, 1866, 151–162 CrossRef CAS PubMed.
- Y. Huang, G. Sun, X. Sun, F. Li, L. Zhao, R. Zhong and Y. Peng, Cancers, 2020, 12, 3332 CrossRef CAS PubMed.
- G. Cheng, Q. Zhang, J. Pan, Y. Lee, O. Ouari, M. Hardy, M. Zielonka, C. R. Myers, J. Zielonka, K. Weh, A. C. Chang, G. Chen, L. Kresty, B. Kalyanaraman and M. You, Nat. Commun., 2019, 10, 2205 CrossRef PubMed.
- K. Fizazi, N. Shore, T. L. Tammela, A. Ulys, E. Vjaters, S. Polyakov, M. Jievaltas, M. Luz, B. Alekseev, I. Kuss, C. Kappeler, A. Snapir, T. Sarapohja and M. R. Smith, N. Engl. J. Med., 2019, 380, 1235–1246 CrossRef CAS PubMed.
- L. J. Scott, Drugs, 2017, 77, 1029–1034 CrossRef PubMed.
- D. Becerra and J.-C. Castillo, RSC Adv., 2025, 15, 7018–7038 RSC.
- Y. Zhang, C. Wu, N. Zhang, R. Fan, Y. Ye and J. Xu, Int. J. Mol. Sci., 2023, 24, 12724 CrossRef CAS PubMed.
- D. Atukuri, Chem. Biol. Drug Des., 2022, 100, 376–388 CrossRef CAS PubMed.
- A. Donaire-Arias, A. M. Montagut, R. Puig de la Bellacasa, R. Estrada-Tejedor, J. Teixidó and J. I. Borrell, Molecules, 2022, 27, 2237 CrossRef CAS PubMed.
- R. F. Barghash, W. M. Eldehna, M. Kovalová, V. Vojáčková, V. Kryštof and H. A. Abdel-Aziz, Eur. J. Med. Chem., 2022, 227, 113952 CrossRef CAS PubMed.
- A. Papastathopoulos, N. Lougiakis, I. K. Kostakis, P. Marakos, N. Pouli, H. Pratsinis and D. Kletsas, Eur. J. Med. Chem., 2021, 218, 113387 CrossRef CAS PubMed.
- V. Giannouli, N. Lougiakis, I. K. Kostakis, N. Pouli, P. Marakos, A.-L. Skaltsounis, S. Nam, R. Jove, D. Horne, R. Tenta, H. Pratsinis and D. Kletsas, Bioorg. Med. Chem. Lett., 2016, 26, 5229–5233 CrossRef CAS PubMed.
- M. Sklepari, N. Lougiakis, A. Papastathopoulos, N. Pouli, P. Marakos, V. Myrianthopoulos, T. Robert, S. Bach, E. Mikros and S. Ruchaud, Chem. Pharm. Bull., 2017, 65, 66–81 CrossRef CAS PubMed.
- Y. Ma, G. Sun, D. Chen, X. Peng, Y.-L. Chen, Y. Su, Y. Ji, J. Liang, X. Wang, L. Chen, J. Ding, B. Xiong, J. Ai, M. Geng and J. Shen, J. Med. Chem., 2015, 58, 2513–2529 CrossRef CAS PubMed.
- X. Li, T. Yang, M. Hu, Y. Yang, M. Tang, D. Deng, K. Liu, S. Fu, Y. Tan, H. Wang, Y. Chen, C. Zhang, Y. Guo, B. Peng, W. Si, Z. Yang and L. Chen, Bioorg. Chem., 2022, 121, 105669 CrossRef CAS PubMed.
- L. A. Smyth, T. P. Matthews and I. Collins, Bioorg. Med. Chem., 2011, 19, 3569–3578 CrossRef CAS PubMed.
- S. L. Li, Y. Zhou, W. Q. Lu, Y. Zhong, W. L. Song, K. D. Liu, J. Huang, Z. J. Zhao, Y. F. Xu, X. F. Liu and H. L. Li, J. Chem. Inf. Model., 2011, 51, 2939–2947 CrossRef CAS PubMed.
- B. Razmienė, E. Řezníčková, V. Dambrauskienė, R. Ostruszka, M. Kubala, A. Žukauskaitė, V. Kryštof, A. Šačkus and E. Arbačiauskienė, Molecules, 2021, 26, 6747 CrossRef PubMed.
- B. Razmienė, V. Vojáčková, E. Řezníčková, L. Malina, V. Dambrauskienė, M. Kubala, R. Bajgar, H. Kolářová, A. Žukauskaitė, E. Arbačiauskienė, A. Šačkus and V. Kryštof, Bioorg. Chem., 2022, 119, 105570 CrossRef PubMed.
- V. Milišiūnaitė, E. Arbačiauskienė, E. Řezníčková, R. Jorda, V. Malínková, A. Žukauskaitė, W. Holzer, A. Šačkus and V. Kryštof, Eur. J. Med. Chem., 2018, 150, 908–919 CrossRef PubMed.
- D. Sahu, P. S. R. Sreekanth, P. K. Behera, M. K. Pradhan, A. Patnaik, S. Salunkhe and R. Cep, Eur. J. Med. Chem. Rep., 2024, 12, 100210 CAS.
- E. P. Gillis, K. J. Eastman, M. D. Hill, D. J. Donnelly and N. A. Meanwell, J. Med. Chem., 2015, 58, 8315–8359 CrossRef CAS PubMed.
- D. F. O'Brien and J. W. Gates, J. Org. Chem., 1966, 31, 1538–1542 CrossRef.
- U. J. Vogelbacher, M. Keil, R. Klintz, J. Wahl, H. Wingert, H. Koenig, M. Rack, R. Goetz and J. H. Teles, 1996.
- Y. Yoshimoto, S. Arimori and Y. Matsuzaki, WO2013162072A1, 2013.
- A. Bieliauskas, S. Krikštolaitytė, W. Holzer and A. Šačkus, Arkivoc, 2018, 2018, 296–307 Search PubMed.
- V. Milišiūnaitė, E. Arbačiauskienė, A. Bieliauskas, G. Vilkauskaitė, A. Šačkus and W. Holzer, Tetrahedron, 2015, 71, 3385–3395 CrossRef.
- E. Arbačiauskienė, V. Martynaitis, S. Krikštolaitytė, W. Holzer and A. Šačkus, Arkivoc, 2011, 1–21 Search PubMed.
- G. Vilkauskaitė, A. Šačkus and W. Holzer, Eur. J. Org. Chem., 2011, 2011, 5123–5133 CrossRef.
- I. Alkorta, J. Elguero and C. Roussel, Comput. Theor. Chem., 2011, 966, 334–339 CrossRef CAS.
- J. Kalenik and Z. Pawełka, J. Mol. Liq., 2005, 121, 63–68 CrossRef CAS.
- C. Blanchet-Boiteux, P. Friant-Michel, A. Marsura, J.-B. Regnouf-de-Vains and M. F. Ruiz-López, J. Mol. Struct.:THEOCHEM, 2007, 811, 169–174 CrossRef CAS.
- A. V. Afonin, Russ. J. Org. Chem., 2012, 48, 682–685 CrossRef CAS.
- G. Desiraju and T. Steiner, The Weak Hydrogen Bond, Oxford University Press, 2001 Search PubMed.
- A. Crncec and H. Hochegger, FEBS Lett., 2019, 593, 2868–2888 CrossRef CAS PubMed.
- Y.-H. Ling, C. Tornos and R. Perez-Soler, J. Biol. Chem., 1998, 273, 18984–18991 CrossRef CAS PubMed.
- N. Hura, A. Naaz, S. S. Prassanawar, S. K. Guchhait and D. Panda, ACS Omega, 2018, 3, 1955–1969 CrossRef CAS PubMed.
- G. Li, Y. Wang, L. Li, Y. Ren, X. Deng, J. Liu, W. Wang, M. Luo, S. Liu and J. Chen, Eur. J. Med. Chem., 2020, 202, 112519 CrossRef CAS PubMed.
- E. A. Borrego, C. D. Guerena, A. Y. Schiaffino Bustamante, D. A. Gutierrez, C. A. Valenzuela, A. P. Betancourt, A. Varela-Ramirez and R. J. Aguilera, Cells, 2024, 13, 1225 CrossRef CAS PubMed.
- G. Wang, W. Liu, Z. Peng, Y. Huang, Z. Gong and Y. Li, Bioorg. Chem., 2020, 103, 104141 CrossRef CAS PubMed.
- L. M. Harwood and C. J. Moody, Experimental Organic Chemistry: Principles and Practice, Blackwell Scientific Publications, 1989 Search PubMed.
- M. C. Burla, R. Caliandro, M. Camalli, B. Carrozzini, G. L. Cascarano, L. De Caro, C. Giacovazzo, G. Polidori, D. Siliqi and R. Spagna, J. Appl. Crystallogr., 2007, 40, 609–613 CrossRef CAS.
- G. M. Sheldrick, Acta Crystallogr., Sect. C:Struct. Chem., 2015, 71, 3–8 Search PubMed.
- T. Mueller, M. G. Hoffmann, E. Buscato Arsequell, H. Jakobi, D. Schmutzler, C. H. Rosinger, A. B. Machettira and E. Asmus, US20230126893A1, 2023.
- CCDC 2524805: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qr8b3.
Footnote |
| † Vaida Aleksienė and Eva Řezníčková contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |