Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
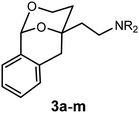
Forcing the phenyl moiety into the axial position by embedding the 2-phenyl-1,3-dioxane system in a tricyclic benzomorphan scaffold: design, synthesis and biological evaluation
Nina
Schulte-Gauczinski
,
Judith
Schmidt
,
Jens
Köhler
,
Dirk
Schepmann
and
Bernhard
Wünsch
 *
*
Universität Münster, Institut für Pharmazeutische und Medizinische Chemie, Corrensstraße 48, D-48149 Münster, Germany. E-mail: wuensch@uni-muenster.de; Tel: +49-251-8333311
First published on 17th February 2026
Abstract
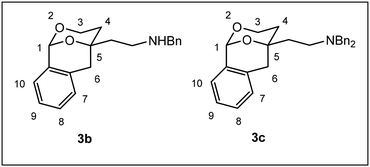
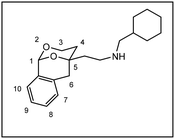
The relative configuration and the substitution pattern control the interaction of 2-(2-phenyl1,3-dioxan-4-yl)ethan-1-amines with σ1 receptors or the PCP binding site of NMDA receptors. In order to investigate the influence of the orientation of the phenyl moiety in 2-position on the receptor interaction, the 2-phenyl-1,3-dioxane system was embedded in a tricyclic benzomorphan scaffold (3) fixing the phenyl moiety in an axial orientation relative to the 1,3-dioxane ring. The key step of the synthesis of tricyclic amines 3 was the addition of lithiated 2-methylbenzamide 7 at pentanone 6 to afford the tertiary alcohol 8. Lactone formation (9), DIBAH reduction (10) and intramolecular transacetalization led to the tricyclic alcohol 11, which was converted into a series of twelve primary, secondary and tertiary amines 3a–m. Although the primary amine 3a is structurally related to the potent PCP antagonist 2a, it did not interact with the PCP binding site of the NMDA receptor. The missing ethyl moiety and/or an unfavorable orientation of the phenyl moiety might be responsible for the lost PCP affinity of 3a. As observed for the flexible 1,3-dioxanes 1b and 2b, introduction of a benzyl moiety at the amino group resulted in high σ1 receptor affinity of 3b. In accordance with σ1 pharmacophore models, two small or two large substituents at the amino moiety were less tolerated by the σ1 receptor, whereas an additional small methyl moiety increased the σ1 affinity of 3h and 3j. With respect to σ1 receptor affinity and selectivity over the σ2 subtype, the methylated cyclohexylmethylamine 3j (Ki(σ1) = 6.4 nM, 9-fold selectivity) represents the most promising ligand. The highest ligand-lipophilicity efficiency (LLE) was obtained for the secondary cyclohexylmethylamine 3d (LLE = 6.7). However, the highest metabolic stability (phase I metabolism) was determined for the benzylamine 3b (89% intact after incubation for 90 min).
1. Introduction
This project deals with target hopping, i.e., a switch from open channel blockers of N-methyl-D-aspartate (NMDA) receptors to σ1 receptor ligands.The heterotetrameric NMDA receptor belongs to the class of ligand-gated ion channels. The pore of the NMDA receptor contains binding sites for Mg2+ ions and phencyclidine (1-(1-phenylcyclohexyl)piperidine, PCP, Fig. 1).1–3 Addressing the PCP binding site with ligands leads to inhibition of the ion flux through the open channel.1–3 NMDA receptor inhibitors interacting with the PCP binding site are termed open-channel blockers, as the ion channel has to be opened by the agonists glutamate and glycine before the ligands can enter the channel pore and reach their binding site. Very potent open-channel blockers, such as phencyclidine, MK-801, and dexoxadrol (Fig. 1)1–3 are associated with desired analgesic and anesthetic activity, but also with strong undesired psychotomimetic and hallucinogenic effects. However, some low affinity open-channel blockers are clinically used as dissociative anesthetic (ketamine) and antitussive (dextromethorphan) as well as for the treatment of Parkinson's (amantadine) and Alzheimer's disease (memantine).1–3 The structure of the NMDA receptor has been solved by X-ray crystal structure analysis.4,5
 | ||
| Fig. 1 Prototypical ligands interacting with the PCP binding site of the NMDA receptor and/or with σ receptors. | ||
The σ1 receptor is a chaperone predominantly located in the membrane of the endoplasmic reticulum controlling Ca2+ flux. It is involved in neurodegenerative and neuropsychiatric diseases such as schizophrenia, depression and addiction as well as Alzheimer's diseases. Moreover, inhibition of the σ1 receptor can be exploited for the treatment of neuropathic pain and cancer.6–8 In the forced swimming test, σ1 receptor knock-out mice showed a depression-like behavior.9,10 In 2016, the σ1 receptor was crystallized for the first time exhibiting an unexpected structure with only one transmembrane helix.11 Two years later, structures of the σ1 receptor in complex with prototypical ligands haloperidol and (+)-pentazocine were reported.12
σ receptors and the PCP binding site of NMDA receptors are historically related. After refuting the hypothesis that the σ receptor is an opioid receptor subtype,13 it was postulated that the PCP binding site and the σ receptor are identical.14 This hypothesis arose from the observation that prototypical open channel blockers of the NMDA receptor, e.g., phencyclidine, interacted with high affinity with σ receptors as well. Moreover, prototypical σ ligands such as racemic SKF-10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 047 (rac-N-allylnormetazocine, Fig. 1) could also inhibit the NMDA receptor by interacting with the PCP binding site.14 However, identification of ligands inhibiting selectively the NMDA receptor by interaction with the PCP binding site without addressing the σ receptor (e.g., dexoxadrol) and ligands interacting selectively with σ1 receptors but not with the PCP binding site (e.g., haloperidol, Fig. 1) led to a clear differentiation of σ receptors and the PCP binding site. Moreover, the distribution patterns of both proteins in the central nervous system are quite different.15
047 (rac-N-allylnormetazocine, Fig. 1) could also inhibit the NMDA receptor by interacting with the PCP binding site.14 However, identification of ligands inhibiting selectively the NMDA receptor by interaction with the PCP binding site without addressing the σ receptor (e.g., dexoxadrol) and ligands interacting selectively with σ1 receptors but not with the PCP binding site (e.g., haloperidol, Fig. 1) led to a clear differentiation of σ receptors and the PCP binding site. Moreover, the distribution patterns of both proteins in the central nervous system are quite different.15
The confusion regarding binding affinity of benzomorphans originated from their absolute configuration: dextrorotatory (S,S,S)-configured N-allyl-normetazocine ((+)-SKF-10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 047) shows high affinity towards the σ1 receptor subtype, whereas the levorotatory (R,R,R)-configured enantiomer (−)-SKF-10
047) shows high affinity towards the σ1 receptor subtype, whereas the levorotatory (R,R,R)-configured enantiomer (−)-SKF-10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 047 inhibits the NMDA receptor associated cation channel. Small N-substituents (e.g., H or CH3) of (R,R,R)-(−)-benzomorphans increase the PCP binding affinity and decrease the affinity towards opioid receptors.16
047 inhibits the NMDA receptor associated cation channel. Small N-substituents (e.g., H or CH3) of (R,R,R)-(−)-benzomorphans increase the PCP binding affinity and decrease the affinity towards opioid receptors.16
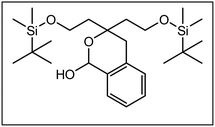
Derived from the PCP ligands dexoxadrol and etoxadrol 2,4-disubstituted 1,3-dioxanes 1 and 2 have been developed.17,18 Depending on the substitution pattern and the orientation of the phenyl moiety in 2-position, 1 and 2 showed high affinity and selectivity either for the σ1 receptor or the PCP binding site of the NMDA receptor. Whereas the benzaldehyde-derived primary amine 1a did neither interact with σ1 receptors nor the PCP binding site, the analogous benzylamine 1b revealed high σ1 receptor affinity and selectivity (>170-fold) over the PCP binding site. The propiophenone-derived 1,3-dioxane 2a with a primary amino moiety showed strong interaction with the PCP binding site and high selectivity over the σ1 receptor (>750-fold). Introduction of a benzyl moiety at the primary amine of 2a led to the secondary amine 2b with high σ1 receptor affinity and high selectivity over the PCP binding site (>500-fold)17 (Fig. 2).
 | ||
| Fig. 2 Design of conformationally restricted 1,3-dioxanes 3 with the phenyl ring fixed in axial orientation. | ||
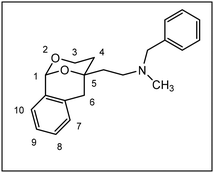
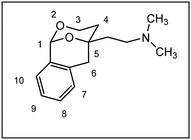
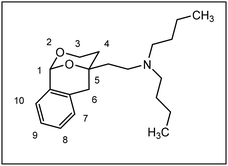
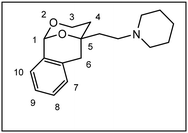
Herein, we report the synthesis and biological evaluation of conformationally restricted analogs of 1 and 2, in which the 2-phenyl-1,3-dioxane system is embedded in a tricyclic benzomorphan scaffold. In the tricyclic acetals 3 the phenyl moiety in “2-position” of the 1,3-dioxane ring is forced to adopt the axial orientation. This axial orientation of the phenyl moiety is not possible for benzaldehyde-derived acetals 1, since the phenyl moiety always adopts the equatorial orientation due to thermodynamic reasons. On the other hand, the “axial orientation” (related to the 1,3-dioxane ring) of the phenyl ring in tricyclic compounds 3 corresponds to the axial orientation in the propiophenone-derived 1,3-dioxanes 2. However, the ethyl moiety at the acetalic center of 2 is missing in the tricyclic acetals 3. The conformationally restricted acetals 3 should provide relationships between structural modifications and affinity towards σ receptors and NMDA receptors. In particular, insights should be gained about the relevance of the orientation of the phenyl moiety and the presence of an additional ethyl substituent in 2-position of the 1,3-dioxane ring. Variation of the substituents at the amnio moiety should further broaden this SAR study.
2. Results and discussion
2.1. Synthesis
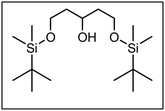
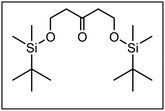
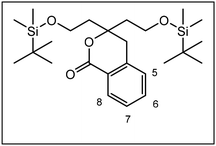
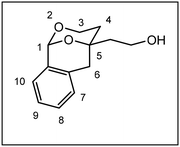
The synthesis of the conformationally restricted 1,3-dioxanes 3 started with pentane-1,3,5-triol (4).19 The primary alcoholic groups of 4 were selectively protected by the tert-butyldimethylsilyl (TBS) protective group to afford the bis-silyl ether 5. Subsequently, the remaining secondary alcohol 5 was oxidized with Dess Martin Periodinane (DMP)20 to give the ketone 6 in 97% yield as key intermediate of this synthesis (Scheme 1).The next step, i.e., the nucleophilic addition of 2-methylbenzamide 7 to the ketone 6 represents the key step of the synthesis. For this purpose, benzamide 7 was deprotonated with sec-BuLi at the CH3 moiety and the resulting methyllithium species reacted with the ketone 6 to provide the tertiary alcohol 8 in 92% yield. Heating of the hydroxy amide 8 without solvent at 186 °C for 16 h led to the δ-lactone 9, which was reduced with diisobutylaluminum hydride (DIBAH)21 to provide the cyclic hemi acetal 10 in 94% yield. Treatment of the hemi acetal 10 with diluted HCl led to cleavage of both silyl ethers, but only one primary alcohol could react with the hemi acetal to form the tricyclic acetal 11. During this reaction step, the symmetry of the molecules was lost leading to chiral compounds (Scheme 1).
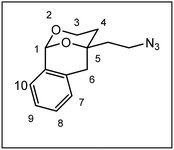
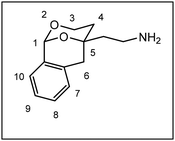
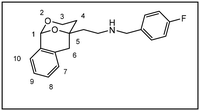
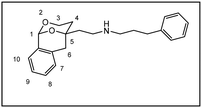
After establishment of the tricyclic system 11, the remaining primary alcohol was converted into various amines 3. At first, a Mitsunobu reaction with Zn(N3)2·2 pyridine in the presence of PPh3 and diisopropyl azodicarboxylate (DIAD) transformed the primary alcohol 11 into azide 12, which was reduced with H2 and Pd/C to afford the primary amine 3a. Finally, reductive alkylation of the primary amine 3a with aldehydes and NaBH(OAc)3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22 provided secondary and tertiary amines 3b–m. Stoichiometric amounts of aldehyde led to secondary amines (e.g.;3d, 3f, 3g), intermediate amounts of benzaldehyde provided a mixture of secondary and tertiary amines 3b and 3c, whereas an excess of aldehyde completely alkylated the primary amine 3a affording tertiary amines 3e, 3k nd 3l. The piperidine ring of 3m was established by reductive alkylation of the primary amine 3a with glutaraldehyde and NaBH(OAc)3.22 Reductive methylation with formaldehyde and NaBH(OAc)3 converted the secondary amines 3b and 3d into tertiary methylamines 3h and 3j, respectively.
22 provided secondary and tertiary amines 3b–m. Stoichiometric amounts of aldehyde led to secondary amines (e.g.;3d, 3f, 3g), intermediate amounts of benzaldehyde provided a mixture of secondary and tertiary amines 3b and 3c, whereas an excess of aldehyde completely alkylated the primary amine 3a affording tertiary amines 3e, 3k nd 3l. The piperidine ring of 3m was established by reductive alkylation of the primary amine 3a with glutaraldehyde and NaBH(OAc)3.22 Reductive methylation with formaldehyde and NaBH(OAc)3 converted the secondary amines 3b and 3d into tertiary methylamines 3h and 3j, respectively.
2.2. Receptor affinity
The affinity of the amines 3a–m towards the PCP binding site of the NMDA receptor and towards σ1 and σ2 receptors was determined in radioligand receptor binding studies. In the PCP assay, pig brain cortex was used as receptor material and [3H](+)-MK-801 as radioligand.17,23 [3H](+)-pentazocine and [3H]di-o-tolylguanidine served as radioligands and guinea pig brain and rat liver as receptor material in the σ1 and σ2 receptor assay, respectively.24,25 The recorded affinities of amines 3 are summarized in Table 1 together with the Ki values of some reference compounds.| Entry | Compd | NR2 | K i ± SEMa,b (nM) | ||
|---|---|---|---|---|---|
| PCP | σ 1 | σ 2 | |||
| a K i values represent the mean of at least three independent experiments. b For ligands with low affinity, the Ki value was determined only once. | |||||
| 1 | 3a | NH2 | >1000 | >1000 | >1000 |
| 2 | 3b | NHBn | >1000 | 69 ± 9 | 60 ± 23 |
| 3 | 3c | NBn2 | >1000 | >1000 | >1000 |
| 4 | 3d | NHCH2C6H11 | >1000 | 15 ± 4 | 26 ± 16 |
| 5 | 3e | N(CH2C6H11)2 | >1000 | 280 | 375 |
| 6 | 3f | NHCH2C6H4(4-F) | >1000 | 47 ± 29 | 30 ± 11 |
| 7 | 3g | NHCH2CH2CH2Ph | >1000 | 242 | 62 ± 8 |
| 8 | 3h | N(CH3)Bn | >1000 | 15 ± 3 | 137 |
| 9 | 3j | N(CH3)CH2C6H11 | >1000 | 6.4 ± 2.0 | 55 ± 11 |
| 10 | 3k | N(CH3)2 | >1000 | >1000 | >1000 |
| 11 | 3l | N(C4H9)2 | >1000 | 3410 | 298 |
| 12 | 3m | N(CH2)5 | >1000 | 766 | 381 |
| 13 | Dexoxadrol | 25 ± 4 | — | — | |
| 14 | Phencyclidine | 59 ± 12 | — | — | |
| 15 | (+)-Pentazocine | — | 5.7 ± 2.2 | — | |
| 16 | Haloperidol | — | 6.3 ± 1.6 | 78 ± 2.3 | |
| 17 | Di-o-tolylguanidine | — | 89 ± 29 | 58 ± 18 | |
The primary amine 3a did not show high affinity for the PCP binding site of the NMDA receptor. The structurally related lead compound 2a exhibited a strong interaction with the PCP binding site of the NMDA receptor (Ki = 13 nM, see Fig. 1). Both compounds 2a and 3a are primary amines with an axially oriented phenyl moiety in 2-position of the 1,3-dioxane ring. However, 2a has an additional ethyl moiety in 2-position of the 1,3-dioxane ring, which is not present in the tricyclic amine 3a. The missing ethyl moiety or the slightly different orientation of the axially oriented phenyl moiety in 2-postion of the 1,3-dioxnae ring might be responsible for the complete loss of PCP affinity of primary amine 3a.
Introduction of various substituents at the amino moiety did not lead to PCP affinity of 3b–m. This observation is in good accordance with literature describing the highest PCP affinity for primary amines for these types of ligands.17
In agreement with literature (see Fig. 1), the primary amine 3a did not bind at σ1 and σ2 receptors. However, as shown for the lead compounds 1b and 2b, the introduction of a benzyl moiety increased the σ1 affinity of 3b (Ki = 69 nM) considerably. A cyclohexylmethyl moiety instead of the benzyl moiety increased the σ1 affinity to 15 nM (3d). A further increase of the σ1 affinity could be obtained by methylation of the secondary amines 3b and 3d: the analogous methylamines 3h (Ki = 15 nM) and 3j (Ki = 6.4 nM) revealed the highest σ1 affinity of this class of compounds. Two large (3c, 3e) or two small substituents (3k. 3m) at the amino moiety did not lead to high σ1 affinity. A larger alkyl chain between the N-atom and the terminal phenyl moiety reduced the σ1 affinity of the phenylpropyl derivative 3g (Ki = 242 nM).
The secondary amines 3b, 3d, and 3f show very similar affinities towards both σ1 and σ2 receptors. However, selectivity for the σ1 receptor was gained by introduction of an additional CH3 moiety at the amino group. The methylated benzylamine 3h and the methylated cyclohexylmethylamine 3j displayed 9-fold selectivity for the σ1 receptor over the σ2 subtype.
2.3. Physicochemical and pharmacokinetic properties of selected ligands
In order to get an idea about the lipophilicity of the tricyclic compounds, the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 value of selected ligands was determined using the micro-shake flask method.26,27 In brief, the compounds were distributed between n-octanol and MOPS buffer pH 7.4 layers and the amount of the compound in the buffer layer was determined by LC-MS.
D7.4 value of selected ligands was determined using the micro-shake flask method.26,27 In brief, the compounds were distributed between n-octanol and MOPS buffer pH 7.4 layers and the amount of the compound in the buffer layer was determined by LC-MS.
The recorded log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 values are summarized in Table 2. The benzylamine 3b and the cyclohexylmethylamine 3d show the same log
D7.4 values are summarized in Table 2. The benzylamine 3b and the cyclohexylmethylamine 3d show the same log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 value of 1.1. The additional methyl moiety at the amino group of 3j increased the log
D7.4 value of 1.1. The additional methyl moiety at the amino group of 3j increased the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 value by approx. one log unit. The amount of the primary amine 3a in the aqueous layer was higher than its amount in the n-octanol layer leading to a negative log
D7.4 value by approx. one log unit. The amount of the primary amine 3a in the aqueous layer was higher than its amount in the n-octanol layer leading to a negative log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 value of −1.9 (Table 2).
D7.4 value of −1.9 (Table 2).
| Compd | NR2 | −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ki(σ1) Ki(σ1) |
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 ± SD (n = 3) D7.4 ± SD (n = 3) |
LLEa | Plasma protein bindingb | Metabolic stability ± SD (n = 3)c |
|---|---|---|---|---|---|---|
a LLE (ligand-lipophilicity efficiency) = −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ki − log Ki − log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4.
b Interaction with human serum albumin was recorded by HPAC analysis.
c Amount (in %) of parent compound after incubation with mouse liver microsomes and NADPH for 90 min.
d Mean of two values.
e Imipramine was used as reference compound to prove the activity of the microsomes and NADPH in the metabolic experiments. D7.4.
b Interaction with human serum albumin was recorded by HPAC analysis.
c Amount (in %) of parent compound after incubation with mouse liver microsomes and NADPH for 90 min.
d Mean of two values.
e Imipramine was used as reference compound to prove the activity of the microsomes and NADPH in the metabolic experiments.
|
||||||
| 3a | NH2 | <6 | −1.9d | 42 ± 0.2 | 87 ± 4 | |
| 3b | NHBn | 7.2 | 1.1 ± 0.04 | 6.1 | 75 ± 1 | 89 ± 6 |
| 3d | NHCH2C6H11 | 7.8 | 1.1 ± 0.05 | 6.7 | 77 ± 1 | 32 ± 4 |
| 3j | N(CH3)CH2C6H11 | 8.2 | 2.0 ± 0.1 | 6.2 | 78 ± 1 | 45 ± 4 |
| Imipraminee | 14 ± 1 | |||||
The recorded log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 values were used to calculate the ligand-lipophilicity efficiency (LLE), which modulates the biological activity of a ligand by its lipophilicity (LLE = −log
D7.4 values were used to calculate the ligand-lipophilicity efficiency (LLE), which modulates the biological activity of a ligand by its lipophilicity (LLE = −log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ki − log
Ki − log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4).28,29 Due to its increased σ1 affinity, the LLE value of the cyclohexylmethyl derivative 3d (LLE = 6.7) is higher than the LLE value of the benzyl derivative 3b (LLE = 6.1). However, the increased σ1 affinity of the methylated derivative 3j is compensated by its increased lipophilicity resulting in a reduced LLE value of 6.2. The LLE value for the primary amine 3a was not calculated, since 3a did not show any interactions with the σ1 receptor even at a concentration of 1 µM (Table 2).
D7.4).28,29 Due to its increased σ1 affinity, the LLE value of the cyclohexylmethyl derivative 3d (LLE = 6.7) is higher than the LLE value of the benzyl derivative 3b (LLE = 6.1). However, the increased σ1 affinity of the methylated derivative 3j is compensated by its increased lipophilicity resulting in a reduced LLE value of 6.2. The LLE value for the primary amine 3a was not calculated, since 3a did not show any interactions with the σ1 receptor even at a concentration of 1 µM (Table 2).
The plasma protein binding of the amines 3a, 3b, 3d and 3j was determined by high performance affinity chromatography (HPAC) using a stationary phase coated with human serum albumin. The retention time correlates with the affinity towards human serum albumin, which represents the main component of human blood.30,31 The plasma protein binding of the secondary amines 3b and 3d and the tertiary amine 3j is very similar (75–78%). However, the very polar primary amine 3a exhibited a considerably lower binding to human serum albumin of 42% (Table 2).
In order to investigate the phase I metabolic stability, selected amines were incubated with mouse liver microsomes and NADPH at 37 °C for 90 min. The amount of the residual parent compound was determined by LC-MS.26,32 The benzylamine 3b revealed high metabolic stability, since 89% of the parent compound were intact after 90 min. The cyclohexylmethylamines 3d and 3j were faster metabolized and only 32% and 45% remained unchanged after 90 min, respectively.
3. Conclusion
The 1,3-dioxane 2a bearing an axially oriented phenyl moiety at 2-position and a primary amino moiety at the ethyl substituent in 4-position showed high affinity towards the PCP binding sit of the NMDA receptor (Ki = 13 nM), but no affinity at σ1 and σ2 receptors (Fig. 1). Although structurally related to 2a, the conformationally restricted 1,3-dioxane 3a with a primary amino moiety at the ethyl substituent did not interact with the PCP binding site. We hypothesize that the missing 2-ethyl moiety of 2a and/or an unfavorable orientation of the conformationally restricted phenyl moiety are the reason for the reduced PCP affinity of the primary amine 3a.Independent of the orientation of the phenyl moiety in 2-position, both 1,3-dioxanes 1b and 2b with a benzylamino moiety displayed high σ1 affinity (Fig. 1). The same effect was observed for the benzylamine 3b showing also high σ1 affinity (Ki = 69 nM).
The results obtained for this novel class of σ1 ligands correlate well with pharmacophore models for σ1 receptor ligands. According to these models, potent σ1 ligands should contain a central amino moiety substituted with two lipophilic substituents.33–35 The introduction of an additional large substituent at the amino group as in 3c and 3e led to considerably reduced σ1 affinity. On the other hand, two small substituents at the amino moiety (e.g., compounds 3k, 3l, 3m) are not sufficient to bind with high affinity at σ1 receptors. However, a small methyl moiety in addition to the large benzyl or cyclohexylmethyl moiety at the amino group resulted in increased σ1 affinity of 3h and 3j.
The highest σ1 affinity was observed for the cyclohexylmethyl substituted amines 3d (Ki = 15 nM) and 3j (Ki = 6.4 nM). Due to their low lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 = 1.1 and 2.0, respectively), both compounds exhibit high LLE values of 6.7 and 6.2, respectively. However, in the presence of mouse liver microsomes and NMDPH (phase I metabolism), both amines 3d and 3j were rapidly metabolized. In contrast, the less potent benzylamine 3b (Ki(σ1) = 69 nM, LLE = 6.1) revealed high metabolic stability as 89% of the parent compound remained unchanged upon incubation with liver microsomes and NADPH over 90 min.
D7.4 = 1.1 and 2.0, respectively), both compounds exhibit high LLE values of 6.7 and 6.2, respectively. However, in the presence of mouse liver microsomes and NMDPH (phase I metabolism), both amines 3d and 3j were rapidly metabolized. In contrast, the less potent benzylamine 3b (Ki(σ1) = 69 nM, LLE = 6.1) revealed high metabolic stability as 89% of the parent compound remained unchanged upon incubation with liver microsomes and NADPH over 90 min.
In conclusion, the methylated cyclohexylmethylamine 3j showed the highest σ1 affinity (Ki = 6.4 nM) and the highest selectivity over the σ2 subtype (9-fold). The secondary cyclohexylmethylamine 3d exhibited the highest LLE value (6.7) and the benzylamine 3b was the metabolically most stable σ1 ligand of this series (89% intact after 90 min).
4. Experimental
4.1. Synthetic procedures
Pentane-1,3,5-triol19 (4, 3 g, 25 mmol), tert-butyldimethylsilyl chloride (7.5 g, 50 mmol) and imidazole (10.2 g, 150 mmol) were dissolved in DMF (30 mL) in a Schlenk flask under N2 atmosphere. The reaction mixture was stirred at rt for 6 h. After addition of water (50 mL) the organic layer was extracted with ethyl acetate (4 × 50 mL). The organic layer was then washed with brine (100 mL), dried (Na2SO4), concentrated in vacuo and the residue was purified by fc (∅ = 8 cm, h = 19.5 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 0.2
cyclohexane = 0.2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9.8, V = 65 mL, Rf = 0.08 (ethyl acetate
9.8, V = 65 mL, Rf = 0.08 (ethyl acetate![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 0.4
cyclohexane = 0.4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9.6)). Colorless oil, yield 6.1 g (71%). C17H40O3Si2 (348.3). 1H NMR (CDCl3): δ [ppm] = 0.07 (s, 12H, 2 × Si(CH3)2), 0.89 (s, 18H, 2 × SiC(CH3)3), 1.63–1.75 (m, 4H, CH2CH2OSi), 3.65 (d, J = 2.3 Hz, 1H, OH), 3.76–3.87 (m, 4H, CH2OSi), 3.95–4.02 (m, 1H, CH). IR: v [cm−1] = 3524 (b, ν, O–H), 2954 (w, ν, C–H, alkyl). MS (EI): m/z = 349 [M + H]+. Elemental analysis: calcd C 58.56, H 11.56; found C 58.54, H 11.80.
9.6)). Colorless oil, yield 6.1 g (71%). C17H40O3Si2 (348.3). 1H NMR (CDCl3): δ [ppm] = 0.07 (s, 12H, 2 × Si(CH3)2), 0.89 (s, 18H, 2 × SiC(CH3)3), 1.63–1.75 (m, 4H, CH2CH2OSi), 3.65 (d, J = 2.3 Hz, 1H, OH), 3.76–3.87 (m, 4H, CH2OSi), 3.95–4.02 (m, 1H, CH). IR: v [cm−1] = 3524 (b, ν, O–H), 2954 (w, ν, C–H, alkyl). MS (EI): m/z = 349 [M + H]+. Elemental analysis: calcd C 58.56, H 11.56; found C 58.54, H 11.80.
Under N2 atmosphere, a solution of alcohol 5 (2.2 g, 6.4 mmol) in CH2Cl2 (90 mL) was added to a solution of Dess Martin Periodinane (3.3 g, 7.7 mmol) in CH2Cl2 (75 mL). The reaction mixture was stirred for 4.5 h at rt. Et2O (150 mL) and 1 M NaOH (63 mL) were added and the mixture was stirred for another 15 min. The Et2O layer was separated and washed with NaOH (100 mL) and water (100 mL) followed by extraction of the combined aqueous layers with Et2O (4 × 200 mL). Finally, the organic layer was dried (Na2SO4), the solvent was removed in vacuo and the residue was purified by fc (∅ = 6 cm, h = 15.5 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 0.3
cyclohexane = 0.3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9.7, V = 30 mL, Rf = 0.17). Colorless oil, yield 2.2 g (97%). C17H38O3Si2 (346.2). 1H NMR (CDCl3): δ [ppm] = 0.05 (s, 12H, 2 × Si(CH3)2), 0.87 (s, 18H, 2 × SiC(CH3)3), 2.65 (t, J = 6.4 Hz, 4H, CH2CH2OSi), 3.88 (t, J = 6.4 Hz, 4H, CH2OSi). IR: v [cm−1] = 2955, 2857 (w, ν, C–H, alkyl), 1715 (s, ν, C
9.7, V = 30 mL, Rf = 0.17). Colorless oil, yield 2.2 g (97%). C17H38O3Si2 (346.2). 1H NMR (CDCl3): δ [ppm] = 0.05 (s, 12H, 2 × Si(CH3)2), 0.87 (s, 18H, 2 × SiC(CH3)3), 2.65 (t, J = 6.4 Hz, 4H, CH2CH2OSi), 3.88 (t, J = 6.4 Hz, 4H, CH2OSi). IR: v [cm−1] = 2955, 2857 (w, ν, C–H, alkyl), 1715 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). MS (EI): m/z = 347 [M + H]+, 289 [M − C(CH3)3]+. Elemental analysis: calcd C 58.90, H 11.05; found C 59.28, H 11.34.
O). MS (EI): m/z = 347 [M + H]+, 289 [M − C(CH3)3]+. Elemental analysis: calcd C 58.90, H 11.05; found C 59.28, H 11.34.
Under N2 atmosphere diethylamide 7 (368 mg, 1.92 mmol) was dissolved in abs. THF (125 mL) and cooled down to −78 °C. Then, 1.3 M sec-butyllithium in cyclohexane/hexane = 92
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8 (2.0 mL, 2.12 mmol) was added dropwise to the solution and the color changed from colorless to orange. After 1.5 h ketone 6 (1.0 g, 2.9 mmol) dissolved in THF (5 mL) was added using a syringe pump over 10 min. The reaction mixture turned yellow and was allowed to warm to −50 °C during 2.5 h. Water (50 mL) was added at −50 °C and after addition of brine (50 mL), the aqueous layer was extracted with chloroform (4 × 100 mL). The organic layer was dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 6 cm, h = 21 cm, ethyl acetate
8 (2.0 mL, 2.12 mmol) was added dropwise to the solution and the color changed from colorless to orange. After 1.5 h ketone 6 (1.0 g, 2.9 mmol) dissolved in THF (5 mL) was added using a syringe pump over 10 min. The reaction mixture turned yellow and was allowed to warm to −50 °C during 2.5 h. Water (50 mL) was added at −50 °C and after addition of brine (50 mL), the aqueous layer was extracted with chloroform (4 × 100 mL). The organic layer was dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 6 cm, h = 21 cm, ethyl acetate![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 2
cyclohexane = 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8, V = 10 mL, Rf = 0.42). Colorless oil, yield 949 mg (92%). C29H55NO4Si2 (537.4). 1H NMR (CDCl3): δ [ppm] = 0.05 (s, 12H, 2 × Si(CH3)2), 0.88 (s, 18H, 2 × SiC(CH3)3), 1.04 (t, J = 7.1 Hz, 3H, CH2CH3), 1.26 (t, J = 7.1 Hz, 3H, CH2CH3), 1.56–1.80 (m, 4H, CH2CH2OSi), 2.55–2.74 (m, 1H, CH2OSi), 2.86–3.05 (m, 1H, CH2OSi), 3.09–3.14 (m, 2H, CH2CH3), 3.31–3.51 (m, 1H, CH2OSi), 3.58–3.75 (m, 1H, CH2OSi), 3.76–3.92 (m, 4H, CH2CH3 and CH2Ph), 4.79 (s, 1H, OH), 7.16 (dd, J = 7.6/1.4 Hz, 1H, CHarom.), 7.22 (td, J = 7.5/1.3 Hz, 1H, CHarom.), 7.31 (td, J = 7.6/1.6 Hz, 1H, CHarom.), 7.53 (dd, J = 7.7/0.9 Hz, 1H, CHarom.). 13C NMR (CDCl3): δ [ppm] = −5.2 (6C, Si(CH3)3), 13.0 (1C, CH2CH3), 14.1 (1C, CH2CH3), 26.1 (6C, SiC(CH3)3), 39.3 (2C, OCH2CH2), 41.6 (1C, OCH2), 43.0 (1C, OCH2), 43.3 (1C, CH2CH3), 60.1 (1C, CH2Ph), 60.2 (1C, CH2CH3), 73.5 (1C, Cq), 125.7 (1C, CHarom), 126.1 (1C, CHarom), 128.4 (1C, CHarom), 132.2 (1C, CHarom), 135.3 (1C, Cqarom), 137.5 (1C, Cqarom), 172.0 (1C, C
8, V = 10 mL, Rf = 0.42). Colorless oil, yield 949 mg (92%). C29H55NO4Si2 (537.4). 1H NMR (CDCl3): δ [ppm] = 0.05 (s, 12H, 2 × Si(CH3)2), 0.88 (s, 18H, 2 × SiC(CH3)3), 1.04 (t, J = 7.1 Hz, 3H, CH2CH3), 1.26 (t, J = 7.1 Hz, 3H, CH2CH3), 1.56–1.80 (m, 4H, CH2CH2OSi), 2.55–2.74 (m, 1H, CH2OSi), 2.86–3.05 (m, 1H, CH2OSi), 3.09–3.14 (m, 2H, CH2CH3), 3.31–3.51 (m, 1H, CH2OSi), 3.58–3.75 (m, 1H, CH2OSi), 3.76–3.92 (m, 4H, CH2CH3 and CH2Ph), 4.79 (s, 1H, OH), 7.16 (dd, J = 7.6/1.4 Hz, 1H, CHarom.), 7.22 (td, J = 7.5/1.3 Hz, 1H, CHarom.), 7.31 (td, J = 7.6/1.6 Hz, 1H, CHarom.), 7.53 (dd, J = 7.7/0.9 Hz, 1H, CHarom.). 13C NMR (CDCl3): δ [ppm] = −5.2 (6C, Si(CH3)3), 13.0 (1C, CH2CH3), 14.1 (1C, CH2CH3), 26.1 (6C, SiC(CH3)3), 39.3 (2C, OCH2CH2), 41.6 (1C, OCH2), 43.0 (1C, OCH2), 43.3 (1C, CH2CH3), 60.1 (1C, CH2Ph), 60.2 (1C, CH2CH3), 73.5 (1C, Cq), 125.7 (1C, CHarom), 126.1 (1C, CHarom), 128.4 (1C, CHarom), 132.2 (1C, CHarom), 135.3 (1C, Cqarom), 137.5 (1C, Cqarom), 172.0 (1C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). IR: v [cm−1] = 2951 (s, ν, C–H, alkyl), 1613 (s, ν, C
O). IR: v [cm−1] = 2951 (s, ν, C–H, alkyl), 1613 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 772 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C29H55NO4Si2H [M + H] 538.3742, found 538.3754. HPLC: Compound 8 was not stable during HPLC analysis.
O), 772 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C29H55NO4Si2H [M + H] 538.3742, found 538.3754. HPLC: Compound 8 was not stable during HPLC analysis.
Diethylamide 8 (1.0 g, 1.93 mmol) was heated in a silicone bath to 186 °C overnight. The residue was purified by fc (∅ = 4 cm, h = 17.5 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 5
cyclohexane = 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95, V = 20 mL, Rf = 0.6). Colorless oil, yield 736 mg (82%). C25H44O4Si2 (464.3). 1H NMR (CDCl3): δ [ppm] = 0.02, 0.03 (2s, 12H, Si(CH3)2), 0.87 (s, 18H, SiC(CH3)3), 2.00 (t, J = 6.5 Hz, 4H, CH2CH2OSi), 3.21 (s, 2H, ArCH2), 3.79 (td, J = 6.4/1.8 Hz, 4H, CH2OSi), 7.19 (d, J = 7.5 Hz, 1H, 5-CHarom), 7.36 (td, J = 7.7/1.1 Hz, 1H, 7-CHarom), 7.52 (td, J = 7.5/1.4 Hz, 1H, 6-CHarom), 8.07 (dd, J = 7.7/1.1 Hz, 1H, 8-CHarom). 13C NMR (CDCl3): δ [ppm] = −5.3 (2C, Si(CH3)2), −5.3 (2C, Si(CH3)2), 18.4 (2C, C(CH3)3), 26.1 (6C, SiC(CH3)3), 36.9 (1C, CH2Ph), 40.3 (2C, CH2CH2OSi), 58.9 (2C, CH2OSi), 84.0 (1C, C-3), 125.2 (1C, CHarom), 127.5 (1C, CHarom), 128.3 (1C, CHarom), 130.0 (1C, CHarom), 134.0 (1C, Cqarom), 138.5 (1C, Cqarom), 165.1 (1C, C
95, V = 20 mL, Rf = 0.6). Colorless oil, yield 736 mg (82%). C25H44O4Si2 (464.3). 1H NMR (CDCl3): δ [ppm] = 0.02, 0.03 (2s, 12H, Si(CH3)2), 0.87 (s, 18H, SiC(CH3)3), 2.00 (t, J = 6.5 Hz, 4H, CH2CH2OSi), 3.21 (s, 2H, ArCH2), 3.79 (td, J = 6.4/1.8 Hz, 4H, CH2OSi), 7.19 (d, J = 7.5 Hz, 1H, 5-CHarom), 7.36 (td, J = 7.7/1.1 Hz, 1H, 7-CHarom), 7.52 (td, J = 7.5/1.4 Hz, 1H, 6-CHarom), 8.07 (dd, J = 7.7/1.1 Hz, 1H, 8-CHarom). 13C NMR (CDCl3): δ [ppm] = −5.3 (2C, Si(CH3)2), −5.3 (2C, Si(CH3)2), 18.4 (2C, C(CH3)3), 26.1 (6C, SiC(CH3)3), 36.9 (1C, CH2Ph), 40.3 (2C, CH2CH2OSi), 58.9 (2C, CH2OSi), 84.0 (1C, C-3), 125.2 (1C, CHarom), 127.5 (1C, CHarom), 128.3 (1C, CHarom), 130.0 (1C, CHarom), 134.0 (1C, Cqarom), 138.5 (1C, Cqarom), 165.1 (1C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). IR: v [cm−1] = 2951 (s, ν, C–H, alkyl), 1720 (s, ν, C
O). IR: v [cm−1] = 2951 (s, ν, C–H, alkyl), 1720 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 752 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C25H44O4Si2H [M + H] 465.2864, found 465.2848, calcd for C25H44O4Si2 − C(CH3)3 + H [M − C(CH3) + H] 407.2074, found 407.2051. Purity (HPLC): 97.1%, tR = 28.81 min.
O), 752 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C25H44O4Si2H [M + H] 465.2864, found 465.2848, calcd for C25H44O4Si2 − C(CH3)3 + H [M − C(CH3) + H] 407.2074, found 407.2051. Purity (HPLC): 97.1%, tR = 28.81 min.
Lactone 9 was dissolved in CH2Cl2 (33 mL) and cooled down to −78 °C. 1M DIBAH solution in toluene (1.7 mL, 1.7 mmol) was added and the reaction mixture was stirred at −78 °C for 1 h and 10 min. After addition of saturated NaHCO3 solution (20 mL) at −78 °C and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 25 mL). The organic layer was dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 4 cm, h = 20 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 5
cyclohexane = 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95, V = 20 mL, Rf = 0.33 (ethyl acetate
95, V = 20 mL, Rf = 0.33 (ethyl acetate![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 1.9)). Colorless oil, yield 598 mg (93%). C25H46O4Si2 (466.3). 1H NMR (CDCl3): δ [ppm] = 0.017, 0.021, 0.027, 0.030 (4s, 12H, Si(CH3)2), 0.87, 0.88 (2s, 18H, SiC(CH3)3), 1.80 (t, J = 7.3 Hz, 2H, CH2CH2OSi), 1.92 (t, J = 6.6 Hz, 1H, CH2CH2OSi), 1.93 (t, J = 6.9 Hz, 1H, CH2CH2OSi), 2.80 (d, J = 15.8 Hz, 1H, CH2Ph), 2.99 (d, J = 15.8 Hz, 1H, CH2Ph), 3.74 (t, J = 7.0 Hz, 2H, CH2OSi), 3.76 (t, J = 6.8 Hz, 2H, CH2OSi), 5.52 (s, 1H, CH), 7.06–7.08 (m, 1H, CHarom), 7.16–7.18 (m, 1H, CHarom), 7.21–7.26 (m, 2H, CHarom). IR: v [cm−1] = 3406 (b, ν, O–H), 2951 (s, ν, C–H, alkyl), 748 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C25H46O4Si2 − OH [M − OH] 449.2947, found 449.2950. HPLC: Compound 10 was not stable during HPLC analysis.
cyclohexane = 1.9)). Colorless oil, yield 598 mg (93%). C25H46O4Si2 (466.3). 1H NMR (CDCl3): δ [ppm] = 0.017, 0.021, 0.027, 0.030 (4s, 12H, Si(CH3)2), 0.87, 0.88 (2s, 18H, SiC(CH3)3), 1.80 (t, J = 7.3 Hz, 2H, CH2CH2OSi), 1.92 (t, J = 6.6 Hz, 1H, CH2CH2OSi), 1.93 (t, J = 6.9 Hz, 1H, CH2CH2OSi), 2.80 (d, J = 15.8 Hz, 1H, CH2Ph), 2.99 (d, J = 15.8 Hz, 1H, CH2Ph), 3.74 (t, J = 7.0 Hz, 2H, CH2OSi), 3.76 (t, J = 6.8 Hz, 2H, CH2OSi), 5.52 (s, 1H, CH), 7.06–7.08 (m, 1H, CHarom), 7.16–7.18 (m, 1H, CHarom), 7.21–7.26 (m, 2H, CHarom). IR: v [cm−1] = 3406 (b, ν, O–H), 2951 (s, ν, C–H, alkyl), 748 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C25H46O4Si2 − OH [M − OH] 449.2947, found 449.2950. HPLC: Compound 10 was not stable during HPLC analysis.
Lactol 10 (1.3 g, 2.8 mmol) was dissolved in abs. THF (104 mL) and acidified with 1M HCl (pH 1, 11 mL). The reaction mixture was stirred overnight at rt. After removing the THF in vacuo, brine (25 mL) was added and the aqueous layer was extracted with chloroform (4 × 50 mL). The residue was dried (Na2SO4) and purified by fc (∅ = 8 cm, h = 18 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 1
cyclohexane = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1–>6
1–>6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4, V = 65 mL, Rf = 0.28). Colorless oil, yield 613 mg (99%). C13H16O3 (220.1). 1H NMR (CDCl3): δ [ppm] = 1.49 (d broad, J = 13.3 Hz, 1H, 4-Heq), 1.92 (td, J = 5.7/2.8 Hz, 2H, CH2CH2OH), 2.30–2.39 (m, 1H, 4-Hax), 2.76 (d, J = 17.5 Hz, 1H, CH2Ph), 3.23 (d, J = 17.5 Hz, 1H, CH2Ph), 3.66–3.70 (m, 2H, 3-Hax and 3-Heq), 3.91 (ddd, J = 6.4/5.3/1.4 Hz, 2H, CH2OH), 5.94 (s, 1H, 1-H), 7.14 (d, J = 7.6 Hz, 1H, CHarom), 7.17 (dd, J = 7.5/1.5 Hz, 1H, CHarom), 7.24 (t, J = 11.1 Hz, 1H, CHarom), 7.30 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.8 (1C, C-6), 45.0 (1C, CH2CH2OH), 57.0 (1C, C-3), 58.9 (1C, CH2OH), 72.8 (1C, C-5), 94.2 (1C, C-1), 126.6 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.7 (1C, CHarom), 131.9 (1C, Cqarom), 135.0 (1C, Cqarom). IR: v [cm−1] = 3406 (b, ν, O–H), 2943 (s, ν, C–H, alkyl), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C13H16O3H [M + H] 221.1172, found 221.1201. Purity (HPLC): 99.7%, tR = 13.65 min.
4, V = 65 mL, Rf = 0.28). Colorless oil, yield 613 mg (99%). C13H16O3 (220.1). 1H NMR (CDCl3): δ [ppm] = 1.49 (d broad, J = 13.3 Hz, 1H, 4-Heq), 1.92 (td, J = 5.7/2.8 Hz, 2H, CH2CH2OH), 2.30–2.39 (m, 1H, 4-Hax), 2.76 (d, J = 17.5 Hz, 1H, CH2Ph), 3.23 (d, J = 17.5 Hz, 1H, CH2Ph), 3.66–3.70 (m, 2H, 3-Hax and 3-Heq), 3.91 (ddd, J = 6.4/5.3/1.4 Hz, 2H, CH2OH), 5.94 (s, 1H, 1-H), 7.14 (d, J = 7.6 Hz, 1H, CHarom), 7.17 (dd, J = 7.5/1.5 Hz, 1H, CHarom), 7.24 (t, J = 11.1 Hz, 1H, CHarom), 7.30 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.8 (1C, C-6), 45.0 (1C, CH2CH2OH), 57.0 (1C, C-3), 58.9 (1C, CH2OH), 72.8 (1C, C-5), 94.2 (1C, C-1), 126.6 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.7 (1C, CHarom), 131.9 (1C, Cqarom), 135.0 (1C, Cqarom). IR: v [cm−1] = 3406 (b, ν, O–H), 2943 (s, ν, C–H, alkyl), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C13H16O3H [M + H] 221.1172, found 221.1201. Purity (HPLC): 99.7%, tR = 13.65 min.
Alcohol 11 (649 mg, 2.94 mmol) was dissolved in dry THF (49 mL). Then Zn(N3)2·2 pyridine (907 mg, 2.94 mmol), PPh3 (1.5 g, 5.9 mmol) and THF (31 mL) were added. DIAD (1.2 mL, 5.9 mmol) was added dropwise and the reaction mixture was stirred at rt for 19 h. After addition of 1 M NaOH (50 mL) and brine (25 mL), the aqueous layer was extracted with ethyl acetate (4 × 50 mL). The organic layer was dried (Na2SO4) and after evaporation of the solvent the residue was purified by fc (∅ = 8 cm, h = 17.5 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane = 4
cyclohexane = 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6, V = 65 mL, Rf = 0.74 (cyclohexane
6, V = 65 mL, Rf = 0.74 (cyclohexane![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 6
ethyl acetate = 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4)). Colorless oil, yield 702 mg (97%). C13H15N3O2 (245.1). 1H NMR (CDCl3): δ [ppm] = 1.53 (d, J = 13.4 Hz, 1H, 4-Heq), 1.94 (td, J = 7.1/0.9 Hz, 2H, CH2CH2N3), 2.15–2.24 (m, 1H, 4-Hax), 2.78 (d, J = 17.4 Hz, 1H, CH2Ph), 3.12 (d, J = 17.4 Hz, 1H, CH2Ph), 3.50 (t, J = 7.4 Hz, 2H, CH2N3), 3.60–3.70 (m, 2H, 3-Hax and 3-Heq), 5.93 (s, 1H, 1-H), 7.13 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (dd, J = 7.5/1.3 Hz, 1H, CHarom), 7.24 (t, J = 7.4 Hz, 1H, CHarom), 7.29 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.46 (1C, C-4), 36.51 (1C, C-6), 42.3 (1C, CH2CH2N3), 46.1 (1C, CH2N3), 56.9 (1C, C-3), 70.5 (1C, C-5), 94.4 (1C, C-1), 126.7 (2C, CHarom), 127.3 (1C, CHarom), 128.7 (1C, CHarom), 132.0 (1C, Cqarom), 134.7 (1C, Cqarom). IR: v [cm−1] = 2951 (s, ν, C–H, alkyl), 2091 (s, ν, N
4)). Colorless oil, yield 702 mg (97%). C13H15N3O2 (245.1). 1H NMR (CDCl3): δ [ppm] = 1.53 (d, J = 13.4 Hz, 1H, 4-Heq), 1.94 (td, J = 7.1/0.9 Hz, 2H, CH2CH2N3), 2.15–2.24 (m, 1H, 4-Hax), 2.78 (d, J = 17.4 Hz, 1H, CH2Ph), 3.12 (d, J = 17.4 Hz, 1H, CH2Ph), 3.50 (t, J = 7.4 Hz, 2H, CH2N3), 3.60–3.70 (m, 2H, 3-Hax and 3-Heq), 5.93 (s, 1H, 1-H), 7.13 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (dd, J = 7.5/1.3 Hz, 1H, CHarom), 7.24 (t, J = 7.4 Hz, 1H, CHarom), 7.29 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.46 (1C, C-4), 36.51 (1C, C-6), 42.3 (1C, CH2CH2N3), 46.1 (1C, CH2N3), 56.9 (1C, C-3), 70.5 (1C, C-5), 94.4 (1C, C-1), 126.7 (2C, CHarom), 127.3 (1C, CHarom), 128.7 (1C, CHarom), 132.0 (1C, Cqarom), 134.7 (1C, Cqarom). IR: v [cm−1] = 2951 (s, ν, C–H, alkyl), 2091 (s, ν, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N
N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C13H15N3O2 − N2 + H [M − N2 + H] 218.1181, found 218.1161. Purity (HPLC): 97.3%, tR = 19.11 min.
N), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C13H15N3O2 − N2 + H [M − N2 + H] 218.1181, found 218.1161. Purity (HPLC): 97.3%, tR = 19.11 min.
Azide 12 (702 mg, 2.86 mmol) was dissolved in ethyl acetate (27 mL) and Pd/C (112 mg, 16%) was added. The mixture was stirred under a H2 atmosphere (1 atm) overnight. It was filtered through Celite®, the filtrate was concentrated in vacuo and the residue was purified by fc (∅ = 4.5 cm, h = 18 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 50
NH3 = 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50
50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, V = 20 mL, Rf = 0.28 (ethyl acetate: methanol = 8
1, V = 20 mL, Rf = 0.28 (ethyl acetate: methanol = 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2)). Colorless oil, yield 472 mg (75%). C13H17NO2 (219.1). 1H NMR (CDCl3): δ [ppm] = 1.50 (d, J = 13.3 Hz, 1H, 4-Heq), 1.81 (t, J = 7.8 Hz, 2H, CH2CH2NH2), 2.17–2.25 (m, 1H, 4-Hax), 2.74 (d, J = 17.4 Hz, 1H, CH2Ph), 2.90 (t, J = 7.5 Hz, 2H, CH2NH2), 3.12 (d, J = 17.4 Hz, 1H, CH2Ph), 3.59–3.69 (m, 2H, 3-Hax and 3-Heq), 5.92 (s, 1H, 1-H), 7.12 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (dd, J = 7.4/1.3 Hz, 1H, CHarom), 7.22 (t, J = 7.3 Hz, 1H, CHarom), 7.28 (td, J = 7.4/1.7 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.7 (1C, C-6), 36.8 (1C, CH2CH2NH2), 47.8 (1C, CH2NH2), 57.7 (1C, C-3), 71.3 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.6 (1C, CHarom), 132.3 (1C, Cqarom), 135.2 (1C, Cqarom). IR: v [cm−1] = 3364 (b, ν, N–H), 2940 (s, ν, C–H, alkyl), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C13H17NO2 + H [M + H] 220.1332, found 220.1312. Purity (HPLC): 99.9%, tR = 10.18 min.
2)). Colorless oil, yield 472 mg (75%). C13H17NO2 (219.1). 1H NMR (CDCl3): δ [ppm] = 1.50 (d, J = 13.3 Hz, 1H, 4-Heq), 1.81 (t, J = 7.8 Hz, 2H, CH2CH2NH2), 2.17–2.25 (m, 1H, 4-Hax), 2.74 (d, J = 17.4 Hz, 1H, CH2Ph), 2.90 (t, J = 7.5 Hz, 2H, CH2NH2), 3.12 (d, J = 17.4 Hz, 1H, CH2Ph), 3.59–3.69 (m, 2H, 3-Hax and 3-Heq), 5.92 (s, 1H, 1-H), 7.12 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (dd, J = 7.4/1.3 Hz, 1H, CHarom), 7.22 (t, J = 7.3 Hz, 1H, CHarom), 7.28 (td, J = 7.4/1.7 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.7 (1C, C-6), 36.8 (1C, CH2CH2NH2), 47.8 (1C, CH2NH2), 57.7 (1C, C-3), 71.3 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.6 (1C, CHarom), 132.3 (1C, Cqarom), 135.2 (1C, Cqarom). IR: v [cm−1] = 3364 (b, ν, N–H), 2940 (s, ν, C–H, alkyl), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C13H17NO2 + H [M + H] 220.1332, found 220.1312. Purity (HPLC): 99.9%, tR = 10.18 min.
NEt3 (40 mg, 0.39 mmol) was added to a solution of 3a-HCl (50 mg, 0.20 mmol) in CH2Cl2 (6 mL) to obtain the free primary amine. Then, benzaldehyde (30 µL, 0.29 mmol) and NaBH(OAc)3 (83 mg, 0.39 mmol) were added. The reaction mixture was stirred overnight at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 20 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 1.5 cm, h = 13.0 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 4
NH3 = 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 96.9
96.9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 5 mL). The fraction with the Rf = 0.55 of the first fc purification was purified by another fc (∅ = 1.5 cm, h = 15 cm, cyclohexane
0.1, V = 5 mL). The fraction with the Rf = 0.55 of the first fc purification was purified by another fc (∅ = 1.5 cm, h = 15 cm, cyclohexane![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate
ethyl acetate![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N,N-dimethylethylamine = 94
N,N-dimethylethylamine = 94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6
6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 5 mL).
0.1, V = 5 mL).
Compound 3b (Rf = 0.33 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 2
ethyl acetate = 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8)): colorless oil, yield 9 mg (16%). C20H23NO2 (309.2). 1H NMR (CDCl3): δ [ppm] = 1.49 (d, J = 13.3 Hz, 1H, 4-Heq), 1.80 (s broad, 1H, NH), 1.88 (t, J = 7.9 Hz, 2H, CH2CH2NH), 2.16–2.24 (m, 1H, 4-Hax), 2.74 (d, J = 17.5 Hz, 1H, 6-H), 2.84 (t, J = 7.4 Hz, 2H, CH2NH), 3.12 (d, J = 17.5 Hz, 1H, 6-H), 3.60–3.66 (m, 2H, 3-Hax and 3-Heq), 3.82 (s, 2H, CH2Ph), 5.93 (s, 1H, 1-H), 7.11 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (d, J = 7.4 Hz, 1H, CHarom), 7.21–7.33 (m, 7H, CHarom).
8)): colorless oil, yield 9 mg (16%). C20H23NO2 (309.2). 1H NMR (CDCl3): δ [ppm] = 1.49 (d, J = 13.3 Hz, 1H, 4-Heq), 1.80 (s broad, 1H, NH), 1.88 (t, J = 7.9 Hz, 2H, CH2CH2NH), 2.16–2.24 (m, 1H, 4-Hax), 2.74 (d, J = 17.5 Hz, 1H, 6-H), 2.84 (t, J = 7.4 Hz, 2H, CH2NH), 3.12 (d, J = 17.5 Hz, 1H, 6-H), 3.60–3.66 (m, 2H, 3-Hax and 3-Heq), 3.82 (s, 2H, CH2Ph), 5.93 (s, 1H, 1-H), 7.11 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (d, J = 7.4 Hz, 1H, CHarom), 7.21–7.33 (m, 7H, CHarom).
13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.7 (1C, C-6), 43.8 (1C, CH2CH2NH), 43.8 (1C, CH2NH), 54.3 (1C, CH2Ph), 57.1 (1C, C-3), 71.4 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.1 (2C, CHarom), 127.3 (1C, CHarom), 128.3 (2C, CHarom), 128.6 (2C, CHarom), 132.2 (1C, Cqarom), 135.2 (1C, Cqarom), 140.2 (1C, Cqarom). IR: v [cm−1] = 3329 (w, ν, N–H), 3028 (s, ν, C–H, aryl), 2932 (s, ν, C–H, alkyl), 1605 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, aryl), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C20H23NO2H [M + H] 310.1802, found 310.1802. Purity (HPLC): 99.5%, tR = 15.90 min.
C, aryl), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C20H23NO2H [M + H] 310.1802, found 310.1802. Purity (HPLC): 99.5%, tR = 15.90 min.
Compound 3c (Rf = 0.55 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 2
ethyl acetate = 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8)): colorless oil, yield 35 mg (45%). C27H29NO2 (399.2). 1H NMR (CDCl3): δ [ppm] = 1.29 (d, J = 13.6 Hz, 1H, 4-Heq), 1.79 (ddd, J = 9.1/6.5/2.6 Hz, 2H, CH2CH2N), 1.94 (td, J = 12.4/7.2 Hz, 1H, 4-Hax), 2.47 (d, J = 17.2 Hz, 1H, 6-H), 2.49–2.53 (m, 2H, CH2N), 2.85 (d, J = 17.5 Hz, 1H, 6-H), 3.40–3.54 (m, 6H, CH2Ph and 3-Hax and 3-Heq), 5.77 (s, 1H, 1-H), 6.92 (d, J = 7.4 Hz, 1H, CHarom), 7.03 (d, J = 7.4 Hz, 1H, CHarom), 7.07–7.16 (m, 4H, CHarom), 7.20 (t, J = 7.4 Hz, 4H, CHarom), 7.27 (d, J = 7.1 Hz, 4H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.3 (1C, C-6), 36.6 (1C, C-4), 41.1 (1C, CH2CH2N), 47.1 (1C, CH2N), 57.0 (2C, CH2Ph), 58.4 (1C, C-3), 71.2 (1C, C-5), 94.3 (1C, C-1), 126.4 (1C, CHarom), 126.6 (1C, CHarom), 127.0 (1C, CHarom), 127.3 (1C, CHarom), 128.3 (4C, CHarom), 128.5 (2C, CHarom), 129.0 (4C, CHarom), 132.3 (1C, Cqarom), 135.2 (1C, Cqarom), 139.7 (2C, Cqarom). IR: v [cm−1] = 3059 (s, ν, C–H, aryl), 2936 (s, ν, C–H, alkyl), 1601 (s, ν, C
8)): colorless oil, yield 35 mg (45%). C27H29NO2 (399.2). 1H NMR (CDCl3): δ [ppm] = 1.29 (d, J = 13.6 Hz, 1H, 4-Heq), 1.79 (ddd, J = 9.1/6.5/2.6 Hz, 2H, CH2CH2N), 1.94 (td, J = 12.4/7.2 Hz, 1H, 4-Hax), 2.47 (d, J = 17.2 Hz, 1H, 6-H), 2.49–2.53 (m, 2H, CH2N), 2.85 (d, J = 17.5 Hz, 1H, 6-H), 3.40–3.54 (m, 6H, CH2Ph and 3-Hax and 3-Heq), 5.77 (s, 1H, 1-H), 6.92 (d, J = 7.4 Hz, 1H, CHarom), 7.03 (d, J = 7.4 Hz, 1H, CHarom), 7.07–7.16 (m, 4H, CHarom), 7.20 (t, J = 7.4 Hz, 4H, CHarom), 7.27 (d, J = 7.1 Hz, 4H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.3 (1C, C-6), 36.6 (1C, C-4), 41.1 (1C, CH2CH2N), 47.1 (1C, CH2N), 57.0 (2C, CH2Ph), 58.4 (1C, C-3), 71.2 (1C, C-5), 94.3 (1C, C-1), 126.4 (1C, CHarom), 126.6 (1C, CHarom), 127.0 (1C, CHarom), 127.3 (1C, CHarom), 128.3 (4C, CHarom), 128.5 (2C, CHarom), 129.0 (4C, CHarom), 132.3 (1C, Cqarom), 135.2 (1C, Cqarom), 139.7 (2C, Cqarom). IR: v [cm−1] = 3059 (s, ν, C–H, aryl), 2936 (s, ν, C–H, alkyl), 1601 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, aryl), 733 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C27H29NO2H [M + H] 400.2271, found 400.2223; calcd for C27H29NO2 − CH2Ph [M − CH2Ph] 308.1651, found 308.1649. Purity (HPLC): 98.6%, tR = 19.61 min.
C, aryl), 733 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C27H29NO2H [M + H] 400.2271, found 400.2223; calcd for C27H29NO2 − CH2Ph [M − CH2Ph] 308.1651, found 308.1649. Purity (HPLC): 98.6%, tR = 19.61 min.
Cyclohexanecarbaldehyde (45 µL, 0.37 mmol) and NaBH(OAc)3 (158 mg, 0.74 mmol) were added to a solution of the primary amine 3a (82 mg, 0.37 mmol) in CH2Cl2 (11.5 mL). The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 20 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 3 cm, h = 1.5 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 20
NH3 = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80
80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.13 (methanol
0.1, V = 10 mL, Rf = 0.13 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 41 mg (35%). C20H29NO2 (315.2). 1H NMR (CDCl3): δ [ppm] = 0.85–0.94 (m, 2H, CH(CH2)5), 1.12–1.28 (m, 3H, CH(CH2)5), 1.49 (d, J = 13.2 Hz, 1H, 4-Heq), 1.64–1.75 (m, 5H, CH(CH2)5), 1.86 (t, J = 7.8 Hz, 2H, CH2CH2N), 2.17–2.24 (m, 2H, 4-Hax, CH(CH2)5), 2.47 (d, J = 6.6 Hz, 2H, CH2CH(CH2)5), 2.74 (d, J = 17.5 Hz, 1H, 6-H), 2.79 (t, J = 7.4 Hz, 2H, CH2N), 3.12 (d, J = 17.5 Hz, 1H, 6-H), 3.62–3.69 (m, 2H, 3-Hax and 3-Heq), 5.91 (s, 1H, 1-H), 7.11 (d, J = 7.4 Hz, 1H, CHarom), 7.16 (d, J = 7.4 Hz, 1H, CHarom), 7.21 (t, J = 7.2 Hz, 1H, CHarom), 7.27 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 26.1 (2C, CH(CH2)5), 26.7 (2C, CH(CH2)5), 31.5 (1C, CH(CH2)5), 36.6 (1C, C-4), 36.7 (1C, C-6), 37.8 (1C, CH(CH2)5), 43.7 (1C, CH2CH2NH), 44.4 (1C, CH2NH), 56.9 (1C, CH2CH(CH2)5), 57.0 (1C, C-3), 71.3 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.6 (1C, CHarom), 132.2 (1C, Cqarom), 135.2 (1C, Cqarom). IR: v [cm−1] = 2920 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C20H29NO2H [M + H] 316.2271, found 316.2234; calcd for C20H29NO2 − Cy [M − Cy] 232.1338, found 232.1306. Purity (HPLC): 95.6%, tR = 17.15 min.
80)). Colorless oil, yield 41 mg (35%). C20H29NO2 (315.2). 1H NMR (CDCl3): δ [ppm] = 0.85–0.94 (m, 2H, CH(CH2)5), 1.12–1.28 (m, 3H, CH(CH2)5), 1.49 (d, J = 13.2 Hz, 1H, 4-Heq), 1.64–1.75 (m, 5H, CH(CH2)5), 1.86 (t, J = 7.8 Hz, 2H, CH2CH2N), 2.17–2.24 (m, 2H, 4-Hax, CH(CH2)5), 2.47 (d, J = 6.6 Hz, 2H, CH2CH(CH2)5), 2.74 (d, J = 17.5 Hz, 1H, 6-H), 2.79 (t, J = 7.4 Hz, 2H, CH2N), 3.12 (d, J = 17.5 Hz, 1H, 6-H), 3.62–3.69 (m, 2H, 3-Hax and 3-Heq), 5.91 (s, 1H, 1-H), 7.11 (d, J = 7.4 Hz, 1H, CHarom), 7.16 (d, J = 7.4 Hz, 1H, CHarom), 7.21 (t, J = 7.2 Hz, 1H, CHarom), 7.27 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 26.1 (2C, CH(CH2)5), 26.7 (2C, CH(CH2)5), 31.5 (1C, CH(CH2)5), 36.6 (1C, C-4), 36.7 (1C, C-6), 37.8 (1C, CH(CH2)5), 43.7 (1C, CH2CH2NH), 44.4 (1C, CH2NH), 56.9 (1C, CH2CH(CH2)5), 57.0 (1C, C-3), 71.3 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.6 (1C, CHarom), 132.2 (1C, Cqarom), 135.2 (1C, Cqarom). IR: v [cm−1] = 2920 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C20H29NO2H [M + H] 316.2271, found 316.2234; calcd for C20H29NO2 − Cy [M − Cy] 232.1338, found 232.1306. Purity (HPLC): 95.6%, tR = 17.15 min.
NEt3 (40 mg, 0.39 mmol) was added to a solution of 3a-HCl (50 mg, 0.20 mmol) in CH2Cl2 (6 mL) to obtain the free primary amine. Then, cyclohexanecarbaldehyde (36 µL, 0.29 mmol) and NaBH(OAc)3 (83 mg, 0.39 mmol) were added. The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 15 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 1.5 cm, h = 13 cm, ethyl acetate
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cyclohexane
cyclohexane![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N,N-dimethylethylamine = 6
N,N-dimethylethylamine = 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 94
94![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 5 mL, Rf = 0.34 (methanol
0.1, V = 5 mL, Rf = 0.34 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 41 mg (51%). C27H41NO2 (411.3). 1H NMR (CDCl3): δ [ppm] = 0.77–0.85 (m, 5H, CH(CH2)5), 1.13–2.25 (m, 6H, CH(CH2)5), 1.38 (s broad, 2H, CH(CH2)5), 1.51 (d, J = 13.3 Hz, 1H, 4-Heq), 1.67–1.80 (m, 11H, CH(CH2)5), 2.10–2.21 (m, 5H, CH2CH2N, CH2CH(CH2)5, 4-Hax), 2.49 (d, J = 7.8 Hz, 2H, CH2N), 2.73 (d, J = 17.5 Hz, 1H, 6-H), 3.16 (d, J = 17.5 Hz, 1H, 6-H), 3.62–3.65 (m, 2H, 3-Hax and 3-Heq), 5.93 (s, 1H, 1-H), 7.12 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (d, J = 7.4 Hz, 1H, CHarom), 7.21 (t, J = 6.9 Hz, 1H, CHarom), 7.27 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 26.4 (4C, CH(CH2)5), 27.1 (2C, CH(CH2)5), 32.1 (4C, CH(CH2)5), 36.4 (2C, CH(CH2)5), 36.6 (1C, C-6), 36.8 (1C, C-4), 41.4 (1C, CH2CH2N), 48.9 (1C, CH2N), 57.1 (1C, C-3), 62.7 (2C, CH2CH(CH2)5), 71.2 (1C, C-5), 94.4 (1C, C-1), 126.4 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.5 (1C, CHarom), 132.4 (1C, Cqarom), 135.4 (1C, Cqarom). IR: v [cm−1] = 2920 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C27H41NO2H [M + H] 412.3210, found 412.3201. Purity (HPLC): 96.1%, tR = 22.02 min.
80)). Colorless oil, yield 41 mg (51%). C27H41NO2 (411.3). 1H NMR (CDCl3): δ [ppm] = 0.77–0.85 (m, 5H, CH(CH2)5), 1.13–2.25 (m, 6H, CH(CH2)5), 1.38 (s broad, 2H, CH(CH2)5), 1.51 (d, J = 13.3 Hz, 1H, 4-Heq), 1.67–1.80 (m, 11H, CH(CH2)5), 2.10–2.21 (m, 5H, CH2CH2N, CH2CH(CH2)5, 4-Hax), 2.49 (d, J = 7.8 Hz, 2H, CH2N), 2.73 (d, J = 17.5 Hz, 1H, 6-H), 3.16 (d, J = 17.5 Hz, 1H, 6-H), 3.62–3.65 (m, 2H, 3-Hax and 3-Heq), 5.93 (s, 1H, 1-H), 7.12 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (d, J = 7.4 Hz, 1H, CHarom), 7.21 (t, J = 6.9 Hz, 1H, CHarom), 7.27 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 26.4 (4C, CH(CH2)5), 27.1 (2C, CH(CH2)5), 32.1 (4C, CH(CH2)5), 36.4 (2C, CH(CH2)5), 36.6 (1C, C-6), 36.8 (1C, C-4), 41.4 (1C, CH2CH2N), 48.9 (1C, CH2N), 57.1 (1C, C-3), 62.7 (2C, CH2CH(CH2)5), 71.2 (1C, C-5), 94.4 (1C, C-1), 126.4 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.5 (1C, CHarom), 132.4 (1C, Cqarom), 135.4 (1C, Cqarom). IR: v [cm−1] = 2920 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C27H41NO2H [M + H] 412.3210, found 412.3201. Purity (HPLC): 96.1%, tR = 22.02 min.
4-Fluorobenzaldehyde (30 µL, 0.28 mmol) and NaBH(OAc)3 (119 mg, 0.56 mmol) were added to a solution of the primary amine 3a (62 mg, 0.28 mmol) in CH2Cl2 (8.5 mL). The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 20 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 2 cm, h = 20 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 5
NH3 = 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95
95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.46 (methanol
0.1, V = 10 mL, Rf = 0.46 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 50 mg (55%). C20H22FNO2 (327.4). 1H NMR (CDCl3): δ [ppm] = 1.46 (d, J = 13.2 Hz, 1H, 4-Heq), 1.84 (t, J = 8.0 Hz, 2H, CH2CH2N), 1.84 (s broad, 1H, NH), 2.13–2.22 (m, 1H, 4-Hax), 2.71 (d, J = 17.5 Hz, 1H, 6-H), 2.80 (t, J = 7.4 Hz, 2H, CH2N), 3.09 (d, J = 17.5 Hz, 1H, 6-H), 3.57–3.66 (m, 2H, 3-Hax and 3-Heq), 3.75 (s, 2H, CH2Ph), 5.90 (s, 1H, 1-H), 6.96 (t, J = 8.8 Hz, 2H, CHarom), 7.09 (d, J = 7.5 Hz, 1H, CHarom), 7.14 (dd, J = 7.5/1.2 Hz, 1H, CHarom), 7.20 (t, J = 7.0 Hz, 1H, CHarom), 7.24–7.28 (m, 3H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.7 (1C, C-4), 36.7 (1C, C-6), 43.8 (2C, CH2CH2N), 53.6 (1C, CH2Ph), 57.1 (1C, C-3), 71.4 (1C, C-5), 94.4 (1C, C-1), 115.3 (1C, CHarom), 115.5 (1C, CHarom), 126.6 (1C, CHarom), 126.7 (1C, CHarom), 127.4 (1C, CHarom), 128.7 (1C, CHarom), 129.8 (1C, CHarom), 129.9 (1C, CHarom), 132.3 (1C, Cqarom), 135.2 (1C, Cqarom), 160.9 (1C, Cqarom), 163.2 (1C, Cqarom). IR: v [cm−1] = 3333 (b, ν, N–H), 2932 (s, ν, C–H, alkyl), 1601 (s, ν, C
80)). Colorless oil, yield 50 mg (55%). C20H22FNO2 (327.4). 1H NMR (CDCl3): δ [ppm] = 1.46 (d, J = 13.2 Hz, 1H, 4-Heq), 1.84 (t, J = 8.0 Hz, 2H, CH2CH2N), 1.84 (s broad, 1H, NH), 2.13–2.22 (m, 1H, 4-Hax), 2.71 (d, J = 17.5 Hz, 1H, 6-H), 2.80 (t, J = 7.4 Hz, 2H, CH2N), 3.09 (d, J = 17.5 Hz, 1H, 6-H), 3.57–3.66 (m, 2H, 3-Hax and 3-Heq), 3.75 (s, 2H, CH2Ph), 5.90 (s, 1H, 1-H), 6.96 (t, J = 8.8 Hz, 2H, CHarom), 7.09 (d, J = 7.5 Hz, 1H, CHarom), 7.14 (dd, J = 7.5/1.2 Hz, 1H, CHarom), 7.20 (t, J = 7.0 Hz, 1H, CHarom), 7.24–7.28 (m, 3H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.7 (1C, C-4), 36.7 (1C, C-6), 43.8 (2C, CH2CH2N), 53.6 (1C, CH2Ph), 57.1 (1C, C-3), 71.4 (1C, C-5), 94.4 (1C, C-1), 115.3 (1C, CHarom), 115.5 (1C, CHarom), 126.6 (1C, CHarom), 126.7 (1C, CHarom), 127.4 (1C, CHarom), 128.7 (1C, CHarom), 129.8 (1C, CHarom), 129.9 (1C, CHarom), 132.3 (1C, Cqarom), 135.2 (1C, Cqarom), 160.9 (1C, Cqarom), 163.2 (1C, Cqarom). IR: v [cm−1] = 3333 (b, ν, N–H), 2932 (s, ν, C–H, alkyl), 1601 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, aryl), 1099 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C20H22FNO2H [M + H] 328.1707, found 328.1731. Purity (HPLC): 99.3%, tR = 16.25 min.
C, aryl), 1099 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C20H22FNO2H [M + H] 328.1707, found 328.1731. Purity (HPLC): 99.3%, tR = 16.25 min.
3-Phenylpropionaldehyde (37 µL, 0.28 mmol) and NaBH(OAc)3 (119 mg, 0.56 mmol) were added to a solution of the primary amine 3a (62 mg, 0.28 mmol) in CH2Cl2 (8.5 mL). The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 20 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 2 cm, h = 20 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 5
NH3 = 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95
95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1–>10
0.1–>10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90
90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.19 (methanol
0.1, V = 10 mL, Rf = 0.19 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 44 mg (46%). C22H27NO2 (337.5). 1H NMR (CDCl3): δ [ppm] = 1.50 (d, J = 13.4 Hz, 1H, 4-Heq), 2.02–2.10 (m, 4H, CH2CH2N and CH2CH2Ph), 2.20–2.28 (m, 1H, 4-Hax), 2.74 (t, J = 8.0 Hz, 2H, CH2CH2CH2Ph), 2.78 (d, J = 17.7 Hz, 1H, 6-H), 2.88 (t, J = 7.2 Hz, 2H, NCH2CH2CH2Ph), 3.03 (t, J = 7.5 Hz, 2H, CH2N), 3.17 (d, J = 17.5 Hz, 1H, 6-H), 3.64–3.70 (m, 2H, 3-Hax and 3-Heq), 4.64 (s broad, 1H, NH), 5.97 (s, 1H, 1-H), 7.16–7.37 (m, 9H, CHarom). 13C NMR (CDCl3): δ [ppm] = 29.9 (1C, CH2CH2Ph), 33.3 (1C, CH2Ph), 36.4 (1C, C-4), 36.5 (1C, C-6), 41.5 (1C, CH2CH2N), 43.8 (1C, CH2N), 48.8 (1C, CH2N), 56.9 (1C, C-3), 71.0 (1C, C-5), 94.3 (1C, C-1), 126.2 (1C, CHarom), 126.6 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.5 (2C, CHarom), 128.6 (2C, CHarom), 128.7 (1C, CHarom), 131.9 (1C, Cqarom), 134.8 (1C, Cqarom), 141.2 (1C, Cqarom). IR: v [cm−1] = 3329 (s, ν, N–H), 2932 (s, ν, C–H, alkyl), 1605 (s, ν, C
80)). Colorless oil, yield 44 mg (46%). C22H27NO2 (337.5). 1H NMR (CDCl3): δ [ppm] = 1.50 (d, J = 13.4 Hz, 1H, 4-Heq), 2.02–2.10 (m, 4H, CH2CH2N and CH2CH2Ph), 2.20–2.28 (m, 1H, 4-Hax), 2.74 (t, J = 8.0 Hz, 2H, CH2CH2CH2Ph), 2.78 (d, J = 17.7 Hz, 1H, 6-H), 2.88 (t, J = 7.2 Hz, 2H, NCH2CH2CH2Ph), 3.03 (t, J = 7.5 Hz, 2H, CH2N), 3.17 (d, J = 17.5 Hz, 1H, 6-H), 3.64–3.70 (m, 2H, 3-Hax and 3-Heq), 4.64 (s broad, 1H, NH), 5.97 (s, 1H, 1-H), 7.16–7.37 (m, 9H, CHarom). 13C NMR (CDCl3): δ [ppm] = 29.9 (1C, CH2CH2Ph), 33.3 (1C, CH2Ph), 36.4 (1C, C-4), 36.5 (1C, C-6), 41.5 (1C, CH2CH2N), 43.8 (1C, CH2N), 48.8 (1C, CH2N), 56.9 (1C, C-3), 71.0 (1C, C-5), 94.3 (1C, C-1), 126.2 (1C, CHarom), 126.6 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.5 (2C, CHarom), 128.6 (2C, CHarom), 128.7 (1C, CHarom), 131.9 (1C, Cqarom), 134.8 (1C, Cqarom), 141.2 (1C, Cqarom). IR: v [cm−1] = 3329 (s, ν, N–H), 2932 (s, ν, C–H, alkyl), 1605 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, aryl), 1099 (s, δ, C–O), 737 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C22H27NO2H [M + H] 338.2115, found 338.2127; calcd for C22H27NO2 − CH2CH2CH2Ph [M − CH2CH2CH2Ph] 218.1181, found 218.1178. Purity (HPLC): 98.2%, tR = 17.69 min.
C, aryl), 1099 (s, δ, C–O), 737 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C22H27NO2H [M + H] 338.2115, found 338.2127; calcd for C22H27NO2 − CH2CH2CH2Ph [M − CH2CH2CH2Ph] 218.1181, found 218.1178. Purity (HPLC): 98.2%, tR = 17.69 min.
Formalin solution (37%, 114 µL, 4.10 mmol) and NaBH(OAc)3 (130 mg, 0.62 mmol) were added to a solution of the benzylamine 3b (63 mg, 0.21 mmol) in CH2Cl2 (7 mL). The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 20 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 1.5 cm, h = 15 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 5
NH3 = 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95
95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.62 (methanol
0.1, V = 10 mL, Rf = 0.62 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 3
ethyl acetate = 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 7)). Colorless oil, yield 64 mg (97%). C21H25NO2 (323.2). 1H NMR (CDCl3): δ [ppm] = 1.52 (d, J = 13.3 Hz, 1H, 4-Heq), 1.89 (dd, J = 8.9/6.6 Hz, 2H, CH2CH2N), 2.13–2.19 (m, 1H, 4-Hax), 2.23 (s, 3H, CH3), 2.58 (td, J = 7.0/3.0 Hz, 2H, CH2N), 2.73 (d, J = 17.5 Hz, 1H, 6-H), 3.10 (d, J = 17.5 Hz, 1H, 6-H), 3.52 (s, 2H, CH2Ph), 3.57–3.65 (m, 2H, 3-Hax and 3-Heq), 5.93 (s, 1H, 1-H), 7.10 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (d, J = 7.4 Hz, 1H, CHarom), 7.20–7.36 (m, 7H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.7 (1C, C-6), 41.6 (1C, CH2CH2N), 42.4 (1C, NCH3), 51.3 (1C, CH2N), 57.1 (1C, C-3), 62.5 (1C, CH2Ph), 71.2 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.1 (1C, CHarom), 127.3 (1C, CHarom), 128.4 (2C, CHarom), 128.5 (1C, CHarom), 129.2 (2C, CHarom), 132.3 (1C, Cqarom), 135.3 (1C, Cqarom), 139.1 (1C, Cqarom). IR: v [cm−1] = 3024 (s, ν, C–H, aryl), 2951 (s, ν, C–H, alkyl), 1604 (s, ν, C
7)). Colorless oil, yield 64 mg (97%). C21H25NO2 (323.2). 1H NMR (CDCl3): δ [ppm] = 1.52 (d, J = 13.3 Hz, 1H, 4-Heq), 1.89 (dd, J = 8.9/6.6 Hz, 2H, CH2CH2N), 2.13–2.19 (m, 1H, 4-Hax), 2.23 (s, 3H, CH3), 2.58 (td, J = 7.0/3.0 Hz, 2H, CH2N), 2.73 (d, J = 17.5 Hz, 1H, 6-H), 3.10 (d, J = 17.5 Hz, 1H, 6-H), 3.52 (s, 2H, CH2Ph), 3.57–3.65 (m, 2H, 3-Hax and 3-Heq), 5.93 (s, 1H, 1-H), 7.10 (d, J = 7.4 Hz, 1H, CHarom), 7.17 (d, J = 7.4 Hz, 1H, CHarom), 7.20–7.36 (m, 7H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-4), 36.7 (1C, C-6), 41.6 (1C, CH2CH2N), 42.4 (1C, NCH3), 51.3 (1C, CH2N), 57.1 (1C, C-3), 62.5 (1C, CH2Ph), 71.2 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.1 (1C, CHarom), 127.3 (1C, CHarom), 128.4 (2C, CHarom), 128.5 (1C, CHarom), 129.2 (2C, CHarom), 132.3 (1C, Cqarom), 135.3 (1C, Cqarom), 139.1 (1C, Cqarom). IR: v [cm−1] = 3024 (s, ν, C–H, aryl), 2951 (s, ν, C–H, alkyl), 1604 (s, ν, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, aryl), 1134 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C21H25NO2H [M + H] 324.1958, found 324.1938; calcd for C21H25NO2 − Bn [M − Bn] 232.1338, found 323.1306; calcd for C21H25NO2 − CH2CH2N(CH3)(Bn) [M − CH2CH2N(CH3)(Bn)] 175.0759, found 175.0753. Purity (HPLC): 99.5%, tR = 16.50 min.
C, aryl), 1134 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C21H25NO2H [M + H] 324.1958, found 324.1938; calcd for C21H25NO2 − Bn [M − Bn] 232.1338, found 323.1306; calcd for C21H25NO2 − CH2CH2N(CH3)(Bn) [M − CH2CH2N(CH3)(Bn)] 175.0759, found 175.0753. Purity (HPLC): 99.5%, tR = 16.50 min.
Formalin solution (37%, 52 µL, 1.88 mmol) and NaBH(OAc)3 (60 mg, 0.28 mmol) were added to a solution of the cyclohexylmethylamine 3d (30 mg, 0.09 mmol) in CH2Cl2 (3 mL). The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with chloroform (4 × 20 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 1.5 cm, h = 17 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 5
NH3 = 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95
95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.25 (methanol
0.1, V = 10 mL, Rf = 0.25 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 27 mg (87%). C21H31NO2 (329.2). 1H NMR (CDCl3): δ [ppm] = 0.86–0.90 (m, 2H, CH(CH2)5), 1.10–1.25 (m, 3H, CH(CH2)5), 1.45–1.50 (m, 1H, CH(CH2)5), 1.54 (d, J = 8.0 Hz, 1H, 4-Heq), 1.64–1.72 (m, 3H, CH(CH2)5), 1.75–1.79 (m, 2H, CH(CH2)5), 1.82–1.86 (m, 2H, CH2CH2N), 2.14–2.22 (m, 3H, CH2CH(CH2)5 and 4-Hax), 2.25 (s, 3H, CH3), 2.52–2.55 (m, 2H, CH2N), 2.76 (d, J = 17.5 Hz, 1H, 6-H), 3.14 (d, J = 17.5 Hz, 1H, 6-H), 3.59–3.68 (m, 2H, 3-Hax and 3-Heq), 5.92 (s, 1H, 1-H), 7.12 (d, J = 7.4 Hz, 1H, CHarom), 7.16 (d, J = 7.4 Hz, 1H, CHarom), 7.22 (t, J = 7.2 Hz, 1H, CHarom), 7.27 (td, J = 7.3/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 26.2 (2C, CH(CH2)5), 26.9 (2C, CH(CH2)5), 32.0 (1C, CH(CH2)5), 35.8 (1C, CH(CH2)5), 36.6 (1C, C-6), 36.7 (1C, C-4), 41.0 (1C, CH2CH2N), 43.1 (1C, CH2N), 52.2 (1C, NCH3), 57.1 (1C, C-3), 65.1 (1C, CH2CH(CH2)5), 71.2 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.5 (1C, CHarom), 132.3 (1C, Cqarom), 135.3 (1C, Cqarom).
80)). Colorless oil, yield 27 mg (87%). C21H31NO2 (329.2). 1H NMR (CDCl3): δ [ppm] = 0.86–0.90 (m, 2H, CH(CH2)5), 1.10–1.25 (m, 3H, CH(CH2)5), 1.45–1.50 (m, 1H, CH(CH2)5), 1.54 (d, J = 8.0 Hz, 1H, 4-Heq), 1.64–1.72 (m, 3H, CH(CH2)5), 1.75–1.79 (m, 2H, CH(CH2)5), 1.82–1.86 (m, 2H, CH2CH2N), 2.14–2.22 (m, 3H, CH2CH(CH2)5 and 4-Hax), 2.25 (s, 3H, CH3), 2.52–2.55 (m, 2H, CH2N), 2.76 (d, J = 17.5 Hz, 1H, 6-H), 3.14 (d, J = 17.5 Hz, 1H, 6-H), 3.59–3.68 (m, 2H, 3-Hax and 3-Heq), 5.92 (s, 1H, 1-H), 7.12 (d, J = 7.4 Hz, 1H, CHarom), 7.16 (d, J = 7.4 Hz, 1H, CHarom), 7.22 (t, J = 7.2 Hz, 1H, CHarom), 7.27 (td, J = 7.3/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 26.2 (2C, CH(CH2)5), 26.9 (2C, CH(CH2)5), 32.0 (1C, CH(CH2)5), 35.8 (1C, CH(CH2)5), 36.6 (1C, C-6), 36.7 (1C, C-4), 41.0 (1C, CH2CH2N), 43.1 (1C, CH2N), 52.2 (1C, NCH3), 57.1 (1C, C-3), 65.1 (1C, CH2CH(CH2)5), 71.2 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.5 (1C, CHarom), 132.3 (1C, Cqarom), 135.3 (1C, Cqarom).
IR: v [cm−1] = 2920 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C21H31NO2H [M + H] 330.2540, found 330.2504; calcd for C21H31NO2 − CH2Cy [M − CH2Cy] 232.1338, found 232.1391. Purity (HPLC): 99.2%, tR = 17.98 min.
NEt3 (100 µL, 145 mg, 0.72 mmol) was added to a suspension of 3a-HCl (62 mg, 0.24 mmol) in CH2Cl2 (7 mL) to obtain the free primary amine 3a. Then, formalin solution (37%, 133 µL, 4.83 mmol) and NaBH(OAc)3 (154 mg, 0.72 mmol) were added. The reaction mixture was stirred overnight at rt. Saturated NaHCO3 solution (10 mL) and brine (10 mL) were added and the aqueous layer was extracted with chloroform (4 × 20 mL). The combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 2 cm, h = 15.5 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 10
NH3 = 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90
90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, V = 10 mL, Rf = 0.13 (methanol
1, V = 10 mL, Rf = 0.13 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 2
ethyl acetate = 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8)). Yellow oil, yield 30 mg (50%). C15H21NO2 (247.3). 1H NMR (CDCl3): δ [ppm] = 1.53 (d, J = 8.0 Hz, 1H, 4-Heq), 1.81–1.85 (m, 2H, CH2CH2N), 2.14–2.22 (m, 1H, 4-Hax), 2.27 (m, 6H, N(CH3)2), 2.46 (t, J = 7.5 Hz, 2H, CH2N), 2.76 (d, J = 17.4 Hz, 1H, 6-H), 3.11 (d, J = 17.5 Hz, 1H, 6-H), 3.58–3.65 (m, 2H, 3-Hax and 3-Heq), 5.92 (s, 1H, 1-H), 7.11 (d, J = 7.4 Hz, 1H, CHarom), 7.16 (dd, J = 7.5/1.4 Hz, 1H, CHarom), 7.21 (t, J = 7.1 Hz, 1H, CHarom), 7.27 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-6), 36.6 (1C, C-4), 41.8 (1C, CH2CH2N), 45.6 (2C, N(CH3)2), 53.5 (1C, CH2N), 57.0 (1C, C-3), 71.0 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.6 (1C, CHarom), 132.3 (1C, Cqarom), 135.1 (1C, Cqarom). IR: v [cm−1] = 2947 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C15H21NO2 + H [M + H] 248.1645, found 248.1666. Purity (HPLC): 99.0%, tR = 12.09 min.
8)). Yellow oil, yield 30 mg (50%). C15H21NO2 (247.3). 1H NMR (CDCl3): δ [ppm] = 1.53 (d, J = 8.0 Hz, 1H, 4-Heq), 1.81–1.85 (m, 2H, CH2CH2N), 2.14–2.22 (m, 1H, 4-Hax), 2.27 (m, 6H, N(CH3)2), 2.46 (t, J = 7.5 Hz, 2H, CH2N), 2.76 (d, J = 17.4 Hz, 1H, 6-H), 3.11 (d, J = 17.5 Hz, 1H, 6-H), 3.58–3.65 (m, 2H, 3-Hax and 3-Heq), 5.92 (s, 1H, 1-H), 7.11 (d, J = 7.4 Hz, 1H, CHarom), 7.16 (dd, J = 7.5/1.4 Hz, 1H, CHarom), 7.21 (t, J = 7.1 Hz, 1H, CHarom), 7.27 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 36.6 (1C, C-6), 36.6 (1C, C-4), 41.8 (1C, CH2CH2N), 45.6 (2C, N(CH3)2), 53.5 (1C, CH2N), 57.0 (1C, C-3), 71.0 (1C, C-5), 94.4 (1C, C-1), 126.5 (1C, CHarom), 126.6 (1C, CHarom), 127.3 (1C, CHarom), 128.6 (1C, CHarom), 132.3 (1C, Cqarom), 135.1 (1C, Cqarom). IR: v [cm−1] = 2947 (s, ν, C–H, alkyl), 1103 (s, δ, C–O), 764 (s, δ, 1,2-disubst. aryl). MS (APCI): calcd for C15H21NO2 + H [M + H] 248.1645, found 248.1666. Purity (HPLC): 99.0%, tR = 12.09 min.
Butyraldehyde (33 µL, 0.36 mmol) and NaBH(OAc)3 (154 mg, 0.73 mmol) were added to a solution of the primary amine 3a (80 mg, 0.36 mmol) in CH2Cl2 (11 mL). The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with CHCl3 (4 × 10 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 2 cm, h = 17 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 10
NH3 = 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90
90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.43 (methanol
0.1, V = 10 mL, Rf = 0.43 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 8 mg (7%). C21H33NO2 (331.5). 1H NMR (CDCl3): δ [ppm] = 0.94 (t, J = 7.3 Hz, 6H, NCH2CH2CH2CH3), 1.29–1.39 (m, 4H, NCH2CH2CH2CH3), 1.53–1.60 (m, 5H, NCH2CH2CH2CH3, 4-Heq), 1.94–1.97 (m, 2H, CH2CH2N), 2.13–2.22 (m, 1H, 4-Hax), 2.59–2.74 (m, 4H, NCH2CH2CH2CH3), 2.79 (d, J = 17.7 Hz, 1H, 6-H), 2.81–2.94 (m, 2H, CH2N), 3.13 (d, J = 17.7 Hz, 1H, 6-H), 3.63–3.66 (m, 2H, 3-Hax and 3-Heq), 5.91 (s, 1H, 1-H), 7.13 (d, J = 7.5 Hz, 1H, CHarom), 7.17 (dd, J = 7.4/1.1 Hz, 1H, CHarom), 7.23 (t, J = 7.2 Hz, 1H, CHarom), 7.29 (td, J = 7.5/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 14.0 (2C, NCH2CH2CH2CH3), 20.7 (2C, NCH2CH2CH2CH3), 27.6 (2C, NCH2CH2CH2CH3), 36.4 (1C, C-6), 36.7 (1C, C-4), 40.0 (1C, CH2CH2N), 47.8 (1C, CH2N), 53.3 (2C, NCH2CH2CH2CH3), 57.0 (1C, C-3), 70.9 (1C, C-5), 94.4 (1C, C-1), 126.60 (1C, CHarom), 126.63 (1C, CHarom), 127.4 (1C, CHarom), 128.7 (1C, CHarom), 132.1 (1C, Cqarom), 134.9 (1C, Cqarom). IR: v [cm−1] = 2955 (s, C–H), 1609 (s, C–Carom), 1103 (s, C–O), 764 (s, 1,2-disubst. benzene). MS (APCI): calcd for C21H33NO2H [M + H] 332.2584, found 332.2598; calcd for C21H33NO2 − butyl [M − butyl] 274.1807, found 274.1806. Purity (HPLC): 96.3%, tR = 18.18 min.
80)). Colorless oil, yield 8 mg (7%). C21H33NO2 (331.5). 1H NMR (CDCl3): δ [ppm] = 0.94 (t, J = 7.3 Hz, 6H, NCH2CH2CH2CH3), 1.29–1.39 (m, 4H, NCH2CH2CH2CH3), 1.53–1.60 (m, 5H, NCH2CH2CH2CH3, 4-Heq), 1.94–1.97 (m, 2H, CH2CH2N), 2.13–2.22 (m, 1H, 4-Hax), 2.59–2.74 (m, 4H, NCH2CH2CH2CH3), 2.79 (d, J = 17.7 Hz, 1H, 6-H), 2.81–2.94 (m, 2H, CH2N), 3.13 (d, J = 17.7 Hz, 1H, 6-H), 3.63–3.66 (m, 2H, 3-Hax and 3-Heq), 5.91 (s, 1H, 1-H), 7.13 (d, J = 7.5 Hz, 1H, CHarom), 7.17 (dd, J = 7.4/1.1 Hz, 1H, CHarom), 7.23 (t, J = 7.2 Hz, 1H, CHarom), 7.29 (td, J = 7.5/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 14.0 (2C, NCH2CH2CH2CH3), 20.7 (2C, NCH2CH2CH2CH3), 27.6 (2C, NCH2CH2CH2CH3), 36.4 (1C, C-6), 36.7 (1C, C-4), 40.0 (1C, CH2CH2N), 47.8 (1C, CH2N), 53.3 (2C, NCH2CH2CH2CH3), 57.0 (1C, C-3), 70.9 (1C, C-5), 94.4 (1C, C-1), 126.60 (1C, CHarom), 126.63 (1C, CHarom), 127.4 (1C, CHarom), 128.7 (1C, CHarom), 132.1 (1C, Cqarom), 134.9 (1C, Cqarom). IR: v [cm−1] = 2955 (s, C–H), 1609 (s, C–Carom), 1103 (s, C–O), 764 (s, 1,2-disubst. benzene). MS (APCI): calcd for C21H33NO2H [M + H] 332.2584, found 332.2598; calcd for C21H33NO2 − butyl [M − butyl] 274.1807, found 274.1806. Purity (HPLC): 96.3%, tR = 18.18 min.
NEt3 (90 µL, 0.65 mmol) was added to a solution of 3a-HCl (83 mg, 0.33 mmol) in CH2Cl2 (10 mL) to obtain the free primary amine 3a. Then, glutaraldehyde (31 µL, 0.33 mmol) and NaBH(OAc)3 (138 mg, 0.65 mmol) were added. The reaction mixture was stirred for 18 h at rt. After addition of a saturated NaHCO3 solution (10 mL) and brine (10 mL), the aqueous layer was extracted with CHCl3 (4 × 10 mL). Finally, the combined organic layers were dried (Na2SO4) and the solvent was removed in vacuo. The residue was purified by fc (∅ = 2 cm, h = 18 cm, methanol
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2
CH2Cl2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH3 = 10
NH3 = 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90
90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1, V = 10 mL, Rf = 0.17 (methanol
0.1, V = 10 mL, Rf = 0.17 (methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate = 20
ethyl acetate = 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 80)). Colorless oil, yield 30 mg (32%). C18H25NO2 (287.2). 1H NMR (CDCl3): δ [ppm] = 1.52–1.56 (m, 3H, 4-Heq, 4-CH2 pip), 1.77–1.85 (m, 4H, 3-CH2, 5-CH2 pip), 2.05–2.21 (m, 3H, 4-Hax, CH2CH2N), 2.62–2.86 (m, 6H, CH2N, 2-CH2, 6-CH2 pip), 2.79 (d, J = 17.8 Hz, 1H, 6-H), 3.13 (d, J = 17.8 Hz, 1H, 6-H), 3.62–3.65 (m, 2H, 3-Hax and 3-Heq), 5.89 (s, 1H, 1-H), 7.11–7.16 (m, 2H, CHarom), 7.21 (t, J = 7.1 Hz, 1H, CHarom), 7.28 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 23.4 (1C, C-4 pip), 24.4 (2C, C-3, C-5 pip), 36.3 (1C, C-6), 36.7 (1C, C-4), 38.7 (1C, CH2CH2N), 52.8 (1C, CH2N), 54.3 (2C, C-2, C-6 pip), 56.9 (1C, C-3), 70.7 (1C, C-5), 94.4 (1C, C-1), 126.6 (2C, CHarom), 127.4 (1C, CHarom), 128.7 (1C, CHarom), 132.0 (1C, Cqarom), 134.8 (1C, Cqarom). IR: v [cm−1] = 2932 (s, C–H), 1099 (s, C–O), 764 (s, 1,2-disubst. benzene). MS (APCI): calcd for C18H25NO2H [M + H] 288.1958, found 288.1981. Purity (HPLC): 97.4%, tR = 13.89 min.
80)). Colorless oil, yield 30 mg (32%). C18H25NO2 (287.2). 1H NMR (CDCl3): δ [ppm] = 1.52–1.56 (m, 3H, 4-Heq, 4-CH2 pip), 1.77–1.85 (m, 4H, 3-CH2, 5-CH2 pip), 2.05–2.21 (m, 3H, 4-Hax, CH2CH2N), 2.62–2.86 (m, 6H, CH2N, 2-CH2, 6-CH2 pip), 2.79 (d, J = 17.8 Hz, 1H, 6-H), 3.13 (d, J = 17.8 Hz, 1H, 6-H), 3.62–3.65 (m, 2H, 3-Hax and 3-Heq), 5.89 (s, 1H, 1-H), 7.11–7.16 (m, 2H, CHarom), 7.21 (t, J = 7.1 Hz, 1H, CHarom), 7.28 (td, J = 7.4/1.6 Hz, 1H, CHarom). 13C NMR (CDCl3): δ [ppm] = 23.4 (1C, C-4 pip), 24.4 (2C, C-3, C-5 pip), 36.3 (1C, C-6), 36.7 (1C, C-4), 38.7 (1C, CH2CH2N), 52.8 (1C, CH2N), 54.3 (2C, C-2, C-6 pip), 56.9 (1C, C-3), 70.7 (1C, C-5), 94.4 (1C, C-1), 126.6 (2C, CHarom), 127.4 (1C, CHarom), 128.7 (1C, CHarom), 132.0 (1C, Cqarom), 134.8 (1C, Cqarom). IR: v [cm−1] = 2932 (s, C–H), 1099 (s, C–O), 764 (s, 1,2-disubst. benzene). MS (APCI): calcd for C18H25NO2H [M + H] 288.1958, found 288.1981. Purity (HPLC): 97.4%, tR = 13.89 min.
4.2. Receptor binding studies
4.3. Determination of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) D7.4 values, plasma protein binding and metabolic stability in vitro
D7.4 values, plasma protein binding and metabolic stability in vitro
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 values were recorded using the micro shake flask method reported in ref. 26 and 27. Plasma protein binding was recorded by HPAC.30,31 Metabolic stability was determined by incubation with mouse liver microsomes.26,32 Details are given in the SI.
D7.4 values were recorded using the micro shake flask method reported in ref. 26 and 27. Plasma protein binding was recorded by HPAC.30,31 Metabolic stability was determined by incubation with mouse liver microsomes.26,32 Details are given in the SI.
Conflicts of interest
The authors declare no conflict of interest.Abbreviations
| DIAD | Diisopropyl azodicarboxylate |
| DIBAH | Diisobutylaluminum hydride |
| DMP | Dess-Martin-Periodinane |
| HPAC | High performance affinity chromatography |
| LLE | Ligand-lipophilicity efficiency |
| LC-MS | Liquid chromatography combined with mass spectrometry |
| MOPS | 3-Morpholinopropanesulfonic acid |
| NADPH | Nicotinamide adenine dinucleotide phosphate |
| NMDA | N-Methyl-D-aspartate |
| NMR | Nuclear magnetic resonance |
| PCP | 1-(1-Phenylcyclohexyl)piperidine (phencyclidine) |
| SEM | Standard error of the mean |
| TBS | tert-Butyldimethylsilyl |
| THF | Tetrahydrofuran |
Data availability
All data will be made available on request to the corresponding author of this manuscript.Supplementary information (SI): general synthetic methods, the method to determine the purity of the compounds, details of the receptor binding studies including the assays for σ1 and σ2 receptor affinity as well as the assay to determine the affinity towards the PCP binding site of the NMDA receptor. Experimental details to determine log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D7.4 values, plasma protein binding and metabolic stability are given. Moreover, 1H and 13C NMR spectra as well as HPLC traces for all test compounds. See DOI: https://doi.org/10.1039/d6ob00129g.
D7.4 values, plasma protein binding and metabolic stability are given. Moreover, 1H and 13C NMR spectra as well as HPLC traces for all test compounds. See DOI: https://doi.org/10.1039/d6ob00129g.
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (DFG), which is gratefully acknowledged.References
- R. Dingledine, K. Borges, D. Bowie and S. F. Traynelis, The Glutamate Receptor Ion Channels, Pharmacol. Rev., 1999, 51(1), 7–61, DOI:10.1016/S0031-6997(24)01394-2.
- S. F. Traynelis, L. P. Wollmuth, C. J. McBain, F. S. Menniti, K. M. Vance, K. K. Ogden, K. B. Hansen, H. Yuan, S. J. Myers and R. Dingledine, Glutamate Receptor Ion Channels: Structure, Regulation, and Function, Pharmacol. Rev., 2010, 62(3), 405–496, DOI:10.1124/pr.109.002451.
- H. Stark, U. Reichert and S. Graßmann, Structure, function and potential therapeutic significance of NMDA receptors. Part 2: Therapeutic concepts and new receptor ligands, Pharm. Unserer Zeit, 2000, 29, 228–236 Search PubMed.
- E. Karakas and H. Furukawa, Crystal structure of heterotetrameric NMDA receptor ion channel, Science, 2014, 344, 992–997 CrossRef CAS PubMed.
- E. Karakas, N. Simorowski and H. Furukawa, Subunit arrangement and phenylethanolamine binding in GluN1/GluN2B NMDA receptors, Nature, 2011, 475, 249–253, DOI:10.1038/nature10180.
- C. G. Rousseaux and S. Greene, Sigma receptors [sRs] : biology in normal and diseased states, J. Recept. Signal Transduction, 2016, 36, 327–388 CAS.
- T. Maurice and T. P. Su, The pharmacology of sigma-1 receptors, Pharmacol. Ther., 2009, 124(2), 195–206 CrossRef CAS PubMed.
- S. Collina, R. Gaggeri, A. Marra, A. Bassi, S. Negrinotti, F. Negri and D. Rossi, Sigma receptor modulators: a patent review, Expert Opin. Ther. Pat., 2013, 23, 597–613 CrossRef CAS PubMed.
- F. Langa, X. Codony, V. Tovar, A. Lavado, E. Giménez, P. Cozar, M. Cantero, A. Dordal, E. Hernández, R. Pérez, X. Monroy, D. Zamanillo, X. Guitart and L. Montoliu, Generation and phenotypic analysis of sigma receptor type I (σ1) knockout mice, Eur. J. Neurosci., 2003, 18, 2188–2196 CrossRef PubMed.
- V. Sabino, P. Cottone, S. L. Parylak, L. Steardo and E. P. Zorrilla, Sigma-1 receptor knockout mice display a depressive-like phenotype, Behav. Brain Res., 2009, 198, 472–476 CrossRef CAS PubMed.
- H. R. Schmidt, S. D. Zheng, E. Gurpinar, A. Koehl, A. Manglik and A. C. Kruse, Crystal structure of the human sigma 1 receptor, Nature, 2016, 532(7600), 527–530 CrossRef CAS PubMed.
- H. R. Schmidt, R. M. Betz, R. O. Dror and A. C. Kruse, Structural basis for sigma(1) receptor ligand recognition, Nat. Struct. Mol. Biol., 2018, 25(10), 981–987 CrossRef CAS PubMed.
- W. R. Martin, C. G. Eades, J. A. Thompson, R. E. Huppler and P. E. Gilbert, The effects of morphine- and nalorphine- like drugs in the nondependent and morphine-dependent chronic spinal dog, J. Pharmacol. Exp. Ther., 1976, 197, 517–532 CrossRef CAS PubMed.
- S. B. Hellewell and W. D. Bowen, A sigma-like binding site in rat pheochromocytoma (PC12) cells: decreased affinity for (+)-benzomorphans and lower molecular weight suggest a different sigma receptor form from that of guinea pig brain, Brain Res., 1990, 527, 244–253 CrossRef CAS PubMed.
- S. B. Hellewell, A. Bruce, G. Feinstein, J. Orringer, W. Williams and W. D. Bowen, Rat liver and kidney contain high densities of sigma 1 and sigma 2 receptors: characterization by ligand binding and photoaffinity labeling, Eur. J. Pharmacol., 1994, 268, 9–18 CrossRef CAS PubMed.
- F. I. Carroll, P. Abraham, K. Parham, X. Bai, X. Zhang, G. A. Brine, S. W. Mascarella, B. R. Martin, E. L. May, C. Sauss, L. Di Paolo, P. Wallace, J. M. Walker and W. D. Bowen, Enantiomeric N-substituted N-normetazocines: a comparative study of affinities at sigma, PCP, and mu opioid receptors, J. Med. Chem., 1992, 35, 2812–2818 CrossRef CAS PubMed.
- J. Köhler, K. Bergander, J. Fabian, D. Schepmann and B. Wünsch, Enantiomerically pure 1,3-dioxanes as highly selective NMDA and σ1 receptor ligands, J. Med. Chem., 2012, 55, 8953–8957 CrossRef PubMed.
- T. Utech, J. Köhler, H. Buschmann, J. Holenz, J. M. Vela and B. Wünsch, Synthesis and pharmacological evaluation of a potent and selective σ1 receptor antagonist with high antiallodynic activity, Arch. Pharm. Chem. Life Sci., 2011, 344, 415–421 CrossRef CAS PubMed.
- E. Quick, D. Schepmann and B. Wünsch, Synthesis and structure-affinity relationships of σ receptor ligands with 1,3-dioxane structure, Pharmaceuticals, 2025, 18, 1300, DOI:10.3390/ph18091300.
- R. J. Boeckman and K. M. George, 1,1,1-Triacetoxy-1,1-dihydro-1,2-benziodoxol-3(1 H)-one, in Encyclopedia of Reagents for Organic Synthesis, John Wiley & Sons, Ltd, Chichester, UK, 2009, p. 15. DOI:10.1002/047084289x.rt157m.pub2.
- E. Winterfeldt, Synthesis, 1975, 617–630 CrossRef CAS.
- A. F. Abdel-Magid and S. J. A. Mehrman, A Review on the Use of Sodium Tiacetoxyborohydride in the Reductive Amination of Ketones and Aldehydes, Org. Process Res. Dev., 2006, 10, 971–1031 CrossRef CAS.
- S. Gawaskar, D. Schepmann, A. Bonifazi, D. Robaa, W. Sippl and B. Wünsch, Benzo[7]annulene-based GluN2B selective NMDA receptor antagonists: Surprising effect of a nitro group in 2-position, Bioorg. Med. Chem. Lett., 2015, 25(24), 5748–5751 CrossRef CAS PubMed.
- N. Kopp, G. Civenni, D. Marson, E. Laurini, S. Pricl, C. V. Catapano, H.-U. Humpf, C. Almansa, F. R. Nieto, D. Schepmann and B. Wünsch, Chemoenzymatic synthesis of 2,6-disubsttuted tetrahydropyrans with high σ1 receptor affinity, antitumor and analgesic activity, Eur. J. Med. Chem., 2021, 219, 113443 CrossRef CAS PubMed.
- K. Miyata, D. Schepmann and B. Wünsch, Synthesis and σ receptor affinity of regioisomeric spirocyclic furopyridines, Eur. J. Med. Chem., 2014, 83, 709–716 CrossRef CAS PubMed.
- F. Börgel, F. Galla, K. Lehmkuhl, D. Schepmann, S. M. Ametamey and B. Wünsch, Pharmacokinetic properties of enantiomerically pure GluN2B selective NMDA receptor antagonists with 3-benzazepine scaffold, J. Pharm. Biomed. Anal., 2019, 172, 214–222 CrossRef PubMed.
- F. Galla, C. Bourgeois, K. Lehmkuhl, D. Schepmann, M. Soeberdt, T. Lotts, C. Abels, S. Ständer and B. Wünsch, Effects of polar κ receptor agonists designed for the periphery on ATP-induced Ca2+ release from keratinocytes, MedChemComm, 2016, 7, 317–326, 10.1039/C5MD00414D.
- P. D. Leeson and B. Springthorpe, The influence of drug-like concepts on decision-making in medicinal chemistry, Nat. Rev. Drug Discovery, 2007, 6(11), 881–890, DOI:10.1038/nrd2445.
- A. Tarcsay, K. Nyíri and G. M. Keseru, Impact of lipophilic efficiency on compound quality, J. Med. Chem., 2012, 55, 1252–1260 CrossRef CAS PubMed.
- V. Butsch, F. Börgel, F. Galla, K. Schwegmann, S. Hermann, M. Schäfers, B. Riemann, B. Wünsch and S. Wagner, Design, (Radio)Synthesis, and in Vitro and in Vivo evaluation of highly selective and potent matrix metalloproteinase 12 (MMP-12) inhibitors as radiotracers for positron emission tomography, J. Med. Chem., 2018, 61, 4115–4134, DOI:10.1021/acs.jmedchem.8b00200.
- C. P. Konken, K. Heßling, I. Thale, S. Schelhaas, J. Dabel, S. Maskri, E. Bulk, T. Budde, O. Koch, A. Schwab, M. Schäfers and B. Wünsch, Imaging of the calcium activated potassium channel 3.1 (KCa3.1) in vivo using a senicapoc-derived PET tracer, Arch. Pharm., 2022, e2200388, DOI:10.1002/ardp.202200388.
- C. Wiese, E. Große Maestrup, F. Galla, D. Schepmann, A. Hiller, S. Fischer, F.-A. Ludwig, W. Deuther-Conrad, C. K. Donat, P. Brust, L. Büter, U. Karst and B. Wünsch, . Comparison of in silico, electrochemical, in vitro and in vivo metabolism of a homologous series of (radio)fluorinated σ1 receptor ligands designed for positron emission tomography, ChemMedChem, 2016, 11, 2445–2458, DOI:10.1002/cmdc.201600366.
- S. Brune, D. Schepmann, K. H. Klempnauer, D. Marson, V. Dal Col, E. Laurini, M. Fermeglia, B. Wünsch and S. Pricl, The sigma enigma: in vitro/in silico site-directed mutagenesis studies unveil σ1 receptor ligand binding, Biochemistry, 2014, 53(18), 2993–3003 CrossRef CAS PubMed.
- E. Laurini, V. Dal Col, M. G. Mamolo, D. Zampieri, P. Posocco, M. Fermeglia, L. Vio and S. Pricl, Homology model and docking-based virtual screening for ligands of the σ1 receptor, ACS Med. Chem. Lett., 2011, 2(11), 834–839 CrossRef CAS PubMed.
- E. Rack, R. Fröhlich, D. Schepmann and B. Wünsch, Design, synthesis and pharmacological evaluation of spirocyclic σ1 receptor ligands with excocyclic amino moiety (increased distance 1), Bioorg. Med. Chem., 2011, 19, 3141–3151 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |