Open Access Article
Open Access ArticlePeptides as multifunctional linchpins in targeted drug conjugates
Xue
Li
a,
Yuechen
Qian
a,
Jiongjia
Cheng
 b and
Qian
Chu
b and
Qian
Chu
 *a
*a
aDepartment of Medicinal Chemistry, School of Pharmacy, China Pharmaceutical University, Nanjing, 211198, China. E-mail: qianchu@cpu.edu.cn
bSchool of Environmental Science, Nanjing Xiaozhuang University, Nanjing, 211171, China
First published on 9th March 2026
Abstract
Peptide–drug conjugates (PDCs) represent an emerging class of targeted therapeutics engineered to enhance drug specificity and minimize systemic toxicity, showing significant promise for treating complex diseases. The peptide component is central to this strategy, fulfilling multiple critical roles. Primarily, it acts as a high-affinity homing device, selectively directing the conjugate to receptors overexpressed on target cells to minimize off-target effects. Beyond targeting, peptides are also being innovatively engineered as enzymatically cleavable linkers to enable selective drug release within a specific pathological microenvironment. Despite this potential, the clinical translation of PDCs is hindered by challenges including instability in circulation, limited tissue penetration, and insufficient targeting selectivity. This review discusses strategic advances in peptide discovery, modification, and optimization to overcome these barriers. We further provide future perspectives on constructing next-generation PDCs, underscoring their potential as highly effective precision medicines.
1. Introduction
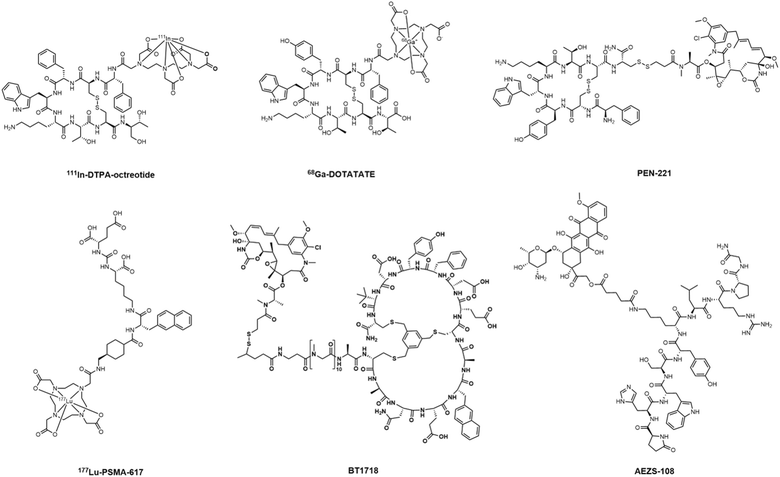
Peptide–drug conjugates have emerged as a promising targeted therapeutic platform, demonstrating considerable potential for treating complex diseases like cancer, drug-resistant infections, and autoimmune disorders.1–3 Over the past two decades, studies in this field have steadily increased, as evidenced by a consistent annual increase in PDC-related publications (Fig. 1). Notably, the clinical translation of PDCs is advancing rapidly, highlighted by recent FDA approvals and a pipeline of representative candidates in various stages of clinical development (Table 1 and Scheme 1).4–6| Name | Peptide | Receptor | Source of peptides | Peptide modification | Linker | Payload | Half-life | Indication | Status |
|---|---|---|---|---|---|---|---|---|---|
| a Sourced from https://pdcdb.idrblab.net, https://www.fda.gov, https://conjupepdb.ttk.hu, and https://clinicaltrials.gov. | |||||||||
| 111In-DTPA-octreotide11 | Octreotide | SSTR | Synthetic analog of natural hormone | 1. Cyclization | Amide | Indium-111 | 67.8 h | Neuroendocrine tumors | Approved (1994) |
| 2. D-Amino acid substitution | |||||||||
| 68Ga-DOTATATE12 | 3-Tyr-octreotate | SSTR | Synthetic analog of natural hormone | 1. Cyclization | Amide | Gallium-68 | 68 min | Neuroendocrine tumors | Approved (2016) |
| 2. D-Amino acid substitution | |||||||||
| 177Lu-DOTATATE13 | 3-Tyr-octreotate | SSTR | Synthetic analog of natural hormone | 1. Cyclization | Amide | Lutetium-177 | 71 h | Neuroendocrine tumors | Approved (2018) |
| 2. D-Amino acid substitution | |||||||||
| 64Cu-DOTATATE14 | 3-Tyr-octreotate | SSTR | Synthetic analog of natural hormone | 1. Cyclization | Amide | Copper-64 | 12.7 h | Neuroendocrine tumors | Approved (2020) |
| 2. Unnatural amino acid substitution | |||||||||
| 68Ga-PSMA-11 (ref. 15) | PSMA-11 | FOLH1 | Rational design | N/A | Aminocaproic acid | Gallium-68 | 68 min | Prostate cancer | Approved (2020) |
| 177Lu-PSMA-617 (ref. 16) | PSMA-617 | FOLH1 | Rational design | N/A | (S)-2-(4-(Aminomethyl)cyclohexane-1-carboxamido)-3-(naphthalen-2-yl)propanoic acid | Lutetium-177 | 41.6 h | Prostate cancer | Approved (2022) |
| ANG1005 (ref. 17) | Angiopep-2 | LRP1 | Synthetic analog of natural peptide | N/A | Succinate ester | Paclitaxel | 3.56 h | Recurrent brain metastases from breast cancer | Phase 3 |
| BT8009 (ref. 18) | Bicycle toxin conjugate precursor | Nectin-4 | Phage display | 1. Bicyclization | Val-Cit-PABC | Monomethyl auristatin E | <1 h | Urothelial cancer; non-small cell lung cancer; breast cancer | Phase 2/3 |
| 2. Unnatural amino acid substitution | |||||||||
| NGR-hTNF19,20 | NGR-peptide (CNGRC) | CD13 | Phage display | Cyclization | Amide | hTNF | 1–2 h | Malignant pleural mesothelioma | Phase 3 |
| BT1718 (ref. 21) | BT1718 bicycle peptide | MMP14 | Phage display | Bicyclization | Disulfide | Maytansine | 0.3 h | Solid tumors | Phase 2 |
| EP-100 (ref. 22) | LHRH | LHRH-R | Natural product | N/A | Amide | CLIP-71 | 7.1–15.9 min | Ovarian cancer | Phase 2 |
| CBX-12 (ref. 23) | Unique variant of pHLIP | Low pH and TOP1 | Synthetic analog of natural peptide | N/A | Disulfide | Exatecan | 14–22 h | Platinum resistant or refractory ovarian cancer | Phase 2 |
| PEN-221 (ref. 24) | [Tyr3, Cys8] octreotate amide | SSTR | Synthetic analog of natural hormone | 1. Cyclization | Disulfide | DM1 | 2.33 h | Advanced neuroendocrine, small cell lung cancer | Phase 1/2 |
| 2. D-Amino acid substitution | |||||||||
| AEZS-108 (ref. 25 and 26) | [D-Lys6]-LH-RH | LHRH-R | Synthetic analog of natural peptide | D-Amino acid substitution | Glutaric acid | Doxorubicin | 2 h | Prostate cancer | Phase 1/2 |
| BMTP-11 (ref. 27 and 28) | IL-11Rα-binding peptide (CGRRAGGSC) | IL-11Rα | Phage display | Cyclization | Amide | Apoptosis-inducing peptidomimetic D-(KLAKLAK)2 | N/A | Prostate cancer with bone metastasis | Phase 1 |
| TH1902 (ref. 29) | TH19P01 | SORT1 | N/A | Unnatural amino acid substitution | Succinate ester | Docetaxel | 1.44 h | Ovarian and endometrial cancer | Phase 1 |
| BT5528 (ref. 30) | Bicyclic peptide | EphA2 | Phage display | 1. Bicyclization | Val-Cit-PABC | Monomethyl auristatin E | 0.4–0.7 h | Solid tumors | Phase 1 |
| 2. Unnatural amino acid substitution | |||||||||
A typical PDC molecule comprises three core components: a targeting peptide, a bioactive payload, and a connecting linker (Fig. 1).4,7 The peptide acts as a homing device, directing the conjugate to specific receptors overexpressed on target cells to facilitate internalization. The payload is typically a cytotoxic drug, radionuclide, or imaging agent, responsible for the intended therapeutic or diagnostic activity. These components are covalently linked via a spacer, usually a biodegradable molecule strategically designed to maintain integrity in circulation while undergoing selective cleavage in target cells.8 Upon administration, the PDC circulates in the bloodstream until the peptide ligand binds to its cognate receptor. This binding event triggers receptor-mediated endocytosis, internalizing the entire conjugate. Inside the acidic and enzyme-rich environment of the endosome or lysosome, the linker is selectively cleaved, releasing the active payload directly inside the target cell.9,10 This targeted mechanism of action ensures high localized drug concentration at the disease site while minimizing systemic exposure and off-target toxicity to healthy tissues.
As an essential element in PDC molecules, the peptide plays multifaceted and critical roles. Its primary function is to act as a homing device, which is engineered to have high affinity and specificity for receptors overexpressed on target cells, such as cancer cells. Besides, peptides can also be designed to function as enzymatically cleavable linkers in PDCs, enabling highly specific drug release triggered by unique cellular environments. Compared to larger protein-based conjugates like antibody–drug conjugates (ADCs), the small molecular weight of peptides confers distinct advantages.31,32 It enables superior penetration into tumor tissue and more uniform distribution. Peptides are also typically less immunogenic, lowering the risk of adverse immune reactions. Pharmacokinetically, PDCs are cleared primarily via the renal system, which can reduce hepatic metabolic burden and potentially offer a more favorable safety profile.33 From a manufacturing standpoint, the established solid-phase peptide synthesis (SPPS) protocol allows for the cost-effective production of PDCs with high homogeneity and purity, facilitating their development.
Despite their promise, the therapeutic potential of PDCs is limited by several challenges, including poor circulatory stability, insufficient cellular permeability, and inadequate targeting potency and specificity.34 Fortunately, the inherent structural flexibility of peptides provides a versatile foundation for addressing these limitations. To enhance resistance to proteolytic degradation, strategies such as peptide cyclization and D-amino acid substitution are effectively employed.35 Furthermore, the rational incorporation of cell-penetrating peptides (CPPs) can significantly improve intracellular delivery and targeting precision.36,37 In addition, advances in peptide discovery technologies are pivotal for addressing challenging protein targets.38,39 Techniques like phage display enable the efficient screening of high-affinity ligands, while mRNA display offers access to exceptionally diverse libraries for novel homing peptide discovery. Most recently, artificial intelligence and computer-aided design are rapidly transforming the field. This shift is moving the peptide design paradigm toward the de novo generation of candidates with optimized stability, affinity, and pharmacokinetic properties.
This review highlights the essential role of peptides in advancing PDCs. Our focus extends beyond their well-characterized function as targeting ligands to systematically examine their involvement in facilitating targeted delivery, enhancing tissue and cellular penetration, and enabling controlled drug release via specialized linkers. By doing so, we aim to provide useful information for better construction of PDCs in future clinical applications.
2. Development of homing peptides
Homing peptides are the essential targeting elements of PDCs. These peptides are mostly derived from natural bioactive sequences, identified through high-throughput screening, or rationally engineered via computational methods (Fig. 2).2.1 Homing peptides obtained from bioactive sources
During the early stages of PDC development, homing peptides were primarily derived from bioactive sources.38,40 These naturally occurring peptides participate in various physiological and pathological processes, and inherently possess robust and selective cell-targeting properties, making them promising candidates for PDC applications.In addition to full-length bioactive peptides, truncated natural peptides also provide a versatile platform for PDC development by retaining the minimal active fragment required for receptor recognition. This strategy offers significant advantages, including lower synthesis costs, improved physicochemical properties, and enhanced suitability for further optimization.52 For example, SOR-C13 is a truncated peptide derived from soricidin that binds with high affinity to transient receptor potential of vanilloid type 6 (TRPV6) calcium channels.53,54 This peptide was engineered into a first-in-class, bispecific ligand drug conjugate, CBP-1008, which simultaneously targets TRPV6 and the folate receptor α (FRα) to deliver the cytotoxic payload MMAE. In clinical trials, CBP-1008 has shown encouraging efficacy and a favorable safety profile in ovarian cancer patients, underscoring the promise of truncated bioactive peptides in PDC therapeutics.55
2.2 Homing peptides discovered by screening technologies
While serendipitous discovery has successfully identified promising bioactive peptides for PDC development, it cannot meet the field's growing demands. As high-throughput screening and display technologies have revolutionized the discovery of small molecule drugs and biologic therapies, they are emerging as preferred strategies for identifying peptide ligands in PDC research.56A representative case is the DOX conjugate of E1-3, a heptapeptide identified from a phage display library for its selective binding to medulloblastoma cells. This peptide–doxorubicin conjugate exhibited potent antitumor cytotoxicity (IC50 = 25 nM) with a remarkable selectivity profile, showing 430-fold and 34-fold greater potency for tumor cells over non-tumor fibroblasts and primary brain astrocytes, respectively.60 In another case, Jiang et al. leveraged their phage-display-derived heptapeptide P7 to create a docetaxel conjugate (DTX-P7) for targeting non-small cell lung cancer (NSCLC). DTX-P7 exerts its effect by degrading the DYRK1A kinase to force quiescent cancer stem-like cells (CSLCs) to re-enter the cell cycle, thereby sensitizing them to death. This strategy demonstrated superior efficacy in vivo, where DTX-P7 reduced tumor growth by 93.2%, significantly surpassing the 35.9% reduction by free DTX.61,62 The phage display strategy also enables the identification of peptide ligands with complex constrained structures. A notable success is a bicyclic peptide that selectively targets membrane type I matrix metalloproteinase (MT1-MMP), a protein overexpressed in multiple tumors. Conjugating this peptide to the anti-tubulin agent DM1 via a cleavable disulfide linker produced BT1718, which showed significant anti-tumor efficacy in both MT1-MMP-dependent cellular assays and xenograft mouse models, and has progressed into clinical trials.63 Taken together, phage display identifies high-affinity, target-specific peptides that serve as the essential targeting component in PDCs, enabling precise cytotoxic drug delivery to tumor cells.
2.3 Homing peptides by rational design
Advances in structural biology, computational methods, and artificial intelligence have established rational design as an efficient strategy for peptide ligand discovery.67,68 Initially guided by empirical structure–activity relationships (SAR), this approach often uses a native substrate as a template, with SAR data guiding optimizations to enhance potency and selectivity. A successful example is the development of a neuropeptide Y (NPY) analogue. It was determined that an aromatic residue at position 7 of NPY enhanced Y1 receptor (Y1R) selectivity, while Pro34 was known to reduce affinity for the Y2 receptor. Incorporating these insights, [F7,P34]-NPY exhibited exceptional subnanomolar affinity for Y1R and a remarkable selectivity.69,70 Leveraging this high-affinity ligand, Wittrisch et al. constructed a PDC by linking this Y1R-preferring peptide to tesaglitazar, a PPARα/γ dual agonist, via a cleavable linker. The resulting conjugate (tesa-NPY) facilitates NPY1R-mediated internalization and targeted delivery to adipocytes, and improved glycemic control and lipid parameters in db/db mice, demonstrating a superior therapeutic potential.71Computer-aided drug design, such as molecular docking and dynamics simulations, has become a cornerstone of modern peptide therapeutic development.72,73 By providing atomic-level insights into peptide–protein interactions, this approach guides the rational optimization of binding affinity and selectivity. A representative application involved designing a selective matrix metallopeptidase 2 (MMP2) peptide inhibitor, which used SYBYL software to engineer hydrophobic contacts with the binding pocket. The isolated peptide exhibited robust MMP2 inhibition and high selectivity. Building on this design, a PDC was constructed by attaching a DOX derivative for targeted cytotoxicity and anti-metastasis.74
Artificial intelligence and machine learning are introducing a new paradigm for multi-property peptide engineering. Hsueh et al. utilized peptide microarrays to generate a large dataset on melanin binding, which was used to train a super learner model. Meanwhile, a complementary model was built to predict cell penetration and cytotoxicity from existing peptide databases. The integration of these models allowed for the rational design of peptides that simultaneously exhibit high melanin affinity, effective cell penetration, and low cytotoxicity.75 As proof-of-principle, the top-performing peptide (HR97) was conjugated to the drug brimonidine. A single injection of this conjugate in rabbits resulted in a 10-fold increase in melanin binding and a 17-fold greater reduction in intraocular pressure compared to the free drug.
3. Strategies for enhancing the targeting of PDCs
Precise targeting capability is a fundamental feature of an effective PDC, as cell-specific recognition is critical for minimizing off-target toxicity and expanding the therapeutic window. Emerging strategies aimed at enhancing the targeting specificity of PDCs have been developed, with bispecific targeting approaches and the integration of covalent warheads emerging as two particularly promising directions.3.1 Application of bispecific targeting
By simultaneously targeting two distinct tumor biomarkers, the bispecific strategy enhances therapeutic efficacy via synergistic effects. It not only reduces drug resistance caused by single-target downregulation but also improves tumor selectivity, thereby optimizing both therapeutic potency and precision.76 For example, peptides targeting a single human epidermal growth receptor 2 (HER2) domain often suffer from insufficient binding affinity, limiting their therapeutic efficacy. To overcome this, a novel bispecific fusion peptide was engineered to simultaneously target HER2 domains II and IV by integrating core fragments from pertuzumab and trastuzumab. The resulting bispecific peptide demonstrated a binding affinity nearly an order of magnitude higher than its single-domain counterpart. The subsequently constructed PDC showed superior cytotoxicity against HER2-positive cells compared to free chemotherapeutic drugs. Furthermore, in the SK-BR-3 tumor-bearing mouse model, the PDC achieved enhanced antitumor activity and significantly reduced off-target toxicity.773.2 Incorporation of covalent warheads
The incorporation of covalent warheads into PDCs is another emerging strategy to enhance therapeutic action by forming irreversible bonds with targets, thereby significantly extending drug-target residence time.78,79 The recent developed sulfur(VI) fluoride exchange (SuFEx) chemistry is a leading bioorthogonal strategy for this purpose. The sulfonyl fluoride group in SuFEx is highly stable under physiological conditions, yet becomes reactive only when in proximity to specific nucleophilic residues (e.g., tyrosine, histidine, lysine) on a target protein.80,81 This unique mechanism achieves a delicate balance between stability and reactivity, making it a powerful platform for developing selective covalent PDC molecules. For example, Wang et al. incorporated an unnatural amino acid with sulfonyl fluorine side chain into the mRNA display library, and identified CP-N1-N3, a macrocyclic peptide that irreversibly binds to the tumor-associated protein nectin-4 via the SuFEx reaction. By conjugating CP-N1-N3 to the microtubule inhibitor MMAE, they constructed the covalent PDC CP-N1-MMAE, which demonstrated potent cytotoxicity and induced cell death in nectin-4 overexpressed MDA-MB-468 cancer cells.82However, covalent PDCs rely on irreversible binding to target proteins through reactions between electrophilic warheads and nucleophilic residues. While this strategy can extend the duration of action and outcompete endogenous ligands, it also raises the risk of off-target protein modification, potentially leading to immune-mediated toxicity. Furthermore, prolonged target occupancy may disrupt normal protein turnover and result in accumulation in non-target tissues over time. Therefore, the design of covalent warheads in PDCs requires a delicate balance between reactivity and safety. Excessive electrophilicity increases the likelihood of non-specific reactions, causing off-target toxicity. Besides, overly active warheads can also react with abundant serum proteins, accelerating clearance and reducing tumor delivery. Conversely, insufficient reactivity compromises target engagement, undermining the rationale for covalent design. Beyond reactivity, the spatial arrangement of the warhead within the PDC scaffold can also significantly influence binding kinetics and biological outcomes. Suboptimal linker length, molecular orientation, or steric hindrance can trap the conjugate in non-productive binding modes, preventing successful covalent bond formation. These interdependent design parameters underscore the need for iterative optimization to achieve selective and durable target engagement while minimizing systemic toxicity.
4. Enzyme-cleavable peptide linkers for controlled release
In addition to their function as homing devices, peptides are uniquely suited to act as enzymatically cleavable linkers in PDCs. A major advantage of peptide linkers is their capacity to maintain stability in systemic circulation while allowing for selective drug release in the tumor microenvironment.8,83 This selectivity is achieved because protease activity is minimized in the blood by endogenous inhibitors and a neutral pH, whereas the tumor microenvironment is enriched with specific proteases such as cathepsin B and matrix metalloproteases (MMPs). Cleavage by these enzymes enables spatially controlled drug release, which enhances therapeutic index by maximizing efficacy at target site and minimizing systemic exposure (Fig. 3). | ||
| Fig. 3 Chemical structures of different enzyme-responsive peptide linkers. A) Cathepsin B-responsive linker, B) MMP-responsive linker, and C) PSA-responsive linker. | ||
Cathepsin B is a lysosomal cysteine protease that is significantly elevated in various cancer cells and participates in cancer progression, metastasis, and invasion.84 This characteristic makes it a proper tool for enzyme-responsive linker in PDC design.85 In cathepsin B-cleavable PDCs, the cleavable linker typically contains a specific cathepsin B cleavage site (such as the dipeptide sequences Val-Cit or Val-Ala) and a self-immolative spacer group (e.g., PABC, p-aminobenzyl carbamate). The core principle is to utilize the highly expressed cathepsin B protease within tumor cell to cleave the specific peptide sequence, which in turn triggers 1,6-elimination of PABC to release the active drug.9 Notably, this strategy enhances tumor killing efficacy while reducing toxicity to normal tissues. Schuster et al. developed GnRH-III–drug conjugates by linking the GnRH targeting peptide to PTX or Dau with cathepsin B-cleavable dipeptide linkers (Val-Ala or Val-Cit). These conjugates demonstrated significant antiproliferative activity against A2780 ovarian cancer cells, which express high levels of the GnRH receptor.86 Structure–activity relationship studies revealed that strategic amino acid substitutions can significantly modulate conjugate stability while preserving enzyme-cleavable properties. For instance, appending a glutamate residue at the N-terminus of a Val-Cit linker markedly enhances its stability in mice. Further substitution of valine with glycine results in even greater stability, as ADCs incorporating this optimized tripeptide linker demonstrated improved antitumor activity. Beyond Val-Cit and its analogs, the tetrapeptide Gly-Phe-Leu-Gly (GFLG) has also been widely used as cathepsin B-responsive sequence in PDCs. For example, in a PDC targeting the human Y1 receptor (hY1R) on breast cancer cells, methotrexate (MTX) was conjugated to the [F7,P34]-NPY peptide using either a stable amide bond or the cathepsin B-cleavable GFLG linker. As a result, the GFLG-linked conjugates outperformed their amide-bonded counterparts in both drug accumulation and cytotoxicity, highlighting the critical role of a cleavable linker for targeted delivery.87
MMPs are another class of enzyme stimulator for triggering cytotoxic drug release in targeted therapies. They are a family of zinc-dependent endopeptidases that degrade extracellular matrix components, which are overexpressed in the tumor microenvironment and play a key role in cancer progression.88 Several peptide substrates for MMP-2 and MMP-9, such as GPLGIAG, PLGLAG, and PVGLIG, have been widely used as cleavable linkers in PDCs.89,90 For instance, one PDC conjugated PTX to a CPP SynB3 via the MMP-2-sensitive PVGLIG linker. This design enabled targeted PTX release in glioblastoma, significantly suppressing tumor cell proliferation, migration, and invasion, whereas the non-cleavable control failed to do.91 Similarly, to mitigate DOX cardiotoxicity in HER2-positive breast cancer, a PDC was developed linking DOX to a HER2-targeting peptide via an MMP-2-cleavable GPLGLAGDD linker. In vitro, MMP-2 cleaved the linker with 80.8% efficiency within two hours. This cleavage was inhibited by MMP-2-specific inhibitors, confirming the enzyme-targeted release mechanism.92
Prostate-specific antigen (PSA) is a serine protease enzyme produced predominantly by the epithelial cells of the prostate gland. It is actively secreted by prostate tumors into the immediate microenvironment so that the concentration of active PSA is much higher in the vicinity of prostate cancer cells than in the general circulation.93,94 This difference has been leveraged to design PSA-cleavable linkers (e.g., SSKYQSL) for PDCs targeting prostate cancer. For example, conjugating a phosphoinositide 3-kinases (PI3K) inhibitor (TGX-D1) to a targeting KYL peptide via a PSA-cleavable linker produced KYL-TGX. This conjugate demonstrated robust toxicity in prostate cancer cells. Notably, the conjugate exhibited enhanced targeting and significantly greater tumor accumulation in mouse xenograft models compared to the free drug.95
A comparative analysis of these peptide-based cleavable linkers in PDCs reveals distinct mechanistic profiles that influence payload release kinetics, tumor selectivity, and therapeutic index (Table 2). The Val-Cit/Val-Ala linker exhibits high plasma stability due to its dipeptide structure, while its self-immolative PABC spacer facilitates rapid drug release upon enzymatic cleavage. However, its reliance on cathepsin B, which shows variable expression across tumors, can lead to heterogeneous payload release. The GFLG tetrapeptide serves as an alternative cathepsin B substrate with distinct cleavage kinetics. The extended spacer length may enhance enzymatic accessibility, albeit at the cost of increased molecular weight and synthetic complexity. In addition, GFLG exhibits slower cleavage rates than the Val-Cit linker, potentially enabling more sustained intracellular release. MMP-cleavable linkers leverage on tumor microenvironment, as MMP-2 and MMP-9 are secreted extracellularly by tumor cells, enabling pericellular payload release. However, MMP substrate promiscuity often raises off-target cleavage concerns, and the physiological roles of MMPs in normal tissues may lead to non-tumor activation.
| Type of enzyme | Representative peptides | Cleavage site | Key features |
|---|---|---|---|
| Cathepsin B | Val-Cit | Tumor endosomes | • Clinically validated |
| Val-Ala | • Heterogeneous payload release | ||
| • Strong hydrophobicity leads to easy aggregation | |||
| GFLG | Tumor endosomes | • Enhanced enzyme accessibility | |
| • Sustained intracellular release | |||
| • Increased molecular weight and synthetic complexity | |||
| MMPs | GPLGIAG | Tumor extracellular microenvironment | • High specificity for MMP-2/9 |
| PLGLAG | • High off-target risk in non-tumor tissues | ||
| PVGLIG | |||
| PSA | SSKYQSL | Prostate tumor extracellular microenvironment | • PSA enzyme selectivity |
| • Prone to being affected by PSA expression levels | |||
| • Restrict in prostate cancer treatment | |||
PSA-cleavable linkers offer exceptional selectivity, given PSA's predominate prostate expression. But the tumor heterogeneity in PSA levels also constrains its application on prostate cancer indications.
While these protease-cleavable linkers have shown great promise in PDC development, several critical challenges hinder their clinical translation.96 A fundamental issue is achieving the delicate balance between stability in circulation and efficient cleavage at the target site, as premature drug release can result in off-target toxicity and reduced efficacy.97 Moreover, the heterogeneous expression of target enzymes across different tumor types and cellular environments leads to inconsistent cleavage efficiency, causing unpredictable drug release and variable pharmacokinetic outcomes. To address these limitations, developing peptide linkers responsive to multiple stimuli, such as pH or redox changes, in addition to enzymatic cleavage, may offer a pathway toward more efficient and reliable drug release profiles.
5. Strategies for enhancing the stability of PDCs
PDCs represent a promising therapeutic class, yet their clinical advancement has been significantly limited by inherent stability issues. A key challenge is their short plasma half-life, which stems from rapid proteolytic degradation and renal clearance. This pharmacokinetic instability not only curtails therapeutic efficacy but also elevates the risk of premature payload release and subsequent systemic toxicity.98,99 To overcome these barriers, numerous strategies have been pursued to improve the pharmacological properties of PDCs (Fig. 4 and Table 3).| Strategy | Modification | Mechanism of stabilization | Representative PDCs |
|---|---|---|---|
| Cyclization | Monocyclization | • Stabilizes bioactive conformation | 99mTc-3PRGD2, BGC0222, BT8009, BT5528 |
| Bicyclization | • Sterically hinders exopeptidase access | ||
| D-Amino acids substitution | Replacement with D-amino acids | • Stereochemical inversion resulting in proteolytic resistance | 111In-DTPA-octreotide |
| 64Cu-DOTATATE | |||
| PEGylation | Polymer conjugation | • Increase molecular weight | BGC0222 |
| OEGylation | • Delay renal clearance | ||
| • Forms steric barrier against proteolysis | |||
| Terminal blocking | N-term acetylation | • Blocking recognition by external peptidases | KYL-TGX |
| N-term methylation | |||
| C-term amidation | |||
| Albumin binding enhancement | Fatty acid/maleimide/iodophenyl conjugation | • Form stable complexes with HSA, enabling FcRn-mediated recycling and evasion of renal clearance | II-3, [177Lu]Lu-7, MPD3 |
| • Leverages the EPR effect to promote tumor accumulation | |||
| Nano-delivery systems | Self-assembled peptide nanoparticles | • Enhancing drug stability through physical encapsulation or covalent conjugation | BA-NFs, CRB-FFE-YSV |
5.1 Chemical modifications on peptides
Another representative study of cyclized PDCs involved the fusion of the RGD peptide with the pro-apoptotic KLA motif to form a heptapeptide (RGDKLAK). Cyclizing this peptide significantly improved its serum stability by 1.5-fold compared to its linear counterparts. Furthermore, when the cyclic RGDKLAK peptide was conjugated to paclitaxel (PTX) using a pH-sensitive succinic acid linker, the resulting compound demonstrated superior antitumor efficacy over free PTX, without inducing significant toxicity.106,107
Bicyclic peptides surpass monocyclic peptides by confining the peptide backbone within two independent rings, thereby exerting greater conformational constraint. This structural rigidity confers significant advantages, including enhanced proteolytic stability and improved tissue penetration.108,109 Leveraging these benefits, bicyclic peptides have recently emerged as promising scaffolds for PDCs. So far, several bicyclic PDCs are advancing through clinical trials. BT8009, for instance, is a conjugate of a nectin-4-binding bicyclic peptide and the cytotoxic payload MMAE, connected by a cleavable Val-Cit linker. Preclinical studies showed marked antitumor efficacy and a favorable tolerability profile, prompting a currently ongoing phase III trial in advanced or metastatic solid tumors.110,111 In addition, BT5528, an EphA2-targeting bicyclic PDC, is also under investigation in phase II trials for solid tumors.30,112
N-terminal acetylation is another established strategy to improve PDC's stability. By adding an acetyl group to block the peptide's terminal amine, this modification efficiently shields the peptide from recognition and cleavage by aminopeptidases.120,121 The efficacy of this approach was demonstrated by the development of KYL-TGX. Acetylation of the KYL peptide enhanced the conjugate's stability, ensuring that its structure and function remained intact for effective delivery to the tumor site.95
Complementing the aforementioned modifications, a range of additional approaches, such as the use of polysialic acid (PSA), hydroxyethyl starch (HES), N-methylation, and C-terminal amidation, have proven effective in mitigating the renal clearance of peptides.2 The expanding repertoire of chemical modifications provides essential strategies for enhancing PDC stability and unlocking their full therapeutic potential.
Peptide modifications are potent chemical strategies to enhance PDC metabolic stability, yet each strategy introduces distinct trade-offs that must be carefully balanced against target binding affinity. For example, conformational constraint from cyclization can distort the peptide's bioactive conformation, reducing receptor engagement. In addition, cyclization typically leads to a decrease in polar surface area and an increase in hydrophobicity, which may compromise aqueous solubility. Similarly, D-amino acid substitution introduces stereochemical inversion, which may disrupt peptide's native secondary structure and potentially abrogate critical binding interactions. From a synthetic and manufacturing perspective, chemically modified peptides are generally more challenging and expensive to produce than their wildtype counterparts. Therefore, the successful application of modified peptides in PDC construction requires empirical optimization to achieve the demands of both proteolytic stability and target engagement.
5.2 Enhancing albumin binding
Human serum albumin (HSA), with a circulation half-life of approximately 19 days, represents a promising carrier for enhancing the pharmacokinetics of PDC molecules. This prolonged half-life is attributed to its molecular weight (∼66.5 kDa), which surpasses the renal filtration threshold, and to FcRn recycling, which avoids lysosomal degradation. By incorporating an albumin-binding moiety, PDC molecules can form a stable complex with HSA, evading rapid renal clearance and gaining an extended plasma half-life. Furthermore, this strategy capitalizes on the natural biodistribution of albumin, leveraging the enhanced permeability and retention (EPR) to solid tumors to promote targeted drug accumulation.122–124 While the EPR effect is widely acknowledged as a valid biological mechanism that facilitates the passive accumulation of macromolecules, such as ADCs and nanoparticles, its effectiveness for PDCs is still being questioned.125 Due to their relatively small size, usually under 5 kDa, PDCs alone might not leverage the EPR effect as efficiently as larger macromolecules. However, binding of PDCs to serum albumin substantially increases their molecular size, potentially enabling their tumor accumulation through the EPR pathway. Currently, several albumin-binding moieties have been employed to enhance the stability of PDCs (Fig. 5). | ||
| Fig. 5 Schematic illustration of the albumin-binding strategy (top) and examples of typical albumin-binding groups (bottom). | ||
A common strategy to promote albumin binding is the conjugation of a fatty acid chain. For instance, Zhang and colleagues developed peptide mimetics targeting the receptor tyrosine kinase-like orphan receptor 1 (ROR1) and linked them to anticancer drugs like gemcitabine and DOX. To enhance the pharmacokinetics of PDC molecules, they incorporated a fatty acid chain to facilitate albumin binding. This approach yielded a leading candidate, II-3, which exhibited exceptional internalization efficiency, anticancer potency, and a dramatically improved half-life (increased from 5.6 minutes to 2.6 hours). Notably, this conjugation strategy did not impair the targeting capability, as II-3 maintained a high binding affinity (Kd = 1.72 × 10−9 M) and specificity for ROR1.126
In addition to fatty acids, a variety of albumin-binding moieties have been developed to stabilize PDCs and improve their pharmacokinetics. For example, Zha et al. created [177Lu]Lu-7 by conjugating a albumin binder (a pegylated p-iodophenylbutanoyl group) to a PSMA-targeting ligand. This modification enabled stable, noncovalent binding to albumin, which significantly prolonged its blood circulation half-life. As a result, [177Lu]Lu-7 demonstrated prolonged circulation, higher tumor uptake, and better retention than its unmodified counterpart.127 In another approach, Cho et al. designed the PDC MPD3 to target PTEN-loss cancers. MPD3 consists of KGDEVD peptide linked to docetaxel with a maleimide-based albumin-binding moiety (N-(ε-maleimidocaproyloxy)succinimide ester) conjugated to the side chain of the peptide. This dual strategy promoted intracellular drug delivery and resulted in a significantly prolonged half-life compared to the free drug.128
5.3 Forming nano-delivery systems
Self-assembled peptide nanoparticles (SAPNs) are a promising drug delivery platform due to their biocompatibility, ease of synthesis, and long half-life. They improve drug stability by either physically encapsulating therapeutics or serving as a covalent scaffold for PDCs, particularly effective for hydrophobic drugs.129,130Zeng and colleagues developed a promising peptide nano-delivery system, baicalin-peptide supramolecular self-assembled nanofibers (BA-NFs).131 This system was constructed by conjugating baicalin (BA) to the targeting peptide ARVYIHPF and co-assembling it with the self-assembling peptide. In an aqueous solution, these components form spiral nanofibers that significantly enhance the solubility and stability of BA (Fig. 6). By selectively targeting the angiotensin II type I receptor (AT1R), BA-NFs function as an efficient cardiac-targeted anti-ferroptosis agent. In a mouse model of doxorubicin-induced cardiomyopathy (DIC), BA-NFs demonstrated effective cardiac accumulation, improved cardiac function, reduced fibrosis, and good biocompatibility.
 | ||
| Fig. 6 The structure of self-assembled baicalin-peptide nanofibers (BA-NFs) and its application in drug delivery. | ||
In another case, Yang et al. developed a self-assembling, supramolecular hydrogel (designated as CRB-FFE-YSV) by covalently conjugating the drug chlorambucil (CRB) and the peptide drug tyroservatide (YSV) to the FFE peptide scaffold.132 This formulation self-assembled into a nanofibrous hydrogel that exhibited remarkable stability, resisting degradation by proteinase K and extending the drug's enzymatic half-life from less than one hour to over four hours. As a result, the construct achieved potent tumor suppression, reducing the tumor volume to approximately 38% of that observed in the PBS control group.
6. Strategies for enhancing the pharmacokinetics of PDCs
Pharmacokinetics, which describes a drug's absorption, distribution, metabolism, and excretion (ADME), is fundamental to therapeutic success. It bridges the critical gap between drug discovery and clinical application, directly determining efficacy and safety.133,134 Therefore, optimizing pharmacokinetics properties of PDCs is paramount to their successful development.6.1 Improving cell membrane penetration and blood–brain barrier transportation
The development of PDCs is hampered by poor cell penetration and an inability to cross the blood–brain barrier (BBB). The inherent physicochemical properties of PDCs, such as high molecular weight, hydrophilicity, and charge, hinder passive diffusion through lipid membranes. Crossing the BBB is even more challenging, as it requires active transport mechanisms to bypass its stringent, selective endothelial layer.135 Cell-penetrating peptides represent a promising technological avenue due to their unique ability to cross cellular membranes and, in some instances, the impermeable blood–brain barrier.136,137 By serving as delivery vectors, CPPs can shuttle therapeutic payloads into the brain for treatment of brain cancers, neurodegenerative diseases, and central nervous system (CNS) infections. Therefore, conjugating CPPs to PDCs offers a promising strategy to overcome these critical delivery barriers in PDC development.138To implement this strategy, Deng's team designed LTP-1, a PDC where PTX is conjugated to a bifunctional peptide. The peptide features luteinizing hormone-releasing hormone (LHRH) as a cancer-targeting ligand and the optimized CPP T2 for membrane penetration.31 This design conferred enhanced tumor targeting and cellular internalization. LTP-1 achieved twofold greater uptake in LHRH-overexpressing MCF-7 cells than free PTX. More significantly, LTP-1 retained potent activity against PTX-resistant A2780/PTX cells, where free PTX was ineffective.
Angiopep-2 (ANG) is a peptide ligand that binds to the LRP1 receptor on brain endothelial cells, enabling BBB crossing via receptor-mediated transcytosis. A novel PDC ANG1005 was constructed accordingly, which utilizes ANG to deliver three PTX molecules across the BBB and into malignant cells. In a phase II trial involving heavily pretreated patients with breast cancer and recurrent brain metastases, ANG1005 demonstrated promising intracranial disease control and prolonged overall survival. Currently, a phase III study of ANG1005 has been initiated to further confirm its therapeutic efficacy.17 In another study, Li et al. designed a dual-targeting PDC, ANG-TAT-PTX. This conjugate links PTX to both ANG for LRP-1 receptor-mediated BBB transcytosis and the HIV-1 TAT peptide for enhanced cellular uptake. In murine glioma models, the ANG-TAT platform achieved 1.8-fold higher tumor accumulation than ANG alone and 5.6-fold more than free PTX. Consequently, ANG-TAT-PTX treatment significantly prolonged survival in the glioblastoma mouse model, demonstrating the critical role CPPs in boosting tumor penetration and therapeutic efficacy.139
6.2 Improving ADME profiles by machine learning
Recently, artificial intelligence and other computer technology are emerging strategies to predict the absorption, distribution, metabolism, excretion, and toxicity (ADMET) properties.140,141 Several artificial intelligence platforms have become essential tools in drug design, enhancing the success rate of drug optimization. By employing the physiologically-based pharmacokinetic (PBPK) modeling software GastroPlus®, Vale et al. evaluate the pharmacokinetic parameters of four CPPs (P1–P4) and three antineoplastic agents for predicting in vitro synergy.142 The simulation revealed that the combination of P2–P4 peptides with 5-fluorouracil (5-FU) was predicted to exhibit the highest synergistic potential in HT-29 colorectal cancer cells, which was confirmed by subsequent cell-based studies. ADMETlab 3.0 is an online prediction platform designed for drug development. Its multi-task DMPNN modeling method and large-scale data enable efficient, accurate, and robust comprehensive ADMET profiling. The introduction of API module support to streamline the process of batch evaluation, and implementation of an uncertainty estimation module is vital for reliable selection of candidate compounds for subsequent experimental phases.143 In addition to the aforementioned tools, several other ADMET predictors have also been developed, with notable examples including OptADMET, pBRICS, SwissADME and FAF-Drugs.144–147So far, the ADME profiling of PDCs remains incompletely established, largely because of the complex interplay between the homing peptide, linker chemistry, and cytotoxic payload.148 Currently, artificial intelligence tools are trained primarily on small-molecule datasets, and their reliability in predicting the ADMET properties of structurally complex PDCs remains to be fully validated. A primary bottleneck is the lack of high-quality and unbiased experimental data. Public databases contain insufficient structural and activity information for peptides, particularly those containing non-canonical amino acids and cyclic peptides that are essential for enhancing PDC stability. Furthermore, existing publications and datasets present bias towards positive results, whereas the negative data are equally critical for training reliable models. Another considerable challenge with AI modeling involves the limited capability to predict complex biological systems. Accurate prediction of peptide binding, stability and functional effects solely from peptide sequence or molecule structure is not a trivial task. For example, peptide dynamics and conformational changes often occur under different physiological conditions, such as temperature, pH and cellular microenvironment, resulting in extremely challenging for comprehensive computational analysis.149 Nevertheless, the computational tools mentioned above show significant promise for PDC development, and addressing these limitations is essential to fully realize the translational potential of AI in PDC discovery.
7. Translational challenges of PDCs
While peptide–drug conjugates present significant promise for therapeutic applications, their clinical limitations remain a major concern. For instance, Pepaxto (melphalan flufenamide) had its accelerated approval withdrawn after a confirmatory trial failed to demonstrate clinical benefit or confirm its safety profile.150 Similarly, several other candidates have also been discontinued during clinical development due to inherent drawbacks. For example, AEZS-108 was found to undergo rapid hydrolysis due to cleavage of its ester bond by carboxylesterases, with half-lives of approximately 20 minutes in mouse serum and 100–120 minutes in human serum. This resulted in poor enzymatic stability in circulation, leading to its clinical failure.26 These setbacks highlight the pressing need to enhance critical features of PDCs, such as target specificity, bioactivity, stability, linker chemistry, and pharmacokinetics, which are extensively examined in this study.Another significant challenge in the clinical translation of PDCs involves their selectivity, stability, and potential off-target toxicity. Target selectivity remains a primary concern, since many receptors overexpressed on tumor cells, such as integrins and somatostatin receptors, are also present at physiological levels in healthy tissues. Consequently, homing peptides may not effectively distinguish between malignant cells and normal tissues, leading to PDC accumulation in non-malignant cells and unintended adverse effects. Moreover, unlike antibody–drug conjugates (ADCs), which typically achieve nanomolar or sub-nanomolar affinity, PDCs often exhibit only moderate target affinity in the micromolar range, further elevating the off-target risks.151 Stability presents another critical challenge in PDC development. Excessive plasma stability can impair intracellular drug activation, whereas insufficient stability leads to systemic toxicity and a shortened half-life. These stability issues directly contribute to off-target toxicity, as unstable PDCs may release their cytotoxic payload into the bloodstream, inadvertently affecting healthy proliferating cells. Nevertheless, ongoing innovations in peptide engineering, linker chemistry, and targeting strategies are steadily advancing the field towards next-generation PDCs with improved therapeutic profiles.
The clinical translation of peptide–drug conjugates is also profoundly influenced by manufacturing scalability and compliance with regulatory expectations. Although solid-phase peptide synthesis (SPPS) is the preferred method for peptide synthesis in research, it encounters considerable challenges when scaling up for industrial production.152 Sequence-dependent risks, such as hydrophobic stretches, can dramatically reduce coupling efficiency during large-scale synthesis, compromising yield and purity. Subsequent solution-phase conjugation of linker-payload moieties adds further complexity, demanding high conversion efficiency while rigorously minimizing payload-related impurities. Therefore, critical CMC (chemistry, manufacturing and controls) regulations are essential for PDC development, which includes purification strategy development, analytical method validation for impurity profiling, and immunogenicity assessment. Importantly, PDC manufacturing requires additional regulation, as linker stability and conjugation specificity introduce extra quality considerations. Thus, the success of PDC manufacturing depends on a coordinated strategy that aligns peptide synthesis, conjugation chemistry, and analytical development to satisfy both scalability and stringent regulatory standards.
8. Future perspectives
Looking forward, the future trajectory of PDCs is focused on evolving beyond simple drug-delivery systems into intelligent, next-generation therapeutics with enhanced precision, efficacy, and safety. As an essential element in PDC design, peptides hold immense promise in propelling this transformation.A primary area of innovation lies in the development of advanced homing peptides with highly efficient targeting and robust tissue penetration. Beyond the well-established phage-display and mRNA display techniques, several high-throughput screening technologies, such as DNA-encoded libraries (DELs) and the one bead one compound (OBOC) platform, have already proven effective in discovering novel peptide ligands, thereby expanding the toolbox for PDC development. Furthermore, the strategic incorporation of non-canonical amino acids not only increases peptide diversity but also enhances their drug-like properties, offering solutions to longstanding challenges such as in vivo stability, pharmacokinetics, and immunogenicity.
Currently, PDC payloads are predominantly limited to cytotoxic drugs, radionuclides, or imaging agents, nearly all of which are small molecules. Peptide drugs, however, represent a maturing yet still innovative therapeutic class, offering precise targeting across a growing range of diseases. Integrating peptide drugs as the therapeutic payload within PDCs could unlock the full potential of peptide-based targeted therapies, moving beyond conventional cytotoxic approaches toward more sophisticated and specific mechanisms of action.
Lastly, the clinical translation of PDCs will be expedited by advances in computational design and AI-driven discovery. Artificial intelligence is transforming PDC development from a trial-and-error process into a rational design approach. By simultaneously optimizing peptide targeting, linker stability, and payload delivery through predictive modeling, AI is expected to dramatically accelerate the development timeline and enhance the safety and efficacy of PDC candidates.
9. Conclusion
PDCs represent a maturing therapeutic modality, with expanding clinical trials positioned to validate their potential as a versatile and precise platform in next-generation targeted medicine. Crucially, the peptide component within PDCs is far more than a passive delivery vehicle. Instead, it is an active pharmaceutical ingredient, which directly governs tissue specificity, drives cellular internalization, steers intracellular trafficking, and is a primary determinant of therapeutic efficacy. Empowered by advances in structural biology, combinatorial chemistry, high-throughput screening, and artificial intelligence, modern peptide engineering has transformed PDCs into highly promising and rationally designed therapeutics.Author contributions
Xue Li: writing – original draft, conceptualization. Yuechen Qian: writing – original draft, data curation. Jiongjia Cheng: writing – review & editing, conceptualization. Qian Chu: writing – review & editing, supervision, conceptualization.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This work was supported by the National Natural Science Foundation of China (22277141, Q. Chu; 22207055, J. Cheng).References
- Y. Wang, A. G. Cheetham, G. Angacian, H. Su, L. Xie and H. Cui, Adv. Drug Delivery Rev., 2017, 110–111, 112–126 CrossRef CAS PubMed.
- S. F. A. Rizvi, H. Zhang and Q. Fang, Med. Res. Rev., 2024, 44, 2420–2471 CrossRef PubMed.
- Y. Feng, T. Li, S. Li, Z. Liu, Z. Tang, C. Chen, C. Zhou, T. Lu and J. Chen, Eur. J. Med. Chem., 2026, 304, 118482 CrossRef CAS PubMed.
- C. Fu, L. Yu, Y. Miao, X. Liu, Z. Yu and M. Wei, Acta Pharm. Sin. B, 2023, 13, 498–516 CrossRef CAS PubMed.
- M. Wang, J. Liu, M. Xia, L. Yin, L. Zhang, X. Liu and Y. Cheng, Eur. J. Med. Chem., 2024, 265, 116119 CrossRef CAS PubMed.
- H. Jin, P. Yang, H. Min, J. Song and Y. Qi, Cancer Lett., 2026, 638, 218174 CrossRef CAS PubMed.
- J. Ma, X. Wang, Y. Hu, J. Ma, Y. Ma, H. Chen and Z. Han, J. Med. Chem., 2025, 68, 9037–9056 CrossRef CAS PubMed.
- M. Alas, A. Saghaeidehkordi and K. Kaur, J. Med. Chem., 2021, 64, 216–232 CrossRef CAS PubMed.
- J. D. Bargh, A. Isidro-Llobet, J. S. Parker and D. R. Spring, Chem. Soc. Rev., 2019, 48, 4361–4374 RSC.
- E. Pérez-Herrero and A. Fernández-Medarde, Acta Pharm. Sin. B, 2021, 11, 2243–2264 CrossRef PubMed.
- G. Weckbecker, F. Raulf, B. Stolz and C. Bruns, Pharmacol. Ther., 1993, 60, 245–264 CrossRef CAS PubMed.
- A. R. Haug, R. Cindea-Drimus, C. J. Auernhammer, M. Reincke, B. Wangler, C. Uebleis, G. P. Schmidt, B. Goke, P. Bartenstein and M. Hacker, J. Nucl. Med., 2012, 53, 1686–1692 CrossRef CAS PubMed.
- J. Strosberg, G. El-Haddad, E. Wolin, A. Hendifar, J. Yao, B. Chasen, E. Mittra, P. L. Kunz, M. H. Kulke, H. Jacene, D. Bushnell, T. M. O'Dorisio, R. P. Baum, H. R. Kulkarni, M. Caplin, R. Lebtahi, T. Hobday, E. Delpassand, E. Van Cutsem, A. Benson, R. Srirajaskanthan, M. Pavel, J. Mora, J. Berlin, E. Grande, N. Reed, E. Seregni, K. Öberg, M. Lopera Sierra, P. Santoro, T. Thevenet, J. L. Erion, P. Ruszniewski, D. Kwekkeboom and E. Krenning, N. Engl. J. Med., 2017, 376, 125–135 CrossRef CAS PubMed.
- E. S. Delpassand, D. Ranganathan, N. Wagh, A. Shafie, A. Gaber, A. Abbasi, A. Kjaer, I. Tworowska and R. Nunez, J. Nucl. Med., 2020, 61, 890–896 CrossRef CAS PubMed.
- U. Hennrich and M. Eder, Pharmaceuticals, 2021, 14, 713 CrossRef CAS PubMed.
- O. Sartor, J. de Bono, K. N. Chi, K. Fizazi, K. Herrmann, K. Rahbar, S. T. Tagawa, L. T. Nordquist, N. Vaishampayan, G. El-Haddad, C. H. Park, T. M. Beer, A. Armour, W. J. Perez-Contreras, M. DeSilvio, E. Kpamegan, G. Gericke, R. A. Messmann, M. J. Morris, B. J. Krause and V. Investigators, N. Engl. J. Med., 2021, 385, 1091–1103 CrossRef CAS PubMed.
- P. Kumthekar, S. C. Tang, A. J. Brenner, S. Kesari, D. E. Piccioni, C. Anders, J. Carrillo, P. Chalasani, P. Kabos, S. Puhalla, K. Tkaczuk, A. A. Garcia, M. S. Ahluwalia, J. S. Wefel, N. Lakhani and N. Ibrahim, Clin. Cancer Res., 2020, 26, 2789–2799 CrossRef CAS PubMed.
- A. Cigliola, V. Tateo, M. Ravasi, G. Di Maria, S. Manzo, B. A. Maiorano, C. Mercinelli and A. Necchi, Expert Opin. Invest. Drugs, 2025, 34, 371–378 CrossRef CAS PubMed.
- B. Valentinis, S. Porcellini, C. Asperti, M. Cota, D. Zhou, P. Di Matteo, G. Garau, C. Zucchelli, N. R. Avanzi, G. P. Rizzardi, M. Degano, G. Musco and C. Traversari, Int. J. Mol. Sci., 2019, 20, 4511 CrossRef CAS PubMed.
- H. W. van Laarhoven, W. Fiedler, I. M. Desar, J. J. van Asten, S. Marreaud, D. Lacombe, A. S. Govaerts, J. Bogaerts, P. Lasch, J. N. Timmer-Bonte, A. Lambiase, C. Bordignon, C. J. Punt, A. Heerschap and C. M. van Herpen, Clin. Cancer Res., 2010, 16, 1315–1323 CrossRef CAS PubMed.
- H. Harrison, G. Bennett, D. Blakeley, A. Brown, S. Campbell, L. H. Chen, R. J. Lutz, S. Pavan, K. van Rietschoten, D. Teufel, P. U. Park and K. Lee, Cancer Res., 2017, 77(13_Supplement), 5144 CrossRef.
- K. K. Curtis, J. Sarantopoulos, D. W. Northfelt, G. J. Weiss, K. M. Barnhart, J. K. Whisnant, C. Leuschner, H. Alila, M. J. Borad and R. K. Ramanathan, Cancer Chemother. Pharmacol., 2014, 73, 931–941 CrossRef CAS PubMed.
- S. Gayle, T. Paradis, K. Jones, J. Vasquez and V. M. Paralkar, Immunotherapy, 2022, 14, 1467–1480 CrossRef CAS PubMed.
- B. H. White, K. Whalen, K. Kriksciukaite, R. Alargova, T. Au Yeung, P. Bazinet, A. Brockman, M. DuPont, H. Oller, C. A. Lemelin, P. Lim Soo, B. Moreau, S. Perino, J. M. Quinn, G. Sharma, R. Shinde, B. Sweryda-Krawiec, R. Wooster and M. T. Bilodeau, J. Med. Chem., 2019, 62, 2708–2719 CrossRef CAS PubMed.
- J. Engel, G. Emons, J. Pinski and A. V. Schally, Expert Opin. Invest. Drugs, 2012, 21, 891–899 CrossRef CAS PubMed.
- G. Emons, M. Kaufmann, G. Gorchev, V. Tsekova, C. Grundker, A. R. Gunthert, L. C. Hanker, M. Velikova, H. Sindermann, J. Engel and A. V. Schally, Gynecol. Oncol., 2010, 119, 457–461 CrossRef CAS PubMed.
- K. Karjalainen, D. E. Jaalouk, C. Bueso-Ramos, L. Bover, Y. Sun, A. Kuniyasu, W. H. Driessen, M. Cardo-Vila, C. Rietz, A. J. Zurita, S. O'Brien, H. M. Kantarjian, J. E. Cortes, G. A. Calin, E. Koivunen, W. Arap and R. Pasqualini, Clin. Cancer Res., 2015, 21, 3041–3051 CrossRef CAS PubMed.
- V. O. Lewis, E. Devarajan, M. Cardo-Vila, D. G. Thomas, E. S. Kleinerman, S. Marchio, R. L. Sidman, R. Pasqualini and W. Arap, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 8065–8070 CrossRef CAS PubMed.
- J. C. Currie, M. Demeule, C. Charfi, A. Zgheib, A. Larocque, B. A. Danalache, A. Ouanouki, R. Beliveau, C. Marsolais and B. Annabi, Cancers, 2022, 14, 1877 CrossRef CAS PubMed.
- G. Bennett, A. Brown, G. Mudd, P. Huxley, K. Van Rietschoten, S. Pavan, L. Chen, S. Watcham, J. Lahdenranta and N. Keen, Mol. Cancer Ther., 2020, 19, 1385–1394 CrossRef CAS PubMed.
- X. Deng, R. Mai, C. Zhang, D. Yu, Y. Ren, G. Li, B. Cheng, L. Li, Z. Yu and J. Chen, Eur. J. Med. Chem., 2021, 213, 113050 CrossRef CAS PubMed.
- Y. Yang, S. Wang, P. Ma, Y. Jiang, K. Cheng, Y. Yu, N. Jiang, H. Miao, Q. Tang, F. Liu, Y. Zha and N. Li, Cancer Lett., 2023, 552, 215969 CrossRef CAS PubMed.
- I. Vhora, S. Patil, P. Bhatt and A. Misra, Adv. Protein Chem. Struct. Biol., 2015, 98, 1–55 CAS.
- B. M. Cooper, J. Iegre, D. H. O’ Donovan, M. Ölwegård Halvarsson and D. R. Spring, Chem. Soc. Rev., 2021, 50, 1480–1494 RSC.
- L. Gong, H. Zhao, Y. Liu, H. Wu, C. Liu, S. Chang, L. Chen, M. Jin, Q. Wang, Z. Gao and W. Huang, Acta Pharm. Sin. B, 2023, 13, 3659–3677 CrossRef CAS PubMed.
- H. Derakhshankhah and S. Jafari, Biomed. Pharmacother., 2018, 108, 1090–1096 CrossRef CAS PubMed.
- A. A. Khakshur, E. Khodaverdi, H. Kamali and A. Nokhodchi, Pharm. Dev. Technol., 2025, 30, 521–547 CrossRef CAS PubMed.
- M. Muttenthaler, G. F. King, D. J. Adams and P. F. Alewood, Nat. Rev. Drug Discovery, 2021, 20, 309–325 CrossRef CAS PubMed.
- K. Sharma, K. K. Sharma, A. Sharma and R. Jain, Drug Discovery Today, 2023, 28, 103464 CrossRef CAS PubMed.
- S. You, G. McIntyre and T. Passioura, Expert Opin. Drug Discovery, 2024, 19, 961–973 CrossRef CAS PubMed.
- D. R. Nassel, Prog. Neurobiol., 2002, 68, 1–84 CrossRef PubMed.
- Z. Zhang, H. Han, J. Zhao, Z. Liu, L. Deng, L. Wu, J. Niu, Y. Guo, G. Wang, X. Gou, C. Li, C. Li and C. M. Liu, Mol. Hortic., 2025, 5, 7 CrossRef CAS PubMed.
- V. P. Martinovich and K. U. Baradzina, Russ. J. Bioorg. Chem., 2022, 48, 221–232 CrossRef CAS.
- C. P. Capper, J. M. Rae and R. J. Auchus, Horm. Cancer, 2016, 7, 149–164 CrossRef CAS PubMed.
- N. Sayyad, E. I. Vrettos, T. Karampelas, C. M. Chatzigiannis, K. Spyridaki, G. Liapakis, C. Tamvakopoulos and A. G. Tzakos, Eur. J. Med. Chem., 2019, 166, 256–266 CrossRef CAS PubMed.
- F. Schneider, W. Tomek and C. Gründker, Theriogenology, 2006, 66, 691–709 CrossRef CAS PubMed.
- I. Ranđelović, S. Schuster, B. Kapuvári, G. Fossati, C. Steinkühler, G. Mező and J. Tóvári, Int. J. Mol. Sci., 2019, 20, 4763 CrossRef PubMed.
- K. Fodor, N. Dobos, A. Schally, Z. Steiber, G. Olah, E. Sipos, L. Szekvolgyi and G. Halmos, Oncotarget, 2020, 11, 175–187 CrossRef PubMed.
- R. He, B. Finan, J. P. Mayer and R. D. DiMarchi, Molecules, 2019, 24, 1855 CrossRef CAS PubMed.
- A. N. Mamelak, S. Rosenfeld, R. Bucholz, A. Raubitschek, L. B. Nabors, J. B. Fiveash, S. Shen, M. B. Khazaeli, D. Colcher, A. Liu, M. Osman, B. Guthrie, S. Schade-Bijur, D. M. Hablitz, V. L. Alvarez and M. A. Gonda, J. Clin. Oncol., 2006, 24, 3644–3650 CrossRef CAS PubMed.
- S. McGonigle, U. Majumder, D. Kolber-Simonds, J. Wu, A. Hart, T. Noland, K. TenDyke, D. Custar, D. Li, H. Du, M. H. D. Postema, W. G. Lai, N. C. Twine, M. Woodall-Jappe and K. Nomoto, Cell Commun. Signaling, 2019, 17, 67 CrossRef PubMed.
- X. Wang, X. Hong, F. Chen and K. J. Wang, Front. Cell. Infect. Microbiol., 2022, 12, 928220 CrossRef CAS PubMed.
- C. V. Bowen, D. DeBay, H. S. Ewart, P. Gallant, S. Gormley, T. T. Ilenchuk, U. Iqbal, T. Lutes, M. Martina, G. Mealing, N. Merkley, S. Sperker, M. J. Moreno, C. Rice, R. T. Syvitski and J. M. Stewart, PLoS One, 2013, 8, e58866 CrossRef CAS PubMed.
- H. Xue, Y. Wang, T. J. MacCormack, T. Lutes, C. Rice, M. Davey, D. Dugourd, T. T. Ilenchuk and J. M. Stewart, J. Cancer, 2018, 9, 3196–3207 CrossRef PubMed.
- J. F. Gong, J. Zhang, N. Li, L. Shen, L. Y. Wu, Y. Z. Zhang, Y. Huang, Y. Wang, J. Y. Wu, G. Lou, Y. Q. Chen, S. X. Luo, Y. Ba, X. Wang, Y. L. Zheng, G. L. Li, Z. S. Tong, R. B. Huang, Y. Teng and X. C. Hu, J. Clin. Oncol., 2023, 41, 5577 CrossRef.
- A. M. Davis, A. T. Plowright and E. Valeur, Nat. Rev. Drug Discovery, 2017, 16, 681–698 CrossRef CAS PubMed.
- G. P. Smith, Science, 1985, 228, 1315–1317 CrossRef CAS PubMed.
- S. Jahandar-Lashaki, S. Farajnia, A. Faraji-Barhagh, Z. Hosseini, N. Bakhtiyari and L. Rahbarnia, Mol. Biotechnol., 2025, 67, 2161–2184 CrossRef CAS PubMed.
- P. E. Saw and E. W. Song, Protein Cell, 2019, 10, 787–807 CrossRef PubMed.
- K. C. Tjandra, N. McCarthy, L. Yang, A. J. Laos, G. Sharbeen, P. A. Phillips, H. Forgham, S. M. Sagnella, R. M. Whan, M. Kavallaris, P. Thordarson and J. A. McCarroll, J. Med. Chem., 2020, 63, 2181–2193 CrossRef CAS PubMed.
- Y. Jiang, N. Yang, H. Zhang, B. Sun, C. Hou, C. Ji, J. Zheng, Y. Liu and P. Zuo, J. Controlled Release, 2016, 221, 26–36 CrossRef CAS PubMed.
- Y. Jiang, W. Huang, X. Sun, X. Yang, Y. Wu, J. Shi, J. Zheng, S. Fan, J. Liu, J. Wang, Z. Liang, N. Yang, Z. Liu and Y. Liu, J. Hematol. Oncol., 2022, 15, 73 CrossRef CAS PubMed.
- G. Bennett, M. Rigby, B. Lutz, P. Park and N. Keen, Mol. Cancer Ther., 2018, 17, B135 CrossRef.
- Y. Fan, R. Feng, X. Zhang, Z. L. Wang, F. Xiong, S. Zhang, Z. F. Zhong, H. Yu, Q. W. Zhang, Z. Zhang, Y. Wang and G. Li, Acta Pharm. Sin. B, 2024, 14, 3362–3384 CrossRef CAS PubMed.
- G. Kamalinia, B. J. Grindel, T. T. Takahashi, S. W. Millward and R. W. Roberts, Chem. Soc. Rev., 2021, 50, 9055–9103 RSC.
- S. Li and R. W. Roberts, Chem. Biol., 2003, 10, 233–239 CrossRef CAS PubMed.
- B. Xu, Q. Dong, C. Yu, H. Chen, Y. Zhao, B. Zhang, P. Yu and M. Chen, Antioxidants, 2024, 13, 479 CrossRef CAS PubMed.
- P. Manzanares, M. Gandia, S. Garrigues and J. F. Marcos, Nutrients, 2019, 11, 2545 CrossRef CAS PubMed.
- R. M. Söll, M. C. Dinger, I. Lundell, D. Larhammer and A. G. Beck-Sickinger, Eur. J. Biochem., 2001, 268, 2828–2837 CrossRef PubMed.
- R. Kufka, R. Rennert, G. N. Kaluđerović, L. Weber, W. Richter and L. A. Wessjohann, Beilstein J. Org. Chem., 2019, 15, 96–105 CrossRef CAS.
- S. Wittrisch, N. Klöting, K. Mörl, R. Chakaroun, M. Blüher and A. G. Beck-Sickinger, Mol. Metab., 2020, 31, 163–180 CrossRef CAS PubMed.
- M. H. Cardoso, R. Q. Orozco, S. B. Rezende, G. Rodrigues, K. G. N. Oshiro, E. S. Candido and O. L. Franco, Front. Microbiol., 2019, 10, 3097 CrossRef PubMed.
- N. Lalani, S. Tivari, V. Jain and Y. Jadeja, Pept. Sci., 2024, 116, e24343 CrossRef CAS.
- C. Qian, J. Wang, Y. Qian, R. Hu, J. Zou, C. Zhu, Y. Zhu, S. Qi, X. Jia, L. Wu, W. Li and Z. Chen, Nano Lett., 2020, 20, 4153–4161 CrossRef CAS.
- H. T. Hsueh, R. T. Chou, U. Rai, W. Liyanage, Y. C. Kim, M. B. Appell, J. Pejavar, K. T. Leo, C. Davison, P. Kolodziejski, A. Mozzer, H. Kwon, M. Sista, N. M. Anders, A. Hemingway, S. V. K. Rompicharla, M. Edwards, I. Pitha, J. Hanes, M. P. Cummings and L. M. Ensign, Nat. Commun., 2023, 14, 2509 CrossRef CAS PubMed.
- Y. Yang, Y. Mou, L. X. Wan, S. Zhu, G. Wang, H. Gao and B. Liu, Med. Res. Rev., 2024, 44, 2600–2623 CrossRef CAS PubMed.
- J. Zhou, X. Bian, Z. Kan, Z. Cai, Y. Jiang, Z. Wang, Y. Li, W. Shi and H. Qian, J. Med. Chem., 2022, 65, 15123–15139 CrossRef CAS PubMed.
- N. V. Mehta and M. S. Degani, Drug Discovery Today, 2023, 28, 103799 CrossRef CAS PubMed.
- M. Gehringer and S. A. Laufer, J. Med. Chem., 2019, 62, 5673–5724 CrossRef CAS PubMed.
- N. Wang, B. Yang, C. Fu, H. Zhu, F. Zheng, T. Kobayashi, J. Liu, S. Li, C. Ma, P. G. Wang, Q. Wang and L. Wang, J. Am. Chem. Soc., 2018, 140, 4995–4999 CrossRef CAS.
- J. Dong, L. Krasnova, M. G. Finn and K. B. Sharpless, Angew. Chem., Int. Ed., 2014, 53, 9430–9448 CrossRef CAS PubMed.
- R. Wang, S. Ran, J. Guo, D. Hu, X. Feng, J. Zhou, Z. Zhang, F. Liang, J. Shang, L. Bu, K. Wang, J. Mao, H. Luo and R. Wang, Acta Pharm. Sin. B, 2025, 15, 5474–5485 CrossRef CAS PubMed.
- M. Fajar, R. Maharani and U. Supratman, Bioorg. Med. Chem., 2026, 133, 118510 CrossRef CAS PubMed.
- D. Musil, D. Zucic, D. Turk, R. A. Engh, I. Mayr, R. Huber, T. Popovic, V. Turk, T. Towatari and N. Katunuma, et al. , EMBO J., 1991, 10, 2321–2330 CrossRef CAS.
- E. I. Vrettos, G. Mező and A. G. Tzakos, Beilstein J. Org. Chem., 2018, 14, 930–954 CrossRef CAS PubMed.
- S. Schuster, É. Juhász, G. Halmos, I. Neundorf, C. Gennari and G. Mező, Int. J. Mol. Sci., 2022, 23, 5071 CrossRef CAS PubMed.
- D. Böhme, J. Krieghoff and A. G. Beck-Sickinger, J. Med. Chem., 2016, 59, 3409–3417 CrossRef.
- Z. A. Rashid and S. K. Bardaweel, Int. J. Mol. Sci., 2023, 24, 12133 CrossRef CAS PubMed.
- X. Zhang, X. Wang, W. Zhong, X. Ren, X. Sha and X. Fang, Int. J. Nanomed., 2016, 11, 1643–1661 CAS.
- L. Zhu, T. Wang, F. Perche, A. Taigind and V. P. Torchilin, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 17047–17052 CrossRef CAS PubMed.
- D. Hua, L. Tang, W. Wang, S. Tang, L. Yu, X. Zhou, Q. Wang, C. Sun, C. Shi, W. Luo, Z. Jiang, H. Li and S. Yu, Adv. Sci., 2021, 8, 2001960 CrossRef CAS PubMed.
- Y. You, Z. Xu and Y. Chen, Drug Delivery, 2018, 25, 448–460 CrossRef CAS PubMed.
- K. W. Watt, P. J. Lee, T. M'Timkulu, W. P. Chan and R. Loor, Proc. Natl. Acad. Sci. U. S. A., 1986, 83, 3166–3170 CrossRef CAS PubMed.
- S. R. Denmeade, L. J. Sokoll, D. W. Chan, S. R. Khan and J. T. Isaacs, Prostate, 2001, 48, 1–6 CrossRef CAS PubMed.
- A. Barve, A. Jain, H. Liu, W. Jin and K. Cheng, Nanomedicine, 2016, 12, 2373–2381 CrossRef CAS.
- Y. Lei, M. Zheng, P. Chen, C. Seng Ng, T. Peng Loh and H. Liu, ChemMedChem, 2025, 20, e202500262 CrossRef CAS PubMed.
- D. Su and D. Zhang, Front. Pharmacol., 2021, 12, 687926 CrossRef CAS PubMed.
- K. Jadhav, A. Abhang, E. B. Kole, D. Gadade, A. Dusane, A. Iyer, A. Sharma, S. K. Rout, A. D. Gholap, J. Naik, R. K. Verma and S. Rojekar, Bioengineering, 2025, 12, 481 CrossRef CAS PubMed.
- D. Wang, F. Yin, Z. Li, Y. Zhang and C. Shi, J. Nanobiotechnol., 2025, 23, 305 CrossRef PubMed.
- C. K. Wang, J. E. Swedberg, S. E. Northfield and D. J. Craik, J. Phys. Chem. B, 2015, 119, 15821–15830 CrossRef CAS.
- Q. N. Vu, R. Young, H. K. Sudhakar, T. Gao, T. Huang, Y. S. Tan and Y. H. Lau, RSC Med. Chem., 2021, 12, 887–901 RSC.
- S. J. Bogdanowich-Knipp, S. Chakrabarti, T. D. Williams, R. K. Dillman and T. J. Siahaan, J. Pept. Res., 1999, 53, 530–541 CrossRef CAS PubMed.
- S. J. Bogdanowich-Knipp, D. S. Jois and T. J. Siahaan, J. Pept. Res., 1999, 53, 523–529 CrossRef CAS.
- L. Xiao, S. Yu, W. Xu, Y. Sun and J. Xin, Korean J. Radiol., 2023, 24, 1142–1150 CrossRef PubMed.
- Y. Q. Huang, J. D. Yuan, H. F. Ding, Y. S. Song, G. Qian, J. L. Wang, M. Ji and Y. Zhang, Eur. J. Med. Chem., 2018, 158, 82–90 CrossRef CAS PubMed.
- S. F. A. Rizvi, N. Abbas, H. Zhang and Q. Fang, J. Med. Chem., 2023, 66, 8324–8337 CrossRef CAS PubMed.
- S. F. A. Rizvi, S. Mu, Y. Wang, S. Li and H. Zhang, Biomed. Pharmacother., 2020, 127, 110179 CrossRef CAS PubMed.
- D. Z. Feng, L. H. Liu, Y. Q. Shi, P. Du, S. T. Xu, Z. Y. Zhu, J. Y. Xu and H. Yao, Chin. Chem. Lett., 2023, 34, 108026 CrossRef CAS.
- S. Ullrich and C. Nitsche, Pept. Sci., 2024, 116, e24326 CrossRef CAS.
- M. Rigby, G. Bennett, L. Chen, G. E. Mudd, H. Harrison, P. J. Beswick, K. Van Rietschoten, S. M. Watcham, H. S. Scott, A. N. Brown, P. U. Park, C. Campbell, E. Haines, J. Lahdenranta, M. J. Skynner, P. Jeffrey, N. Keen and K. Lee, Mol. Cancer Ther., 2022, 21, 1747–1756 CrossRef CAS PubMed.
- G. E. Mudd, H. Scott, L. Chen, K. van Rietschoten, G. Ivanova-Berndt, K. Dzionek, A. Brown, S. Watcham, L. White, P. U. Park, P. Jeffrey, M. Rigby and P. Beswick, J. Med. Chem., 2022, 65, 14337–14347 CrossRef CAS PubMed.
- B. Bashir, J. S. Wang, G. Falchook, E. Fontana, H. T. Arkenau, L. Carter, R. Galot, B. Basu, A. Greystoke, V. Subbiah, D. L. Richardson, H. Orr, G. Bennett, R. Sharma, H. Xu, P. Paganoni, C. Xu, C. Campbell and M. McKean, J. Clin. Oncol., 2024, 42, 3443–3452 CrossRef CAS.
- F. Accardo, B. Prandi, L. Dellafiora, T. Tedeschi and S. Sforza, Food Chem., 2024, 458, 140175 CrossRef CAS PubMed.
- A. G. Harris, Gut, 1994, 35, S1–S4 CrossRef CAS PubMed.
- M. Werle and A. Bernkop-Schnürch, Amino Acids, 2006, 30, 351–367 CrossRef CAS PubMed.
- R. Haubner, R. Gratias, B. Diefenbach, S. L. Goodman, A. Jonczyk and H. Kessler, J. Am. Chem. Soc., 1996, 118, 7461–7472 CrossRef CAS.
- H. Su, F. Wang, W. Ran, W. Zhang, W. Dai, H. Wang, C. F. Anderson, Z. Wang, C. Zheng, P. Zhang, Y. Li and H. Cui, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 4518–4526 CrossRef CAS.
- M. Yang, D. Xu, L. Jiang, L. Zhang, D. Dustin, R. Lund, L. Liu and H. Dong, Chem. Commun., 2014, 50, 4827–4830 RSC.
- Y. Cai, B. Zhu, X. Shan, L. Zhou, X. Sun, A. Xia, B. Wu, Y. Yu, H. H. Zhu, P. Zhang and Y. Li, Adv. Mater., 2024, 36, e2306676 CrossRef.
- S. Cui, C. Guo, L. Yan, Y. He and L. Wu, Bioorg. Chem., 2025, 158, 108337 CrossRef CAS PubMed.
- R. Ree, S. Varland and T. Arnesen, Exp. Mol. Med., 2018, 50, 1–13 CrossRef CAS PubMed.
- A. Ullah, G. Shin and S. I. Lim, Drug Discovery Today, 2023, 28, 103738 CrossRef CAS PubMed.
- A. Zorzi, S. J. Middendorp, J. Wilbs, K. Deyle and C. Heinis, Nat. Commun., 2017, 8, 16092 CrossRef CAS PubMed.
- F. Kratz, J. Controlled Release, 2008, 132, 171–183 CrossRef CAS PubMed.
- Y. Nakamura, A. Mochida, P. L. Choyke and H. Kobayashi, Bioconjugate Chem., 2016, 27, 2225–2238 CrossRef CAS PubMed.
- Y. Zhang, Y. Fan, S. Liu, Y. Guan, J. Wan, Q. Ren, J. Wang, L. Zhong, Z. Hu, W. Shi and H. Qian, J. Med. Chem., 2024, 67, 10967–10985 CrossRef CAS PubMed.
- Z. Zha, S. R. Choi, L. Li, R. Zhao, K. Ploessl, X. Yao, D. Alexoff, L. Zhu and H. F. Kung, J. Med. Chem., 2022, 65, 13001–13012 CrossRef CAS PubMed.
- Y. S. Cho, H. R. Kim, S. J. Park, S. W. Chung, Y. G. Ko, J. H. Yeo, J. Lee, S. K. Kim, J. U. Choi, S. Y. Kim and Y. Byun, Biomaterials, 2022, 289, 121783 CrossRef CAS PubMed.
- Y. Fang and H. Wang, Pharmaceutics, 2022, 14, 9037–9056 Search PubMed.
- M. A. Beach, U. Nayanathara, Y. Gao, C. Zhang, Y. Xiong, Y. Wang and G. K. Such, Chem. Rev., 2024, 124, 5505–5616 CrossRef CAS PubMed.
- Y. Zeng, X. Liao, Y. Guo, F. Liu, F. Bu, J. Zhan, J. Zhang, Y. Cai and M. Shen, J. Controlled Release, 2024, 366, 838–848 CrossRef CAS PubMed.
- L. Yang, C. Zhang, C. Ren, J. Liu, Y. Zhang, J. Wang, F. Huang, L. Zhang and J. Liu, ACS Appl. Mater. Interfaces, 2019, 11, 331–339 CrossRef CAS PubMed.
- H. Zou, P. Banerjee, S. S. Y. Leung and X. Yan, Front. Pharmacol., 2020, 11, 997 CrossRef CAS PubMed.
- Y. Xiong, M. N. Samtani and D. Ouellet, Adv. Drug Delivery Rev., 2025, 217, 115503 CrossRef CAS PubMed.
- J. Bart, H. J. Groen, N. H. Hendrikse, W. T. van der Graaf, W. Vaalburg and E. G. de Vries, Cancer Treat. Rev., 2000, 26, 449–462 CrossRef CAS PubMed.
- S. Stalmans, N. Bracke, E. Wynendaele, B. Gevaert, K. Peremans, C. Burvenich, I. Polis and B. De Spiegeleer, PLoS One, 2015, 10, e0139652 CrossRef PubMed.
- S. M. Ghorai, A. Deep, D. Magoo, C. Gupta and N. Gupta, Pharmaceutics, 2023, 15, 1999 CrossRef CAS PubMed.
- X. Zhou, Q. R. Smith and X. Liu, Wiley Interdiscip. Rev.: Nanomed. Nanobiotechnol., 2021, 13, e1695 CrossRef PubMed.
- Y. Li, X. Zheng, M. Gong and J. Zhang, Oncotarget, 2016, 7, 79401–79407 CrossRef PubMed.
- K. Zhang, X. Yang, Y. Wang, Y. Yu, N. Huang, G. Li, X. Li, J. C. Wu and S. Yang, Nat. Med., 2025, 31, 45–59 CrossRef CAS PubMed.
- H. Han, B. Shaker, J. H. Lee, S. Choi, S. Yoon, M. Singh, S. Basith, M. Cui, S. Ahn, J. An, S. Kang, M. S. Yeom and S. Choi, J. Chem. Inf. Model., 2025, 65, 3215–3225 CrossRef CAS PubMed.
- N. Vale, M. Pereira, J. Santos, C. Moura, L. Marques and D. Duarte, Int. J. Mol. Sci., 2022, 24, 69 CrossRef PubMed.
- L. Fu, S. Shi, J. Yi, N. Wang, Y. He, Z. Wu, J. Peng, Y. Deng, W. Wang, C. Wu, A. Lyu, X. Zeng, W. Zhao, T. Hou and D. Cao, Nucleic Acids Res., 2024, 52, W422–W431 CrossRef PubMed.
- J. Yi, S. Shi, L. Fu, Z. Yang, P. Nie, A. Lu, C. Wu, Y. Deng, C. Hsieh, X. Zeng, T. Hou and D. Cao, Nat. Protoc., 2024, 19, 1105–1121 CrossRef CAS PubMed.
- S. R. Vangala, S. R. Krishnan, N. Bung, R. Srinivasan and A. Roy, J. Chem. Inf. Model., 2023, 63, 5066–5076 CrossRef CAS PubMed.
- B. Bakchi, A. D. Krishna, E. Sreecharan, V. B. J. Ganesh, M. Niharika, S. Maharshi, S. B. Puttagunta, D. K. Sigalapalli, R. R. Bhandare and A. B. Shaik, J. Mol. Struct., 2022, 1259, 132712 CrossRef CAS.
- D. Lagorce, L. Bouslama, J. Becot, M. A. Miteva and B. O. Villoutreix, Bioinformatics, 2017, 33, 3658–3660 CrossRef CAS PubMed.
- D. E. Zhang, T. He, T. Shi, K. Huang and A. Peng, Front. Pharmacol., 2025, 16, 1553853 CrossRef CAS PubMed.
- C. Agoni, R. Fernandez-Diaz, P. B. Timmons, A. Adelfio, H. Gomez and D. C. Shields, Biomolecules, 2025, 15, 524 CrossRef CAS PubMed.
- A. Armstrong, F. Coburn, Y. Nsereko and O. Al Musaimi, J. Pept. Sci., 2025, 31, e70040 CrossRef CAS PubMed.
- X. Zhang, X. Wang, S. Shi and D. Guo, Biomark. Res., 2025, 13, 144 CrossRef PubMed.
- V. Martin, P. H. G. Egelund, H. Johansson, S. Thordal Le Quement, F. Wojcik and D. Sejer Pedersen, RSC Adv., 2020, 10, 42457–42492 RSC.
| This journal is © The Royal Society of Chemistry 2026 |