Open Access Article
Open Access ArticleHyaluronan like polysaccharide based nanodrugs with enhanced CD44 avidity for image-guided drug delivery to breast cancer
Chia-Wei Yangab,
A. K. M. Atique Ullah ab and
Xuefei Huang
ab and
Xuefei Huang *abc
*abc
aDepartment of Chemistry, Michigan State University, East Lansing, MI 48824, USA
bInstitute for Quantitative Health Science and Engineering, Michigan State University, East Lansing, MI 48824, USA
cDepartment of Biomedical Engineering, Michigan State University, East Lansing, MI 48824, USA. E-mail: huangxu2@msu.edu
First published on 14th January 2026
Abstract
Targeted drug delivery is an exciting strategy to treat cancer and reduce the harmful side effects of chemotherapy to normal tissues. CD44 has been an attractive target for drug delivery due to its expression on many types of tumor cells including breast cancer. To better target the CD44-expressing breast cancer cells, we report the design and synthesis of a new nanodrug, G2-Sal-ICG, comprising a hyaluronan (HA)-like polysaccharide (G2) conjugated with the anticancer drug salinomycin (Sal) and the imaging agent indocyanine green (ICG). This nanodrug targets CD44 receptors and allows for near-infrared fluorescence imaging (NIR-FI) to track drug delivery in real-time. G2-Sal-ICG had significantly higher affinity with CD44+ cancer cells than the corresponding nanodrug of unmodified HA conjugated with Sal and ICG (HA-Sal-ICG). In vitro studies demonstrated that the nanodrug can be released in typical lysosomal conditions, leading to effective cancer cell killing. In vivo experiments with an orthotopic breast cancer model demonstrated the superior tumor-targeting capability of G2-Sal-ICG and its ability to significantly suppress tumor growth. Our findings suggested G2-Sal-ICG is a promising theranostic nanodrug for CD44-targeted therapy, combining efficient drug delivery with real-time non-invasive imaging, thus highlighting its potential for clinical applications in breast cancer treatment.
Introduction
Chemotherapy is one of the main strategies for cancer treatment. Despite significant advances, a major drawback of chemotherapy is the dose limiting side effects of the drugs on normal tissues. To reduce the toxicities, an attractive approach is to target receptors on the tumor cell surface, which can selectively deliver drugs to the tumor site and enhance the therapeutic efficacy.1,2 CD44 is a cell surface receptor that is highly expressed in many cancers, and plays important roles in cancer development and metastasis.3–6 In addition, CD44 has been regarded as a biomarker for the purported cancer stem cells (CSCs),7,8 which are a subset of cancer cells with unique characteristics contributing to tumor heterogeneity and resistance to therapy.9 Therefore, CD44 is an appealing target for selective drug delivery to tumor sites.Multiple strategies have been developed to target CD44 for drug delivery, which include the usage of monoclonal antibodies, peptides, and polysaccharides.10–12 Anti-CD44 monoclonal antibodies reached human clinical trials for treatment of CD44+ cancer.13,14 Unfortunately, serious toxicities with a fatal outcome were observed in these studies. The polysaccharide hyaluronan (HA), the major endogenous ligand for CD44, is an appealing alternative.15,16 With its high compatibility and ready availability, HA has been investigated in a variety of delivery systems by enhancing cellular uptake through CD44-mediated endocytosis.17–20 For effective drug delivery, the binding affinity of the targeting agent to tumor is an important parameter. Recently, we developed a panel of new HA derivatives with their carboxyl groups modified through the four-component Ugi reaction. Ugi reaction is a multicomponent reaction that forms structures with two amide bonds by combining an amine, an aldehyde or ketone, a carboxylic acid, and an isocyanide in a single reaction.21 A lead HA-like polysaccharide named G2 (formed through the reaction of HA, 3-phenyl-1-propylamine, formaldehyde, and cyclohexyl isocyanide) was identified, which has a significantly higher affinity with CD44 receptors than native unmodified HA. However, the potential of G2 in CD44 targeted drug delivery has not yet been explored.
Another important area for development is the method to monitor the drug delivery process. Image-guided drug delivery merges the precision of imaging techniques with the targeted delivery of therapeutic agents.22 By integrating imaging modalities with nanocarrier-based drug delivery systems, researchers can visualize tumor sites non-invasively and monitor drug distribution in real time.23–27 Moreover, image-guided drug delivery can facilitate early determination of treatment efficacy, enabling timely adjustments in therapy.28,29
Herein, we describe the synthesis of a new nanodrug composed of the HA like polysaccharide G2, near infrared fluorophore indocyanine green (ICG)30–34 and a drug molecule salinomycin (G2-Sal-ICG). Salinomycin was chosen due to its remarkable anticancer properties.35 Emerging research highlights salinomycin's ability to kill CSCs,36–39 and to improve the effectiveness of radiation and various chemotherapy drugs.40–42 We found that G2-Sal-ICG nanodrug could significantly enhance the efficacy of salinomycin and reduce the growth of CD44 tumor in vivo in a mouse model of breast cancer. Furthermore, with the incorporation of ICG, the nanodrugs enabled near-infrared fluorescence imaging (NIR-FI) of the drug delivery process.
Results and discussion
Synthesis and characterization of nanodrug G2-Sal-ICG
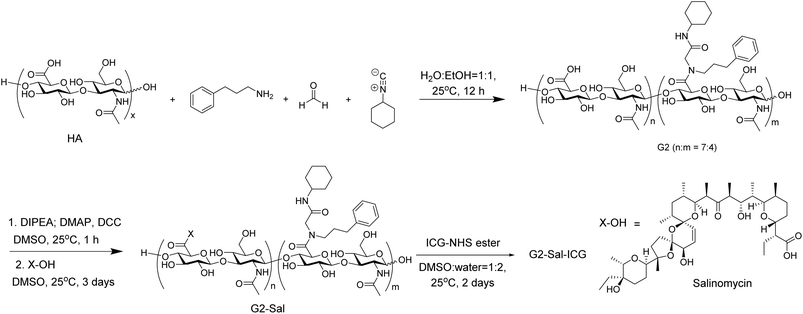
Synthesis of G2-Sal-ICG started from the preparation of the HA derivative G2 through the Ugi reaction (Scheme 1),21 which was formed by reacting HA (16 kDa) with 3-phenyl-1-propylamine, formaldehyde, and cyclohexyl isocyanide.21 The degree of carboxyl group modification in G2 was 36.6% per disaccharide repeating unit as determined by 1H NMR (Fig. S1). G2 was then conjugated to Sal through ester bond formation promoted by N,N-dicyclohexylcarbodiimide (DCC), leading to G2-Sal, which was purified through extensive dialysis. The degree of drug conjugation in G2-Sal was 34% per disaccharide repeating unit, based on analysis of integrations of pertinent signals in 1H NMR spectrum of the conjugate (Fig. S2), indicating around 13.6 drug molecules were conjugated per G2 molecule. In contrast, incubation of G2 and Sal without DCC led to the recovery of G2 supporting the formation of the covalent bonds between G2 and Sal promoted by DCC. To compare the affinity of G2-Sal to CD44, HA-Sal was synthesized using the same method with a similar degree of drug conjugation as G2-Sal (Fig. S3). G2-Sal and HA-Sal were conjugated with ICG32 with a similar amount of the fluorophore per chain, leading to G2-Sal-ICG and HA-Sal-ICG (Scheme 1).Nanodrug formation and characterization
To form the nanodrug, G2-Sal-ICG was dispersed in water at various concentrations, then ultrasonicated at room temperature for 1 h. The critical association concentration of G2-Sal-ICG to form self-assembled nanoparticles was determined to be 0.01 mg mL−1 (Table S1). HA-Sal-ICG nanoparticles were formed in a similar manner. The morphology of G2-Sal-ICG nanoparticles was characterized by scanning electron microscope (SEM), which showed a spherical shape with an average diameter of 265 nm (Fig. 1a). The hydrodynamic size of G2-Sal-ICG nanodrugs was 273 nm (Fig. 1b). No significant size variations were observed for the G2-Sal-ICG nanodrugs in water, PBS or fetal bovine serum containing media suggesting serum stability of these particles in biological media (Fig. S4), which aligns well with other HA based nanodrugs.43–45 In addition, storage stability of the particles was tested. G2-Sal-ICG nanoparticles were lyophilized, stored at 4 °C, and analyzed after 6 months. The particle sizes remained the same over 6 months. This compared well with the one to four month stabilities of lyophilized HA based nanoparticles reported in the literature.46–48In addition of sizes of the G2-Sal-ICG nanodrugs, other characteristics of the particles were measured. The zeta potential of G2-Sal-ICG nanodrugs was −16.7 ± 0.6 mV in PBS with a concentration of 0.5 mg mL−1. The negative surface charge was presumably due to the free carboxyl moieties of HA as the hydrophilic HA is most likely exposed on the external surface to the aqueous environment with the more hydrophobic Sal and ICG in the core of the nanodrugs. The fluorescence emission maximum peak of G2-Sal-ICG was blue-shifted from 810 to 795 nm, compared to free ICG in solution (Fig. 1c). The absorption peak of G2-Sal-ICG was found to be broader than that for ICG. These observations were attributed to the nanoaggregation of ICG with the formation of nanospheres.49,50 UV spectra of serially diluted G2-Sal-ICG nanodrug solutions showed a linear correlation between fluorescence signals and nanodrug concentrations (Fig. 1d), suggesting the potential for nanodrug quantification with NIR-FI. The X-ray diffraction (XRD) analysis showed that the G2-Sal-ICG nanodrug is in amorphous form (Fig. S5). HA-Sal-ICG nanodrug was characterized and found to have similar sizes and zeta potential values as G2-Sal-ICG (Fig. S6 and Table S2).
G2-Sal-ICG could bind with CD44-expressing breast cancer cells
Breast cancer is one of the leading causes of cancer associated death of women.51,52 To develop a new breast cancer treatment, we examined the ability of G2-Sal-ICG nanodrug to target breast cancer cells. CD44-expressing 4T1 breast cancer cells were incubated with G2-Sal-ICG nanodrug and HA-Sal-ICG nanodrug in parallel at 37 °C for 2 h, followed by three PBS washes to remove unbonded nanodrugs. The cells were then examined by flow cytometry (Fig. 2a). G2-Sal-ICG nanodrug treated cancer cells showed a significantly higher median fluorescence intensity (MFI) compared to cells incubated with HA-Sal-ICG nanodrug, indicating the stronger binding of G2-Sal-ICG by cancer cells. To support the role of HA (or G2) in CD44 targeting, 4T1 cells were incubated with a mixture of G2-Sal-ICG nanodrug and free HA. The presence of free HA during incubation significantly reduced the MFI of the cells as compared to the G2-Sal-ICG nanodrug group (Fig. 2a) likely because HA competed with G2-Sal-ICG for cell binding.To better quantify the binding of nanodrug, fluorescence measurements were performed. 4T1 cells were cultured in a 12-well plate overnight, then incubated with G2-Sal-ICG or HA-Sal-ICG nanodrug at 37 °C for 2 h. After thorough washing with PBS, the cells were detached with trypsin with fluorescence intensities measured. The G2-Sal-ICG nanodrug group showed 2.6 times higher fluorescence intensities compared to the HA-Sal-ICG nanodrug group (Fig. 2b). When 4T1 cells were incubated with G2-Sal-ICG nanodrug at 4 °C, the amount of intracellular nanodrug was significantly reduced compared to that at 37 °C (Fig. 2b). This is consistent with the idea that the nanoparticle uptake was through CD44 mediated endocytosis, an energy-dependent process. Incubation of cells with G2-Sal-ICG or HA-Sal-ICG nanodrug for various durations showed that nanodrug accumulation reached its maximum after 8 h (Fig. S7).
To investigate the distribution of nanodrugs in the cells, 4T1 cells were incubated with G2-Sal-ICG nanodrug at 37 °C for 2 h, followed by lysotracker red and nuclear staining with DAPI, and imaging by confocal microscopy (Fig. 2c). The ICG signals were co-localized with lysotracker red signals, indicating the presence of G2-Sal-ICG nanodrug in the lysosome (Fig. 2c, first row). Furthermore, cells incubated with the HA-Sal-ICG nanodrug or the G2-Sal-ICG nanodrug with free HA were imaged following the same protocol using the identical setting (Fig. 2c, second and third row). As shown in Fig. 2c, in the ICG channel, the G2-Sal-ICG nanodrug treated cells had stronger signals compared to the HA-Sal-ICG nanodrug and the G2-Sal-ICG nanodrug/free HA groups. These results supported the stronger binding and higher uptake of G2-Sal-ICG nanodrug by cancer cells compared to HA-Sal-ICG nanodrug as well as the cellular binding dependence on HA.
To understand the rate of drug release from the nanodrug (presumably through hydrolysis of the ester bond linkage53–55), G2-Sal was incubated under various conditions, including pH 7.4, 5.5, and the presence of hyaluronidase (HAdase) at 37 °C to mimic lysosomal conditions. At pH 7.4, after 36 hours, 14% of the cargo Sal was released from the nanodrug formulation. In comparison, lowering the pH to 5.5, a value typical of that in lysosomes,56–58 the amount of drug released in 36 hours was significantly higher at 51% (Fig. S8). A further increase in the drug release percentage was observed with the addition of HAdase, which reached 62% suggesting HAdase could help degrade the nanoparticles. The results also indicated that the G2-Sal is relatively stable in neutral conditions than acidic conditions.
The nanodrugs were cytotoxic against 4T1 cells
To compare the efficacy of nanodrug, 4T1 cells were incubated with G2-Sal, HA-Sal, and salinomycin at various drug concentrations (1 µM and 10 µM) at 37 °C followed by 3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) cell viability assays. At 24 h (Fig. 3a), the viabilities of salinomycin treated groups were about 20% lower compared to the G2-Sal and HA-Sal groups at both 1 and 10 µM concentrations. Interestingly, when cells were incubated for 36 hours, G2-Sal and HA-Sal killed significantly more cancer cells as compared to the salinomycin only group (Fig. 3b). The delay of cell killing observed in both G2-Sal and HA-Sal groups could be because the nanodrugs need to be degraded in lysosomes to release the free drug for cell killing.59 For comparison with CD44+ tumor cells, we performed MTS assays using HC11 mammary epithelial cells, which are low in CD44 expression level. HC11 cells were incubated with various concentrations of G2-Sal-ICG nanodrugs. The results showed that the nanodrugs had no significant impact on cell viability at all concentrations tested up to 450 µM equivalent of drug concentration (Fig. S9).G2-Sal-ICG enabled image-guided drug delivery in an orthotopic breast cancer model
With ICG conjugated, G2-Sal-ICG and HA-Sal-ICG nanodrugs could enable NIR-FI imaging in vivo for image-guided drug delivery. To evaluate the performance of nanodrugs, 4T1 cells (105) were orthotopically inoculated to the mammary gland of female BALB/c mice. The tumor bearing mice were randomized into four groups (n = 5 each), and G2-Sal-ICG (8 mg kg−1 salinomycin), HA-Sal-ICG (8 mg kg−1 salinomycin), salinomycin (8 mg kg−1, the equivalent salinomycin dose as nanodrug group), and PBS were administrated intravenously to mice respectively at day 7 post-tumor inoculation. To detect the accumulation of nanodrugs, the G2-Sal-ICG and HA-Sal-ICG groups were imaged at 1 h, 4 h, and 24 h post injection (Fig. 4). For the G2-Sal-ICG nanodrug treated group, the highest fluorescence signals were observed in tumor area (indicated with white arrow) at 4 h (Fig. 4a and b). In contrast, for the HA-Sal-ICG group, only weak signals were observed in tumor area (indicated with white arrow) during the full imaging period (Fig. 4c), and the average fluorescence signals showed no significant changes during the 24 h observing period (Fig. 4d). The G2-Sal-ICG group showed 3 times stronger fluorescence signals than the HA-Sal-ICG group in tumor area at 4 h post injection, indicating that G2-Sal-ICG accumulated more in tumor sites in vivo.G2-Sal-ICG significantly reduced tumor growth
The tumor volume of the mice in all groups were measured by a caliper (length × width2/2) every three days (Fig. 5a). The G2-Sal-ICG nanodrug treated group showed significantly lower average tumor volume compared to the HA-Sal-ICG, salinomycin, and PBS groups, indicating better suppression of tumor growth by the G2-Sal-ICG nanodrug.To further investigate the effects of nanodrugs, mice from different study groups were sacrificed at 27 days post-inoculation and the tumors were dissected and disaggregated to single cells by triple enzyme digestion methods. The disaggregated cells from various study groups were stained with an anti-CD44 monoclonal antibody then analyzed by flow cytometry. The tumor cells from G2-Sal-ICG nanodrug treated group showed a significantly lower CD44 expression compared to the tumor cells from HA-Sal-ICG nanodrug, salinomycin, and PBS group (Fig. 5b), indicating the G2-Sal-ICG killed more CD44 expressing cancer cells, a selective cell killing.
The tumor from the G2-Sal-ICG treatment group showed lower levels of CD44 expression in immunohistochemistry (IHC) tissue staining compared to HA-Sal-ICG, salinomycin free drug, and PBS control groups (Fig. 6 and S10). These results are consistent with the flow cytometry data, indicating greater CD44+ breast cancer cell killing proportionally by G2-Sal-ICG. This is presumably because G2-Sal-ICG could better target CD44 high breast cancer and suppress the growth of these cells, leading to the lower percentage of CD44 positive cells in residual tumor.
 | ||
| Fig. 6 Expressions of CD44 in dissected tumors from different treatment groups (n = 3). Statistical analysis was performed through one-way ANOVA analysis. **, p < 0.01; ***, p < 0.001. | ||
Discussion
CD44 is a major receptor of HA and has been found to be overexpressed in breast cancer cells.3 HA like polysaccharide G2 was developed, which has a stronger interaction with CD44 compared to HA.21 With this new G2 ligand in hand, we investigated it for the first time as a targeting molecule for image-guided drug delivery toward CD44 expressing breast cancer. For synthesis of CD44 targeting nanodrug, salinomycin was covalently conjugated to G2 through biodegradable ester linkage and can be released from the nanoparticles under conditions similar to those in lysosomes.53–55 We further conjugated ICG to G2-Sal, leading to the possibility of using NIR-FI to monitor the delivery of G2-Sal-ICG nanodrug. ICG is an FDA approved clinical imaging agent30–33,60–62 that increases the translational potential of G2-Sal-ICG nanodrug. The G2-Sal-ICG formed the nanostructure with hydrophobic core in aqueous solution through sonication due to the hydrophobicity of incorporated salinomycin.The incorporation of G2 into the nanodrug led to significant enhancement of tumor binding as G2-Sal-ICG gave 2.8 times better targeting than HA-Sal-ICG based on flow cytometry results. Pretreatment of cells with free HA resulted in partial inhibition of G2-Sal-ICG nanodrug accumulation, which supports the involvement of HA/CD44 receptor-mediated mechanism of nanodrug uptake. Moreover, a lower level of nanodrug accumulation was observed with HA-Sal-ICG in cell-based fluorescence assay, suggesting the stronger binding of G2-Sal-ICG to cancer cells.
Cellular biodistribution of nanodrugs showed the nanodrugs were taken up by 4T1 cancer cells and accumulated in lysosomes. The G2-Sal-ICG group had stronger fluorescence signals in cells as compared to HA-Sal-ICG group. Moreover, the partial inhibition of G2-Sal-ICG nanodrug accumulation was observed in free HA pretreatment group, consistent with the receptor-mediated mechanism of nanodrug uptake.
When evaluated in vitro for its toxicity, the nanodrugs showed delayed killing of cancer cells in the MTS assay. The reason could be the nanodrugs need to be degraded first to release the drug to kill the cancer cells.59 Presumably, nanodrugs were internalized by receptor-mediated endocytosis then transferred into lysosomes, where they were degraded by the acidic pH and hyaluronidase and subsequently released the drug intracellularly.
Anti-cancer drugs commonly suffered from the solubility issue in water due to their high hydrophobicity.63 Salinomycin is originally used in veterinary medicine, has garnered significant attention for its potent anti-cancer properties. However, its clinical translation has been hampered by extremely poor aqueous solubility, posing substantial challenges for formulation and systemic delivery. With its higher affinity to CD44, G2 based drug delivery system could not only be used to overcome the solubility challenge for salinomycin but also possess the ability to target CD44-expressing cancer cells. The labile ester linkage between G2 and salinomycin allowed the drug to be released under the lysosomal condition, potentially reducing the systemic toxicity.
To target cancer precisely, a variety of drug delivery systems have been developed.64,65 However, the typical drug delivery system could not track the drug in vivo. When comparing the targeting efficiency among different nanodrugs, the abilities to monitor the delivery process would be very informative.28,29 In our work, the ICG contained in the nanodrugs enabled non-invasive NIR-FI tracking of drug delivery in vivo, to help confirm that the nanodrugs were targeted to breast tumor in real time.
One limitation to this study is that only CD44 is targeted as a biomarker for tumor. As tumor is heterogeneous,66 targeting a single receptor may not eliminate all tumor cells. We have observed that the residual tumor in G2-Sal-ICG treated mice had lower levels of CD44 expressed. Thus, to further enhance tumor treatment efficacy, it may be important to develop drug delivery systems that can target multiple tumor associated receptors besides CD44.
Conclusions
Salinomycin and ICG conjugated G2-Sal-ICG nanodrug was synthesized for breast cancer image-guided drug delivery. The G2-Sal-ICG nanodrug enabled CD44-expressing cancer cell targeting through the CD44 binding and drug release under lysosomal conditions to kill the cancer cells. Moreover, we demonstrated that G2-Sal-ICG targets 4T1 breast cancer cells in the orthotopic mouse model for NIR-FI and drug delivery, enabling imaging and therapeutics simultaneously. Tumor volume measurements indicated significantly better suppression of tumor growth with G2-Sal-ICG administration. Flow cytometry and histological analyses confirmed a greater reduction of CD44-expressing cancer cells. Our findings demonstrated G2-Sal-ICG as a nanodrug to target CD44 expressing tumor cells, combining efficient drug delivery with real-time imaging capabilities. Thus, G2-Sal-ICG nanodrug is a promising candidate for breast cancer image-guided drug delivery.Materials and methods
Materials
Sodium hyaluronate (16 kDa) was purchased from Lifecore Biomedicals. Dimethyl sulfoxide (DMSO), N,N-dimethylamino pyridine (DMAP), 3-phenyl-1-propylamine, hydrochloric acid, formaldehyde, N,N-diisopropylethylamine (DIPEA), N,N-dicyclohexylcarbodiimide (DCC), sodium carbonate, salinomycin, Dulbecco's phosphate-buffered saline (DPBS), RPMI 1640, fetal bovine serum (FBS), penicillin–streptomycin (Pen–Strep), D-luciferin were purchased from Sigma-Aldrich (Sigma-Aldrich, USA). ATCC-formulated RPMI-1640 cell culture media was purchased from ATCC (American Type Culture Collection, USA). ICG-NHS ester was purchased from Ruixibiotech (Ruixibiotech, China). CellTiter 96 Aqueous One solution was purchased from Promega. DAPI and lysotracker red were purchased from Thermo Fisher Scientific (Thermo Fisher Scientific, USA). The dialysis tube (MWCO: 3.5 kDa), centrifugal filter (MWCO 10 kDa), and centrifugal filter (MWCO 100 kDa) were purchased from MilliporeSigma (MilliporeSigma, USA). Anti-CD44 APC/Cy7 (IM7, BioLegend catalog # 103027) was acquired from BioLegend (BioLegend, USA). 4T1 and HC11 cells were acquired from American Tissue Culture Collection (ATCC, USA).Synthesis of HA derivative: G2
Sodium hyaluronate (100 mg, 16 kDa), MilliQ water (6 mL), and ethanol (4 mL) were mixed and stirred until all solid dissolved. 3-Phenyl-1-propylamine in ethanol (200 µL, 0.25 mmol) was added then hydrochloric acid (1.4 M) solution was added to adjust the pH to 4. Formaldehyde in ethanol (100 µL, 0.125 mmol) and cyclohexyl isocyanide in ethanol (200 µL, 0.25 mmol) were added and the solution was stirred at 25 °C for 1 h. Sodium carbonate solution (2 M) was added to adjust the pH to 12 and the reaction was stirred at 25 °C for 48 h. The reaction mixture was then dialyzed with dialysis tube (MWCO: 3.5 kDa) against MilliQ water for 72 h over 9 times of water change. The recovery yield of G2 is nearly quantitative.Syntheses of G2-Sal and HA-Sal
G2 (100 mg, 0.25 mmol carboxyl groups) and DIPEA (42 µL, 0.25 mmol) were dissolved in anhydrous DMSO (5 mL). DMAP (2 mg) and DCC (16 mg, 0.075 mmol) were added, and the solution was stirred at 25 °C for 1 h. Then, salinomycin (56 mg, 0.075 mmol) was added and the reaction was stirred at 25 °C for 72 h. The solution was dialyzed with a dialysis tube (7.5 kDa cutoff) against MilliQ water for 24 h. To ensure the unreacted reagent in the mixture was completely removed, the reaction mixture was further centrifuged through centrifugal filters (MWCO 10 kDa) with 20% ethanol and water, leading to the G2-salinomycin conjugate (G2-Sal). The recovery yield of G2-Sal is nearly quantitative. The HA and salinomycin conjugate (HA-Sal) was produced following the same synthetic method as G2-Sal. The recovery yield of HA-Sal is nearly quantitative.Synthesis of G2-Sal-ICG and HA-Sal-ICG
G2-Sal (10 mg) was mixed with the ICG-NHS ester (0.06 mg) in water/DMSO (3 mL, 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1), and the mixture was stirred at 25 °C for 24 h in the dark. The resulting mixture was centrifuged with centrifugal filters (MWCO 10 kDa) to remove the unreacted ICG-NHS ester, leading to G2-Sal-ICG. The HA-Sal-ICG was formed following the same synthetic method as G2-Sal-ICG. The products were lyophilized and stored in a 4 °C refrigerator.
1), and the mixture was stirred at 25 °C for 24 h in the dark. The resulting mixture was centrifuged with centrifugal filters (MWCO 10 kDa) to remove the unreacted ICG-NHS ester, leading to G2-Sal-ICG. The HA-Sal-ICG was formed following the same synthetic method as G2-Sal-ICG. The products were lyophilized and stored in a 4 °C refrigerator.
Self-assembly of G2-Sal-ICG and HA-Sal-ICG nanoparticles and characterization
G2-Sal-ICG or HA-Sal-ICG (1 mg) was dissolved in 1 mL of deionized (DI) water then sonicated with a bath sonicator for 1 h at 25 °C to form uniform nanoparticles. The hydrodynamic diameter and zeta potential were measured with a concentration of 0.5 mg mL−1 in PBS by DLS using Zetasizer Nano zs apparatus (Malvern, U.K.). The G2-Sal-ICG and HA-Sal-ICG were imaged with SEM JEOL 7500F (JEOL USA Inc., USA). Absorption and emission of ICG, G2-Sal-ICG and HA-Sal-ICG nanoparticles were measured by a SpectraMax M3 plate reader (Molecular Devices LLC, USA). XRD results were performed with Bruker D8 DaVinci diffractometer (Bruker Corporation, USA) equipped with Cu X-ray radiation operating at 40 kV and 40 mA. Peak intensities were obtained by counting with the Lynxeye detector every 0.02° at sweep rates of 2.4° 2θ min−1. The sample rotated at 5° per minute.Cell culture
RPMI 1640 and ATCC-formulated RPMI-1640 cell culture media were supplemented with 10% FBS and 1% Pen-Strep. 4T1 mouse breast cancer cells were cultured with supplemented RPMI 1640 at 37 °C with 5% CO2. HC11 cells were cultured with supplemented ATCC-formulated RPMI-1640 at 37 °C with 5% CO2. The 4T1 cells and HC11 cells were detached by trypsin-EDTA (0.25%) solution after reaching 90% confluence.Nanodrug cell uptake by flow cytometry & fluorescence reading
To compare cancer cell targeting ability of nanodrugs, 105 of 4T1 cells were cultured in each well of 12-well plate overnight. The cells were incubated with G2-Sal-ICG or HA-Sal-ICG (0.5 mg mL−1 of nanodrug) for 2 h at 37 °C then washed with PBS for three times. For the HA co-treatment group, the cells were incubated with G2-Sal-ICG (0.5 mg mL−1 of nanodrug) and free HA (2 mg mL−1) mixture for 2 h at 37 °C, followed by the same washing steps. For the low temperature group, the cells were incubated with G2-Sal-ICG (0.5 mg mL−1 of nanodrug) for 2 h at 4 °C then followed by the same washing steps. The cells were detached by trypsin-EDTA solution then placed in flow cytometry tubes and stored on ice until flow cytometry analysis. For fluorescence reading, the detached cells were placed in a 96-well plate and the fluorescence intensities were measured by a SpectraMax M3 plate reader with excitation at 700 nm and emission at 820 nm.Kinetics of cellular uptake of nanodrug
To compare the kinetics of nanodrug uptake by cells, the 4T1 cells were cultured in a 12-well plate for overnight. The cells were incubated with G2-Sal-ICG or HA-Sal-ICG (0.5 mg mL−1 of nanodrug) at 37 °C for 1 h, 2 h, 4 h, 8 h, 12 h, 24 h, and 36 h. The cells were then washed with PBS three times and detached by trypsin-EDTA solution. The cell suspensions were placed in a 96-well plate and fluorescence intensities were measured by a SpectraMax M3 plate reader with excitation at 700 nm and emission at 820 nm.In vitro cellular targeting
To perform cellular targeting, 104 of 4T1 cells were cultured in a Lab-Tek chamber slide overnight at 37 °C with 5% CO2. The cells were incubated with G2-Sal-ICG or HA-Sal-ICG (0.5 mg mL−1 of nanodrug) for 2 h at 37 °C with 5% CO2 followed by lysotracker red staining for 1 h. After two washes with PBS, the cells were fixed with 4% paraformaldehyde (PFA) then stained with DAPI for 30 min. The slides were covered with cover slips then sealed with nail paint. The cellular fluorescence images were measured by a Leica THUNDER Imagers (excitation wavelength at 750 nm and emission wavelength at 795 nm) (Leica Microsystems, Germany).Kinetics of drug release from nanodrug in vitro
To estimate the rate of drug release, G2-Sal or HA-Sal nanodrugs were placed in a 5 mL Eppendorf tube and dissolved in PBS (4 mL) under various conditions, including pH 7.4, pH 5.5, pH 7.4 with hyaluronidase (1 mg mL−1, 400–1000 units per mg solid), and pH 5.5 with hyaluronidase. The tubes were placed on a rotator in a 37 °C incubator and 200 µL of the solution was taken out for analysis at 1 h, 2 h, 4 h, 8 h, 12 h, 24 h, and 36 h respectively. The testing solution was filtered with centrifugal filter (MWCO: 10 kDa) to remove the nanodrug or degraded polymers. The released salinomycin in the filtrate was quantified by UV-vis spectrometer through reading the absorption at 280 nm.Cytotoxicity of nanodrugs
To evaluate the cytotoxicity of nanodrugs, 4T1 cells (5000 cells) were placed in each well of 96-well plate and cultured for overnight at 37 °C with 5% CO2. The cells were then incubated with G2-Sal-ICG, HA-Sal-ICG, or salinomycin at the same drug concentration (1 µM, 10 µM). To check the viability of each group, the cells were incubated with CellTiter 96 Aqueous One solution according to the manufacturer's protocol. The absorption values were measured at 490 nm with the SpectraMax M3 plate reader. To evaluate the biocompatibility of the G2-Sal-ICG nanodrug, MTS assays were performed according to the manufacturer's protocol using HC11 cells incubated with G2-Sal-ICG nanodrugs at concentrations of 0.2, 0.4, 0.8, and 1.0 mg mL−1.Orthotopic breast cancer models
All mice were kept in the University Laboratory Animal Resources Facility of Michigan State University. All the experimental procedures and guidelines for animal study were performed under approval of Institutional Animal Care and Use Committee (IACUC) of Michigan State University (Protocol #: 202100095). Female 6 weeks old BALB/c mice were purchased from Charles River Laboratories. To build an orthotopic breast cancer model, 4T1 cells (105) were injected into mammary gland of the mouse under anesthesia.Near-infrared fluorescence imaging in vivo
To acquire in vivo NIR-FI images, the mouse was mounted with tape on a Trilogy Pearl system (LI-COR Biosciences) under anesthesia. The parameter of Trilogy Pearl was set at exposure time: 500 ms; excitation: 785 nm; signal detection: 820 nm. The in vivo NIR-FI images were processed by ImageJ.Therapeutic efficacy
4T1 cells were injected into mammary gland of female BALB/c mice on day 0. The nanodrugs (1 mg mL−1 drug concentration in PBS), salinomycin (dissolved in DMSO at 10 mg mL−1 and then diluted to 1 mg mL−1 in PBS), and PBS were injected to mice intravenously on day 7 (n = 5 for each group). Tumor sizes were measured every 3 days with a caliper. Tumor volumes were calculated using the modified formula: (length × width2)/2.Tumor disaggregation and analysis
Mice were euthanized on day 27 and the tumors were dissected. The tumor was minced into small pieces then digested using 1 mg mL−1 collagenase, 10 mg mL−1 hyaluronidase, and 0.02 mg mL−1 DNase of 5 mL of Hank's balanced salt solution (HBSS) on a rotator for 2 h at 25 °C. The cells suspension was filtered with a 70 µm cell strainer and the solution was centrifuged at 1200 rpm for 8 min. The supernatant was removed, and the cell pellet was washed with HBSS for three times.Tumor cells (105) from different groups were placed in a V-bottom shape 96-well plate. To check the expression of CD44, cells were stained with anti-CD44 APC/Cy7 according to the manufacturer's protocol and washed with PBS for three times then analyzed by flow cytometry.
Histological analysis
Dissected tumors were fixed in 10% neutral buffered formalin (NBF) and then processed, and vacuum infiltrated with paraffin on the Sakura VIP 2000 tissue processor followed by embedding. Paraffin blocks were sectioned at 5 µm for all tissue staining. For CD44 IHC staining, slides were blocked for nonspecific binding with Rodent Block M for 20 min, followed by polyclonal rabbit anti-CD44 antibody (catalog #: ab157107) dilutions (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200) incubations for 1 h at room temperature. Slides were then incubated with rabbit on rodent HRP micro polymer (catalog #: RMR 622 G, H, L) for 20 min with reaction developed utilizing Romulin AEC chromogen for 5 min. Slides were counterstained with CAT hematoxylin in a 1
200) incubations for 1 h at room temperature. Slides were then incubated with rabbit on rodent HRP micro polymer (catalog #: RMR 622 G, H, L) for 20 min with reaction developed utilizing Romulin AEC chromogen for 5 min. Slides were counterstained with CAT hematoxylin in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 ratio for 1 min and were analyzed with a Nikon Eclipse Ci microscope with a Nikon DS-Fi3 camera (Nikon Instruments Inc., Japan).
5 ratio for 1 min and were analyzed with a Nikon Eclipse Ci microscope with a Nikon DS-Fi3 camera (Nikon Instruments Inc., Japan).
Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra07975f.Acknowledgements
We are grateful for the financial support from National Institutes of Health (R01CA225105 and R35GM158263), and Michigan State University.References
- M. T. Manzari, Y. Shamay, H. Kiguchi, N. Rosen, M. Scaltriti and D. A. Heller, Nat. Rev. Mater., 2021, 6, 351–370 CrossRef CAS PubMed.
- D. Rosenblum, N. Joshi, W. Tao, J. M. Karp and D. Peer, Nat. Commun., 2018, 9, 1410 CrossRef PubMed.
- H. Xu, M. Niu, X. Yuan, K. Wu and A. Liu, Exp. Hematol. Oncol., 2020, 9, 36 CrossRef CAS.
- Q. Guo, C. Yang and F. Gao, FEBS J., 2022, 289, 7970–7986 CrossRef CAS.
- C. Chen, S. Zhao, A. Karnad and J. W. Freeman, J. Hematol. Oncol., 2018, 11, 64 CrossRef.
- G. Honeth, P.-O. Bendahl, M. Ringnér, L. H. Saal, S. K. Gruvberger-Saal, K. Lövgren, D. Grabau, M. Fernö, Å. Borg and C. Hegardt, Breast Cancer Res., 2008, 10, R53 CrossRef.
- R. Thapa and G. D. Wilson, Stem Cell. Int., 2016, 2016, 2087204 CrossRef.
- Y. Yan, X. Zuo and D. Wei, Stem Cells Transl. Med., 2015, 4, 1033–1043 CrossRef CAS.
- S. Bisht, M. Nigam, S. S. Kunjwal, P. Sergey, A. P. Mishra and J. Sharifi-Rad, Stem Cell. Int., 2022, 2022, 9653244 Search PubMed.
- P. Kesharwani, R. Chadar, A. Sheikh, W. Y. Rizg and A. Y. Safhi, Front. Pharmacol, 2021, 12, 800481 CrossRef CAS.
- I. H. Sahin and J. Klostergaard, Expert Opin. Ther. Targets, 2015, 19, 1587–1591 CrossRef CAS.
- S. Li, H. Wang, S. Xiong, J. Liu and S. Sun, Pharmaceuticals, 2024, 17, 832 CrossRef CAS PubMed.
- U. Rupp, E. Schoendorf-Holland, M. Eichbaum, F. Schuetz, I. Lauschner, P. Schmidt, A. Staab, G. Hanft, J. Huober, H. P. Sinn, C. Sohn and A. Schneeweiss, Anticancer Drugs, 2007, 18, 477–485 Search PubMed.
- H. Riechelmann, A. Sauter, W. Golze, G. Hanft, C. Schroen, K. Hoermann, T. Erhardt and S. Gronau, Oral Oncol., 2008, 44, 823–829 CrossRef CAS PubMed.
- S. Misra, V. C. Hascall, R. R. Markwald and S. Ghatak, Front. Immunol., 2015, 6, 201 Search PubMed.
- L. Lapcík, Jr., L. Lapcík, S. De Smedt, J. Demeester and P. Chabrecek, Chem. Rev., 1998, 98, 2663–2684 CrossRef PubMed.
- S. Banerji, A. J. Wright, M. Noble, D. J. Mahoney, I. D. Campbell, A. J. Day and D. G. Jackson, Nat. Struct. Mol. Biol., 2007, 14, 234–239 CrossRef CAS.
- K. Liu and X. Huang, Carbohydr. Res., 2022, 511, 108500 CrossRef CAS PubMed.
- N. V. Rao, H. Y. Yoon, H. S. Han, H. Ko, S. Son, M. Lee, H. Lee, D.-G. Jo, Y. M. Kang and J. H. Park, Expet Opin. Drug Deliv., 2016, 13, 239–252 Search PubMed.
- I. S. Bayer, Molecules, 2020, 25, 2649 CrossRef CAS PubMed.
- K. Liu, W. M. McCue, C.-W. Yang, B. C. Finzel and X. Huang, Carbohydr. Polym., 2023, 300, 120255 Search PubMed.
- I. P. Marjolein and L. M. Timo, Adv. Drug Deliv. Rev., 2023, 192, 114621 CrossRef.
- Y. Sun, H. S. Kim, S. Kang, Y. J. Piao, S. Jon and W. K. Moon, Adv. Healthcare Mater., 2018, 7, 1800266 Search PubMed.
- W. Cai, H. Gao, C. Chu, X. Wang, J. Wang, P. Zhang, G. Lin, W. Li, G. Liu and X. Chen, ACS Appl. Mater. Interfaces, 2017, 9, 2040–2051 Search PubMed.
- R. Chakravarty, H. Hong and W. Cai, Mol. Pharm., 2014, 11, 3777–3797 Search PubMed.
- H. He, X. Zhang, L. Du, M. Ye, Y. Lu, J. Xue, J. Wu and X. Shuai, Adv. Drug Delivery Rev., 2022, 186, 114320 Search PubMed.
- Y. Li, J. Zhang, L. Zhu, M. Jiang, C. Ke, H. Long, R. Lin, C. Ye, X. Zhou, Z.-X. Jiang and S. Chen, Adv. Healthcare Mater., 2023, 12, 2300941 CrossRef CAS.
- B. Guo, A. M. Sofias, T. Lammers and J. Xu, Adv. Drug Deliv. Rev., 2024, 206, 115188 CrossRef CAS.
- M. I. Priester and T. L. M. ten Hagen, Adv. Drug Delivery Rev., 2023, 192, 114621 CrossRef CAS.
- T. Desmettre, J. M. Devoisselle and S. Mordon, Surv. Ophthalmol., 2000, 45, 15–27 Search PubMed.
- M. Ogawa, N. Kosaka, P. L. Choyke and H. Kobayashi, Cancer Res., 2009, 69, 1268–1272 CrossRef CAS.
- J. T. Alander, I. Kaartinen, A. Laakso, T. Patila, T. Spillmann, V. V. Tuchin, M. Venermo and P. Valisuo, Int. J. Biomed. Imaging, 2012, 2012, 940585 Search PubMed.
- K. G. Gareev, K. Y. Babikova, V. N. Postnov, E. B. Naumisheva and D. V. Korolev, J. Phys.:Conf. Ser., 2017, 917, 042008 CrossRef.
- I. Akrida, N. V. Michalopoulos, M. Lagadinou, M. Papadoliopoulou, I. Maroulis and F. Mulita, Cancers, 2023, 15, 5755 CrossRef PubMed.
- P. B. Gupta, T. T. Onder, G. Jiang, K. Tao, C. Kuperwasser, R. A. Weinberg and E. S. Lander, Cell, 2009, 138, 645–659 CrossRef CAS.
- C. Naujokat and R. Steinhart, BioMed Res. Int., 2012, 2012, 950658 Search PubMed.
- T. T. Mai, A. Hamaï, A. Hienzsch, T. Cañeque, S. Müller, J. Wicinski, O. Cabaud, C. Leroy, A. David, V. Acevedo, A. Ryo, C. Ginestier, D. Birnbaum, E. Charafe-Jauffret, P. Codogno, M. Mehrpour and R. Rodriguez, Nat. Chem., 2017, 9, 1025–1033 CrossRef CAS.
- H. Wang, H. Zhang, Y. Zhu, Z. Wu, C. Cui and F. Cai, Front. Oncol., 2021, 11, 654428 CrossRef CAS.
- J. Zhou, S. Liu, Y. Wang, W. Dai, H. Zou, S. Wang, J. Zhang and J. Pan, Mol. Cancer, 2019, 18, 159 CrossRef.
- Y. Zhang, H. Zhang, X. Wang, J. Wang, X. Zhang and Q. Zhang, Biomaterials, 2012, 33, 679–691 Search PubMed.
- D. Qi, Y. Liu, J. Li, J. H. Huang, X. Hu and E. Wu, Med. Res. Rev., 2022, 42, 1037–1063 CrossRef CAS.
- Y. Zhao, K. Wang, Y. Zheng, X. Zeng, Y. C. Lim and T. Liu, Front. Chem., 2021, 8, 601649 CrossRef.
- M. Hamdi, E. Elmowafy, H. M. Abdel-Bar, A. M. ElKashlan, K. T. Al-Jamal and G. A. S. Awad, Int. J. Biol. Macromol., 2022, 217, 731–747 CrossRef CAS.
- Y. S. Yang, Y. Zhao, J. S. Lan, Y. N. Kang, T. Zhang, Y. Ding, X. Y. Zhang and L. Lu, Int. J. Nanomed., 2018, 13, 4361–4378 CrossRef CAS.
- L. Isert, I. Gialdini, T. M. H. Ngo, G. Loiudice, D. C. Lamb and O. M. Merkel, Nanoscale, 2025, 17, 16256–16273 RSC.
- S.-J. Yang, J.-A. Pai, C.-J. Yao, C.-H. Huang, J. L. Y. Chen, C.-H. Wang, K.-C. Chen and M.-J. Shieh, Cancer Nanotechnol., 2023, 14, 1 CrossRef CAS.
- B. Lin, P. Lin, X. Zhang, Y. Liao, Y. Yu, X. Xu and X. Wang, Int. J. Nanomed., 2024, 19, 14105–14124 Search PubMed.
- Y. Shao, W. Luo, Q. Guo, X. Li, Q. Zhang and J. Li, Drug Des., Dev. Ther., 2019, 13, 2043–2055 Search PubMed.
- S. Sarkar, B. Kanchibotla, J. D. Nelson, J. D. Edwards, J. Anderson, G. C. Tepper and S. Bandyopadhyay, Nano Lett., 2014, 14, 5973–5978 CrossRef CAS.
- Z. Zhao, B. He, H. Nie, B. Chen, P. Lu, A. Qin and B. Z. Tang, Chem. Commun., 2014, 50, 1131–1133 RSC.
- R. L. Siegel, K. D. Miller, N. S. Wagle and A. Jemal, Ca-Cancer J. Clin., 2023, 73, 17 CrossRef.
- X. Sun, Y. Xu, Q. Guo, N. Wang, B. Wu, C. Zhu, W. Zhao, W. Qiang and M. Zheng, Langmuir, 2022, 38, 1360 Search PubMed.
- R. K. Mittapalli, X. Liu, C. E. Adkins, M. I. Nounou, K. A. Bohn, T. B. Terrell, H. S. Qhattal, W. J. Geldenhuys, D. Palmieri, P. S. Steeg, Q. R. Smith and P. R. Lockman, Mol. Cancer Ther., 2013, 12, 2389–2399 CrossRef CAS PubMed.
- S. Chu, X. Shi, Y. Tian and F. Gao, Front. Oncol., 2022, 12, 855019 CrossRef CAS.
- P. Thummarati, J. Suksiriworapong, K. Sakchaisri, T. Nawroth, P. Langguth, B. Roongsawang and V. B. Junyaprasert, J. Drug Delivery Sci. Technol., 2022, 77, 103883 CrossRef CAS.
- Q. Wang, L. Zhou, L. Qiu, D. Lu, Y. Wu and X.-B. Zhang, Analyst, 2015, 140, 5563–5569 RSC.
- D. E. Johnson, P. Ostrowski, V. Jaumouillé and S. Grinstein, J. Cell Biol., 2016, 212, 677–692 CrossRef CAS PubMed.
- M. S. Klempner and B. Styrt, J. Clin. Invest., 1983, 72, 1793–1800 CrossRef CAS PubMed.
- J. Ngai, P. MacMillan, B. R. Kingston, Z. P. Lin, B. Ouyang and W. C. W. Chan, J. Controlled Release, 2023, 353, 988–1001 Search PubMed.
- Z. Starosolski, R. Bhavane, K. B. Ghaghada, S. A. Vasudevan, A. Kaay and A. Annapragada, PLoS One, 2017, 12, 0187563 CrossRef.
- C.-W. Yang, K. Liu, C.-Y. Yao, B. Li, A. Juhong, Z. Qiu and X. Huang, ACS Appl. Nano Mater., 2022, 5, 18912–18920 Search PubMed.
- C.-W. Yang, K. Liu, C.-Y. Yao, B. Li, A. Juhong, A. K. M. A. Ullah, H. Bumpers, Z. Qiu and X. Huang, ACS Appl. Mater. Interfaces, 2024, 16, 27055–27064 CrossRef CAS.
- S. Kim, Y. Shi, J. Y. Kim, K. Park and J.-X. Cheng, Expet Opin. Drug Deliv., 2010, 7, 49–62 Search PubMed.
- H. Duan, Y. Liu, Z. Gao and W. Huang, Acta Pharm. Sin. B, 2021, 11, 55–70 CrossRef CAS PubMed.
- S. Shrestha, A. Banstola, J.-H. Jeong, J. H. Seo and S. Yook, J. Controlled Release, 2022, 348, 518–536 CrossRef CAS PubMed.
- I. Dagogo-Jack and A. T. Shaw, Nat. Rev. Clin. Oncol., 2018, 15, 81–94 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |