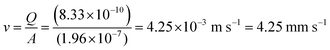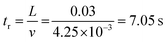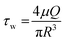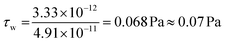Open Access Article
Open Access ArticleA low-cost liquid crystal display (LCD)-printed microfluidic device: fabrication and application in the biodegradation of petroleum hydrocarbons in petroleum refinery sludge
Amlan Ashish†
 ab,
Ankita Debnath†cd,
Poulomi Biswas
ab,
Ankita Debnath†cd,
Poulomi Biswas a,
Ramkrishna Sen
a,
Ramkrishna Sen c and
Gorachand Dutta
c and
Gorachand Dutta *a
*a
aNanoBiosensors and Biodevices Lab, School of Medical Sciences and Technology, Indian Institute of Technology Kharagpur, Kharagpur-721302, India. E-mail: g.dutta@smst.iitkgp.ac.in
bAdvanced Technology Development Centre (ATDC), Indian Institute of Technology Kharagpur, Kharagpur-721302, India
cDepartment of Bioscience and Biotechnology, Institute of Technology Kharagpur, Kharagpur-721302, India
dALS Testing Services India Pvt Ltd, Hyderabad, India
First published on 7th May 2026
Abstract
Liquid crystal display (LCD) printing is evolving into a ground-breaking technique in the biological domain, demonstrating the benefits of fabricating low-cost, high-resolution, complex geometry in bulk compared to the time-consuming and labour-intensive lithography techniques. However, the exploration of 3D-printed microfluidic devices for bioremediation and biofilm formation remains underrepresented in the current literature. This article describes the validation of a cost-effective LCD-printed microfluidic device that was employed as a biofilm-assisted carrier for the remediation of petroleum hydrocarbons in refinery sludge. Our technology surpasses current bioremediation approaches by providing a degrading surface that differs from standard methods relying solely on particulate carriers. A novel microbial consortium comprising four hydrocarbon-degrading strains (Dietzia sp. IRB191, Dietzia sp. IRB192, Staphylococcus sp. BSM19, and Stenotrophomonas sp. IRB19), isolated from refinery sludge, was enforced to form a biofilm on the microchannel's surface. The microfluidic device was fabricated by optimizing the printing layer height, curing timelight intensity and post-processing techniques. Micro-CT scanning was performed to analyse the deviation in the overall dimensions of the microchannels. Microscopy suggested an increase in extracellular polymeric substance (EPS) accumulation and roughness value with the increase in incubation time. The roughness value increased from 93 nm to 162 nm after a duration of 15 days. The microfluidic device exhibited a tensile strength of 18.57 MPa, which shows the structural stability of the fabricated device. Approximately 82% ± 6% of total hydrocarbons from the petroleum refinery waste were degraded during a period of 15 days, as confirmed by gravimetric analysis and gas chromatography-mass spectrometry (GC-MS) measurements.
1. Introduction
Microfluidic devices are a group of miniature, efficient and highly controlled devices, commonly known as Lab-on-a-chip (LOC) devices,1–8 that hold great potential for studying and optimising the growth of microorganisms. They help in microbial screening, biodegradation analysis, and can ultimately lead to faster and more effective cleanup strategies.The devices comprise channels connected to chambers and reservoirs, forming closed networks that enable researchers to replicate the microenvironment of many complicated biological phenomena and describe changes in environmental variables at the micron range more precisely than with conventional culture plates. LOC-based microdevices are less often utilised in molecular biology, resulting in limited research on their application for microbial cultivation and other activities. Presently, there is a surge in the use of LOC for investigations into bacterial behaviour in response to external stimuli, encompassing taxis mechanisms, drug screening, and biofilm development. 3D-printed miniaturized chip-based LOC devices offer an innovative approach to studying biofilm formation and the effects of various parameters such as nutrient concentration, environmental conditions, and shear stress. The detection of environmental contaminants can be coupled with these microfluidic devices to find a solution. 3D-printing, a kind of additive manufacturing process, is emerging as a revolutionary approach in the medical and industrial domains for the fabrication of microfluidic devices, implants and visualization of prototypes.8–16 Basically, it can be used to produce complex structures with minimal material waste in a shorter time. Liquid crystal display (LCD) printing/resin printing is a technique used to create cost-effective 3D-printed geometries with high dimensional accuracy and resolution by exposing the resin to a UV light source.2,3,6,17–23 The advancement of LOC will facilitate the identification of bacterial biofilm populations capable of decomposing petroleum hydrocarbons, thereby positively influencing bioremediation processes. Furthermore, the innovative LOC technology, as a compact system, facilitates interactions between bacterial biofilms and hydrodynamic conditions (such as shear stress) and may also be employed in the development of portable and user-friendly devices for analyzing effluents or contaminated soil or water samples.
Petroleum hydrocarbons (PHs) are major emerging pollutants in the environment, having a notoriously harmful effect not only on mankind but also on animal and marine ecosystems. These pollutants are recalcitrant in nature as they are bio-accumulated through the food chain of marine ecosystems, thereby having a longer shelf life.24,25 Concomitant spillage of these pollutants from underground reservoirs during petroleum processing and transportation from storage tanks, etc.,26 cannot be completely prevented, thus making effective remediation the need of the hour. For the bioremediation, many indigenous bacterial species with effective hydrocarbon-degradation potential have been identified from petroleum-polluted streams.27 However, there is a lack of evidence of a single bacterial species that can completely degrade total PHs. This has led to the utilization of microbial consortia for bio-stimulation and bio-augmentation mediated degradation of PHs.28 We have compared our carrier-mediated bioremediation of total PHs to other carriers in Table 1.
| Carrier | Microorganisms immobilized | % TPH degradation | No. of treatment days | Reference |
|---|---|---|---|---|
| Wheat bran biochar | Pseudomonas, Acinetobacter, and Sphingobacterium | 58.31 | 84 | 29 |
| Wheat bran | Hydrocarbonoclastic consortia of Pseudoalteromonas sp., Oceanobacillus sp., Nesiotobacter sp., Photobacterium sp., Ruegeria sp., Enterobacter sp., Haererehalobacter sp., Exiguobacterium sp. and Acinetobacter sp. | 80-90 | 10 | 30 |
| Polylactic acid and polycaprolactone electrospun membranes | Alcanivorax borkumensis SK2, Gordonia sp. SoCg, Oleibacter marinus 5, Nocardia sp. SoB | 66 | 10 | 31 |
| Activated carbon and zeolite | Hydrocarbon-degrading bacterial consortium | 48.89 | 30 | 32 |
| Peanut hull powder | Indigenous hydrocarbon-degrading bacterial consortium | 61 | 90 | 33 |
| Biochar | Pseudomonas aeruginosa, Acinetobacter radioresistens | 53 | 84 | 34 |
| Cinnamon shells | Pseudomonas sp. YT-11 | 69.94 | 10 | 35 |
| Peanut shells | Pseudomonas sp. YT-11 | 64.41 | 10 | 35 |
| Calcium alginate (CA)-chitosan compound with a coating of puffed foxtail millet (PFM) | Acinetobacter sp. F9 | 90 | 7 | 36 |
| 3D-printed platform carrying a biofilm of TPH-degrading bacteria | Microbial consortium of Dietzia sp. IRB191, Stenotrophomonas sp. IRB19, Dietzia sp. IRB192 and Staphylococcus sp. BSM19 in the form of biofilm on 3D-printed surface | 81 | 15 | Present study |
One of the main challenges in current bioremediation approaches is gaining a clear understanding of microbial activity. Typically, laboratory tests are used to demonstrate the capability of native microorganisms to degrade pollutants, which often requires either bioaugmentation or biostimulation. However, these tests are traditionally conducted at a macroscale using conventional microbiological techniques, which are time-consuming and expensive due to the need for large amounts of reagents and materials. Furthermore, there is a growing demand for technologies that can enable effective, real-time monitoring and control of bioremediation processes directly in contaminated environments, to enhance the efficiency and success of microbial treatment strategies. The advancement of 3D-printed microfluidic technology holds promise for enhancing the identification of bacterial biofilms capable of breaking down petroleum-based pollutants, leading to more efficient and well-monitored bioremediation processes. As a compact and miniaturized platform, these systems offer unique benefits by enabling biofilm formation inside the device and controlled interactions between microbial biofilms and fluid dynamics, such as shear stress. Additionally, this technology has the potential to be adapted into portable, user-friendly biosensors for analyzing wastewater or contaminated soil samples in the field. In the future, this technology could be explored through the development of material coatings on the interiors of refinery tanks and on ship surfaces, where biofilm-mediated degradation may potentially occur.
In this work, a bacterial biofilm-coated 3D-printed microfluidic device was conceptualized for active biodegradation and bio-deterioration of petroleum hydrocarbon-contaminated refinery sludge. Promising results in terms of total petroleum hydrocarbon (TPH) degradation were obtained at the end of two weeks. A bacterial consortium was allowed to form a biofilm on the 3D-printed microchannel, which was then used for TPH degradation on the 15th day. The biofilm formed on the 3D-printed surface has degraded 81 ± 2% of TPH obtained from petroleum refinery sludge (PRS) after 15 days of treatment. GC MS chromatograms revealed a noticeable decrease in the concentration of long chain hydrocarbons (C50–C30) and its convertion to shorter chain hydrocarbons (C10–C20). This strategy prevents the dilution and damage of microbial cells in polluted water as the microorganisms remain in the biofilm, encased in extracellular polymeric substances. These biofilms, in turn, remain attached to the surface of the 3D-printed microchannels, thereby acting as smart tools for degrading total PHs. This study represents an ultra-low-cost LCD-printed PEGDA microfluidic device that can be employed as a reusable biofilm immobilization carrier as an alternative to existing particle-based carrier techniques for treating real petroleum refinery sludge (TPH mixture). The device used a novel microbial consortium comprising four hydrocarbon-degrading strains (Dietzia sp. IRB191, Dietzia sp. IRB192, Staphylococcus sp. BSM19, and Stenotrophomonas sp. IRB19), isolated from refinery sludge, which is further supported by micro-CT dimensional metrology and mechanical property optimization.
2. Materials and methods
2.1. Materials and equipment
All the media components used for culturing, biofilm formation, and TPH degradation studies were purchased from Merck Lifesciences Private Limited, India. Petroleum refinery sludge was collected from IOCL Haldia, West Bengal, India. Four bacterial strains (Dietzia sp. IRB191, Dietzia sp. IRB192, Staphylococcus sp. BSM19, and Stenotrophomonas sp. IRB19) were isolated from petroleum refinery sludge. Solvents used for TPH extraction were purchased from Sisco Research Laboratories Pvt. Ltd, India, and Sigma-Aldrich. GC MS PerkinElmer model no Clarus SQ 8T (United Kingdom) was used to analyse TPH degradation. Poly(ethylene glycol)diacrylate (PEGDA, MW250) and phenylbis(2,4,6-trimethylbenzoyl)phosphine oxide (Irgacure 819), which served as monomer and photoinitiator, respectively, were procured from Sigma-Aldrich. Isopropanol was purchased from Sigma to post-process the microfluidic device. The device fabrication was carried out using a low-cost liquid crystal display (LCD) based Photon Mono M5 printer purchased from Anycubic (China).2.2. Resin formulation
The resin was formulated by mixing PEGDA monomer, with an average molecular weight of 250, and 1 wt% phenylbis(2,4,6-trimethylbenzoyl)phosphine oxide (Irgacure-819), which acts as the photoinitiator. PEGDA-based resin formulations are widely utilised for device fabrication owing to their favourable curing kinetics and mechanical stability. However, it is important to select the correct ratio of monomer to PI for the fabrication of a functional and defect-free microfluidic device. In this study, an optimised and practical formulation of the PEGDA-based resin has been proposed. The mixed solution container was covered with aluminum foil to avoid direct exposure to light and magnetically stirred for 30 min to ensure proper mixing. A schematic of the crosslinking of the PEGDA with Irgacure 819 is shown in Fig. 1.PEGDA, a PEG derivative, has reactive acrylate groups that were replaced with hydroxyl groups from PEG, and was considered as the monomer. The transformation of the polymer solution into a three-dimensional long-chain configuration is accomplished via crosslinking. Various methodologies for cross-linking PEGDA encompass physical, chemical, and enzymatic methods, or a combination of these approaches. The crosslinking technique significantly affects the mechanical and physicochemical properties of the 3D-printed microfluidic device. Free radical photopolymerization is employed for this conversion, an exceptionally efficient method that rapidly transforms a liquid monomer into a solid polymer while allowing precise control over the material's properties, including mechanical strength, flexibility and biodegradability. It occurs at ambient temperature, rendering it appropriate for several applications, including 3D printing, coatings, and adhesives. The polymerisation of PEGDA with Irgacure 819 is initiated by the generation of free radicals, following exposure to light, often ultraviolet (UV) or visible light. A photoinitiator absorbs light energy (photons), undergoes a photochemical reaction, and decomposes to produce free radicals during this process. Subsequently, the produced free radicals target the double-bonded carbon (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) in the acrylate groups and initiate a chain polymerisation reaction that forms a crosslinked polymer network in which individual PEGDA molecules are interconnected.
C) in the acrylate groups and initiate a chain polymerisation reaction that forms a crosslinked polymer network in which individual PEGDA molecules are interconnected.
2.3. 3D-printing of the microfluidic device
The microfluidic chip was designed using CAD software (Autodesk Fusion 360), and the file was saved in STL format for printing on the Anycubic Photon mono M5 (LCD printer). LCD printing is a technique used for creating cost-effective 3D-printed geometries with high dimensional accuracy and resolution by exposing the resin to a UV light source.2,3,6,17–23 LCD-based printing is a subset of the vat polymerization technique that uses a photocurable liquid resin that selectively cures each layer when exposed to a light source of a specific wavelength. An array of LEDs shines the pattern onto the bottom of the resin tank through an LCD screen. The process facilitates crosslinking of an entire layer of resin at a time, rather than curing point by point, which results in a drastic reduction in printing time. The microfluidic device was printed using a layer-by-layer technique that induced a roughness on the surface of the device, which facilitates initial attachment of bacteria and subsequent formation of the biofilm on its surface.This printer has a 405 nm wavelength, 45 W power rating, 12K LCD of 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 520 × 5120 pixels, and a pixel size of 19 µm. The dimensions of the printer are 460 mm × 270 mm × 290 mm, a resin vat of 170 mm × 120 mm, and a build volume of 200 mm × 80 mm × 123 mm. A screw-based mechanism was used to move the printing platform. The design file was converted into a sliced file using Chitubox software, with the optimized values of part orientation, exposure time, support, layer height, etc. The platform was printed with a 2 s per layer (exposure time), while the bottom layer was exposed for 18 s, with a lift speed of 120 mm min−1 and a retract speed of 100 mm min−1, with every layer having a height of 50 µm. The 3D-printed microfluidic device was fabricated in 30 min with a minimal resin consumption of 2.48 mL. Prior to commencing the printing process, the 3D printer was calibrated to make a reference position, and subsequently, a calculated volume of resin was poured into the vat tray. After the completion of printing, the microfluidic device was gently rinsed with isopropanol for 10 min in the washing station and then post-cured using a UV lamp (40 W) to remove any excess resin and improve its longevity. The printed microfluidic device was constructed with a length of 40 mm, a width of 20 mm, and a height of 3 mm. The overall dimensions of the device and the microchannel are shown in Fig. 2a, and the printing process is shown schematically in Fig. 2b. The device is cost-effective (USD 0.28 per device) and can be reused after cleaning with IPA.
520 × 5120 pixels, and a pixel size of 19 µm. The dimensions of the printer are 460 mm × 270 mm × 290 mm, a resin vat of 170 mm × 120 mm, and a build volume of 200 mm × 80 mm × 123 mm. A screw-based mechanism was used to move the printing platform. The design file was converted into a sliced file using Chitubox software, with the optimized values of part orientation, exposure time, support, layer height, etc. The platform was printed with a 2 s per layer (exposure time), while the bottom layer was exposed for 18 s, with a lift speed of 120 mm min−1 and a retract speed of 100 mm min−1, with every layer having a height of 50 µm. The 3D-printed microfluidic device was fabricated in 30 min with a minimal resin consumption of 2.48 mL. Prior to commencing the printing process, the 3D printer was calibrated to make a reference position, and subsequently, a calculated volume of resin was poured into the vat tray. After the completion of printing, the microfluidic device was gently rinsed with isopropanol for 10 min in the washing station and then post-cured using a UV lamp (40 W) to remove any excess resin and improve its longevity. The printed microfluidic device was constructed with a length of 40 mm, a width of 20 mm, and a height of 3 mm. The overall dimensions of the device and the microchannel are shown in Fig. 2a, and the printing process is shown schematically in Fig. 2b. The device is cost-effective (USD 0.28 per device) and can be reused after cleaning with IPA.
2.4. Fourier-transform infrared spectroscopy analysis
Fourier-transform infrared spectroscopy (FTIR) was performed with Agilent Cary 630 equipment to establish the composition and determine the functional groups present on the 3D-printed surface. The spectral data were acquired within the wavelength region of 4000–500 cm−1. In order to prepare the sample, a glass slide was thoroughly cleaned with isopropanol and DI water and then air-dried. A volume (200 µL) of resin was then injected onto a surface and exposed to ultraviolet (UV) radiation within a controlled environment for 10 minutes to accelerate the process of curing and the formation of a thinner resin layer. The cured coating was then extracted from the glass slide in order to conduct the study.2.5. X-ray photoelectron spectroscopic analysis
The elemental composition of the material was determined by employing the surface-based analysis technique called X-ray photoelectron spectroscopy (XPS). This determination is based on the binding energy associated with the elements present on the material's surface. A block with dimensions of 5 mm × 5 mm × 1 mm was fabricated using the 3D printer, and the sample was cleaned using DI water. A fraction of the sample was taken and used for X-ray photoelectron spectroscopy (XPS) analysis.2.6. Atomic force microscopic analysis
Atomic force microscopy (AFM) using an Agilent Technologies (USA) machine was employed to investigate the surface roughness of the 3D-printed carrier before and after formation of the biofilm. Blocks of 4 mm × 4 mm × 1 mm were 3D-printed and then cleaned with isopropanol and DI water to remove uncured surface resin and left to dry. Following this, four strains of oil-eating bacteria were allowed to grow as a biofilm on the surface, and roughness measurements were taken on the 3rd, 7th, and 15th day. Surface imaging of the printed surface was conducted in tapping mode (intermittent contact) using a silicon nitride cantilever tip.2.7. Micro-CT analysis of the 3D-printed microfluidic device
Micro-computed tomography (Micro-CT) was employed to evaluate the internal structure and dimensional precision of the microfluidic device in a non-destructive manner. This X-ray-based technique generates multiple two-dimensional X-ray projections, which are then reconstructed into cross-sectional images of the object. Such an approach enables detailed observation of internal features in materials and biological specimens without physically altering or cutting them, preserving the samples for additional analysis. The microfluidic device was scanned using a micro-CT system (GE Phoenix v|tome|x, Germany), operating at 85 kV and 100 µA. VGSTUDIO MAX software was used to accurately determine the channel's height and width, along with the inlet and outlet dimensions, which were then compared to the original measurements specified in the CAD model.2.8. Development of a microbial consortium and the viability of the biofilm-coated microchannel
Four hydrocarbon-degrading bacterial strains, Stenotrophomonas sp. IRB19, Dietzia sp. IRB191, Staphylococcus sp. BSM19, and Dietzia sp. IRB192, previously isolated and characterized by Behera et al.,37 were used to develop a consortium by inoculating an equal volume of each, to study the biofilm formation. A dextrose mineral salts medium (DMSM) with the following composition was used: 2% (w/v) dextrose, 668 mg L−1 Na2HPO4·7H2O, 435 mg L−1 K2HPO4, 170 mg L−1 KH2PO4, 850 mg L−1 NH4Cl, 27.5 mg L−1 CaCl2·2H2O, 0.25 mg L−1 FeCl3, and 22.5 mg L−1 MgSO4·7H2O. The consortium was allowed to flow into the microfluidic channel along with DMSM to attach and develop the biofilm on the surface of the microchannel. The microfluidic device was sterilised by UV irradiation for 20–30 min prior to addition to the medium. Similarly, a negative control was made without the addition of the microbial consortium. The viability of the biofilm on the 3D-printed microfluidic surface was studied on the 3rd, 7th and 15th day. To study the viability, the microchannels were subjected to a washing step using 0.1 M phosphate buffer at pH 7.4 and flowing the same buffer. The suspension was then serially diluted and plated to obtain the cell concentration as cfu per mL per surface. The same procedure was repeated for the negative control. Also, in order to confirm the viability of the biofilm, the microchannels were directly placed on nutrient agar plates. On confirming biofilm viability, they were subsequently applied for TPH degradation.2.9. Total hydrocarbon degradation by the biofilm-coated microchannels
The experimental setup for the microfluidic hydrocarbon degradation of petroleum sludge is illustrated in Fig. 3a, and the degradation mechanism is shown in Fig. 3b. The microfluidic device was connected with a dual syringe pump (Harvard 33 DDS Dual) to flow the bacterial consortium, growth media, and petroleum sludge with the required flow rates. An open-loop syringe pump system with 100 mL syringes was used, and the syringes were replaced periodically 11 times to accommodate a total volume requirement of 1080 mL to complete the experiment for a period of 15 days. The flow was resumed after syringe refilling to maintain quasi-continuous operation. Considering the channel width of 0.5 mm and length of 30 mm, operating at a flow rate of 50 µL min−1, the average velocity was 4.25 mm s−1, resulting in a residence time of approximately 7 s per pass. The corresponding wall shear stress was approximately 0.07 Pa, which lies within the low-shear-rate regime, which is suitable for stable microbial attachment and biofilm-mediated biodegradation. The calculations are given below. | ||
| Fig. 3 (a) Experimental setup of the microfluidic hydrocarbon degradation of petroleum sludge and (b) degradation mechanism. | ||
Cross-sectional area (A)
| A = πr2 = π(2.5 × 10−4)2 |
Average velocity (ν)
Residual time (tr)
Wall shear stress (τw)
For fully developed laminar flow inside a circular microchannel
To avoid clogging, the sludge solution was passed through a coarse filter prior to syringe loading. Also, lower flow rates and laminar flow minimised the clogging issue. Additionally, the microchannel dimensions were kept relatively large (i.e., 0.5 mm instead of 0.1 or 0.2 mm).
The entire degradation process occurs in two steps: (i) biofilm formation by the bacterial consortium and (ii) degradation of sludge. Initially, the microchannels were sterilized by flushing 70% ethanol and type I water, followed by 20 min UV irradiation. During the biofilm formation step, inlet 1 was used to flow the liquid consortium, and DMSM media was allowed to pass through inlet 2 with a flow rate of 50 µL min−1 using a syringe pump, providing sufficient time to adhere to the microchannel surface and form the biofilm. After completion of the biofilm-formation process, 0.1 M phosphate buffer (pH 7.4) was used to eliminate loosely attached cells or culture exudates and residual medium components. For the hydrocarbon degradation step, the same media composition was utilised with a single modification, where 2% (w/v) PRS was used as the sole carbon source instead of dextrose. After a period of 15 days of biofilm formation, the media without dextrose was allowed to flow through inlet 1 and the PRS was passed along inlet 2. The TPH degradation was investigated on the 3rd, 7th and 15th day using a gravimetric method, as reported by Behera et al.,37,38 and gas chromatography-mass spectral analysis. TPH degradation was studied using the biofilm-coated microchannels as a tool for bioremediation, without the addition of any other microbial cells in the form of inoculum. Biofilm-coated microchannels during TPH degradation were also studied by microscopy to assess changes in the 3D-printed channel's surface with respect to the control. The negative control was made with a UV-sterilised microchannel that did not exhibit any biofilm formation. TPH extraction from PRS involved liquid–liquid extraction using hexane, dichloromethane and chloroform. TPH was extracted from untreated PRS in the negative control and PRS obtained after treatment with the biofilm-coated surface. The experiment utilised a PerkinElmer Clarus SQ 8T gas chromatography-mass spectrometry (GC-MS) instrument with an Elite 5MS column. The temperature was ramped from 35 °C to 280 °C with a rate of 10 °C per minute, while the injector temperature was maintained at 280 °C throughout the analysis.
2.10. Morphological characterization of the biofilm-coated 3D-printed microfluidic device
To examine the formation of the biofilm on the microchannel surface, scanning electron microscopy was employed. Samples were collected after 168 and 360 h of cultivating the microbial consortium on the biofilm development media. The biofilm-coated microchannels were first washed with 0.1 M phosphate buffer, following by 4% formaldehyde fixation for 1 h. After the fixing step, formaldehyde was decanted, and the microchannels were again washed with phosphate buffer. Subsequently, the film was dehydrated by exposure to a gradient of ethanol concentrations starting from 30%, 50%, 70% and 90% for 10 min each with a final dehydration in absolute ethanol for 1 h. The dehydrated microchannels were then stored overnight in a vacuum desiccator, followed by gold coating and imaging. As a control, platforms were taken from the media without any microbial inoculation. Both FESEM-JEOL and FESEM-MERLIN were utilized to observe the biofilm on the 3D-printed surface.3. Results and discussion
3.1. Material characterisation
Fourier-transform infrared (FTIR) spectroscopy enables the acquisition of valuable insights pertaining to the chemical bonding processes that occur during the polymerization phase of printing and to investigate the chemical structure and functional groups present in poly(ethylene glycol)diacrylate (PEGDA). Fourier-transform infrared attenuated total reflection (FTIR-ATR) spectra of the 3D-printed microfluidic device are shown in Fig. 4a. The spectra reveal the vibrational frequencies of certain functional groups when exposed to infrared radiation. The FTIR spectrum of PEGDA typically exhibits several characteristic peaks corresponding to its acrylate, ester, and ether functionalities. One of the most prominent peaks is the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching vibration (∼1720 cm−1), which arises from the ester groups in the acrylate moiety. The C
O stretching vibration (∼1720 cm−1), which arises from the ester groups in the acrylate moiety. The C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibration (∼1635 cm−1) is also observed, representing the carbon–carbon double bonds of the acrylate functional groups, which play a key role in photopolymerization. Additionally, the spectrum shows strong C–O–C stretching vibrations (∼1100–1200 cm−1) associated with the ether linkages in the PEG backbone, contributing to PEGDA's flexibility and hydrophilicity. Peaks around 2800–3000 cm−1 correspond to C–H stretching from aliphatic groups. When Irgacure 819 is added, additional peaks may appear, particularly in the aromatic C
C stretching vibration (∼1635 cm−1) is also observed, representing the carbon–carbon double bonds of the acrylate functional groups, which play a key role in photopolymerization. Additionally, the spectrum shows strong C–O–C stretching vibrations (∼1100–1200 cm−1) associated with the ether linkages in the PEG backbone, contributing to PEGDA's flexibility and hydrophilicity. Peaks around 2800–3000 cm−1 correspond to C–H stretching from aliphatic groups. When Irgacure 819 is added, additional peaks may appear, particularly in the aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching region (∼1600 cm−1) and P
C stretching region (∼1600 cm−1) and P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching (∼1250–1300 cm−1) due to the phosphine oxide structure of the photoinitiator. After UV curing or polymerization, the disappearance or significant reduction of the C
O stretching (∼1250–1300 cm−1) due to the phosphine oxide structure of the photoinitiator. After UV curing or polymerization, the disappearance or significant reduction of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C peak (∼1635 cm−1) serves as an indicator of successful crosslinking. FTIR analysis is thus essential for confirming the chemical composition of PEGDA, monitoring polymerization efficiency, and detecting any structural modifications in applications such as 3D printing, tissue engineering, and microfluidics.
C peak (∼1635 cm−1) serves as an indicator of successful crosslinking. FTIR analysis is thus essential for confirming the chemical composition of PEGDA, monitoring polymerization efficiency, and detecting any structural modifications in applications such as 3D printing, tissue engineering, and microfluidics.
X-ray photoelectron spectroscopy (XPS) is a valuable technique for analyzing the surface chemistry of poly(ethylene glycol)diacrylate (PEGDA) resin. It is crucial for evaluating the chemical composition, crosslinking behavior, and surface modifications of PEGDA-based materials, making it an essential tool for applications in biomedical engineering, microfluidics, and 3D printing. The overall XPS spectrum of the PEGDA resin-based microfluidic device is shown in Fig. 4b. The XPS spectra typically exhibit peaks for carbon (C 1s) and oxygen (O 1s). The C 1s spectrum is usually deconvoluted into three main components: C–C/C–H (∼285 eV) from the aliphatic hydro–carbon backbone, C–O (∼286.5 eV) corresponding to ether (–O–) groups from the PEG segments, and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (∼288–289 eV) related to ester (–COO–) functionalities from the acrylate groups, as shown in Fig. 4c. The O 1s peak appears around 532–533 eV, reflecting contributions from the ether and ester linkages (Fig. 4d). In the presence of irgacure 819 was confirmed by presence of phosphorus peak and the same is deconvoluted to P
O (∼288–289 eV) related to ester (–COO–) functionalities from the acrylate groups, as shown in Fig. 4c. The O 1s peak appears around 532–533 eV, reflecting contributions from the ether and ester linkages (Fig. 4d). In the presence of irgacure 819 was confirmed by presence of phosphorus peak and the same is deconvoluted to P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (∼133.5 eV) that refers to the phosphine oxide functional group and the interacting phosphine oxide with PEGDA matrix (–P–C) appeared at ∼134.5 eV. The polymerization of the resin was confirmed by the absence of an acrylate (C
O (∼133.5 eV) that refers to the phosphine oxide functional group and the interacting phosphine oxide with PEGDA matrix (–P–C) appeared at ∼134.5 eV. The polymerization of the resin was confirmed by the absence of an acrylate (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) peak in the spectra. Furthermore, a slight shift in the intensity of the C
C) peak in the spectra. Furthermore, a slight shift in the intensity of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (288.5 eV) and C–O (286.5 eV) peaks is observed due to the polymerization process.
O (288.5 eV) and C–O (286.5 eV) peaks is observed due to the polymerization process.
The surface roughness of the 3D-printed microchannels was analyzed using atomic force microscopy, as depicted in Fig. 4. The image scale has been altered such that higher phases appear lighter and lower phases appear darker. A scan area of 30 µm was considered for observation. 3D printing is an additive manufacturing process, where the part is printed layer by layer, resulting in a surface roughness instead of a completely smooth surface. It implies that the roughness (97.33 ± 4.51 nm) is an intrinsic property of the 3D-printed microchannels, as shown in Fig. 4e. Taking advantage of the roughness, the biofilm starts growing on the surface of the 3D-printed platform.39,40 After 3 days of incubation, the roughness value decreased to 33.33 ± 3.06 nm due to the initial adhesion of bacterial cells onto the intrinsically rough surface of the 3D-printed microchannels and gradual growth of the biofilm, as shown in Fig. 4f. Over time, more EPS is deposited, which, in turn, increases the roughness value. Roughness values of 87.00 ± 4.36 nm and 157.33 ± 4.51 nm were observed on the 7th and 15th day of incubation, as illustrated in Fig. 4g and h. (n = 3, p < 0.001). For the quantitative measurement, the roughness factors are listed in Table S1, and the standard deviations are shown in Fig. S1.
The 3D-printed microfluidic device was scanned to measure the internal dimensions of the microchannels, inlets, and outlets, as well as the overall dimensions of the device. The length and the breadth of the device were calculated to be 39.06 mm (with a deviation of 0.94 mm) and 19.38 mm (with a deviation of 0.62 mm), as shown in Fig. 5a and b. The angle between the y channels has a tolerance of 0.15°, and the width of the y channel was measured to be 0.49 mm (Fig. 5c and d). The width of the middle channel was 0.50 mm, and the diameter of the inlets/outlets was 1.99 mm, with a deviation of 0.01 mm (Fig. 5e and f). To support micro-CT data, the widths of the microchannels were measured using scanning electron microscopy (Fig. S2a and b), and the height of the microchannel was measured with surface profilometry, as illustrated in Fig. S2c.
UV curing as a post-processing tool has a significant impact on the mechanical properties and overall performance of the 3D-printed microfluidic device. During the curing process, exposure to ultraviolet light initiates further polymerization of the printed resin, enhancing crosslinking within the material. This results in increased stiffness, strength, and dimensional stability of the printed parts. As curing time increases, mechanical properties such as tensile strength and modulus improve, while the ductility tends to decrease due to the formation of a more rigid polymer network. Additionally, UV curing enhances the part's surface hardness and reduces its tackiness, making it more suitable for functional applications. However, excessive UV exposure can lead to brittleness and reduced toughness, highlighting the importance of optimizing curing time to balance strength and flexibility in the LCD-printed components. When uncured, the device has the lowest strength (4.99 ± 0.31 MPa), indicating a soft and flexible device with a Young's modulus of 156.4 MPa and toughness of 0.181 MJ m−3. The strength and modulus gradually increased to 21.47 ± 1.03 MPa and 1298.9 MPa, respectively, upon further curing for 10 min. Nonetheless, after a curing time of 5 min, a decline in the toughness value is observed with only a slight improvement in device strength. Hence, we selected 5 min as the optimised curing time, giving a strength of 17.56 ± 1.16 MPa (n = 3, p < 0.001). An image of the UTM machine and the mechanical strength data are shown in Fig. 5g and h, detailed parameters are listed in Table S2, and the deviations in the strength values are shown in Fig. S3.
3.2. Total petroleum hydrocarbon degradation by the biofilm-coated 3D-printed microchannel
The viability of the biofilm formed on the surface was studied by the standard plate counting method, and results were obtained in terms of cfu per mL per surface. It was observed that the viability of cells on the biofilm coated on the 3D-printed surface increased with the number of days (Fig. 6a). This was due to the fact that a greater number of cells were attached to the surface when incubated for 15 days (1.5 × 103 cfu per mL per surface). Although an increase in the number of cells was observed from day 3 to day 7 of biofilm formation, it reached an almost stationary phase from day 7 to day 15. The mature biofilm obtained over a period of 15 days was mostly comprised of an increased population density of bacterial cells.41 Moreover, on plating the biofilm-forming consortia on nutrient agar plates, different types of colonies with varying morphology, texture, colour, and size were seen (Fig. 6b). This indicates the simultaneous growth and subsequent biofilm formation by the microbial consortia. | ||
| Fig. 6 (a) Viability of cells in the biofilm-coated microchannel and control, and (b) growth of the microbial consortium showing the co-culture of different bacteria on the same nutrient plate. | ||
The formation of biofilm on the surface of the 3D-printed microchannels was monitored by electron microscopy. The electron micrograph taken after 168 hours (Fig. 7a) shows confluent biofilm formation over the 3D-printed surface. At a higher magnification of 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 00× (Fig. 7b), it was possible to observe a network composed of individual rod-shaped cells in the biofilm. These cells are primarily connected to one another through EPS, which maintains the structural integrity of the biofilm42 along with facilitating irreversible attachment of the bacterial cells in the biofilm with the substrate.43 Rod-shaped cells of the biofilm were observed to be adhered to each other under 30
00× (Fig. 7b), it was possible to observe a network composed of individual rod-shaped cells in the biofilm. These cells are primarily connected to one another through EPS, which maintains the structural integrity of the biofilm42 along with facilitating irreversible attachment of the bacterial cells in the biofilm with the substrate.43 Rod-shaped cells of the biofilm were observed to be adhered to each other under 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000× magnification, as shown in Fig. 7c. In contrast, no such biofilm formation can be seen on the control surface (Fig. 7f), which only shows the intrinsic structural features of the 3D-printed surface. Furthermore, a dense matrix of biofilm growth (Fig. 7d) was observed after 360 hours of biofilm formation over the 3D-printed surface. The cells were seen to be densely connected to each other (Fig. 7e) after 360 hours of biofilm formation, with thick EPS accumulation, corresponding to a higher rate of biofilm formation with time. Biofilm formation is generally favoured on rough hydrophobic substrates, which generate a conditioning layer allowing for modification of the substratum's surface, facilitating attachment of cells onto them. Moreover, the exopolymers or EPS produced by the bacterial cells attach to the substrates to form the conditioning layer, which further helps in cell adhesion.
000× magnification, as shown in Fig. 7c. In contrast, no such biofilm formation can be seen on the control surface (Fig. 7f), which only shows the intrinsic structural features of the 3D-printed surface. Furthermore, a dense matrix of biofilm growth (Fig. 7d) was observed after 360 hours of biofilm formation over the 3D-printed surface. The cells were seen to be densely connected to each other (Fig. 7e) after 360 hours of biofilm formation, with thick EPS accumulation, corresponding to a higher rate of biofilm formation with time. Biofilm formation is generally favoured on rough hydrophobic substrates, which generate a conditioning layer allowing for modification of the substratum's surface, facilitating attachment of cells onto them. Moreover, the exopolymers or EPS produced by the bacterial cells attach to the substrates to form the conditioning layer, which further helps in cell adhesion.
During our investigation, we observed biofilm formation starting from day 3 of the bacterial consortium incubation, with a continuous rise detected until day 15. Upon reaching day 15, a stationary phase becomes apparent (as shown in Fig. 6a), indicating that the creation of the biofilm reached a saturation point after a certain period of time.43 The SEM images in Fig. 7f reveal the inherent structural roughness of the printed microchannel surface, which is a favourable characteristic for cell adhesion. The formation of a conditioning layer contributes to the increase in surface roughness, facilitating the creation of a dense biofilm over the 3D-printed microchannel. Furthermore, the biofilm exhibited a noticeable growth in the form of progressively thicker and denser layers, observed from day 5 to day 7. Following the successful establishment of a biofilm, these 3D-printed microchannels were employed to investigate the degradation of petroleum hydrocarbons.
These bacterial cells adhere to the surface in the form of a biofilm as the 3D-printed microchannel surface is incubated in a glucose mineral salt medium in the presence of the microbial consortium. The absence of a biofilm layer on the blank sample prevented the PRS particles from adhering to the 3D-printed surface during the experiment. With this observation, it can be said that the biofilm coating on the surface is responsible for hydrophobicity, because of which the PRS particles are getting attached to it. As previously reported,24 biofilms confer overall hydrophobicity to the cell surface, which facilitates the utilization of TPH by their degradation.24 Therefore, PRS attachment to the biofilm-coated surface results in the ultimate solubilisation of TPH present in PRS, which leads to their utilization by bacterial cells as a source of nutrients for growth.
The compositional analysis of PRS was previously reported using solvent–solvent extraction.19 The TPH content of the PRS sludge was found to be 180.6 ± 3.4 g kg−1. Asphaltene content was found to be 50% of TPH, and the maltene content was 50%, containing aromatic, aliphatic and polar components. As observed in Fig. 8, TPH degradation gradually increased from day 0 to day 15 as compared to the control. After day 15, approximately 82% ± 6% of the TPH was found to be degraded in treated samples, compared with 3.85 ± 0.21 g mL−1 TPH extracted from the control samples, as seen in Table 2. There is almost no increase in degradation of TPH beyond 15 days (extended up to 30 days). Hence, 15 days is the optimum time required for the degradation process. This indicates that TPH degradation mostly occurs due to microbial activity in the biofilm-coated microfluidic surface, whereas an almost negligible amount of degradation was seen for the UV-sterilised surface (control).
| TPH degradation by biofilm coated resin | ||
|---|---|---|
| Sample | TPH extracted (g mL−1) | % Degradation |
| Control (untreated microchannel surface) | 3.85 ± 0.21 | 0 |
| 12 hours | 3.75 ± 0.20 | 2.59 ± 0.14 |
| 72 hours | 2.2 ± 0.28 | 42.85 ± 4.13 |
| 168 hours | 1 ± 0.27 | 74.02 ± 5.65 |
| 360 hours | 0.7 ± 0.28 | 82 ± 6 |
| 480 hours | 0.65 ± 0.20 | 82 ± 1 |
| 720 hours | 0.65 ± 0.20 | 82 ± 1 |
This indicates that TPH degradation mostly occurs due to microbial activity in the biofilm-coated microchannel, whereas in the control with the UV-sterilised microchannel and autoclaved PRS, an almost negligible amount of degradation was seen (Fig. 8). Biofilm-containing microbial cells start growing on PRS-containing medium, utilizing PRS as the sole carbon and nitrogen source, thereby degrading the hydrocarbons present in PRS. These hydrocarbons are mostly metabolised by the consortium of bacteria to meet their energetic demands by acting as a source of nutrients.28 The development of the consortium helps to achieve this degradation at a faster rate than when using a single bacterial species. Moreover, the development of a microbial consortium is important when targeting total hydrocarbon degradation, as single bacterial species are not capable of utilizing the mixture of hydrocarbons.
In addition, the gravimetric results were supported by the GC-MS chromatograms (Fig. 9a–d). When compared to the negative control (untreated), displayed in Fig. 9a, the number of peaks in the treated samples reduced significantly from day 3 to day 7 of treatment (Fig. 9b and c). On the 15th day, as seen in Fig. 9d, only a few peaks were detectable in the treated sample. Hydrocarbon degradation was progressively visible from day 3 through day 15, when compared with the negative control (day 0), as peak intensities were seen to decrease with the duration of treatment. Analysis of GC MS data shows that the control or untreated sample exhibited high concentrations of C10–C50 hydrocarbons (Table S3). On treatment with the biofilm-coated surface on the 3rd day, C41–C50 peaks were not seen. On the 15th day of the treatment, only C10–C20 (Tables S4 and S5) peaks, which pertained mostly to aromatics, were visible. This persistence of some aromatic hydrocarbons may be attributed to preferential utilization of aliphatic hydrocarbons by the consortium at a faster rate, probably owing to the greater bioavailability of the aliphatics as carbon sources.
4. Conclusion
This study centers on the fabrication of a low-cost and portable LCD-printed microfluidic device employing a custom-made resin that facilitates biofilm-based bioremediation of petroleum hydrocarbons. The biofilm is formed by a microbial consortium consisting of four strains isolated from petroleum sludge (Dietzia sp. IRB191, Dietzia sp. IRB192, Staphylococcus sp. BSM19, and Stenotrophomonas sp. IRB19). It provides an added advantage in terms of providing a platform for biodegradation as compared to the conventional approaches. This is crucial for future enhancements of PAH bioremediation strategies employing bacterial biofilms and biostimulation techniques. The fabricated microfluidic device has a dimensional error of 0.01 mm and a mechanical strength of 18.57 MPa, as confirmed from micro-CT and mechanical testing. With the aid of the biofilm-coated 3D-printed microfluidic carrier platform, almost 82% ± 6% of TPH was removed from PRS through 15 days of treatment. Correspondingly, there is an enhancement in the roughness value to 162 nm after 15 days, owing to the accumulation of EPS, as confirmed by SEM analysis. A reduction of C50–C30 hydrocarbons to C10–C20 was observed from the GC-MS results. This indicates that long-chain hydrocarbons were degraded by this biofilm-coated 3D-printed microfluidic device. Our study has opened new avenues for carrier-mediated bioaugmentation, which can be effectively applied to biologically remediate oil spills in marine environments, rather than directly introducing bacterial cells that tend to readily dilute off. Thus, this study demonstrates an efficient and fool proof strategy for carrier-bound biofilm-based hydrocarbon degradation using a low-cost LCD-printed microfluidic device fabricated using a custom-made resin. The approach offers a new, cost-effective tool for other potential applications in bioremediation, such as wastewater treatment and removal of pollutants from industrial effluents, along with oil recovery in MEOR (microbial enhanced oil recovery).Conflicts of interest
There is no conflicts to declare.Data availability
Data are available upon request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra06174a.
Acknowledgements
The authors gratefully acknowledge the Indian Council of Medical Research, Government of India (IIT/SRIC/IDK-PHASE-II/2024/02 and IIT/SRIC/IDK-PHASE-II/2024/03) for the support. The authors duly acknowledge Mr Sundipan Bhowmick of the P. K. Sinha Centre for Bioenergy and Renewables, IIT Kharagpur, for his support during GC-MS experiments. Novel hydrocarbon-degrading strains were isolated by Dr Ipsita Dipamitra Behera from petroleum refinery sludge collected from Indian Oil Corporation Limited, Haldia, West Bengal, India. RS is thankful to DST for financial support (No. DST/TMD (EWO)/OWUIS-2018/RS-18). The authors thank the analytical facility of the Department of Biotechnology, IIT Kharagpur, for the instrumentation facility. AD acknowledges the Department of Biotechnology, Govt. of India, for providing financial support through a DBT-JRF fellowship (Fellow No. DBT/2018/IIT-KH/1012), New Delhi, India. The author acknowledge the support from Mr. Sayak Mitra during the revision of the manuscript.References
- G. M. Whitesides, Nature, 2006, 442, 368–373 CrossRef CAS PubMed.
- H. Gong, M. Beauchamp, S. Perry, A. T. Woolley and G. P. Nordin, RSC Adv., 2015, 5, 106621–106632 RSC.
- C. I. Rogers, K. Qaderi, A. T. Woolley and G. P. Nordin, Biomicrofluidics, 2015, 9, 1–9 CrossRef PubMed.
- C. M. B. Ho, S. H. Ng, K. H. H. Li and Y. J. Yoon, Lab Chip, 2015, 15, 3627–3637 RSC.
- Z. Luo, H. Zhang, R. Chen, H. Li, F. Cheng, L. Zhang, J. Liu, T. Kong, Y. Zhang and H. Wang, Microsyst. Nanoeng., 2023, 9, 1–13 Search PubMed.
- Z. Weng, X. Huang, S. Peng, L. Zheng and L. Wu, Nat. Commun., 2023, 14, 1–9 Search PubMed.
- S. Waheed, J. M. Cabot, N. P. Macdonald, T. Lewis, R. M. Guijt, B. Paull and M. C. Breadmore, Lab Chip, 2016, 16, 1993–2013 RSC.
- N. Bhattacharjee, A. Urrios, S. Kang and A. Folch, Lab Chip, 2016, 16, 1720–1742 RSC.
- S. Waheed, J. M. Cabot, N. P. Macdonald, T. Lewis, R. M. Guijt, B. Paull and M. C. Breadmore, Lab Chip, 2016, 16, 1993–2013 RSC.
- S. Razavi Bazaz, O. Rouhi, M. A. Raoufi, F. Ejeian, M. Asadnia, D. Jin and M. Ebrahimi Warkiani, Sci. Rep., 2020, 10, 1–14 Search PubMed.
- C. Chen, B. T. Mehl, A. S. Munshi, A. D. Townsend, D. M. Spence and R. S. Martin, Anal. Methods, 2016, 8, 6005–6012 RSC.
- G. G. Morbioli, N. C. Speller and A. M. Stockton, Anal. Chim. Acta, 2020, 1135, 150–174 CrossRef CAS PubMed.
- E. J. Carrasco-Correa, E. F. Simó-Alfonso, J. M. Herrero-Martínez and M. Miró, TrAC, Trends Anal. Chem., 2021, 136, 11–13 CrossRef.
- A. K. Au, W. Huynh, L. F. Horowitz and A. Folch, Angew. Chem., Int. Ed., 2016, 55, 3862–3881 CrossRef CAS PubMed.
- H. Gong, A. T. Woolley and G. P. Nordin, Lab Chip, 2016, 16, 2450–2458 RSC.
- P. F. Jacobs, J. Manuf. Syst., 1993, 12, 430–433 CrossRef.
- H. Gong, B. P. Bickham, A. T. Woolley and G. P. Nordin, Lab Chip, 2017, 17, 2899–2909 RSC.
- A. Amini, R. M. Guijt, T. Themelis, J. De Vos and S. Eeltink, J. Chromatogr. A, 2023, 1692, 1–14 CrossRef PubMed.
- E. Skliutas, M. Lebedevaite, S. Kasetaite, S. Rekštytė, S. Lileikis, J. Ostrauskaite and M. Malinauskas, Sci. Rep., 2020, 10, 1–9 Search PubMed.
- J. L. Sanchez Noriega, N. A. Chartrand, J. C. Valdoz, C. G. Cribbs, D. A. Jacobs, D. Poulson, M. S. Viglione, A. T. Woolley, P. M. Van Ry, K. A. Christensen and G. P. Nordin, Nat. Commun., 2021, 12, 1–13 Search PubMed.
- A. Pongwisuthiruchte, S. T. Dubas, C. Aumnate and P. Potiyaraj, Sci. Rep., 2022, 12, 1–10 Search PubMed.
- Q. Ge, B. Jian and H. Li, Forces Mech., 2022, 6, 1–12 Search PubMed.
- H. B. Musgrove, M. A. Catterton and R. R. Pompano, Anal. Chim. Acta, 2022, 1209, 339842 CrossRef CAS PubMed.
- Vandana and S. Das, J. Hazard. Mater., 2023, 457, 131795 CrossRef CAS PubMed.
- D. Sarkar, M. Ferguson, R. Datta and S. Birnbaum, Environ. Pollut., 2005, 136, 187–195 CrossRef CAS PubMed.
- R. M. Atlas and T. C. Hazen, Environ. Sci. Technol., 2011, 45, 6709–6715 CrossRef CAS PubMed.
- N. Das and P. Chandran, Biotechnol. Res. Int., 2011, 2011, 1–13 Search PubMed.
- X. Xu, W. Liu, S. Tian, W. Wang, Q. Qi, P. Jiang, X. Gao, F. Li, H. Li and H. Yu, Front. Microbiol., 2018, 9, 1–11 Search PubMed.
- S. Guo, X. Liu and J. Tang, Chemosphere, 2022, 286, 131663 CrossRef CAS PubMed.
- A. Ganesh Kumar, N. Nivedha Rajan, R. Kirubagaran and G. Dharani, Mar. Pollut. Bull., 2019, 146, 741–750 CrossRef CAS PubMed.
- V. Catania, F. Lopresti, S. Cappello, R. Scaffaro and P. Quatrini, New Biotechnol., 2020, 58, 25–31 CrossRef CAS PubMed.
- Y. Liang, X. Zhang, D. Dai and G. Li, Int. Biodeterior. Biodegrad., 2009, 63, 80–87 CrossRef CAS.
- Y. Xu and M. Lu, J. Hazard. Mater., 2010, 183, 395–401 CrossRef CAS PubMed.
- P. Galitskaya, L. Akhmetzyanova and S. Selivanovskaya, Biogeosciences, 2016, 13, 5739–5752 CrossRef CAS.
- X. Fu, H. Wang, Y. Bai, J. Xue, Y. Gao, S. Hu, T. Wu and J. Sun, Environ. Sci. Ecotechnology, 2020, 2, 100028 CrossRef PubMed.
- D. Hou, X. Shen, Q. Luo, Y. He, Q. Wang and Q. Liu, Mar. Pollut. Bull., 2013, 67, 146–151 CrossRef CAS PubMed.
- I. D. Behera, G. Basak, R. R. Kumar, R. Sen and B. C. Meikap, J. Environ. Sci. Health, Part A: Toxic/Hazard. Subst. Environ. Eng., 2020, 56, 226–239 Search PubMed.
- I. D. Behera, M. Nayak, S. Biswas, B. C. Meikap and R. Sen, J. Environ. Manage., 2021, 292, 112746 CrossRef CAS PubMed.
- Y. Cai, C. Yu, S. Zhong, G. Chen and L. Liu, J. Environ. Chem. Eng., 2023, 11, 110101 CrossRef CAS.
- S. Al-Amshawee, M. Y. B. M. Yunus, J. G. Lynam, W. H. Lee, F. Dai and I. H. Dakhil, Environ. Technol. Innovation, 2021, 21, 101233 CrossRef.
- J. A. Cho, Y. J. Roh, H. R. Son, H. Choi, J. W. Lee, S. J. Kim and C. H. Lee, Sci. Rep., 2022, 12, 1–11 Search PubMed.
- P. Di Martino, AIMS Microbiol., 2018, 4, 274–288 CAS.
- G. S. Lorite, C. M. Rodrigues, A. A. de Souza, C. Kranz, B. Mizaikoff and M. A. Cotta, J. Colloid Interface Sci., 2011, 359, 289–295 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to the work. |
| This journal is © The Royal Society of Chemistry 2026 |