Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Designing jasmine lactone copolymer micelles for drug delivery: influence of ionic group density and chain length
Vishal
Kumar
ab,
Elisa
Léonard†
ab,
Atefeh
Atefi
ac,
Jessica M.
Rosenholm
 a,
Carl-Eric
Wilén
b,
Shakhawath
Hossain
a,
Carl-Eric
Wilén
b,
Shakhawath
Hossain
 *d and
Kuldeep K.
Bansal
*d and
Kuldeep K.
Bansal
 *ab
*ab
aPharmaceutical Sciences Laboratory, Faculty of Science and Engineering Åbo Akademi University, Biocity, Tykistökatu 6A, 20520 Turku, Finland. E-mail: kuldeep.bansal@abo.fi
bLaboratory of Molecular Science and Engineering, Faculty of Science and Engineering, Åbo Akademi University, Aurum, Henrikinkatu 2, 20500 Turku, Finland
cFaculty of Science and Engineering, Cell Biology, Åbo Akademi University, Biocity, Tykistökatu 6A, 20520 Turku, Finland
dDepartment of Pharmacy and The Swedish Drug Delivery Center (SweDeliver), Uppsala University, Uppsala 751 23, Sweden. E-mail: shakhawath.hossain@uu.se
First published on 7th January 2026
Abstract
Functionalized amphiphilic polymers have been widely applied as drug delivery vehicles due to their ability to self-assemble into micelles that enhance the solubility, stability, and bioavailability of poorly water-soluble drugs. Among these, poly-jasmine lactone (PJL), a recently developed amphiphilic copolymer, offers the opportunity to functionalize with versatile functional groups via facile thiol–ene chemistry. In this study, we synthesized and compared anionic functionalized block copolymers of PJL (mPEG-b-PJL-COOH) having varying numbers of the –COOH group to assess the effect on the encapsulation efficiency, particle size, drug release behavior, and cytotoxicity. Our results demonstrate that increasing the number of anionic groups did not improve the encapsulation efficiency of model drugs but sustained the drug release profile. Ex vivo hemolytic studies were also performed to evaluate pH-dependent cell lysis as an indirect indicator of the endosomal escape capability of the prepared micelles. Coarse-grained molecular dynamics simulations also revealed that increasing the number of –COOH groups altered the structural properties of the lipid bilayer. Moreover, the aggregation of –COOH units within the lipid bilayer may represent the molecular mechanism underlying the higher cytotoxicity observed with a greater number of –COOH groups.
Introduction
The advancement of high-throughput screening technologies shortens the drug development process, but drug's physicochemical properties, such as solubility, stability, and potential threat of toxicity, limit their progress as effective clinical drug formulations.1 Various strategies, including use of polymeric nanoparticles as drug delivery carriers, have been explored to overcome these limitations. In this context, amphiphilic block copolymers are gaining the interest of scientists because they can self-assemble and form a core–shell type of nanoparticle called polymeric micelles.2 Despite the several advantages polymeric micelles hold, their low drug loading and potential instability limit their applications in drug delivery.3 To overcome these problems, polymer structures with functionally active groups have been developed to enhance the drug loading and stability through ionic or covalent bonds.4,5 Furthermore, such functional polymers allow the generation of controlled, targeted, and stimuli-responsive drug delivery systems.6,7 For instance, quercetin and vancomycin encapsulated in surface-modified Pluronic F127 micelles demonstrated reduced burst release and improved antibacterial activity against S. Aureus compared to free vancomycin.8 Similarly, γ-functionalized PCL poly{γ-2-[2-(2-methoxyethoxy)ethoxy]ethoxy-ε-caprolactone}-b-poly(γ-benzyloxy-ε-caprolactone) or (PME3CL-b-PBnCL) showed the higher loading and delayed release of doxorubicin compared to poly{γ-2-[2-(2-methoxyethoxy)ethoxy]ethoxy-ε-caprolactone}-b-poly(γ-(4-ethoxylphenyl)-ε-caprolactone) or (PME3CL-b-PEtOPhCL), which possesses higher structural rigidity and larger steric hindrance due to the aromatic functional group.9Moreover, the hydrophobicity of the functional groups on such polymers affects their properties, such as drug loading capacity. Recently, He et al. reported that the hydrophobic group on the hydrophobic block (HOOC-PNAM-PBMA-Phen) enhanced the loading and release performance of paclitaxel in comparison to the hydrophobic group on the hydrophilic block (HOOC-PBMA-PNAM-Phen) containing a higher number of carboxylic groups.10 Similarly, Yuan et al. demonstrated that the doxorubicin-loaded micelles of chitosan-cystamine-acetylamine (LC-Cys-CA) with a higher hydrophobic chain length showed superior loading and improved cellular internalization in A549 and L929 cells in comparison to the chitosan-cystamine-octylamine (LC-Cys-OA) having a lower hydrophobic chain length.11
Furthermore, the application of such functionalized amphiphilic polymers is not limited to solubility enhancement; research is also exploring the potential of these functionalized amphiphilic polymers for intracellular delivery and controlled release of bioactive molecules, enabling them to act within the cells. Such macromolecules usually translocated into cells by endocytosis are encapsulated within the endosomes and are trafficked to the lysosomal compartment for maturation, diminishing the therapeutic response.12 Therefore, a delivery vehicle that can escape the endosomes and avoid lysosomal degradation is crucial for effective drug delivery to the target organelles of cells. Cationic lipids like 1,2-di-O-octadecenyl-3-trimethylammonium propane and polymers such as polyethylenimine can escape endosomes but pose challenges including cellular damage and toxicity.13,14 In contrast, ionizable lipids like 1,2-di-O-octadecenyl-3-dimethylaminopropane remain noncharged, enhancing safety and endosomal escape. However, they may disrupt cellular signalling and metabolism, leading to cellular toxicity, imposing continued research to develop vehicles that can accomplish effective endosomal escape with less toxicity.15 In this regard, our research group is exploring the anionic (–COOH) functionalized, patent-protected poly-jasmine lactone (mPEG-b-PJL-COOH) to improve formulation performances.16–18
Molecular dynamics (MD) simulation is a powerful computational method used in drug development to ascertain the atomic behavior of drug molecules and their interaction with the excipients and human cells. Additionally, it provides information about energetic processes, such as binding, conformational changes, and aggregation of drug formulations, that are difficult to analyze experimentally. MD simulation is a rapid, cost-effective method in the drug development process.19,20
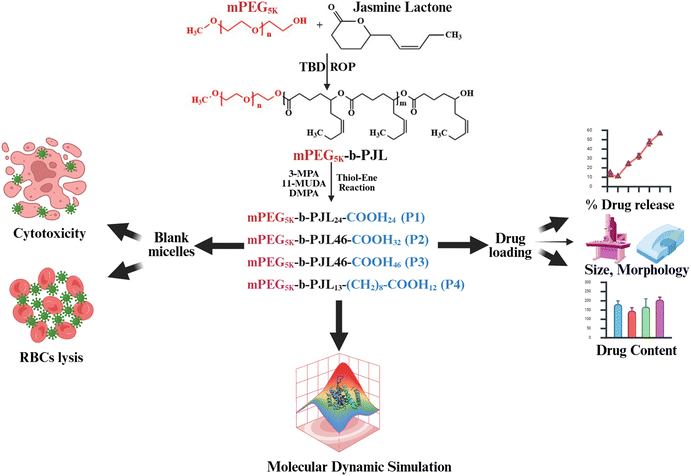
Thus, in this study, we aimed to evaluate the effect of the varied number of the anionic (–COOH) group on the loading and release of the interactively favored basic drug trimethoprim (TMP) and the unfavored acidic photosensitizer chlorine e6 (Ce6). Amphiphilic block copolymers of the jasmine lactone were synthesized and functionalized to achieve varied numbers of anionic (–COOH) groups and chain lengths (Fig. 1 shows the synthetic scheme with experimental work). Additionally, to assess the effect of the hydrophobicity on the drug loading and release performance, one polymer was functionalized using a functional group having a comparatively high aliphatic CH2 chains adjacent to the –COOH group. Moreover, we investigated the pH-responsive membrane disruption properties on RBCs. In addition, molecular dynamics simulations were performed to validate and understand how the polymer structural difference will affect drug loading and analyze the polymer–cell membrane interaction mechanism.
Results and discussion
Synthesis of the block copolymer of jasmine lactone (mPEG-b-PJL)
Two different block copolymers, polymer A (PA) and polymer B (PB), were synthesized by the ring-opening polymerization technique (Scheme S1) with varied monomer/initiator ratios following the reported procedure.21 The average molar mass was calculated by 1HNMR analysis (Fig. S1A and B) by interpreting and comparing the protons of the methylene group of mPEG5K (initiator) at 3.3 ppm and the protons of PJL at 4.9 ppm with respect to the protons of the methylene group of PJL at 0.98 ppm (Table 1). The number-average molar mass (Mn), weight-average molar mass (Mw), and mass distribution (polydispersity (Đ), Mw/Mn) obtained for polymers by high-performance size exclusion chromatography (HPSEC) (Fig. S2) are depicted in Table 1. The characterization results suggested successful synthesis and purification of polymers.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) I (monomer to initiator ratio), Mn (number-average molar mass), Mw (weight-average molar mass), and Đ (polydispersity), NMR (nuclear magnetic resonance), CMC (critical micelle concentration), NA (not applicable), NE (not evaluated)
I (monomer to initiator ratio), Mn (number-average molar mass), Mw (weight-average molar mass), and Đ (polydispersity), NMR (nuclear magnetic resonance), CMC (critical micelle concentration), NA (not applicable), NE (not evaluated)
| SN | Polymers | M![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) I I |
% Rxn | Number of COOH groups | Mol wt. by 1HNMR (kDa) | Mol wt. (Mn) by HPSEC (kDa) | Mol wt. (Mw) by HPSEC (kDa) | Đ (Mw/Mn) | CMC (μg mL−1) |
|---|---|---|---|---|---|---|---|---|---|
| a Results are published previously by our group but included to understand the difference between each polymer used in this study.16,21 | |||||||||
| 1 | PA | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
54 | NA | 7.5 | 6.0 | 11.4 | 1.8 | NE |
| 2 | PB | 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
70 | NA | 12.9 | 16.3 | 19.4 | 1.1 | NE |
| 3 | P1a | NA | 100 | 24 | 11.4 | 9.8 | 16.9 | 1.4 | 10.2 ± 3.9 |
| 4 | P2 | NA | 70 | 32 | 16.2 | 14.8 | 32.9 | 2.2 | 10.7 ± 2.9 |
| 5 | P3 | NA | 100 | 46 | 17.5 | 15.3 | 22.4 | 1.4 | 12.7 ± 3.9 |
| 6 | P4 | NA | 90 | 12 | 9.8 | 13.8 | 34.7 | 2.4 | 11.4 ± 2.7 |
Functionalization of mPEG-b-PJL to mPEG-b-PJL-COOH
To assess the effect of the anionic (–COOH) groups on the drug loading, release, particle size, RBC rupture, and cytotoxicity, the synthesized block copolymers mPEG-b-PJL were functionalized to three divergent polymers (acronyms P2, P3, and P4) having different numbers of ionic groups and chain lengths by the thiol–ene click reaction (Schemes S2 and S3) at room temperature, and the degree of functionalization was controlled by reaction time and monitored via1HNMR by interpreting and comparing the changes in double bond peaks (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, site of reaction) at 5.3 and 5.5 ppm. The consumption of the unsaturated carbon peak suggested the functionalization of the pendant chain of PJL and the attachment of the thiol (SH) group of 3-mercaptopropionic acid or 11-mercaptoundecanoic acid to form the new (C–S–C) bond.22 The molar mass and the number of ionic groups of the functionalized polymers P2, P3, and P4 were determined by 1HNMR spectroscopy (Table 1 and Fig. S3A, S3B, and S4) following the reported procedure of the previously synthesized and anionic (–COOH) functionalized polymer mPEG-b-PJL-COOH, which was included (acronym P1) to compare with the polymers P2, P3, and P4. Furthermore, molecular weight and its distribution were assessed by HPSEC (Table 1 and Fig. S5).
C, site of reaction) at 5.3 and 5.5 ppm. The consumption of the unsaturated carbon peak suggested the functionalization of the pendant chain of PJL and the attachment of the thiol (SH) group of 3-mercaptopropionic acid or 11-mercaptoundecanoic acid to form the new (C–S–C) bond.22 The molar mass and the number of ionic groups of the functionalized polymers P2, P3, and P4 were determined by 1HNMR spectroscopy (Table 1 and Fig. S3A, S3B, and S4) following the reported procedure of the previously synthesized and anionic (–COOH) functionalized polymer mPEG-b-PJL-COOH, which was included (acronym P1) to compare with the polymers P2, P3, and P4. Furthermore, molecular weight and its distribution were assessed by HPSEC (Table 1 and Fig. S5).
Assessment of critical micelle concentration
Critical micelle concentration (CMC) represents the concentration at which the amphiphilic polymers start assembling into micelles forming a core–shell structure.23 Upon administration into the body, the micelles undergo dilution by large blood volume/GI fluid and disrupt the self-assembly of micelles, leading to premature drug release.24 Therefore, designing an amphiphilic polymer with a low CMC value is essential for enhancing the micelle stability. To evaluate the potential of the synthesized polymers in drug delivery systems as micellar formulations, their CMC was assessed by the pyrene fluorescence probe method. Pyrene is a hydrophobic fluorescent probe that prefers to be in a hydrophobic environment in solution, so as micelles form, it moves to the hydrophobic core of the micelles, resulting in a shift in the intensity ratio, which correlates with the formation of the micelles.16,25 The CMC value for P1 (10.2 ± 3.9) has been established earlier and was found to be non-significantly different from the CMC of P2, P3, and P4 (Fig. S6) (Table 1), suggesting that the acidic functionalization compensated the hydrophobicity of P2 and P3, while the less hydrophobic block of P4 could be compensated by the hydrophobicity of aliphatic CH2 chains adjacent to the COOH group, making its CMC non-significant compared to other polymers synthesized in this study.Preparation and characterization of TMP and chlorine e6 (Ce6) loaded micelles
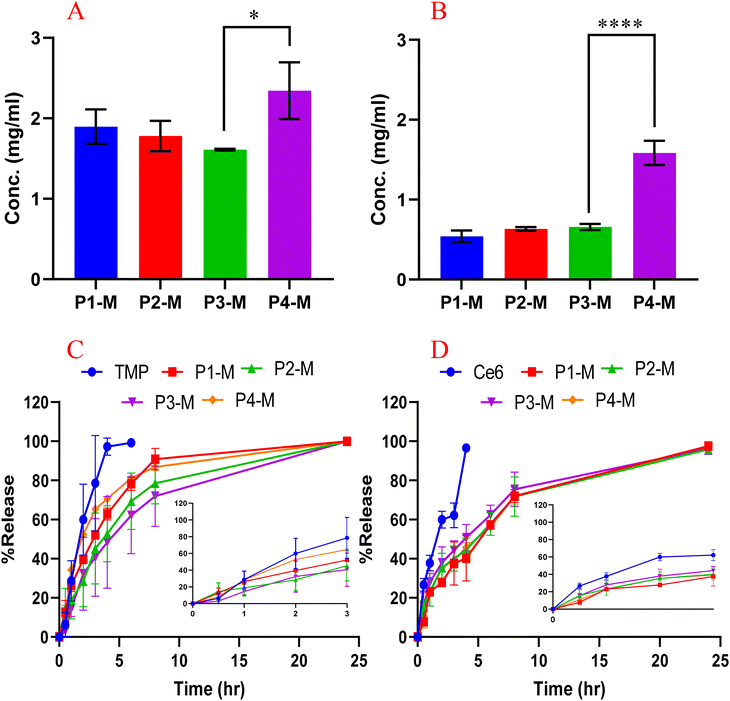
To assess the influence of carboxylic acid (–COOH) group number and polymer structure on the encapsulation efficiency, hydrophobic and interactively favored positively charged TMP and unfavored, negatively charged Ce6 were loaded in the polymeric micelles (P1-M, P2-M, P3-M, and P4-M) using the polymers P1, P2, P3, and P4, respectively. The results (Fig. 2A) revealed that the prepared micelles (P4-M) significantly enhanced the aqueous solubility of TMP and Ce6 up to 7.5-fold and 273-fold higher than their aqueous solubility i.e. 0.31 ± 0.01 mg mL−1 and 0.0052 ± 0.0004 mg mL−1, respectively. The drug content and encapsulation efficiency, which provide a measure of how effectively the drug is incorporated into the micelles, were calculated, and they follow the order P4-M > P1-M > P2-M > P3-M (based on mean values). No significant differences between P1-M, P2-M, and P3-M were observed despite the presence of different numbers of the –COOH groups. This result indicates that an increase in the number of the –COOH groups may not significantly impact the encapsulation efficiency of the micelles on the tested drugs. Although TMP is a basic drug, we observed that the hydrophobic interaction is dominant compared to the ionic interaction, as P4-M has the lowest number of –COOH groups, exhibiting significantly higher drug loading. This high loading could be attributed to the longer aliphatic CH2 chain adjacent to the –COOH groups, which imparts higher hydrophobicity to the micellar core, favoring a stronger interaction with TMP.The drug loading results of Ce6 (Fig. 2B) showed similar trends of the encapsulation, and P4-M again demonstrates the highest encapsulation efficiency and comparatively high drug content compared to other micelles.
The hydrodynamic sizes of the prepared micelles were measured using DLS, and the results are presented in Table 2 and Fig. 3. Notably, the PdI of the TMP-loaded micelles is higher than that of the Ce6-loaded micelles, which could be attributed to the second peak in intensity distribution (Fig. 3). Therefore, the size and morphology of the TMP-loaded micelles were reconfirmed by TEM (Fig. 4), and the TEM images analyzed by Image J (Fig. S7) did not reveal the presence of large-sized particles in P1-M, but the P2-M, P3-M, and P4-M samples showed the broad size range as seen in the intensity distribution by DLS.26 However, the size distribution by volume in DLS reveals that only 1.1, 0.9, and 5.5% of micelles in P2-M, P3-M, and P4-M respectively were of large size. Furthermore, the TEM images revealed the spherical shape of micelles with sizes of 17 ± 3 nm and 19 ± 5 nm for P1-M and P2-M, while the sizes for P3-M and P4-M were found to be 20 ± 5 nm and 23 ± 6 nm respectively. Overall, the prepared micelles remained in sub-50 nm size, suggesting their potential use as a carrier for drug delivery applications.
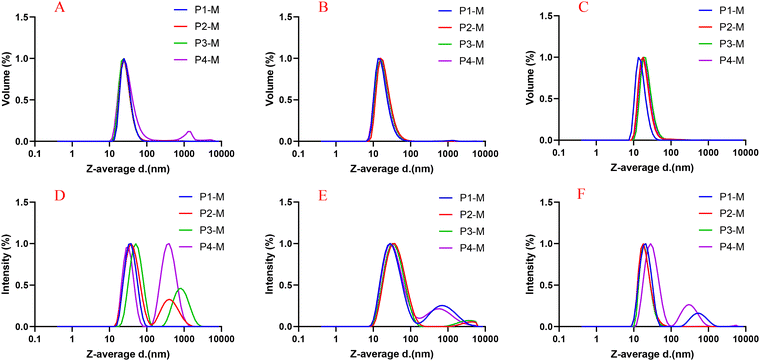 | ||
| Fig. 3 Normalized size distribution curve by volume and intensity of the TMP loaded (A and D), Ce6 loaded (B and E) and blank micelles (C and F) by DLS. | ||
| Micelles | TMP content (wt.%) (±SD) | TMP EE (%) (±SD) | Ce6 content (wt.%) (±SD) | Ce6 EE (%) (±SD) | TMP loaded Z-average size (d nm) (±SD) | Ce6 loaded Z-average size (d nm) (±SD) | Blank Z-average size (d nm) (±SD) | TMP loaded PdI (±SD) | Ce6 loaded PdI (±SD) | Blank PdI (±SD) |
|---|---|---|---|---|---|---|---|---|---|---|
| TMP (TMP), Ce6 (chlorine e6), EE (encapsulation efficiency), d nm (diameter in nanometer), and PdI (polydispersity index). | ||||||||||
| P1-M | 8.2 ± 0.05 | 59.7 ± 4.5 | 2.9 ± 0.3 | 29.9 ± 3.9 | 37 ± 2 | 37 ± 1 | 23 ± 7 | 0.19 ± 0.04 | 0.32 ± 0.05 | 0.32 ± 0.03 |
| P2-M | 8.1 ± 0.07 | 59.3 ± 6.2 | 3.0 ± 0.1 | 31.3 ± 1 | 41 ± 5 | 43 ± 3 | 33 ± 8 | 0.41 ± 0.02 | 0.2 ± 0.03 | 0.3 ± 0.02 |
| P3-M | 7.5 ± 0.04 | 53.7 ± 0.3 | 3.0 ± 0.3 | 31.2 ± 3.4 | 44 ± 1 | 38 ± 3 | 36 ± 9 | 0.42 ± 0.1 | 0.23 ± 0.02 | 0.32 ± 0.02 |
| P4-M | 10.4 ± 1.4 | 78.1 ± 11.7 | 6.6 ± 0.5 | 71.0 ± 6.8 | 103 ± 13 | 35 ± 1 | 35 ± 8 | 0.62 ± 0.03 | 0.46 ± 0.03 | 0.36 ± 0.02 |
In vitro drug release
To investigate the release profile of TMP and Ce6 from micelles, an in vitro drug release test was performed in a relevant medium. As shown in Fig. 2C, the free drug dissolved in methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DI water (1
DI water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) was completely released through the dialysis tube within 4 h. In contrast, all micellar formulations demonstrated a sustained release, and 100% release was observed within 24 h. Notably, there was no clear difference observed in the first 3 h of release of TMP from micelles compared to free drug. This could be attributed to the burst release and release of the un-encapsulated drug, as we did not separate the un-encapsulated drug soluble in water. The P1-M, P2-M, and P4-M released 51.5 ± 1%, 54.4 ± 2.6%, and 64.6 ± 1.1% of their TMP content, respectively. Interestingly, formulation P3-M displayed a markedly reduced burst effect, releasing only 27.6 ± 2.9% of TMP within the 3 h, lower than P4-M, for instance. This slow release may be attributed to the higher number of carboxylic groups in the P3 structure, which interchangeably interact with the TMP. Despite this interaction, 100% of drug release was observed from all formulations, suggesting that the presence of ionic groups on a polymeric chain can facilitate the sustained release of the encapsulated drug but did not hinder the release. A similar release pattern for Ce6 was observed as free Ce6 released completely through the dialysis membrane within 4 h, whereas the micellar formulations sustained the release, reaching 100% release within 24 h (Fig. 2D). No statistical differences were observed among the release profiles of the micellar formulations, indicating that the release kinetics were independent of the number of ionic groups in the polymer structure.
1) was completely released through the dialysis tube within 4 h. In contrast, all micellar formulations demonstrated a sustained release, and 100% release was observed within 24 h. Notably, there was no clear difference observed in the first 3 h of release of TMP from micelles compared to free drug. This could be attributed to the burst release and release of the un-encapsulated drug, as we did not separate the un-encapsulated drug soluble in water. The P1-M, P2-M, and P4-M released 51.5 ± 1%, 54.4 ± 2.6%, and 64.6 ± 1.1% of their TMP content, respectively. Interestingly, formulation P3-M displayed a markedly reduced burst effect, releasing only 27.6 ± 2.9% of TMP within the 3 h, lower than P4-M, for instance. This slow release may be attributed to the higher number of carboxylic groups in the P3 structure, which interchangeably interact with the TMP. Despite this interaction, 100% of drug release was observed from all formulations, suggesting that the presence of ionic groups on a polymeric chain can facilitate the sustained release of the encapsulated drug but did not hinder the release. A similar release pattern for Ce6 was observed as free Ce6 released completely through the dialysis membrane within 4 h, whereas the micellar formulations sustained the release, reaching 100% release within 24 h (Fig. 2D). No statistical differences were observed among the release profiles of the micellar formulations, indicating that the release kinetics were independent of the number of ionic groups in the polymer structure.
Cytotoxicity assay of blank micelles
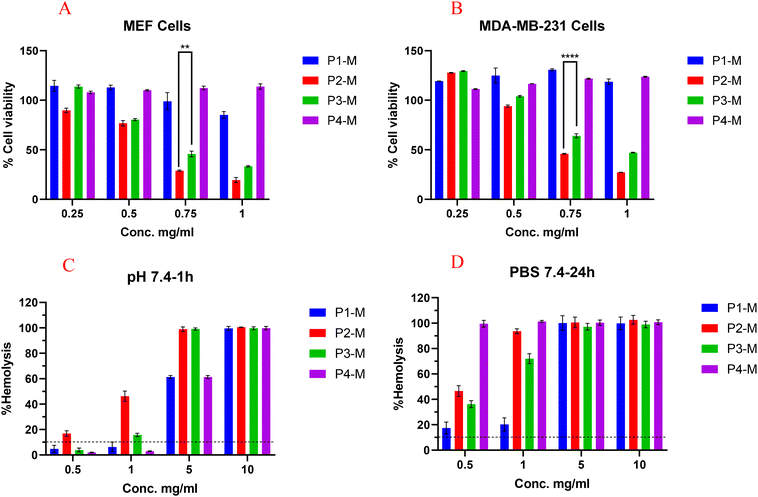
Cytotoxicity assessment is critical for verifying the safety and biocompatibility of polymeric excipients in drug delivery systems, with cell culture assays offering initial indications of potential toxicity. In this work, the cytotoxicity of the blank micelles of the synthesized polymers was evaluated on MEF (primary cell) and MDA-MB-231 cell lines by using the Alamar Blue cell viability assay. The results (Fig. 5A) demonstrated that in MEF cells, P4-M showed excellent biocompatibility up to the tested concentration (1 mg mL−1). In contrast, P1-M was found to be biocompatible up to 0.75 mg ml−1, but at 1 mg ml−1 it caused a 15% reduction in cell viability, while P2-M and P3-M exhibited a dose-dependent toxicity, where P2-M exhibited mild cytotoxicity with a 10% reduction in viability at the tested lower concentration (0.25 mg mL−1). P3-M caused noticeable cytotoxicity at 0.5 mg mL−1, likely due to its higher number of COOH groups. In our previous study, we compared functionalized (mPEG-b-PJL-COOH) and non-functionalized (mPEG-b-PJL) polymers and found that –COOH incorporation increases polymeric micelles’ cytotoxicity.16 The present results further support this trend, as polymers with less –COOH groups (P1-M and P4-M) exhibit lower toxicity than P3-M. The cytotoxicity results on the MDA-MB-231 (Fig. 5B) cell line reveal a similar phenomenon where P4-M exhibits biocompatibility up to the highest tested concentration of 1 mg mL−1. A similar toxicity trend was observed for P1-M, whereas P2-M and P3-M induced marked cytotoxic effects, resulting in approximately 55% and 35% reductions in cell viability, respectively, at 0.75 mg mL−1. In general, the toxicity trend seems similar in both cell lines, where P2-M and P3-M were found to be toxic above 0.5 mg mL−1 concentration and demonstrate higher toxicity in MEF compared to MDA-MD-231, probably due to the higher proliferation rate of cancer cells. The concentration-dependent toxicity observed for P2-M and P3-M could be directly attributed to the presence of higher -COOH groups compared to P1 and P4.Ex vivo hemolytic study of blank micelles
A hemolytic study is the assessment of the potential toxicity of the compounds on red blood cells, which provides insight into the safety of the excipients intended to be used intravenously.27,28 We analyze the hemocompatibility of blank polymeric micelles up to 10 mg ml−1. As shown in Fig. 5C, after 1 h incubation at physiological pH, blank micelles of P1-M and P4-M were found to be hemocompatible up to 1 mg ml−1, suggesting good short-term blood compatibility. However, at 24 h (Fig. 5D), all the polymers cause marked hemolysis even at the tested lower concentration i.e., 0.5 mg ml−1.In addition, their cell membrane disruption potential at low concentrations was investigated at pH 7.4, 6.0, 5.5, and 5.0. The purpose of this study was to understand the vesicle rupture potential of anionic micelles in an environment mimicking the different cellular compartments (endosomes/lysosomes), which provides an estimate relevant to endosomal escape.29 The principle of this assay relies on the pH-dependent interactions of micelles (steered by ionic groups) that disrupt the membrane of RBCs, resulting in the release of hemoglobin. Ideal excipients with the potential of endosomal escape should not cause disruption of more than 10% at physiological pH, but 100% disruption of the RBCs should be observed at acidic endosomal pH.16
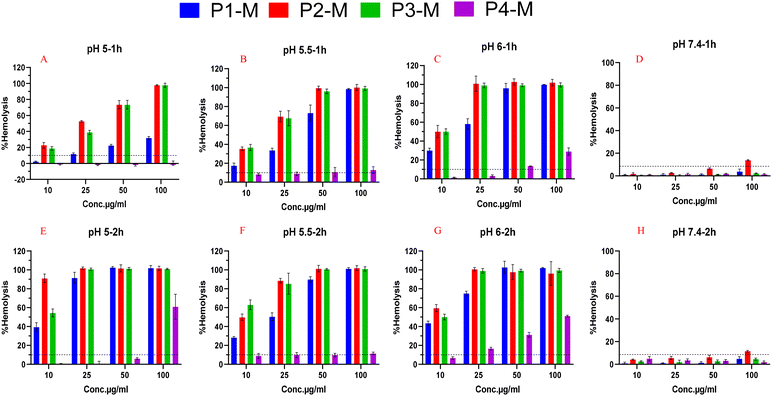
Upon incubation of micelles for up to 2 h at pH 6 (early endosomes) to pH 5 (late endosomes) (where endosomal escape should occur), it was demonstrated (Fig. 6) that P2-M and P3-M cause 100% cell lysis at 25 μg mL−1, while causing minimal hemolytic activity at physiological pH (7.4). Additionally, P1-M also demonstrated sufficient RBC disruption, but at a comparatively higher concentration (50 μg mL−1). In contrast, P4-M needed the highest concentration (500 µg mL−1) to achieve RBC disruption, suggesting its lower membrane-disruptive potential. The higher membrane-disrupting behaviour of P2-M and P3-M at relatively lower concentrations could be due to the higher number of –COOH groups, which, at endosomal pH, accept hydrogen ions (H+) and become protonated, thereby enhancing the hydrophobicity, increasing the interaction with the membrane, and promoting the endosomal permeability. These results indicated their potential application in gene delivery and are supported by previously published studies, where Wang et al. developed pH-responsive –COOH-terminated methacrylic acid and butyl methacrylate-based block copolymers that are prone to disrupting lipid membranes at endo/lysosomal pH.30,31 In addition, Chen et al. reported that hydrophobic amino acid grafts containing –COOH groups significantly enhance pH-responsive membrane disruption and facilitate endosomal destabilization.32 Overall, P2-M and P3-M were found to be the most suitable polymeric micelles, which disrupt the RBC membrane at low concentrations at all tested endosomal pH and could be promising alternative candidates to commercially available cationic polymers such as polyethylenimine (PEI) and poly(beta-amino ester) (pBAE), which demonstrate noticeable toxicity.33,34
Coarse-grained molecular dynamics simulations
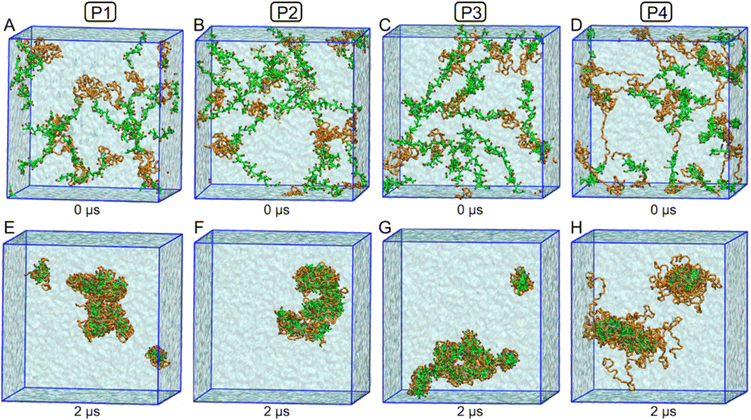
To investigate the aggregation behavior, we randomly placed multiple polymer molecules in a simulation box to approximately mimic the 20 mg mL−1 concentration. Due to differences in molecular weight, the number of molecules varied for each polymer: 14 for P1, 10 for P2, 8 for P3, and 18 for P4. Each simulation was run for 2 microseconds. During the simulations, we observed that the polymers started assembling to form micelles rapidly, within a few hundred nanoseconds. Fig. 7 shows the initial snapshots of the polymers randomly placed in the simulation box (A–D), along with the final snapshots illustrating the micellar structures formed by each polymer (E–H).
To understand the structural characteristics of the micelles, we calculated the average number of contacts between the PEG and COOH units of the polymers, both among themselves and with surrounding water beads (Fig. S8). Across all systems, we found that COOH–COOH contacts were significantly more frequent than other interactions. This indicates that the COOH units preferentially aggregate to form the hydrophobic core of the micelle, while the PEG chains are oriented toward the water interface, stabilizing the micelle in solution.
Further analysis was performed on the solvent-accessible surface area (SASA) of each polymer during the simulation, with a specific focus on the hydrophobic SASA. For this calculation, we excluded the hydrophilic mPEG beads and the charged beads representing the carboxylate groups of the −COOH units. As expected, the formation of micelles led to a reduction in the hydrophobic SASA of the –COOH regions across all polymer systems (Fig. S9A). Among the polymers, P3 exhibited the highest total hydrophobic SASA, primarily due to the higher number of –COOH units present (Table S1 for detailed polymer and –COOH unit counts). P1 and P2 showed similar SASA values, corresponding to their comparable numbers of –COOH units, while P4 exhibited the lowest values due to having the fewest –COOH units. Interestingly, when we normalized SASA per –COOH unit (Fig. S9B), we found that –COOH groups with longer aliphatic CH2 chains adjacent to the carboxyl group had the highest SASA values. Among the systems, P4 (with long-chain –COOH units) showed the highest per-unit SASA, followed by P1. The lowest values were observed for P3 and P2, respectively.
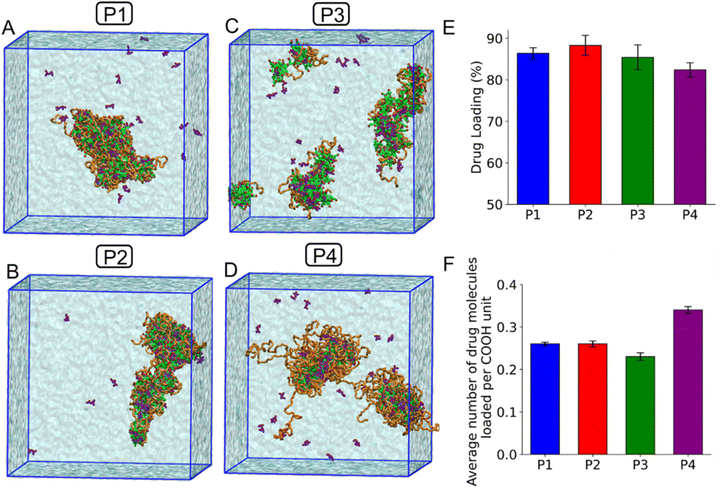
We then extracted the final snapshots of the four polymeric systems, as shown in Fig. 7(E–H) for polymers P1 to P4, respectively. To each system, we added 3 mg of TMP molecules randomly within the simulation box (Fig. S10) and conducted simulations for 1 microsecond. During the simulations, the drug molecules began interacting with the polymeric micelles and gradually attached to them. We monitored the number of TMP molecules bound to the polymers over time and observed that the number of attached drug molecules reached a plateau around 500 ns. The final configurations of the drug-loaded polymeric micelles are shown in Fig. 8(A–D) for the P1, P2, P3, and P4 systems, respectively.
We also quantified the percentage of drug loading for each polymer system by identifying whether the TMP molecules were interacting with mPEG units, –COOH units, or a combination of both. Across all systems, we found that mPEG units alone accounted for only a small fraction of the drug binding (Fig. S11). –COOH only interactions loaded ∼10–40% of the drug molecules, whereas the combination of COOH and mPEG interactions dominated the overall drug loading. This aligns with previous findings showing that mPEG chains are hydrophilic and tend to interact preferentially with water molecules rather than hydrophobic molecules.35
To evaluate drug loading more quantitatively, we calculated the average number of TMP molecules bound to the polymers over the last 200 ns of the simulations (Fig. 8E). The overall percentage of drug loading ranged between 82% and 89%, with the highest loading observed for the P2 polymer. The trend followed the order: P2 > P1 > P3 > P4. Interestingly, both the observed trend and the drug loading efficiency of TMP molecules differ from experimental results, likely due to the use of coarse-grained models and the limited system size in our simulations. Coarse-grained simulations simplify molecular details such as hydrogen bonding, specific electrostatic interactions, and the geometry of binding sites, which can impact the accuracy of drug–polymer interactions. Moreover, the hydrophobic–hydrophilic balance and solubility behavior of small drug molecules like TMP may not be fully represented. For example, the calculated log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value for the coarse-grained model of TMP used in this study was approximately 1.4 times higher than experimental values, which could contribute to deviations from the experimentally observed loading efficiencies.
P value for the coarse-grained model of TMP used in this study was approximately 1.4 times higher than experimental values, which could contribute to deviations from the experimentally observed loading efficiencies.
Nevertheless, the results reveal that the P4 system, despite having the fewest –COOH units, achieved a TMP loading level comparable to those of the other polymers. In fact, the total number of –COOH units in P1, P2, and P3 systems (summed across all molecules) was approximately 1.5, 1.5, and 1.7 times higher than in P4, respectively. To further understand this, we calculated the number of TMP molecules loaded per –COOH unit (Fig. 8F). We found that each –COOH unit in the P4 system loaded 1.35 times more TMP than the –COOH units in P1 and P2 and 1.5 times more than those in P3. The trend followed the order: P4 > P1 > P2 > P3. Overall, in agreement with the experimental results, the simulations also suggest that the –COOH units in the P4 polymer, consisting of longer aliphatic CH2 chains adjacent to the carboxyl group, show superior drug-loading capacity.
We then performed 2-μs long simulations by placing the polymers above the membrane surface in the aqueous phase. Each polymer was introduced either as a monomer or in a small micellar form. In the monomer setup, P1, P2, and P3 contained one monomer each, whereas P4 contained two monomers; this yielded 24, 32, 46, and 24 –COOH units for P1, P2, P3, and P4, respectively. In the micelle setup, the systems comprised four P1, three P2, three P3, and six P4 molecules, corresponding to 96, 96, 138, and 72 –COOH units, respectively. The initial configurations, showing the placement of polymer monomers and micelles above the membrane surface, are shown in Fig. S12 and S13, respectively. Note that the effective polymer concentration in the micelle simulations was approximately 9 mg mL−1, which is higher than the concentrations used in the experimental in vitro cytotoxicity studies in this work. However, since the experimental conditions were also above the polymers’ CMC, the use of pre-formed micelles in our simulations still realistically reflects the conditions relevant to cellular interactions and potential toxicity.
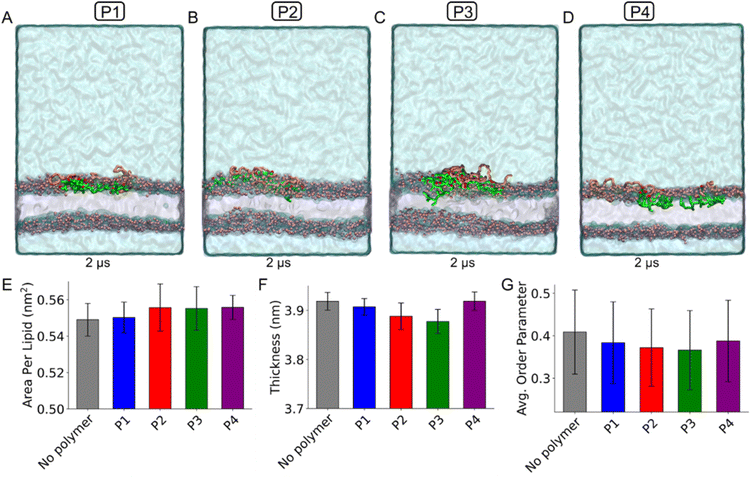
During the simulations, we observed that the polymers, regardless of their monomeric or micellar form, started interacting with the membrane surface and gradually adsorbed onto its surface. The final snapshots of the systems, showing the polymer monomers and micelles adsorbed at the membrane interface, are presented in Fig. 9(A–D) and Fig. S14(A–D), respectively. These snapshots reveal that some beads from the –COOH units partially inserted beneath the membrane head group region, while the negatively charged carboxylate beads remained near the head group interface. The mPEG beads predominantly stayed at the membrane–water interface, where they interacted with the surrounding water molecules, membrane head groups, and other polymer beads.
The insertion of –COOH units into the membrane altered key structural properties: area per lipid (APL), membrane thickness, and lipid tail order parameters, as shown in Fig. 9E, F and G for simulations using polymers in the monomer form. The polymer interactions with the membrane surface, including insertion of –COOH units below the lipid head groups, led to an increased area per lipid (APL; the average lateral area occupied by each lipid, where higher values indicate looser packing), reduced bilayer thickness, and decreased tail order parameters relative to the membrane-only system. These changes indicate a loss of membrane compactness in the presence of polymers.37,38 The extent of these effects reflects each polymer's membrane-disrupting potential, with P3 causing the greatest disruption, as evidenced by the lowest thickness and order values. Overall, the disruption ability follows the trend P3 > P2 > P1 > P4, as indicated in Fig. 9E–G. Simulations using polymer micelles showed a similar trend in membrane structural properties, as presented in Fig. S14E–G. Compared to systems with polymers in the monomer form, the micelle-containing systems exhibited higher APL, lower membrane thickness, and reduced tail order parameters. These effects are primarily due to the greater number of polymer molecules at the membrane surface and the increased insertion of –COOH units into the membrane.
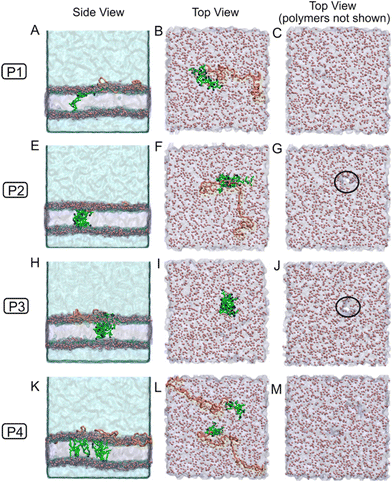
It is important to note that when COOH groups are attached to the membrane surface, their extended presence in the hydrophobic environment is expected to induce a pKa shift. This phenomenon has been previously observed for fatty acids with carboxylic acid head groups and aliphatic CH2 chains. In such cases, the pKa of the carboxyl group can shift from ∼4.5 in aqueous solution to between 5.5 and 7.5 in membrane environments, depending on the fatty acid chain length and its capacity to remain on the membrane surface.39 To reflect this shift, we used the final snapshots from the polymer–membrane interaction simulations (Fig. 10A–D and Fig. S14A–D) and modified the protonation state of the –COOH units, converting carboxylates to carboxylic acids. This was modeled by replacing the negatively charged bead representing the carboxylate group with a neutral bead. We then performed an additional 3 microseconds of simulation.
Through these extended simulations, the mPEG segments of the polymers remained near the membrane surface, maintaining interactions with the membrane head group beads or nearby water molecules. In contrast, the –COOH units were inserted more deeply into the membrane. Fig. 10 shows the final snapshots from different perspectives (side and top views) for systems with polymers in the monomeric form. These snapshots reveal that the higher number of –COOH units in the P2 and P3 systems aggregated within the membrane, forming clusters that spanned the full membrane thickness. This led to pronounced membrane disruption, including visible pore formation (Fig. 10E–J). In contrast, the P1 monomer system contained fewer –COOH units, which were insufficient to form large clusters capable of inducing similar disruption. However, when polymer micelles were used, the snapshots (Fig. S15A–J) showed pore formation in the P1 system as well. In these cases, aggregation of –COOH units enabled P1 to form clusters large enough to span the membrane and form pores, similar to P2 and P3.
In contrast, the –COOH units in the P4 polymers exhibited a distinct mode of interaction. Two consecutive COOH units appeared to span the membrane, with their carboxylic acid beads aligned with the membrane head group beads, one interacting with the upper leaflet and the other with the lower. The carboxylic acid beads and their adjacent aliphatic CH2 beads were vertically aligned with the lipid head groups and tails. A similar vertical orientation has been observed for fatty acids, which insert into membranes in a comparable fashion.38,40 Although this unique insertion pattern of P4 –COOH units (unlike those in P1, P2, and P3) did alter membrane structural properties, the ordered alignment prevented the formation of disordered clusters that span the membrane, thereby inhibiting pore formation in both monomeric and micellar systems, as shown in Fig. 10K–M and Fig. S15K–M, respectively.
Overall, the membrane-disrupting ability of the polymers followed the trend P3 > P2 > P1 > P4. P3 and P2 caused the most significant structural changes, including visible pore formation driven by aggregation of –COOH units in their neutral form. P1 induced moderate disruption, with limited effects in the charged state, and pore formation was observed in the micellar form with neutral –COOH groups. In contrast, P4 caused minimal disruption, as its –COOH units inserted in an ordered fashion that prevented clustering and pore formation. These findings underscore how the polymer structure and aggregation behavior influence membrane interactions and potential toxicity. Moreover, the simulations demonstrate the utility of coarse-grained molecular dynamics simulations in capturing polymer–membrane interactions and revealing molecular mechanisms underlying these effects.
Conclusion
The synthesized, functionalized amphiphilic block copolymers improve the solubility and encapsulation of the hydrophobic drugs, highlighting their potential as effective drug delivery vehicles. The results of this study demonstrate the number of ionic (–COOH) functional groups and the hydrophobic chain length of the functional molecules significantly influence micellar behavior. However, in contrast to our belief, increasing the –COOH groups on the polymer chain did not improve the drug loading capacity of the basic drug. Amphiphilic block copolymers with a higher number of –COOH groups (P3) exhibited sustained drug release, while those with fewer –COOH groups containing long carbon chains (P4) showed higher drug encapsulation efficiency and a greater in vitro safety profile. Furthermore, the polymers having a higher number of ionic groups (P2 and P3) significantly demonstrated pronounced membrane-disruptive activity towards red blood cells and could be used for the intracellular delivery of biological therapeutics. These experimental observations are supported by molecular dynamics simulations, which provide molecular-level insight into drug–polymer and polymer–cell membrane interactions. The simulations show that –COOH groups of the polymers interact strongly with the membrane and, when protonated, can aggregate within the bilayer. Such aggregation has the potential to generate membrane pores and disrupt membrane integrity, offering a mechanistic explanation for the inherent cytotoxicity of the polymers. However, further research is needed to evaluate the potential application of these next-generation nanocarriers as intracellular drug delivery vehicles.Author contributions
K. K.B.: conceptualization; V. K. and S. H. prepared the manuscript; K. K. B., J. M. R., and C. E. W. reviewed and edited the manuscript.; V. K. and E. L. synthesized and characterized the materials; S. H. and A. A. performed the molecular dynamics and cell toxicity study respectively; V. K., S. H., K. K. B., E. L., and A. A. performed the investigation and data analysis; K. K. B., J. M. R., S. H., and C. E. W. provided resources.; K. K. B., J. M. R., and C. E. W.: project administration, funding acquisition and supervision. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
The authors declare no conflict of interest.Abbreviations
| PA | mPEG5K-b-PJL16 |
| PB | mPEG5K-b-PJL48 |
| P1 | mPEG5K-b-PJL24-COOH24 |
| P2 | mPEG5K-b-PJL46-COOH32 |
| P3 | mPEG5K-b-PJL46-COOH46 |
| P4 | mPEG5K-b-PJL13-(CH2)8-COOH12 |
| 3MPA | 3-Mercaptopropionic acid |
| 11MUDA | 11-Mercaptoundecanoic acid |
| Trimethoprim | TMP |
| Ce6 | Chlorine E6 |
Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: tables, figures, NMR spectra and further experimental details. See DOI: https://doi.org/10.1039/d5py01016k.Any raw data files needed in another format are available from the corresponding author on request.
Acknowledgements
V. K. acknowledges the National Overseas Scholarship, Ministry of Social Justice and Empowerment, Government of India for providing funding for his personal PhD scholarship. The computations/data handling were enabled by resources provided by the National Academic Infrastructure for Supercomputing in Sweden (NAISS), partially funded by the Swedish Research Council through grant agreement no. 2022-06725. Financial support from VINNOVA (2019-00048 and 2024-03851) for the Swedish Drug Delivery Center (SweDeliver) is gratefully acknowledged.This research was partly funded by Business Finland Research-to-Business project Jasmine PRO (1609/31/2021). Additionally, this study is part of the activities of Åbo Akademi Foundation (SÅA) funded by the Centre of Excellence in Research “Materials-driven solutions for combating antimicrobial resistance (MADNESS)”. The activities are also aligned with the strategic research profiling area “Solutions for Health” at Åbo Akademi University [Academy of Finland, # 336355].
References
- R. Roy, S. K. Singh, N. Ahmad and S. Misra, Challenges and advancements in high-throughput screening strategies for cancer therapeutics, Global Transl. Med., 2024, 3(1), 2448 Search PubMed.
- Y. Zhuo, Y. G. Zhao and Y. Zhang, Enhancing Drug Solubility, Bioavailability, and Targeted Therapeutic Applications through Magnetic Nanoparticles, Molecules, 2024, 29(20), 4854 Search PubMed.
- Y. Lu, E. Zhang, J. Yang and Z. Cao, Strategies to improve micelle stability for drug delivery, Nano Res., 2018, 11(10), 4985–4998 CrossRef.
- P. Weingarten, M. T. Y. Lin, M. Kränzlein, A. Fietz, I. Kachel and J. Hurst, et al., Phosphorous-containing, amphiphilic ABB′ copolymers as siRNA nanocarriers with enhanced stability, reduced in vitro cytotoxicity, and efficient knockdown ability for the treatment of ocular diseases, RSC Appl. Polym., 2025, 3(2), 381–390 RSC.
- M. Chehelgerdi, M. Chehelgerdi, O. Q. B. Allela, R. D. C. Pecho, N. Jayasankar and D. P. Rao, et al., Progressing nanotechnology to improve targeted cancer treatment: overcoming hurdles in its clinical implementation, Mol. Cancer, 2023, 22, 169 CrossRef PubMed.
- R. Salah Othman, S. Zarei, H. Rezaei Haghighat, A. Afshar Taromi and H. A. Khonakdar, Recent Advances in Smart Polymeric Micelles for Targeted Drug Delivery, Polym. Adv. Technol., 2025, 36(4), e70180 CrossRef CAS.
- A. H. Maboudi, M. H. Lotfipour, M. Rasouli, M. H. Azhdari, R. MacLoughlin and S. Bekeschus, et al., Micelle-based nanoparticles with stimuli-responsive properties for drug delivery, Nanotechnol. Rev., 2024, 13(1), 20230218 Search PubMed.
- N. Akhlaghi and G. Najafpour-Darzi, Amino-Functionalized Pluronic F127 micelles as a dual drug delivery nanostructure for controlled therapeutics release, J. Mol. Liq., 2024, 400, 124489 CrossRef CAS.
- H. Wang, H. Polara, A. Bhadran, T. Shah, G. K. Babanyinah and Z. Ma, et al., Effect of aromatic substituents on thermoresponsive functional polycaprolactone micellar carriers for doxorubicin delivery, Front. Pharmacol., 2024, 15, 1356639 Search PubMed.
- H. He, N. Huang, Z. Qiu, L. He, J. Guo and M. Xu, et al., Effects of polymer terminal group inside micelle core on paclitaxel loading promoting and burst release suppressing, J. Gastrointest. Oncol., 2023, 14(4), 1659–1668 Search PubMed.
- Y. Yuan, W. Tan, Y. Mi, L. Wang, Z. Qi and Z. Guo, Effect of Hydrophobic Chain Length in Amphiphilic Chitosan Conjugates on Intracellular Drug Delivery and Smart Drug Release of Redox-Responsive Micelle, Mar. Drugs, 2024, 22(1), 18 CrossRef CAS.
- Y. Wang, V. Ukwattage, Y. Xiong and G. K. Such, Advancing endosomal escape of polymeric nanoparticles: towards improved intracellular delivery, Mater. Horiz., 2025, 12(11), 3622–3632 RSC.
- J. Wang, Y. Ding, K. Chong, M. Cui, Z. Cao and C. Tang, et al., Recent Advances in Lipid Nanoparticles and Their Safety Concerns for mRNA Delivery, Vaccines, 2024, 12(10), 1148 Search PubMed.
- J. Casper, S. H. Schenk, E. Parhizkar, P. Detampel, A. Dehshahri and J. Huwyler, Polyethylenimine (PEI) in gene therapy: Current status and clinical applications, J. Controlled Release, 2023, 362, 667–691 CrossRef CAS.
- K. Mrksich, M. S. Padilla and M. J. Mitchell, Breaking the final barrier: Evolution of cationic and ionizable lipid structure in lipid nanoparticles to escape the endosome, Adv. Drug Delivery Rev., 2024, 214, 115446 CrossRef CAS PubMed.
- A. Ali, R. Bhadane, A. A. Asl, C. E. Wilén, O. Salo-Ahen and J. M. Rosenholm, et al., Functional block copolymer micelles based on poly (jasmine lactone) for improving the loading efficiency of weakly basic drugs, RSC Adv., 2022, 12(41), 26763–26775 RSC.
- J. Verma, V. Kumar, C. E. Wilen, J. M. Rosenholm and K. K. Bansal, Reactive Oxygen Species-Regulated Conjugates Based on Poly(jasmine) Lactone for Simultaneous Delivery of Doxorubicin and Docetaxel, Pharmaceutics, 2024, 16(9), 1164 CrossRef CAS.
- R. Di, K. K. Bansal, J. M. Rosenholm, H. Grohganz and T. Rades, Utilizing the allyl-terminated copolymer methoxy(poly(ethylene glycol))-block-poly(jasmine lactone) in the development of amorphous solid dispersions: A comparative study of functionalized and non-functionalized polymer, Int. J. Pharm., 2024, 657, 124175 CrossRef CAS PubMed.
- O. M. H. Salo-Ahen, I. Alanko, R. Bhadane, A. M. J. J. Bonvin, R. V. Honorato and S. Hossain, et al., Molecular Dynamics Simulations in Drug Discovery and Pharmaceutical Development, Processes, 2021, 9(1), 71 CrossRef CAS.
- M. De Vivo, M. Masetti, G. Bottegoni and A. Cavalli, Role of Molecular Dynamics and Related Methods in Drug Discovery, J. Med. Chem., 2016, 59(9), 4035–4061 CrossRef CAS PubMed.
- K. K. Bansal, E. Özliseli, A. Rosling and J. M. Rosenholm, Synthesis and Evaluation of Novel Functional Polymers Derived from Renewable Jasmine Lactone for Stimuli-Responsive Drug Delivery, Adv. Funct. Mater., 2021, 31(33), 2101998 CrossRef CAS.
- C. E. Hoyle and C. N. Bowman, Thiol–Ene Click Chemistry, Angew. Chem., Int. Ed., 2010, 49(9), 1540–1573 Search PubMed.
- A. Mustafai, M. Zubair, A. Hussain and A. Ullah, Recent Progress in Proteins-Based Micelles as Drug Delivery Carriers, Polymers, 2023, 15(4), 836 CrossRef CAS.
- I. Negut and B. Bita, Polymeric Micellar Systems—A Special Emphasis on “Smart” Drug Delivery, Pharmaceutics, 2023, 15(3), 976 CrossRef CAS PubMed.
- K. K. Bansal, D. Kakde, L. Purdie, D. J. Irvine, S. M. Howdle and G. Mantovani, et al., New biomaterials from renewable resources – amphiphilic block copolymers from δ-decalactone, Polym. Chem., 2015, 6(40), 7196–7210 RSC.
- S. K. Filippov, R. Khusnutdinov, A. Murmiliuk, W. Inam, L. Y. Zakharova and H. Zhang, et al., Dynamic light scattering and transmission electron microscopy in drug delivery: a roadmap for correct characterization of nanoparticles and interpretation of results, Mater. Horiz., 2023, 10(12), 5354–5370 Search PubMed.
- S. Sethi, Medha, S. Thakur and B. S. Kaith, Preliminary in vitro hemocompatibility assessment of biopolymeric hydrogels for versatile biomedical applications, Polym. Bull., 2024, 81(5), 4499–4522 CrossRef CAS.
- K. K. Bansal, J. Gupta, A. Rosling and J. M. Rosenholm, Renewable poly(δ-decalactone) based block copolymer micelles as drug delivery vehicle: in vitro and in vivo evaluation, Saudi Pharm. J., 2018, 26(3), 358–368 CrossRef.
- S. Deng, H. Shao, H. Shang, L. Pang, X. Chen and J. Cao, et al., Development of a Cationic Polymeric Micellar Structure with Endosomal Escape Capability Enables Enhanced Intramuscular Transfection of mRNA-LNPs, Vaccines, 2025, 13(1), 25 CrossRef CAS.
- S. Wang, S. Wannasarit, P. Figueiredo, G. Molinaro, Y. Ding and A. Correia, Intracellular Delivery of Budesonide and Polydopamine Co-Loaded in Endosomolytic Poly(butyl methacrylate-co-methacrylic acid) Grafted Acetalated Dextran for Macrophage Phenotype Switch from M1 to M2, Adv. Therap., 2021, 4, 2000058 Search PubMed.
- S. Wang and R. Chen, pH-Responsive, Lysine-Based, Hyperbranched Polymers Mimicking Endosomolytic Cell-Penetrating Peptides for Efficient Intracellular Delivery, Chem. Mater., 2017, 29(14), 5806–5815 Search PubMed.
- R. Chen, S. Khormaee, M. E. Eccleston and N. K.H. Slater, The role of hydrophobic amino acid grafts in the enhancement of membrane-disruptive activity of pH-responsive pseudo-peptides, Biomaterials, 2009, 30(10), 1954–1961 CrossRef CAS.
- K. Yu, J. Zhao, C. Yu, F. Sun, Y. Liu and Y. Zhang, et al., Role of Four Different Kinds of Polyethylenimines (PEIs) in Preparation of Polymeric Lipid Nanoparticles and Their Anticancer Activity Study, J. Cancer, 2016, 7(7), 872–882 CrossRef CAS PubMed.
- J. R. Magaña Rodriguez, M. Guerra-Rebollo, S. Borrós and C. Fornaguera, Nucleic acid-loaded poly(beta-aminoester) nanoparticles for cancer nano-immuno therapeutics: the good, the bad, and the future, Drug Delivery Transl. Res., 2024, 14(12), 3477–3493 CrossRef PubMed.
- D. Altun, X. He, C. Bergström, M. Hubert and S. Hossain, Molecular dynamics simulations of a hexagonal liquid crystal phase to study drug partitioning and release mechanisms, Colloids Surf., B, 2026, 258, 115240 CrossRef CAS.
- R. Robeyns, A. Sisto, E. Iturrospe, K. M. da Silva, M. van de Lavoir and V. Timmerman, et al., The Metabolic and Lipidomic Fingerprint of Torin1 Exposure in Mouse Embryonic Fibroblasts Using Untargeted Metabolomics, Metabolites, 2024, 14(5), 248 CrossRef CAS.
- D. Altun, P. Larsson, C. A. S. Bergström and S. Hossain, Molecular dynamics simulations of lipid composition and its impact on structural and dynamic properties of skin membrane, Chem. Phys. Lipids, 2024, 265, 105448 Search PubMed.
- R. Kneiszl, S. Hossain and P. Larsson, In Silico-Based Experiments on Mechanistic Interactions between Several Intestinal Permeation Enhancers with a Lipid Bilayer Model, Mol. Pharmaceutics, 2022, 19(1), 124–137 CrossRef CAS PubMed.
- A. A. Pashkovskaya, M. Vazdar, L. Zimmermann, O. Jovanovic, P. Pohl and E. E. Pohl, Mechanism of Long-Chain Free Fatty Acid Protonation at the Membrane-Water Interface, Biophys. J., 2018, 114(9), 2142–2151 Search PubMed.
- S. Hossain, P. Joyce, A. Parrow, S. Jõemetsa, F. Höök and P. Larsson, et al., Influence of Bile Composition on Membrane Incorporation of Transient Permeability Enhancers, Mol. Pharmaceutics, 2020, 17(11), 4226–4240 CrossRef CAS.
Footnote |
| † Present address: Department of chemistry, Toulouse INP, ENSIACET 4 allee, Emile Monso, 31030 Toulouse France. |
| This journal is © The Royal Society of Chemistry 2026 |