Open Access Article
Open Access ArticlePyrazole/pyrimidine derivatives endowed with azobenzenes as dual EGFRT790M and VEGFR-2 inhibitors: anticancer, docking, synthesis, design and ADMET assessments
Kurls E. Anwer a,
Mohamed A. Abdelgawad*b,
Felemban Athary Abdulhaleem M.c,
Nour E. A. Abd El-Sattar
a,
Mohamed A. Abdelgawad*b,
Felemban Athary Abdulhaleem M.c,
Nour E. A. Abd El-Sattar ad,
Ahmed El-morsyef,
Tamer Nasr
ad,
Ahmed El-morsyef,
Tamer Nasr gh,
Mohammed Elmowafyi,
Khaled El-Adl
gh,
Mohammed Elmowafyi,
Khaled El-Adl *jk and
Naglaa M. Ahmed
*jk and
Naglaa M. Ahmed l
l
aDepartment of Chemistry, Faculty of Science, Ain Shams University, Abbassia, Cairo, Egypt
bDepartment of Pharmaceutical Chemistry, College of Pharmacy, Jouf University, Sakaka, 72388, Saudi Arabia. E-mail: mhmdgwd@ju.edu.sa
cDepartment of Biology, College of Sciences, Umm Al-Qura University, Makkah 21955, Saudi Arabia
dBasic & Medical Sciences Department, Faculty of Dentistry, Alryada University for Science & Technology, Egypt
ePharmaceutical Organic Chemistry Department, Faculty of Pharmacy (Boys), Al-Azhar University, Nasr City, Cairo, 11884, Egypt
fPharmaceutical Chemistry Department, College of Pharmacy, The Islamic University, Najaf, Iraq
gDepartment of Pharmaceutical Chemistry, Faculty of Pharmacy, Capital University (Formerly Helwan University), Ain-Helwan, Cairo 11795, Egypt
hMedicinal Chemistry Department, Faculty of Pharmacy, Egypt-Japan University of Science and Technology (E-JUST), Alexandria, 21934, Egypt
iDepartment of Pharmaceutics, College of Pharmacy, Jouf University, Sakaka 72388, Saudi Arabia
jPharmaceutical Chemistry Department, Faculty of Pharmacy, Heliopolis University for Sustainable Development, Cairo, Egypt. E-mail: khaled.eladl@hu.edu.eg; eladlkhaled74@yahoo.com
kPharmaceutical Medicinal Chemistry and Drug Design Department, Faculty of Pharmacy (Boys), Al-Azhar University, Nasr City, Cairo, 11884, Egypt. E-mail: eladlkhaled74@azhar.edu.eg
lPharmaceutical Organic Chemistry Department, Faculty of Pharmacy, Capital University (Formerly Helwan University), Ain-Helwan, Cairo 11795, Egypt
First published on 10th March 2026
Abstract
New pyrazole/pyrimidine derivatives endowed with azobenzenes were synthesized using microwave and traditional methods. Our compounds were assessed for cytotoxicity against HepG2, MCF-7, HCT-116 and A549 cell lines as dual inhibitors of EGFRT790M and VEGFR-2. A docking study was carried out to show the binding affinities and orientations of our derivatives in the active sites of VEGFR-2 and EGFRT790M. The data of the docking study were highly correlated with the biological data. The HCT116 and A549 cell lines were extremely affected by our derivatives. Derivative 12 showed the greatest activity against A549, HepG2, MCF-7 and HCT116 cells, with IC50 = 5.12, 6.77, 5.85 and 5.25 µM, respectively. It showed higher cytotoxicity than erlotinib (IC50 = 5.49, 7.73, 8.20 and 13.91 µM, respectively) and lower cytotoxicity than sorafenib (IC50 = 4.04, 4.00, 5.58 and 5.05 µM, respectively) against the tested cell lines. The cytotoxicity of the highly active derivatives 5, 6, 8, 9, 10 and 12 against the MCF-10 healthy cell lines was evaluated. The assessed derivatives showed low cytotoxicity against MCF-10 cells, with IC50 = 50.90–55.50 µM. Additionally, all derivatives were assessed as dual VEGFR-2 and EGFRT790M inhibitors. Compounds 12, 8 and 10 displayed very good inhibitions toward VEGFR-2, with IC50 = 0.90, 0.95 and 1.00 µM, respectively. Similarly, structures 12, 8, 10, 5 and 9 showed strong EGFRT790M inhibitions, with IC50 = 0.25, 0.30, 0.33, 0.35 and 0.40 µM, respectively. In addition, in silico ADMET predictions were calculated for the highly active derivatives 8, 10 and 12 and correlated to Lipinski's rule of five using erlotinib and sorafenib as standard ligands. The results presented our derivatives as promising candidates for advanced manipulations to get more potent anticancer agents with advanced VEGFR-2 and EGFRT790M inhibitions.
1. Introduction
Cancer, one of the most serious diseases to threaten human life, is characterized as a rebound system of uncontrolled cellular growth.1 Angiogenesis is essential for the survival and spread of tumor cells, which is a primary cause of cancer-related deaths. As a result, the importance of angiogenesis inhibition in cancer treatment is growing.2 Signal transduction leads to endothelial cell proliferation, migration, survival, and new vessel formation involved in angiogenesis.3 Through signal transduction, receptor tyrosine kinases control intra- and intercellular signaling. The regulation of vital cellular functions, such as growth, differentiation, survival, metabolism, and proliferation, is greatly influenced by these proteins.4EGFR and VEGFR-2, as tyrosine kinases, are associated with several diseases and diverse cancer growth groups. They are closely related and share common signaling pathways, which are segmented sequentially. The interrelationship between VEGFR-2 and EGFR has been well established: VEGFR-2 signaling pathway inhibition enhances the anticancer activities of EGFR inhibitors, while the activation of VEGFR-2 is independent of EGFR signaling, which is blocked by EGFR inhibitors. Therefore, concurrent blocking of VEGFR-2 and EGFR signaling pathways appears to be a promising strategy in cancer treatment.5,6
VEGFR-2 inhibitors can be categorized into three types. Type I inhibitors occupy the ATP-binding region, forming a hydrogen bond with the hinge region amino acid Cys919. Type II inhibitors occupy the ATP binding site and extend over the gate area into the adjacent allosteric hydrophobic back pocket. Type III inhibitors block the receptor through hydrophobic interactions, and they accommodate the allosteric hydrophobic back pocket of VEGFR-2. Type II inhibitors are preferred over type I inhibitors due to their high affinity and selectivity. Moreover, type II inhibitors exhibit an increased drug-target residence time, resulting in prolonged TK suppression.7 Therefore, diverse strategies have been employed to develop novel type II VEGFR-2 inhibitors (Fig. 1A).
Moreover, the ATP-binding pocket of EGFR-TK consists of five main regions. (a) Adenine binding pocket contains key amino acids, which can form hydrogen bonds with the adenine ring; (b) sugar region (hydrophilic ribose pocket); (c) hydrophobic region I plays an important role in inhibitor selectivity; (d) hydrophobic region II may be exploited for inhibitor specificity; (e) phosphate binding region can be used for improving inhibitor pharmacokinetics (Fig. 1B).8
N-heterocyclic compounds are essential building blocks of RNA and DNA and exert a significant role in the biochemical processes of living organisms.9,10 They are recognizable scaffolds in drug discovery due to their biostability, structural variability and ability to form H bonds with biomolecules. Approximately 60% of FDA-approved small-molecule drugs include N-heterocyclic rings, emphasizing their importance in medicinal chemistry. Among them, pyrazole derivatives appear to provide specific value due to their extensive therapeutic potential, particularly in anti-cancer agent development.9,11
Pyrazole is an attractive choice as a substrate for the synthesis of several broad-spectrum biologically active pharmaceutical drugs. Globally, there are more than 50 pyrazole-based synthetic medicines on the market. These pyrazole-based marketed drugs target a varied range of clinical disorders, including rheumatoid arthritis, cystic fibrosis, sickle cell disease, non-small cell lung cancer (NSCLC), and hereditary angioedema.12 Furthermore, a variety of pyrazole-based compounds have been recognized in cancer treatments as dual inhibitors of EGFR and VEGFR-2.13,14
Moreover, numerous drugs featuring pyrimidines (N-heterocycles) as the principal structural scaffold were recently FDA-approved as potential medications for a variety of cancer-related disorders. Most of these compounds inhibit VEGFR-2 and EGFR, which are vital receptor tyrosine kinases associated with the tumor microenvironment, to prevent tumor growth.15
Azobenzenes, which contain the N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond, have been reported as a multi-targeted scaffold in medicinal chemistry.16 It has a variety of uses, including as antimicrobial, anti-inflammatory, antiviral, antidiabetic, anti-tuberculosis and anti-tumor agents.17 Several in vitro and in vivo studies have confirmed that azo compounds could potentially act as anticancer agents. Giampietro et al. reported the synthesis, biological evaluation, and docking studies of a series of phenyldiazenyl sulfonamide compounds as aromatase inhibitors.18 Moreover, Bustos et al. synthesized a series of 12 phenyldiazenyl pyrazoles and tested them against a wide library of cancer cell lines, including leukemia, colon, and brain cancer, showing that the compounds have anticancer activity.19 All compounds showed a certain degree of growth inhibition against different cell lines. Highly selective growth inhibition activity was observed against the lung cancer, kidney cancer and leukemia cell lines. With the aim of identifying new anti-breast cancer compounds, Gomha et al. synthesized a series of thiazole-benzofuran phenyldiazenyl compounds. These molecules were evaluated for their anticancer activity against the human breast carcinoma (MCF-7) cell lines, compared with the doxorubicin drug.20 In 2024, Tang et al. synthesized different azobenzene coumarin derivatives, where the coumarin moiety was linked to substituted phenyl and pyridine groups via amide bonds. These derivatives showed different degrees of anticancer activities against HeLa, A549, MCF-7, and HepG-2 cell lines.21 Additionally, some azobenzene derivatives have been reported to show anti-pancreatic cancer activity.22
N bond, have been reported as a multi-targeted scaffold in medicinal chemistry.16 It has a variety of uses, including as antimicrobial, anti-inflammatory, antiviral, antidiabetic, anti-tuberculosis and anti-tumor agents.17 Several in vitro and in vivo studies have confirmed that azo compounds could potentially act as anticancer agents. Giampietro et al. reported the synthesis, biological evaluation, and docking studies of a series of phenyldiazenyl sulfonamide compounds as aromatase inhibitors.18 Moreover, Bustos et al. synthesized a series of 12 phenyldiazenyl pyrazoles and tested them against a wide library of cancer cell lines, including leukemia, colon, and brain cancer, showing that the compounds have anticancer activity.19 All compounds showed a certain degree of growth inhibition against different cell lines. Highly selective growth inhibition activity was observed against the lung cancer, kidney cancer and leukemia cell lines. With the aim of identifying new anti-breast cancer compounds, Gomha et al. synthesized a series of thiazole-benzofuran phenyldiazenyl compounds. These molecules were evaluated for their anticancer activity against the human breast carcinoma (MCF-7) cell lines, compared with the doxorubicin drug.20 In 2024, Tang et al. synthesized different azobenzene coumarin derivatives, where the coumarin moiety was linked to substituted phenyl and pyridine groups via amide bonds. These derivatives showed different degrees of anticancer activities against HeLa, A549, MCF-7, and HepG-2 cell lines.21 Additionally, some azobenzene derivatives have been reported to show anti-pancreatic cancer activity.22
In 2024, Sandor and colleagues introduced a set of quinazoline–thiazole hybrid compounds with diazene spacers as potential antiproliferative and anti-angiogenic agents. The majority of these compounds exhibited enhanced antiproliferative activity (IC50 = 1.83–4.24 µM) against HepG2 cells compared to sorafenib (IC50 = 6.28 µM). The interaction with the VEGFR-2 kinase domain was evaluated using computational methods, such as molecular docking and molecular dynamics simulations. The compound series demonstrated a notable resemblance to sorafenib in terms of binding orientation within the VEGFR-2 active site, with compound I (Fig. 2) forming the most stable complex with VEGFR-2, in comparison to sorafenib. The highest free energy was observed for I (−71.23 ± 5.29 kcal mol−1), which closely approximated the value for sorafenib (−69.39 ± 3.63 kcal mol−1).23
In 2020, Srour and colleagues created another set of compounds based on a combination of benzimidazole and thiazole with the diazene moiety to act as EGFR inhibitors. Compound II (Fig. 2) exhibited notable efficacy as an EGFR tyrosine kinase inhibitor and achieved anti-breast cancer treatment, with IC50 values of 109.71 ± 3.55 nM and 6.30 ± 0.37 µM, respectively.24
Kinase inhibitors have significant therapeutic potential in oncology, as they specifically target abnormal kinases responsible for the malignancy of cancer cells. By disturbing the signaling pathways that cancer cells rely on to divide and survive, these inhibitors can efficiently prevent tumor growth.25
Certain kinase inhibitors have been designed to target receptor tyrosine kinases (RTKs), which are often mutated or overexpressed in cancers, such as breast, lung and colorectal cancers.26 Among the many classes of kinase inhibitors, pyrazoles27 and pyrimidines28 have received considerable attention due to their potent inhibition of kinases closely associated with cancer and other diseases.
These products bind to the ATP-binding site of kinases, thereby preventing the transfer of phosphate required for the signal transduction pathways of cancer cell survival and proliferation.29 Moreover, various pyrimidines and pyrazoles have been designated as EGFR and VEGFR-2 inhibitors.30 Furthermore, numerous phenyldiazene-containing compounds have been described as anticancer agents that inhibit both VEGFR-2 and EGFRT790M receptors.31
Based on former findings and as an extension of our research program in developing and synthesizing various anticancer derivatives,32–36 new pyrazoles and pyrimidines containing azobenzenes were prepared and assessed for cytotoxicity against HepG2, A549, MCF-7 and HCT-116 cell lines.
The selectivity of the active compounds was also evaluated on normal human MCF-10 cell lines. Additionally, all compounds were assessed for both EGFRT790M and VEGFR-2 inhibition. Docking experiments were also conducted in the EGFRT790M and VEGFR-2 active sites to predict their probable binding modes and orientations.
2. Structure-based and rationale design
New pyrazoles and pyrimidines substituted with azobenzenes were designed and synthesized according to the structural requirements for both VEGFR-2 and EGFRT790M inhibition.The new derivatives were obtained through bioisosteric modification of VEGFR-2 inhibitors (sorafenib and pazopanib) at four different parts.37–39 The hydrophobic substituted phenyl tails of sorafenib and pazopanib that occupy the hydrophobic pocket were replaced by unsubstituted phenyl groups. The hydrogen bond donor (HBD) urea and/or amino spacers were replaced by HBA diazene (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) groups that can bind to Glu885 and/or Asp1046. The central hydrophobic phenoxy and/or N-methylpyrimidine linkers of sorafenib and pazopanib, respectively, were replaced by azobenzene moieties. The substituted hetero-aromatic systems, specifically the N-methylpicolinamide and/or the 2,3-dimethyl-2H-indazol-6-yl moieties of sorafenib and pazopanib, respectively, were replaced by substituted pyrazoles 3–8 and/or pyrimidines 9–12, respectively (Fig. 3).
N) groups that can bind to Glu885 and/or Asp1046. The central hydrophobic phenoxy and/or N-methylpyrimidine linkers of sorafenib and pazopanib, respectively, were replaced by azobenzene moieties. The substituted hetero-aromatic systems, specifically the N-methylpicolinamide and/or the 2,3-dimethyl-2H-indazol-6-yl moieties of sorafenib and pazopanib, respectively, were replaced by substituted pyrazoles 3–8 and/or pyrimidines 9–12, respectively (Fig. 3).
 | ||
| Fig. 3 Pharmacophoric and structural similarities of some designed compounds and VEGFR-2 inhibitors. | ||
In the same manner, our new derivatives were obtained through modifications of the EGFR inhibitors (erlotinib and gefitinib) at four positions.40,41 The hydrophobic bis(2-methoxyethoxy) and 3-morpholinopropoxy tails of erlotinib and gefitinib, respectively, that occupy the hydrophobic region II were replaced by the terminal hydrophobic azobenzene moieties. The HBD NH group was replaced by the NH2 group. The hetero-aromatic quinazoline scaffold that occupies the adenine pocket was replaced by the hetero-aromatic pyrazole and/or pyrimidine rings linked to azobenzenes. The lipophilic 3-ethynylphenyl and 3-chloro-4-fluorophenyl heads of erlotinib and gefitinib, respectively, that occupy hydrophobic region I were replaced by different aliphatic and/or aromatic groups (Fig. 4).
3. Results and discussion
3.1. Chemistry
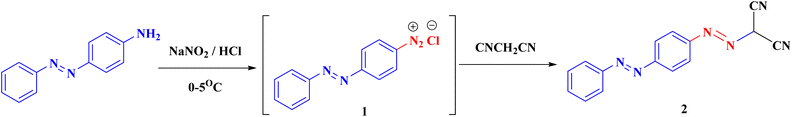
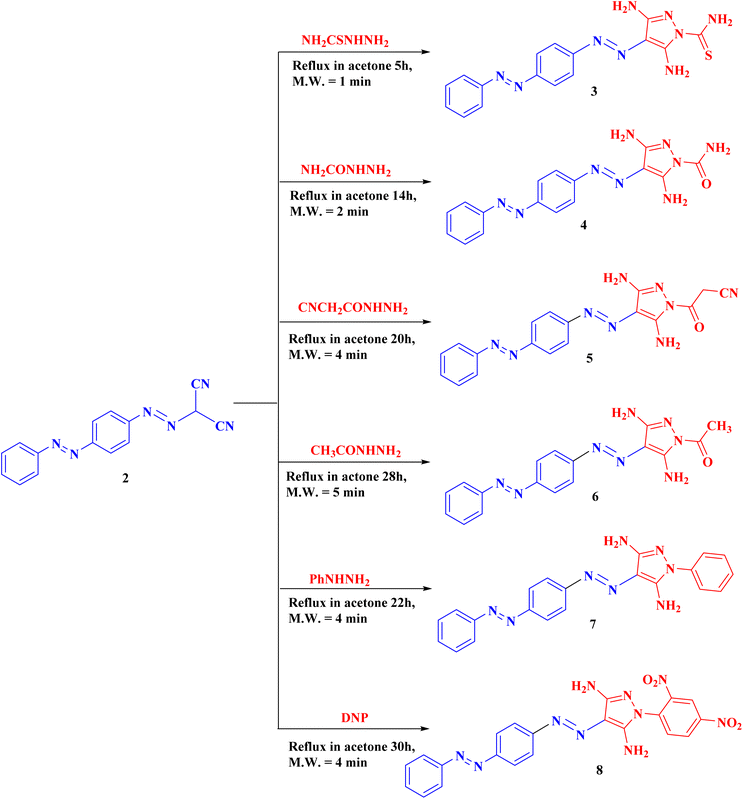
Our derivatives were synthesized following the reactions described in Schemes 1–3. The reaction of 4-(phenyldiazenyl)aniline with NaNO2 in the presence of HCl at 0–5 °C gives the diazonium chloride salt 1, which couples with malononitrile to yield the key compound 2-(4-(phenyldiazenyl)phenyldiazenyl)-malononitrile (2) (Scheme 1) according to the described technique.42–44Derivative 2 was cyclized with the appropriate thiosemicarbazide, semicarbazide, 2-cyanoacetohydrazide, acetohydrazide, phenyl hydrazine, or 2,4-dinitrophenyl hydrazine (DNP) in acetone at reflux temperature for 5–30 h to give the corresponding targeted pyrazole derivatives 3–8 (Scheme 2). The structure of compound 5 produced IR absorption bands at 3443 (NH2), 2194 (CN) and 1657 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). The 1H NMR spectrum of compound 5 showed one singlet signal for CH2 at 2.41. Additionally, two D2O exchangeable singlet signals were observed for the 2NH2 groups at 5.86 and 6.30. Moreover, the 13C-NMR data support the structure.
O). The 1H NMR spectrum of compound 5 showed one singlet signal for CH2 at 2.41. Additionally, two D2O exchangeable singlet signals were observed for the 2NH2 groups at 5.86 and 6.30. Moreover, the 13C-NMR data support the structure.
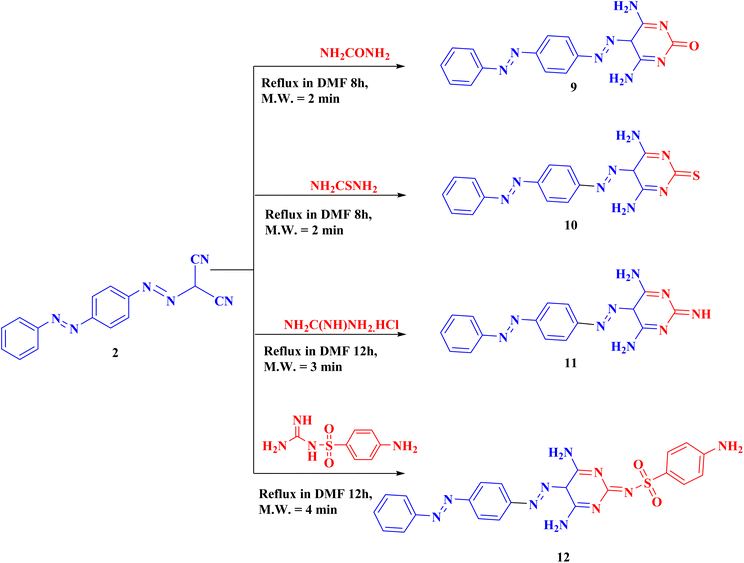
Derivative 2 was used to synthesize the novel pyrimidines (9–12) through cyclization reactions with the suitable urea, thiourea, guanidine and/or sulfaguanidine in DMF using a water bath for 8–12 h to obtain the corresponding products 9–12 (Scheme 3).
3.2. Microwave and conventional techniques assessment
A comparative evaluation between microwave-assisted and conventional methods was conducted using several green chemistry efficiency parameters, including yield economy (YE), atomic economy (AE), reaction mass efficiency (RME), and optimum efficiency (OE). The reaction times and yields obtained for both methods were compared45–49 (Table 1).| Compound no. | Time “min” | Yield (%) | YE | OE | RME | AE | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| M.W.b | Conv.a | M.W. | Conv. | M.W. | Conv. | M.W. | Conv. | M.W. | Conv. | ||
| a Conv.: conventional technique.b M.W.: microwave irradiation technique. | |||||||||||
| 3 | 1 | 300 | 90 | 54 | 90.00 | 0.1800 | 0.7766 | 0.3883 | 77.66 | 38.83 | 100 |
| 4 | 2 | 840 | 92 | 51 | 46.00 | 0.0607 | 0.7996 | 0.4431 | 72.48 | 40.17 | 90.65 |
| 5 | 4 | 1200 | 91 | 53 | 22.75 | 0.0442 | 0.7875 | 0.4587 | 78.75 | 45.87 | 100 |
| 6 | 5 | 1680 | 90 | 52 | 18.00 | 0.0310 | 0.7714 | 0.4457 | 77.14 | 44.57 | 100 |
| 7 | 4 | 1320 | 90 | 52 | 22.50 | 0.0394 | 0.7814 | 0.4515 | 78.14 | 45.15 | 100 |
| 8 | 4 | 1800 | 92 | 53 | 23.00 | 0.0294 | 0.8193 | 0.4720 | 81.93 | 47.20 | 100 |
| 9 | 2 | 480 | 90 | 54 | 45.00 | 0.1125 | 0.7386 | 0.4431 | 73.86 | 44.31 | 100 |
| 10 | 2 | 480 | 92 | 52 | 46.00 | 0.1083 | 0.7612 | 0.4303 | 76.12 | 43.03 | 100 |
| 11 | 3 | 720 | 90 | 51 | 30.00 | 0.0708 | 0.7514 | 0.4258 | 67.81 | 38.42 | 90.24 |
| 12 | 4 | 720 | 91 | 51 | 22.75 | 0.0708 | 0.7916 | 0.4436 | 79.16 | 44.36 | 100 |
The yield economy (YE) was employed to identify the most efficient method for reaction accomplishment and was calculated according to the following equation:
| YE = yield (%)/time (min) |
The reaction mass efficiency (RME) was calculated using the equation:
| RME = (weight of isolated product)/(total weight of reactants) |
In addition, the optimum efficiency (OE) was employed as a comparative parameter between the two methods and was calculated as follows:
| OE = (RME/AE) × 100 |
While atomic economy (AE) represents the theoretical maximum efficiency of a reaction, RME reflects the actual mass efficiency. The AE values for both the microwave and conventional methods are identical, as the same target molecules were synthesized from identical reactants under different reaction conditions, as shown in Table 1.
3.3. Docking studies
The Molsoft program was used to assess the binding affinities and orientations of our derivatives inside the active sites of both VEGFR-2 (PDB ID: 4ASD)50,51 and EGFRT790M (PDB ID: 3W2O).44,52| Compound | ΔG [kcal mol−1] | Compound | ΔG [kcal mol−1] |
|---|---|---|---|
| 3 | −75.00 | 9 | −81.91 |
| 4 | −68.21 | 10 | −90.88 |
| 5 | −83.99 | 11 | −79.79 |
| 6 | −80.36 | 12 | −106.48 |
| 7 | −79.91 | Sorafenib | −99.50 |
| 8 | −91.33 |
Sorafenib presented −99.50 kcal mol−1 and formed five H bonds with Cysteine919 (2.51 Å and 2.10 Å), Glutamate885 (2.75 Å and 1.77 Å) and Aspartate1046 (1.50 Å). The 3-trifluoromethyl-4-chlorophenyl tail inhibited the hydrophobic pocket constructed by Isoleucine892, Isoleucine888, Histidine1026, Glutamate885, Aspartate1046 and Cysteine1045. Moreover, the substituted hetero-aromatic pyridine ring inhibited the pocket formed by Cysteine919, Phenylalanine918, Glutamate917, Valine848, Lysine920, Leucine1035, and Leucine840. Furthermore, the central phenoxy linker inhibited the hydrophobic pocket formed by Leucine1035, Cysteine1045, Valine848, and Lysine868 (Fig. 5).
Compound 12 exhibited −106.48 kcal mol−1 and formed 6H bonds with Cys919 (2.53 Å), Glu917 (1.61 Å and 2.42 Å), Asp1046 (1.97 Å), Asn923 (2.57 Å) and Leu840 (2.49 Å). The distal phenyldiazene tail inhibited the hydrophobic domain formed by Ile1025, Ile888, Asp1046, Hie1026, Glu885, Cys1045 and Ile892. Moreover, the aromatic 4-aminophenyl ring inhibited the pocket formed by Cys919, Lys920, Leu1035, Phe918, Asn923, and Leu840. Furthermore, the central azobenzene linker inhibited the pocket formed by Cys1045, Val848, Lys868, Glu917 and Asp1046 (Fig. 6).
In the same manner, compound 8 showed nearly the same binding orientation as 12, with −91.33 kcal mol−1, and it formed 7 hydrogen bonds with Asp1046 (2.08 Å and 2.34 Å), Cys919 (1.66 Å and 2.38 Å), Gly922 (2.44 Å) and Glu917 (1.72 Å and 2.30 Å) (Fig. 7).
Similarly, derivative 10 showed nearly the same orientation as 12, with −90.88 kcal mol−1, and it formed 4H bonds with Asp1046 (1.69 Å and 2.72 Å), Cys919 (1.97 Å) and Glu917 (1.68 Å) (Fig. 8).
| Compound | ΔG [kcal mol−1] | Compound | ΔG [kcal mol−1] |
|---|---|---|---|
| 3 | −57.55 | 9 | −76.16 |
| 4 | −64.31 | 10 | −79.46 |
| 5 | −77.54 | 11 | −72.40 |
| 6 | −72.04 | 12 | −99.31 |
| 7 | −67.67 | Erlotinib | −82.77 |
| 8 | −80.68 |
Erlotinib displayed −82.77 kcal mol−1 and formed four H-bonds with Thr854 (2.99 Å), Cys797 (2.05 Å), Methionine793 (1.82 Å), and Valine726 (2.97 Å). The 3-ethynylphenyl head inhibited the hydrophobic area I produced by Ile759, Phe723, Gly724, Asp855, Thr854, Val726, Glu762, Leu777, Met790 and Glu791. Similarly, the 2-methoxyethoxy end inhibited the hydrophobic groove II produced by Leu844, Met793, Leu718, Pro794 and Val845 (Fig. 9).
Derivative 12 showed −99.31 kcal mol−1 and formed 9H bonds with Met793 (2.98 Å), Glu762 (1.96 Å and 2.28 Å), Ala743 (2.76 Å and 2.96 Å), Lys745 (1.71 Å), Asp855 (2.26 Å and 2.95 Å) and Thr854 (2.99 Å). The azobenzene side chain inhibited the hydrophobic area II produced by Pro794, Met793, Leu718, Val726, Val845, Glu791, Met790, Ala743 and Leu844. Moreover, the sulfonylaniline head inhibited the hydrophobic area I formed by Gly724, Glu762, Asp855, Phe723, Ile759 and Thr854 (Fig. 10).
Also, derivative 8 exhibited nearly the same orientation as 12 (−80.68 kcal mol−1). It formed 6H bonds with Met793 (2.95 Å), Lys745 (2.51 Å), Glu791 (2.13 Å and 2.38 Å), Asp855 (2.77 Å) and Thr854 (2.29 Å) (Fig. 11).
Derivative 10 displayed nearly the same orientation as 12, with −79.46 kcal mol−1, and it formed 5H bonds with Met793 (2.92 Å and 2.96 Å), Lys745 (2.98 Å), Glu791 (2.64 Å) and Thr854 (2.06 Å) (Fig. 12).
3.4. Biological testing
| Comp. | IC50a (µM) | ||||||
|---|---|---|---|---|---|---|---|
| HCT116 | MCF-7 | HepG2 | A549 | MCF-10 | VEGFR-2 | EGFRT790M | |
| a IC50 values are the mean ± S.D. of three separate experiments.b NT = not tested. | |||||||
| 3 | 15.75 ± 1.5 | 15.25 ± 1.5 | 15.90 ± 1.5 | 17.50 ± 1.5 | NTb | 2.20 ± 0.50 | 1.60 ± 0.15 |
| 4 | 18.30 ± 1.5 | 18.50 ± 1.5 | 19.60 ± 1.5 | 15.20 ± 5.5 | NTb | 2.45 ± 0.15 | 1.45 ± 0.25 |
| 5 | 7.80 ± 0.5 | 7.90 ± 0.5 | 7.93 ± 0.8 | 7.20 ± 0.5 | 55.00 ± 2.9 | 1.30 ± 0.15 | 0.35 ± 0.15 |
| 6 | 6.80 ± 0.5 | 7.55 ± 0.5 | 9.75 ± 1.0 | 7.55 ± 0.5 | 50.90 ± 2.9 | 1.08 ± 0.15 | 0.54 ± 0.15 |
| 7 | 8.25 ± 0.5 | 8.60 ± 0.5 | 10.25 ± 1.5 | 8.80 ± 0.8 | NTb | 1.50 ± 0.15 | 0.62 ± 0.15 |
| 8 | 6.00 ± 0.5 | 7.15 ± 0.5 | 7.35 ± 0.5 | 6.10 ± 0.5 | 53.50 ± 2.9 | 0.95 ± 0.15 | 0.30 ± 0.15 |
| 9 | 7.25 ± 0.5 | 7.38 ± 0.5 | 8.06 ± 0.5 | 6.95 ± 0.5 | 52.95 ± 2.9 | 1.12 ± 0.50 | 0.40 ± 0.15 |
| 10 | 6.50 ± 0.5 | 7.35 ± 0.5 | 11.90 ± 1.1 | 7.50 ± 0.5 | 52.66 ± 2.9 | 1.00 ± 0.15 | 0.33 ± 0.15 |
| 11 | 10.25 ± 1.1 | 10.80 ± 1.0 | 7.70 ± 0.5 | 7.33 ± 0.5 | NTb | 1.70 ± 0.50 | 0.50 ± 0.25 |
| 12 | 5.25 ± 0.5 | 5.85 ± 0.5 | 6.77 ± 1.1 | 5.12 ± 0.5 | 55.50 ± 2.9 | 0.90 ± 0.15 | 0.25 ± 0.15 |
| Sorafenib | 5.05 ± 0.50 | 5.58 ± 0.55 | 4.00 ± 0.33 | 4.04 ± 0.33 | NTb | 0.84 ± 0.04 | NTb |
| Erlotinib | 13.91 ± 1.3 | 8.20 ± 0.34 | 7.73 ± 0.67 | 5.49 ± 0.45 | NTb | NTb | 0.24 ± 0.22 |
Against the HCT116 cell lines, compounds 5, 6, 7, 8, 9 and 10, with IC50 = 6.00–8.25 µM, offered very good cytotoxicity. Compound 11, with IC50 = 10.25 µM, offered good cytotoxicity. Additionally, derivatives 3 and 4, with IC50 = 15.75 and 18.30 µM, respectively, showed moderate cytotoxicity.
Against the MCF-7 cell lines, derivatives 5, 6, 7, 8, 9 and 10, with IC50 = 7.15–8.60 µM, offered very good cytotoxicity. Compound 11, with IC50 = 10.80 µM, offered good cytotoxicity. Additionally, derivatives 3 and 4, with IC50 = 15.25 and 18.50 µM, respectively, showed moderate cytotoxicity.
With regard to the HepG2 cell lines, derivatives 5, 6, 8, 9 and 11, with IC50 = 7.85–9.75 µM, showed very good cytotoxicity. Compounds 7 and 10, with IC50 = 10.25 and 11.90 µM, offered good cytotoxicity. Additionally, derivatives 3 and 4, with IC50 = 15.90 and 19.60 µM, respectively, showed moderate cytotoxicity.
Against the A549 cell lines, compounds 5, 6, 7, 8, 9, 10 and 11, with IC50 = 6.10–8.80 µM, offered very good cytotoxicity. Additionally, derivatives 3 and 4, with IC50 = 17.50 and 15.20 µM, respectively, showed moderate cytotoxicity.
Moreover, the cytotoxicity of the significantly potent derivatives 5, 6, 8, 9, 10 and 12 was evaluated against the MCF-10 healthy cell lines. The estimated structures indicated low toxicity against the MCF-10 cells, with IC50 = 50.90–55.50 µM. Compounds 5, 6, 8, 9, 10 and 12 exhibited 8.09-, 6.53-, 8.92-, 7.30-, 8.10- and 10.57-fold higher toxicity against HCT-116 cells, respectively, than against MCF-10 cells. Equivalently, derivatives 5, 6, 8, 9, 10 and 12 exhibited 7.28-, 6.44-, 7.48-, 7.17-, 7.16- and 9.49-fold higher toxicity against MCF-7 cells, respectively, than against healthy MCF-10 cells. In addition, derivatives 5, 6, 8, 9, 10 and 12 exhibited 6.94-, 5.22-, 7.28-, 6.57-, 6.84- and 8.20-fold higher toxicity against HepG2 cells, respectively, than against MCF-10 cells. Also, derivatives 5, 6, 8, 9, 10 and 12 displayed 7.64-, 6.74-, 8.77-, 7.22-, 7.58- and 10.84-fold higher toxicity against A549 cells, respectively, than against MCF-10 cells.
Statistical comparisons relative to sorafenib and erlotinib were performed using pairwise Welch's t-tests (assuming n = 3). The compounds exhibited a broad range of cytotoxic activities, with IC50 values spanning 5.12–19.60 µM. Compounds 3 and 4 were the least potent derivatives, displaying IC50 values generally exceeding 15 µM across all tested cell lines. In contrast, compounds 7, 9, and 11 demonstrated moderate activities, particularly against HCT-116 and MCF-7 cells, with IC50 values in the range of 7.25–10.80 µM. Notably, compounds 5 and 6 showed enhanced cytotoxicity toward HepG2 cells (IC50 = 7.93 ± 0.8 and 9.75 ± 1.0 µM, respectively), and their activity was statistically comparable to that of erlotinib (p > 0.05), indicating potentially relevant efficacy in hepatocellular carcinoma models. Among the evaluated derivatives, compound 12 emerged as the most potent analogue, exhibiting consistently low IC50 values across all cancer cell lines (5.12–6.77 µM). Statistical analysis revealed that its antiproliferative activities against HCT-116 and MCF-7 cells were statistically indistinguishable from those of sorafenib (p > 0.05), indicating comparable potency to the clinically used multikinase inhibitor. Moreover, compound 12 demonstrated significantly greater potency than erlotinib in these cell lines (p < 0.01), supporting its superior cellular efficacy. Compounds 8 and 10 displayed notable cytotoxic activity, particularly against HCT-116 cells (IC50 = 6.00 ± 0.5 and 6.50 ± 0.5 µM, respectively), where both were significantly more potent than erlotinib (p < 0.01). Their activity was statistically comparable to that of sorafenib in selected assays (p > 0.05), supporting their clinically relevant potency, although their overall antiproliferative effects remained slightly weaker than those of compound 12. These findings position compounds 8 and 10 as promising secondary leads. To gain mechanistic insights, the compounds were evaluated for their inhibitory activity against VEGFR-2 and the resistant EGFRT790M kinase. In the EGFRT790M assay, compound 12 exhibited inhibitory potency statistically comparable to that of erlotinib (p > 0.05), confirming the effective inhibition of the clinically relevant resistance mutation. Compounds 8 and 10 also demonstrated low micromolar inhibitory activities, with potencies statistically comparable to that of erlotinib in this assay, albeit with slightly higher IC50 values. In the VEGFR-2 inhibition assay, compound 12 again showed the strongest activity (IC50 = 0.90 ± 0.15 µM), with no statistically significant difference relative to sorafenib (p > 0.05), indicating comparable anti-angiogenic potential. Compounds 8 and 10 displayed measurable VEGFR-2 inhibition, with IC50 values close to that of sorafenib, and no statistically significant differences, supporting partial dual-target engagement, although their inhibitory profiles were less consistent than that of compound 12. Overall, the combined cellular and biochemical data identify compound 12 as the most advanced lead candidate, characterized by high antiproliferative potency, favorable selectivity toward cancer cells, and dual inhibition of VEGFR-2 and EGFRT790M with activity statistically comparable to those of established reference drugs. Compounds 8 and 10 represent promising secondary scaffolds, exhibiting clinically relevant cytotoxicity and kinase inhibition in selected assays.
All compounds displayed strong-to-low inhibitions, with IC50 = 0.90–2.45 µM (Table 4). Compounds 12, 8, 10, 5 and 9 showed strong EGFRT790M inhibitions, with IC50 = 0.25, 0.30, 0.33, 0.35 and 0.40 µM, respectively. Derivative 11 exhibited good inhibition, with IC50 = 0.50 µM. Also, structures 6 and 7 gave moderate inhibitions, with IC50 = 0.54 and 0.62 µM, respectively. Also, structures 3 and 4 showed lower inhibitions, with IC50 = 1.60 and 1.45 µM, respectively.
The first group of pyrazole derivatives includes compounds 3, 4, 5, 6, 7 and 8. In this group, derivative 8, with the dinitrophenyl moiety, displayed higher cytotoxicity than derivative 6 with the acetyl group, derivative 5 with COCH2CN, derivative 7 with an unsubstituted phenyl group, derivative 3 with CSNH2, and derivative 4 with CONH2 against HCT116 and MCF-7 cells. Conversely, the order of activity against HepG2 cells is 8 > 5 > 6 > 7 > 3 > 4, while that against A549 cells is 8 > 5 > 6 > 7 > 4 > 3 (Fig. 13).
The second group of pyrimidine derivatives includes compounds 9, 10, 11 and 12. In this group, derivative 12, with a sulfanilamide moiety, showed higher anticancer activities than pyrimidine-2(5H)-thione derivative 10, pyrimidine-2(5H)-one derivative 9 and pyrimidine-2(5H)-imine derivative 11 against HCT116 and MCF-7 cells. Conversely, the order of activity is 12 > 11 > 9 > 10 against HepG2 cells, while against A549 cells, the order of activity is 12 > 9 > 11 > 10.
In addition, pyrimidine derivative 12, with sulfanilamide, displayed better dual VEGFR-2 and EGFRT790M inhibitions than pyrazole derivative 8 with dinitrophenyl and pyrimidin-2-thione derivative 10. This could be because the sulfanilamide moiety formed two extra H bonds with Asparagine923 and Leucine840 in the ATP binding domain, while dinitrophenyl formed two extra H bonds with Cysteine919 and Glycine922, but pyrimidin-2-thione derivative 10 formed no extra H bonds.
Derivative 5 exhibited good VEGFR-2 inhibition: 5 > 6 > 9. Conversely, compounds 7 and 11 displayed moderate inhibitions, while compounds 3 and 4 displayed low inhibitions (Fig. 13).
In the same manner, compounds 5 and 9 displayed very good EGFRT790M inhibitions, while derivative 11 exhibited good inhibition. Conversely, compounds 6 and 7 displayed moderate inhibitions, while derivatives 3 and 4 showed low inhibitions (Fig. 13).
3.5. In silico ADMET profile
In silico ADMET predictions were carried out for the most potent compounds 8, 10 and 12 and correlated to Lipinski's rule of five.59 Calculations were achieved using the pkCSM descriptor algorithm.60 For good absorption, our derivatives should adhere to at least three of the following rules: (i) no more than 5H-bond donors, (ii) a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P less than 5, (iii) a molecular weight less than 500, and (iv) no more than 10H-bond acceptors. Sorafenib violated one rule, and derivative 8 violated two rules, but erlotinib did not violate any rules.
P less than 5, (iii) a molecular weight less than 500, and (iv) no more than 10H-bond acceptors. Sorafenib violated one rule, and derivative 8 violated two rules, but erlotinib did not violate any rules.
Derivatives 8 and 10 exhibited excellent human intestinal absorption (97.026 and 89.224, respectively), and derivative 12 exhibited very good absorption (80.283), indicating that they can cross numerous biological membranes61 (Table 5). So through GIT, they might display a meaningfully high bioavailability. Our compounds can reach the CNS (CNS permeability range: −2.043 to −2.379), which may be attributed to their high lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 3.5672–5.6839).
P = 3.5672–5.6839).
| Parameter | 8 | 10 | 12 | Sorafenib | Erlotinib |
|---|---|---|---|---|---|
| Physicochemical properties | |||||
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
5.6839 | 3.5672 | 3.6094 | 5.5497 | 3.4051 |
| Molecular weight | 472.425 | 350.411 | 488.537 | 464.831 | 393.443 |
| Donors | 2 | 2 | 3 | 3 | 1 |
| Acceptors | 12 | 7 | 9 | 4 | 7 |
| Rotatable bonds | 7 | 4 | 6 | 5 | 10 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Absorption | |||||
| Human intest. absorption | 97.026 | 89.224 | 80.283 | 89.043 | 94.58 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Distribution | |||||
| CNS permeability | −2.043 | −2.101 | −2.379 | −2.007 | −3.216 |
| BBB permeability | −1.71 | −0.835 | −1.182 | −1.684 | −0.745 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Metabolism | |||||
| CYP2D6 substrate | No | No | No | No | No |
| CYP3A4 substrate | Yes | Yes | Yes | Yes | Yes |
| Inhibition of CYP1A2 | No | Yes | Yes | Yes | Yes |
| Inhibition of CYP2C19 | Yes | No | Yes | Yes | Yes |
| Inhibition of CYP2C9 | Yes | No | Yes | Yes | Yes |
| Inhibition of CYP2D6 | No | No | Yes | No | No |
| Inhibition of CYP3A4 | Yes | No | No | Yes | Yes |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Excretion | |||||
| Clearance | −0.536 | 0.321 | −0.658 | −0.219 | 0.702 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Toxicity | |||||
| Chronic toxic activity (LOAEL) | 1.061 | 0.884 | 2.172 | 1.198 | 1.37 |
| Acute toxic activity (LD50) | 2.566 | 2.415 | 2.687 | 2.538 | 2.393 |
| Hepatotoxic effect | Yes | No | Yes | Yes | Yes |
It is commonly known that sorafenib and erlotinib can hinder drug metabolism through CYP3A4 inhibition, but our derivatives 10 and 12 cannot. The predicted elimination can be correlated with the total clearance, which is important for estimating dosing intervals. Erlotinib presented higher clearance rates than sorafenib and our compounds. Erlotinib needs shorter dosing intervals as it is removed more quickly. In contrast to erlotinib, our compounds indicated broader dosing windows and prolonged half-life, as they have lower clearance rates. Hepatotoxicity was demonstrated by sorafenib, erlotinib and compounds 8 and 12, while derivative 10 did not exhibit this unwanted hepatotoxic effect. Lastly, the acute toxic doses of erlotinib and sorafenib are similar to those of our compounds. Moreover, the oral chronic toxic doses of compound 12 are higher than those of both erlotinib and sorafenib, while derivatives 8 and 10 exhibited lower doses.
4. Conclusion
Novel pyrazole/pyrimidine derivatives substituted with azobenzenes were synthesized using microwave and traditional methods. Our compounds were assessed for cytotoxicity against HepG2, MCF-7, HCT-116 and A549 cells as dual inhibitors for EGFRT790M and VEGFR-2. A docking study was carried out to show the binding affinities and orientations of our derivatives in the active sites of VEGFR-2 and EGFRT790M. The data of the docking study were highly correlated with the biological data, where high binding affinities towards both VEGFR-2 and EGFRT790M receptors were obtained for compounds 12, 8 and 10. These compounds exhibited the highest cytotoxicity and VEGFR-2/EGFRT790M inhibition assays. The obtained data concluded that the presence of azobenzene linkers augmented the cytotoxicity of pyrazole and pyrimidine derivatives, which may be attributed to the hydrophobic interactions between this azobenzene scaffold and the hydrophobic pockets in the EGFRT790M and VEGFR-2 receptors. The HCT116 and A549 cell lines were extremely affected by our derivatives. Compound 12 exhibited the highest potency against A549, HepG2, MCF-7, and HCT116 cancer cell lines, with IC50 = 5.12, 6.77, 5.85, and 5.25 µM, respectively. It presented higher cytotoxicity than erlotinib (IC50 = 5.49, 7.73, 8.20, and 13.91 µM, respectively) and showed lower cytotoxicity than sorafenib (IC50 = 4.04, 4.00, 5.58, and 5.05 µM, respectively). Against the HCT116 cell lines, compounds 5, 6, 7, 8, 9 and 10, with IC50 = 6.00–8.25 µM, offered very good cytotoxicity. Against the MCF-7 cell lines, derivatives 5, 6, 7, 8, 9 and 10, with IC50 = 7.15–8.60 µM, offered very good cytotoxicity. With regard to the HepG2 cell lines, derivatives 5, 6, 8, 9 and 11, with IC50 = 7.85–9.75 µM, offered very good cytotoxicity. Against the A549 cell lines, compounds 5, 6, 7, 8, 9, 10 and 11, with IC50 = 6.10–8.80 µM, offered very good cytotoxicity. Moreover, the cytotoxicity of the significantly potent derivatives 5, 6, 8, 9, 10 and 12 against the MCF-10 healthy cell lines was evaluated. The estimated structures indicated low toxicity against MCF-10 cells, with IC50 = 50.90–55.50 µM. Compounds 5, 6, 8, 9, 10 and 12 exhibited 8.09-, 6.53-, 8.92-, 7.30-, 8.10- and 10.57-fold higher toxicity against HCT-116 cells, respectively, than against MCF-10 cells. Equivalently, derivatives 5, 6, 8, 9, 10 and 12 exhibited 7.28-, 6.44-, 7.48-, 7.17-, 7.16- and 9.49-fold higher toxicity against MCF-7 cells, respectively, than against healthy MCF-10 cells. In addition, derivatives 5, 6, 8, 9, 10 and 12 exhibited 6.94-, 5.22-, 7.28-, 6.57-, 6.84- and 8.20-fold higher toxicity against HepG2 cells, respectively, than against MCF-10 cells. Also, derivatives 5, 6, 8, 9, 10 and 12 exhibited 7.64-, 6.74-, 8.77-, 7.22-, 7.58- and 10.84-fold higher toxicity against A549 cells, respectively, than against MCF-10 cells. Furthermore, all compounds were tested for dual EGFRT790M and VEGFR-2 inhibitions. Amongst them, derivatives 12, 8 and 10 showed very good VEGFR-2 inhibitions, with IC50 = 0.90, 0.95 and 1.00 µM, respectively. Similarly, structures 12, 8, 10, 5 and 9 showed strong EGFRT790M inhibitions, with IC50 = 0.25, 0.30, 0.33, 0.35 and 0.40 µM, respectively. In addition, in silico ADMET predictions were calculated for the highly active derivatives 8, 10 and 12 and correlated to Lipinski's rule of five, using erlotinib and sorafenib as standard ligands. The results presented our derivatives as promising candidates for advanced manipulations to get more potent anticancer agents with advanced VEGFR-2 and EGFRT790M inhibitions.5. Experimental
5.1. Chemistry
Microwave technique was carried out using “10 mL” borosilicate glass vials under 2–5 bar pressure at 120 °C in an Anton Paar Monowave 300 for 1–5 min. The spectra of 1H NMR and 13C NMR were recorded on Varian Mercury 300 MHz and 100 MHz-NMR spectrometers, respectively. 2-(4-(Phenyldiazenyl)phenyldiazenyl)-malononitrile (2) was obtained through a recognized method.42–445.1.1.1. 3,5-Diamino-4-(4-(phenyldiazenyl)phenyldiazenyl)-1H-pyrazole-1-carbothioamide (3). Reddish brown crystal; m.p. 268–270 °C; IR (cm−1): 3395, 3304 (NH2), 1617 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1582, 1516 (N
N), 1582, 1516 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1224 (C
N), 1224 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 5.86 (s, 2H, NH2, D2O exchangeable), 6.28 (s, 2H, NH2, D2O exchangeable), 7.031–7.52 (m, 9H, Ar-H), 10.81 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 76.3, 122.2, 122.9, 123.1, 123.2, 124.1, 124.4, 129.1, 129.4, 129.7, 130.0, 132.0, 138.8, 139.0, 147.8 and 182.9; anal. calcd for C16H15N9S (365) C, 52.60; H, 4.11; N, 34.52; S, 8.77, found: C, 52.49; H, 4.27; N, 34.61; S, 8.63.
S); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 5.86 (s, 2H, NH2, D2O exchangeable), 6.28 (s, 2H, NH2, D2O exchangeable), 7.031–7.52 (m, 9H, Ar-H), 10.81 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 76.3, 122.2, 122.9, 123.1, 123.2, 124.1, 124.4, 129.1, 129.4, 129.7, 130.0, 132.0, 138.8, 139.0, 147.8 and 182.9; anal. calcd for C16H15N9S (365) C, 52.60; H, 4.11; N, 34.52; S, 8.77, found: C, 52.49; H, 4.27; N, 34.61; S, 8.63.
5.1.1.2. 3,5-Diamino-4-(4-(phenyldiazenyl)phenyldiazenyl)-1H-pyrazole-1-carboxamide (4). Brown crystal; m.p. > 300 °C; IR (cm−1): 3402, 3304, 3207 (NH2), 1686 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1606 (C
O), 1606 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1511 (N
N), 1511 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 5.92 (s, 2H, NH2, D2O exchangeable), 6.22 (s, 2H, NH2, D2O exchangeable), 7.03–7.52 (m, 9H, Ar-H), 8.73 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 71.6, 119.1, 121.2, 121.3, 122.2, 127.0, 129.0, 129.4, 138.4, 138.8, 144.1, 153.1, 153.7, 154.3 and 158.7; anal. calcd for C16H15N9O (349): C, 55.01; H, 4.30; N, 36.10; found: C, 54.89; H, 4.44; N, 36.17.
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 5.92 (s, 2H, NH2, D2O exchangeable), 6.22 (s, 2H, NH2, D2O exchangeable), 7.03–7.52 (m, 9H, Ar-H), 8.73 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 71.6, 119.1, 121.2, 121.3, 122.2, 127.0, 129.0, 129.4, 138.4, 138.8, 144.1, 153.1, 153.7, 154.3 and 158.7; anal. calcd for C16H15N9O (349): C, 55.01; H, 4.30; N, 36.10; found: C, 54.89; H, 4.44; N, 36.17.
5.1.1.3. 3-(3,5-Diamino-4-(4-(phenyldiazenyl)phenyldiazenyl)-1H-pyrazol-1-yl)-3-oxopropanenitrile (5). Reddish brown crystal; m.p. 254–256 °C; IR (cm−1): 3443 (NH2), 2194 (CN), 1657 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1626 (C
O), 1626 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1599, 1559 (N
N), 1599, 1559 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.41 (s, 2H, CH2), 5.86 (s, 2H, NH2, D2O exchangeable), 6.30 (s, 2H, NH2, D2O exchangeable), 7.03–7.54 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 21.6, 71.4, 119.9, 122.5, 127.8, 128.9, 129.0, 129.2, 129.3, 129.5, 129.8, 130.0, 131.9, 138.4, 138.8, 140.0, 147.7 and 152.8; anal. calcd for C18H15N9O (373): C, 57.91; H, 4.02; N, 33.78; found: C, 57.98; H, 3.92; N, 33.69.
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.41 (s, 2H, CH2), 5.86 (s, 2H, NH2, D2O exchangeable), 6.30 (s, 2H, NH2, D2O exchangeable), 7.03–7.54 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 21.6, 71.4, 119.9, 122.5, 127.8, 128.9, 129.0, 129.2, 129.3, 129.5, 129.8, 130.0, 131.9, 138.4, 138.8, 140.0, 147.7 and 152.8; anal. calcd for C18H15N9O (373): C, 57.91; H, 4.02; N, 33.78; found: C, 57.98; H, 3.92; N, 33.69.
5.1.1.4. 1-(3,5-Diamino-4-(4-(phenyldiazenyl)phenyldiazenyl)-1H-pyrazol-1-yl)ethan-1-one (6). Orange crystal; m.p. 232–234 °C; IR (cm−1): 3425 (NH2), 1650 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1563 (C
O), 1563 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N and N
N and N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.90 (s, 3H, CH3), 6.96 (s, 4H, 2NH2, D2O exchangeable), 7.13–7.54 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 21.6, 63.8, 121.2, 122.3, 123.4, 126.0, 128.9, 129.3, 129.8, 129.9, 131.5, 138.8, 147.7, 152.8 and 152.9; anal. calcd for C17H16N8O (348): C, 58.62; H, 4.60; N, 32.18; found: C, 58.58; H, 4.62; N, 32.23.
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.90 (s, 3H, CH3), 6.96 (s, 4H, 2NH2, D2O exchangeable), 7.13–7.54 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 21.6, 63.8, 121.2, 122.3, 123.4, 126.0, 128.9, 129.3, 129.8, 129.9, 131.5, 138.8, 147.7, 152.8 and 152.9; anal. calcd for C17H16N8O (348): C, 58.62; H, 4.60; N, 32.18; found: C, 58.58; H, 4.62; N, 32.23.
5.1.1.5. 1-Phenyl-4-(4-(phenyldiazenyl)phenyldiazenyl)-1H-pyrazole-3,5-diamine (7). Orange crystal; m.p. > 300 °C; IR (cm−1): 3474, 3395, 3305 (NH2), 1617 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1594, 1516 (N
N), 1594, 1516 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 6.88 (s, 2H, NH2, D2O exchangeable), 6.94 (s, 2H, NH2, D2O exchangeable), 7.03–7.70 (m, 14H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 63.6, 118.9, 121.2, 121.3, 122.2, 122.6, 123.8, 124.2, 127.9, 129.1, 129.3, 135.9, 138.5, 138.8, 147.0, 147.1, 147.4, 152.7, 153.8 and 153.9; anal. calcd for C21H18N8 (382): C, 65.97; H, 4.71; N, 29.32, found: 66.04; H, 4.65; N, 29.31.
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 6.88 (s, 2H, NH2, D2O exchangeable), 6.94 (s, 2H, NH2, D2O exchangeable), 7.03–7.70 (m, 14H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 63.6, 118.9, 121.2, 121.3, 122.2, 122.6, 123.8, 124.2, 127.9, 129.1, 129.3, 135.9, 138.5, 138.8, 147.0, 147.1, 147.4, 152.7, 153.8 and 153.9; anal. calcd for C21H18N8 (382): C, 65.97; H, 4.71; N, 29.32, found: 66.04; H, 4.65; N, 29.31.
5.1.1.6. 1-(2,4-Dinitrophenyl)-4-(4-(phenyldiazenyl)phenyldiazenyl)-1H-pyrazole-3,5-diamine (8). Black crystal; m.p. > 300 °C; IR (cm−1): 3423 (NH2), 1512 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1538 (N
N), 1538 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 7.13–7.85 (m, 12H, Ar-H), 8.04 (s, 2H, NH2, D2O exchangeable), 8.06 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 71.3, 118.4, 119.0, 120.0, 121.3, 122.0, 122.9, 123.1, 123.2, 124.1, 124.4, 129.1, 129.4, 129.7, 130.0, 132.0, 138.8, 139.0, 147.8, 151.0 and 151.6; anal. calcd for C21H16N10O4 (472): C, 53.39; H, 3.39; N, 29.66; found: C, 53.46; H, 3.30; N, 29.71.
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 7.13–7.85 (m, 12H, Ar-H), 8.04 (s, 2H, NH2, D2O exchangeable), 8.06 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 71.3, 118.4, 119.0, 120.0, 121.3, 122.0, 122.9, 123.1, 123.2, 124.1, 124.4, 129.1, 129.4, 129.7, 130.0, 132.0, 138.8, 139.0, 147.8, 151.0 and 151.6; anal. calcd for C21H16N10O4 (472): C, 53.39; H, 3.39; N, 29.66; found: C, 53.46; H, 3.30; N, 29.71.
5.1.2.1. 4,6-Diamino-5-(4-(phenyldiazenyl)phenyldiazenyl)pyrimidin-2(5H)-one (9). Black crystal; m.p. 280–282 °C; IR (cm−1): 3369, 3262, 3180 (NH2), 1643 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1619 (C
O), 1619 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1531 (N
N), 1531 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.36 (s, 1H, CH–N
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.36 (s, 1H, CH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 5.92 (s, 2H, NH2, D2O exchangeable), 6.22 (s, 2H, NH2, D2O exchangeable), 7.03–7.52 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 71.9, 114.6, 118.4, 121.3, 127.8, 129.0, 138.4, 144.0, 148.0, 153.1, 153.7, 154.2 and 158.8; anal. calcd for C16H14N8O (334): C, 57.49; H, 4.19; N, 33.53; found: C, 57.54; H, 4.23; N, 33.47.
N), 5.92 (s, 2H, NH2, D2O exchangeable), 6.22 (s, 2H, NH2, D2O exchangeable), 7.03–7.52 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 71.9, 114.6, 118.4, 121.3, 127.8, 129.0, 138.4, 144.0, 148.0, 153.1, 153.7, 154.2 and 158.8; anal. calcd for C16H14N8O (334): C, 57.49; H, 4.19; N, 33.53; found: C, 57.54; H, 4.23; N, 33.47.
5.1.2.2. 4,6-Diamino-5-(4-(phenyldiazenyl)phenyldiazenyl)pyrimidine-2(5H)-thione (10). Black crystal; m.p. 274–276 °C; IR (cm−1): 3397 (NH2), 1625, 1617 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1555 (N
N), 1555 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1266 (C
N), 1266 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.40 (s, 1H, CH–N
S); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.40 (s, 1H, CH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 5.82 (s, 2H, NH2, D2O exchangeable), 6.32 (s, 2H, NH2, D2O exchangeable), 6.87–7.54 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 79.2, 121.2, 121.3, 122.2, 127.0, 129.0, 129.4, 138.4, 138.8, 144.1, 147.9, 153.1, 153.7, 154.3 and 178.7; anal. calcd for C16H14N8S (350): C, 54.86; H, 4.00; N, 32.00; S, 9.14; found: C, 54.79; H, 4.06; N, 31.92; S, 9.23.
N), 5.82 (s, 2H, NH2, D2O exchangeable), 6.32 (s, 2H, NH2, D2O exchangeable), 6.87–7.54 (m, 9H, Ar-H); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 79.2, 121.2, 121.3, 122.2, 127.0, 129.0, 129.4, 138.4, 138.8, 144.1, 147.9, 153.1, 153.7, 154.3 and 178.7; anal. calcd for C16H14N8S (350): C, 54.86; H, 4.00; N, 32.00; S, 9.14; found: C, 54.79; H, 4.06; N, 31.92; S, 9.23.
5.1.2.3. 2-Imino-5-(4-(phenyldiazenyl)phenyldiazenyl)-2,5-dihydropyrimidine-4,6-diamine (11). Brown crystal; m.p. 264–266 °C; IR (cm−1): 3461, 3417 (NH2 and NH), 1512 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1538 (N
N), 1538 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.31 (s, 1H, CH–N
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.31 (s, 1H, CH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 6.84 (s, 2H, NH2, D2O exchangeable), 6.87 (s, 2H, NH2, D2O exchangeable), 7.15–7.99 (m, 9H, Ar-H), 9.14 (s, 1H, NH, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 65.7, 120.5, 121.2, 122.7, 127.0, 128.8, 132.1, 132.2, 135.5, 136.1, 143.2, 144.2, 150.4, 151.3, 155.3, and 156.3; anal. calcd for C16H15N9 (333): C, 57.66; H, 4.50; N, 37.84, found: C, 57.69; H, 4.51; N, 37.80.
N), 6.84 (s, 2H, NH2, D2O exchangeable), 6.87 (s, 2H, NH2, D2O exchangeable), 7.15–7.99 (m, 9H, Ar-H), 9.14 (s, 1H, NH, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 65.7, 120.5, 121.2, 122.7, 127.0, 128.8, 132.1, 132.2, 135.5, 136.1, 143.2, 144.2, 150.4, 151.3, 155.3, and 156.3; anal. calcd for C16H15N9 (333): C, 57.66; H, 4.50; N, 37.84, found: C, 57.69; H, 4.51; N, 37.80.
5.1.2.4. 4-Amino-N-(4,6-diamino-5-(4-(phenyldiazenyl)phenyldiazenyl)pyrimidin-2(5H)-ylidene) benzenesulfonamide (12). Reddish brown crystal; m.p. 226–228 °C; IR (cm−1): 3439 (NH2), 1620 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1539 (N
N), 1539 (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.41 (s, 1H, CH–N
N); 1H-NMR (300 MHz, DMSO-d6) δ (ppm): 2.41 (s, 1H, CH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 5.86 (s, 2H, NH2, D2O exchangeable), 6.30 (s, 2H, NH2, D2O exchangeable), 6.86–7.89 (m, 13H, Ar-H), 7.96 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 66.0, 118.4, 120.1, 120.7, 121.0, 121.3, 122.5, 127.8, 129.0, 129.3, 130.3, 131.5, 138.4, 139.0, 154.0, 162.4, 162.4, 162.8, 166.1 and 166.5; anal. calcd for C22H20N10SO2 (488): C, 54.10; H, 4.10; N, 28.69; S, 6.56, found: C, 54.03; H, 4.14; N, 28.73; S, 6.48.
N), 5.86 (s, 2H, NH2, D2O exchangeable), 6.30 (s, 2H, NH2, D2O exchangeable), 6.86–7.89 (m, 13H, Ar-H), 7.96 (s, 2H, NH2, D2O exchangeable); 13C-NMR (300 MHz, DMSO-d6) δ (ppm): 66.0, 118.4, 120.1, 120.7, 121.0, 121.3, 122.5, 127.8, 129.0, 129.3, 130.3, 131.5, 138.4, 139.0, 154.0, 162.4, 162.4, 162.8, 166.1 and 166.5; anal. calcd for C22H20N10SO2 (488): C, 54.10; H, 4.10; N, 28.69; S, 6.56, found: C, 54.03; H, 4.14; N, 28.73; S, 6.48.
Conflicts of interest
There is no interest to declare.Data availability
The data supporting this article are available upon request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra08997b.
Acknowledgements
This work was funded by the Deanship of Graduate Studies and Scientific Research at Jouf University under grant no. DGSSR-2025-01-01639.References
- A. A. Alotaibi, H. H. Asiri, A. F. M. M. Rahman and M. M. Alanazi, Novel pyrrolo[2,3-d]pyrimidine derivatives as multi-kinase inhibitors with VEGFR-2 selectivity, J. Saudi Chem. Soc., 2023, 27(5), 101712, DOI:10.1016/j.jscs.2023.101712.
- I. A. Y. Ghannam, A. M. El Kerdawy, M. M. Mounier, M. T. Abo-elfadl and I. H. Ali, Novel 2-oxo-2-phenylethoxy and benzyloxy diaryl urea hybrids as VEGFR-2 inhibitors: Design, synthesis, and anticancer evaluation, Arch. Pharm., 2023, 356(2), 2200341, DOI:10.1002/ardp.202200341.
- N. S. Aparna, S. S. Aswani, M. S. Mohan, P. T. Boban and S. Kamalamma, Sesamol inhibits LPS induced angiogenesis via downregulating VEGFA/VEGFR2 signalling both in vitro and in vivo, Pharmacol. Res. Mod. Chin. Med., 2023, 9, 100302, DOI:10.1016/j.prmcm.2023.100302.
- U. Acar Çevik, I. Celik, Ş. Görgülü, Z. D. Şahin Inan, H. E. Bostancı, Y. Özkay and Z. A. Kaplacıklı, New benzimidazole-oxadiazole derivatives as potent VEGFR-2 inhibitors: Synthesis, anticancer evaluation, and docking study, Drug Dev. Res., 2024, 85(4), e22218, DOI:10.1002/ddr.22218 , PMID: 38825827..
- N. M. Saleh, M. S. A. El-Gaby, K. El-Adl and N. E. A. Abd El-Sattar, Design, green synthesis, molecular docking and anticancer evaluations of diazepam bearing sulfonamide moieties as VEGFR-2 inhibitors, Bioorg. Chem., 2020, 104, 104350, DOI:10.1016/j.bioorg.2020.104350.
- J. Zhu, T. Guo, Z. Wang and Y. Zhao, Triggered azobenzene-based prodrugs and drug delivery systems, J. Controlled Release, 2022, 345, 475–493, DOI:10.1016/j.jconrel.2022.03.041.
- K. El-Adl, A.-H. Abdel-Rahman, A. M. Omar, M. Alswah and N. M. Saleh, Arch. Pharm., 2021, e2100237, DOI:10.1002/ardp.202100237.
- V. Gandin, A. Ferrarese, M. Dalla Via, C. Marzano, A. Chilin and G. Marzaro, Targeting kinases with anilinopyrimidines: discovery of N-phenyl-N′-[4-(pyrimidin-4-ylamino)phenyl]urea derivatives as selective inhibitors of class III receptor tyrosine kinase subfamily, Sci. Rep., 2015, 5, 16750 CrossRef CAS PubMed.
- D. M. Wanode and P. B. Khedekar, Synthetic advances and SAR insights of pyrazole-based VEGFR-2 kinase inhibitors, Med. Drug Discovery, 2025, 28, 100227, DOI:10.1016/j.medidd.2025.100227.
- H. S. H. Mohamed and S. A. Ahmed, Reviewing of Synthesis and Computational Studies of Pyrazolo Pyrimidine Derivatives, J. Chem. Rev., 2019, 1(3), 183–232 CrossRef.
- L. M. Moreno, J. Quiroga, R. Abonia, A. Lauria, A. Martorana, H. Insuasty and B. Insuasty, Synthesis, biological evaluation, and in silico studies of novel chalcone- and pyrazoline-based 1,3,5-triazines as potential anticancer agents, RSC Adv., 2020, 10, 34114, 10.1039/D0RA06799G.
- G. Li, Y. Cheng, C. Han, C. Song, N. Huang and Y. Du, Pyrazole-containing pharmaceuticals: target, pharmacological activity, and their SAR studies, RSC Med. Chem., 2022, 13(11), 1300–1321, 10.1039/d2md00206j.
- N. M. Saleh, M. G. El-Gazzar, H. M. Aly and R. A. Othman, Novel Anticancer Fused Pyrazole Derivatives as EGFR and VEGFR-2 Dual TK Inhibitors, Front. Chem., 2020, 7, 917, DOI:10.3389/fchem.2019.00917.
- S. M. Alhamaky, N. A. Khalil, A. K. A. Bass, N. Osama and M. S. A. Hassan, Design, Synthesis, Docking Studies, and Investigation of Dual EGFR/VEGFR-2 Inhibitory Potentials of New Pyrazole and Pyrazolopyridine Derivatives, Drug Dev. Res., 2025, 86(1), e70056, DOI:10.1002/ddr.70056.
- S. Kumar and B. Narasimhan, Therapeutic potential of heterocyclic pyrimidine scaffolds, Chem. Cent. J., 2018, 12, 386–398 Search PubMed.
- B. De Filippis, A. Della Valle, A. Ammazzalorso, C. Maccallini, G. Tesse, R. Amoroso, A. Mollica and L. Giampietro, Azobenzene as Multi-Targeted Scaffold in Medicinal Chemistry, Molecules, 2024, 29(24), 5872, DOI:10.3390/molecules29245872.
- I. Karakaya, Synthesis and Characterization of Azobenzene Derived from 8-aminoquinoline in Aqueous Media, J. Turk. Chem. Soc., Sect. A, 2022, 9(1), 85–114 CrossRef CAS.
- L. Giampietro, M. Gallorini, N. Gambacorta, A. Ammazzalorso, B. De Filippis, A. Della Valle, M. Fantacuzzi, C. Maccallini, A. Mollica and A. Cataldi, et al., Synthesis, structure-activity relationships and molecular docking studies of phe-nyldiazenyl sulfonamides as aromatase inhibitors, Eur. J. Med. Chem., 2021, 224, 113737, DOI:10.1016/j.ejmech.2021.113737.
- C. Bustos, L. Alvarez-Thon, E. Molins, I. Moreno-Villoslada, G. Vallejos-Contreras, C. Sánchez, X. Zarate, D. M.-L. Carey and E. Schott, Tuning the molecular/electronic structure of new substituted pyrazoles: Synthesis, biological trials, theoretical approaches and Hammett correlations, J. Mol. Struct., 2018, 1171, 349–361, DOI:10.1016/j.molstruc.2018.05.088.
- S. M. Gomha, A. O. Abdelhamid, N. A. Abdelrehem and S. M. Kandeel, Efficient Synthesis of New Benzofuran-based Thiazoles and Investigation of their Cytotoxic Activity Against Human Breast Carcinoma Cell Lines, J. Heterocycl. Chem., 2018, 55, 995–1001, DOI:10.1002/jhet.3131.
- J. Tang, W. Lu, J. Hu, Y. Jia, X. He, L. Li, S. Yang, Y. Wang and L. Xu, Design, synthesis and biological evaluation of Novel 6-azophenylcoumarin-3-formamido derivatives and a copper(II) complex, J. Mol. Struct., 2024, 1301, 137448, DOI:10.1016/j.molstruc.2023.137448.
- K. Shinzawa, D. Kageta, R. J. Nash, G. W. J. Fleet, T. Imahori and A. Kato, Azobenzene derivatives show anti-cancer activity against pancreatic cancer cells only under nutrient starvation conditions via G0/G1 cell cycle arrest, Tetrahedron, 2021, 85, 132077, DOI:10.1016/j.tet.2021.132077.
- A. Sandor, I. Fizesan, I. Ionut, G. Marc, C. Moldovan, I. Oniga, A. Pîrnău, L. Vlase, A. E. Petru, I. Macasoi and O. Oniga, Discovery of A Novel Series of Quinazoline–Thiazole Hybrids as Potential Antiproliferative and Anti-Angiogenic Agents, Biomolecules, 2024, 14(2), 218, DOI:10.3390/biom14020218.
- A. M. Srour, N. S. Ahmed, S. S. Abd El-Karim, M. M. Anwar and S. M. El-Hallouty, Design, synthesis, biological evaluation, QSAR analysis and molecular modelling of new thiazol-benzimidazoles as EGFR inhibitors, Bioorg. Med. Chem., 2020, 28(18), 115657, DOI:10.1016/j.bmc.2020.115657.
- N. E. M. Kaufman, S. Dhingra, S. D. Jois and M. D. G. H. Vicente, Molecular Targeting of Epidermal Growth Factor Receptor (EGFR) and Vascular Endothelial Growth Factor Receptor (VEGFR), Molecules, 2021, 26(4), 1076, DOI:10.3390/molecules26041076.
- C. Tomuleasa, A. B. Tigu and R. Munteanu, et al., Therapeutic advances of targeting receptor tyrosine kinases in cancer, Signal Transduction Targeted Ther., 2024, 9, 201, DOI:10.1038/s41392-024-01899-w.
- G. M. Nitulescu, G. Stancov, O. C. Seremet, G. Nitulescu, D. P. Mihai, C. G. Duta-Bratu, S. F. Barbuceanu and O. T. Olaru, The Importance of the Pyrazole Scaffold in the Design of Protein Kinases Inhibitors as Targeted Anticancer Therapies, Molecules, 2023, 28(14), 5359, DOI:10.3390/molecules28145359.
- R. Dubey, R. Makhija, A. Sharma, A. Sahu and V. Asati, Unveiling the promise of pyrimidine-modified CDK inhibitors in cancer treatment, Bioorg. Chem., 2024, 149, 107508, DOI:10.1016/j.bioorg.2024.107508.
- A. Jug and J. Ilaš, ATP-competitive inhibitors for cancer treatment – kinases and the world beyond, RSC Med. Chem., 2025, 16(9), 4044–4067 RSC , https://api.semanticscholar.org/CorpusID:279604312.
- A. M. El-Naggar, A. M. A. Hassan, E. B. Elkaeed, M. S. Alesawy and A. Al-Karmalawy, Design, synthesis, and SAR studies of novel 4-methoxyphenyl pyrazole and pyrimidine derivatives as potential dual tyrosine kinase inhibitors targeting both EGFR and VEGFR-2, Bioorg. Chem., 2022, 123, 105770, DOI:10.1016/j.bioorg.2022.105770.
- M. Alsulaimany, A. K. B. Aljohani, N. E. A. Abd El-Sattar, S. A. Almadani, O. M. Alatawi, H. Y. Alharbi, M. S. Aljohani, A. H. Al-Shareef, R. Alghamdi, S. M. Tayeb, D. E. Keshek, K. El-Adl and K. E. Anwer, Dual VEGFR-2 and EGFRT790M inhibitors of phenyldiazenes: anticancer evaluations, ADMET, docking, design and synthesis, Future Med. Chem., 2025, 17(3), 287–300, DOI:10.1080/17568919.2025.2453409.
- A. M. Sayed, F. A. Taher, M. R. K. Abdel-Samad, M. S. A. El-Gaby, K. El-Adl and N. M. Saleh, Design, synthesis, molecular docking, in silico ADMET profile and anticancer evaluations of sulfonamide endowed with hydrazone-coupled derivatives as VEGFR-2 inhibitors, Bioorg. Chem., 2021, 108, 104669, DOI:10.1016/j.bioorg.2021.104669.
- N. M. Saleh, A. A. Abdel-Rahman, A. M. Omar, M. M. Khalifa and K. El-Adl, Pyridine-derived VEGFR-2 inhibitors: Rational design, synthesis, anticancer evaluations, in silico ADMET profile, and molecular docking, Arch. Pharm., 2021, 354(8), e2100085, DOI:10.1002/ardp.202100085.
- M. Alsulaimany, F. A. Abdulhaleem M., R. Alghamdi, N. E. A. Abd El-Sattar, S. A. Almadani, W. A. Samman, H. Y. Alharbi, M. S. Aljohani, S. M. Tayeb, T. Nasr, K. El-Adl and K. E. Anwer, ADMET, docking, anticancer evaluations, design and synthesis of pyrazolo[1,5-a]pyrimidines substituted with furan and phenyldiazene as dual EGFRT790M and VEGFR-2 inhibitors, Bioorg. Chem., 2025, 163, 108746, DOI:10.1016/j.bioorg.2025.108746.
- M. Alsulaimany, A. K. B. Aljohani, N. E. A. Abd El-Sattar, S. A. Almadani, O. M. Alatawi, H. Y. Alharbi, M. S. Aljohani, A. H. Al-Shareef, R. Alghamdi, S. M. Tayeb, D. E. Keshek, K. El-Adl and K. E. Anwer, Dual VEGFR-2 and EGFRT790M inhibitors of phenyldiazenes: anticancer evaluations, ADMET, docking, design and synthesis, Future Med. Chem., 2025, 17(3), 287–300, DOI:10.1080/17568919.2025.2453409 , PMID: 39819342, PMCID: PMC11792794..
- A. K. B. Aljohani, S. S. A. El-Hddad, M. Alsulaimany, N. A. Maghrabi, A. M. Alhammad, M. N. Aloufi, L. F. Alahmadi, T. A. Saeedi, Y. Y. Neyaz, K. E. Anwer, N. E. A. Abd El-Sattar and K. El-Adl, Design, nanogel synthesis, anti-proliferative activity and in silico ADMET profile of pyrazoles and pyrimidines as topo-II inhibitors and DNA intercalators, RSC Adv., 2025, 15(13), 10037–10048, 10.1039/d5ra00166h.
- Q.-Q. Xie, et al., Pharmacophore modeling studies of type I and type II kinase inhibitors of Tie2, J. Mol. Graphics Modell., 2009, 27(6), 751–758 CrossRef CAS PubMed.
- K. Lee, et al., Pharmacophore modeling and virtual screening studies for new VEGFR-2 kinase inhibitors, Eur. J. Med. Chem., 2010, 45(11), 5420–5427 CrossRef CAS PubMed.
- V. A. Machado, et al., Synthesis, antiangiogenesis evaluation and molecular docking studies of 1-aryl-3-[(thieno[3,2-b]pyridin-7-ylthio)phenyl]ureas: Discovery of a new substitution pattern for type II VEGFR-2 Tyr kinase inhibitors, Bioorg. Med. Chem., 2015, 23(19), 6497–6509 CrossRef CAS PubMed.
- F. Sangande, E. Julianti and D. H. Tjahjono, Ligand-Based Pharmacophore Modeling, Molecular Docking, and Molecular Dynamic Studies of Dual Tyrosine Kinase Inhibitor of EGFR and VEGFR2, Int. J. Mol. Sci., 2020, 21(20), 7779, DOI:10.3390/ijms21207779.
- S. A. Elmetwally, K. F. Saied, I. H. Eissa and E. B. Elkaeed, Design, synthesis and anticancer evaluation of thieno[2,3-d]pyrimidine derivatives as dual EGFR/HER2 inhibitors and apoptosis inducers, Bioorg. Chem., 2019, 88, 102944, DOI:10.1016/j.bioorg.2019.102944.
- M. M. Mashaly, A. T. Ramadan, B. A. El-Shetary and A. K. Dawoud, Synthesis and Characterization of New Transition and Actinide Metal Complexes of a Hydrazone Ligand. Mixed-Ligand Complexes, Pyrolysis Products, and Biological Activity, Synth. React. Inorg. Met.-Org. Chem., 2004, 34(8), 1319–1348 CrossRef CAS.
- K. E. Anwer, S. S. A. El-Hddad, N. E. A. Abd El-Sattar, A. El-Morsy, F. Khedr, S. Mohamady, D. E. Keshek, S. A. Salama, K. El-Adl and N. S. Hanafy, Five and six membered heterocyclic rings endowed with azobenzene as dual EGFRT790M and VEGFR-2 inhibitors: design, synthesis, in silico ADMET profile, molecular docking, dynamic simulation and anticancer evaluations, RSC Adv., 2023, 13(50), 35321–35338, 10.1039/d3ra06614b.
- M. Alsulaimany, A. K. B. Aljohani, N. E. A. Abd El-Sattar, S. A. Almadani, O. M. Alatawi, H. Y. Alharbi, M. S. Aljohani, A. H. Al-Shareef, R. Alghamdi, S. M. Tayeb, D. E. Keshek, K. El-Adl and K. E. Anwer, Dual VEGFR-2 and EGFRT790M inhibitors of phenyldiazenes: anticancer evaluations, ADMET, docking, design and synthesis, Future Med. Chem., 2025, 17(3), 287–300, DOI:10.1080/17568919.2025.2453409.
- B. M. Essa, A. A. Selim, G. H. Sayed and K. E. Anwer, Conventional and microwave-assisted synthesis, anticancer evaluation, 99mTc-coupling and In-vivo study of some novel pyrazolone derivatives, Bioorg. Chem., 2022, 125, 105846, DOI:10.1016/j.bioorg.2022.105846.
- N. E. A. Abd El-Sattar, E. H. K. Badawy, E. Z. Elrazaz and N. S. M. Ismail, Discovery of pyrano[2,3-d]pyrimidine-2,4-dione derivatives as novel PARP-1 inhibitors: Design, synthesis and antitumor activity, RSC Adv., 2021, 11(8), 4454–4464, 10.1039/D0RA10321G.
- M. K. A. Regal, E. Rafat and N. E. A. Abd El-Sattar, Synthesis, characterization, and dyeing performance of some azo thienopyridine and thienopyrimidine dyes based on wool and nylon, J. Heterocycl. Chem., 2020, 57(3), 1173–1182, DOI:10.1002/jhet.3854.
- K. E. Anwer and G. H. Sayed, Conventional and microwave reactions of 1,3-diaryl-5,4-enaminonitrile-pyrazole derivative with expected antimicrobial and anticancer activities, J. Heterocycl. Chem., 2020, 57(6), 2339–2353, DOI:10.1002/jhet.3946.
- K. E. Anwer, G. H. Sayed and R. M. Ramadan, Synthesis, spectroscopic, DFT calculations, biological activities and molecular docking studies of new isoxazolone, pyrazolone, triazine, triazole and amide derivatives, J. Mol. Struct., 2022, 1256, 132513, DOI:10.1016/j.molstruc.2022.132513.
- F. Khedr, M.-K. Ibrahim, I. H. Eissa, H. S. Abulkhair and K. El-Adl, Phthalazine-based VEGFR-2 inhibitors: Rationale, design, synthesis, in silico, ADMET profile, docking, and anticancer evaluations, Arch. Pharm., 2021, e2100201, DOI:10.1002/ardp.202100201.
- M. McTigue, B. W. Murray, J. H. Chen, Y. Deng, J. Solowiej and R. S. Kania, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 18281, DOI:10.1073/pnas.1207759109.
- S. A. Elmetwally, K. F. Saied, I. H. Eissa and E. B. Elkaeed, Design, synthesis and anticancer evaluation of thieno[2,3-d]pyrimidine derivatives as dual EGFR/HER2 inhibitors and apoptosis inducers, Bioorg. Chem., 2019, 88, 102944, DOI:10.1016/j.bioorg.2019.102944.
- T. Mosmann, Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays, J. Immunol. Methods, 1983, 65(1–2), 55–63, DOI:10.1016/0022-1759(83)90303-4.
- F. M. Freimoser, et al., The MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide] assay is a fast and reliable method for colorimetric determination of fungal cell densities, Appl. Environ. Microbiol., 1999, 65(8), 3727–3729 CrossRef CAS PubMed.
- K. El-Adl, et al., Design, synthesis, molecular docking, and anticancer evaluations of 1-benzylquinazoline-2,4(1H,3H)-dione bearing different moieties as VEGFR-2 inhibitors, Arch. Pharm., 2020, e2000068, DOI:10.1002/ardp.202000068.
- C. Deng, J. Xiong, X. Gu, X. Chen, S. Wu, Z. Wang, D. Wang, J. Tu and J. Xie, Novel recombinant immunotoxin of EGFR specific nanobody fused with cucurmosin, construction and antitumor efficiency in vitro, Oncotarget, 2017, 8(24), 38568–38580 CrossRef PubMed.
- N. A. A. M. Aziz, R. F. George, K. El-Adl and W. R. Mahmoud, Exploration of thiazolidine-2,4-diones as tyrosine kinase inhibitors: Design, synthesis, ADMET, docking, and antiproliferative evaluations, Arch. Pharm., 2022, e2200465, DOI:10.1002/ardp.202200465.
- S. M. Abou-Seri, et al., 1-Piperazinylphthalazines as potential VEGFR-2 inhibitors and anticancer agents: synthesis and in vitro biological evaluation, Eur. J. Med. Chem., 2016, 107, 165–179 CrossRef CAS PubMed.
- C. A. Lipinski, F. Lombardo, B. W. Dominy and P. J. Feeney, Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings, Adv. Drug Delivery Rev., 1997, 23(1–3), 3–25, DOI:10.1016/S0169-409X(96)00423-1.
- D. E. V. Pires, T. L. Blundell and D. B. Ascher, pkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Properties Using Graph-Based Signatures, J. Med. Chem., 2015, 58(9), 4066–4072, DOI:10.1021/acs.jmedchem.5b00104.
- A. Beig, R. Agbaria and A. Dahan, Oral Delivery of Lipophilic Drugs: The Tradeoff between Solubility Increase and Permeability Decrease When Using Cyclodextrin-Based Formulation, PLoS One, 2013, 8(10), e68237, DOI:10.1371/journal.pone.0068237.
| This journal is © The Royal Society of Chemistry 2026 |