Open Access Article
Open Access ArticleMethacrylated pulmonary dECM-enriched GelMA bioinks promote endothelialization and angiogenesis in 3D printed tubular constructs
Nehar
Celikkin
 *ab,
Maria Celeste
Tirelli
*ab,
Maria Celeste
Tirelli
 a,
Fabio
Maiullari
a,
Carolina Gutiérrez
Cisneros
b,
Vineeta
Kaushik
a,
Marina
Volpi
c,
Andreas
Aerts
a,
Fabio
Maiullari
a,
Carolina Gutiérrez
Cisneros
b,
Vineeta
Kaushik
a,
Marina
Volpi
c,
Andreas
Aerts
 d,
Ronny
Mohren
ef,
Michiel
Vandenbosch
e,
Ruth
Cardinaels
d,
Ronny
Mohren
ef,
Michiel
Vandenbosch
e,
Ruth
Cardinaels
 d,
Arn
Mignon
d,
Arn
Mignon
 b and
Marco
Costantini
b and
Marco
Costantini
 a
a
aInstitute of Physical Chemistry Polish Academy of Sciences, Warsaw, Poland. E-mail: neharcelikkin@gmail.com
bSmart Polymeric Biomaterials, Department of Materials Engineering, KU Leuven, Leuven, Belgium
cFaculty of Materials Science and Engineering, Warsaw University of Technology, Warsaw, Poland
dSoft Matter Rheology and Technology, Department of Chemical Engineering, KU Leuven, Leuven, Belgium
eThe Maastricht MultiModal Molecular Imaging (M4I) Institute, Division of Imaging Mass Spectrometry (IMS), Maastricht University, Maastricht, The Netherlands
fDepartment of Human Biology, M3 Research Group, Maastricht University, Maastricht, The Netherlands
First published on 31st December 2025
Abstract
In tissue engineering, developing biomimicking vascular tissue constructs is still significantly challenging due to the tubular architecture of the vessels and the limited availability of inexpensive biomaterials that can support effective endothelialization and angiogenesis. This study presents a robust protocol for designing 3D printed vascular tissue constructs composed of gelatin methacrylate (GelMA) and methacrylated pulmonary decellularized extracellular matrix (dECMMA) composite biomaterial ink. The pepsin-digested pulmonary dECM was identified with crucial ECM proteins responsible for blood vessel and circulatory system development, including collagen, fibrillin-1, annexin A2, and S100A11. The methacryloyl functionalization of dECM to form dECMMA was enabled by covalent crosslinking with GelMA to attain an inexpensive biomaterial ink that can support effective endothelialization and angiogenesis. Structurally intact 3D printed GelMA-dECMMA scaffolds, produced using the Suspended Layer Additive Manufacturing (SLAM) technique, showed improved endothelial cell attachment, proliferation, and organisation into vessel-like structures, strongly outperforming GelMA alone in promoting angiogenesis. Our results demonstrate the GelMA-dECMMA biomaterial ink's ability to create sophisticated biomimetic vascular models, which are promising for disease modelling and regenerative medicine.
Introduction
Vasculature holds essential importance for the vast majority of the tissues in organisms; hence, engineered vascular models are at the forefront of attention in tissue engineering research.1,2 These models are anticipated to address challenges from developing engineered vascularized tissues to understanding significant cardiovascular disease factors, such as thrombosis and stenosis.3 The foundation of developing physiologically relevant vasculature models relies significantly on choosing a suitable biomimetic material and creating the 3D vascular architectures with state-of-the-art biofabrication techniques. When choosing a suitable biomaterial for engineered vascular tissues, like many engineered tissues, gelatin methacrylate (GelMA) stands out regarding its cost, bioactivity, ease of synthesis, tunable mechanical properties hence its versatile use in biofabrication applications.4,5 GelMA also contains matrix metalloproteinase (MMP) sequences that promote enzymatic degradation and tissue regeneration. Its photo reactivity allows spatial and temporal control over stiffness and stability, and its inherent cell-adhesive motifs support endothelial attachment and remodeling, making perfusable vascular constructs more physiologically relevant. However, one of the remaining questions for vascular models is whether GelMA can be further functionalized for vascular tissues and enhanced in biomimicry for endothelialisation.6 One promising and cost-efficient approach, unlike the addition of growth factors (i.e., vascular endothelial growth factor -VEGF), can be the addition of decellularized extracellular matrix (dECM) from vasculature tissue to GelMA.7,8 While the vascular dECM can initially be considered favorable, as it retains a cocktail of angiogenic factors, the isolation of vascular tissue from its surrounding environment is complex, often yielding limited quantities for an effective decellularization process.9 Therefore, a compelling alternative can be to utilize highly vascularized organs from the body, such as pulmonary tissue.10 The lung's complex vascular architecture, comprising large arteries, extensive vascular networks, and a vast array of capillaries, combined with its intrinsic ability to facilitate the formation of new capillaries, offers unique possibilities for enhancing the maturation and development of endothelial cells. Adding pulmonary dECM could enhance the angiogenic potential of GelMA for engineering vasculature. However, a significant challenge arises from their differing processing requirements.11 GelMA is typically processed at a temperature higher than its gelation point (for instance, 18–40 °C for a 5% GelMA solution),12 while dECM requires lower temperatures (approximately 4 °C, which may vary depending on the concentration) to maintain its bioactivity and prevent premature gelation. This disparity poses a problem when trying to combine these two materials effectively. To address this issue, in the present study, we developed a methacrylated form of pulmonary dECM (dECMMA) that was used as a bioadditive to GelMA for the formulation of a composite biomaterial ink, which was 3D printed at room temperature to create tubular scaffolds with enhanced endothelialization and angiogenesis-related features. We comprehensively characterized the porcine pulmonary dECM, including its composition and structural integrity following decellularization. Furthermore, we conducted a proteomic analysis to identify key proteins retained in the dECM derived from pulmonary porcine tissue, shedding light on its potential bioactivity for angiogenesis. Through methacrylation of the pulmonary dECM (dECMMA) we aimed to (i) suppress uncontrolled thermal gelation at room temperature, keeping the biomaterial ink a stable sol for extrusion, and (ii) introduce on-demand photocrosslinkable moieties that can form a covalent interpenetrating network with GelMA upon UV exposure, yielding tunable mechanics and structural integrity. This combination expands the bioprinting window and platform compatibility of dECM biomaterial inks. We investigated further the angiogenic potential of dECMMA as a covalently crosslinked bioadditive aimed at enhancing the bioactivity of GelMA-based bioinks, promoting endothelialization, and recapitulating angiogenesis-associated features in 3D printed in vitro models.Results and discussion
Decellularization and digestion of pulmonary ECM
The porcine lung tissue was chemically decellularized using ionic and non-ionic detergents and then enzymatically digested using pepsin. The decellularization process using SDS and Triton X-100 effectively removed cellular components from porcine lung tissue while preserving the biochemical and structural proteins of the ECM. The multistep decellularization utilizing SDS and Triton X-100 has been chosen in this study, due to the potent ability of SDS to remove cellular components and the capability of Triton X-100 to help preserve the matrix proteins, biochemical cues, and remove SDS. SDS is widely used in a range of decellularization applications due to its high efficacy in removing both cells and nucleic acids, which meets the requirement of eliminating over 90% cellular components from tissues.13 However, SDS also causes undesired effects on the ECM, such as compacting collagen or altering tissue ultrastructure, leading to an increased fibrous appearance compared to native tissue.14 In addition, SDS is ionic, making it more difficult to remove completely, affecting subsequent recellularization and cytocompatibility. Therefore, combining the decellularization protocol with Triton X-100 in the latter stage provided a significant advantage by dedicating half of the overall decellularization time to a milder, nonionic detergent. Use of Triton X-100 not only reduces the negative effects of SDS on the extracellular matrix, preserving essential structural proteins, collagen fibers, and biochemical cues, but also facilitates the removal of residual SDS, resulting in a cytocompatible matrix with preserved structural proteins.15 After decellularization and lyophilization, pepsin-digested dECM (sdECM) was evaluated for its thermo-responsive properties once at pH 7.416 (Fig. 1).sdECM at temperatures below the gelation point retained its liquid-like behavior, transforming into a dECM hydrogel at 37 °C (Fig. 1). To demonstrate the efficacy of removing cell nuclei and preserving base membrane proteins during decellularization, porcine lung tissue has been stained with laminin α1 antibody and DAPI before and after decellularization. Laminins are a family of large ECM glycoproteins and are well known for their integral roles in promoting angiogenesis and modulating cell adhesion, migration, and differentiation. In particular, laminin–containing isoforms, such as laminin-111 (LM-111), are critical for polarized endothelial tissue, where they sustain high turnover and remodeling activities.17
The immunostaining revealed the preserved laminin α1 in the porcine pulmonary ECM, which indicated these matrices can inherently support endothelial cell infiltration and capillary formation for vascular architectures.18 Of note, LM-111 (a laminin α1–α1-containing isoform) is also the major glycoprotein in Matrigel, a commonly used in vitro substrate for studying endothelial cell organization into capillary-like structures.19 Along with the presence of laminin α1, DAPI staining showed the absence of nuclei post-decellularization, confirming the efficacy of our protocol (Fig. 1). Eliminating the detergents used in the decellularization process is essential for the cytocompatible use of the dECMs as biomaterial ink for in vitro studies. Therefore, bulk dECM scaffolds were seeded with 3T3 fibroblasts to evaluate their cytocompatibility. We demonstrated high cell viability (>95%) of the fibroblasts, highlighting the efficient removal of residual detergents from the decellularization process and cytocompatibility of the produced dECM matrix. Elongated and extensively spread cytoskeletal structures of 3T3s revealed extensive active colonization and actin filament extensions through the scaffold. Additionally, the uniform distribution of actin suggested that the dECM scaffold provided a supportive surface for cell attachment and cytoskeletal organization conducive to migration in the cytocompatible microenvironment and tissue remodeling.
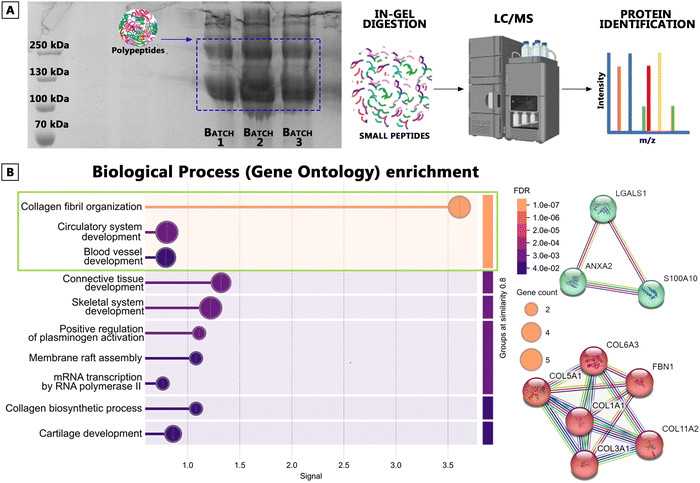
Proteomic insights into angiogenetic potential of pulmonary dECM
The sdECM was further evaluated using SDS-PAGE to examine the effect of digestion on the protein size distribution. The representative image of the resulting SDS-PAGE gel (Fig. 2A) demonstrated the size distribution with a large proportion of high-molecular-weight proteins at 200 kDa and 120 kDa.20 The proteomic analysis performed on these fractions indicated the detection of 71 proteins on the dECM, including lung-derived angiogenic cues such as the structural ECM proteins (Col1, Col3, Col5, Col6, and Col11), fibrillin-1 (FBN1), Galectin-1, S100A11, and annexin A2 (ANXA2) that guide endothelial cell attachment, migration, and vascular remodeling.10 (Fig. 2B) Among them, particularly Col1 and Col3, were significant for vascular tissue integrity and angiogenic sprouting.21 Col3 is a predominant fibrillar collagen found notably in blood vessels, which coexists with Col1. Col3 is responsible for regulating the fibrillogenesis of Col1 and modulating the elastic modulus of composite fibrils.22–24 Similarly, the presence of FBN1 contributes to the regulation of vascular elasticity, whereas it also acts as a binding site for proangiogenic growth factors, such as VEGF and TGF-β, which are essential for sprouting and angiogenesis.25 Thus, FBN1 indirectly regulates proangiogenic growth factor availability to endothelial cells by binding to and sequestering on demand. Annexin A2 and S100A11, the other key proteins detected in the dECM, facilitate ECM degradation, which is also crucial during the angiogenic remodeling and endothelial penetration into the scaffold. Annexin A2 is recognized as an intracellular, calcium-dependent phospholipid-binding protein; however, it could be externalized on the cell surface, where it facilitates plasminogen activation. As a receptor for S100A11, annexin A2 mediates the generation of plasmin and maintains vascular patency.26,27 S100A11, in turn, supports dynamic blood vessel remodeling through regulated plasmin production.28 These integral proteins collectively established a microenvironment conducive to endothelial adhesion, growth, migration, capillary formation, and microvessel stabilization. This confirms that the dECM can provide physical support and molecular signals promoting endothelialization and angiogenesis.Functionalization with methacryloyl groups and physicochemical characterization of dECMMA
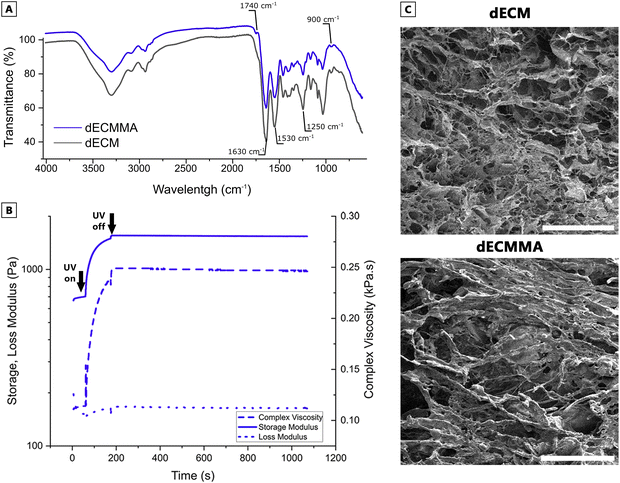
While GelMA and dECM are well-established biomaterials, here, it is aimed to tailor a biocomposite biomaterial ink composed of the covalent integration of dECMMA with GelMA to fabricate 3D-printed vascular constructs. dECM is inherently challenging to modify and process due to its complex biochemical composition. dECM's thermoresponsive fibrillogeneisis offers biocompatible crosslinking; however, when biofabrication techniques lack precise temperature control, dECM is significantly difficult to handle, typically resulting in premature crosslinking or clogging of extruders. The pulmonary dECM was functionalized with methacryloyl (MA) groups to produce dECMMA, enabling covalent interpenetration with GelMA to enhance its endothelialization and angiogenic potential and decoupling uncontrolled thermal gelation from printability, which allows room-temperature extrusion and rapid photocrosslinking without cooling. FTIR analysis revealed that the dECM and dECMMA presented peaks specific to collagen, glycoproteins, and glycosaminoglycans. For instance, glycosidic linkages in saccharide rings were observed as a band at 900 cm−1, indicating the presence of glycosaminoglycans and polysaccharide components of proteoglycans.29 Additionally, the Amide I band at 1600–1700 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching in proteins), Amide II band around 1500–1600 cm−1 (N–H bending and C–N stretching in proteins), and Amide III vibration at 1200–1350 cm−1, strongly indicated the structural integrity of collagen both in dECM and dECMMA.30 The presence of the carbonyl (C
O stretching in proteins), Amide II band around 1500–1600 cm−1 (N–H bending and C–N stretching in proteins), and Amide III vibration at 1200–1350 cm−1, strongly indicated the structural integrity of collagen both in dECM and dECMMA.30 The presence of the carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) group peak at 1740 cm−1 solely in dECMMA spectra is a strong indicator of the successful integration of methacryloyl moieties onto dECM (Fig. 3A).31 The degree of methacrylation of lung dECMMA was also quantified by 1H NMR spectroscopy using the lysine ε-CH2 proton signal (I Lys, 2.95–3.05 ppm) as a molecular indicator of functionalization. The ε-CH2 signal decreases upon methacrylation due to the substitution of lysine amine groups with methacrylate moieties. Quantitative comparison of the lysine ε-CH2 integral, normalized to the invariant aliphatic region (0.90–1.40 ppm), yielded a degree of methacrylation of 34%, consistent across replicate spectra. These results confirm efficient incorporation of methacrylate groups into the lung ECM backbone and validate the chemical modification strategy used to generate dECMMA (Fig. S2).
O) group peak at 1740 cm−1 solely in dECMMA spectra is a strong indicator of the successful integration of methacryloyl moieties onto dECM (Fig. 3A).31 The degree of methacrylation of lung dECMMA was also quantified by 1H NMR spectroscopy using the lysine ε-CH2 proton signal (I Lys, 2.95–3.05 ppm) as a molecular indicator of functionalization. The ε-CH2 signal decreases upon methacrylation due to the substitution of lysine amine groups with methacrylate moieties. Quantitative comparison of the lysine ε-CH2 integral, normalized to the invariant aliphatic region (0.90–1.40 ppm), yielded a degree of methacrylation of 34%, consistent across replicate spectra. These results confirm efficient incorporation of methacrylate groups into the lung ECM backbone and validate the chemical modification strategy used to generate dECMMA (Fig. S2).
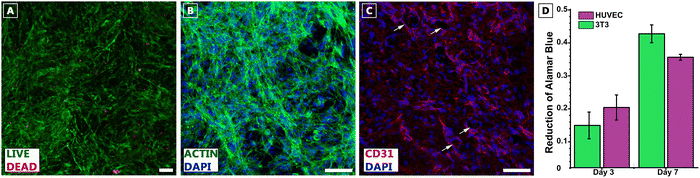
Additionally, the photorheological kinetics were analyzed to confirm photoresponsive behavior of dECMMA by monitoring changes in its storage (G′) and loss modulus (G″) during UV-light exposure at 20 °C for a period of 2 min. At the start of the test, G′ > G″, indicating a weak elastic-dominated physical gel -likely from reversible collagen fibril/ionic associations in dECMMA. This weak network remains injectable because its yield stress is low; under small-amplitude oscillation, it stores elastic energy (G′ > G″), but under large strain. Upon UV irradiation, G′ and the complex viscosity (η*) increased ∼10-fold while G″ remained approximately constant, evidencing a strong rise in elastic network connectivity with little change in viscous dissipation. After UV exposure, the elevated G′ and η* plateaued irreversibly, evidencing covalent photocrosslinking of methacrylate groups rather than reversible physical restructuring. As the sample displayed gel-like behavior already at baseline (G′ > G″), a critical gel point cannot be defined from a G′/G″ crossover. Instead, we report ∼20 s as the apparent photocuring onset, defined by the onset of the rapid increase in G′ (and η*) that locks the network covalently. Notably, the dECMMA solution exhibited a gelation time of 20 seconds, which is similar to the gelation time of 15 seconds for dECMMA derived from liver tissue that has been previously reported.32 To identify the structural differences between the dECM and dECMMA, SEM imaging was performed (Fig. 3C) on crosslinked, bulk freeze-dried scaffolds. Both dECM and dECMMA exhibited interconnected porous networks, although the dECMMA was observed to be a denser fibrillar matrix compared to the dECM scaffold. The differences in the fibrillar texture can be attributed to the differences in the crosslinking mechanisms of dECM (thermal crosslinking) and dECMMA (photocrosslinking). Following the physicochemical characterization, the biological potential of dECMMA was evaluated by investigating its ability to support cellular viability, attachment, and angiogenesis. UV crosslinked, freeze-dried dECMMA scaffolds seeded with 3T3 fibroblasts demonstrated high cell viability (>90%), indicating the successful removal of cytotoxic byproducts of the methacrylation reaction (Fig. 4A). Furthermore, phalloidin staining of 3T3 fibroblasts confirmed robust cell attachment and spreading, with no significant changes compared to dECM scaffolds (Fig. 1B) and well-organized cytoskeletal structures throughout the scaffold (Fig. 4B). In a parallel experiment, HUVECs were also seeded on the scaffolds to evaluate the potential use of the scaffolds for endothelial tissue formation. HUVECs exhibited good attachment and growth on the dECMMA substrates, as presented by the expression of CD31 and the formation of small lumen-like structures on the scaffold surface (Fig. 4C). The presence of these structures indicated that dECMMA shows advantageous material characteristics for HUVECs.
In addition to qualitative analysis, the cellular metabolic activity of 3T3 and HUVECs was quantitatively evaluated through Alamar Blue reduction, revealing sustained growth of both HUVECs and 3T3 fibroblasts over 7 days (Fig. 4D). Notably, both 3T3 fibroblasts and HUVECs demonstrated a steady increase in metabolic activity, indicating their ability to proliferate on dECMMA substrates.
3D printing of tubular constructs with enhanced endothelialization and angiogenic potential
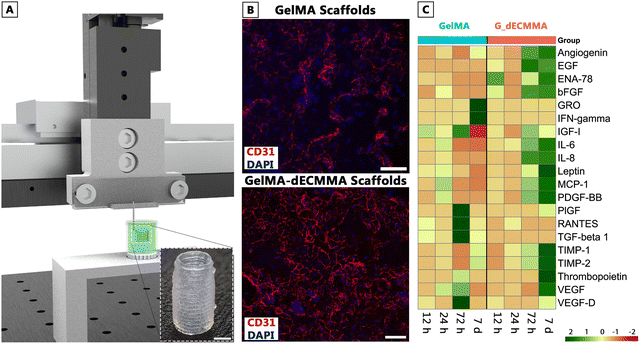
To obtain architecturally relevant vascular constructs, we designed and printed tubular scaffolds. The GelMA and GelMA-dECMMA biomaterial inks were extruded with microfluidic pumps through a 21G needle (needle nominal inner diameter is 0.514 mm) attached to a custom-made 3D printing head. Due to the low viscosity of the biomaterial inks, we employed the Suspended Layer Additive Manufacturing (SLAM) technique.33 The biomaterial inks were deposited and stabilized within the agarose fluid–gel bath, which functioned as a temporary support medium during the printing process.34 Agarose bath, known for its self-healing properties, exhibited fluid–gel behavior with rapid elastic recovery under shear stress, helping the 3D printed tubular constructs retain their architecture without collapsing due to its slurry-like nature.33 (Fig. 5A) Thanks to the support of the agarose bath, the effective printing resolution in the XY plane ∼500 µm, with a vertical (Z axis) resolution of 250 µm was achieved. The 3D printed tubular scaffolds were then physically crosslinked at 4 °C for 10 min, covalently crosslinked with UV exposure, and collected from the agarose bath. The brief incubation at 4 °C promotes scaffold integrity and a mechanical baseline for the tubular scaffolds. The physically crosslinked interpenetrating network of GelMA and dECMMA is then rapidly photocrosslinked thanks to the established methacrylate network. This dual mechanism provided process flexibility and reproducibility of tubular self-standing and perfusable vascular constructs (Video S1). Well-defined and reproducible tubular scaffolds that closely mimic native human-sized veins (5 mm diameter with 0.5 mm wall thickness) were seeded with HUVECs to assess endothelialization and angiogenic potential. Fig. 5B illustrates immunofluorescent staining for the endothelial marker CD31 (red) and nuclei (DAPI, blue). Over 7 days of culture, more endothelial networks were observed when GelMA-dECMMA scaffolds were compared to GelMA alone, according to quantitative morphometry of CD31 immunostaining. The HUVECs cultured on GelMA-dECMMA scaffolds showed approximately 1.7-fold increase in surface endothelialization, with 9.6% CD31-positive coverage compared to 5.6% in GelMA. Importantly, these biological improvements were not attributable to differences in bulk mechanics; compression testing confirmed no significant differences in compressive modulus between GelMA and GelMA–dECMMA hydrogels at the concentrations used (Fig. S3). Thus, the enhanced endothelial response derives from the bioactive biochemical composition of the dECMMA component rather than stiffness-mediated effects. Furthermore, dECMMA significantly improved endothelial network architecture, increasing vessel-mesh density, total tubule length, and branch-point counts, collectively demonstrating more extensive and intricate 3D endothelial organization (Fig. S4).The GelMA-dECMMA scaffolds show visibly enhanced CD31 coverage compared to GelMA scaffolds, indicative of more robust endothelial cell attachment and proliferation. Despite being an established indicator, CD31 expression alone was insufficient to claim improvement of endothelialization and angiogenic potential by adding dECMMA in GelMA scaffolds. Therefore, we have analyzed the angiogenic cytokines of HUVECs seeded on GelMA-dECMMA and GelMA scaffolds in the basal media to understand the effect of the dECMMA from a broader perspective. Several key correlations emerged when analyzing angiogenic cytokines from the heat map in conjunction with the proteomic findings. The heat map data in Fig. 5C demonstrate that the expression of VEGF and PDGF-BB was significantly higher in GelMA-dECMMA (G_dECMMA) scaffolds compared to GelMA alone, particularly at 72 hours and 7 days, suggesting the contribution of dECMMA in enhancing endothelial proliferation and angiogenesis, particularly at later culture time points, as the dECMMA is covalently bonded to GelMA.
For instance, increased VEGF detected on day 7 could be associated with the presence of FBN1 on dECM, which serves as a VEGF reservoir, storing and releasing growth factor on-demand to endothelial cells.25,35 Additionally, annexin A2 may have also facilitated the VEGF release from the dECMMA, promoting the pro-angiogenic effects of the scaffolds. The increased expression of bFGF and angiogenin in G_dECMMA scaffolds can again be related to the presence of Col1, Col3, Col5, Col6, and Col11, which created a highly favorable microenvironment for endothelial sprouting and migration.36 Furthermore, the elevated expression of PDGF-BB in G_dECMMA scaffolds at later time points (Day 7), which is essential for vascular stabilization and pericyte recruitment, also suggests that G_dECMMA scaffolds promote endothelial proliferation and vascular maturation, making them particularly suitable for long-term tissue integration. Furthermore, the downregulation of inflammatory markers also highlights the cytocompatibility of dECMMA, reinforcing it as a potential additive for enhanced endothelialization and angiogenic potential to GelMA.
Conclusions & future perspectives
The proteomic analysis and cytocompatibility characterization of porcine-derived pulmonary dECM revealed that it is a bioactive and cytocompatible additive that supports endothelial function, angiogenesis, and vascular maturation. The presence of structural ECM proteins (collagens, fibrillin-1, annexin A2) and metabolic regulators (Fig. S1) creates a comprehensive biochemical environment conducive to vascular formation. Methacryloyl functionalization enables covalent bonding of dECMMA to GelMA, ensuring on-demand availability of dECM components to HUVECs while decoupling uncontrolled thermal gelation from printability, thus allowing room-temperature extrusion and rapid photo-locking. Once integrated, the proteomic findings together with angiogenic marker expression confirm that GelMA–dECMMA scaffolds promote VEGF-driven endothelial proliferation, bFGF-mediated migration, and PDGF-BB-associated vascular stabilization (Fig. S5). Although the printed constructs are at the large-vessel scale, our biological assays intentionally focus on microvascular processes: endothelial activation, upregulation of angiogenic proteins, and cytokine secretion that govern endothelialization, host integration, and early neovascular recruitment around both micro- and macrovasculature. In this context, the 5 mm tubular constructs primarily serve to demonstrate that the dECMMA-based bioink maintains printability, shape fidelity, and lumen integrity under macroscale fabrication, while still supporting microvascular-like endothelial network formation. This geometry was selected as a representative model to confirm that the material can sustain self-supporting architectures without collapse, and, given the current ∼250 µm printing resolution of our extrusion setup, it represents an optimal compromise between construct stability and biological relevance, enabling accurate handling, crosslinking, and subsequent biological analysis. Our results therefore provide a biochemical and biological proof-of-concept for dECMMA as a bioactive matrix in a mechanically stable, macroscale printed format, and motivate future studies on the detailed mechanical robustness of GelMA–dECMMA scaffolds under physiological flow and pressure, as well as on long-term stability, biodegradation kinetics, sustained bioactivity, and extended proteomic and release-profile analyses.Experimental
All primary and secondary antibodies, cell culture media, penicillin–streptomycin (P/S), fetal bovine serum (FBS), 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), 4′,6-diamidino-2-phenylindole (DAPI), and phalloidin were purchased from ThermoFischer. All chemicals were purchased from Merck (Sigma-Aldrich) unless indicated otherwise. Confocal imaging was performed with a Nikon A1R confocal microscope.Decellularization of porcine lung tissue
Lungs from domestic pigs (Sus scrofa) were obtained from local slaughterhouses, and 12 lobes from 6 different animals were pooled to reduce the batch-to-batch variability and enhance the reproducibility. Porcine lungs were decellularized in a 3L container according to a previously established protocol.20 In brief, 500 g porcine lung tissue was dissected into 1–2 cm3 pieces and frozen at −80 °C. Before decellularization, the dissected tissue pieces were defrosted at room temperature, immersed in H2O, underwent constant mixing at 800 rpm with an overhead stirrer (IKA, Ministar 40), and were rinsed externally with H2O to remove excess blood. The dissected lung pieces were then sequentially treated and incubated in a series of reagents at room temperature, including 0.3% w/v sodium dodecyl sulfate (72 h, with changes every 24 h), 3% w/v Triton x-100 (72 h, with changes every 24 h), 1 M NaCl (6 h) and H2O (6 h). The lung tissue was washed with H2O between each treatment step to remove the previous reagent and cellular debris. Following decellularization, the remaining tissue was loosely packed in sterile Petri dishes, frozen at −80 °C, and then lyophilized. The dried acellular tissue was then enzymatically digested. Dried dECM was suspended at 30 mg mL−1 in 0.5 M acetic acid solution containing 3 mg mL−1 pepsin from porcine gastric mucosa (3200 U mg−1). Enzymatic digestion was performed at room temperature under constant agitation (overhead stirrer at 1000 rpm) under a laminar hood for 72 h. The digested solution was centrifuged to pellet insoluble proteins; the supernatant was collected as the solubilized dECM (sdECM). The sdECM was placed on ice, and the pH of the solution was adjusted to 7.4 with 5 M NaOH under constant agitation. Following pH adjustment, sdECM was poured into sterile Petri dishes, frozen at −80 °C, lyophilized, and stored at −20 °C until further use.Protocol for laminin staining and assessment of decellularization
To quantitatively evaluate the decellularization, the fixed pulmonary tissue samples and fixed dECM hydrogels were stained with DAPI and prominent protein Laminin. To crosslink sdECM pre-gel, frozen sdECM (at pH 7.4) was thawed on ice, and the 20 µL volume of sdECM was transferred to PDMS molds (disk diameter of 500 µm, thickness 200 µm) and then incubated at 37 °C for 30 min to induce gelation. The tissue samples and dECM hydrogels were fixed with 4% paraformaldehyde (PFA), and then permeabilized with 0.3% (v/v) Triton X-100 in phosphate-buffered saline (PBS). Fixed tissue samples and dECM hydrogels were blocked with 1% (w/v) bovine serum albumin (BSA) to inhibit the non-specific binding. Following the BSA blocking step, the samples were incubated with anti-laminin (Sigma L9393) produced in rabbit (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50) overnight. After the primary antibody staining, samples were washed and incubated in Texas Red anti-rabbit (1
50) overnight. After the primary antibody staining, samples were washed and incubated in Texas Red anti-rabbit (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000) and 4′,6-diamidino-2-phenylindole (DAPI, 1
1000) and 4′,6-diamidino-2-phenylindole (DAPI, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500) solution. After washing, the samples were imaged with a confocal microscope. Laminin and cellular nuclei detection were evaluated and presented using image analysis software (ImageJ, National Institutes of Health, USA).
500) solution. After washing, the samples were imaged with a confocal microscope. Laminin and cellular nuclei detection were evaluated and presented using image analysis software (ImageJ, National Institutes of Health, USA).
Evaluation of cytocompatibility
The cytocompatibility (according to the ISO 10993-5 guidelines for biological evaluation of medical devices) of the dECM hydrogels was evaluated using 3T3 murine fibroblast cells purchased from Sigma-Aldrich. 3T3 cells were cultured in DMEM (high glucose medium) supplemented with 10% FBS and 1% P/S at 37 °C and 5% CO2. The sdECM concentration was adjusted to 2 mg mL−1 with cold PBS, and the sdECM pregels were crosslinked at 37 °C. After the gelation, dECM hydrogels were seeded with 500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per cm2 (50
000 cells per cm2 (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per scaffold). To obtain a qualitative assessment of the cell's viability, a live/dead assay was performed on day 3 of culture to detect any acute cytotoxicity due to the residue from the decellularization process. The 3T3 seeded dECM hydrogels were stained with LIVE/DEAD™ Cell Imaging Kit (488/570) (Invitrogen) according to the manufacturer's protocol. Following the staining, the samples were visualized with confocal microscopy. To evaluate the cell attachment, the samples were stained with phalloidin to visualize the actin filaments on the cell cytoskeleton. On day 3, the 3T3 seeded dECM hydrogels were fixed, permeabilized, and blocked in 1% (w/v) BSA for unspecific binding. The samples then were incubated with Alexa Fluor 488 Phalloidin (1
000 cells per scaffold). To obtain a qualitative assessment of the cell's viability, a live/dead assay was performed on day 3 of culture to detect any acute cytotoxicity due to the residue from the decellularization process. The 3T3 seeded dECM hydrogels were stained with LIVE/DEAD™ Cell Imaging Kit (488/570) (Invitrogen) according to the manufacturer's protocol. Following the staining, the samples were visualized with confocal microscopy. To evaluate the cell attachment, the samples were stained with phalloidin to visualize the actin filaments on the cell cytoskeleton. On day 3, the 3T3 seeded dECM hydrogels were fixed, permeabilized, and blocked in 1% (w/v) BSA for unspecific binding. The samples then were incubated with Alexa Fluor 488 Phalloidin (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40) and DAPI (1
40) and DAPI (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500), the cell cytoskeleton and nuclei were visualized using confocal microscope.
500), the cell cytoskeleton and nuclei were visualized using confocal microscope.
Proteomic analysis
The dECM was directly lysed in 5 M urea buffer and in-gel fractionation was performed via SDS-PAGE electrophoresis. Protein bands were cut into equal pieces. In-gel digestion: protein bands were manually excised on a glass plate from the gel and processed on a MassPREP digestion robot (Waters, Manchester UK). A 50 mM ammonium bicarbonate solution in 50% v/v ACN was used for destaining. Cysteines were reduced with 10 mM DTT in 100 mM ammonium bicarbonate for 30 min followed by alkylation with 55 mM iodoacetamide in 100 mM ammonium bicarbonate for 20 min. Spots were washed with 100 mM ammonium bicarbonate to remove excess reagents and were subsequently dehydrated with 100% ACN. Trypsin (6 ng mL−1) in 50 mM ammonium bicarbonate was added to the gel plug and incubated at 37 °C for 5 h. The peptides were extracted with 1% v/v formic acid/2% v/v ACN and subsequently with 1% v/v formic acid/50% v/v ACN. Liquid chromatography – mass spectrometry: peptide separation was performed on a Thermo Scientific (Dionex) Ultimate 3000 Rapid Separation UHPLC system equipped with an Acclaim PepMap C18 analytical column (2 um, 100 Å, 75 µm × 150 mm). Peptide samples were first desalted on an online installed C18 trapping column. After desalting peptides were separated on the analytical column with a 90 minutes linear gradient from 5% to 35% Acetonitrile (ACN) with 0.1% FA at 300 nL min−1 flow rate. The UHPLC system was coupled to a Q Exactive HF mass spectrometer (Thermo Scientific). Data was acquired in DIA mode with full ms scans between m/z 385–1015 at a resolution of 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and MS/MS scans at 30
000 and MS/MS scans at 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. Mass spectrometric raw data analysis: the LC-MS DIA data were analyzed with DIA-NN version 1.8.1. The SwissProt Human database Homo sapiens (SwissProt TaxID = 9606) was used. The database search was performed with the following settings. Enzyme was trypsin, a maximum of 2 missed cleavages, minimum peptide length of 7 and a static modification of cysteine carbamidomethylation.
000. Mass spectrometric raw data analysis: the LC-MS DIA data were analyzed with DIA-NN version 1.8.1. The SwissProt Human database Homo sapiens (SwissProt TaxID = 9606) was used. The database search was performed with the following settings. Enzyme was trypsin, a maximum of 2 missed cleavages, minimum peptide length of 7 and a static modification of cysteine carbamidomethylation.
Synthesis of methacrylated dECM (dECMMA)
dECMMA was synthesized modifying a protocol previously published.37 Briefly, 5 g of freeze-dried sdECM was dispersed in 100 mL of 0.1 M sodium carbonate buffer and 12 µL of methacrylic anhydride (94%) was slowly added to the solution. During the reaction, the pH of the solution was adjusted to and kept at 8 with 5 N NaOH to avoid the formation of the methacrylic acid byproduct, and the solution was gently stirred at 4 °C for 24 h in the dark. Then, 400 mL of deionized water was added to stop the reaction, followed by three days of dialysis against 0.01 mM hydrochloric acid solution using a cellulose acetate tube (MWCO: 12 kDa). The dECMMA solution was lyophilized and stored at −20 °C until further processing.Physiochemical characterization of dECMMA
Fourier transform infrared (FTIR) spectroscopy infrared transmission spectra were collected using a Spectrum 100 spectrometer from PerkinElmer. For spectra acquisition, 1 cm−1 resolution was used in the 4000–600 cm−1 wavenumber range. Photorheology rheology measurements were performed on an Anton Paar MCR702 rheometer consisting of a glass bottom lower plate and a sandblasted upper plate with a diameter of 8 mm. The following procedure was performed three times for each hydrogel-type. The 30 mg mL−1 dECMMA solution containing 0.05 mg mL−1 2-hydroxy-4′-(2-hydroxyethoxy)-2-methylpropiophenone (Irgacure®2959) sample was injected on the bottom plate and the upper plate was lowered to a height of 101 µm. Left-over sample was scrapped away and the plate was lowered to the actual measuring height of 100 µm. Oscillatory shear was applied at a frequency of 1 Hz and a strain of 1% to determine the shear storage modulus (G′) and shear loss modulus (G″) during following steps: First, the moduli of the uncured samples were determined during 1 minute. Second, a UV-curing step was performed in which the samples were irradiated, OmniCure S2000, for two minutes at an intensity of 29 mW cm−2 during which the moduli were measured. Third, the moduli were measured over 20 minutes after the UV-curing step. For each hydrogel-type, a frequency sweep was executed in the frequency range of 0.0159 Hz to 15.9 Hz at a strain of 1%. Morphology assessment with scanning electron microscopy high-resolution electron micrographs were obtained with a Philips XL30 FEG scanning electron microscope (SEM) (Philips, The Netherlands) operating at an accelerating voltage of 5 kV and collecting secondary electron emissions. Samples were kept in a vacuum chamber overnight before imaging. Before measurement, they were fixed onto carbon stickers a 5 nm Pt80/Pd20 alloy coating was applied via plasma sputtering.Biological evaluation of dECMMA for endothelialization
To evaluate the biological performance of dECMMA, cytocompatibility test (ISO 10993-5 guidelines for biological evaluation of medical devices) was performed with 3T3 and HUVECs. Initially, dECMMA pre-gels were prepared at 30 mg mL−1 in sterile PBS. After the complete dispersion of the dECMMA, 0.05 mg mL−1 Irgacure® 2959 was added to the pre-gel and the 20 µL volume of pre-gel dispersion is transferred to PDMS molds (disks diameter of 500 µm, thickness 200 µm) and photocrosslinked exposing to 12.5 mW cm−2 UV-lamp (λ = 365 nm) for 60 s. The crosslinked gels were too soft to transfer from the molds, hence the gels were frozen and lyophilized at −80 °C to easily transfer the scaffolds from the molds for in vitro experiments. 3T3 cells were cultured in DMEM (high glucose medium) supplemented with 10% FBS and 1% P/S and HUVECs were cultured in Endothelial Cell Growth Medium (PeloBiotech, Germany) at 37 °C and 5% CO2. 3T3 or HUVEC cells were seeded on the lyophilized scaffolds at a cell density of 500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per cm2 (50
000 cells per cm2 (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per scaffold). To evaluate the cell viability and cell attachment Alexa Fluor 488 Phalloidin (1
000 cells per scaffold). To evaluate the cell viability and cell attachment Alexa Fluor 488 Phalloidin (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40) and DAPI (1
40) and DAPI (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500) staining was performed on the scaffolds seeded with 3T3. To evaluate the endothelium formation Alexa FluorR 647 Conjugate CD31 (PECAM-1,Cell Signaling Technology) (1
500) staining was performed on the scaffolds seeded with 3T3. To evaluate the endothelium formation Alexa FluorR 647 Conjugate CD31 (PECAM-1,Cell Signaling Technology) (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40) and DAPI (1
40) and DAPI (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500) staining was performed on the scaffolds seeded with HUVECs. To monitor the proliferation of the cells growing on the scaffolds, Alamar Blue assay (BioRad) was performed on day 3 and day 7. The viability of the 3T3 or HUVEC cell were normalized to the positive control of the cells treated with Triton X-100.
500) staining was performed on the scaffolds seeded with HUVECs. To monitor the proliferation of the cells growing on the scaffolds, Alamar Blue assay (BioRad) was performed on day 3 and day 7. The viability of the 3T3 or HUVEC cell were normalized to the positive control of the cells treated with Triton X-100.
3D printing of tubular constructs with enhanced endothelialization and angiogenic potential
20 mg dECMMA and 40 mg GelMA (GelMA was synthesized in-house, briefly Type A porcine skin gelatin (Sigma-Aldrich, St. Luis, MO) was modified with methacrylic anhydride according to the protocol reported previously to achieve 57 ± 2% degree of functionalization.) were simultaneously dissolved in 1 mL PBS. After complete dissolution of GeLMA and dECMMA 0.1 mg Irgacure® 2959 was then added. The biomaterial ink solution was then filled into 5 mL Hamilton glass syringes and printed using a custom-made 3D bioprinter equipped with a microfluidic pump (neMESYS, Cetoni GmbH). The 3D cylinders (diameter: 5 mm, height: 7 mm) were designed using 3D Builder (Microsoft). To create tubular constructs, the cylinders were sliced using Slic3r software without the inner architecture, resulting in printing only the perimeter, and the g-code for the tubes was generated, considering a 0.5 mm diameter printing nozzle and a 0.25 mm layer thickness. The biomaterial ink was extruded through a custom-made polycarbonate holder equipped with a 21G metallic needle for precise deposition into agarose fluid–gel. The agarose fluid gel was prepared, as previously described, by autoclaving 0.5% w/v agarose solution and cooling down the solution to room temperature at under constant stirring (700 rpm) for 12 h.38 Following the extrusion, the prints in the agarose fluid–gel were incubated in fridge at 4 °C for 5 minutes and then exposed to UV light (12.5 mW cm−2, λ = 365 nm) for 60 seconds, for permanent crosslinking of the tubular constructs. After the photocrosslinking, the 3D structures were removed from the agarose bath and rinsed with PBS. 3D printed tubular GelMA and GelMA-dECMMA scaffolds were sterilized under a laminar hood, exposed to UV light for 1 h, and transferred to 1.5 mL sterile Eppendorf tubes for further use for in vitro experiments. The printed samples were imaged using a motorized 2D/3D microscope (MRCL700 3D Image Pro, MicroQubic AG, Switzerland).HUVEC culture on tubular scaffolds, immunofluorescence staining and evaluation of angiogenic markers
For evaluation of endothelial layer formation, HUVECs were collected and suspended in Endothelial Cell Growth Medium (PeloBiotech, Germany) at a cell density of 107 cells per mL whereby 500 µL HUVEC cell suspension was added to each Eppendorf holding printed scaffolds (5 × 106 cells per scaffold). The tubes were placed in a rotator for 2 h in the incubator for homogeneous seeding of the HUVECs on the tubular scaffolds. The HUVEC-coated GelMA and GelMA-dECMMA scaffolds were then transferred to well plates and cultured over 10 days in 12 well plates with 3 ml of Endothelial Cell Growth Medium. On day 10, the samples were fixed, and CD31 and DAPI staining was performed. Maximum-intensity projections of the CD31/DAPI confocal stacks were analysed in Fiji, the red channel was thresholded to generate binary masks, skeletonised and quantified with the Angiogenesis Analyzer plug-in to obtain total tubule length, branch-point number, and mesh density. Lumen fraction was calculated after binary closing of the masks to delineate void spaces. To analyze the effect of dECMMA on the expression of angiogenic markers, GelMA and GelMA-dECMMA bulk scaffolds (prepared in PDMS molds, disks with a diameter of 500 µm, thickness of 200 µm, crosslinked at 4 °C for 5 minutes and then exposed to UV light (12.5 mW cm−2, λ = 365 nm) for 60 seconds, sterilized with 30 min UV exposure) were seeded with HUVECs at a cell density of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per scaffold. The GelMA and GelMA-dECMMA scaffolds were cultured for 7 days in 3 mL supplement-free Endothelial Cell Basal Medium (PeloBiotech, Germany). The detection and evaluation of the angiogenic markers were performed using Human Angiogenesis Array (RayBiotech, Array C1) at 12 h, 24 h, 72 h and day 7. The detection and evaluation of the angiogenic markers were performed according to the manufacturers’ manual.
000 cells per scaffold. The GelMA and GelMA-dECMMA scaffolds were cultured for 7 days in 3 mL supplement-free Endothelial Cell Basal Medium (PeloBiotech, Germany). The detection and evaluation of the angiogenic markers were performed using Human Angiogenesis Array (RayBiotech, Array C1) at 12 h, 24 h, 72 h and day 7. The detection and evaluation of the angiogenic markers were performed according to the manufacturers’ manual.
Statistical analysis
Samples were tested in triplicate, and data are reported as in mean value ± standard deviation. Network prediction and pathway enrichment of differentially expressed proteins were carried out with the STRING database.39 For statistical and data processing, GraphPad Prism software was used. Statistical analysis was performed using either a two-way ANOVA with Tukey's post hoc test or a t-test when only two parameters were compared. A p value of less than 0.05 was considered statistically significant (*).Author contributions
NC: writing – original draft, visualization, methodology, investigation, analysis, funding acquisition, conceptualization, data curation; MCT: writing – original draft, visualization, investigation; FM: methodology, investigation; CGC: methodology, investigation; AA and RC: methodology, investigation; MV: revision, methodology, investigation, VK: methodology, investigation RM and MV: methodology, investigation; MC and AM: conceptualization, methodology, writing – review.Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article, including: Fig. 1Bhttps://doi.org/10.18150/SA7UQQ (Laminin and DAPI staining for decellularized matrix). Fig. 2https://doi.org/10.18150/XQ6DS5 (Proteomic analysis of decelularized porcine lung). Fig. 3https://doi.org/10.18150/KCGEFM, https://doi.org/10.18150/RPFZLD (Physicochemical Characterization of dECMMA). Fig. 4https://doi.org/10.18150/A44N2F (Cytocompatibility of 3T3 fibroblast and HUVECs_). Fig. 5https://doi.org/10.18150/VNBJJL Angiogenesic markers - HUVECs cultured on GelMA and GelMA-dECMMA scaffolds are available at RepOD at https://repod.icm.edu.pl/.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ma00860c.
Acknowledgements
This study was supported by the National Science Centre Poland (NCN) within the PRELUDIUM 19 Project No. 2020/37/N/ST5/03272, SONATINA 8 2023/48/C/ST5/00230 and by FWO Grant for a scientific stay in Flanders Grant No. V507423N.References
- J. Lim, H.-W. Fang, S. Bupphathong, P.-C. Sung, C.-E. Yeh, W. Huang and C.-H. Lin, The Edifice of Vasculature-on-Chips: A Focused Review on the Key Elements and Assembly of Angiogenesis Models, ACS Biomater. Sci. Eng., 2024, 10(6), 3548–3567 Search PubMed
.
- J. Zhu, Y. Wang, L. Zhong, F. Pan and J. Wang, Advances in Tissue Engineering of Vasculature through Three-dimensional Bioprinting, Dev. Dyn., 2021, 250(12), 1717–1738 CrossRef PubMed
.
- A. Shakeri, Y. Wang, Y. Zhao, S. Landau, K. Perera, J. Lee and M. Radisic, Engineering Organ-on-a-Chip Systems for Vascular Diseases, Arterioscler., Thromb., Vasc. Biol., 2023, 43(12), 2241–2255 CrossRef CAS PubMed
.
- T. Jin, Z. Fu, L. Zhou, L. Chen, J. Wang, L. Wang, S. Yan, T. Li and P. Jin, GelMA Loaded with Platelet Lysate Promotes Skin Regeneration and Angiogenesis in Pressure Ulcers by Activating STAT3, Sci. Rep., 2024, 14(1), 18345 CrossRef PubMed
.
- Z. Jia, L. Chen, D. Gu, X. Li, T. Wen and W. Li, Lentinan-Loaded GelMA Hydrogel Accelerates Diabetic Wound Healing through Enhanced Angiogenesis and Immune Microenvironment Modulation, Int. J. Biol. Macromol., 2024, 264, 130716 CrossRef CAS PubMed
.
- S. Bupphathong, J. Lim, H.-W. Fang, H.-Y. Tao, C.-E. Yeh, T.-A. Ku, W. Huang, T.-Y. Kuo and C.-H. Lin, Enhanced Vascular-like Network Formation of Encapsulated HUVECs and ADSCs Coculture in Growth Factors Conjugated GelMA Hydrogels, ACS Biomater. Sci. Eng., 2024, 10(5), 3306–3315 CrossRef CAS PubMed
.
- S. P. Parthiban, D. Rana, E. Jabbari, N. Benkirane-Jessel and M. Ramalingam, Covalently Immobilized VEGF-Mimicking Peptide with Gelatin Methacrylate Enhances Microvascularization of Endothelial Cells, Acta Biomater., 2017, 51, 330–340 CrossRef
.
- B. Byambaa, N. Annabi, K. Yue, G. Trujillo-de Santiago, M. M. Alvarez, W. Jia, M. Kazemzadeh-Narbat, S. R. Shin, A. Tamayol and A. Khademhosseini, Bioprinted Osteogenic and Vasculogenic Patterns for Engineering 3D Bone Tissue, Adv. Healthcare Mater., 2017, 6(16), 1700015 CrossRef PubMed
.
- X. Wang, V. Chan and P. R. Corridon, Decellularized Blood Vessel Development: Current State-of-the-Art and Future Directions, Front. Bioeng. Biotechnol., 2022, 10, 951644 Search PubMed
.
- E. A. Calle, R. C. Hill, K. L. Leiby, A. V. Le, A. L. Gard, J. A. Madri, K. C. Hansen and L. E. Niklason, Targeted Proteomics Effectively Quantifies Differences between Native Lung and Detergent-Decellularized Lung Extracellular Matrices, Acta Biomater., 2016, 46, 91–100 CrossRef CAS PubMed
.
- J. M. Seok, M. Ahn, D. Kim, J. S. Lee, D. Lee, M. J. Choi, S. J. Yeo, J. H. Lee, K. Lee, B. S. Kim and S. A. Park, Decellularized Matrix Bioink with Gelatin Methacrylate for Simultaneous Improvements in Printability and Biofunctionality, Int. J. Biol. Macromol., 2024, 262, DOI:10.1016/j.ijbiomac.2024.130194
.
- A. T. Young, O. C. White and M. A. Daniele, Rheological Properties of Coordinated Physical Gelation and Chemical Crosslinking in Gelatin Methacryloyl (GelMA) Hydrogels, Macromol. Biosci., 2020, 20(12), 2000183 Search PubMed
.
- A. Gilpin and Y. Yang, Decellularization Strategies for Regenerative Medicine: From Processing Techniques to Applications, BioMed Res. Int., 2017, 2017, DOI:10.1155/2017/9831534
.
- L. J. White, A. J. Taylor, D. M. Faulk, T. J. Keane, L. T. Saldin, J. E. Reing, I. T. Swinehart, N. J. Turner, B. D. Ratner and S. F. Badylak, The Impact of Detergents on the Tissue Decellularization Process: A ToF-SIMS Study, Acta Biomater., 2017, 50, 207–219 Search PubMed
.
- D. C. Sullivan, S.-H. Mirmalek-Sani, D. B. Deegan, P. M. Baptista, T. Aboushwareb, A. Atala and J. J. Yoo, Decellularization Methods of Porcine Kidneys for Whole Organ Engineering Using a High-Throughput System, Biomaterials, 2012, 33(31), 7756–7764 CrossRef CAS PubMed
.
- N. Celikkin; (Poland), N. S. C. Rheological Testing Decellularized Chicken Heart, SHeep Heart, SHeep Lung and Porcine Lung. V1 ed. RepOD, 2024 DOI:10.18150/DJBUSD.
- J. Dixelius, L. Jakobsson, E. Genersch, S. Bohman, P. Ekblom and L. Claesson-Welsh, Laminin-1 Promotes Angiogenesis in Synergy with Fibroblast Growth Factor by Distinct Regulation of the Gene and Protein Expression Profile in Endothelial Cells, J. Biol. Chem., 2004, 279(22), 23766–23772 CrossRef CAS PubMed
.
- D. S. Grant and H. K. Kleinman, Regulation of Capillary Formation by Laminin and Other Components of the Extracellular Matrix, EXS, 1997, 79, 317–333, DOI:10.1007/978-3-0348-9006-9_13
.
- C. Hughes, L.-M. Postovit and G. Lajoie, Matrigel: A Complex Protein Mixture Required for Optimal Growth of Cell Culture, Proteomics, 2010, 10, 1886–1890, DOI:10.1002/pmic.200900758
.
- R. A. Pouliot, B. M. Young, P. A. Link, H. E. Park, A. R. Kahn, K. Shankar, M. B. Schneck, D. J. Weiss and R. L. Heise, Porcine Lung-Derived Extracellular Matrix Hydrogel Properties Are Dependent on Pepsin Digestion Time, Tissue Eng., Part C, 2020, 26(6), 332–346 Search PubMed
.
- Y. Fei, Z. Ling, Q. Tong and J. Wang, Apoptotic Extracellular Vesicles from Supernumerary Tooth-Derived Pulp Stem Cells Transfer COL1A1 to Promote Angiogenesis via PI3K/Akt/VEGF Pathway, Int J Nanomed., 2024, 6811–6828 Search PubMed
.
- E. L. Doherty, W. Y. Aw, E. C. Warren, M. Hockenberry, C. P. Whitworth, G. Krohn, S. Howell, B. O. Diekman, W. R. Legant and H. T. Nia, Patient-Derived Extracellular Matrix Demonstrates Role of COL3A1 in Blood Vessel Mechanics, Acta Biomater., 2023, 166, 346–359 Search PubMed
.
- Z. Ma, C. Mao, Y. Jia, Y. Fu and W. Kong, Extracellular Matrix Dynamics in Vascular Remodeling, Am. J. Physiol.: Cell Physiol., 2020, 319(3), C481–C499 CrossRef CAS PubMed
.
- X. Liu, H. Wu, M. Byrne, S. Krane and R. Jaenisch, Type III Collagen Is Crucial for Collagen I Fibrillogenesis and for Normal Cardiovascular Development, Proc. Natl. Acad. Sci. U. S. A., 1997, 94(5), 1852–1856 CrossRef CAS PubMed
.
- F. Alonso, Y. Dong, L. Li, T. Jahjah, J. W. Dupuy, I. Fremaux, D. P. Reinhardt and E. Génot, Fibrillin-1 Regulates Endothelial Sprouting during Angiogenesis, Proc. Natl. Acad. Sci. U. S. A., 2023, 120(23) DOI:10.1073/pnas.2221742120
.
- T. M. Sveeggen, C. A. Abbey, R. L. Smith, M. L. Salinas, R. S. Chapkin and K. J. Bayless, Annexin A2 Modulates Phospholipid Membrane Composition Upstream of Arp2 to Control Angiogenic Sprout Initiation, FASEB J., 2023, 37(1) DOI:10.1096/fj.202201088R
.
- J. König, J. Prenen, B. Nilius and V. Gerke, The Annexin II-P11 Complex Is Involved in Regulated Exocytosis in Bovine Pulmonary Artery Endothelial Cells, J. Biol. Chem., 1998, 273(31), 19679–19684 CrossRef PubMed
.
- E. R. Martinez, Y. Franko, R. Franko, G. A. Ferronato, A. E. S. Viana, E. Windenbach, J. B. Stoeckl, T. Frohlich and M. A. M. M. Ferraz, Developing and Characterising Bovine Decellularized Extracellular Matrix Hydrogels to Biofabricate Female Reproductive Tissues, Acta Biomater., 2025, 196, 152–170, DOI:10.1016/j.actbio.2025.03.009
.
- N. Mainreck, S. Brézillon, G. D. Sockalingum, F. X. Maquart, M. Manfait and Y. Wegrowski, Rapid Characterization of Glycosaminoglycans Using a Combined Approach by Infrared and Raman Microspectroscopies, J. Pharm. Sci., 2011, 100(2), 441–450, DOI:10.1002/jps.22288
.
- A. Vásquez-Rivera, H. Oldenhof, A. Hilfiker and W. F. Wolkers, Spectral Fingerprinting of Decellularized Heart Valve Scaffolds, Spectrochim. Acta, Part A, 2019, 214, 95–102 CrossRef PubMed
.
- S. Ramesh and M. Ramalingam, Aqueous-Mediated Synthesis and Characterization of Gelatin Methacryloyl for Biomedical Applications, Biointerface Res. Appl. Chem, 2022, 12(5), 6269–6279 CAS
.
- A. Almalla, L. Elomaa, N. Fribiczer, T. Landes, P. Tang, Z. Mahfouz, B. Koksch, K. H. Hillebrandt, I. M. Sauer, D. Heinemann, S. Seiffert and M. Weinhart, Chemistry Matters: A Side-by-Side Comparison of Two Chemically Distinct Methacryloylated DECM Bioresins for Vat Photopolymerization, Biomater. Adv., 2024, 160, 213850, DOI:10.1016/j.bioadv.2024.213850
.
- J. J. Senior, M. E. Cooke, L. M. Grover and A. M. Smith, Fabrication of Complex Hydrogel Structures Using Suspended Layer Additive Manufacturing (SLAM), Adv. Funct. Mater., 2019, 29(49), 1904845 CrossRef CAS
.
- M. Marcotulli, M. C. Tirelli, M. Volpi, J. Jaroszewicz, C. Scognamiglio, P. Kasprzycki, K. Karnowski, W. Święszkowski, G. Ruocco and M. Costantini, Microfluidic 3D Printing of Emulsion Ink for Engineering Porous Functionally Graded Materials, Adv. Mater. Technol., 2023, 8(5), 2201244 CrossRef CAS
.
- L. Li, J. Huang and Y. Liu, The Extracellular Matrix Glycoprotein Fibrillin-1 in Health and Disease, Front. Cell Dev. Biol., 2024, 11, 1302285 CrossRef PubMed
.
- A. Irastorza, P. Vázquez-Aristizabal, P. Guerrero, K. de la Caba and A. Izeta, Comparative Proteomic Analysis of the Composition of Decellularized Extracellular Matrix (DECM) and DECM-Based Inks as Compared to the Native Tissue, bioRxiv, 2024, preprint DOI:10.1101/2024.09.23.614437.
- M. Jung, Y. Han, C. Woo and C. S. Ki, Pulmonary Tissue-Mimetic Hydrogel Niches for Small Cell Lung Cancer Cell Culture, J. Mater. Chem. B, 2021, 9(7), 1858–1866 RSC
.
- M. Marcotulli, M. C. Tirelli, M. Volpi, J. Jaroszewicz, C. Scognamiglio, P. Kasprzycki, K. Karnowski, W. Święszkowski, G. Ruocco and M. Costantini, Microfluidic 3D Printing of Emulsion Ink for Engineering Porous Functionally Graded Materials, Adv. Mater. Technol., 2023, 8(5), 2201244 CrossRef CAS
.
- D. Szklarczyk, R. Kirsch, M. Koutrouli, K. Nastou, F. Mehryary, R. Hachilif, A. L. Gable, T. Fang, N. T. Doncheva and S. Pyysalo, The STRING Database in 2023: Protein–Protein Association Networks and Functional Enrichment Analyses for Any Sequenced Genome of Interest, Nucleic Acids Res., 2023, 51(D1), D638–D646 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |