 Open Access Article
Open Access ArticleReaction yield oscillates over reaction time in first-order chemical reactions
Yu Harabuchi†
ab,
Tomohiko Yokoyama†c,
Keisuke Katayamac,
Satoshi Maeda abd,
Taihei Oki*abe and
Satoru Iwata
abd,
Taihei Oki*abe and
Satoru Iwata *abc
*abc
aInstitute for Chemical Reaction Design and Discovery (WPI-ICReDD), Hokkaido University, Kita 21, Nishi 10, Kita-ku, Sapporo, Hokkaido 001-0021, Japan. E-mail: oki@icredd.hokudai.ac.jp; iwata@mist.i.u-tokyo.ac.jp
bJST, ERATO Maeda Artificial Intelligence in Chemical Reaction Design and Discovery Project, Kita 10, Nishi 8, Kita-ku, Sapporo, Hokkaido 060-0810, Japan
cDepartment of Mathematical Informatics, Graduate School of Information Science and Technology, The University of Tokyo, Hongo 7-3-1, Bunkyo-ku, Tokyo 113-8656, Japan
dDepartment of Chemistry, Faculty of Science, Hokkaido University, Kita 10, Nishi 8, Kita-ku, Sapporo, Hokkaido 060-0810, Japan
eCenter for Advanced Intelligence Project, RIKEN, Nihombashi 1-4-1, Chuo-ku, Tokyo 103-0027, Japan
First published on 9th February 2026
Abstract
The yield of a first-order chemical reaction normally has a unimodal trend over reaction time. In other words, once the population of a product decreases, it will normally never increase again by elongating the reaction time. However, in analyzing the full reaction path network of the Strecker reaction, we encountered a nontraditional behavior, where the population of an intermediate increased again after decreasing, even in a first-order reaction. Furthermore, simulations on model reaction path networks suggested that the reaction yield can oscillate multiple times over the reaction time. We mathematically prove that the number of reaction yield oscillations is at most NEQ/2, where NEQ denotes the number of equilibrium structures in the reaction path network. We also show that this upper bound is tight by constructing a model network that exhibits the maximum number of yield oscillations. This study provides new theoretical insight into reaction optimization strategies aimed at controlling product yields.
1. Introduction
Oscillating phenomena often appear in nonequilibrium dynamics, in both biology and chemistry. In biology, ATP (adenosine triphosphate)-driven proton pumps undergo repetitive cycles of conformational changes powered by ATP hydrolysis, and exhibit cyclic behavior that is crucial for maintaining cellular functions.1 Another prominent example in biology is glycolytic oscillation, a phenomenon in yeast where the concentrations of metabolic intermediates fluctuate periodically, playing a crucial biological role.2,3 In chemistry, the Belousov–Zhabotinsky (BZ) reaction is a well-known example of temporal oscillations, famous for its dramatic, periodic color changes in solution and the formation of complex spatial patterns like spiral waves.4,5 It serves as a paradigm for studying how nonlinear feedback mechanisms can lead to self-organization in chemical systems.6These phenomena are generally regarded as characteristic of open nonequilibrium systems that exchange energy or matter with their environment, and continue to motivate broad research efforts.7–14 In contrast, oscillatory changes in product yield within isolated, closed equilibrium systems have received little attention. It has generally been assumed that the concentration of products in such systems evolves monotonically towards equilibrium, without exhibiting oscillating or non-monotonic behaviour over time.
Under the assumption of a first-order reaction, reaction kinetics are described based on the elementary processes. Computational approaches play a critical role in the analysis of elementary processes.15–20 An elementary step (also referred to as a reaction path) represents a transition from one EQ to another EQ via a TS, as shown in Fig. 1a. Recently, it has become possible to systematically explore many equilibrium (EQ) and transition state (TS) structures on potential energy surfaces, and to construct the reaction path network (Fig. 1b) consisting of thousands of reaction paths.21–32 Such networks have also been used to study reaction mechanisms, as well as to design materials33 and chemical reactions.34,35
Transition state theory36–38 estimates the rate constants corresponding to each reaction path at a specific temperature. Using these rate constants, kinetic simulations37,39–42 predict the population (denoted by reaction yields for the products) of each EQ starting from the given reactants at specific times. The chemical kinetics is assumed to follow the first-order rate equation:
 | (1) |
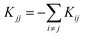 . In the canonical ensemble, the rate constant Kij is given by
. In the canonical ensemble, the rate constant Kij is given by
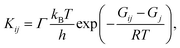 | (2) |
First-order chemical reactions usually show a unimodal trend over time in their reaction yield. In other words, once the reaction yield of a species decreases, it will normally never increase again. However, we have encountered cases that behaved differently from those in analyzing a reaction path network of the Strecker reaction45,46 shown in Fig. 2a. The reaction path network includes 9203 EQs and 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 156 TSs, and Fig. 2b represents the simplified graph network in which different nodes represent different chemical species. Fig. 2c describes the population variations for one of the intermediates, EQ1076. The change of population is simulated by numerically solving the ordinary differential equation (ODE). As the coefficient matrix was extremely ill-conditioned, we employed the Lanczos algorithm for eigendecomposition with 200-digit precision floating-point arithmetic, following the approach of Iwata, Oki, and Sakaue.44 As shown in Fig. 2c, the reaction yield of EQ1076 once decreases at around 10−8 seconds and increases again at around 10−2 seconds over the reaction time. In other words, yield oscillations are observed in a large reaction path network.
156 TSs, and Fig. 2b represents the simplified graph network in which different nodes represent different chemical species. Fig. 2c describes the population variations for one of the intermediates, EQ1076. The change of population is simulated by numerically solving the ordinary differential equation (ODE). As the coefficient matrix was extremely ill-conditioned, we employed the Lanczos algorithm for eigendecomposition with 200-digit precision floating-point arithmetic, following the approach of Iwata, Oki, and Sakaue.44 As shown in Fig. 2c, the reaction yield of EQ1076 once decreases at around 10−8 seconds and increases again at around 10−2 seconds over the reaction time. In other words, yield oscillations are observed in a large reaction path network.
 | ||
| Fig. 2 (a) Reaction scheme and (b) reaction path network of the Strecker reaction.45,46 Three-dimensional structures of reactant, intermediate, and product are shown in Fig. 1c. Part (c) shows the variation of reaction yields over time computed by solving the ODE. The initial population is set to 1.0 for the reactant, EQ1126, and the simulation is done at 300 K. The horizontal axis represents the reaction time in log scale, and the vertical axis is for the population. The red curve indicates the time evolution of the population for the EQ1076. | ||
In response, we investigated this behavior, both numerically and mathematically. In the process, we also obtained insights into the conditions under which these reaction yields oscillate. We then provide a mathematical upper bound on the number of oscillations, and demonstrate its tightness by constructing a network whose yield oscillates as many times as the upper bound. These findings provide a valuable perspective on how yield oscillations may affect the study of actual chemical reaction development.
2. Results and discussion
We design simple model networks to elucidate how reaction yields can oscillate. Fig. 3a–d show a simple model network with NEQ = 4, which demonstrates the oscillation of the reaction yield on EQ1(one of the products). Fig. 3a indicates the initial situation in which 100% of the population is attributed to EQ0, which is unstable (1000 kJ mol−1), and there are three product minima, EQ1–EQ3, having the same energy of 0 kJ mol−1. Immediately after the reaction begins, the population of EQ0 with the zero barrier paths bifurcates into EQ1 and EQ2 in a 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 split, as illustrated in Fig. 3b. Consequently, the population of EQ1 increases to approximately 50%. The activation barriers between EQ1 and EQ2, and between EQ1 and EQ3, are 100 kJ mol−1 and 10 kJ mol−1, respectively. Thus, the reaction from EQ1 to EQ3 proceeds much faster than the process between EQ1 and EQ2, and the population of EQ1 (≈50%) in Fig. 3b is divided into two nodes, EQ1 and EQ3. After the equilibration between EQ1 and EQ3, the population of EQ1 and EQ3 becomes around 25% as shown in Fig. 3c. Finally, after the process between EQ1 and EQ2 proceeds, the equilibration among EQ1–EQ3 results in around 33% of the population on EQ1–EQ3. In summary, the population of EQ1 starts at 0% and first increases to 50%, then decreases to 25% and finally increases again to 33%. This model network verifies the occurrence of a yield oscillation.
50 split, as illustrated in Fig. 3b. Consequently, the population of EQ1 increases to approximately 50%. The activation barriers between EQ1 and EQ2, and between EQ1 and EQ3, are 100 kJ mol−1 and 10 kJ mol−1, respectively. Thus, the reaction from EQ1 to EQ3 proceeds much faster than the process between EQ1 and EQ2, and the population of EQ1 (≈50%) in Fig. 3b is divided into two nodes, EQ1 and EQ3. After the equilibration between EQ1 and EQ3, the population of EQ1 and EQ3 becomes around 25% as shown in Fig. 3c. Finally, after the process between EQ1 and EQ2 proceeds, the equilibration among EQ1–EQ3 results in around 33% of the population on EQ1–EQ3. In summary, the population of EQ1 starts at 0% and first increases to 50%, then decreases to 25% and finally increases again to 33%. This model network verifies the occurrence of a yield oscillation.
The model reaction path network in Fig. 3e has the same structure of the network as Fig. 3a–3d, but with the energy modified to achieve a realistic timescale in the real world. Fig. 3f shows the population change simulated by solving the ODE. As shown in Fig. 3f, the reaction yields vary as approximately 0%, 38.2%, 20.4%, and 50.0% along different time scales of 0, 5, 300, and 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 seconds. This demonstrates the possible situation of reaction yield oscillation in the actual experiment.
000 seconds. This demonstrates the possible situation of reaction yield oscillation in the actual experiment.
Fig. 3g shows a reaction path network in which the reaction starts with 100% population at the high-energy equilibrium point (EQ0). Fig. 3h illustrates the variation in the EQ1 population (reaction yield) as a result of numerically solving the ODE on the network. In the initial stage of reaction (before 10−12 s), the initial population of EQ0 (100%) spreads to EQ1–EQ5, via TS0–TS4, respectively. After the time scale of 10−12 s, this population on EQ1–EQ5 flows into EQ1 and spreads to EQ6–EQ9, sequentially. In other words, the population on EQ1 changes by the following factors: decrease by spreading from EQ1 to EQ6, increase by flowing into EQ1 from EQ2, decrease by spreading to EQ6 and EQ7, increase by flowing into EQ1 from EQ2 and EQ3, decrease by spreading to EQ6–8, …, and so on. This model network, which includes NEQ = 10, demonstrates the population of EQ1 increasing or decreasing nine times, i.e., NEQ − 1 times. These results led us to analyze the maximum number of times of this oscillation from a mathematical point of view.
We mathematically show that the number of yield oscillations of every EQ is at most [NEQ/2], where [·] denotes the rounding down into an integer, indicating NEQ − 1 increases/decreases of the population in the EQ. This upper bound is straightforwardly obtained by using classical Descartes' rule for the number of roots of an exponential polynomial. This gives rise a natural question: is this upper bound tight, i.e., is there a network that oscillates exactly [NEQ/2] times? We affirmatively answer this question by constructing such a model network, which is illustrated in Fig. 4. This model network in Fig. 4 was constructed as one of the simple extensions of the system illustrated in Fig. 3a–d, including a single reactant with high energy and NEQ − 1 equilibrium states (EQs) that all have identical energies. In this network, the population originating from one reactant is distributed among the NEQ − 1 EQs and gradually converges to chemical equilibrium. All the precise statements and proofs can be found in the SI.
 | ||
| Fig. 4 A model reaction path network for the mathematical verification of reaction yield oscillation. A detailed discussion is shown in the supporting information. | ||
In actual organic synthesis experiments, there are two constraints on measurement accuracy to observe yield oscillations. One is the observation error in reaction yields, and the other is the experimentally distinguishable time scale difference. The error in the organic synthesis reaction yield is usually around 5–10%. Thus, the gap between the lower and the higher population during the oscillation requires more than 10%. Also, to observe a yield oscillation, the time scale of the population up/down during the yield oscillation should be about 100 times different. Fig. 5 illustrates one chemically plausible scenario with abstracted molecular structures (EQ0–EQ3). The reaction path network is identical to that shown in Fig. 3e. In Fig. 5, the reaction proceeds from the reactant (EQ0) to an intermediate (EQ2) and one of the product isomers (EQ1) in the initial stage, increasing the EQ1 yield. Subsequently, the population of EQ1 converges to an equilibrium between EQ1 and EQ3 (another product isomer), leading to a decrease in the EQ1 yield and an increase in the EQ3 yield. Finally, the population in EQ2 is distributed into EQ1 and EQ3, resulting in a subsequent increase in the EQ1 yield. In addition, this simple example demonstrates that yield oscillations can be observed by solving a master equation consisting of four equilibrium states and four reaction paths (corresponding to eight rate constants).
 | ||
| Fig. 5 Schematic illustration of a system that exhibits oscillatory behavior in the yield of EQ1 over time. The reaction network is identical to that shown in Fig. 3e. | ||
Additionally, we need to control the reaction barriers among EQ0–EQ3 to satisfy the experimental measurement requirements. As shown in Fig. 5 and 3f, when the reaction yield varies three times (up-down-up), the yield needs to be measured over a wide range of timescales (around 106). This means that we need to measure the yield of reactions with short (sub-second) and long (106 seconds) reaction times, as shown in Fig. 3e and f. Additionally, several reaction barriers of elementary processes in a chemical reaction must be designed for the specific reaction timescales. Consequently, it might be difficult to realise yield oscillations in the actual system within the first-order reaction process. On the other hand, recent advances in the experimental techniques for conducting precise time-controlled experiments, such as flow chemical synthesis, may open the way to achieving the reaction yield oscillation. The experimental demonstration will break the assumption that the concentration of products evolves towards equilibrium without non-monotonic behaviour over time in a first-order reaction.
3. Conclusion
In this study, we demonstrated the possibility of reaction yield oscillation by controlling the reaction path network under the assumption of a first-order reaction. The analyses of reaction yields based on transition state theory38 for the actual reaction path network shown in the SCAN platform46 exhibited the behavior of a reaction yield oscillation on a specific EQ. We mathematically proved that the number of reaction yield oscillations is at most [NEQ/2]. Furthermore, we constructed a model network that attains this tight upper bound on the number of yield oscillations.This study investigates a particular property of master-equation solutions, namely, the oscillatory behavior of the solutions. The present discussion is limited to reaction systems composed of first-order reactions. However, it further suggests that this discussion can be extended to general first-order ODE systems that satisfy the detailed balance condition. Maximizing the reaction yield of a desired product remains one of the central challenges in chemistry. Therefore, the present findings provide promising insights for developing strategies to optimize and control reaction yields.
Conflicts of interest
There are no conflicts to declare.Data availability
The molecular geometries of EQs and TSs, the rate constant matrix of the reaction path network of the Strecker reaction, are available in the SCAN database45,46 at https://scan.sci.hokudai.ac.jp/. All information about the model networks is provided in Fig. 3.Supplementary information (SI): the mathematical proof of the reaction yield oscillation is described in the supporting information (PDF). See DOI: https://doi.org/10.1039/d5ra08614k.
Acknowledgements
This study was funded by the Japan Science and Technology Agency (JST) through CREST (Grant Number JPMJCR24Q2), ERATO (Grant Number JPMJER1903), and FOREST (Grant Number JPMJFR2221), and by Japan Society for the Promotion of Science (JSPS) through WPI Program and KAKENHI (Grant-in-Aid for Transformative Research Areas (B), Grant Number JP23B203).References
- P. D. Boyer, Annu. Rev. Biochem., 1997, 66, 717–749 CrossRef CAS PubMed.
- A. Ghosh and B. Chance, Biochem. Biophys. Res. Commun., 1964, 16, 174–181 Search PubMed.
- B. Hess and A. Boiteux, Annu. Rev. Biochem., 1971, 40, 237–258 CrossRef CAS PubMed.
- B. P. Belousov, Sb. Ref. Po Radiatsionnoi Meditsine, 1959, pp. 145–147..
- A. M. Zhabotinskii, Biofizika, 1964, 9, 306–311 CAS.
- R. J. Field, E. Koros and R. M. Noyes, J. Am. Chem. Soc., 1972, 94, 8649–8664 CrossRef CAS.
- M. Feinberg, Foundations of Chemical Reaction Network Theory, Springer International Publishing, Cham, 2019, vol. 202. Search PubMed.
- P. L. Gentili and J.-C. Micheau, J. Photochem. Photobiol., C, 2020, 43, 100321 CrossRef CAS.
- A. I. Hanopolskyi, V. A. Smaliak, A. I. Novichkov and S. N. Semenov, ChemSystemsChem, 2021, 3, e2000026 Search PubMed.
- S. Ghosh, M. G. Baltussen, N. M. Ivanov, R. Haije, M. Jakštaitė, T. Zhou and W. T. S. Huck, Chem. Rev., 2024, 124, 2553–2582 CrossRef CAS PubMed.
- N. Vassena and P. F. Stadler, Proc. R. Soc. A, 2024, 480, 20230694 CrossRef.
- J. Kaur and J. P. Barham, Chem.–Eur. J., 2025, e02075 CrossRef PubMed.
- M. A. Budroni, M. Rustici and F. Rossi, ChemPhysChem, 2025, 26, e202400841 CrossRef CAS PubMed.
- A. Blokhuis, P. F. Stadler and N. Vassena, arXiv, preprint, arXiv:2508.15273, 2025, DOI:10.48550/arXiv.2508.15273.
- H. B. Schlegel, J. Comput. Chem., 2003, 24, 1514–1527 CrossRef CAS PubMed.
- K. N. Houk and P. H.-Y. Cheong, Nature, 2008, 455, 309–313 CrossRef CAS PubMed.
- W. Thiel, Angew. Chem., Int. Ed., 2014, 53, 8605–8613 CrossRef CAS PubMed.
- W. M. C. Sameera, S. Maeda and K. Morokuma, Acc. Chem. Res., 2016, 49, 763–773 CrossRef CAS PubMed.
- K. N. Houk and F. Liu, Acc. Chem. Res., 2017, 50, 539–543 Search PubMed.
- S. Ahn, M. Hong, M. Sundararajan, D. H. Ess and M.-H. Baik, Chem. Rev., 2019, 119, 6509–6560 Search PubMed.
- S. Maeda, K. Ohno and K. Morokuma, Phys. Chem. Chem. Phys., 2013, 15, 3683–3701 RSC.
- P. M. Zimmerman, J. Comput. Chem., 2015, 36, 601–611 CrossRef CAS PubMed.
- Y. V. Suleimanov and W. H. Green, J. Chem. Theory Comput., 2015, 11, 4248–4259 Search PubMed.
- L.-P. Wang, R. T. McGibbon, V. S. Pande and T. J. Martinez, J. Chem. Theory Comput., 2016, 12, 638–649 CrossRef CAS PubMed.
- A. L. Dewyer, A. J. Argüelles and P. M. Zimmerman, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2018, 8, e1354 Search PubMed.
- G. N. Simm, A. C. Vaucher and M. Reiher, J. Phys. Chem. A, 2019, 123, 385–399 Search PubMed.
- R. Van de Vijver and J. Zádor, Comput. Phys. Commun., 2020, 248, 106947 CrossRef CAS.
- S. Maeda and Y. Harabuchi, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2021, 11, e1538 Search PubMed.
- T. A. Young, J. J. Silcock, A. J. Sterling and F. Duarte, Angew. Chem., Int. Ed., 2021, 60, 4266–4274 Search PubMed.
- M. Liu, A. Grinberg Dana, M. S. Johnson, M. J. Goldman, A. Jocher, A. M. Payne, C. A. Grambow, K. Han, N. W. Yee, E. J. Mazeau, K. Blondal, R. H. West, C. F. Goldsmith and W. H. Green, J. Chem. Inf. Model., 2021, 61, 2686–2696 CrossRef CAS PubMed.
- E. Martínez-Núñez, G. L. Barnes, D. R. Glowacki, S. Kopec, D. Peláez, A. Rodríguez, R. Rodríguez-Fernández, R. J. Shannon, J. J. P. Stewart, P. G. Tahoces and S. A. Vazquez, J. Comput. Chem., 2021, 42, 2036–2048 CrossRef PubMed.
- J. P. Unsleber, S. A. Grimmel and M. Reiher, J. Chem. Theory Comput., 2022, 18, 5393–5409 Search PubMed.
- J. Jiang, Z. J. Wang, R. Staub, Y. Harabuchi, A. Varnek, J. P. Gong and S. Maeda, Chem. Sci., 2025, 16, 14278–14285 RSC.
- T. Mita, Y. Harabuchi and S. Maeda, Chem. Sci., 2020, 11, 7569–7577 RSC.
- H. Hayashi, H. Katsuyama, H. Takano, Y. Harabuchi, S. Maeda and T. Mita, Nat. Synth., 2022, 1, 804–814 CrossRef CAS.
- A. Fernández-Ramos, J. A. Miller, S. J. Klippenstein and D. G. Truhlar, Chem. Rev., 2006, 106, 4518–4584 Search PubMed.
- S. J. Klippenstein, V. S. Pande and D. G. Truhlar, J. Am. Chem. Soc., 2014, 136, 528–546 CrossRef CAS PubMed.
- P. Ptáček, F. Šoukal and T. Opravil, Introduction to the Transition State Theory, in Introducing the Effective Mass of Activated Complex and the Discussion on the Wave Function of This Instanton, InTech, 2018. Search PubMed.
- W. H. Press, S. A. Teukolsky, W. T. Vetterling and B. P. Flannery, Numerical Recipes in Chapter 11, Cambridge Univ. Press, Cambridge, 2nd edn, 1992, pp. 456–495. Search PubMed.
- J. I. Steinfeld, J. S. Francisco and W. L. Hase, Chemical Kinetics and Dynamics, Prentice Hall, Upper Saddle River, NJ, 2nd edn, 1999. Search PubMed.
- S. Kozuch and S. Shaik, Acc. Chem. Res., 2011, 44, 101–110 CrossRef CAS PubMed.
- L. E. Rush, P. G. Pringle and J. N. Harvey, Angew. Chem., Int. Ed., 2014, 53, 8672–8676 CrossRef CAS PubMed.
- Y. Sumiya, Y. Nagahata, T. Komatsuzaki, T. Taketsugu and S. Maeda, J. Phys. Chem. A, 2015, 119, 11641–11649 CrossRef CAS PubMed.
- S. Iwata, T. Oki and S. Sakaue, SIAM J. Sci. Comput., in press Search PubMed.
- Y. Sumiya, Y. Harabuchi, Y. Nagata and S. Maeda, JACS Au, 2022, 2, 1181–1188 CrossRef CAS PubMed.
- M. Kuwahara, Y. Harabuchi, S. Maeda, J. Fujima and K. Takahashi, Digital Discovery, 2023, 2, 1104–1111 Search PubMed.
Footnote |
| † Equally contributed as the first author. |
| This journal is © The Royal Society of Chemistry 2026 |


