 Open Access Article
Open Access ArticleFunctionalization of carbon nanospheres with curcumin, polyethylene glycol and folic acid: potential use as drug carriers
Luis De Sales,
Sergio A. Bernal-Chávez * and
Jessica Campos-Delgado
* and
Jessica Campos-Delgado
Universidad de las Américas Puebla, Escuela de Ciencias, Ex Hacienda Sta. Catarina Mártir S/N, San Andrés Cholula, Puebla 72810, Mexico. E-mail: sergio.bernal@udlap.mx
First published on 23rd January 2026
Abstract
This study reports the synthesis of carbon nanospheres (CNS) modified by anchoring curcumin (CUR), polyethylene glycol (PEG) and folic acid (FA) on their surface as a drug delivery system (DDS) for potential applications in cancer therapy. CNS were synthesized via chemical vapor deposition (CVD) while CUR and PEG were anchored onto their surface by impregnation, and FA conjugation was achieved using N-hydroxysuccinimide (NHS) and 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) as coupling agents. The modified CNS underwent comprehensive characterization using scanning electron microscopy (SEM), Fourier-transform infrared (FTIR) spectroscopy, Raman spectroscopy, thermogravimetric analysis (TGA), X-ray diffraction (XRD), zeta potential measurements, and release profile analysis. The functionalization of CNS was successfully achieved, yielding particles with an average size of 150 nm, a polydispersity index (PDI) of 0.215, zeta potential of −27.7 mV and loading efficiency (LE%) of 63.63 ± 1.61%. Release profile experiments demonstrated a sustained pH-responsive drug release of 3.3% at pH 7.4 vs. 4.3% at pH 5.8 after 48 h.
1. Introduction
Cancer is driven by genetic mutations that cause uncontrolled proliferation of abnormal cells, leading to tissue dysfunction. In 2022, GLOBOCAN reported ∼20 million new cases worldwide.1 Current therapies (chemotherapy, targeted agents and immunotherapies) have improved outcomes but remain limited by adverse effects such as cardiac effects and hypothyroidism, high cost and reduced patient quality of life. The demand for more effective approaches has accelerated research in nanotechnology.2 In this context, nanotechnology enables the design of Drug Delivery Systems (DDS) with improved solubility, stability and pharmacokinetics. These systems use various mechanisms, such as enhanced permeability and retention effect in tumors, and active targeting through receptor-mediated uptake.3,4 Common nanocarriers include natural polymers (chitosan, alginate), polymeric micelles, liposomes and dendrimers, which are effective for hydrophobic drugs but face immune clearance and opsonization issues. Inorganic nanoparticles (gold, iron oxide, silica) offer versatile surface chemistry but raise biocompatibility concerns.5In relation to these nanomaterials, carbon nanostructures have emerged as promising DDS components due to their structural diversity and tunable properties. Carbon's sp, sp2, and sp3 hybridization yields multiple allotropes (nanospheres, nanotubes, graphene, graphene oxide and fullerenes) each with distinct physicochemical features.6 Although carbon nanomaterials are utilized as scaffolds for drug delivery systems, biosensors, and bioimaging diagnostics, they exhibit limitations regarding hydrophilicity, however, surface functionalization with polymers such as polyethylene glycol (PEG), improves dispersibility, evades the reticuloendothelial system, and reduces toxicity.7 Non-covalent interactions (van der Waals, electrostatic interactions, or π–π stacking) enable efficient drug loading.8 Functionalized carbon nanostructures have been developed for delivery of anticancer agents (doxorubicin, cisplatin), nucleic acids (microRNAs), and dual drug–gene systems.9–12
On the other hand, a potential drug for use in the carbon nanomaterials functionalization is curcumin (CUR), the principal bioactive of Curcuma longa, displays anti-inflammatory, antioxidant, antimicrobial and antitumoral activities. CUR shows limited solubility in water. However, it shows good solubility in several organic solvents, including ethanol, acetone, and chloroform. It modulates nuclear factor-κB (NF-κB) and mitogen-activated protein kinase (MAPK) signaling pathways, scavenges free radicals via phenolic groups and inhibits angiogenesis and tumor proliferation.13–15 Moreover, CUR exhibits significant antimicrobial activity against a range of bacteria, fungi, and viruses, indicating its potential role in combating infections. However, CUR's low absorption, rapid metabolism and instability at physiological pH severely limit clinical use. Bioavailability is in the nanomolar range even at 14 g doses, and its half-life at 37 °C is <10 min, with rapid degradation into ferulic acid, vanillin and related products.16,17
CUR, despite its broad therapeutic potential, exhibits extremely low bioavailability due to its rapid intestinal and hepatic metabolism. Even after high doses (10–12 g), serum levels remain minimal (typically undetectable or only reaching nanomolar concentrations in isolated cases) evidencing its poor absorption. This pharmacokinetic limitation has driven the development of nanoscale DDS. Notably, nanoparticle-based formulations achieve higher intratumoral levels and improved antitumor activity compared to free CUR.18,19 In this respect, formulations strategies—liposomes, polymeric carriers and carbon nanomaterials-based systems—have been explored to overcome these drawbacks. Conversely, the application of carbon nanomaterials has been more limited. In this field, advances include carbon nanotubes loaded with CUR for photothermal-triggered drug release (temperature-responsive drug release), and graphene based nanocarriers engineered to respond to external stimuli such as near-infrared irradiation. Enabling controlled release, improved tumor accumulation, and synergetic antitumor activity.15,20–24 Folic acid (FA) plays a fundamental role in nucleotide biosynthesis and cells proliferation, and its elevated demand on tumors results in receptor upregulation. In addition, folate receptors (FR) have gained increasing attention as therapeutic targets in oncology because they are overexpressed in nearly 40% of solid tumors, while their presence in healthy tissues remains very limited. In breast cancer, FR overexpression has been identified in HER2 positive tumors and triple negative breast cancer tumor type, positioning this receptor as a compelling alternative for the development of targeted drug delivery strategies.25 Consequently, functionalization of nanomaterials with FA constitutes an uptake for selective tissues. This approach has been widely explored and has proven to increase therapeutic efficacy and reduce systemic toxicity in cancer treatment.26,27
Circulating nanoparticles bypass the liver first and accumulate in tumors via the enhanced permeability mechanism, although their rapid clearance by the reticuloendothelial system (RES) remains a major limitation. To improve biodistribution (considering some nanoparticles can induce cytotoxicity through oxidative stress) their surfaces are frequently functionalized with polymers such as PEG. This coating enhances nanoparticles stability and biocompatibility, reduces protein adsorption and subsequent RES sequestration, and increases the hydrophilicity of otherwise hydrophobic nanomaterials, thereby improving their dispersion and solubility in physiological environments.28,29 Recent advances in carbon-based nanomaterials have demonstrated their strong potential as platform for targeted and stimulus-responsive DDS, particularly for CUR. Magnetite nanoparticles coated with β-cyclodextrin and poly(propylene glycol) have enabled FA mediated targeting and sustained CUR release, while nitrogen/sulphur-doped carbon dots functionalized with FA have served as fluorescent, biocompatible nanocarriers capable of drug loading and selective uptake via FR recognition.30,31 Similarly, multiwalled carbon nanotubes functionalized with carboxyl groups and FA have shown improved dispersion, pH-responsive release, and enhanced cytotoxicity of CUR against ovarian cancer cells. Together, these studies highlight the relevance of carbon nanostructures, folate targeting, and pH-triggered drug liberation as key strategies to overcome CUR poor bioavailability and limited systemic stability.32 Building on these advances, our work introduces a distinct CNS synthesized via chemical vapor deposition (CVD) system, which differs from previous platforms by combining a spherical, non-tubular carbon core with PEG mediated hydrophilization and enhanced colloidal stability, functionalized with CUR, PEG and FA for potential cancer therapy. Fig. 1 depicts a schematic illustration of our platform, which is designed to enhance CUR attachment, stability and pharmacokinetics through surface modification. To the best of our knowledge, the synthesis, characterization and release profiles of alike surface-modified CNS have not been previously reported.
 | ||
| Fig. 1 General synthesis mechanism of carbon based-drug delivery system and proposed biomedical application. | ||
2. Experimental
2.1. Materials
Commercially available chemicals have been used. Anhydrous Toluene (Merck®) and absolute ethanol (Meyer®) were used as organic solvents during the CNS synthesis by CVD method, along with ultra-high purity argon (99.999%, INFRA®) for a non-reactive environment. Poly(ethylene glycol) ethyl ether (Mn750, PEG, Aldrich®) served as the hydrophilic modifier for surface functionalization. CUR (≥81%, Sigma-Aldrich®) was employed as the model hydrophobic drug for loading onto the CNS surface. Polysorbate 80 (Hyce®) was used as a non-ionic surfactant to enhance CNS dispersion. The coupling agents 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) (Sigma-Aldrich®) and N-hydroxy succinimide (NHS) (Sigma®) were utilized to activate carboxyl groups and facilitate covalent conjugation of FA (Sigma®) onto the nanosystem.2.2. Synthesis of carbon nanospheres
CNS were synthesized in a horizontal atmospheric pressure chemical vapor deposition (APCVD) reactor, following the protocol of Patiño-Guillén et al.33 with minor modifications. A precursors solution consisting of 3 wt% ethanol in toluene was prepared and placed in a glass bubbler. The addition of ethanol aimed to introduce oxygen-rich functional groups (–OH, –C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, –COO) onto the surface of the nanospheres during synthesis, thereby enhancing their functionalization potential and improving their hydrophilicity. Argon was supplied at 1 L min−1 as a carrier gas to transport the vaporized precursor into a quartz reaction tube positioned inside a tubular furnace preheated to 1100 °C. For each batch, 15 mL of the precursor solution were placed into the bubbler, and the synthesis was carried out during 15 min. Upon completion, precursor supply was interrupted, and the furnace was allowed to cool down to ambient temperature under continuous argon flow (0.2 L min−1).
O, –COO) onto the surface of the nanospheres during synthesis, thereby enhancing their functionalization potential and improving their hydrophilicity. Argon was supplied at 1 L min−1 as a carrier gas to transport the vaporized precursor into a quartz reaction tube positioned inside a tubular furnace preheated to 1100 °C. For each batch, 15 mL of the precursor solution were placed into the bubbler, and the synthesis was carried out during 15 min. Upon completion, precursor supply was interrupted, and the furnace was allowed to cool down to ambient temperature under continuous argon flow (0.2 L min−1).
The light black matte deposit of CNS was collected from the deposition zone located outside the heating region. Each run yielded ∼100 mg of material.
2.3. CUR loading
Ten milligrams of carbon nanospheres were dispersed in 15 mL of absolute ethanol by ultrasonication (10 min). CUR (50 mg) and one drop of polysorbate 80 were added, the mixture was stirred at 1000 rpm for 24 h at 25 °C in the absence of light. The suspension was centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 30 min. The supernatant was retained for quantification of non-attached CUR, and the pellet was frozen at −80 °C (Thermo Scientific Ultra-freezer) for 24 h before lyophilization (Labcono 6 L freeze dryer).34 Initial and final weight measurements were conducted to assess the loading efficiency (LE%) and was calculated as:
000 rpm for 30 min. The supernatant was retained for quantification of non-attached CUR, and the pellet was frozen at −80 °C (Thermo Scientific Ultra-freezer) for 24 h before lyophilization (Labcono 6 L freeze dryer).34 Initial and final weight measurements were conducted to assess the loading efficiency (LE%) and was calculated as:
 | (1) |
Loading capacity (LC%) was calculated as:
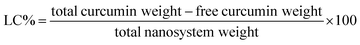 | (2) |
2.4. PEGylation
A non-covalent PEGylation was performed by dispersing 10 mg of CUR-loaded CNS in 15 mL of ultrapure water using ultrasonication (10 min). PEG methyl ether (Mn = 750; 10 mg) and one drop of polysorbate 80 were added. The dispersion was stirred at 1000 rpm for 8 h at 25 °C. Following centrifugation (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm, 30 min). The pellet was frozen at −80 °C and lyophilized.35 Initial and final weight measurements were conducted to assess the functionalization efficiency (FE%) and was determined as:
000 rpm, 30 min). The pellet was frozen at −80 °C and lyophilized.35 Initial and final weight measurements were conducted to assess the functionalization efficiency (FE%) and was determined as:
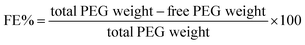 | (3) |
Loading capacity (LC%) was calculated as:
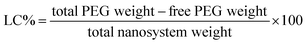 | (4) |
2.5. FA conjugation
FA was conjugated following a modified protocol described by Jagiełło et al.36 Briefly, an aqueous solution of 0.3 µg mL−1 EDC in hydrochloric acid and separate solution of 0.25 µg mL−1 NHS in ultrapure water were prepared. Equal volumes of these two solutions were combined to obtain the activating mixture, which was then added to the previously PEGylated CNS. The suspension was maintained under continuous stirring for 24 h at room temperature. Subsequently, the product was washed three times with ultrapure water, after which 60 mg of FA were added to the aqueous suspension and stirred for an additional 24 h. The resulting material was again washed three times, frozen at −80 °C, and lyophilized. The amount of FA conjugated to the CNS was determined gravimetrically according to the following equation:
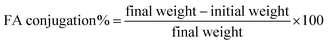 | (5) |
2.6. Scanning electron microscopy (SEM)
Samples were mounted on aluminum stubs using double-sided carbon tape. Micrographs were obtained with a Maia 3 Tescan SEM equipment, particle size distribution was determined using Image J software.2.7. Raman spectroscopy
Raman spectra were recorded using a Horiba XploRa microRaman spectrometer in backscattering mode, with a 532 nm excitation laser, 600 g mm−1 grating, and a 100× objective lens. Laser power was minimized to prevent CUR photodegradation.2.8. Fourier transform infrared (FTIR) spectroscopy
FTIR spectra were collected using an Agilent Cary 630 spectrometer equipped with a diamond attenuated total reflectance (ATR) (1 mm sampling area). The ATR crystal was cleaned with isopropyl alcohol before each measurement.2.9. Zeta potential measurements
Zeta potential (ζ) was measured at 25 °C using a Nanotrac Wave II analyzer (Microtrac) at a concentration of 0.1 mg mL−1 in ultrapure water.2.10. UV-vis CUR quantification
The amount of CUR in the samples was quantified using ultraviolet-visible (UV-vis) spectrophotometry with a Multiskan Sky_1530-800220C plate reader. Prior to sample analysis, a calibration curve of CUR standard solutions in ethanol was prepared at concentrations ranging from 10 to 100 µg mL−1. The calibration curve was evaluated for linearity (R2 ≥ 0.999), repeatability, accuracy, and specificity.2.11. Thermogravimetric analysis (TGA)
TGA was performed with a STA 2500 Regulus thermobalance (Netzsch) under N2 flow. Approximately 10 mg of sample were heated from 25 °C to 600 °C at a rate of 10 °C min−1 in alumina crucibles.2.12. X-ray diffraction (XRD)
XRD patterns were acquired using a Panalytical Empyrean DY1450 equipment (Cu Kα radiation, λ = 1.5406 Å) under standard scan parameters and a BRUKER D8-ADVANCE diffractometer equipped with a LynxEye Detector (Cu Kα, λ = 1.5406 Å) operated at 40 kV and 40 mA.2.13. Release profile
Dialysis bags were loaded with 10 mg of functionalized CNS (equivalent to 250 µg mL−1 CUR). Bags were immersed in 50 mL of phosphate-buffered saline (PBS, pH 7.4 or 5.8) under constant agitation. Aliquots (1 mL) were withdrawn at 0.5, 1, 2, 4, 6, 8, 24 and 48 h, and replaced with equal volumes of PBS. CUR concentration was measured at 421 nm by UV-vis spectroscopy.3. Results and discussion
SEM was used to evaluate the morphology of the pristine and functionalized nanomaterials, as well as to determine the particle size distribution of each formulation. SEM images (Fig. 2A) illustrate the CNS synthesized via APCVD, they exhibit average particle sizes of ∼100 nm and ∼200 nm. | ||
| Fig. 2 SEM micrographs. (A) Pristine CNS, (B) CUR-loaded CNS, (C) PEGylated CNS and (D) FA-conjugated NCS (DDS). | ||
All nanoparticles remain below the 500 nm threshold, which is considered optimal for DDS.37 Functionalized formulations (Fig. 2B–D) showed no significant changes in morphology, maintaining spherical shape and similar dimensions (125–130 nm).
SEM results confirm that APCVD produces CNS within the optical nanometric range for tumor targeting via the enhanced permeability and retention effect in tumor vasculature. This aligns with reports from Li et al. where particles under 1200 nm showed superior tumor accumulation and penetration depth compared to larger counterparts.38 The preservation of morphology after CUR, PEG and FA conjugation indicates that the functionalization process does not compromise structural integrity.
Dynamic light scattering (DLS) analysis was conducted to measure the hydrodynamic diameter and zeta potential of the CNS before and after functionalization, enabling the assessment of surface charge, colloidal stability and dispersion quality of the DDS. DLS revealed that the pristine CNS exhibits an average hydrodynamic diameter of ∼200 nm, with narrow size distribution reflected by polydispersity index (PDI) of 0.056 and zeta potential of −31.66 mV. These values indicate a stable colloidal suspension and suggest the presence of oxygen-rich functional groups introduced through ethanol incorporation during synthesis. In contrast, CUR loaded CNS and PEGylated CNS displayed an increased hydrodynamic diameter ∼290 nm and ∼314 nm, a PDI of 0.221 and 0.215 respectively. CNS-CUR shows a zeta potential of −25.0 mV, meanwhile CNS-CUR-PEG exhibited a zeta potential of −27.7 mV.33 This shift is consistent with surface modification by PEG, whose hydroxyl groups contribute to a less negative surface charge. The addition of PEG not only altered colloidal stability but also enhanced dispersion and hydrophilicity, properties that are highly advantageous for biomedical applications by improving membrane permeability and extending systemic circulation time.37 Finally, the FA-functionalized CNS system exhibited a mean diameter of 321 nm, a PDI of 0.6 and a zeta potential of −30.6 mV. Although a slight variation in charge was observed, the minimal size difference relative to PEGylated CNS can be attributed to the small molecular nature of FA, with is unlikely to induce substantial changes in hydrodynamic size. Importantly, the DDS preserved a net negative surface charge throughout the functionalization process, ensuring colloidal stability and compatibility with biological environments.33
The LE% of CUR in CNS was determined to be 63.63 ± 1.61%, with a corresponding LC% of 72.92 ± 13.22%. In relation to the PEGylation process, the FE% reached 91.3 ± 3.41%, while the associated LC% was calculated as 46.56 ± 1.95%. Consistent with the adsorption mechanisms reported for carbon nano systems, the adsorption efficiencies are comparable or higher, primarily due to π–π stacking interactions between aromatic structures and the graphitic domains of CNS.39 On the other hand, the efficiency of FA conjugation was 22.6%.
XRD was employed to examine the crystalline structure of the precursor materials and to identify changes in characteristic diffraction peaks resulting from the incorporation of CUR, PEG and FA onto the CNS surface. XRD pattern of pristine CNS in the region depicted (Fig. 3 black line), displayed one diffraction peak at ∼25° which can be assigned to the (002) lattice plane. Although the presence of this band unequivocally points to the presence of sp2 hybridization, the width of the feature (being a broad band) indicates small crystallite sizes and a low degree of graphitization, as discussed by Patiño-Guillén and co-authors.33
According to the literature, X-ray diffraction measurement of crystalline PEG should present two intense diffraction peaks ∼19° and ∼23°, instead, our PEG sample displays a broad diffraction peak in the depicted area, centered slightly above 20° (orange line in Fig. 3), this is probably due to the low average molecular weight (Mn 750) of our sample, compared to the high Mn in ref. 32 and 33 (6000 and 8000, respectively).40,41 Such low Mn hinders the ability of the polymer to crystallize; hence the diffraction pattern results in a broad line typical of small crystallite sizes. CUR exhibits distinct diffraction peaks at 2θ values of ∼17°, ∼18°, ∼24°, ∼26° and ∼27°.
The positions of these reflections correspond to characteristic diffraction planes of CUR crystalline lattice, associated with molecular features such as aromatic rings and keto–enol functional groups, which contribute to the compound's high crystallinity. Upon incorporation into nano systems, these peaks tend to decrease in intensity or even disappear, indicating a reduction in crystallinity. This phenomenon is particularly relevant for drug delivery applications, as more amorphous state often enhances dissolution and controlled release behavior. In the diffraction patterns of CNS-CUR-PEG-FA, only the reflection near 18° is clearly observed, suggestion a disruption of crystalline structure of CUR. Similar behavior has been reported by Winijiani et al., who observed vanishing sharp and intense diffraction peaks of pure CUR after incorporation into micellar systems.42–44 FA diffraction peaks were observed at ∼10°, ∼13°, ∼16° and ∼22°, being the feature around 10° the most intense (Fig. 3 dark yellow line). These peaks correspond to the ordered arrangement of pterin and p-aminobenzoic acid moieties with a relatively wide interplanar planes. They constitute the primary structural components of the folate molecule. In CNS-CUR-PEG-FA the above listed features are present in the diffractogram (violet line in Fig. 3), indicating the successful incorporation of FA in our system. Another noteworthy feature in the XRD spectrum of CNS-CUR-PEG-FA is a low intensity bump in the region of the (002) graphitic plane, confirming the presence of CNS.
Hence, we can conclude by our XRD analysis that the conjugated system possesses features of CNS, CUR and FA. Although no feature in the conjugated system can be related to the presence of PEG, its presence is evidenced by Raman spectroscopy and TGA.
Raman and FTIR analyses were carried out to verify the preservation of functional groups from each precursor material following every modification step, as well as to elucidate the type of interactions occurring on the CNS surface of the pristine CNS after functionalization. FTIR spectra (Fig. 4) of pristine CNS showed negative (inverted) bands in the region between 1900 and 2300 cm−1, corresponding to CO2 asymmetric stretching modes. Air was used as blank for the ATR-FTIR measurements, so the inverted bands indicate that the solid CNS contain less CO2 than the corresponding background. A slight band appears around 2700 cm−1, characteristic of O–H vibrations (alcohol groups).33 CUR spectrum exhibited characteristic vibrations; phenolic O–H (∼3500 cm−1), aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C (∼1620 cm−1), benzene ring (∼1590 cm−1) and C–O–C (∼1000 cm−1).42 PEG spectra displayed alkane C–H (∼2880 cm−1), O–H stretching (∼3400 cm−1), C
C (∼1620 cm−1), benzene ring (∼1590 cm−1) and C–O–C (∼1000 cm−1).42 PEG spectra displayed alkane C–H (∼2880 cm−1), O–H stretching (∼3400 cm−1), C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (∼1700 cm−1) and C
O (∼1700 cm−1) and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C (∼1460 cm−1).43 FA spectra shows C
C (∼1460 cm−1).43 FA spectra shows C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching of carboxylic groups (∼1670 cm−1), aromatic C
O stretching of carboxylic groups (∼1670 cm−1), aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching (∼1600 cm−1), C–N bending (∼1400 cm−1) and stretching from O–H carboxylic acid groups (∼3400–3600 cm−1).44 The CNS-CUR-PEG spectrum incorporated the characteristic peaks of both CUR and PEG, confirming successful conjugation. Collectively, the FTIR spectra confirms the progressive incorporation CUR, PEG and FA onto the CNS surface, with characteristic peaks of each component preserved in the hybrid systems.
C stretching (∼1600 cm−1), C–N bending (∼1400 cm−1) and stretching from O–H carboxylic acid groups (∼3400–3600 cm−1).44 The CNS-CUR-PEG spectrum incorporated the characteristic peaks of both CUR and PEG, confirming successful conjugation. Collectively, the FTIR spectra confirms the progressive incorporation CUR, PEG and FA onto the CNS surface, with characteristic peaks of each component preserved in the hybrid systems.
 | ||
| Fig. 4 FT-IR spectra. CNS: carbon nanospheres, CUR: curcumin, PEG: polyethylene glycol and FA: folic acid. | ||
Raman spectra of pristine CNS (Fig. 5 black line) exhibited features at ∼1580 cm−1 and ∼1350 cm−1, these bands are typical of graphitic materials possessing sp2 hybridization (G band) along with symmetry breaking elements or defects (D bands), respectively.33 CUR showed C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and C
C and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O vibrations (1600–1630 cm−1).45 The modified CNS show the D and G bands, confirming the presence of the carbon nanostructure as core of the formulation. Identifiable bands in the PEG spectra includes C–H stretching of methylene groups (∼2880–2940 cm−1), CH2 bending (∼1440–1460 cm−1) and the C–O–C stretching band (∼1090–1120 cm−1).
O vibrations (1600–1630 cm−1).45 The modified CNS show the D and G bands, confirming the presence of the carbon nanostructure as core of the formulation. Identifiable bands in the PEG spectra includes C–H stretching of methylene groups (∼2880–2940 cm−1), CH2 bending (∼1440–1460 cm−1) and the C–O–C stretching band (∼1090–1120 cm−1).
 | ||
| Fig. 5 Raman spectra of the synthesized systems. CNS: carbon nanospheres, CUR: curcumin, PEG: polyethylene glycol and FA: folic acid. | ||
The systems with PEG (CNS-CUR-PEG, CNS-PEG, CNS-CUR-PEG-FA) exhibit a low intensity broad band just below 3000 cm−1 which can be attributed to the main feature in the PEG spectrum corresponding to the CH2 stretching, confirming PEGylation of the nanosystem.46 FA Raman spectrum shows aromatic stretching and C–N bending (∼1600 cm−1), a C–H rocking vibration (∼1385–1400 cm−1) and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching (∼700 cm−1).47,48
C stretching (∼700 cm−1).47,48
FTIR and Raman analyses provide direct evidence of CUR, PEG and FA incorporation. The coexistence of their characteristic vibrations along with those of CNS indicates π–π stacking and hydrogen bonding mechanisms also described for phenolic compounds, graphene and CNS.10 Such non-covalent interactions are advantageous as they preserve CUR bioactivity while ensuring stable loading.
TGA was used to determine the thermal stability of each nanosystem after successive functionalization stages and to evaluate whether new thermally stable interactions or bonds were formed, supporting its suitability as a DDS. TGA profile (Fig. 6) of CNS showed minimal weight loss, ∼5%, we intentionally display its TGA curve from 70% to 100% to perceive the small mass decrease for this sample, confirming the stability of the nanospheres due to their high carbon content.49 CUR degradation evidences a major mass loss at ∼250–350 °C, loosing around 70% of mass in this temperature range. PEG degradation occurred mainly in the range 350–400 °C and the loss of mass was ∼60%. FA TGA profile exhibited an initial loss close to 100 °C that can be attributed to water evaporation. A subsequent weight reduction ∼125–150 °C corresponds to the glutamic acid moiety, followed by a further loss around 250 °C associated to pterin and p-aminobenzoic acid components, from this temperature a constant mass loss is observed, remaining only 25% of the total mass at the end of the thermal treatment. For the CNS-CUR system, a ∼10% mass loss around 100 °C was evidenced (water evaporation), at higher temperatures, around 300 °C (showing the degrading behaviour of CUR) a mass loss of ∼50% was observed. This mass loss is in excellent agreement with the amount of CUR loaded to our system (LE% = 63.63%).
The π–π stacking interactions between CUR and CNS surface are relatively weak, facilitating gradual drug desorption and release. Similarly, the CNS-PEG formulation exhibited a mass loss at the degradation temperature shown for PEG: 350–400 °C, however, the mass loss was around 40% not aligned with its FE% (91%), suggesting higher stability, which can be an indicator of the strong PEG and CNS attachment. CNS-CUR-PEG system reveals a slight continuous mass loss (∼10%), then around 350–400 °C a second mass reduction can be noticed, which is also minimal (∼10%), at this range PEG shows its degradation, confirming the presence of PEG but evidencing higher stability of the formulation since we do not observe the CUR-related mass loss in the range 250–350 °C. TGA data show that the functionalized CNS-CUR-PEG-FA system degrades much more slowly and is largely more stable. We do not find the CUR-related degradation (250–350 °C), nor the PEG-related mass loss (350–400 °C). A steady minimal mass loss (∼10%) is evidenced up to a temperature of 525 °C, where a 10% mass reduction is registered. This high temperature degradation was also observed for FA, this is direct evidence of the anchoring of FA molecules at the surface of our system. The thermal stability behaviour may be attributed to the covalent coupling of FA via EDC-NHS chemistry, which constitutes a stable bond, contributing to a more robust nanostructure less prone to thermal degradation of the internal components (CUR and PEG). Thermal stability is crucial for long-term storage, which is an important characteristic in DDS, validating thus this potential application for our system.
In vitro release study was conducted in phosphate-buffered saline (PBS) to assess the drug-release behavior of the system under physiologically relevant pH conditions. Table 1 summarizes R2 values obtained for each release profile analysis under the evaluated conditions of PBS at pH 7.4 (physiological) and PBS at 5.3 (lysosomal). The corresponding graphs of the four different models used to fit the data can be found in Fig. S1 in the SI. Among the kinetic models tested, the Korsmeyer–Peppas model provided the best fit, which is typically associated with polymeric systems but is also applicable to long controlled-release mechanism exhibiting Fickian diffusion.50 This finding is consistent with previous reports for non-mesoporous carbon-based materials with surface chemistries comparable to the CNS synthesized in this work, which also followed the Korsmeyer–Peppas model.51,52 Furthermore, under PBS conditions at both pH 5.3 and 7.4, the release of CUR (∼4.3% and ∼3.3% cumulative percentages, respectively) indicates a slow and sustained release profile (Fig. 7).
This sustained release behavior can be partially attributed to the presence of the PEG layer on the CNS surface. PEGylation introduces a hydrophilic steric barrier that stabilizes the nanosystem and limits premature desorption of CUR from the carbon surface, thereby reducing burst release and favoring a diffusion-controlled release mechanism. The hydrated PEG chains create a physical barrier that hinders direct contact between the adsorbed drug and the surrounding medium, which has been widely reported to slow the release of hydrophobic molecules from nanocarriers in aqueous medium.53–55 Moreover, the strong π–π stacking and hydrophobic interactions between CUR phenolic groups and CNS further contribute to drug retention, a phenomenon widely described for hydrophobic molecules loaded onto carbon nanomaterials surfaces such as graphene oxide and carbon nanotubes.56
While the cumulative amount of curcumin released after 48 h (∼0.5 µM) is below the IC50 values commonly reported for free curcumin in standard in vitro cytotoxicity assays, the present system was not designed for rapid drug dumping or acute cytotoxic exposure. Instead, the low cumulative release reflects a sustained, diffusion-controlled profile characteristic of adsorption-driven carbon nanocarriers and further modulated by PEGylation. Several studies have demonstrated CUR exerts antitumor effects in breast cancer cell lines at concentrations below 5 µM, with reported effective doses as low as ∼1.32 µM. In this context, the sustained and controlled release profile observed is relevant.57–60 Therefore, increasing the dose of DDS does not aim to promote rapid drug release, but rather to ensure prolonged maintenance of CUR at therapeutically relevant concentrations. Moreover, folic-acid functionalization is expected to enhance cellular uptake via folate receptor-mediated internalization, potentially increasing the effective intracellular drug concentration beyond that measured in the release medium. Therefore, the observed release profile is consistent with a controlled, long-term delivery strategy rather than a high-dose cytotoxic burst. Future studies will focus on evaluating the biological response of cancer cells to the DDS, including cellular uptake and intracellular drug accumulation, to better correlate release kinetics with therapeutic efficacy.
It is important to consider that these assays were conducted under physiological-mimicking conditions. In this fact, the system also exhibited clearly pH-dependent behavior, with greater diffusion of CUR under acidic conditions, consistent with previous reports in carbon nanomaterials in which CUR release is enhanced at low pH following adsorption onto carbon nanotubes surface.32,61 This enhanced release at acidic pH is especially relevant for cancer therapy, as tumor microenvironments typically exhibit pH values between 5.3 and 6.5.62 Collectively, these results indicate that the synthesized DDS, combine favorable physicochemical properties (optimal size, structural stability and responsive release with formulations adaptability) making them promising candidates for sustained, pH-sensitive cancer drug delivery. Acidic conditions favored the stability of CUR, supporting the potential of this DDS as a pH-responsive, prolonged-release platform for biomedical applications.
Several studies have shown that PEGylation enhances the biocompatibility and colloidal stability of nanosystems by preserving the structural integrity and biological activity of sensitive molecules, imparting antioxidant properties, and reducing nonspecific interactions with cell membranes. This behavior aligns with our observations in the CNS-CUR-PEG-FA system, where PEGylation improved nanoparticle dispersion, increased structural stability and maintained the spectroscopic and thermal signatures of CUR. Consistent with reports on hemoglobin-encapsulated PEG nanoparticles (where PEG preserves biomolecular functionality) our formulation exhibited a slow and sustained drug release profile, this limited diffusion indicates that the PEG layer acts a hydrophilic barrier that restricts CUR desorption, imparting characteristic diffusional control. The higher release observed under acidic conditions is also in agreement with studies reporting improved permeability of PEGylated nanoplatforms in acidified tumor microenvironment, further supporting the therapeutic potential of this system for targeted, prolonged DDS.63
4. Conclusions
This work reports the successful synthesis and characterization of a controlled DDS based on CNS obtained via APCV. The optimized synthesis yielded nanostructures with an average particle size of ∼150 nm, a range particularly favorable for biomedical applications due to its potential to enhance tumor tissue penetration. Comprehensive characterization of the functionalized CNS confirmed the efficient incorporation of CUR and its stable interaction with the CNS, loading efficiency of 63% and sustained release <5% after 48 h pH 5.3, which means its release profile fits to a Korsmeyer–Peppas profile (R2 = 0.9053), optimal for controlled and prolonged DDS, besides a pH responsive system was proved. Additionally, PEGylation reached a functionalization efficiency of 91.3% and a FA conjugation of 22.6%. Future studies should focus on the antitumor efficacy, biocompatible profile, stability, and sterilization protocols of the proposed DDS to validate its in vitro application and facilitate its eventual use in a clinical setting.Author contributions
Luis de Sales: experiments, data analysis, writing original draft; Sergio Bernal: supervision, draft review; Jessica Campos-Delgado: conceptualization, supervision, data analysis, draft review and edition.Conflicts of interest
There are no conflicts to declare.Data availability
Data is available upon request.Supplementary information (SI): the graphs of kinetic models fitting for drug release. See DOI: https://doi.org/10.1039/d5ra08562d.
Acknowledgements
L. de S. thanks CONAHCYT for graduate studies scholarship. We thank Dr Kolosovas and LANAFQB from CIACYT, UASLP, for XRD measurements. In addition, we thank Dr Herrera from Facultad de Farmacia, UAEM for XRD measurements.Notes and references
- J. Ferlay, M. Ervik, F. Lam, M. Laversanne, M. Colombet, L. Mery, A. Znaor, I. Soerjomataram and F. Bray, Cancer Today, International Agency for Research on Cancer, 2022 Search PubMed.
- C. E. P. Smith and V. Prasad, Am. Fam. Physician, 2021, 103, 155–163 Search PubMed.
- B. Hosnedlova, M. Kepinska, C. Fernandez, Q. Peng, B. Ruttkay-Nedecky, H. Milnerowicz and R. Kizek, Chem. Rec., 2019, 19, 502–522 CrossRef CAS PubMed.
- S. Bayda, M. Adeel, T. Tuccinardi, M. Cordani and F. Rizzolio, Molecules, 2019, 25, 112 CrossRef PubMed.
- J. K. Patra, G. Das, L. F. Fraceto, E. V. R. Campos, M. D. P. Rodriguez-Torres, L. S. Acosta-Torres, L. A. Diaz-Torres, R. Grillo, M. K. Swamy, S. Sharma, S. Habtemariam and H. S. Shin, J. Nanobiotechnol., 2018, 16(1), 71 CrossRef PubMed.
- A. M. Monaco and M. Giugliano, Beilstein J. Nanotechnol., 2014, 5, 1849–1863 CrossRef PubMed.
- E. M. Alosime, Discover Nano, 2023, 18, 12 CrossRef CAS PubMed.
- W. Liu and G. Speranza, C, 2019, 5, 72 CAS.
- R. Li, Z. Bao, P. Wang, Y. Deng, J. Fan, X. Zhu, X. Xia, Y. Song, H. Yao and D. Li, Polymers, 2023, 15, 3333 CrossRef CAS PubMed.
- L. Wang, X. Xu, L. Chu, C. Meng, L. Xu, Y. Wang, Q. Jiao, T. Huang, Y. Zhao, X. Liu, J. Li, B. Zhou and T. Wang, Biomed. Pharmacother., 2023, 168, 115836 CrossRef CAS PubMed.
- A. Masotti, M. R. Miller, A. Celluzzi, L. Rose, F. Micciulla, P. W. F. Hadoke, S. Bellucci and A. Caporali, Nanomedicine, 2016, 12, 1511 CrossRef CAS PubMed.
- H. Zare, S. Ahmadi, A. Ghasemi, M. Ghanbari, N. Rabiee, M. Bagherzadeh, M. Karimi, T. J. Webster, M. R. Hamblin and E. Mostafavi, Int. J. Nanomed., 2021, 16, 1681 CrossRef PubMed.
- L. Camacho-Barquero, I. Villegas, J. M. Sánchez-Calvo, E. Talero, S. Sánchez-Fidalgo, V. Motilva and C. Alarcón de la Lastra, Int. Immunopharmacol., 2007, 7, 333–342 CrossRef CAS PubMed.
- H. Ni, W. Jin, T. Zhu, J. Wang, B. Yuan, J. Jiang, W. Liang and Z. Ma, J. Spinal Cord Med., 2015, 38, 199 CrossRef PubMed.
- J. Sharifi-Rad, Y. El Rayess, A. A. Rizk, C. Sadaka, R. Zgheib, W. Zam, S. Sestito, S. Rapposelli, K. Neffe-Skocińska, D. Zielińska, B. Salehi, W. N. Setzer, N. S. Dosoky, Y. Taheri, M. El Beyrouthy, M. Martorell, E. A. Ostrander, H. A. R. Suleria, W. C. Cho, A. Maroyi and N. Martins, Front. Pharmacol., 2020, 11, 550909 Search PubMed.
- M. Sathyabhama, L. C. Priya Dharshini, A. Karthikeyan, S. Kalaiselvi and T. Min, Biomolecules, 2022, 12, 1405 CrossRef CAS PubMed.
- R. R. Kotha and D. L. Luthria, Molecules, 2019, 24, 2930 CrossRef CAS PubMed.
- F. Silvestre, C. Santos, V. Silva, A. Ombredane, W. Pinheiro, L. Andrade, M. Garcia, T. Pacheco, G. Joanitti, G. Luz and M. Carneiro, Pharmaceuticals, 2023, 16, 943 CrossRef CAS PubMed.
- M. Heger, R. F. van Golen, M. Broekgaarden and M. C. Michel, Pharmacol. Rev., 2014, 66, 222–307 CrossRef PubMed.
- M. Gera, N. Sharma, M. Ghosh, D. L. Huynh and S. Jin, Oncotarget, 2017, 8, 66680–66698 CrossRef PubMed.
- Y. Chen, Y. Lu, R. J. Lee and G. Xiang, Int. J. Nanomed., 2020, 15, 3099 CrossRef CAS PubMed.
- D. L. Pouliquen, K. G. Trošelj and R. J. Anto, Pharmaceutics, 2023, 15, 1612 CrossRef CAS PubMed.
- Z. Song, R. Feng, M. Sun, C. Guo, Y. Gao, L. Li and G. Zhai, J. Colloid Interface Sci., 2011, 354, 116–123 CrossRef CAS PubMed.
- M. Rezayi, P. Mahmoodi, H. Langari, B. Behnam and A. Sahebkar, Curr. Med. Chem., 2019, 27, 6849–6863 CrossRef PubMed.
- D. J. O'Shannessy, E. B. Somers, J. Maltzman, R. Smale and Y. S. Fu, SpringerPlus, 2012, 1, 1–9 Search PubMed.
- A. Granja, C. Nunes, C. T. Sousa and S. Reis, Biomed. Pharmacother., 2022, 154, 113525 CrossRef CAS PubMed.
- N. Norton, B. Youssef, D. W. Hillman, A. Nassar, X. J. Geiger, B. M. Necela, H. Liu, K. J. Ruddy, M. Y. C. Polley, J. N. Ingle, F. J. Couch, E. A. Perez, M. C. Liu, J. M. Carter, R. A. Leon-Ferre, J. C. Boughey, E. B. Somers, K. R. Kalari, D. W. Visscher, M. P. Goetz and K. L. Knutson, npj Breast Cancer, 2020, 6, 4 CrossRef CAS PubMed.
- S. W. Kim and D. Khang, Int. J. Nanomed., 2015, 10, 3989–4008 CAS.
- H. Sharma and S. Mondal, Int. J. Mol. Sci., 2020, 21, 6280 CrossRef CAS PubMed.
- M. Salem, Y. Xia, A. Allan, S. Rohani and E. R. Gillies, RSC Adv., 2015, 5, 37521–37532 RSC.
- P. Das, S. Ganguly, T. Agarwal, P. Maity, S. Ghosh, S. Choudhary, S. Gangopadhyay, T. K. Maiti, S. Dhara, S. Banerjee and N. C. Das, Mater. Chem. Phys., 2019, 237, 121860 CrossRef CAS.
- A. Chattaraj, Y. Mishra, A. A. A. Aljabali and V. Mishra, Carbon Trends, 2025, 19, 100464 CrossRef CAS.
- G. Patiño-Guillén, A. Arceta-Lozano, J. A. Falcón-Montes, E. García-Díaz, J. N. Díaz De León, R. Vazquez-Duhalt, G. Gao, M. Méndez-Rojas and J. Campos-Delgado, Nanotechnology, 2021, 32(8), 085602 CrossRef PubMed.
- M. Roopesh, D. Davis, M. S. Jyothi, M. Vandana, B. S. Thippeswamy, G. Hegde, T. P. Vinod and R. S. Keri, RSC Adv., 2023, 13, 24320 RSC.
- N. Hadidi, F. Kobarfard, N. Nafissi-Varcheh and R. Aboofazeli, Int. J. Nanomed., 2011, 6, 737 CAS.
- J. Jagiełło, M. Kuśmierz, E. Kijeńska-Gawrońska, M. Winkowska-Struzik, W. Święszkowski and L. Lipińska, Mater. Today Commun., 2021, 26, 102056 CrossRef.
- S. Poh, V. Chelvam and P. S. Low, Nanomedicine, 2015, 10, 1439 CrossRef CAS PubMed.
- H. Wang, X. Li, Z. Ma, D. Wang, L. Wang, J. Zhan, L. She and F. Yang, Int. J. Nanomed., 2016, 11, 1793 CAS.
- F. Sahne, M. Mohammadi and G. D. Najafpour, ACS Biomater. Sci. Eng., 2019, 5, 2595–2609 CrossRef CAS PubMed.
- X. Fu, W. Kong, Y. Zhang, L. Jiang, J. Wang and J. Lei, RSC Adv., 2015, 5, 68881–68889 RSC.
- M. K. Barron, T. J. Young, K. P. Johnston and R. O. Williams, AAPS PharmSciTech, 2003, 4(2), E12 CrossRef PubMed.
- H. M. Asif, F. Zafar, K. Ahmad, A. Iqbal, G. Shaheen, K. A. Ansari, S. Rana, R. Zahid and S. Ghaffar, Sci. Rep., 2023, 13, 10274 CrossRef CAS PubMed.
- S. Sundararajan, A. Kumar, B. C. Chakraborty, A. B. Samui and P. S. Kulkarni, Sustainable Energy Fuels, 2018, 2, 688–697 RSC.
- X. Zhao, J. Zhang, L. Shi, M. Xian, C. Dong and S. Shuang, RSC Adv., 2017, 67, 42159–42167 RSC.
- P. R. K. Mohan, G. Sreelakshmi, C. V. Muraleedharan and R. Joseph, Vib. Spectrosc., 2012, 62, 77–84 CrossRef CAS.
- V. V. Kuzmin, V. S. Novikov, L. Y. Ustynyuk, K. A. Prokhorov, E. A. Sagitova and G. Y. Nikolaeva, J. Mol. Struct., 2020, 1217, 128331 CrossRef CAS.
- M. Trukawka, K. Cendrowski, W. Konicki and E. Mijowska, Appl. Sci., 2020, 10(18), 6465 CrossRef CAS.
- R. A. R. Teixeira, F. R. A. Lima, P. C. Silva, L. A. S. Costa and A. C. Sant'Ana, Spectrochim Acta, Part A, 2019, 223, 117305 CrossRef CAS PubMed.
- P. Karna, M. Ghimire, S. Mishra, S. Karna, P. Karna, M. Ghimire, S. Mishra and S. Karna, OAlib, 2017, 4, 1–7 Search PubMed.
- L. Ahmed, R. Atif, T. S. Eldeen, I. Yahya, A. Omara and M. Eltayeb, Study the Using of Nanoparticles as Drug Delivery System Based on Mathematical Models for Controlled Release, Int. J. Latest Technol. Eng. Manage. Appl. Sci., 2019, 8, 52 Search PubMed.
- R. P. Morais, G. B. Novais, L. S. Sangenito, A. L. S. Santos, R. Priefer, M. Morsink, M. C. Mendonça, E. B. Souto, P. Severino and J. C. Cardoso, Int. J. Mol. Sci., 2020, 21, 4557 CrossRef CAS PubMed.
- Y. Huang, Y. Yuan, Z. Zhou, J. Liang, Z. Chen and G. Li, Appl. Surf. Sci., 2014, 292, 378–386 CrossRef CAS.
- T. Samkange, S. D'Souza, K. Obikeze and A. Dube, J. Pharm. Pharmacol., 2019, 71, 1497–1507 CrossRef CAS PubMed.
- E. Esposito, I. Christoforou, A. Kalatzis, A. Siamidi, M. Vlachou, S. Pispas and N. Pippa, Nanomaterials, 2025, 15, 1762 CrossRef PubMed.
- E. Basavci, E. Demirel, M. Elhassan and Y. Yuksel Durmaz, Eur. Polym. J., 2024, 215, 113211 CrossRef CAS.
- C. Jin, H. Zheng and J. Chen, Pharmaceuticals, 2025, 18, 1245 CrossRef CAS PubMed.
- M. M. Yallapu, M. Jaggi and S. C. Chauhan, Curr. Pharm. Des., 2013, 19, 1994–2010 CAS.
- C. L. A. Passos, R. M. Polinati, C. Ferreira, N. A. N. dos Santos, D. G. V. Lima, J. L. da Silva and E. Fialho, Sci. Rep., 2023, 13(1), 13446 CrossRef CAS PubMed.
- S. Hu, Y. Xu, L. Meng, L. Huang and H. Sun, Exp. Ther. Med., 2018, 16, 1266 Search PubMed.
- K. Zhai, A. Brockmüller, P. Kubatka, M. Shakibaei and D. Büsselberg, Biomolecules, 2020, 10, 1469 CrossRef CAS PubMed.
- A. Mohammadi, M. S. Hosseini, F. Bagheri, H. Safari, Y. Shadfar, A. Sharafi, H. Rezaeejam, A. Aghaei and H. Danafar, Results Chem., 2024, 7, 101370 CrossRef CAS.
- A. Mohammadi, M. S. Hosseini, F. Bagheri, H. Safari, Y. Shadfar, A. Sharafi, H. Rezaeejam, A. Aghaei and H. Danafar, Results Chem., 2024, 7, 101370 CrossRef CAS.
- M. A. Majid, A. Aleem, A. ur R. Khan, M. A. Siddique, M. Z. Manzoor, M. Akhtar, Q. Raza, H. A. Alodaini, M. B. Junaid, M. U. Dad and H. Ullah, Mater. Chem. Phys., 2025, 333, 130309 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |



